Catalase (CAT) Gene Family in Rapeseed (Brassica napus L.): Genome-Wide Analysis, Identification, and Expression Pattern in Response to Multiple Hormones and Abiotic Stress Conditions
Abstract
1. Introduction
2. Results
2.1. Identification of CAT Genes Family in Rapeseed
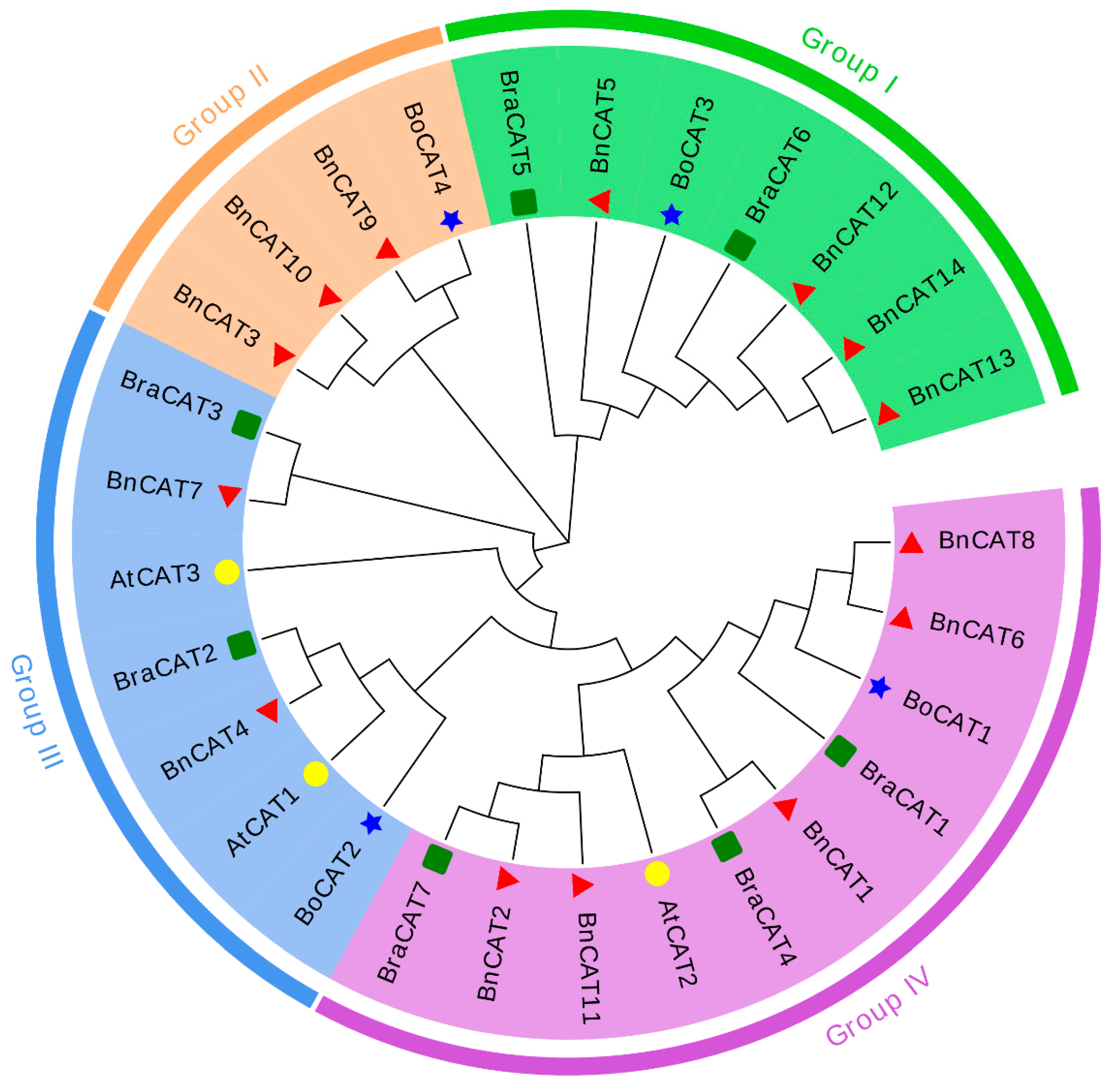
2.2. Phylogenetic Relationships of CAT Genes
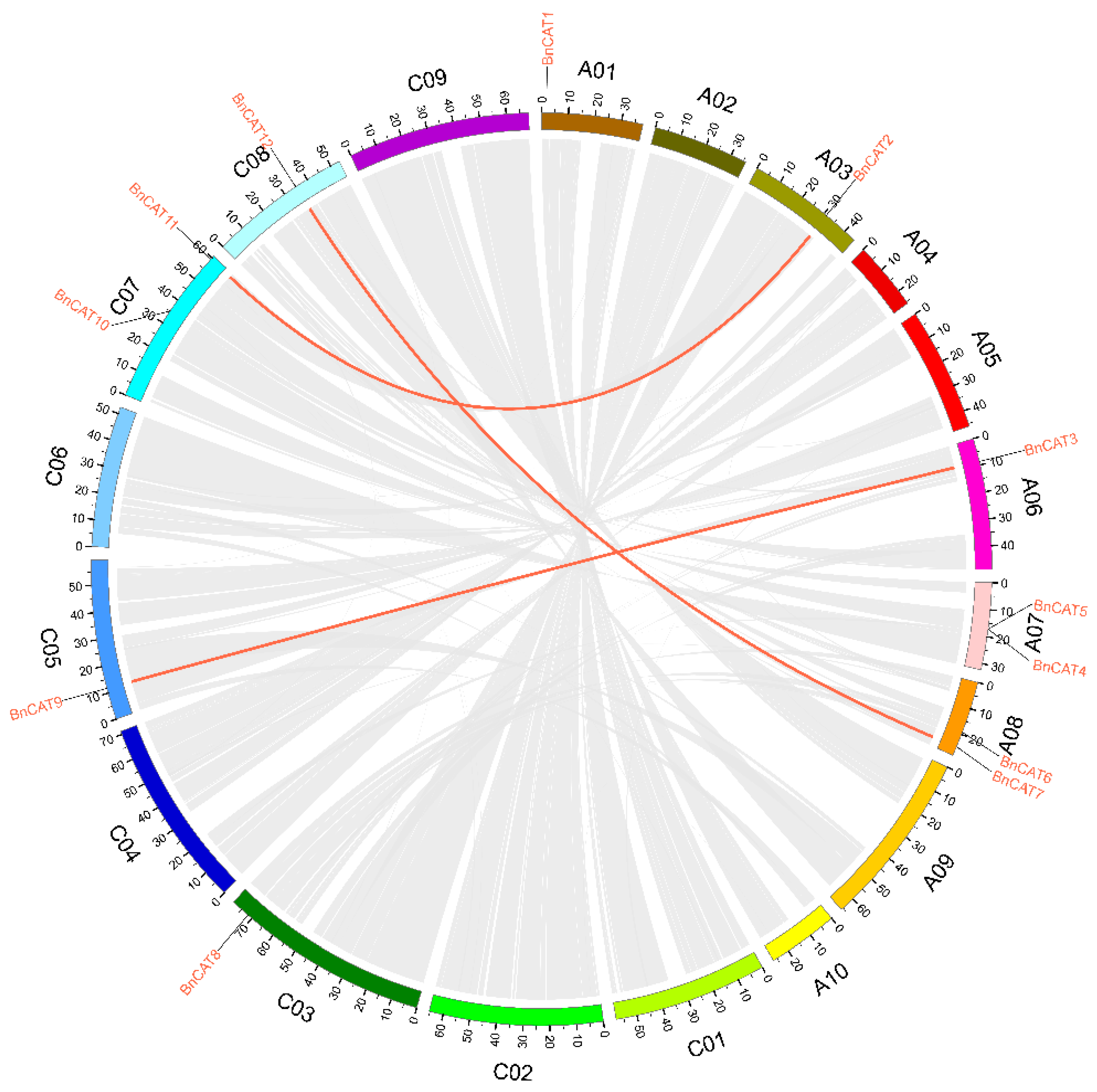
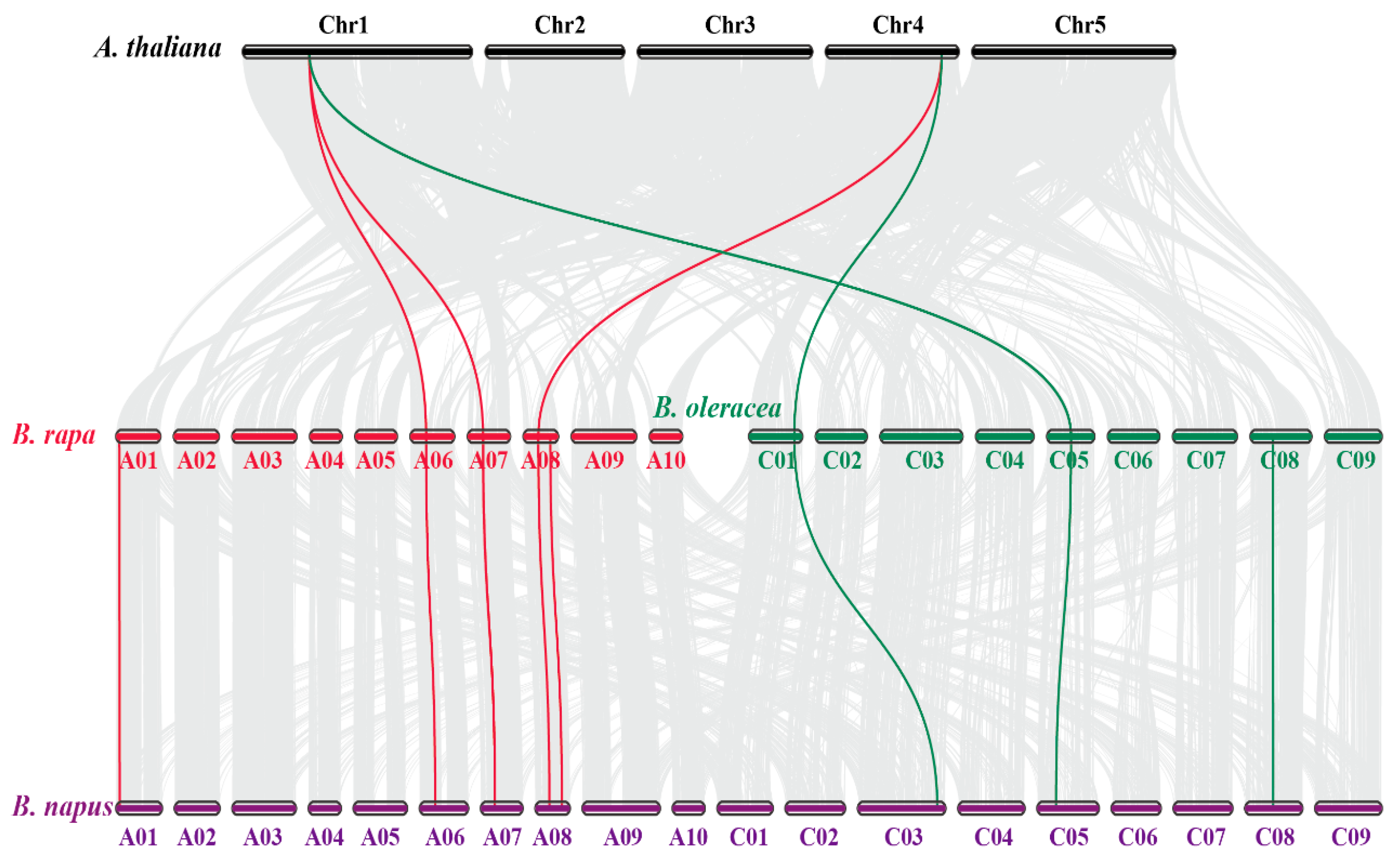
2.3. Synteny Analysis of CAT Genes
2.4. Analysis of BnCAT Gene Structures and Conserved Motifs
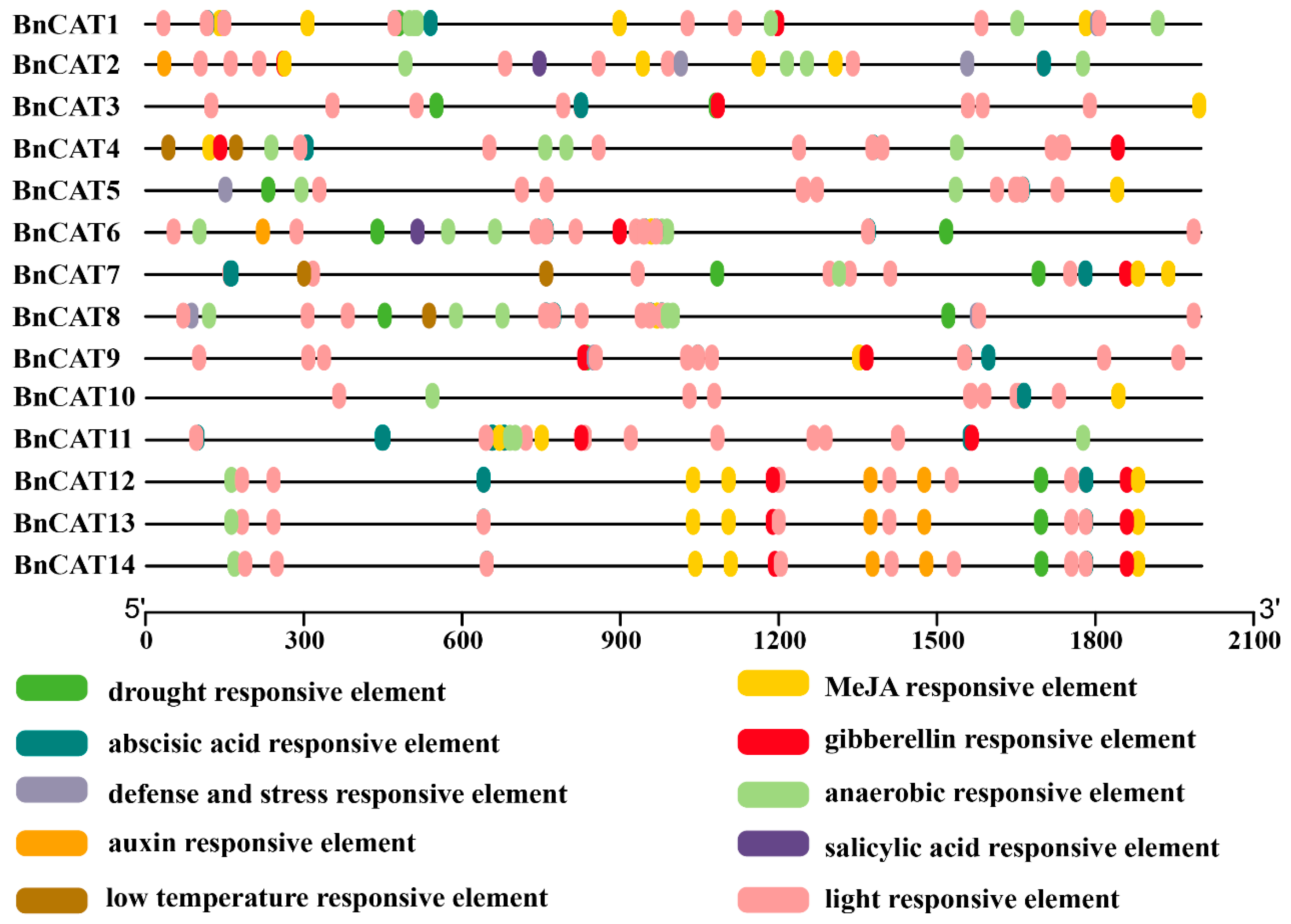
2.5. Cis-Elements in Promoters of BnCAT Genes
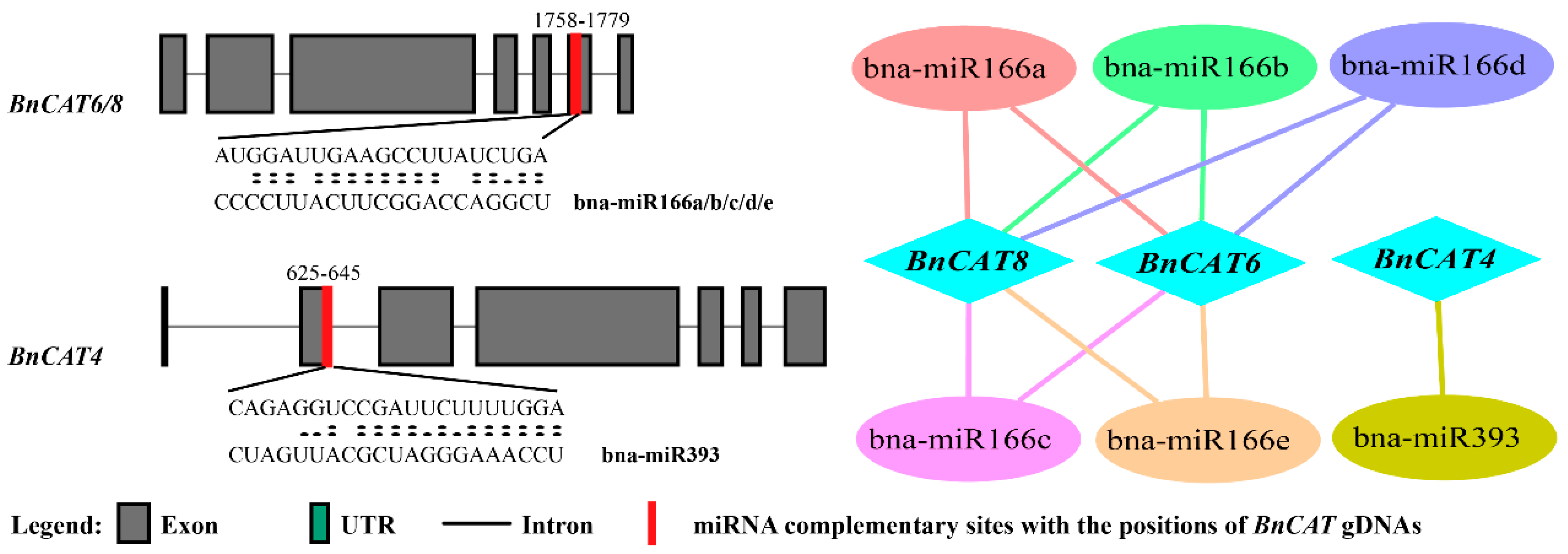
2.6. Genome-Wide Analysis of miRNA Targeting BnCAT Genes
2.7. Functional Annotation Analysis of BnCAT Genes
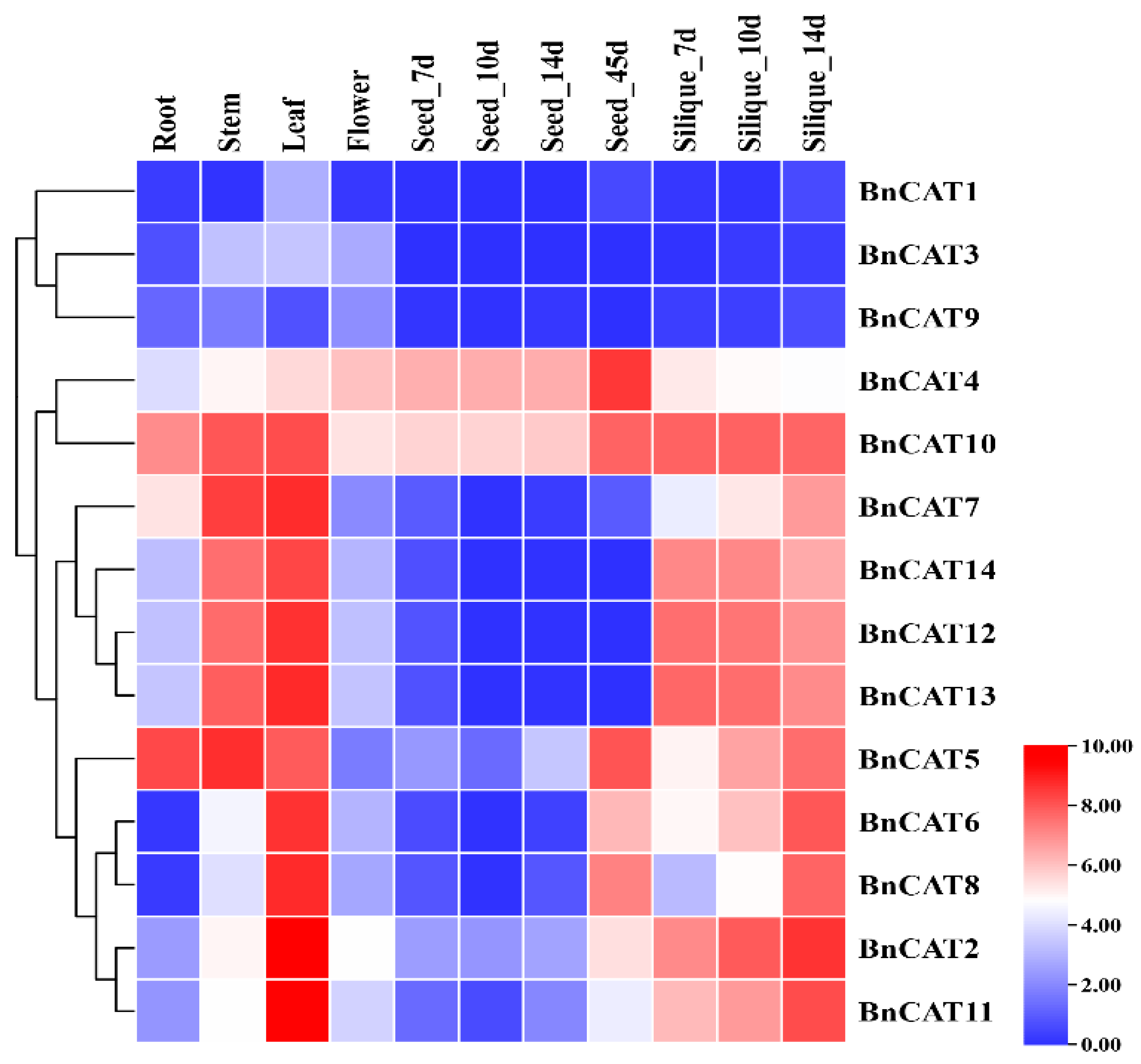
2.8. Expression Profiling of BnCAT Genes in Various Tissues
2.9. Expression Profiles of BnCAT Genes under Phytohormones and Abiotic Stress Conditions
3. Discussion
3.1. Characterization of CAT Gene Family in Rapeseed
3.2. The Critical Role of CAT Genes in ROS Metabolism, Hormone, and Abiotic Stress Responses
3.3. Expression Pattern of CAT Genes in Various Tissues
3.4. miRNA: Key Players in the Regulation of Stress Responses
4. Materials and Methods
4.1. Identification of CAT Family Genes
4.2. Characterization of CAT Family Genes
4.3. Phylogenetic Tree and Synteny Analysis of BnCAT Family Proteins
4.4. Cis-Elements Analysis in the BnCAT Gene Promoters
4.5. Prediction of Putative miRNA Targeting BnCAT Genes and Functional Annotation Analysis
4.6. Expression Analysis of BnCAT Genes in Different Tissues
4.7. Plant Material and Stress Conditions
4.8. RNA Extraction and qRT-PCR Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Raza, A.; Razzaq, A.; Mehmood, S.S.; Zou, X.; Zhang, X.; Lv, Y.; Xu, J. Impact of climate change on crops adaptation and strategies to tackle its outcome: A review. Plants 2019, 8, 34. [Google Scholar] [CrossRef]
- Hasanuzzaman, M.; Bhuyan, M.; Zulfiqar, F.; Raza, A.; Mohsin, S.M.; Mahmud, J.A.; Fujita, M.; Fotopoulos, V. Reactive Oxygen Species and Antioxidant Defense in Plants under Abiotic Stress: Revisiting the Crucial Role of a Universal Defense Regulator. Antioxidants 2020, 9, 681. [Google Scholar] [CrossRef] [PubMed]
- Mittler, R. ROS are good. Trends Plant. Sci. 2017, 22, 11–19. [Google Scholar] [CrossRef]
- Mhamdi, A.; Queval, G.; Chaouch, S.; Vanderauwera, S.; Van Breusegem, F.; Noctor, G. Catalase function in plants: A focus on Arabidopsis mutants as stress-mimic models. J. Exp. Bot. 2010, 61, 4197–4220. [Google Scholar] [CrossRef]
- Anjum, N.A.; Sharma, P.; Gill, S.S.; Hasanuzzaman, M.; Khan, E.A.; Kachhap, K.; Mohamed, A.A.; Thangavel, P.; Devi, G.D.; Vasudhevan, P. Catalase and ascorbate peroxidase—Representative H2O2-detoxifying heme enzymes in plants. Environ. Sci. Pollut. Res. 2016, 23, 19002–19029. [Google Scholar] [CrossRef] [PubMed]
- Kaushal, J.; Mehandia, S.; Singh, G.; Raina, A.; Arya, S.K. Catalase enzyme: Application in bioremediation and food industry. Biocatal. Agric. Biotechnol. 2018, 16, 192–199. [Google Scholar] [CrossRef]
