Heparan Sulfate Deficiency in Cartilage: Enhanced BMP-Sensitivity, Proteoglycan Production and an Anti-Apoptotic Expression Signature after Loading
Abstract
1. Introduction
2. Results
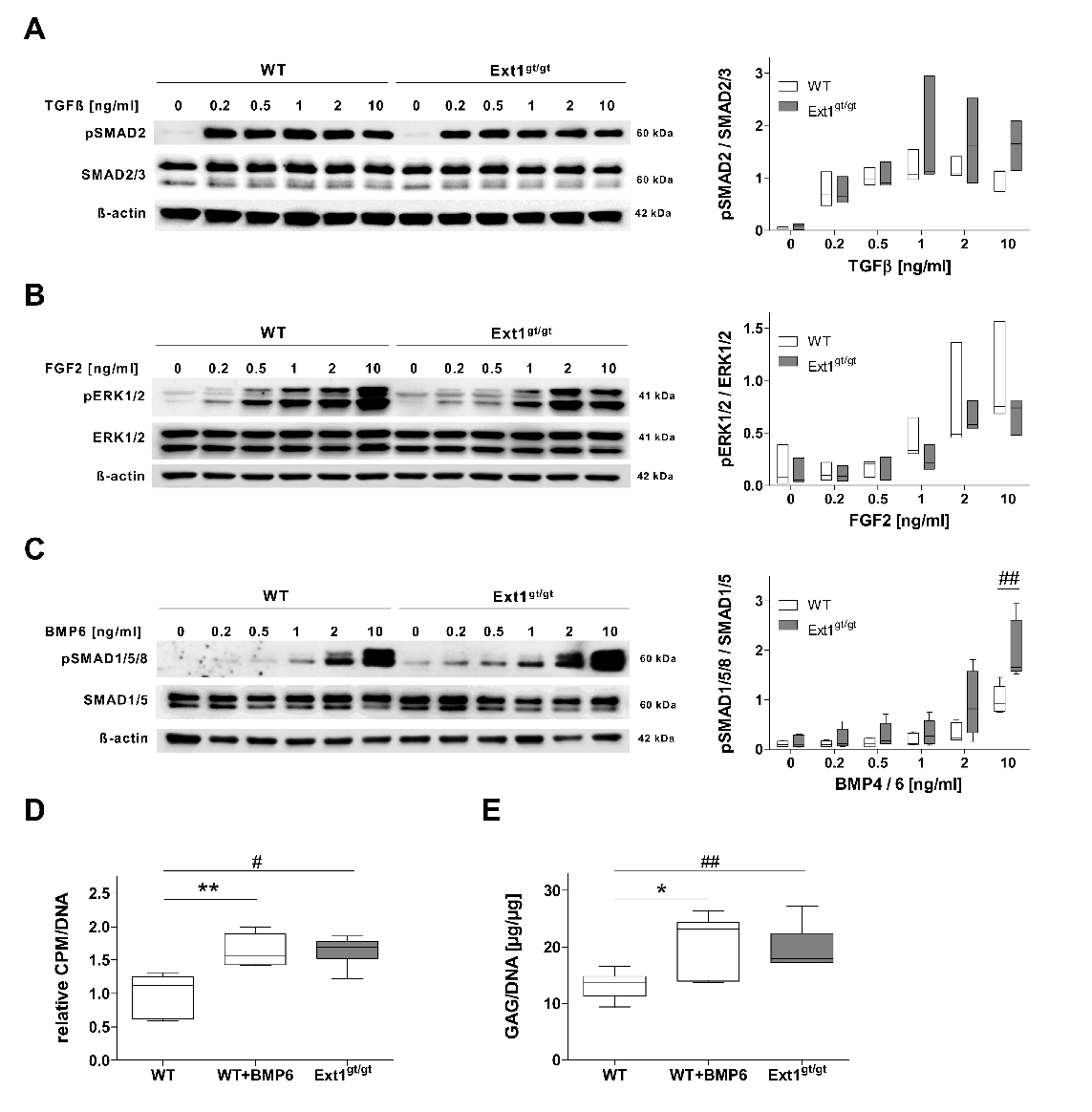
2.1. Elevated Glycosaminoglycan Production in HS-Deficient Engineered Cartilage
2.2. Higher BMP-Sensitivity of HS-Reduced Chondrocytes
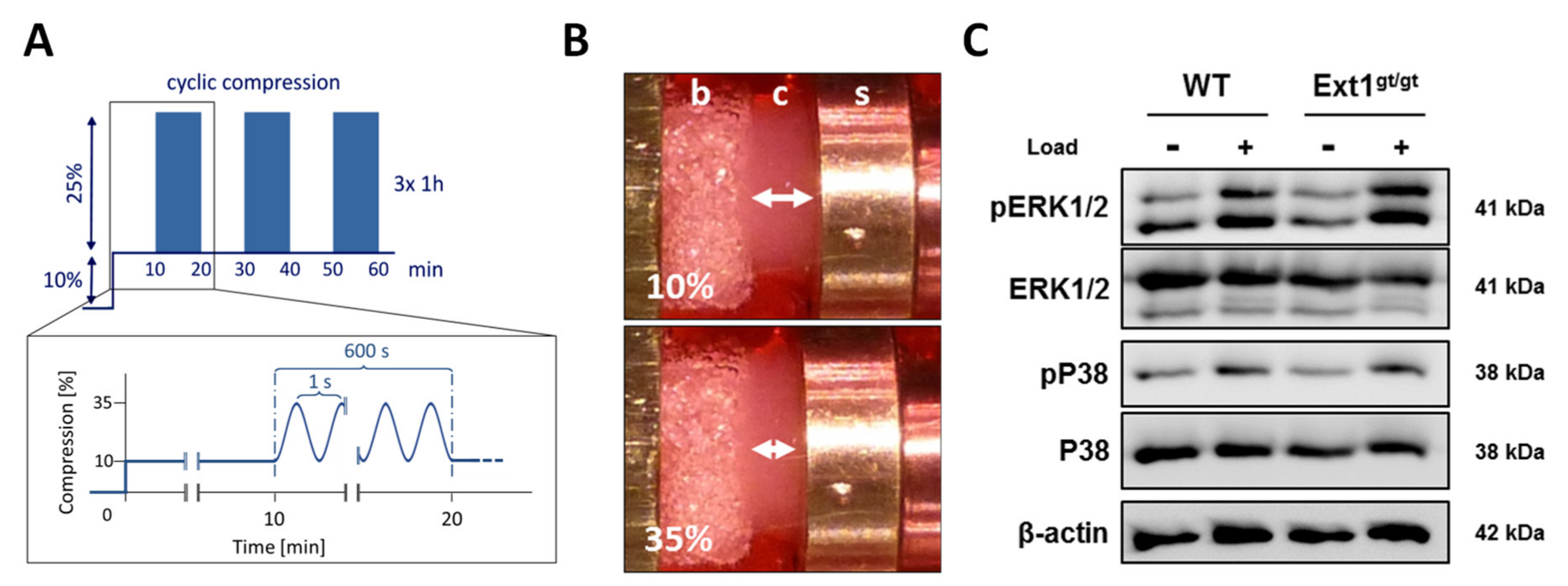
2.3. Similar Load-Induced Activation of ERK1/2 and P38 Pathways
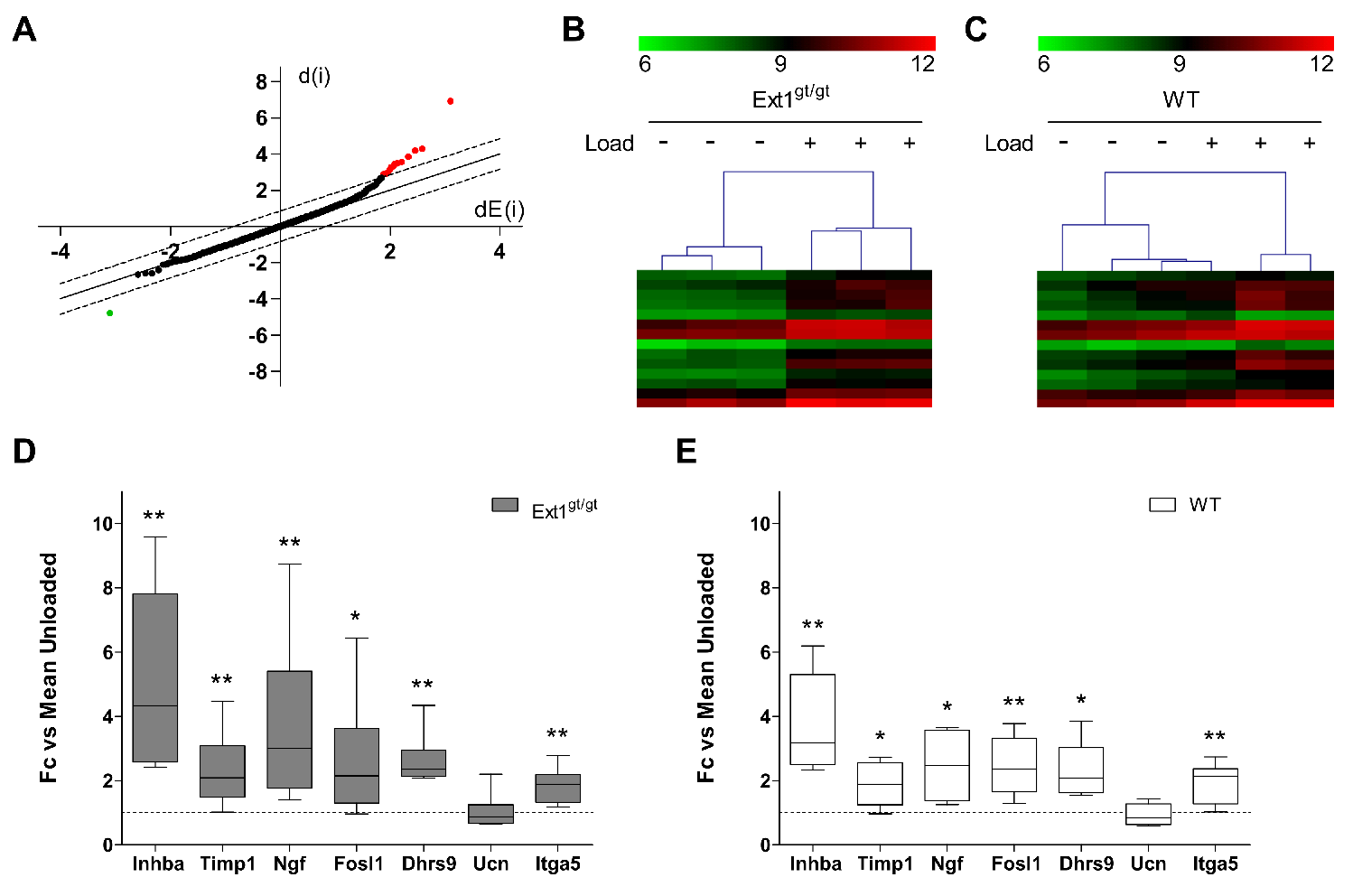
2.4. Global Molecular Characterization of the Loading Response
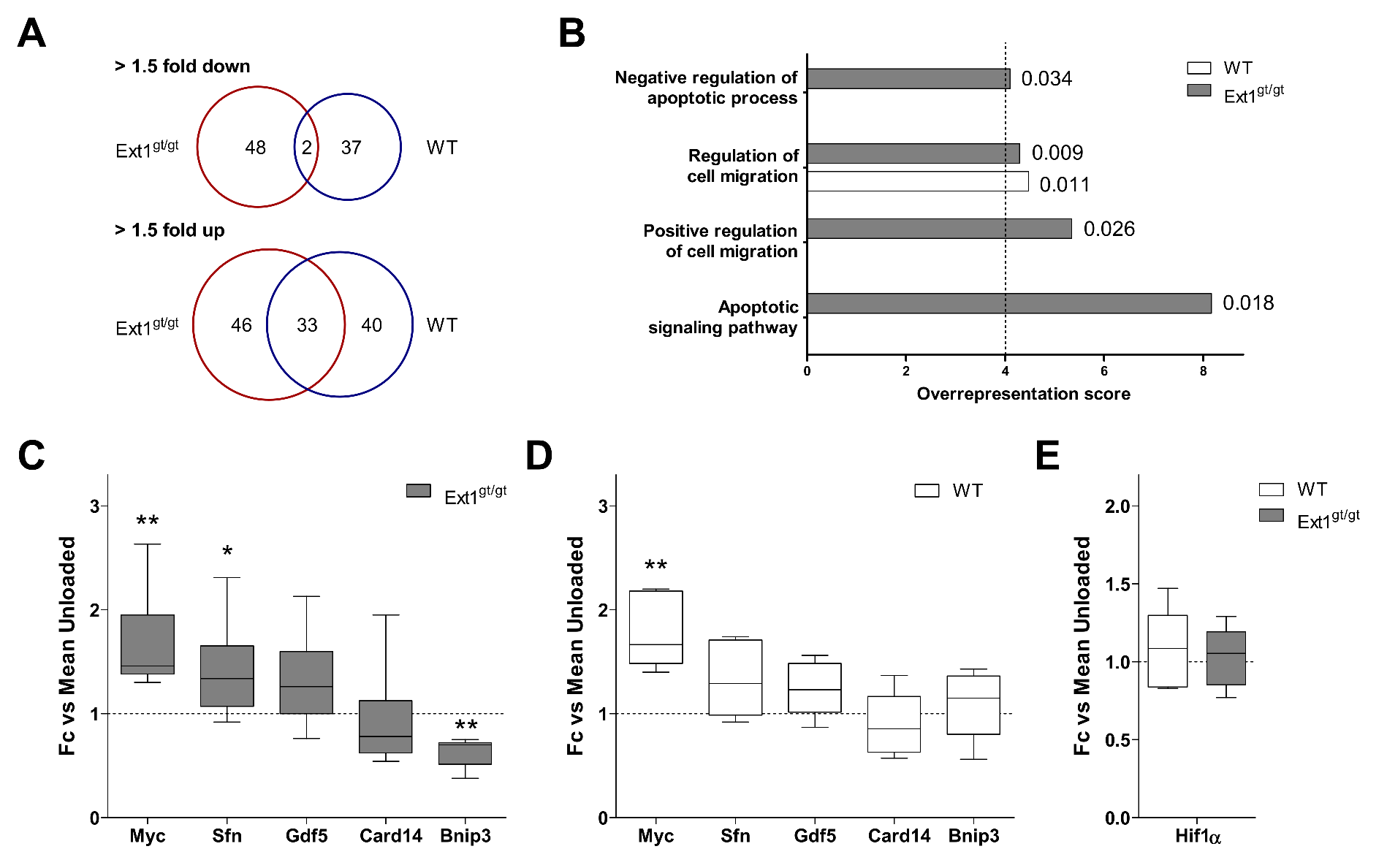
2.5. Induction of an Anti-Apoptotic Expression Signature in HS-Deficient Cartilage by Loading
3. Discussion
4. Materials and Methods
4.1. Transgenic Mice
4.2. Cell Isolation and Culture
4.3. Generation and Culture of Engineered Cartilage
4.4. RNA Isolation and RT-qPCR
4.5. Histology
4.6. GAG and DNA-Quantification
4.7. GAG-Synthesis
4.8. Mechanical Loading
4.9. Protein Lysates and Western Blot
4.10. Caspase 3 Activation and MTT Assay
4.11. Microarray Analysis
4.12. Statistics
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Poole, A.R.; Kojima, T.; Yasuda, T.; Mwale, F.; Kobayashi, M.; Laverty, S. Composition and structure of articular cartilage: A template for tissue repair. Clin. Orthop. Relat. Res. 2001, 391, S26–S33. [Google Scholar] [CrossRef] [PubMed]
- Vannini, F.; Spalding, T.; Andriolo, L.; Berruto, M.; Denti, M.; Espregueira-Mendes, J.; Menetrey, J.; Peretti, G.M.; Seil, R.; Filardo, G. Sport and early osteoarthritis: The role of sport in aetiology, progression and treatment of knee osteoarthritis. Knee Surg. Sports Traumatol. Arthrosc. Off. J. Esska 2016, 24, 1786–1796. [Google Scholar] [CrossRef]
- Valderrabano, V.; Steiger, C. Treatment and Prevention of Osteoarthritis through Exercise and Sports. J Aging Res 2010, 2011, 374653. [Google Scholar] [CrossRef] [PubMed]
- Iijima, H.; Ito, A.; Nagai, M.; Tajino, J.; Yamaguchi, S.; Kiyan, W.; Nakahata, A.; Zhang, J.; Wang, T.; Aoyama, T.; et al. Physiological exercise loading suppresses post-traumatic osteoarthritis progression via an increase in bone morphogenetic proteins expression in an experimental rat knee model. Osteoarthr. Cartil. 2017, 25, 964–975. [Google Scholar] [CrossRef]
