Could MicroRNAs Be Useful Tools to Improve the Diagnosis and Treatment of Rare Gynecological Cancers? A Brief Overview
Abstract
1. Introduction
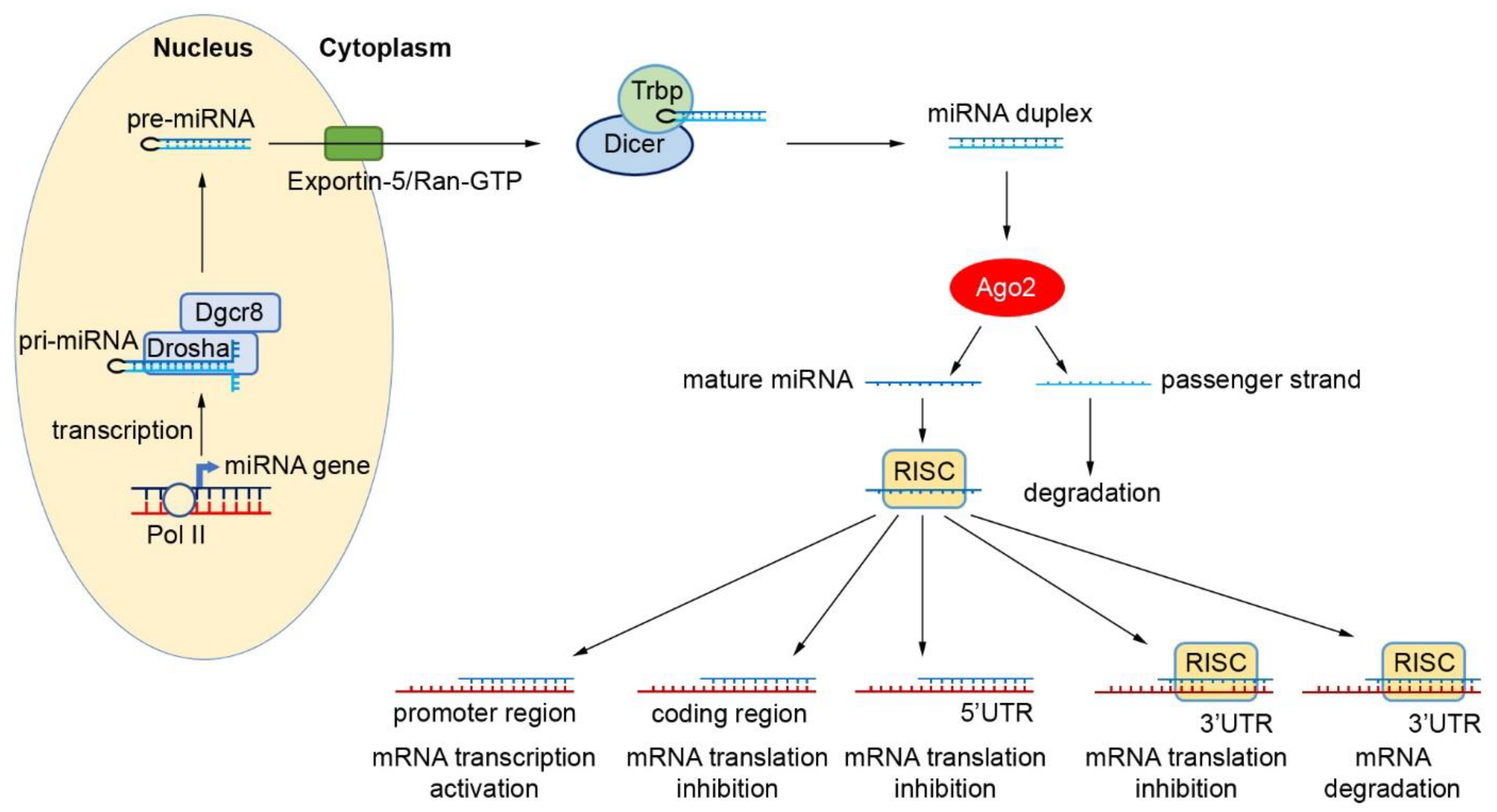
2. Biogenesis and Function of miRNAs
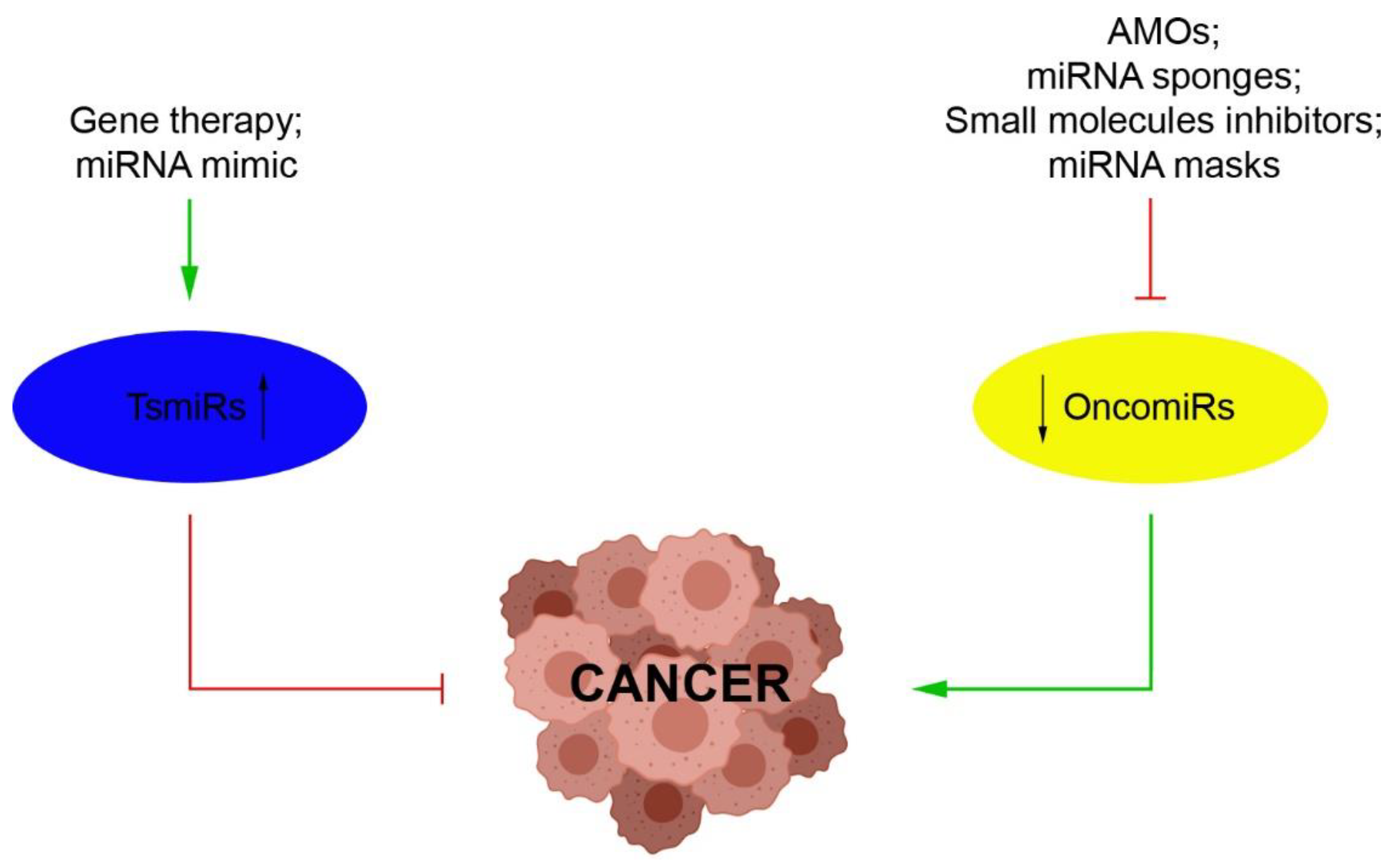
3. miRNAs as Novel Therapeutic Strategies
4. Dysregulation of miRNAs in Rare Gynecological Cancers
4.1. Tubo-Ovarian Cancer
4.1.1. miRNAs in the Rare Types of Epithelial Ovarian Cancers
4.1.2. miRNAs in Nonepithelial Ovarian Cancers
4.2. Uterine Cancer
4.2.1. miRNAs in Uterine Sarcomas
4.2.2. miRNAs in Uterine Carcinosarcomas
4.3. miRNAs in Vulvar Tumors
4.4. miRNAs in Melanoma of the Female Genital Tract
4.5. miRNAs in Gestational Trophoblastic Disease
5. Circulating miRNAs as Potential Biomarkers
6. A Brief Overview on miRNAs and Their Regulated Targets in RGCs
6.1. Up-/Down-Regulated miRNAs and Their Roles in RGCs
6.1.1. miR-9 Induces EMT by CDH1 Targeting
6.1.2. miR-10a Promotes Tumorigenesis by Regulating PTEN, Akt and Wnt Pathways
6.1.3. miR-21 Targeted PDCD4 and PTEN Genes
6.1.4. miR-590-5p Promotes Cellular Malignant Behaviors via the Target Gene TGFβRII
6.1.5. miR-3147 Regulates SMAD4
6.1.6. miR-4712-5p Regulates PTEN and Affects Its Downstream p-AKT, p-GSK3β and Cyclin D1 Signaling Pathways
6.1.7. miR-126 Regulates EGFL7
6.1.8. miR-34a and miR-196b Are Tumor Suppressors in Choriocarcinoma
7. Conclusions and Future Directions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| BEP | bleomycin, etoposide, and cisplatin |
| CSC | Cancer Stem Cell |
| EMT | epithelial–mesenchymal transition |
| ESS | endometrial stromal sarcoma |
| GTD | gestational trophoblastic disease |
| GTN | gestational trophoblastic neoplasia |
| hCG | human chorionic gonadotropin |
| HGSC | high-grade serous ovarian carcinoma |
| HPV | human papillomavirus |
| LMS | leiomyosarcoma |
| mEOC | mucinous ovarian cancer |
| miRNA/miR | micro-RNA |
| OCCC | ovarian clear cell cancer |
| PTEN | phosphatase and tensin homolog deleted on chromosome ten |
| PTN | persistent trophoblastic neoplasia |
| RGC | rare gynecological cancer |
| RISC | RNA-induced silencing complex |
| TIP | paclitaxel, ifosfamide, and cisplatin |
| TKIs | tyrosine kinase inhibitors |
| TRBP | transactivation response RNA binding protein |
| TsmiRs | tumor suppressors miRs |
| UCS | uterine carcinosarcoma |
| UPS | undifferentiated pleomorphic sarcoma |
| VSCC | vulvar squamous cell carcinoma |
| 3′-UTR | 3′ untranslated region |
References
- Srivastava, S.K.; Ahmad, A.; Zubair, H.; Miree, O.; Singh, S.; Rocconi, R.P.; Scalici, J.; Singh, A.P. MicroRNAs in gynecological cancers: Small molecules with big implications. Cancer Lett. 2017, 407, 123–138. [Google Scholar] [CrossRef]
- Maheshwari, A.; Kumar, N.; Mahantshetty, U. Gynecological cancers: A summary of published Indian data. South Asian J. Cancer 2016, 5, 112–120. [Google Scholar] [CrossRef]
- Esserman, L.J.; Thompson, I.M.; Reid, B.; Nelson, P.; Ransohoff, D.F.; Welch, H.G.; Hwang, S.; Berry, D.A.; Kinzler, K.W.; Black, W.C.; et al. Addressing overdiagnosis and overtreatment in cancer: A prescription for change. Lancet Oncol. 2014, 15, e234–e242. [Google Scholar] [CrossRef]
- Mandilaras, V.; Karakasis, K.; Clarke, B.; Amit Oza, A.; Lheureux, S. Rare tumors in gynaecological cancers and the lack of therapeutic options and clinical trials. Expert Opin. Orphan Drugs 2017, 5, 71–83. [Google Scholar] [CrossRef]
- Di Fiore, R.; Suleiman, S.; Ellul, B.; O’Toole, S.A.; Savona-Ventura, C.; Felix, A.; Napolioni, V.; Conlon, N.T.; Kahramanoglu, I.; Azzopardi, M.J.; et al. GYNOCARE Update: Modern Strategies to Improve Diagnosis and Treatment of Rare Gynecologic Tumors-Current Challenges and Future Directions. Cancers 2021, 13, 493. [Google Scholar] [CrossRef] [PubMed]
- Ray-Coquard, I.; Trama, A.; Seckl, M.J.; Fotopoulou, C.; Pautier, P.; Pignata, S.; Kristensen, G.; Mangili, G.; Falconer, H.; Massuger, L.; et al. Rare ovarian tumours: Epidemiology, treatment challenges in and outside a network setting. Eur. J. Surg. Oncol. 2017, 45, 67–74. [Google Scholar] [CrossRef] [PubMed]
- Cancer Research UK. Available online: https://www.cancerresearchuk.org (accessed on 20 February 2021).
- Rare Cancers Europe. Available online: https://www.rarecancerseurope.org (accessed on 17 February 2021).
- Jonas, S.; Izaurralde, E. Towards a molecular understanding of microRNA mediated gene silencing. Nat. Rev. Genet. 2015, 16, 421–433. [Google Scholar] [CrossRef] [PubMed]
- O’Brien, J.; Hayder, H.; Zayed, Y.; Peng, C. Overview of MicroRNA Biogenesis, Mechanisms of Actions, and Circulation. Front. Endocrinol. 2018, 9, 402. [Google Scholar] [CrossRef]
- Kozomara, A.; Birgaoanu, M.; Griffiths-Jones, S. miRBase: From microRNA sequences to function. Nucleic Acids Res. 2019, 47, D155–D162. [Google Scholar] [CrossRef]
- Zhong, X.; Coukos, G.; Zhang, L. miRNAs in human cancer. Methods Mol. Biol. 2012, 822, 295–306. [Google Scholar]
- Rajagopal, T.; Talluri, S.; Venkatabalasubramanian, S.; Dunna, N.R. Multifaceted roles of long non-coding RNAs in triple-negative breast cancer: Biology and clinical applications. Biochem. Soc. Trans. 2020, 48, 2791–2810. [Google Scholar] [CrossRef] [PubMed]
- Tabatabaeian, H.; Peiling Yang, S.; Tay, Y. Non-Coding RNAs: Uncharted Mediators of Thyroid Cancer Pathogenesis. Cancers 2020, 12, 3264. [Google Scholar] [CrossRef]
- Lu, X.; Zhang, Y.; Xie, G.; Ding, Y.; Cong, H.; Xuan, S. Exosomal non coding RNAs: Novel biomarkers with emerging clinical applications in gastric cancer (Review). Mol. Med. Rep. 2020, 22, 4091–4100. [Google Scholar] [CrossRef]