- GONZÁLEZ, E. The C-terminal domain of plant catalases Implications for a glyoxysomal targeting sequence. Eur. J. Biochem. 1991, 199, 211–215. [Google Scholar] [CrossRef] [PubMed]
- Drory, A.; Woodson, W.R. Molecular cloning and nucleotide sequence of a cDNA encoding catalase from tomato. Plant Physiol. 1992, 100, 1605. [Google Scholar] [CrossRef]
- Chen, H.-J.; Wu, S.-D.; Huang, G.-J.; Shen, C.-Y.; Afiyanti, M.; Li, W.-J.; Lin, Y.-H. Expression of a cloned sweet potato catalase SPCAT1 alleviates ethephon-mediated leaf senescence and H2O2 elevation. J. Plant Physiol. 2012, 169, 86–97. [Google Scholar] [CrossRef] [PubMed]
- Skadsen, R.W.; Schulze-Lefert, P.; Herbst, J.M. Molecular cloning, characterization and expression analysis of two catalase isozyme genes in barley. Plant Mol. Biol. 1995, 29, 1005–1014. [Google Scholar] [CrossRef]
- Willekens, H.; Villarroel, R.; Van Montagu, M.; Inzé, D.; Van Camp, W. Molecular identification of catalases from Nicotiana plumbaginifolia (L.). FEBS Lett. 1994, 352, 79–83. [Google Scholar] [CrossRef]
- Guan, L.; Scandalios, J.G. Developmentally related responses of maize catalase genes to salicylic acid. Proc. Nat. Acad. Sci. USA 1995, 92, 5930–5934. [Google Scholar] [CrossRef] [PubMed]
- Esaka, M.; Yamada, N.; Kitabayashi, M.; Setoguchi, Y.; Tsugeki, R.; Kondo, M.; Nishimura, M. cDNA cloning and differential gene expression of three catalases in pumpkin. Plant Mol. Biol. 1997, 33, 141–155. [Google Scholar] [CrossRef] [PubMed]
- Du, Y.Y.; Wang, P.C.; Chen, J.; Song, C.P. Comprehensive functional analysis of the catalase gene family in Arabidopsis thaliana. J. Integr. Plant Biol. 2008, 50, 1318–1326. [Google Scholar] [CrossRef] [PubMed]
- Joo, J.; Lee, Y.H.; Song, S.I. Rice CatA, CatB, and CatC are involved in environmental stress response, root growth, and photorespiration, respectively. J. Plant Biol. 2014, 57, 375–382. [Google Scholar] [CrossRef]
- Hu, L.; Yang, Y.; Jiang, L.; Liu, S. The catalase gene family in cucumber: Genome-wide identification and organization. Genet. Mol. Biol. 2016, 39, 408–415. [Google Scholar] [CrossRef] [PubMed]
- Alam, N.B.; Ghosh, A. Comprehensive analysis and transcript profiling of Arabidopsis thaliana and Oryza sativa catalase gene family suggests their specific roles in development and stress responses. Plant Physiol. Biochem. 2018, 123, 54–64. [Google Scholar] [CrossRef]
- Wang, W.; Cheng, Y.; Chen, D.; Liu, D.; Hu, M.; Dong, J.; Zhang, X.; Song, L.; Shen, F. The catalase gene family in cotton: Genome-wide characterization and bioinformatics analysis. Cells 2019, 8, 86. [Google Scholar] [CrossRef]
- Inaba, J.-I.; Kim, B.M.; Shimura, H.; Masuta, C. Virus-induced necrosis is a consequence of direct protein-protein interaction between a viral RNA-silencing suppressor and a host catalase. Plant Physiol. 2011, 156, 2026–2036. [Google Scholar] [CrossRef]
