Profiling Colorectal Cancer in the Landscape Personalized Testing—Advantages of Liquid Biopsy
Abstract
1. Introduction
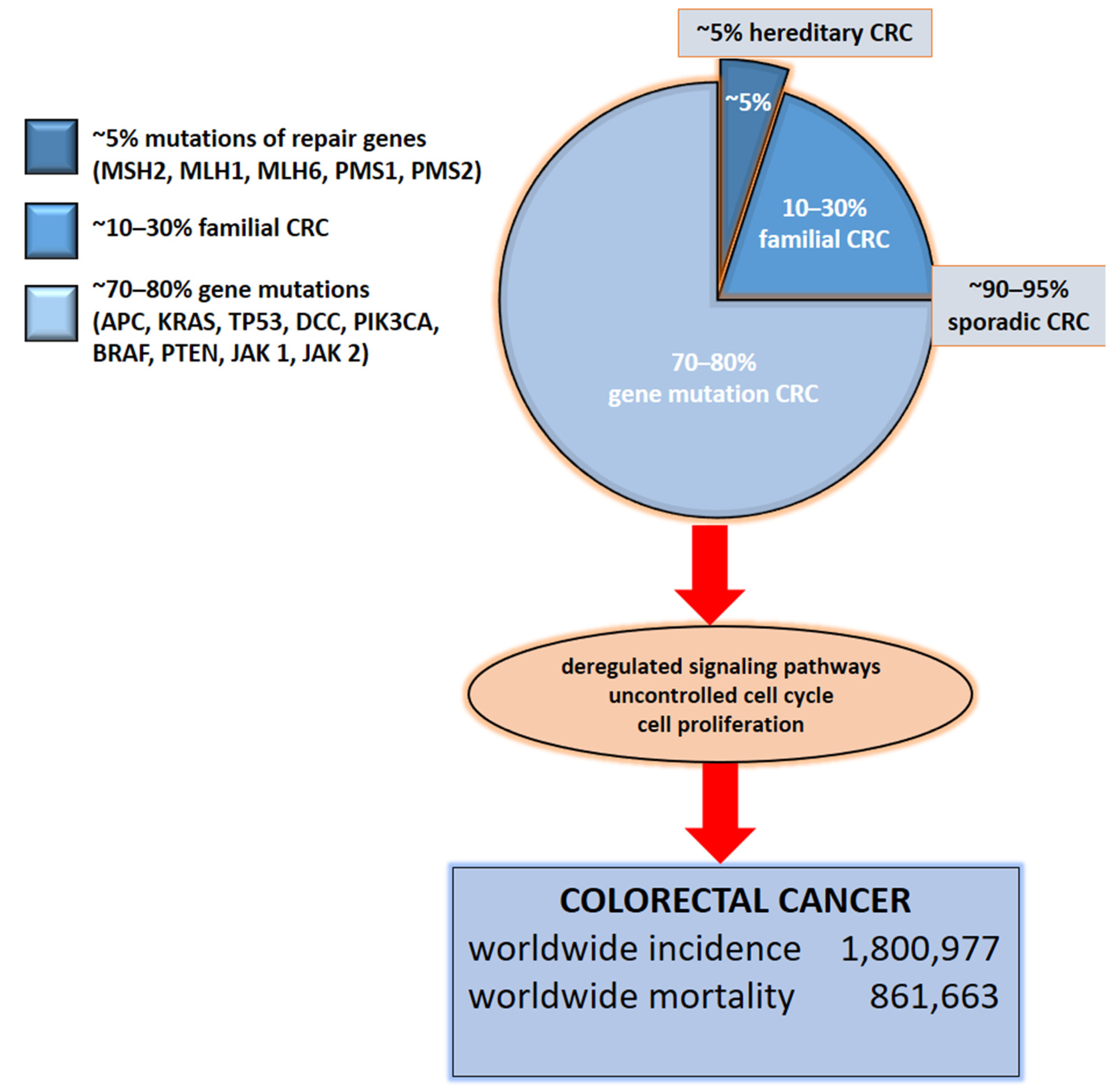
2. Etiology of Colorectal Cancer
3. The impact of Genetic Alterations on Disease Outcome
4. Liquid Biopsy
5. Circulating Tumor DNA (ctDNA)
6. Exosomes
7. Circulating Tumor Cells (CTC)
8. Conclusions
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| APC | adenomatous polyposis coli protein gene |
| BRAF | serine/threonine-protein kinase B-Raf gene |
| CA | carbohydrate antigen |
| CAPNS1 | calpain small subunit 1 |
| CD326 | cluster of differentiation 326 |
| c-MYC | cellular MYC gene |
| CEA | carcinoembryonic antigen |
| cfDNA | cell-free circulating deoxyribonucleic acid |
| CIMP | epigenetic instability |
| CIN | chromosomal instability |
| circRNA | circular ribonucleic acid |
| CRC | colorectal cancer |
| CTC | circulating tumor cells |
| ctDNA | circulating tumor deoxyribonucleic acid |
| DCC | netrin-1 receptor gene |
| DPP IV | dipeptidyl-peptidase IV |
| EpCAM | epithelial cell adhesion molecule |
| Hsp | heat shock protein |
| ILK | integrin-linked protein kinase |
| JAK | Janus kinase |
| KRAS | c-K-ras protein gene |
| lncRNA | long non-coding ribonucleic acid |
| MAPK | mitogen-activated protein kinase |
| MMP9 | matrix metallopeptidase 9 |
| MLH1 | mutL homolog 1 gene |
| MLH6 | mutL homolog 6 gene |
| MSH2 | mutS homolog 2 gene |
| MSI | microsatellite instability |
| NRAS | neuroblastoma RAS |
| NRAS | neuroblastoma RAS gene |
| PD-1 | programmed cell death protein 1 |
| PIK3CA | phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit alpha gene |
| PI3K | phosphatidylinositol 3-kinase |
| PMS1 | PMS1 protein homolog 1 gene |
| PMS2 | PMS1 protein homolog 2 gene |
| PTEN | phosphatase and tensin homolog |
| PTEN | phosphatase and tensin homolog gene |
| SERPINA1 | α-1 antitrypsin |
| SERPINAF1 | α-2 antiplasmin |
| SMAD2 | mothers against decapentaplegic homolog 2 gene |
| SMAD4 | mothers against decapentaplegic homolog 4 gene |
| STAT | signal transducer and activator of transcription |
| TGF-β | transforming growth factor β |
| TP53 | tumor protein p53 gene |
| WNT | wingless-related integration site |
References
- Knight, S.R.; Shaw, A.C.; Pius, R.; Drake, T.M.; Norman, L.; Ademuyiwa, O.A.; Adisa, O.A.; Aguilera, M.L.; Al-Saqqa, S.W.; Al-Slaibi, I.; et al. Global variation in postoperative mortality and complications after cancer surgery: A multicentre, prospective cohort study in 82 countries. Lancet 2021, 397, 387–397. [Google Scholar] [CrossRef]
- Xu, P.; Zhu, Y.; Sun, B.; Xiao, Z. Colorectal cancer characterization and therapeutic target prediction based on microRNA expression profile. Sci. Rep. 2016, 6, 20616. [Google Scholar] [CrossRef]