- Iorio, M.V.; Visone, R.; Di Leva, G.; Donati, V.; Petrocca, F.; Casalini, P.; Taccioli, C.; Volinia, S.; Liu, C.G.; Alder, H.; et al. MicroRNA signatures in human ovarian cancer. Cancer Res. 2007, 67, 8699–8707. [Google Scholar] [CrossRef] [PubMed]
- Miyoshi, K.; Miyoshi, T.; Siomi, H. Many ways to generate microRNA-like small RNAs: Non-canonical pathways for microRNA production. Mol. Genet. Genom. 2010, 284, 95–103. [Google Scholar] [CrossRef] [PubMed]
- Kim, V.N. MicroRNA biogenesis: Coordinated cropping and dicing. Nat. Rev. Mol. Cell Biol. 2005, 6, 376–385. [Google Scholar] [CrossRef]
- Bartel, D.P. MicroRNAs: Genomics, biogenesis, mechanisms and function. Cell 2004, 116, 281–297. [Google Scholar] [CrossRef]
- Abdelfattah, A.M.; Park, C.; Choi, M.Y. Update on non-canonical microRNAs. Biomol. Concepts 2014, 5, 275–287. [Google Scholar] [CrossRef] [PubMed]
- Han, J.; Lee, Y.; Yeom, K.H.; Nam, J.W.; Heo, I.; Rhee, J.K.; Sohn, S.Y.; Cho, Y.; Zhang, B.T.; Kim, V.N. Molecular Basis for the Recognition of Primary microRNAs by the Drosha-DGCR8 Complex. Cell 2006, 125, 887–901. [Google Scholar] [CrossRef]
- Valinezhad Orang, A.; Safaralizadeh, R.; Kazemzadeh-Bavili, M. Mechanisms of miRNA-Mediated Gene Regulation from Common Downregulation to mRNA-Specific Upregulation. Int. J. Genom. 2014, 2014, 970607. [Google Scholar] [CrossRef]
- Yuan, X.; Berg, N.; Lee, J.W.; Le, T.T.; Neudecker, V.; Jing, N.; Eltzschig, H. MicroRNA miR-223 as regulator of innate immunity. J. Leukoc. Biol. 2018, 104, 515–524. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Fan, M.; Zhang, X.; Huang, F.; Wu, K.; Zhang, J.; Liu, J.; Huang, Z.; Luo, H.; Tao, L.; et al. Cellular microRNAs up-regulate transcription via interaction with promoter TATA-box motifs. RNA 2014, 20, 1878–1889. [Google Scholar] [CrossRef]
- Daugaard, I.; Hansen, T.B. Biogenesis and Function of Ago- Associated RNAs. Trends Genet. 2017, 33, 208–219. [Google Scholar] [CrossRef] [PubMed]
- Calin, G.A.; Dumitru, C.D.; Shimizu, M.; Bichi, R.; Zupo, S.; Noch, E.; Aldler, H.; Rattan, S.; Keating, M.; Rai, K.; et al. Frequent deletions and down-regulation of micro- RNA genes miR15 and miR16 at 13q14 in chronic lymphocytic leukemia. Proc. Natl. Acad. Sci. USA 2002, 99, 15524–15529. [Google Scholar] [CrossRef]
- Acunzo, M.; Romano, G.; Wernicke, D.; Croce, C.M. MicroRNA and cancer--a brief overview. Adv. Biol. Regul. 2015, 57, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Lu, J.; Getz, G.; Miska, E.A.; Alvarez-Saavedra, E.; Lamb, J.; Peck, D.; Sweet-Cordero, A.; Ebert, B.L.; Mak, R.H.; Ferrando, A.A.; et al. MicroRNA expression profiles classify human cancers. Nature 2005, 435, 834–838. [Google Scholar] [CrossRef] [PubMed]
- Shimono, Y.; Zabala, M.; Cho, R.W.; Lobo, N.; Dalerba, P.; Qian, D.; Diehn, M.; Liu, H.; Panula, S.P.; Chiao, E.; et al. Downregulation of miRNA-200c links breast cancer stem cells with normal stem cells. Cell 2009, 138, 592–603. [Google Scholar] [CrossRef]
- Calin, G.A.; Croce, C.M. MicroRNA signatures in human cancers. Nat. Rev. Cancer 2006, 6, 857–866. [Google Scholar] [CrossRef]
- Zhang, B.; Pan, X.; Cobb, G.P.; Anderson, T.A. microRNAs as oncogenes and tumor suppressors. Dev. Biol. 2007, 302, 1–12. [Google Scholar] [CrossRef]
- Bhardwaj, A.; Singh, S.; Singh, A.P. MicroRNA-based Cancer Therapeutics: Big Hope from Small RNAs. Mol. Cell Pharmacol. 2010, 2, 213–219. [Google Scholar]
- Tsuda, N.; Ishiyama, S.; Li, Y.; Ioannides, C.G.; Abbruzzese, J.L.; Chang, D.Z. Synthetic microRNA designed to target glioma-associated antigen 1 transcription factor inhibits division and induces late apoptosis in pancreatic tumor cells. Clin. Cancer Res. 2006, 12, 6557–6564. [Google Scholar] [CrossRef] [PubMed]
- Lima, J.F.; Cerqueira, L.; Figueiredo, C.; Oliveira, C.; Azevedo, N.F. Anti-miRNA oligonucleotides: A comprehensive guide for design. RNA Biol. 2018, 15, 338–352. [Google Scholar] [CrossRef] [PubMed]
- Wen, D.; Danquah, M.; Chaudhary, A.K.; Mahato, R.I. Small molecules targeting microRNA for cancer therapy: Promises and obstacles. J. Control. Release 2015, 219, 237–247. [Google Scholar] [CrossRef]
- Gambari, R.; Brognara, E.; Spandidos, D.A.; Fabbri, E. Targeting oncomiRNAs and mimicking tumor suppressor miRNAs: Νew trends in the development of miRNA therapeutic strategies in oncology (Review). Int. J. Oncol. 2016, 49, 5–32. [Google Scholar] [CrossRef]
- Mognato, M.; Celotti, L. MicroRNAs Used in Combination with Anti-Cancer Treatments Can Enhance Therapy Efficacy. Mini Rev. Med. Chem. 2015, 15, 1052–1062. [Google Scholar] [CrossRef] [PubMed]
- Segal, M.; Slack, F.J. Challenges identifying efficacious miRNA therapeutics for cancer. Expert Opin. Drug Discov. 2020, 15, 987–992. [Google Scholar] [CrossRef] [PubMed]
- Yanai, H. Pathology of Epithelial Ovarian Tumors. In Frontiers in Ovarian Cancer Science; Katabuchi, H., Ed.; Springer: Singapore, 2017; pp. 83–113. [Google Scholar]
- Fukunaga, M. Pathology of Non-epithelial Ovarian Tumors. In Frontiers in Ovarian Cancer Science; Katabuchi, H., Ed.; Springer: Singapore, 2017; pp. 115–141. [Google Scholar]
- Kurman, R.J.; Shih, I. Molecular pathogenesis and extraovarian origin of epithelial ovarian cancer–shifting the paradigm. Hum. Pathol. 2011, 42, 918–931. [Google Scholar] [CrossRef] [PubMed]
- Fung-Kee-Fung, M.; Oliver, T.; Elit, L.; Oza, A.; Hirte, H.W.; Bryson, P. Optimal chemotherapy treatment for women with recurrent ovarian cancer. Curr. Oncol. 2007, 14, 195–208. [Google Scholar] [CrossRef] [PubMed]
- Zorn, K.K.; Bonome, T.; Gangi, L.; Chandramouli, G.V.; Awtrey, C.S.; Gardner, G.J.; Barrett, J.C.; Boyd, J.; Birrer, M.J. Gene expression profiles of serous, endometrioid, and clear cell subtypes of ovarian and endometrial cancer. Clin. Cancer Res. 2005, 11, 6422–6430. [Google Scholar] [CrossRef]