- Yong, B.; Wang, X.; Xu, P.; Zheng, H.; Fei, X.; Hong, Z.; Ma, Q.; Miao, Y.; Yuan, X.; Jiang, Y.; et al. Isolation and abiotic stress resistance analyses of a catalase gene from Ipomoea batatas (L.) Lam. BioMed Res. Inter. 2017, 2017. [Google Scholar] [CrossRef] [PubMed]
- Su, Y.; Guo, J.; Ling, H.; Chen, S.; Wang, S.; Xu, L.; Allan, A.C.; Que, Y. Isolation of a novel peroxisomal catalase gene from sugarcane, which is responsive to biotic and abiotic stresses. PLoS ONE 2014, 9, e84426. [Google Scholar] [CrossRef]
- Balestrasse, K.B.; Yannarelli, G.G.; Noriega, G.O.; Batlle, A.; Tomaro, M.L. Heme oxygenase and catalase gene expression in nodules and roots of soybean plants subjected to cadmium stress. Biometals 2008, 21, 433–441. [Google Scholar] [CrossRef]
- Lin, K.-H.; Huang, H.-C.; Lin, C.-Y. Cloning, expression and physiological analysis of broccoli catalase gene and Chinese cabbage ascorbate peroxidase gene under heat stress. Plant Cell Rep. 2010, 29, 575–593. [Google Scholar] [CrossRef] [PubMed]
- Queval, G.; Issakidis-Bourguet, E.; Hoeberichts, F.A.; Vandorpe, M.; Gakière, B.; Vanacker, H.; Miginiac-Maslow, M.; Van Breusegem, F.; Noctor, G. Conditional oxidative stress responses in the Arabidopsis photorespiratory mutant cat2 demonstrate that redox state is a key modulator of daylength-dependent gene expression, and define photoperiod as a crucial factor in the regulation of H2O2-induced cell death. Plant J. 2007, 52, 640–657. [Google Scholar]
- Contento, A.L.; Bassham, D.C. Increase in catalase-3 activity as a response to use of alternative catabolic substrates during sucrose starvation. Plant Physiol. Biochem. 2010, 48, 232–238. [Google Scholar] [CrossRef] [PubMed]
- Polidoros, A.; Mylona, P.; Scandalios, J. Transgenic tobacco plants expressing the maize Cat2 gene have altered catalase levels that affect plant-pathogen interactions and resistance to oxidative stress. Trans. Res. 2001, 10, 555–569. [Google Scholar] [CrossRef] [PubMed]
- Lee, S.H.; An, C.S. Differential expression of three catalase genes in hot pepper (Capsicum annuum L.). Mol. Cells 2005, 20, 247–255. [Google Scholar]
- Niewiadomska, E.; Polzien, L.; Desel, C.; Rozpadek, P.; Miszalski, Z.; Krupinska, K. Spatial patterns of senescence and development-dependent distribution of reactive oxygen species in tobacco (Nicotiana tabacum) leaves. J. Plant Physiol. 2009, 166, 1057–1068. [Google Scholar] [CrossRef] [PubMed]
- Vuosku, J.; Sutela, S.; Kestilä, J.; Jokela, A.; Sarjala, T.; Häggman, H. Expression of catalase and retinoblastoma-related protein genes associates with cell death processes in Scots pine zygotic embryogenesis. BMC Plant Biol. 2015, 15, 88. [Google Scholar] [CrossRef]
- Cui, C.; Wang, J.-J.; Zhao, J.-H.; Fang, Y.-Y.; He, X.-F.; Guo, H.-S.; Duan, C.-G. A Brassica miRNA Regulates Plant Growth and Immunity through Distinct Modes of Action. Mol. Plant 2020, 13, 231–245. [Google Scholar] [CrossRef]
- Khandal, H.; Parween, S.; Roy, R.; Meena, M.K.; Chattopadhyay, D. MicroRNA profiling provides insights into post-transcriptional regulation of gene expression in chickpea root apex under salinity and water deficiency. Sci. Rep. 2017, 7, 1–14. [Google Scholar] [CrossRef]