- Venn, M.; Maroudas, A. Chemical composition and swelling of normal and osteoarthrotic femoral head cartilage. I. Chemical composition. Ann. Rheum. Dis. 1977, 36, 121–129. [Google Scholar] [CrossRef] [PubMed]
- Palukuru, U.P.; McGoverin, C.M.; Pleshko, N. Assessment of hyaline cartilage matrix composition using near infrared spectroscopy. Matrix Biol. J. Int. Soc. Matrix Biol. 2014, 38, 3–11. [Google Scholar] [CrossRef] [PubMed]
- Cs-Szabo, G.; Roughley, P.J.; Plaas, A.H.; Glant, T.T. Large and small proteoglycans of osteoarthritic and rheumatoid articular cartilage. Arthritis Rheum. 1995, 38, 660–668. [Google Scholar] [CrossRef] [PubMed]
- Malemud, C.J.; Papay, R.S.; Hering, T.M.; Holderbaum, D.; Goldberg, V.M.; Haqqi, T.M. Phenotypic modulation of newly synthesized proteoglycans in human cartilage and chondrocytes. Osteoarthr. Cartil. 1995, 3, 227–238. [Google Scholar] [CrossRef][Green Version]
- Venkatesan, N.; Barre, L.; Bourhim, M.; Magdalou, J.; Mainard, D.; Netter, P.; Fournel-Gigleux, S.; Ouzzine, M. Xylosyltransferase-I regulates glycosaminoglycan synthesis during the pathogenic process of human osteoarthritis. PLoS ONE 2012, 7, e34020. [Google Scholar] [CrossRef]
- Bachvarova, V.; Dierker, T.; Esko, J.; Hoffmann, D.; Kjellen, L.; Vortkamp, A. Chondrocytes respond to an altered heparan sulfate composition with distinct changes of heparan sulfate structure and increased levels of chondroitin sulfate. Matrix Biol. J. Int. Soc. Matrix Biol. 2020, 93, 43–59. [Google Scholar] [CrossRef]
- Vynios, D.H.; Papadas Th, A.; Faraos, A.; Mastronikolis, N.S.; Goumas, P.; Tsiganos, C.P. A solid phase assay for the determination of heparan sulfate and its application to normal and cancerous human cartilage samples. J. Immunoass. Immunochem. 2001, 22, 337–351. [Google Scholar] [CrossRef]
- Severmann, A.C.; Jochmann, K.; Feller, K.; Bachvarova, V.; Piombo, V.; Stange, R.; Holzer, T.; Brachvogel, B.; Esko, J.; Pap, T.; et al. An altered heparan sulfate structure in the articular cartilage protects against osteoarthritis. Osteoarthr. Cartil. 2020, 28, 977–987. [Google Scholar] [CrossRef]