- Yoshida, S.; Furukawa, N.; Haruta, S.; Tanase, Y.; Kanayama, S.; Noguchi, T.; Sakata, M.; Yamada, Y.; Oi, H.; Kobayashi, H. Theoretical model of treatment strategies for clear cell carcinoma of the ovary: Focus on perspectives. Cancer Treat. Rev. 2009, 35, 608–615. [Google Scholar] [CrossRef]
- Shibuya, Y.; Tokunaga, H.; Saito, S.; Shimokawa, K.; Katsuoka, F.; Bin, L.; Kojima, K.; Nagasaki, M.; Yamamoto, M.; Yaegashi, N.; et al. Identification of somatic genetic alterations in ovarian clear cell carcinoma with next generation sequencing. Genes Chromosomes Cancer 2018, 57, 51–60. [Google Scholar] [CrossRef]
- Bitler, B.G.; Aird, K.M.; Garipov, A.; Li, H.; Amatangelo, M.; Kossenkov, A.V.; Schultz, D.C.; Liu, Q.; Shih, I.; Conejo-Garcia, J.R.; et al. Synthetic lethality by targeting EZH2 methyltransferase activity in ARID1A-mutated cancers. Nat. Med. 2015, 21, 231–238. [Google Scholar] [CrossRef] [PubMed]
- Miller, R.E.; Brough, R.; Bajrami, I.; Williamson, C.T.; McDade, S.; Campbell, J.; Kigozi, A.; Rafiq, R.; Pemberton, H.; Natrajan, R.; et al. Synthetic lethal targeting of ARID1A-mutant ovarian clear cell tumors with dasatinib. Mol. Cancer Ther. 2016, 15, 1472–8144. [Google Scholar] [CrossRef]
- Bitler, B.G.; Wu, S.; Park, P.H.; Hai, Y.; Aird, K.M.; Wang, Y.; Zhai, Y.; Kossenkov, A.V.; Vara-Ailor, A.; Rauscher, F.J., III; et al. ARID1A-mutated ovarian cancers depend on HDAC6 activity. Nat. Cell Biol. 2017, 19, 962–973. [Google Scholar] [CrossRef]
- Berns, K.; Caumanns, J.J.; Hijmans, E.M.; Gennissen, A.M.C.; Severson, T.M.; Evers, B.; Wisman, G.B.A.; Jan Meersma, G.; Lieftink, C.; Beijersbergen, R.L.; et al. ARID1A mutation sensitizes most ovarian clear cell carcinomas to BET inhibitors. Oncogene 2018, 37, 4611–4625. [Google Scholar] [CrossRef] [PubMed]
- Fujimura, M.; Hidaka, T.; Saito, S. Selective inhibition of the epidermal growth factor receptor by ZD1839 decreases the growth and invasion of ovarian clear cell adenocarcinoma cells. Clin. Cancer Res. 2002, 8, 2448–2454. [Google Scholar] [PubMed]
- Mabuchi, S.; Kawase, C.; Altomare, D.A.; Morishige, K.; Sawada, K.; Hayashi, M.; Tsujimoto, M.; Yamoto, M.; Klein-Szanto, A.J.; Schilder, R.J.; et al. mTOR is a promising therapeutic target both in cisplatin-sensitive and cisplatin-resistant clear cell carcinoma of the ovary. Clin. Cancer Res. 2009, 15, 5404–5413. [Google Scholar] [CrossRef]
- Anglesio, M.S.; Kommoss, S.; Tolcher, M.C.; Clarke, B.; Galletta, L.; Porter, H.; Damaraju, S.; Fereday, S.; Winterhoff, B.J.; Kalloger, S.E.; et al. Molecular characterization of mucinous ovarian tumours supports a stratified treatment approach with HER2 targeting in 19% of carcinomas. J. Pathol. 2013, 229, 111–120. [Google Scholar] [CrossRef] [PubMed]
- Vilming Elgaaen, B.; Olstad, O.K.; Haug, K.B.F.; Brusletto, B.; Sandvik, L.; Staff, A.C.; Gautvik, K.M.; Davidson, B. Global miRNA expression analysis of serous and clear cell ovarian carcinomas identifies differentially expressed miRNAs including miR-200c-3p as a prognostic marker. BMC Cancer 2014, 14, 80. [Google Scholar] [CrossRef]
- Wu, X.; Ruan, Y.; Jiang, H.; Xu, C. MicroRNA-424 inhibits cell migration, invasion, and epithelial mesenchymal transition by downregulating doublecortin-like kinase 1 in ovarian clear cell carcinoma. Int. J. Biochem. Cell Biol. 2017, 85, 66–74. [Google Scholar] [CrossRef]
- Yanaihara, N.; Noguchi, Y.; Saito, M.; Takenaka, M.; Takakura, S.; Yamada, K.; Okamoto, A. MicroRNA Gene Expression Signature Driven by miR-9 Overexpression in Ovarian Clear Cell Carcinoma. PLoS ONE 2016, 11, e0162584. [Google Scholar] [CrossRef] [PubMed]
- Jang, S.G.; Yoo, C.W.; Park, S.Y.; Kang, S.; Kim, H.K. Low expression of miR-449 in gynecologic clear cell carcinoma. Int. J. Gynecol. Cancer 2014, 24, 1558–1563. [Google Scholar] [CrossRef] [PubMed]
- Dai, F.; Zhang, Y.; Chen, Y. Involvement of miR-29b signaling in the sensitivity to chemotherapy in patients with ovarian carcinoma. Hum. Pathol. 2014, 45, 1285–1293. [Google Scholar] [CrossRef] [PubMed]
- Agostini, A.; Brunetti, M.; Davidson, B.; Tropé, C.G.; Eriksson, A.G.Z.; Heim, S.; Panagopoulos, I.; Micci, F. The microRNA miR-192/215 family is upregulated in mucinous ovarian carcinomas. Sci. Rep. 2018, 23, 11069. [Google Scholar] [CrossRef] [PubMed]
- Bower, M.; Fife, K.; Holden, L.; Paradinas, F.J.; Rustin, G.J.; Newlands, E.S. Chemotherapy for ovarian germ cell tumours. Eur. J. Cancer 1996, 32A, 593–597. [Google Scholar] [CrossRef]
- Simone, C.G.; Markham, M.J.; Dizon, D.S. Chemotherapy in ovarian germ cell tumors: A systematic review. Gynecol. Oncol. 2016, 141, 602–607. [Google Scholar] [CrossRef]
- Goulvent, T.; Ray-Coquard, I.; Borel, S.; Haddad, V.; Devouassoux-Shisheboran, M.; Vacher-Lavenu, M.C.; Pujade-Laurraine, E.; Savina, A.; Maillet, D.; Gillet, G.; et al. DICER1 and FOXL2 mutations in ovarian sex cord-stromal tumours: A GINECO Group study. Histopathology 2016, 68, 279–285. [Google Scholar] [CrossRef]
- Ray-Coquard, I.; Weber, B.; Lotz, J.P.; Tournigand, C.; Provencal, J.; Mayeur, D.; Treilleux, I.; Paraiso, D.; Duvillard, P.; Pujade-Lauraine, E.; et al. Management of rare ovarian cancers: The experience of the French website “Observatory for rare malignant tumours of the ovaries” by the GINECO group: Interim analysis of the first 100 patients. Gynecol. Oncol. 2010, 119, 53–59. [Google Scholar] [CrossRef]
- WHO Classification of Tumors Editoral Board. Female Genital Tumors. In WHO Classification of Tumors Series, 5th ed.; IARC: Lyon, France, 2020; Volume 4, Available online: https://publications.iarc.fr/592 (accessed on 20 December 2020).