- Ding, Y.; Ding, L.; Xia, Y.; Wang, F.; Zhu, C. Emerging roles of microRNAs in plant heavy metal tolerance and homeostasis. J. Agric. Food Chem. 2020, 68, 1958–1965. [Google Scholar] [CrossRef] [PubMed]
- Ravichandran, S.; Ragupathy, R.; Edwards, T.; Domaratzki, M.; Cloutier, S. MicroRNA-guided regulation of heat stress response in wheat. BMC Genom. 2019, 20, 488. [Google Scholar] [CrossRef]
- Park, S.-Y.; Grabau, E. Bypassing miRNA-mediated gene regulation under drought stress: Alternative splicing affects CSD1 gene expression. Plant Mol. Biol. 2017, 95, 243–252. [Google Scholar] [CrossRef]
- SHI, G.-q.; FU, J.-y.; RONG, L.-j.; ZHANG, P.-y.; GUO, C.-j.; Kai, X. TaMIR1119, a miRNA family member of wheat (Triticum aestivum), is essential in the regulation of plant drought tolerance. J. Integr. Agric. 2018, 17, 2369–2378. [Google Scholar] [CrossRef]
- Ci, D.; Song, Y.; Tian, M.; Zhang, D. Methylation of miRNA genes in the response to temperature stress in Populus simonii. Front. Plant Sci. 2015, 6, 921. [Google Scholar] [CrossRef]
- Raza, A. Eco-physiological and Biochemical Responses of Rapeseed (Brassica napus L.) to Abiotic Stresses: Consequences and Mitigation Strategies. J. Plant Growth Regul. 2020. [Google Scholar] [CrossRef]
- Raza, A.; Hafeez, M.B.; Zahra, N.; Shaukat, K.; Umbreen, S.; Tabassum, J.; Charagh, S.; Khan, R.S.A.; Hasanuzzaman, M. The Plant Family Brassicaceae: Introduction, Biology, and Importance. In The Plant Family Brassicaceae; Hasanuzzaman, M., Ed.; Springer: Singapore, 2020; pp. 1–43. [Google Scholar]
- He, H.; Lei, Y.; Yi, Z.; Raza, A.; Zeng, L.; Yan, L.; Xiaoyu, D.; Yong, C.; Xiling, Z. Study on the mechanism of exogenous serotonin improving cold tolerance of rapeseed (Brassica napus L.) seedlings. Plant Growth Regul. 2021. [Google Scholar] [CrossRef]
- Song, J.-M.; Guan, Z.; Hu, J.; Guo, C.; Yang, Z.; Wang, S.; Liu, D.; Wang, B.; Lu, S.; Zhou, R.; et al. Eight high-quality genomes reveal pan-genome architecture and ecotype differentiation of Brassica napus. Nat. Plants 2020, 6, 34–45. [Google Scholar] [CrossRef] [PubMed]
- Cavell, A.C.; Lydiate, D.; Parkin, I.; Dean, C.; Trick, M. Collinearity between a 30-centimorgan segment of Arabidopsis thaliana chromosome 4 and duplicated regions within the Brassica napus genome. Genome 1998, 41, 62–69. [Google Scholar] [CrossRef] [PubMed]
- Frugoli, J.A.; McPeek, M.A.; Thomas, T.L.; McClung, C.R. Intron loss and gain during evolution of the catalase gene family in angiosperms. Genetics 1998, 149, 355–365. [Google Scholar] [CrossRef] [PubMed]
- Zou, J.-J.; Li, X.-D.; Ratnasekera, D.; Wang, C.; Liu, W.-X.; Song, L.-F.; Zhang, W.-Z.; Wu, W.-H. Arabidopsis CALCIUM-DEPENDENT PROTEIN KINASE8 and CATALASE3 function in abscisic acid-mediated signaling and H2O2 homeostasis in stomatal guard cells under drought stress. Plant Cell 2015, 27, 1445–1460. [Google Scholar] [CrossRef]
- Hackenberg, T.; Juul, T.; Auzina, A.; Gwiżdż, S.; Małolepszy, A.; Van Der Kelen, K.; Dam, S.; Bressendorff, S.; Lorentzen, A.; Roepstorff, P. Catalase and NO CATALASE ACTIVITY1 promote autophagy-dependent cell death in Arabidopsis. Plant Cell 2013, 25, 4616–4626. [Google Scholar] [CrossRef]