- Echtermeyer, F.; Bertrand, J.; Dreier, R.; Meinecke, I.; Neugebauer, K.; Fuerst, M.; Lee, Y.J.; Song, Y.W.; Herzog, C.; Theilmeier, G.; et al. Syndecan-4 regulates ADAMTS-5 activation and cartilage breakdown in osteoarthritis. Nat. Med. 2009, 15, 1072–1076. [Google Scholar] [CrossRef]
- Shu, C.C.; Jackson, M.T.; Smith, M.M.; Smith, S.M.; Penm, S.; Lord, M.S.; Whitelock, J.M.; Little, C.B.; Melrose, J. Ablation of Perlecan Domain 1 Heparan Sulfate Reduces Progressive Cartilage Degradation, Synovitis, and Osteophyte Size in a Preclinical Model of Posttraumatic Osteoarthritis. Arthritis Rheumatol. (Hobokenn. J.) 2016, 68, 868–879. [Google Scholar] [CrossRef]
- Chanalaris, A.; Clarke, H.; Guimond, S.E.; Vincent, T.L.; Turnbull, J.E.; Troeberg, L. Heparan Sulfate Proteoglycan Synthesis Is Dysregulated in Human Osteoarthritic Cartilage. Am. J. Pathol. 2019, 189, 632–647. [Google Scholar] [CrossRef]
- Beauvais, D.M.; Ell, B.J.; McWhorter, A.R.; Rapraeger, A.C. Syndecan-1 regulates alphavbeta3 and alphavbeta5 integrin activation during angiogenesis and is blocked by synstatin, a novel peptide inhibitor. J. Exp. Med. 2009, 206, 691–705. [Google Scholar] [CrossRef]
- Vuoriluoto, K.; Jokinen, J.; Kallio, K.; Salmivirta, M.; Heino, J.; Ivaska, J. Syndecan-1 supports integrin alpha2beta1-mediated adhesion to collagen. Exp. Cell Res. 2008, 314, 3369–3381. [Google Scholar] [CrossRef]
- Bass, M.D.; Roach, K.A.; Morgan, M.R.; Mostafavi-Pour, Z.; Schoen, T.; Muramatsu, T.; Mayer, U.; Ballestrem, C.; Spatz, J.P.; Humphries, M.J. Syndecan-4-dependent Rac1 regulation determines directional migration in response to the extracellular matrix. J. Cell Biol. 2007, 177, 527–538. [Google Scholar] [CrossRef]
- McQuade, K.J.; Beauvais, D.M.; Burbach, B.J.; Rapraeger, A.C. Syndecan-1 regulates alphavbeta5 integrin activity in B82L fibroblasts. J. Cell Sci. 2006, 119, 2445–2456. [Google Scholar] [CrossRef]
- Langhe, R.P.; Gudzenko, T.; Bachmann, M.; Becker, S.F.; Gonnermann, C.; Winter, C.; Abbruzzese, G.; Alfandari, D.; Kratzer, M.C.; Franz, C.M.; et al. Cadherin-11 localizes to focal adhesions and promotes cell-substrate adhesion. Nat. Commun. 2016, 7, 10909. [Google Scholar] [CrossRef]