- Manchana, T.; Ittiwut, C.; Mutirangura, A.; Kavanagh, J.J. Targeted therapies for rare gynaecological cancers. Lancet Oncol. 2010, 11, 685–693. [Google Scholar] [CrossRef]
- Kommoss, S.; Anglesio, M.S.; Mackenzie, R.; Yang, W.; Senz, J.; Ho, J.; Bell, L.; Lee, S.; Lorette, J.; Huntsman, D.G.; et al. FOXL2 molecular testing in ovarian neoplasms: Diagnostic approach and procedural guidelines. Modern Pathol. 2013, 26, 860–867. [Google Scholar] [CrossRef]
- Jamieson, S.; Fuller, P.J. Molecular pathogenesis of granulosa cell tumors of the ovary. Endocr. Rev. 2012, 33, 109–144. [Google Scholar] [CrossRef]
- Weis-Banke, S.E.; Lerdrup, M.; Kleine-Kohlbrecher, D.; Mohammad, F.; Sidoli, S.; Jensen, O.N.; Yanase, T.; Nakamura, T.; Iwase, A.; Stylianou, A.; et al. Mutant FOXL2C134W Hijacks SMAD4 and SMAD2/3 to Drive Adult Granulosa Cell Tumors. Cancer Res. 2020, 80, 3466–3479. [Google Scholar] [CrossRef]
- Chang, R.K.; Li, X.; Mu, N.; Hrydziuszko, O.; Garcia-Majano, B.; Larsson, C.; Lui, W.O. MicroRNA expression profiles in non-epithelial ovarian tumors. Int. J. Oncol. 2018, 52, 55–66. [Google Scholar] [CrossRef] [PubMed]
- Poynter, J.N.; Bestrashniy, J.R.; Silverstein, K.A.; Hooten, A.J.; Lees, C.; Ross, J.A.; Tolar, J. Cross platform analysis of methylation, miRNA and stem cell gene expression data in germ cell tumors highlights characteristic differences by tumor histology. BMC Cancer 2015, 15, 769. [Google Scholar] [CrossRef]
- Cheng, W.T.; Rosario, R.; Muthukaruppan, A.; Wilson, M.K.; Payne, K.; Fong, P.C.; Shelling, A.N.; Blenkiron, C. MicroRNA profiling of ovarian granulosa cell tumours reveals novel diagnostic and prognostic markers. Clin. Epigenetics 2017, 9, 72. [Google Scholar] [CrossRef] [PubMed]
- Rosario, R.; Blenkiron, C.; Shelling, A.N. Comparative study of microRNA regulation on FOXL2 between adult-type and juvenile-type granulosa cell tumours in vitro. Gynecol. Oncol. 2013, 129, 209–215. [Google Scholar] [CrossRef] [PubMed]
- Tu, J.; Cheung, H.H.; Lu, G.; Chan, C.L.K.; Chen, Z.; Chan, W.Y. microRNA-126 Is a Tumor Suppressor of Granulosa Cell Tumor Mediated by Its Host Gene EGFL7. Front. Oncol. 2019, 9, 486. [Google Scholar] [CrossRef]
- Tu, J.; Cheung, H.H.; Lu, G.; Chen, Z.; Chan, W.Y. MicroRNA-10a promotes granulosa cells tumor development via PTEN-AKT/Wnt regulatory axis. Cell Death Dis. 2018, 9, 1076. [Google Scholar] [CrossRef] [PubMed]
- De Kock, L.; Terzic, T.; McCluggage, W.G.; Stewart, C.J.R.; Shaw, P.; Foulkes, W.D.; Clarke, B.A. DICER1 Mutations Are Consistently Present in Moderately and Poorly Differentiated Sertoli-Leydig Cell Tumors. Am. J. Surg. Pathol. 2017, 41, 1178–1187. [Google Scholar] [CrossRef]
- Huvila, J.; Pors, J.; Thompson, E.F.; Gilks, C.B. Endometrial carcinoma: Molecular subtypes, precursors and the role of pathology in early diagnosis. J. Pathol. 2021, 253, 355–365. [Google Scholar] [CrossRef]
- Mbatani, N.; Olawaiye, A.; Prat, J. Uterine sarcomas. Int. J. Gynecol. Obstet. 2018, 143, 51–58. [Google Scholar] [CrossRef]
- Ferrandina, G.; Aristei, C.; Biondetti, P.R.; Cananzi, F.C.M.; Casali, P.; Ciccarone, F.; Colombo, N.; Comandone, A.; Corvo’, R.; De Iaco, P.; et al. Italian consensus conference on management of uterine sarcomas on behalf of S.I.G.O. (Societa’ italiana di Ginecologia E Ostetricia). Eur. J. Cancer 2020, 139, 149–168. [Google Scholar] [CrossRef] [PubMed]
- Kurman, R.J.; Carcangiu, M.L.; Herrington, C.S.; Young, R.H. WHO Classification of Tumors of Female Reproductive Organs; IARC: Lyon, France, 2014. [Google Scholar]
- Micci, F.; Heim, S.; Panagopoulos, I. Molecular pathogenesis and prognostication of “low-grade’’ and “high-grade” endometrial stromal sarcoma. Genes Chromosomes Cancer 2020, 60, 160–167. [Google Scholar] [CrossRef] [PubMed]
- Spano, J.P.; Soria, J.C.; Kambouchner, M.; Piperno-Neuman, S.; Morin, F.; Morere, J.F.; Martin, A.; Breau, J.L. Long-term survival of patients given hormonal therapy for metastatic endometrial stromal sarcoma. Med. Oncol. 2003, 20, 87–93. [Google Scholar] [CrossRef]
- Tanner, E.J.; Garg, K.; Leitao, M.M., Jr.; Soslow, R.A.; Hensley, M.L. High grade undifferentiated uterine sarcoma: Surgery, treatment, and survival outcomes. Gynecol. Oncol. 2012, 127, 27–31. [Google Scholar] [CrossRef]
- Ray-Coquard, I.; Hatcher, H.; Bompas, E.; Casado, A.; Westermann, A.; Isambert, N.; Casali, P.G.; Pratap, S.; Stark, D.; Valverde, C.; et al. A randomized double-blind phase II study evaluating the role of maintenance therapy with cabozantinib in high-grade uterine sarcoma after stabilization or response to doxorubicin ± ifosfamide following surgery or in metastatic first line treatment (EORTC62113). Int. J. Gynecol. Cancer 2020, 30, 1633–1637. [Google Scholar]
- Hensley, M.L.; Barrette, B.A.; Baumann, K.; Gaffney, D.; Hamilton, A.L.; Kim, J.W.; Maenpaa, J.U.; Pautier, P.; Siddiqui, N.A.; Westermann, A.M.; et al. Gynecologic Cancer InterGroup (GCIG) consensus review: Uterine and ovarian leiomyosarcomas. Int. J. Gynecol. Cancer 2014, 24, S61–S66. [Google Scholar] [CrossRef]
- Van der Graaf, W.T.; Blay, J.Y.; Chawla, S.P.; Kim, D.W.; Bui-Nguyen, B.; Casali, P.G.; Schöffski, P.; Aglietta, M.; Staddon, A.P.; Beppu, Y.; et al. Pazopanib for metastatic soft-tissue sarcoma (PALETTE): A randomised, double-blind, placebo-controlled phase 3 trial. Lancet 2012, 379, 1879–1886. [Google Scholar] [CrossRef]
- Kobayashi, H.; Uekuri, C.; Akasaka, J.; Ito, F.; Shigemitsu, A.; Koike, N.; Shigetomi, H. The biology of uterine sarcomas: A review and update. Mol. Clin. Oncol. 2013, 1, 599–609. [Google Scholar] [CrossRef] [PubMed]
- Gonzalez Dos Anjos, L.; de Almeida, B.C.; Gomes de Almeida, T.; Mourão Lavorato Rocha, A.; De Nardo Maffazioli, G.; Soares, F.A.; Werneck da Cunha, I.; Baracat, E.C.; Carvalho, K.C. Could miRNA Signatures be Useful for Predicting Uterine Sarcoma and Carcinosarcoma Prognosis and Treatment? Cancers 2018, 10, 315. [Google Scholar] [CrossRef] [PubMed]
- Kowalewska, M.; Bakula-Zalewska, E.; Chechlinska, M.; Goryca, K.; Nasierowska-Guttmejer, A.; Danska-Bidzinska, A.; Bidzinski, M. microRNAs in uterine sarcomas and mixed epithelial-mesenchymal uterine tumors: A preliminary report. Tumour Biol. 2013, 34, 2153–2160. [Google Scholar] [CrossRef]
- Ravid, Y.; Formanski, M.; Smith, Y.; Reich, R.; Davidson, B. Uterine leiomyosarcoma and endometrial stromal sarcoma have unique miRNA signatures. Gynecol. Oncol. 2016, 140, 512–517. [Google Scholar] [CrossRef] [PubMed]
- De Almeida, B.C.; Dos Anjos, L.G.; Uno, M.; Cunha, I.W.D.; Soares, F.A.; Baiocchi, G.; Baracat, E.C.; Carvalho, K.C. Let-7 miRNA’s Expression Profile and Its Potential Prognostic Role in Uterine Leiomyosarcoma. Cells 2019, 8, 1452. [Google Scholar] [CrossRef]