- Corpas, F.J.; Barroso, J.B.; González-Gordo, S.; Muñoz-Vargas, M.A.; Palma, J.M. Hydrogen sulfide: A novel component in Arabidopsis peroxisomes which triggers catalase inhibition. J. Integr. Plant Biol. 2019, 61, 871–883. [Google Scholar] [CrossRef]
- Li, J.; Liu, J.; Wang, G.; Cha, J.-Y.; Li, G.; Chen, S.; Li, Z.; Guo, J.; Zhang, C.; Yang, Y. A chaperone function of NO CATALASE ACTIVITY1 is required to maintain catalase activity and for multiple stress responses in Arabidopsis. Plant Cell 2015, 27, 908–925. [Google Scholar] [CrossRef]
- Chiang, C.-M.; Chen, S.-P.; Chen, L.-F.O.; Chiang, M.-C.; Chien, H.-L.; Lin, K.-H. Expression of the broccoli catalase gene (BoCAT) enhances heat tolerance in transgenic Arabidopsis. J. Plant Biochem. Biotechnol. 2014, 23, 266–277. [Google Scholar] [CrossRef]
- Zhou, Y.-B.; Liu, C.; Tang, D.-Y.; Yan, L.; Wang, D.; Yang, Y.-Z.; Gui, J.-S.; Zhao, X.-Y.; Li, L.-G.; Tang, X.-D.; et al. The receptor-like cytoplasmic kinase STRK1 phosphorylates and activates CatC, thereby regulating H2O2 homeostasis and improving salt tolerance in rice. Plant Cell 2018, 30, 1100–1118. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Cui, L.; Xie, Z.; Zhang, Z.; Liu, E.; Peng, X. Two NCA1 isoforms interact with catalase in a mutually exclusive manner to redundantly regulate its activity in rice. BMC Plant Biol. 2019, 19, 105. [Google Scholar] [CrossRef]
- Shahpiri, A.; Deljoonia, R. Enhancement of catalase and superoxide dismutase activities in transgenic Escherichia coli expressing rice metallothionein isoforms. J. BioSci. Biotechnol. 2018, 7, 5–10. [Google Scholar]
- Matsumura, T.; Tabayashi, N.; Kamagata, Y.; Souma, C.; Saruyama, H. Wheat catalase expressed in transgenic rice can improve tolerance against low temperature stress. Physiol. Plant 2002, 116, 317–327. [Google Scholar] [CrossRef]
- Hameed, A.; Naseer, S.; Iqbal, T.; Syed, H.; Haq, M.A. Effects of NaCl salinity on seedling growth, senescence, catalase and protease activities in two wheat genotypes differing in salt tolerance. Pak. J. Bot. 2008, 40, 1043–1051. [Google Scholar]
- Srivalli, B.; Khanna-Chopra, R. Induction of new isoforms of superoxide dismutase and catalase enzymes in the flag leaf of wheat during monocarpic senescence. Biochem. Biophys. Res. Commun. 2001, 288, 1037–1042. [Google Scholar] [CrossRef]
- dos Santos, C.M.; de Almeida Silva, M.; Lima, G.P.P.; Bortolheiro, F.P.d.A.P.; Brunelli, M.C.; de Holanda, L.A.; Oliver, R. Physiological changes associated with antioxidant enzymes in response to sugarcane tolerance to water deficit and rehydration. Sugar Tech. 2015, 17, 291–304. [Google Scholar] [CrossRef]
- Liu, Y.; Hu, X.; Yao, Y.; Xu, L.; Xing, S. Isolation and expression analysis of catalase genes in Erianthus arundinaceus and sugarcane. Sugar Tech. 2016, 18, 468–477. [Google Scholar] [CrossRef]