- Gopal, S.; Sogaard, P.; Multhaupt, H.A.; Pataki, C.; Okina, E.; Xian, X.; Pedersen, M.E.; Stevens, T.; Griesbeck, O.; Park, P.W.; et al. Transmembrane proteoglycans control stretch-activated channels to set cytosolic calcium levels. J. Cell Biol. 2015, 210, 1199–1211. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Echtermeyer, F.; Thilo, F.; Theilmeier, G.; Schmidt, A.; Schulein, R.; Jensen, B.L.; Loddenkemper, C.; Jankowski, V.; Marcussen, N.; et al. The proteoglycan syndecan 4 regulates transient receptor potential canonical 6 channels via RhoA/Rho-associated protein kinase signaling. Arter. Thromb. Vasc. Biol. 2012, 32, 378–385. [Google Scholar] [CrossRef]
- Couchman, J.R. Transmembrane signaling proteoglycans. Annu. Rev. Cell Dev. Biol. 2010, 26, 89–114. [Google Scholar] [CrossRef] [PubMed]
- Mitsou, I.; Multhaupt, H.A.B.; Couchman, J.R. Proteoglycans, ion channels and cell-matrix adhesion. Biochem. J. 2017, 474, 1965–1979. [Google Scholar] [CrossRef] [PubMed]
- Vincent, T.L.; Hermansson, M.A.; Hansen, U.N.; Amis, A.A.; Saklatvala, J. Basic fibroblast growth factor mediates transduction of mechanical signals when articular cartilage is loaded. Arthritis Rheum. 2004, 50, 526–533. [Google Scholar] [CrossRef] [PubMed]
- Vincent, T.L.; McLean, C.J.; Full, L.E.; Peston, D.; Saklatvala, J. FGF-2 is bound to perlecan in the pericellular matrix of articular cartilage, where it acts as a chondrocyte mechanotransducer. Osteoarthr. Cartil. 2007, 15, 752–763. [Google Scholar] [CrossRef]
- Shi, M.; Zhu, J.; Wang, R.; Chen, X.; Mi, L.; Walz, T.; Springer, T.A. Latent TGF-beta structure and activation. Nature 2011, 474, 343–349. [Google Scholar] [CrossRef] [PubMed]
- Forsten-Williams, K.; Chua, C.C.; Nugent, M.A. The kinetics of FGF-2 binding to heparan sulfate proteoglycans and MAP kinase signaling. J. Theor. Biol. 2005, 233, 483–499. [Google Scholar] [CrossRef]
- Lyon, M.; Rushton, G.; Gallagher, J.T. The interaction of the transforming growth factor-betas with heparin/heparan sulfate is isoform-specific. J. Biol. Chem. 1997, 272, 18000–18006. [Google Scholar] [CrossRef]
- Irie, A.; Habuchi, H.; Kimata, K.; Sanai, Y. Heparan sulfate is required for bone morphogenetic protein-7 signaling. Biochem. Biophys. Res. Commun. 2003, 308, 858–865. [Google Scholar] [CrossRef]
- Praxenthaler, H.; Kramer, E.; Weisser, M.; Hecht, N.; Fischer, J.; Grossner, T.; Richter, W. Extracellular matrix content and WNT/beta-catenin levels of cartilage determine the chondrocyte response to compressive load. Biochim. Biophys. Acta Mol. Basis Dis. 2018, 1864, 851–859. [Google Scholar] [CrossRef]