- De Almeida, B.C.; Garcia, N.; Maffazioli, G.; dos Anjos, L.G.; Baracat, E.C.; Carvalho, K.C. Oncomirs Expression Profiling in Uterine Leiomyosarcoma Cells. Int. J. Mol. Sci. 2017, 19, 52. [Google Scholar] [CrossRef] [PubMed]
- Guled, M.; Pazzaglia, L.; Borze, I.; Mosakhani, N.; Novello, C.; Benassi, M.S.; Knuutila, S. Differentiating soft tissue leiomyosarcoma and undifferentiated pleomorphic sarcoma: A miRNA analysis. Genes Chromosomes Cancer 2014, 53, 693–702. [Google Scholar] [CrossRef] [PubMed]
- Stope, M.B.; Cernat, V.; Kaul, A.; Diesing, K.; Koensgen, D.; Burchardt, M.; Mustea, A. Functionality of the Tumor Suppressor microRNA-1 in Malignant Tissue and Cell Line Cells of Uterine Leiomyosarcoma. Anticancer Res. 2018, 38, 1547–1550. [Google Scholar] [PubMed]
- Berton-Rigaud, D.; Devouassoux-Shisheboran, M.; Ledermann, J.A.; Leitao, M.M.; Powell, M.A.; Poveda, A.; Beale, P.; Glasspool, R.M.; Creutzberg, C.L.; Harter, P.; et al. Gynecologic Cancer InterGroup (GCIG) consensus review for uterine and ovarian carcinosarcoma. Int. J. Gynecol. Cancer 2014, 24, S55–S60. [Google Scholar] [CrossRef] [PubMed]
- Cantrell, L.A.; Blank, S.V.; Duska, L.R. Uterine carcinosarcoma: A review of the literature. Gynecol. Oncol. 2015, 137, 581–588. [Google Scholar] [CrossRef]
- Pautier, P.; Floquet, A.; Gladieff, L.; Bompas, E.; Ray-Coquard, I.; Piperno-Neumann, S.; Selle, F.; Guillemet, C.; Weber, B.; Largillier, R.; et al. A randomized clinical trial of adjuvant chemotherapy with doxorubicin, ifosfamide, and cisplatin followed by radiotherapy versus radiotherapy alone in patients with localized uterine sarcomas (SARCGYN study). A study of the French Sarcoma group. Ann. Oncol. 2013, 24, 1099–1104. [Google Scholar] [CrossRef]
- Reed, N.S.; Mangioni, C.; Malmström, H.; Scarfone, G.; Poveda, A.; Pecorelli, S.; Tateo, S.; Franchi, M.; Jobsen, J.J.; Coens, C.; et al. Phase III randomised study to evaluate the role of adjuvant pelvic radiotherapy in the treatment of uterine sarcomas stages I and II: An European Organisation for Research and Treatment of Cancer Gynaecological Cancer Group Study (protocol 55874). Eur. J. Cancer 2008, 44, 808–818. [Google Scholar] [CrossRef] [PubMed]
- Omura, G.A.; Major, F.J.; Blessing, J.A.; Sedlacek, T.V.; Thigpen, J.T.; Creasman, W.T.; Zaino, R.J. A randomized study of adriamycin with and without dimethyl triazenoimidazole carboxamide in advanced uterine sarcomas. Cancer 1983, 52, 626–632. [Google Scholar] [CrossRef]
- Brunetti, M.; Agostini, A.; Staurseth, J.; Davidson, B.; Heim, S.; Micci, F. Molecular characterization of carcinosarcomas arising in the uterus and ovaries. Oncotarget 2019, 10, 3614–3624. [Google Scholar] [CrossRef]
- McConechy, M.K.; Hoang, L.N.; Chui, M.H.; Senz, J.; Yang, W.; Rozenberg, N.; Mackenzie, R.; McAlpine, J.N.; Huntsman, D.G.; Clarke, B.A.; et al. In-depth molecular profiling of the biphasic components of uterine carcinosarcomas. J. Pathol. Clin. Res. 2015, 1, 173–185. [Google Scholar] [CrossRef] [PubMed]
- Cherniack, A.D.; Shen, H.; Walter, V.; Stewart, C.; Murray, B.A.; Bowlby, R.; Hu, X.; Ling, S.; Soslow, R.A.; Broaddus, R.R.; et al. Integrated Molecular Characterization of Uterine Carcinosarcoma. Cancer Cell 2017, 31, 411–423. [Google Scholar] [CrossRef] [PubMed]
- Tseng, J.H.; Bisogna, M.; Hoang, L.N.; Olvera, N.; Rodriguez-Aguayo, C.; Lopez-Berestein, G.; Sood, A.K.; Levine, D.A.; Jelinic, P. miR-200c-driven Mesenchymal-To-Epithelial Transition is a Therapeutic Target in Uterine Carcinosarcomas. Sci. Rep. 2017, 7, 3614. [Google Scholar] [CrossRef] [PubMed]
- Miccò, M.; Sala, E.; Lakhman, Y.; Hricak, H.; Vargas, H.A. Imaging Features of Uncommon Gynecologic Cancers. AJR Am. J. Roentgenol. 2015, 205, 1346–1359. [Google Scholar] [CrossRef][Green Version]
- Olawaiye, A.; Lee, L.M.; Krasner, C.; Horowitz, N. Treatment of squamous cell vulvar cancer with the anti-EGFR tyrosine kinase inhibitor Tarceva. Gynecol. Oncol. 2007, 106, 628–630. [Google Scholar] [CrossRef]
- Richard, S.D.; Krivak, T.C.; Beriwal, S.; Zorn, K.K. Recurrent metastatic vulvar carcinoma treated with cisplatin plus cetuximab. Int. J. Gynecol. Cancer 2008, 18, 1132–1135. [Google Scholar] [CrossRef]
- Plaza, J.A.; Torres-Cabala, C.; Ivan, D.; Prieto, V.G. HER-2/neu expression in extramammary Paget disease: A clinicopathologic and immunohistochemistry study of 47 cases with and without underlying malignancy. J. Cutan. Pathol. 2009, 36, 729–733. [Google Scholar] [CrossRef]
- Woelber, L.; Prieske, K.; Eulenburg, C.; Oliveira-Ferrer, L.; de Gregorio, N.; Klapdor, R.; Kalder, M.; Braicu, I.; Fuerst, S.; Klar, M.; et al. p53 and p16 expression profiles in vulvar cancer: A translational analysis by the Arbeitsgemeinschaft Gynäkologische Onkologie Chemo and Radiotherapy in Epithelial Vulvar Cancer study group. Am. J. Obstet. Gynecol. 2021. [Google Scholar] [CrossRef]
- De Melo Maia, B.; Lavorato-Rocha, A.M.; Rodrigues, L.S.; Coutinho-Camillo, C.M.; Baiocchi, G.; Stiepcich, M.M.; Puga, R.; de A. Lima, L.; Soares, F.A.; Rocha, R.M. microRNA portraits in human vulvar carcinoma. Cancer Prev. Res. 2013, 6, 1231–1241. [Google Scholar] [CrossRef]
- Yang, X.; Wu, X. miRNA expression profile of vulvar squamous cell carcinoma and identification of the oncogenic role of miR-590-5p. Oncol. Rep. 2016, 35, 398–408. [Google Scholar] [CrossRef]
- Yang, S.; Zhao, Y.; Wang, L.; Liu, C.; Lu, Y.; Fang, Z.; Shi, H.; Zhang, W.; Wu, X. MicroRNA-4712-5p promotes proliferation of the vulvar squamous cell carcinoma cell line A431 by targeting PTEN through the AKT/cyclin D1 signaling pathways. Oncol. Rep. 2019, 42, 1689–1698. [Google Scholar] [CrossRef]
- Yang, X.H.; Guo, F. miR-3147 serves as an oncomiR in vulvar squamous cell cancer via Smad4 suppression. Mol. Med. Rep. 2018, 17, 6397–6404. [Google Scholar] [CrossRef] [PubMed]
- Yuan, G.; Wu, L.; Li, B.; An, J. Primary malignant melanoma of the cervix: Report of 14 cases and review of literature. Oncotarget 2017, 8, 73162–73167. [Google Scholar] [CrossRef] [PubMed]
- Min, K.J.; Kim, Y.S.; Hong, J.H.; Lee, J.K.; Yang, D.S. Primary malignant melanoma of uterine cervix: A suggestion of new scheme of treatment combination. Chin. J. Cancer Res. 2014, 26, 351–354. [Google Scholar] [PubMed]
- DiVincenzo, M.; Suarez-Kelly, L.; Moufawad, M.; Ren, C.; Barricklow, Z.; Fadda, P.; Yu, L.; Peters, S.; Gru, A.; Carson, W. 736 MicroRNA expression patterns in melanomas originating from gynecologic sites. J. Immunother. Cancer 2020, 8. [Google Scholar] [CrossRef]