- Bray, F.; Ferlay, J.; Soerjomataram, I.; Siegel, R.L.; Torre, L.A.; Jemal, A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 2018, 68, 394–424. [Google Scholar] [CrossRef] [PubMed]
- Tripathi, S.; Belkacemi, L.; Cheung, M.S.; Bose, R.N. Correlation between Gene Variants, Signaling Pathways, and Efficacy of Chemotherapy Drugs against Colon Cancers. Cancer Inform. 2016, 15, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Rodriguez-Casanova, A.; Costa-Fraga, N.; Bao-Caamano, A.; López-López, R.; Muinelo-Romay, L.; Diaz-Lagares, A. Epigenetic Landscape of Liquid Biopsy in Colorectal Cancer. Front. Cell Dev. Biol. 2021, 9, 622459. [Google Scholar] [CrossRef]
- Miyamoto, Y.; Hiyoshi, Y.; Tokunaga, R.; Akiyama, T.; Daitoku, N.; Sakamoto, Y.; Yoshida, N.; Baba, H. Postoperative complications are associated with poor survival outcome after curative resection for colorectal cancer: A propensity-score analysis. J. Surg. Oncol. 2020, 122, 344–349. [Google Scholar] [CrossRef] [PubMed]
- Howlader, N.; Noone, A.M.; Krapcho, M.; Miller, D.; Bishop, K.; Altekruse, S.F.; Kosary, C.L.; Yu, M.; Ruhl, J.; Tatalovich, Z.; et al. SEER Cancer Statistics Review; National Cancer Institute: Bethesda, MD, USA; pp. 1975–2013, Based on November 2015 SEER data submission, posted to the SEER web site, April 2016.
- Nakamura, M.; Kageyama, S.-I.; Seki, M.; Suzuki, A.; Okumura, M.; Hojo, H.; Motegi, A.; Akimoto, T. Liquid Biopsy Cell-free DNA Biomarkers in Patients with Oligometastatic Colorectal Cancer Treated by Ablative Radiotherapy. Anticancer Res. 2021, 41, 829–834. [Google Scholar] [CrossRef] [PubMed]
- Najmanova, I.; Vopršalová, M.; Saso, L.; Mladěnka, P. The pharmacokinetics of flavanones. Crit. Rev. Food Sci. Nutr. 2020, 60, 3155–3171. [Google Scholar] [CrossRef]
- Buhrmann, C.; Shayan, P.; Brockmueller, A.; Shakibaei, M. Resveratrol Suppresses Cross-Talk between Colorectal Cancer Cells and Stromal Cells in Multicellular Tumor Microenvironment: A Bridge between In Vitro and In Vivo Tumor Microenvironment Study. Molecules 2020, 25, 4292. [Google Scholar] [CrossRef]
- Xie, Y.-H.; Chen, Y.-X.; Fang, J.-Y. Comprehensive review of targeted therapy for colorectal cancer. Signal Transduct. Target. Ther. 2020, 5, 1–30. [Google Scholar] [CrossRef]
- Ab Mutalib, N.-S.; Yusof, N.F.M.; Abdul, S.-N.; Jamal, R. Pharmacogenomics DNA Biomarkers in Colorectal Cancer: Current Update. Front. Pharmacol. 2017, 8, 736. [Google Scholar] [CrossRef] [PubMed]
- Etiology of CRC—Colorectal Cancer Research. Available online: https://www.colorectalcancer.dk/en/our-research/four-areas/crc-etiology/ (accessed on 1 February 2021).