- Scholtes, S.; Kramer, E.; Weisser, M.; Roth, W.; Luginbuhl, R.; Grossner, T.; Richter, W. Global chondrocyte gene expression after a single anabolic loading period: Time evolution and re-inducibility of mechano-responses. J. Cell. Physiol. 2018, 233, 699–711. [Google Scholar] [CrossRef]
- Monaco, G.; El Haj, A.J.; Alini, M.; Stoddart, M.J. Ex Vivo Systems to Study Chondrogenic Differentiation and Cartilage Integration. J. Funct. Morphol. Kinesiol. 2021, 6, 6. [Google Scholar] [CrossRef]
- Hall, A.C. The Role of Chondrocyte Morphology and Volume in Controlling Phenotype-Implications for Osteoarthritis, Cartilage Repair, and Cartilage Engineering. Curr. Rheumatol. Rep. 2019, 21, 38. [Google Scholar] [CrossRef]
- Kreuger, J.; Kjellen, L. Heparan sulfate biosynthesis: Regulation and variability. J. Histochem. Cytochem. Off. J. Histochem. Soc. 2012, 60, 898–907. [Google Scholar] [CrossRef]
- Lin, X.; Wei, G.; Shi, Z.; Dryer, L.; Esko, J.D.; Wells, D.E.; Matzuk, M.M. Disruption of gastrulation and heparan sulfate biosynthesis in EXT1-deficient mice. Dev. Biol. 2000, 224, 299–311. [Google Scholar] [CrossRef]
- Mitchell, K.J.; Pinson, K.I.; Kelly, O.G.; Brennan, J.; Zupicich, J.; Scherz, P.; Leighton, P.A.; Goodrich, L.V.; Lu, X.; Avery, B.J.; et al. Functional analysis of secreted and transmembrane proteins critical to mouse development. Nat. Genet. 2001, 28, 241–249. [Google Scholar] [CrossRef]
- Koziel, L.; Kunath, M.; Kelly, O.G.; Vortkamp, A. Ext1-dependent heparan sulfate regulates the range of Ihh signaling during endochondral ossification. Dev. Cell 2004, 6, 801–813. [Google Scholar] [CrossRef]
- Otsuki, S.; Hanson, S.R.; Miyaki, S.; Grogan, S.P.; Kinoshita, M.; Asahara, H.; Wong, C.H.; Lotz, M.K. Extracellular sulfatases support cartilage homeostasis by regulating BMP and FGF signaling pathways. Proc. Natl. Acad. Sci. USA 2010, 107, 10202–10207. [Google Scholar] [CrossRef]
- Mundy, C.; Yang, E.; Takano, H.; Billings, P.C.; Pacifici, M. Heparan sulfate antagonism alters bone morphogenetic protein signaling and receptor dynamics, suggesting a mechanism in hereditary multiple exostoses. J. Biol. Chem. 2018, 293, 7703–7716. [Google Scholar] [CrossRef]