- Vergani, E.; Dugo, M.; Cossa, M.; Frigerio, S.; Di Guardo, L.; Gallino, G.; Mattavelli, I.; Vergani, B.; Lalli, L.; Tamborini, E.; et al. miR-146a-5p impairs melanoma resistance to kinase inhibitors by targeting COX2 and regulating NFkB-mediated inflammatory mediators. Cell Commun. Signal. 2020, 18, 156. [Google Scholar] [CrossRef] [PubMed]
- Audrito, V.; Serra, S.; Stingi, A.; Orso, F.; Gaudino, F.; Bologna, C.; Neri, F.; Garaffo, G.; Nassini, R.; Baroni, G.; et al. PD-L1 up-regulation in melanoma increases disease aggressiveness and is mediated through miR-17-5p. Oncotarget 2017, 28, 15894–15911. [Google Scholar] [CrossRef] [PubMed]
- Ngan, H.Y. The practicability of FIGO 2000 staging for gestational trophoblastic neoplasia. Int. J. Gynecol. Cancer 2004, 14, 202–205. [Google Scholar] [CrossRef]
- Steigrad, S.J. Epidemiology of gestational trophoblastic diseases. Best Pract. Res. Clin. Obstet. Gynaecol. 2003, 17, 837–847. [Google Scholar] [CrossRef]
- Dhanda, S.; Ramani, S.; Thakur, M. Gestational trophoblastic disease: A multimodality imaging approach with impact on diagnosis and management. Radiol. Res. Pract. 2014, 2014, 842751. [Google Scholar] [CrossRef] [PubMed]
- Nagymanyoki, Z.; Growdon, W.B.; Sarno, J.; Callahan, M.J.; Parast, M.M.; Fulop, V.; Mok, S.C.; Horowitz, N.; Berkowitz, R.S. Vascularization and expression of angiogenic factors in partial and complete molar pregnancies. J. Reprod. Med. 2008, 53, 589–594. [Google Scholar] [PubMed]
- Na, Q.; Wang, D.; Song, W. Underexpression of 4 placenta-associated microRNAs in complete hydatidiform moles. Int. J. Gynecol. Cancer 2012, 22, 1075–1080. [Google Scholar] [CrossRef] [PubMed]
- Zhao, J.R.; Cheng, W.W.; Wang, Y.X.; Cai, M.; Wu, W.B.; Zhang, H.J. Identification of microRNA signature in the progression of gestational trophoblastic disease. Cell Death Dis. 2018, 9, 94. [Google Scholar] [CrossRef] [PubMed]
- Lin, L.H.; Maestá, I.; St Laurent, J.D.; Hasselblatt, K.T.; Horowitz, N.S.; Goldstein, D.P.; Quade, B.J.; Sun, S.Y.; Braga, A.; Fisher, R.A.; et al. Distinct microRNA profiles for complete hydatidiform moles at risk of malignant progression. Am. J. Obstet. Gynecol. 2021, 224, 372.e1–372.e30. [Google Scholar] [CrossRef]
- Chao, A.; Tsai, C.L.; Wei, P.C.; Hsueh, S.; Chao, A.S.; Wang, C.J.; Tsai, C.N.; Lee, Y.S.; Wang, T.H.; Lai, C.H. Decreased expression of microRNA-199b increases protein levels of SET (protein phosphatase 2A inhibitor) in human choriocarcinoma. Cancer Lett. 2010, 291, 99–107. [Google Scholar] [CrossRef]
- Pang, R.T.; Leung, C.O.; Lee, C.L.; Lam, K.K.; Ye, T.M.; Chiu, P.C.; Yeung, W.S. MicroRNA-34a is a tumor suppressor in choriocarcinoma via regulation of Delta-like1. BMC Cancer 2013, 13, 1–10. [Google Scholar] [CrossRef]
- Guo, Z.; Sui, L.; Qi, J.; Sun, Q.; Xu, Y.; Zou, N.; Xie, Y.; Kong, Y. miR-196b inhibits cell migration and invasion through targeting MAP3K1 in hydatidiform mole. Biomed. Pharmacother. 2019, 113, 108760. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.-X.; Zhao, J.-R.; Xu, Y.-Y.; Wu, W.-B.; Zhang, H.-J. miR-21 Is Overexpressed in Hydatidiform Mole Tissues and Promotes Proliferation, Migration, and Invasion in Choriocarcinoma Cells. Int. J. Gynecol. Cancer 2017, 27, 364–374. [Google Scholar] [CrossRef]
- Palma Flores, C.; García-Vázquez, R.; Gallardo Rincón, D.; Ruiz-García, E.; Astudillo de la Vega, H.; Marchat, L.A.; Salinas Vera, Y.M.; López-Camarillo, C. MicroRNAs driving invasion and metastasis in ovarian cancer: Opportunities for translational medicine (Review). Int. J. Oncol. 2017, 50, 1461–1476. [Google Scholar] [CrossRef] [PubMed]
- Prahm, K.P.; Novotny, G.W.; Hogdall, C.; Hogdall, E. Current status on microRNAs as biomarkers for ovarian cancer. APMIS 2016, 124, 337–355. [Google Scholar] [CrossRef]
- Resnick, K.E.; Alder, H.; Hagan, J.P.; Richardson, D.L.; Croce, C.M.; Cohn, D.E. The detection of differentially expressed microRNAs from the serum of ovarian cancer patients using a novel real-time PCR platform. Gynecol. Oncol. 2009, 112, 55–59. [Google Scholar] [CrossRef]
- Cortez, M.A.; Bueso-Ramos, C.; Ferdin, J.; Lopez-Berestein, G.; Sood, A.K.; Calin, G.A. MicroRNAs in body fluids—the mix of hormones and biomarkers. Nat. Rev. Clin. Oncol. 2011, 8, 467–477. [Google Scholar] [CrossRef]
- Kim, Y.K. Extracellular microRNAs as Biomarkers in Human Disease. Chonnam Med. J. 2015, 51, 51–57. [Google Scholar] [CrossRef] [PubMed]
- Mitchell, P.S.; Parkin, R.K.; Kroh, E.M.; Fritz, B.R.; Wyman, S.K.; Pogosova-Agadjanyan, E.L.; Peterson, A.; Noteboom, J.; O’Briant, K.C.; Allen, A.; et al. Circulating microRNAs as stable blood-based markers for cancer detection. Proc. Natl. Acad. Sci. USA 2008, 105, 10513–10518. [Google Scholar] [CrossRef]
- Reid, G.; Kirschner, M.B.; van Zandwijk, N. Circulating microRNAs: Association with disease and potential use as biomarkers. Crit. Rev. Oncol. Hematol. 2011, 80, 193–208. [Google Scholar] [CrossRef]
- Ender, C.; Meister, G. Argonaute proteins at a glance. J. Cell Sci. 2010, 123, 1819–1823. [Google Scholar] [CrossRef]
- Iftikhar, H.; Carney, G.E. Evidence and potential in vivo functions for biofluid miRNAs: From expression profiling to functional testing: Potential roles of extracellular miRNAs as indicators of physiological change and as agents of intercellular information exchange. Bioessays 2016, 38, 367–378. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Xu, S.; Liu, X. MicroRNA profiling of plasma exosomes from patients with ovarian cancer using high-throughput sequencing. Oncol. Lett. 2019, 17, 5601–5607. [Google Scholar] [CrossRef] [PubMed]
- Penyige, A.; Marton, E.; Soltesz, B.; Szilagyi-Bonizs, M.; Poka, R.; Lukacs, J.; Szeles, L.; Nagy, B. Circulating miRNA Profiling in Plasma Samples of Ovarian Cancer Patients. Int. J. Mol. Sci. 2019, 20, 4533. [Google Scholar] [CrossRef]
- Chao, A.; Lai, C.H.; Chen, H.C.; Lin, C.Y.; Tsai, C.L.; Tang, Y.H.; Huang, H.J.; Lin, C.T.; Chen, M.Y.; Huang, K.G.; et al. Serum microRNAs in clear cell carcinoma of the ovary. Taiwan J. Obstet. Gynecol. 2014, 53, 536–541. [Google Scholar] [CrossRef] [PubMed]
- Murray, M.J.; Halsall, D.J.; Hook, C.E.; Williams, D.M.; Nicholson, J.C.; Coleman, N. Identification of microRNAs from the miR-371~373 and miR-302 clusters as potential serum biomarkers of malignant germ cell tumors. Am. J. Clin. Pathol. 2011, 135, 119–125. [Google Scholar] [CrossRef]
- Murray, M.J.; Smith, S.; Ward, D.; Verduci, L.; Nicholson, J.C.; Scarpini, C.G.; Coleman, N. Circulating microRNAs as biomarkers to assist the management of the malignant germ-cell-tumour subtype choriocarcinoma. Transl. Oncol. 2021, 14, 100904. [Google Scholar] [CrossRef]