- Mármol, I.; Sánchez-De-Diego, C.; Dieste, A.P.; Cerrada, E.; Yoldi, M.J.R. Colorectal Carcinoma: A General Overview and Future Perspectives in Colorectal Cancer. Int. J. Mol. Sci. 2017, 18, 197. [Google Scholar] [CrossRef] [PubMed]
- Ahmed, M. Colon Cancer: A Clinician’s Perspective in 2019. Gastroenterol. Res. 2020, 13, 1–10. [Google Scholar] [CrossRef]
- Matijašić, M.; Meštrović, T.; Perić, M.; Paljetak, H. Čipčić; Panek, M.; Bender, D.V.; Kelečić, D.L.; Krznarić, Željko; Verbanac, D. Modulating Composition and Metabolic Activity of the Gut Microbiota in IBD Patients. Int. J. Mol. Sci. 2016, 17, 578. [Google Scholar] [CrossRef]
- Verbanac, D.; Maleš, Ž.; Barišić, K. Nutrition—Acts and myths. Acta Pharm. 2019, 69, 497–510. [Google Scholar] [CrossRef] [PubMed]
- Pino, M.S.; Chung, D.C. The Chromosomal Instability Pathway in Colon Cancer. Gastroenterology 2010, 138, 2059–2072. [Google Scholar] [CrossRef] [PubMed]
- Bišćanin, A.; Ljubičić, N.; Boban, M.; Baličević, D.; Pavić, I.; Bišćanin, M.M.; Budimir, I.; Dorosulic, Z.; Duvnjak, M. CX43 Expression in Colonic Adenomas and Surrounding Mucosa Is a Marker of Malignant Potential. Anticancer Res. 2016, 36, 5437–5442. [Google Scholar] [CrossRef]
- Lao, V.V.; Grady, W.M. Epigenetics and colorectal cancer. Nat. Rev. Gastroenterol. Hepatol. 2011, 8, 686–700. [Google Scholar] [CrossRef]
- Herzig, D.O.; Tsikitis, V.L. Molecular markers for colon diagnosis, prognosis and targeted therapy. J. Surg. Oncol. 2014, 111, 96–102. [Google Scholar] [CrossRef]
- Schatoff, E.M.; Leach, B.I.; Dow, L.E. WNT Signaling and Colorectal Cancer. Curr. Color. Cancer Rep. 2017, 13, 101–110. [Google Scholar] [CrossRef]
- Brocardo, M.; Henderson, B.R. APC shuttling to the membrane, nucleus and beyond. Trends Cell Biol. 2008, 18, 587–596. [Google Scholar] [CrossRef]
- Rennoll, G.Y.S. Regulation ofMYCgene expression by aberrant Wnt/β-catenin signaling in colorectal cancer. World J. Biol. Chem. 2015, 6, 290–300. [Google Scholar] [CrossRef] [PubMed]
- Toon, C.W.; Chou, A.; Clarkson, A.; DeSilva, K.; Houang, M.; Chan, J.C.Y.; Sioson, L.L.; Jankova, L.; Gill, A.J. Immunohistochemistry for Myc Predicts Survival in Colorectal Cancer. PLoS ONE 2014, 9, e87456. [Google Scholar] [CrossRef]
- Chen, J.; Guo, F.; Shi, X.; Zhang, L.; Zhang, A.; Jin, H.; He, Y. BRAF V600E mutation and KRAS codon 13 mutations predict poor survival in Chinese colorectal cancer patients. BMC Cancer 2014, 14, 802. [Google Scholar] [CrossRef]
- Li, W.; Qiu, T.; Zhi, W.; Shi, S.; Zou, S.; Ling, Y.; Shan, L.; Ying, J.; Lu, N. Colorectal carcinomas with KRAS codon 12 mutation are associated with more advanced tumor stages. BMC Cancer 2015, 15, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Canon, J.; Rex, K.; Saiki, A.Y.; Mohr, C.; Cooke, K.; Bagal, D.; Gaida, K.; Holt, T.; Knutson, C.G.; Koppada, N.; et al. The clinical KRAS(G12C) inhibitor AMG 510 drives anti-tumour immunity. Nature 2019, 575, 217–223. [Google Scholar] [CrossRef] [PubMed]
- Porru, M.; Pompili, L.; Caruso, C.; Biroccio, A.; Leonetti, C. Targeting KRAS in metastatic colorectal cancer: Current strategies and emerging opportunities. J. Exp. Clin. Cancer Res. 2018, 37, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Ogino, S.; Nosho, K.; Kirkner, G.J.; Kawasaki, T.; Meyerhardt, J.A.; Loda, M.; Giovannucci, E.L.; Fuchs, C.S. CpG island methylator phenotype, microsatellite instability, BRAF mutation and clinical outcome in colon cancer. Gut 2008, 58, 90–96. [Google Scholar] [CrossRef]
- Goldstein, J.; Tran, B.; Ensor, J.; Gibbs, P.; Wong, H.L.; Wong, S.F.; Vilar, E.; Tie, J.; Broaddus, R.; Kopetz, S.; et al. Multicenter retrospective analysis of metastatic colorectal cancer (CRC) with high-level microsatellite instability (MSI-H). Ann. Oncol. 2014, 25, 1032–1038. [Google Scholar] [CrossRef]