- Zhou, Y.; Liu, S.; Yang, Z.; Yang, Y.; Jiang, L.; Hu, L. CsCAT3, a catalase gene from Cucumis sativus, confers resistance to a variety of stresses to Escherichia coli. Biotechnol. Biotechnol. Equip. 2017, 31, 886–896. [Google Scholar] [CrossRef]
- Murota, K.; Shimura, H.; Takeshita, M.; Masuta, C. Interaction between Cucumber mosaic virus 2b protein and plant catalase induces a specific necrosis in association with proteasome activity. Plant Cell Rep. 2017, 36, 37–47. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q.; Li, C.X.; Fan, X.; Wang, P.; Cui, L. Immobilization of catalase on cotton fabric oxidized by sodium periodate. Biocatal. Biotrans. 2008, 26, 437–443. [Google Scholar] [CrossRef]
- Luo, X.; Wu, J.; Li, Y.; Nan, Z.; Guo, X.; Wang, Y.; Zhang, A.; Wang, Z.; Xia, G.; Tian, Y. Synergistic effects of GhSOD1 and GhCAT1 overexpression in cotton chloroplasts on enhancing tolerance to methyl viologen and salt stresses. PLoS ONE 2013, 8, e54002. [Google Scholar] [CrossRef] [PubMed]
- Duman, Y.A.; Kaya, E. Three-phase partitioning as a rapid and easy method for the purification and recovery of catalase from sweet potato tubers (Solanum tuberosum). Appl. Biochem. Biotechnol. 2013, 170, 1119–1126. [Google Scholar] [CrossRef]
- Afiyanti, M.; Chen, H.-J. Catalase activity is modulated by calcium and calmodulin in detached mature leaves of sweet potato. J. Plant Physiol. 2014, 171, 35–47. [Google Scholar] [CrossRef]
- Xing, Y.; Jia, W.; Zhang, J. AtMEK1 mediates stress-induced gene expression of CAT1 catalase by triggering H2O2 production in Arabidopsis. J. Exp. Bot. 2007, 58, 2969–2981. [Google Scholar] [CrossRef]
- Guo, P.; Li, Z.; Huang, P.; Li, B.; Fang, S.; Chu, J.; Guo, H. A tripartite amplification loop involving the transcription factor WRKY75, salicylic acid, and reactive oxygen species accelerates leaf senescence. Plant Cell 2017, 29, 2854–2870. [Google Scholar] [CrossRef]
- Corpas, F.J.; Barroso, J.B. Lead-induced stress, which triggers the production of nitric oxide (NO) and superoxide anion (O2·-) in Arabidopsis peroxisomes, affects catalase activity. Nitric Oxide 2017, 68, 103–110. [Google Scholar] [CrossRef]
- Vighi, I.; Benitez, L.; do Amaral, M.; Auler, P.; Moraes, G.; Rodrigues, G.; Da Maia, L.; Pinto, L.; Braga, E. Changes in gene expression and catalase activity in Oryza sativa L. under abiotic stress. Genet. Mol. Res. 2016, 15, gmr15048977. [Google Scholar] [CrossRef] [PubMed]
- Wang, W.; Liu, D.; Zhang, X.; Chen, D.; Cheng, Y.; Shen, F. Plant microRNAs in cross-kingdom regulation of gene expression. Int. J. Mol. Sci. 2018, 19, 2007. [Google Scholar] [CrossRef] [PubMed]
- Huang, S.Q.; Xiang, A.L.; Che, L.L.; Chen, S.; Li, H.; Song, J.B.; Yang, Z.M. A set of miRNAs from Brassica napus in response to sulphate deficiency and cadmium stress. Plant Biotechnol. J. 2010, 8, 887–899. [Google Scholar] [CrossRef]
- Zhou, Z.S.; Song, J.B.; Yang, Z.M. Genome-wide identification of Brassica napus microRNAs and their targets in response to cadmium. J. Exp. Bot. 2012, 63, 4597–4613. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.; Chen, L.; Zhang, X.; Liu, T.; Niu, S.; Wen, J.; Yi, B.; Ma, C.; Tu, J.; Fu, T. Identification of miRNAs that regulate silique development in Brassica napus. Plant Sci. 2018, 269, 106–117. [Google Scholar] [CrossRef]