- Yokoi, A.; Matsuzaki, J.; Yamamoto, Y.; Tate, K.; Yoneoka, Y.; Shimizu, H.; Uehara, T.; Ishikawa, M.; Takizawa, S.; Aoki, Y.; et al. Serum microRNA profile enables preoperative diagnosis of uterine leiomyosarcoma. Cancer Sci. 2019, 110, 3718–3726. [Google Scholar] [CrossRef] [PubMed]
- Hasegawa, Y.; Miura, K.; Furuya, K.; Yoshiura, K.; Masuzaki, H. Identification of complete hydatidiform mole pregnancy-associated microRNAs in plasma. Clin. Chem. 2013, 59, 1410–1412. [Google Scholar] [CrossRef] [PubMed]
- Miura, K.; Miura, S.; Yamasaki, K.; Higashijima, A.; Kinoshita, A.; Yoshiura, K.; Masuzaki, H. Identification of pregnancy-associated microRNAs in maternal plasma. Clin. Chem. 2010, 56, 1767–1771. [Google Scholar] [CrossRef] [PubMed]
- Erson-Bensan, A.E. Alternative polyadenylation and RNA-binding proteins. J. Mol. Endocrinol. 2016, 57, F29–F34. [Google Scholar] [CrossRef]
- Sandberg, R.; Neilson, J.R.; Sarma, A.; Sharp, P.A.; Burge, C.B. Proliferating cells express mRNAs with shortened 3′ untranslated regions and fewer microRNA target sites. Science 2008, 320, 1643–1647. [Google Scholar] [CrossRef]
- Mayr, C.; Bartel, D.P. Widespread shortening of 3’UTRs by alternative cleavage and polyadenylation activates oncogenes in cancer cells. Cell 2009, 138, 673–684. [Google Scholar] [CrossRef] [PubMed]
- Akman, B.H.; Can, T.; Erson-Bensan, A.E. Estrogen-induced upregulation and 3’-UTR shortening of CDC6. Nucleic Acids Res. 2012, 40, 10679–10688. [Google Scholar] [CrossRef]
- Akman, H.B.; Oyken, M.; Tuncer, T.; Can, T.; Erson-Bensan, A.E. 3′UTR shortening and EGF signaling: Implications for breast cancer. Hum. Mol. Genet. 2015, 24, 6910–6920. [Google Scholar] [CrossRef]
- Gruber, A.J.; Zavolan, M. Alternative cleavage and polyadenylation in health and disease. Nat. Rev. Genet. 2019, 20, 599–614. [Google Scholar] [CrossRef]
- Singh, P.; Alley, T.L.; Wright, S.M.; Kamdar, S.; Schott, W.; Wilpan, R.Y.; Mills, K.D.; Graber, J.H. Global changes in processing of mRNA 3’ untranslated regions characterize clinically distinct cancer subtypes. Cancer Res. 2009, 69, 9422–9430. [Google Scholar] [CrossRef] [PubMed]
- Stricker, T.P.; Brown, C.D.; Bandlamudi, C.; McNerney, M.; Kittler, R.; Montoya, V.; Peterson, A.; Grossman, R.; White, K.P. Robust stratification of breast cancer subtypes using differential patterns of transcript isoform expression. PLoS Genet. 2017, 13, e1006589. [Google Scholar] [CrossRef]
- Begik, O.; Oyken, M.; Cinkilli Alican, T.; Can, T.; Erson-Bensan, A.E. Alternative Polyadenylation Patterns for Novel Gene Discovery and Classification in Cancer. Neoplasia 2017, 19, 574–582. [Google Scholar] [CrossRef] [PubMed]
- Xue, Z.; Warren, R.L.; Gibb, E.A.; MacMillan, D.; Wong, J.; Chiu, R.; Hammond, S.A.; Yang, C.; Nip, K.M.; Ennis, C.A.; et al. Recurrent tumor-specific regulation of alternative polyadenylation of cancer-related genes. BMC Genom. 2018, 19, 536. [Google Scholar] [CrossRef]
- Chehade, M.; Bullock, M.; Glover, A.; Hutvagner, G.; Sidhu, S. Key MicroRNA’s and Their Targetome in Adrenocortical Cancer. Cancers 2020, 12, 2198. [Google Scholar] [CrossRef]
- Mendonsa, A.M.; Na, T.Y.; Gumbiner, B.M. E-cadherin in contact inhibition and cancer. Oncogene 2018, 37, 4769–4780. [Google Scholar] [CrossRef] [PubMed]
- Vu, T.T.; Stölzel, F.; Wang, K.W.; Röllig, C.; Tursky, M.L.; Molloy, T.J.; Ma, D.D. miR-10a as a therapeutic target and predictive biomarker for MDM2 inhibition in acute myeloid leukemia. Leukemia 2020. [Google Scholar] [CrossRef] [PubMed]
- Jiajie, T.; Yanzhou, Y.; Hoi-Hung, A.C.; Zi-Jiang, C.; Wai-Yee, C. Conserved miR-10 family represses proliferation and induces apoptosis in ovarian granulosa cells. Sci. Rep. 2017, 7, 41304. [Google Scholar] [CrossRef] [PubMed]
- Feng, Y.H.; Tsao, C.J. Emerging role of microRNA-21 in cancer. Biomed. Rep. 2016, 5, 395–402. [Google Scholar] [CrossRef] [PubMed]
- Jiang, X.M.; Xiang, G.; Wang, Y.; Zhang, L.; Yang, X.; Cao, L.; Peng, H.; Xue, P.; Chen, D. MicroRNA-590-5p regulates proliferation and invasion in human hepatocellular carcinoma cells by targeting TGF-β RII. Mol. Cells 2012, 33, 545–551. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Yao, D.; Li, Y.; Chen, H.; He, C.; Ding, N.; Lu, Y.; Ou, T.; Zhao, S.; Li, L.; et al. Serum microRNA expression levels can predict lymph node metastasis in patients with early-stage cervical squamous cell carcinoma. Int. J. Mol. Med. 2013, 32, 557–567. [Google Scholar] [CrossRef]
- Stark, M.S.; Tyagi, S.; Nancarrow, D.J.; Boyle, G.M.; Cook, A.L.; Whiteman, D.C.; Parsons, P.G.; Schmidt, C.; Sturm, R.A.; Hayward, N.K. Characterization of the melanoma miRNAome by deep sequencing. PLoS ONE 2010, 5, e9685. [Google Scholar] [CrossRef]
- Wang, F.Y.; Kang, C.S.; Wang-Gou, S.Y.; Huang, C.H.; Feng, C.Y.; Li, X.J. EGFL7 is an intercellular EGFR signal messenger that plays an oncogenic role in glioma. Cancer Lett. 2017, 384, 9–18. [Google Scholar] [CrossRef]
- Huang, C.; Yuan, X.; Wan, Y.; Liu, F.; Chen, X.; Zhan, X.; Li, X. VE-statin/Egfl7 expression in malignant glioma and its relevant molecular network. Int. J. Clin. Exp. Pathol. 2014, 7, 1022–1031. [Google Scholar]
- Hansen, T.F.; Andersen, R.F.; Olsen, D.A.; Sørensen, F.B.; Jakobsen, A. Prognostic importance of circulating epidermal growth factor-like domain 7 in patients with metastatic colorectal cancer treated with chemotherapy and bevacizumab. Sci. Rep. 2017, 7, 1–9. [Google Scholar] [CrossRef]
- Papaioannou, D.; Shen, C.; Nicolet, D.; McNeil, B.; Bill, M.; Karunasiri, M.; Burke, M.H.; Ozer, H.G.; Yilmaz, S.A.; Zitzer, N.; et al. Prognostic and biological significance of the proangiogenic factor EGFL7 in acute myeloid leukemia. Proc. Natl. Acad. Sci. USA 2017, 114, E4641–E4647. [Google Scholar] [CrossRef]
- Oh, J.; Park, S.H.; Lee, T.S.; Oh, H.K.; Choi, J.H.; Choi, Y.S. High expression of epidermal growth factor-like domain 7 is correlated with poor differentiation and poor prognosis in patients with epithelial ovarian cancer. J. Gynecol. Oncol. 2014, 25, 334–341. [Google Scholar] [CrossRef]
- Yamauchi, M.; Fukuda, T.; Wada, T.; Kawanishi, M.; Imai, K.; Tasaka, R.; Yasui, T.; Sumi, T. Expression of epidermal growth factor-like domain 7 may be a predictive marker of the effect of neoadjuvant chemotherapy for locally advanced uterine cervical cancer. Oncol. Lett. 2016, 12, 5183–5189. [Google Scholar] [CrossRef] [PubMed]
- Andersen, M.; Trapani, D.; Ravn, J.; Sørensen, J.B.; Andersen, C.B.; Grauslund, M.; Santoni-Rugiu, E. Methylation-associated Silencing of microRNA-126 and its Host Gene EGFL7 in Malignant Pleural Mesothelioma. Anticancer Res. 2015, 35, 6223–6229. [Google Scholar]