- Kadowaki, S.; Kakuta, M.; Takahashi, S.; Takahashi, A.; Arai, Y.; Nishimura, Y.; Yatsuoka, T.; Ooki, A.; Yamaguchi, K.; Matsuo, K.; et al. Prognostic value ofKRASandBRAFmutations in curatively resected colorectal cancer. World J. Gastroenterol. 2015, 21, 1275–1283. [Google Scholar] [CrossRef]
- Yaeger, R.; Cercek, A.; O’Reilly, E.M.; Reidy, D.L.; Kemeny, N.; Wolinsky, T.; Capanu, M.; Gollub, M.J.; Rosen, N.; Berger, M.F.; et al. Pilot Trial of Combined BRAF and EGFR Inhibition in BRAF-Mutant Metastatic Colorectal Cancer Patients. Clin. Cancer Res. 2015, 21, 1313–1320. [Google Scholar] [CrossRef]
- Liao, X.; Morikawa, T.; Lochhead, P.; Imamura, Y.; Kuchiba, A.; Yamauchi, M.; Nosho, K.; Qian, Z.R.; Nishihara, R.; Meyerhardt, J.A.; et al. Prognostic Role of PIK3CA Mutation in Colorectal Cancer: Cohort Study and Literature Review. Clin. Cancer Res. 2012, 18, 2257–2268. [Google Scholar] [CrossRef]
- Atreya, C.E.; Sangale, Z.; Xu, N.; Matli, M.R.; Tikishvili, E.; Welbourn, W.; Stone, S.; Shokat, K.M.; Warren, R.S. PTEN expression is consistent in colorectal cancer primaries and metastases and associates with patient survival. Cancer Med. 2013, 2, 496–506. [Google Scholar] [CrossRef] [PubMed]
- Sarli, L.; Bottarelli, L.; Bader, G.; Iusco, D.; Pizzi, S.; Costi, R.; D’Adda, T.; Bertolani, M.; Roncoroni, L.; Bordi, C. Association Between Recurrence of Sporadic Colorectal Cancer, High Level of Microsatellite Instability, and Loss of Heterozygosity at Chromosome 18q. Dis. Colon Rectum 2004, 47, 1467–1482. [Google Scholar] [CrossRef] [PubMed]
- Popat, S.; Houlston, R.S. A systematic review and meta-analysis of the relationship between chromosome 18q genotype, DCC status and colorectal cancer prognosis. Eur. J. Cancer 2005, 41, 2060–2070. [Google Scholar] [CrossRef]
- Popat, S.; Zhao, D.; Chen, Z.; Pan, H.; Shao, Y.; Chandler, I.; Houlston, R.S. Relationship between chromosome 18q status and colorectal cancer prognosis: A prospective, blinded analysis of 280 patients. Anticancer Res. 2007, 27, 627–633. [Google Scholar]
- Said, A.H.; Raufman, J.-P.; Xie, G. The Role of Matrix Metalloproteinases in Colorectal Cancer. Cancers 2014, 6, 366–375. [Google Scholar] [CrossRef] [PubMed]
- Munro, A.J.; Lain, S.; Lane, D.P. P53 abnormalities and outcomes in colorectal cancer: A systematic review. Br. J. Cancer 2005, 92, 434–444. [Google Scholar] [CrossRef]
- Lay, M.; Mariappan, R.; Gotlib, J.; Dietz, L.; Sebastian, S.; Schrijver, I.; Zehnder, J.L. Detection of the JAK2 V617F Mutation by LightCycler PCR and Probe Dissociation Analysis. J. Mol. Diagn. 2006, 8, 330–334. [Google Scholar] [CrossRef]
- Albacker, L.A.; Wu, J.; Smith, P.; Warmuth, M.; Stephens, P.J.; Zhu, P.; Yu, L.; Chmielecki, J. Loss of function JAK1 mutations occur at high frequency in cancers with microsatellite instability and are suggestive of immune evasion. PLoS ONE 2017, 12, e0176181. [Google Scholar] [CrossRef]
- Shin, D.S.; Zaretsky, J.M.; Escuin-Ordinas, H.; Garcia-Diaz, A.; Hu-Lieskovan, S.; Kalbasi, A.; Grasso, C.S.; Hugo, W.; Sandoval, S.; Torrejon, D.Y.; et al. Primary Resistance to PD-1 Blockade Mediated by JAK1/2 Mutations. Cancer Discov. 2017, 7, 188–201. [Google Scholar] [CrossRef] [PubMed]
- Locker, G.Y.; Hamilton, S.; Harris, J.; Jessup, J.M.; Kemeny, N.; Macdonald, J.S.; Somerfield, M.R.; Hayes, D.F.; Bast, R.C. ASCO 2006 Update of Recommendations for the Use of Tumor Markers in Gastrointestinal Cancer. J. Clin. Oncol. 2006, 24, 5313–5327. [Google Scholar] [CrossRef]
- Lee, W.-S.; Baek, J.-H.; Kim, K.K.; Park, Y.H. The prognostic significant of percentage drop in serum CEA post curative resection for colon cancer. Surg. Oncol. 2012, 21, 45–51. [Google Scholar] [CrossRef] [PubMed]
- Su, B.-B. Role of serum carcinoembryonic antigen in the detection of colorectal cancer before and after surgical resection. World J. Gastroenterol. 2012, 18, 2121–2126. [Google Scholar] [CrossRef] [PubMed]