- Ornitz, D.M. FGFs, heparan sulfate and FGFRs: Complex interactions essential for development. Bioessays News Rev. Mol. Cell. Dev. Biol. 2000, 22, 108–112. [Google Scholar] [CrossRef]
- Paine-Saunders, S.; Viviano, B.L.; Economides, A.N.; Saunders, S. Heparan sulfate proteoglycans retain Noggin at the cell surface: A potential mechanism for shaping bone morphogenetic protein gradients. J. Biol. Chem. 2002, 277, 2089–2096. [Google Scholar] [CrossRef]
- Bougault, C.; Aubert-Foucher, E.; Paumier, A.; Perrier-Groult, E.; Huot, L.; Hot, D.; Duterque-Coquillaud, M.; Mallein-Gerin, F. Dynamic compression of chondrocyte-agarose constructs reveals new candidate mechanosensitive genes. PLoS ONE 2012, 7, e36964. [Google Scholar] [CrossRef] [PubMed]
- Wright, M.O.; Nishida, K.; Bavington, C.; Godolphin, J.L.; Dunne, E.; Walmsley, S.; Jobanputra, P.; Nuki, G.; Salter, D.M. Hyperpolarisation of cultured human chondrocytes following cyclical pressure-induced strain: Evidence of a role for alpha 5 beta 1 integrin as a chondrocyte mechanoreceptor. J. Orthop. Res. Off. Publ. Orthop. Res. Soc. 1997, 15, 742–747. [Google Scholar] [CrossRef] [PubMed]
- Chowdhury, T.T.; Salter, D.M.; Bader, D.L.; Lee, D.A. Integrin-mediated mechanotransduction processes in TGFbeta-stimulated monolayer-expanded chondrocytes. Biochem. Biophys Res. Commun. 2004, 318, 873–881. [Google Scholar] [CrossRef]
- Pecchi, E.; Priam, S.; Gosset, M.; Pigenet, A.; Sudre, L.; Laiguillon, M.C.; Berenbaum, F.; Houard, X. Induction of nerve growth factor expression and release by mechanical and inflammatory stimuli in chondrocytes: Possible involvement in osteoarthritis pain. Arthritis Res. Ther. 2014, 16, R16. [Google Scholar] [CrossRef]
- Pufe, T.; Lemke, A.; Kurz, B.; Petersen, W.; Tillmann, B.; Grodzinsky, A.J.; Mentlein, R. Mechanical overload induces VEGF in cartilage discs via hypoxia-inducible factor. Am. J. Pathol. 2004, 164, 185–192. [Google Scholar] [CrossRef]
- Krase, A.; Abedian, R.; Steck, E.; Hurschler, C.; Richter, W. BMP activation and Wnt-signalling affect biochemistry and functional biomechanical properties of cartilage tissue engineering constructs. Osteoarthr. Cartil. 2014, 22, 284–292. [Google Scholar] [CrossRef]