- Buhtz, A.; Springer, F.; Chappell, L.; Baulcombe, D.C.; Kehr, J. Identification and characterization of small RNAs from the phloem of Brassica napus. Plant J. 2008, 53, 739–749. [Google Scholar] [CrossRef] [PubMed]
- Fu, Y.; Mason, A.S.; Zhang, Y.; Lin, B.; Xiao, M.; Fu, D.; Yu, H. MicroRNA-mRNA expression profiles and their potential role in cadmium stress response in Brassica napus. BMC Plant Biol. 2019, 19, 1–20. [Google Scholar] [CrossRef] [PubMed]
- Körbes, A.P.; Machado, R.D.; Guzman, F.; Almerao, M.P.; de Oliveira, L.F.V.; Loss-Morais, G.; Turchetto-Zolet, A.C.; Cagliari, A.; dos Santos Maraschin, F.; Margis-Pinheiro, M. Identifying conserved and novel microRNAs in developing seeds of Brassica napus using deep sequencing. PLoS ONE 2012, 7, e50663. [Google Scholar] [CrossRef] [PubMed]
- Shen, E.; Zou, J.; Hubertus Behrens, F.; Chen, L.; Ye, C.; Dai, S.; Li, R.; Ni, M.; Jiang, X.; Qiu, J. Identification, evolution, and expression partitioning of miRNAs in allopolyploid Brassica napus. J. Exp. Bot. 2015, 66, 7241–7253. [Google Scholar] [CrossRef] [PubMed]
- Casati, P. Analysis of UV-B regulated miRNAs and their targets in maize leaves. Plant Signal. Behav. 2013, 8, e26758. [Google Scholar] [CrossRef] [PubMed]
- Li, S.; Yu, X.; Lei, N.; Cheng, Z.; Zhao, P.; He, Y.; Wang, W.; Peng, M. Genome-wide identification and functional prediction of cold and/or drought-responsive lncRNAs in cassava. Sci. Rep. 2017, 7, 45981. [Google Scholar] [CrossRef] [PubMed]
- Nageshbabu, R.; Jyothi, M.; Sharadamma, N. Expression of miRNAs regulates growth and development of French bean (Phaseolus vulgaris) under salt and drought stress conditions. ISCA J. Biol. Sci. 2013, 2, 52–56. [Google Scholar]
- Zhang, Y.; Zhu, X.; Chen, X.; Song, C.; Zou, Z.; Wang, Y.; Wang, M.; Fang, W.; Li, X. Identification and characterization of cold-responsive microRNAs in tea plant (Camellia sinensis) and their targets using high-throughput sequencing and degradome analysis. BMC Plant Biol. 2014, 14, 271. [Google Scholar] [CrossRef]
- Yang, Z.; Wang, Z.; Yang, C.; Yang, Z.; Li, H.; Wu, Y. Physiological responses and small RNAs changes in maize under nitrogen deficiency and resupply. Genes Genom. 2019, 41, 1183–1194. [Google Scholar] [CrossRef]
- Wang, Z.; Chen, Y.; Fang, H.; Shi, H.; Chen, K.; Zhang, Z.; Tan, X. Selection of reference genes for quantitative reverse-transcription polymerase chain reaction normalization in Brassica napus under various stress conditions. Mol. Genet. Genom. 2014, 289, 1023–1035. [Google Scholar] [CrossRef] [PubMed]


| Gene ID | Gene Name | Genomic Position (bp) | Gene Length (bp) | CDS Length (bp) | Exon | Protein Length (Amino Acids) | Molecular Weight (kDa) | Isoelectric Point (pI) | Sub-Cellular Localization |
|---|---|---|---|---|---|---|---|---|---|
| BnaA01T0032600ZS | BnCAT1 | A01:1801796-1803991 + | 2195 | 1488 | 4 | 495 | 57.30 | 6.5 | Cytoskeleton |