- Tümay, V.; Guner, O.S. The utility and prognostic value of CA 19-9 and CEA serum markers in the long-term follow up of patients with colorectal cancer. A single-center experience over 13 years. Ann. Ital. Chir. 2020, 91, 494–503. [Google Scholar] [PubMed]
- Jung, A.; Kirchner, T. Liquid Biopsy in Tumor Genetic Diagnosis. Dtsch. Aerzteblatt Online 2018, 115, 169–174. [Google Scholar] [CrossRef] [PubMed]
- Domínguez-Vigil, I.G.; Moreno-Martínez, A.K.; Wang, J.Y.; Roehrl, M.H.A.; Barrera-Saldaña, H.A. The dawn of the liquid biopsy in the fight against cancer. Oncotarget 2017, 9, 2912–2922. [Google Scholar] [CrossRef]
- Hood, L. Systems Biology and P4 Medicine: Past, Present, and Future. Rambam Maimonides Med. J. 2013, 4, e0012. [Google Scholar] [CrossRef]
- O’Sullivan, A.; Henrick, B.; Dixon, B.; Barile, D.; Zivkovic, A.; Smilowitz, J.; Lemay, D.; Martin, W.; German, J.B.; Schaefer, S.E. 21st century toolkit for optimizing population health through precision nutrition. Crit. Rev. Food Sci. Nutr. 2018, 58, 3004–3015. [Google Scholar] [CrossRef]
- Taghizadeh, H.; Prager, G.W. Personalized Adjuvant Treatment of Colon Cancer. Visc. Med. 2020, 36, 397–406. [Google Scholar] [CrossRef]
- Chakrabarti, S.; Xie, H.; Urrutia, R.; Mahipal, A. The Promise of Circulating Tumor DNA (ctDNA) in the Management of Early-Stage Colon Cancer: A Critical Review. Cancers 2020, 12, 2808. [Google Scholar] [CrossRef]
- Sholl, L.M.; Oxnard, G.R.; Paweletz, C.P. Traditional Diagnostics versus Disruptive Technology: The Role of the Pathologist in the Era of Liquid Biopsy. Cancer Res. 2020, 80, 3197–3199. [Google Scholar] [CrossRef]
- Wu, T.; Mo, Y.; Wu, C. Prognostic values of CEA, CA19-9, and CA72-4 in patients with stages I-III colorectal cancer. Int. J. Clin. Exp. Pathol. 2020, 13, 1608–1614. [Google Scholar]
- Tsai, W.-S.; Chen, J.-S.; Shao, H.-J.; Wu, J.-C.; Lai, J.-M.; Lu, S.-H.; Hung, T.-F.; Chiu, Y.-C.; You, J.-F.; Hsieh, P.-S.; et al. Circulating Tumor Cell Count Correlates with Colorectal Neoplasm Progression and Is a Prognostic Marker for Distant Metastasis in Non-Metastatic Patients. Sci. Rep. 2016, 6, 24517. [Google Scholar] [CrossRef]
- Osumi, H.; Shinozaki, E.; Yamaguchi, K. Circulating Tumor DNA as a Novel Biomarker Optimizing Chemotherapy for Colorectal Cancer. Cancers 2020, 12, 1566. [Google Scholar] [CrossRef] [PubMed]
- Said, R.; Guibert, N.; Oxnard, G.R.; Tsimberidou, A.M. Circulating tumor DNA analysis in the era of precision oncology. Oncotarget 2020, 11, 188–211. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Chen, J.-Q.; Liu, J.-L.; Tian, L. Exosomes in tumor microenvironment: Novel transporters and biomarkers. J. Transl. Med. 2016, 14, 1–9. [Google Scholar] [CrossRef]
- Han, X.; Wang, J.; Sun, Y. Circulating Tumor DNA as Biomarkers for Cancer Detection. Genom. Proteom. Bioinform. 2017, 15, 59–72. [Google Scholar] [CrossRef] [PubMed]
- Mouliere, F.; El Messaoudi, S.; Pang, D.; Dritschilo, A.; Thierry, A.R. Multi-marker analysis of circulating cell-free DNA toward personalized medicine for colorectal cancer. Mol. Oncol. 2014, 8, 927–941. [Google Scholar] [CrossRef] [PubMed]
- Schøler, L.V.; Reinert, T.; Ørntoft, M.-B.W.; Kassentoft, C.G.; Árnadóttir, S.S.; Vang, S.; Nordentoft, I.; Knudsen, M.; Lamy, P.; Andreasen, D.; et al. Clinical Implications of Monitoring Circulating Tumor DNA in Patients with Colorectal Cancer. Clin. Cancer Res. 2017, 23, 5437–5445. [Google Scholar] [CrossRef] [PubMed]
- Kang, J.-K.; Heo, S.; Kim, H.-P.; Song, S.-H.; Yun, H.; Han, S.-W.; Kang, G.H.; Bang, D.; Kim, T.-Y. Liquid biopsy-based tumor profiling for metastatic colorectal cancer patients with ultra-deep targeted sequencing. PLoS ONE 2020, 15, e0232754. [Google Scholar] [CrossRef] [PubMed]
- Demoret, B.; Gregg, J.; Liebner, D.A.; Tinoco, G.; Lenobel, S.; Chen, J.L. Prospective Evaluation of the Concordance of Commercial Circulating Tumor DNA Alterations with Tumor-Based Sequencing across Multiple Soft Tissue Sarcoma Subtypes. Cancers 2019, 11, 1829. [Google Scholar] [CrossRef] [PubMed]