- Matsumoto, Y.; Matsumoto, K.; Irie, F.; Fukushi, J.; Stallcup, W.B.; Yamaguchi, Y. Conditional ablation of the heparan sulfate-synthesizing enzyme Ext1 leads to dysregulation of bone morphogenic protein signaling and severe skeletal defects. J. Biol. Chem. 2010, 285, 19227–19234. [Google Scholar] [CrossRef]
- Huegel, J.; Mundy, C.; Sgariglia, F.; Nygren, P.; Billings, P.C.; Yamaguchi, Y.; Koyama, E.; Pacifici, M. Perichondrium phenotype and border function are regulated by Ext1 and heparan sulfate in developing long bones: A mechanism likely deranged in Hereditary Multiple Exostoses. Dev. Biol. 2013, 377, 100–112. [Google Scholar] [CrossRef]
- Piombo, V.; Jochmann, K.; Hoffmann, D.; Wuelling, M.; Vortkamp, A. Signaling systems affecting the severity of multiple osteochondromas. Bone 2018, 111, 71–81. [Google Scholar] [CrossRef]
- Jones, K.B.; Piombo, V.; Searby, C.; Kurriger, G.; Yang, B.; Grabellus, F.; Roughley, P.J.; Morcuende, J.A.; Buckwalter, J.A.; Capecchi, M.R.; et al. A mouse model of osteochondromagenesis from clonal inactivation of Ext1 in chondrocytes. Proc. Natl. Acad. Sci. USA 2010, 107, 2054–2059. [Google Scholar] [CrossRef]
- Sgariglia, F.; Candela, M.E.; Huegel, J.; Jacenko, O.; Koyama, E.; Yamaguchi, Y.; Pacifici, M.; Enomoto-Iwamoto, M. Epiphyseal abnormalities, trabecular bone loss and articular chondrocyte hypertrophy develop in the long bones of postnatal Ext1-deficient mice. Bone 2013, 57, 220–231. [Google Scholar] [CrossRef]
- Loeser, R.F. Integrins and chondrocyte-matrix interactions in articular cartilage. Matrix Biol. J. Int. Soc. Matrix Biol. 2014, 39, 11–16. [Google Scholar] [CrossRef]
- Lucchinetti, E.; Bhargava, M.M.; Torzilli, P.A. The effect of mechanical load on integrin subunits alpha5 and beta1 in chondrocytes from mature and immature cartilage explants. Cell Tissue Res. 2004, 315, 385–391. [Google Scholar] [CrossRef]
- Olsen, O.E.; Wader, K.F.; Hella, H.; Mylin, A.K.; Turesson, I.; Nesthus, I.; Waage, A.; Sundan, A.; Holien, T. Activin A inhibits BMP-signaling by binding ACVR2A and ACVR2B. Cell Commun. Signal. Ccs 2015, 13, 27. [Google Scholar] [CrossRef]
- Vernon, L.; Abadin, A.; Wilensky, D.; Huang, C.Y.; Kaplan, L. Subphysiological compressive loading reduces apoptosis following acute impact injury in a porcine cartilage model. Sports Health 2014, 6, 81–88. [Google Scholar] [CrossRef]