| BnaA03T0547900ZS | BnCAT2 | A03:31026573-31028922 − | 2349 | 1479 | 5 | 492 | 56.84 | 6.63 | Peroxisome |
| BnaA06T0144600ZS | BnCAT3 | A06:8685726-8691515 + | 5789 | 2925 | 12 | 974 | 112.30 | 6.78 | Peroxisome |
| BnaA07T0132000ZS | BnCAT4 | A07:17121532-17124087 − | 2555 | 1479 | 5 | 492 | 56.90 | 6.83 | Cytoplasm |
| BnaA07T0132100ZS | BnCAT5 | A07:17124704-17130749 − | 6045 | 1671 | 6 | 556 | 64.83 | 8.42 | Chloroplast |
| BnaA08T0131200ZS | BnCAT6 | A08:18424311-18426301 + | 1990 | 1470 | 4 | 489 | 565.40 | 6.9 | Peroxisome |
| BnaA08T0247000ZS | BnCAT7 | A08:25020319-25023539 − | 3220 | 1479 | 5 | 492 | 56.80 | 7.69 | Peroxisome |
| BnaC03T0740800ZS | BnCAT8 | C03:71851456-71853466 + | 2010 | 1470 | 4 | 489 | 56.52 | 6.77 | Peroxisome |
| BnaC05T0176000ZS | BnCAT9 | C05:11786858-11795441 + | 8583 | 2949 | 12 | 982 | 113.09 | 6.94 | Mitochondrion |
| BnaC07T0194100ZS | BnCAT10 | C07:32771906-32781676 − | 9770 | 3123 | 15 | 1040 | 120.71 | 7.34 | Chloroplast |
| BnaC07T0524400ZS | BnCAT11 | C07:59429784-59432167 − | 2383 | 1479 | 4 | 492 | 56.78 | 6.63 | Peroxisome |
| BnaC08T0260800ZS | BnCAT12 | C08:34921906-34925039 + | 3133 | 1479 | 5 | 492 | 56.80 | 7.69 | Peroxisome |
| Bnascaffold0025T0038400ZS | BnCAT13 | 0025:3787780-3790910 − | 3130 | 1434 | 5 | 477 | 55.19 | 7.72 | Peroxisome |
| Bnascaffold0026T0027200ZS | BnCAT14 | 0026:2854478-2857610 + | 3132 | 1479 | 5 | 492 | 56.80 | 7.69 | Peroxisome |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Raza, A.; Su, W.; Gao, A.; Mehmood, S.S.; Hussain, M.A.; Nie, W.; Lv, Y.; Zou, X.; Zhang, X. Catalase (CAT) Gene Family in Rapeseed (Brassica napus L.): Genome-Wide Analysis, Identification, and Expression Pattern in Response to Multiple Hormones and Abiotic Stress Conditions. Int. J. Mol. Sci. 2021, 22, 4281. https://doi.org/10.3390/ijms22084281
Raza A, Su W, Gao A, Mehmood SS, Hussain MA, Nie W, Lv Y, Zou X, Zhang X. Catalase (CAT) Gene Family in Rapeseed (Brassica napus L.): Genome-Wide Analysis, Identification, and Expression Pattern in Response to Multiple Hormones and Abiotic Stress Conditions. International Journal of Molecular Sciences. 2021; 22(8):4281. https://doi.org/10.3390/ijms22084281
Chicago/Turabian StyleRaza, Ali, Wei Su, Ang Gao, Sundas Saher Mehmood, Muhammad Azhar Hussain, Wenlong Nie, Yan Lv, Xiling Zou, and Xuekun Zhang. 2021. "Catalase (CAT) Gene Family in Rapeseed (Brassica napus L.): Genome-Wide Analysis, Identification, and Expression Pattern in Response to Multiple Hormones and Abiotic Stress Conditions" International Journal of Molecular Sciences 22, no. 8: 4281. https://doi.org/10.3390/ijms22084281
APA StyleRaza, A., Su, W., Gao, A., Mehmood, S. S., Hussain, M. A., Nie, W., Lv, Y., Zou, X., & Zhang, X. (2021). Catalase (CAT) Gene Family in Rapeseed (Brassica napus L.): Genome-Wide Analysis, Identification, and Expression Pattern in Response to Multiple Hormones and Abiotic Stress Conditions. International Journal of Molecular Sciences, 22(8), 4281. https://doi.org/10.3390/ijms22084281