- Perakis, S.; Auer, M.; Belic, J.; Heitzer, E. Advances in Circulating Tumor DNA Analysis. Adv. Clin. Chem. 2017, 80, 73–153. [Google Scholar] [PubMed]
- Zhao, H.; Chen, K.-Z.; Hui, B.-G.; Zhang, K.; Yang, F.; Wang, J. Role of circulating tumor DNA in the management of early-stage lung cancer. Thorac. Cancer 2018, 9, 509–515. [Google Scholar] [CrossRef]
- Rothwell, D.G.; Ayub, M.; Cook, N.; Thistlethwaite, F.; Carter, L.; Dean, E.; Smith, N.; Villa, S.; Dransfield, J.; Clipson, A.; et al. Utility of ctDNA to support patient selection for early phase clinical trials: The TARGET study. Nat. Med. 2019, 25, 738–743. [Google Scholar] [CrossRef]
- El Andaloussi, S.; Mäger, I.; Breakefield, X.O.; Wood, M.J.A. Extracellular vesicles: Biology and emerging therapeutic opportunities. Nat. Rev. Drug Discov. 2013, 12, 347–357. [Google Scholar] [CrossRef] [PubMed]
- Bobrie, A.; Colombo, M.; Krumeich, S.; Raposo, G.; Théry, C. Diverse subpopulations of vesicles secreted by different intracellular mechanisms are present in exosome preparations obtained by differential ultracentrifugation. J. Extracell. Vesicles 2012, 1. [Google Scholar] [CrossRef] [PubMed]
- Record, M.; Carayon, K.; Poirot, M.; Silvente-Poirot, S. Exosomes as new vesicular lipid transporters involved in cell–cell communication and various pathophysiologies. Biochim. Biophys. Acta (BBA) Mol. Cell Biol. Lipids 2014, 1841, 108–120. [Google Scholar] [CrossRef] [PubMed]
- Xiao, H.; Lässer, C.; Shelke, G.V.; Wang, J.; Rådinger, M.; Lunavat, T.R.; Malmhäll, C.; Lin, L.H.; Li, J.; Li, L.; et al. Mast cell exosomes promote lung adenocarcinoma cell proliferation—Role of KIT-stem cell factor signaling. Cell Commun. Signal. 2014, 12, 1–10. [Google Scholar] [CrossRef]
- Borges, F.T.; Melo, S.A.; Özdemir, B.C.; Kato, N.; Revuelta, I.; Miller, C.A.; Ii, V.H.G.; LeBleu, V.S.; Kalluri, R. TGF-β1–Containing Exosomes from Injured Epithelial Cells Activate Fibroblasts to Initiate Tissue Regenerative Responses and Fibrosis. J. Am. Soc. Nephrol. 2012, 24, 385–392. [Google Scholar] [CrossRef]
- Liu, Q.; Rojas-Canales, D.M.; DiVito, S.J.; Shufesky, W.J.; Stolz, D.B.; Erdos, G.; Sullivan, M.L.; Gibson, G.A.; Watkins, S.C.; Larregina, A.T.; et al. Donor dendritic cell–derived exosomes promote allograft-targeting immune response. J. Clin. Investig. 2016, 126, 2805–2820. [Google Scholar] [CrossRef] [PubMed]
- Greening, D.W.; Gopal, S.K.; Mathias, R.A.; Liu, L.; Sheng, J.; Zhu, H.-J.; Simpson, R.J. Emerging roles of exosomes during epithelial–mesenchymal transition and cancer progression. Semin. Cell Dev. Biol. 2015, 40, 60–71. [Google Scholar] [CrossRef] [PubMed]
- Hoshino, A.; Costa-Silva, B.; Shen, T.-L.; Rodrigues, G.; Hashimoto, A.; Mark, M.T.; Molina, H.; Kohsaka, S.; Di Giannatale, A.; Ceder, S.; et al. Tumour exosome integrins determine organotropic metastasis. Nature 2015, 527, 329–335. [Google Scholar] [CrossRef]
- Lv, L.-H.; Wan, Y.-L.; Lin, Y.; Zhang, W.; Yang, M.; Li, G.-L.; Lin, H.-M.; Shang, C.-Z.; Chen, Y.-J.; Min, J. Anticancer Drugs Cause Release of Exosomes with Heat Shock Proteins from Human Hepatocellular Carcinoma Cells That Elicit Effective Natural Killer Cell Antitumor Responses in Vitro. J. Biol. Chem. 2012, 287, 15874–15885. [Google Scholar] [CrossRef]
- Kastelowitz, N.; Yin, H. Exosomes and Microvesicles: Identification and Targeting By Particle Size and Lipid Chemical Probes. ChemBioChem 2014, 15, 923–928. [Google Scholar] [CrossRef]
- Kowal, J.; Tkach, M.; Théry, C. Biogenesis and secretion of exosomes. Curr. Opin. Cell Biol. 2014, 29, 116–125. [Google Scholar] [CrossRef] [PubMed]
- Tauro, B.J.; Greening, D.W.; Mathias, R.A.; Mathivanan, S.; Ji, H.; Simpson, R.J. Two Distinct Populations of Exosomes Are Released from LIM1863 Colon Carcinoma Cell-derived Organoids. Mol. Cell. Proteom. 2013, 12, 587–598. [Google Scholar] [CrossRef]