- Bommer, G.T.; Gerin, I.; Feng, Y.; Kaczorowski, A.J.; Kuick, R.; Love, R.E.; Zhai, Y.; Giordano, T.J.; Qin, Z.S.; Moore, B.B.; et al. p53-mediated activation of miRNA34 candidate tumor-suppressor genes. Curr. Biol. 2007, 17, 1298–1307. [Google Scholar] [CrossRef] [PubMed]
- He, L.; He, X.; Lim, L.P.; de Stanchina, E.; Xuan, Z.; Liang, Y.; Xue, W.; Zender, L.; Magnus, J.; Ridzon, D.; et al. A microRNA component of the p53 tumour suppressor network. Nature 2007, 447, 1130–1134. [Google Scholar] [CrossRef]
- Chang, T.C.; Wentzel, E.A.; Kent, O.A.; Ramachandran, K.; Mullendore, M.; Lee, K.H.; Feldmann, G.; Yamakuchi, M.; Ferlito, M.; Lowenstein, C.J.; et al. Transactivation of miR-34a by p53 broadly influences gene expression and promotes apoptosis. Mol. Cell. 2007, 26, 745–752. [Google Scholar] [CrossRef] [PubMed]
- Bhatia, S.; Kaul, D.; Varma, N. Functional genomics of tumor suppressor miR-196b in T-cell acute lymphoblastic leukemia. Mol. Cell. Biochem. 2011, 346, 103–116. [Google Scholar] [CrossRef] [PubMed]
- Coskun, E.; von der Heide, E.K.; Schlee, C.; Kühnl, A.; Gökbuget, N.; Hoelzer, D.; Hofmann, W.K.; Thiel, E.; Baldus, C.D. The role of microRNA-196a and microRNA-196b as ERG regulators in acute myeloid leukemia and acute T-lymphoblastic leukemia. Leuk. Res. 2011, 35, 208–213. [Google Scholar] [CrossRef]
- How, C.; Hui, A.B.; Alajez, N.M.; Shi, W.; Boutros, P.C.; Clarke, B.A.; Yan, R.; Pintilie, M.; Fyles, A.; Hedley, D.W.; et al. MicroRNA-196b regulates the homeobox B7-vascular endothelial growth factor axis in cervical cancer. PLoS ONE 2013, 8, e67846. [Google Scholar] [CrossRef]
| Malignancy | Sample Type | References | |||
|---|---|---|---|---|---|
| Tissue | Cell Line | ||||
| Upregulated | Downregulated | Upregulated | Downregulated | ||
| Clear cell ovarian cancer | miR-9 miR-34a miR-126 miR-509-3-5p miR-509-3p miR-509-5p miR-510 | miR-29b miR-449 | miR-9 | miR-424 | [53–57] |
| Mucinous ovarian cancer | miR-192 miR-194 miR-215 | [58] | |||
| Ovarian germ cell tumor | miR-122 miR-126 miR-302a miR-302b miR-302c-3p miR-302d miR-371-5p miR-372-3p miR-373 miR-373-3p | miR-199a-5p miR-202-3p miR-214-5p | [68,69] | ||
| Granulosa cell tumors of the ovary | miR-10a | miR-29c-3p miR-126 miR-138-5p miR-184 miR-204-5p miR-328-3p miR-501-3p | miR-17 family miR-10a | miR-126 | [70–73] |
| Sex cord stromal ovary tumor | miR-202c-3p miR-513c-5p | [68] | |||
| Uterine sarcomas (Leiomyosarcoma, Endometrial stromal sarcoma, Mixed epithelial–mesenchymal tumors) | miR-7-5p miR-34c-5p miR-138-5p miR-196a-5p miR-202-3p miR-210-3p miR-301a-3p miR-335-5p miR-372-3p miR-373-3p | miR-1 miR-1-3p miR-10a-5p miR-23-3p miR-23b miR-125a-5p let-7 family | miR-129-5p miR-141-3p miR-148a-3p miR-202-3p miR-203a-3p | miR-1 miR-1-3p miR-125b-1-3p miR-140-5p miR-152-3p miR-21-5p miR-27b-3p miR-485-5p miR-495-3p | [86–92] |
| Uterine carcinosarcoma | miR-184 | let-7a let-7b-5p let-7d miR-16 miR-26a miR-30c miR-124-3p miR-200 family miR-214 | miR-200c | [98,100,101] | |
| Vulvar carcinoma | miR-133a miR-519b | miR-19-b1-5p miR-100-3p miR-223-5p | [107] | ||
| Vulvar squamous cell carcinoma | miR-590-5p miR-182-5p miR-183-5p miR-3147 miR-4712-5p | miR-603 miR-103a-3p miR-107 | miR-182-5p miR-183-5p miR-223-5p miR-590-5p miR-4712-5p miR-3147 | miR-103a-3p miR-107 miR-603 | [108–110] |
| Melanoma of the female genital tract | miR-17-5p miR-19b-3p miR-20a-5p miR-20b-5p miR-146a-5p | miR-15 miR-99a-5p miR-145-5p miR-200a-3p miR-200b-3p miR-494-p miR-1972 | [113] | ||
| Gestational trophoblastic disease | miR-21 miR-181b-5p miR-181d-5p miR-371a-5p miR-518a-3p miR-519d-3p miR-520a-3p miR-934 | miR-199b miR-370-3p miR-517a miR-517b miR-518b miR-519a | miR-21 miR-371a-5p miR-518a-3p | miR-34a miR-196b miR-199b | [120–126] |
| Malignancy | Serum/Plasma miRNA | References | |
|---|---|---|---|
| Upregulated | Downregulated | ||
| Clear cell ovarian cancer | miR-130a miR-138 miR-187 miR-202 | [138] | |
| Ovarian germ cell tumor | miR-302 miR-371~373 Chromosome 19 microRNA cluster (C19MC) miR-371a-3p | [139,140] | |
| Uterine sarcomas (Leiomyosarcoma, Endometrial stromal sarcoma, Mixed epithelial–mesenchymal tumors) | miR-191-5p miR-1246 | [141] | |
| Gestational trophoblastic disease | miR-520b miR-520f miR-520c-3p | [142,143] | |
| miRNA | Up- or Down-Regulated | Validated Targets | Pathway/Process Affected | Cell Line | References |
|---|---|---|---|---|---|
| miR-9 | Upregulated | CDH1 | EMT | OCCC | [55] |
| miR-10a | Upregulated | PTEN | Akt and Wnt pathways | Cancerous granulosa | [73] |
| miR-21 | Upregulated | PDCD4 | Akt pathway | Choriocarcinoma | [126] |
| miR-590-5p | Upregulated | TGFβRII | TGFβ pathway | VSCC | [108] |
| miR-3147 | Upregulated | SMAD4 | TGFβ pathway | VSCC | [110] |
| miR-4712-5p | Upregulated | PTEN | AKT, GSK3β and cyclin D1 signaling pathways | VSCC | [109] |
| miR-34a | Downregulated | DLL1 | Notch pathway | Choriocarcinoma | [124] |
| miR-126 | Downregulated | EGFL7 | PI3K/AKT pathway | Cancerous granulosa | [72] |
| miR-196b | Downregulated | MAP3K1 | Cell migration and invasion | Choriocarcinoma | [125] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Di Fiore, R.; Suleiman, S.; Pentimalli, F.; O’Toole, S.A.; O’Leary, J.J.; Ward, M.P.; Conlon, N.T.; Sabol, M.; Ozretić, P.; Erson-Bensan, A.E.; et al. Could MicroRNAs Be Useful Tools to Improve the Diagnosis and Treatment of Rare Gynecological Cancers? A Brief Overview. Int. J. Mol. Sci. 2021, 22, 3822. https://doi.org/10.3390/ijms22083822
Di Fiore R, Suleiman S, Pentimalli F, O’Toole SA, O’Leary JJ, Ward MP, Conlon NT, Sabol M, Ozretić P, Erson-Bensan AE, et al. Could MicroRNAs Be Useful Tools to Improve the Diagnosis and Treatment of Rare Gynecological Cancers? A Brief Overview. International Journal of Molecular Sciences. 2021; 22(8):3822. https://doi.org/10.3390/ijms22083822
Chicago/Turabian StyleDi Fiore, Riccardo, Sherif Suleiman, Francesca Pentimalli, Sharon A. O’Toole, John J. O’Leary, Mark P. Ward, Neil T. Conlon, Maja Sabol, Petar Ozretić, Ayse Elif Erson-Bensan, and et al. 2021. "Could MicroRNAs Be Useful Tools to Improve the Diagnosis and Treatment of Rare Gynecological Cancers? A Brief Overview" International Journal of Molecular Sciences 22, no. 8: 3822. https://doi.org/10.3390/ijms22083822
APA StyleDi Fiore, R., Suleiman, S., Pentimalli, F., O’Toole, S. A., O’Leary, J. J., Ward, M. P., Conlon, N. T., Sabol, M., Ozretić, P., Erson-Bensan, A. E., Reed, N., Giordano, A., Herrington, C. S., & Calleja-Agius, J. (2021). Could MicroRNAs Be Useful Tools to Improve the Diagnosis and Treatment of Rare Gynecological Cancers? A Brief Overview. International Journal of Molecular Sciences, 22(8), 3822. https://doi.org/10.3390/ijms22083822