- Welt, S.; Ritter, G.; Williams, C.; Cohen, L.S.; John, M.; Jungbluth, A.; Richards, A.E.; Old, L.J.; Kemeny, E.N. Phase I study of anticolon cancer humanized antibody A33. Clin. Cancer Res. 2003, 9, 1338–1346. [Google Scholar]
- Liu, D.; Sun, J.; Zhu, J.; Zhou, H.; Zhang, X.; Zhang, Y. Expression and clinical significance of colorectal cancer stem cell marker EpCAMhigh/CD44+ in colorectal cancer. Oncol. Lett. 2014, 7, 1544–1548. [Google Scholar] [CrossRef] [PubMed]
- Bernhard, O.K.; Greening, D.W.; Barnes, T.W.; Ji, H.; Simpson, R.J. Detection of cadherin-17 in human colon cancer LIM1215 cell secretome and tumour xenograft-derived interstitial fluid and plasma. Biochim. Biophys. Acta (BBA) Proteins Proteom. 2013, 1834, 2372–2379. [Google Scholar] [CrossRef] [PubMed]
- Jurčić, P.; Radulović, P.; Balja, M.P.; Milošević, M.; Krušlin, B. E-cadherin and NEDD9 expression in primary colorectal cancer, metastatic lymph nodes and liver metastases. Oncol. Lett. 2019, 17, 2881–2889. [Google Scholar] [CrossRef]
- Chen, Y.; Xie, Y.; Xu, L.; Zhan, S.; Xiao, Y.; Gao, Y.; Wu, B.; Ge, W. Protein content and functional characteristics of serum-purified exosomes from patients with colorectal cancer revealed by quantitative proteomics. Int. J. Cancer 2017, 140, 900–913. [Google Scholar] [CrossRef]
- Ji, H.; Greening, D.W.; Barnes, T.W.; Lim, J.W.; Tauro, B.J.; Rai, A.; Xu, R.; Adda, C.; Mathivanan, S.; Zhao, W.; et al. Proteome profiling of exosomes derived from human primary and metastatic colorectal cancer cells reveal differential expression of key metastatic factors and signal transduction components. Proteomics 2013, 13, 1672–1686. [Google Scholar] [CrossRef] [PubMed]
- Ogata-Kawata, H.; Izumiya, M.; Kurioka, D.; Honma, Y.; Yamada, Y.; Furuta, K.; Gunji, T.; Ohta, H.; Okamoto, H.; Sonoda, H.; et al. Circulating Exosomal microRNAs as Biomarkers of Colon Cancer. PLoS ONE 2014, 9, e92921. [Google Scholar] [CrossRef] [PubMed]
- Lianidou, E.S.; Strati, A.; Markou, A. Circulating tumor cells as promising novel biomarkers in solid cancers. Crit. Rev. Clin. Lab. Sci. 2014, 51, 160–171. [Google Scholar] [CrossRef] [PubMed]
- Cristofanilli, M.; Budd, G.T.; Ellis, M.J.; Stopeck, A.; Matera, J.; Miller, M.C.; Reuben, J.M.; Doyle, G.V.; Allard, W.J.; Terstappen, L.W.; et al. Circulating Tumor Cells, Disease Progression, and Survival in Metastatic Breast Cancer. N. Engl. J. Med. 2004, 351, 781–791. [Google Scholar] [CrossRef]
- Miller, M.C.; Doyle, G.V.; Terstappen, L.W.M.M. Significance of Circulating Tumor Cells Detected by the CellSearch System in Patients with Metastatic Breast Colorectal and Prostate Cancer. J. Oncol. 2009, 2010, 1–8. [Google Scholar] [CrossRef]
- Yang, C.; Xia, B.-R.; Jin, W.-L.; Lou, G. Circulating tumor cells in precision oncology: Clinical applications in liquid biopsy and 3D organoid model. Cancer Cell Int. 2019, 19, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Mueller, M.M.; Fusenig, N.E. Friends or foes—bipolar effects of the tumour stroma in cancer. Nat. Rev. Cancer 2004, 4, 839–849. [Google Scholar] [CrossRef]
- Hardingham, J.E.; Grover, P.; Winter, M.; Hewett, P.J.; Price, T.J.; Thierry, B. Detection and Clinical Significance of Circulating Tumor Cells in Colorectal Cancer—20 Years of Progress. Mol. Med. 2015, 21, 25–31. [Google Scholar] [CrossRef]
- Alix-Panabières, C.; Pantel, K. Circulating Tumor Cells: Liquid Biopsy of Cancer. Clin. Chem. 2013, 59, 110–118. [Google Scholar] [CrossRef] [PubMed]
- Eliasova, P.; Pinkas, M.; Kolostova, K.; Gurlich, R.; Bobek, V. Circulating tumor cells in different stages of colorectal cancer. Folia Histochem. Cytobiol. 2017, 55, 1–5. [Google Scholar] [CrossRef] [PubMed]

| Biomarker | Signification | Structure | Experience/Implication | Reference |
|---|---|---|---|---|
| CEA | carcinoembryonic antigen | glycoprotein | Validated blood biomarker in clinical practice. Not recommended as sole CRC screening test. Preoperative CEA > 5 mg/mL may correlate with poorer CRC prognosis. Used as postoperative serum testing and monitoring during active CRC treatment every 3 months.Diagnostic sensitivity 54.5%; specificity 98.4%. | Locker et al.,2006 [44] Wu et al., 2020 [55] |
| CA 19-9 | carbohydrate antigen | glycoprotein | Validated blood biomarker in clinical practice. Not recommended as sole screening or monitoring CRC marker. Used as supplementary progress monitoring during pancreatic cancer treatment every 1–3 months. Individual values for each patient. Diagnostic sensitivity 64.4%; specificity 96.8%. | Locker et al., 2006 [44] Wu et al., 2020 [55] |
| CTC | circulating tumor cells | tumor cells | Epithelial marker in the peripheral blood via automatic detection system. Detected in different cancer types. 1-10 CTCs per ml blood were found in patients with metastases but rarely in healthy people. Poor prognosis for CRC patients with ≥ 5 CTC per 7.5 ml blood. Diagnostic sensitivity 62.7%; specificity 82.0%. Multivariate analysis of the disease-free survival data of examined patient group showed that a CTC count ≥5 was an independent prognostic factor of distant metastasis (Hazard ratio = 7.5, 95% CI: 1.6 to 34.7, p = 0.01). | Dominguez-Vigil et al., 2018 [49] Tsai et al., 2016 [56] |
| ctDNA | circulating tumor DNA | small DNA fragments released by tumor cells | Tumor mutation search in the peripheral blood, plasma and serum. Patients with 100 g tumor burden released 3.3% of ctDNA into circulation. In CRC, ctDNA is more sensitive than CEA. KRAS mutations were detected with 87.2% sensitivity and a 99.2% specificity. | Osumi et al., 2020 [57] Said et al., 2020 [58] Dominguez-Vigil et al., 2018 [49] |
| exosomes | nanovesicles | vesicular structures released by different cell types, including tumors | Tumor miRNA molecules in biological fluid like blood and urine. Associated with several types of CRC. Each tumor is characterized by specific protein profile. Positive correlation from miRNA exosomes and proteins with the stage of tumor progression. | Dominguez-Vigil et al., 2018 [49] Wang et al., 2016 [59] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Verbanac, D.; Čeri, A.; Hlapčić, I.; Shakibaei, M.; Brockmueller, A.; Krušlin, B.; Ljubičić, N.; Baršić, N.; Detel, D.; Batičić, L.; et al. Profiling Colorectal Cancer in the Landscape Personalized Testing—Advantages of Liquid Biopsy. Int. J. Mol. Sci. 2021, 22, 4327. https://doi.org/10.3390/ijms22094327
Verbanac D, Čeri A, Hlapčić I, Shakibaei M, Brockmueller A, Krušlin B, Ljubičić N, Baršić N, Detel D, Batičić L, et al. Profiling Colorectal Cancer in the Landscape Personalized Testing—Advantages of Liquid Biopsy. International Journal of Molecular Sciences. 2021; 22(9):4327. https://doi.org/10.3390/ijms22094327
Chicago/Turabian StyleVerbanac, Donatella, Andrea Čeri, Iva Hlapčić, Mehdi Shakibaei, Aranka Brockmueller, Božo Krušlin, Neven Ljubičić, Neven Baršić, Dijana Detel, Lara Batičić, and et al. 2021. "Profiling Colorectal Cancer in the Landscape Personalized Testing—Advantages of Liquid Biopsy" International Journal of Molecular Sciences 22, no. 9: 4327. https://doi.org/10.3390/ijms22094327
APA StyleVerbanac, D., Čeri, A., Hlapčić, I., Shakibaei, M., Brockmueller, A., Krušlin, B., Ljubičić, N., Baršić, N., Detel, D., Batičić, L., Rumora, L., Somborac-Bačura, A., Štefanović, M., Ćelap, I., Demirović, A., Petlevski, R., Petrik, J., Grdić Rajković, M., Hulina-Tomašković, A., ... Barišić, K. (2021). Profiling Colorectal Cancer in the Landscape Personalized Testing—Advantages of Liquid Biopsy. International Journal of Molecular Sciences, 22(9), 4327. https://doi.org/10.3390/ijms22094327