- Lee, M.S.; Trindade, M.C.; Ikenoue, T.; Goodman, S.B.; Schurman, D.J.; Smith, R.L. Regulation of nitric oxide and bcl-2 expression by shear stress in human osteoarthritic chondrocytes in vitro. J. Cell. Biochem. 2003, 90, 80–86. [Google Scholar] [CrossRef]
- Loening, A.M.; James, I.E.; Levenston, M.E.; Badger, A.M.; Frank, E.H.; Kurz, B.; Nuttall, M.E.; Hung, H.H.; Blake, S.M.; Grodzinsky, A.J.; et al. Injurious mechanical compression of bovine articular cartilage induces chondrocyte apoptosis. Arch. Biochem. Biophys. 2000, 381, 205–212. [Google Scholar] [CrossRef]
- Zamora, R.; Alarcon, L.; Vodovotz, Y.; Betten, B.; Kim, P.K.; Gibson, K.F.; Billiar, T.R. Nitric oxide suppresses the expression of Bcl-2 binding protein BNIP3 in hepatocytes. J. Biol. Chem. 2001, 276, 46887–46895. [Google Scholar] [CrossRef]
- Gosset, M.; Berenbaum, F.; Thirion, S.; Jacques, C. Primary culture and phenotyping of murine chondrocytes. Nat. Protoc. 2008, 3, 1253–1260. [Google Scholar] [CrossRef] [PubMed]
- Farndale, R.; Buttle, D.; Barrett, A. Improved quantitation and discrimination of sulphated glycosaminoglycans by use of dimethylmethylene blue. Biochim. Biophys. Acta (BBA) Gen. Subj. 1986, 883, 173–177. [Google Scholar] [CrossRef]
- Tusher, V.G.; Tibshirani, R.; Chu, G. Significance analysis of microarrays applied to the ionizing radiation response. Proc. Natl. Acad. Sci. USA 2001, 98, 5116–5121. [Google Scholar] [CrossRef] [PubMed]

| Gene Symbol | Gene Name | Mean Intensities | ||
|---|---|---|---|---|
| Ctrl | Load | Fold Change | ||
| Inhba | Inhibin beta-A | 253 | 1023 | 4.04 |
| Gjb4 | Gap junction protein, beta 4 | 217 | 763 | 3.52 |
| Gprc5a | G protein-coupled receptor C, 5, a | 245 | 760 | 3.11 |
| Timp1 | Tissue inhibitor of MMPs 1 | 988 | 2518 | 2.55 |
| Cd44 | CD44 antigen | 244 | 590 | 2.42 |
| Ngf | Nerve growth factor | 336 | 811 | 2.41 |
| Plaur | PLG-activator, urokinase receptor | 179 | 409 | 2.29 |
| Srxn1 | Sulfiredoxin 1 | 224 | 469 | 2.09 |
| Dusp1 | Dual specificity phosphatase 1 | 1720 | 3590 | 2.09 |
| Fosl1 | Fos-like antigen 1 | 255 | 528 | 2.07 |
| Dhrs9 | Dehydrogenase/reductase 9 | 101 | 209 | 2.06 |
| Nt5e | 5′-nucleotidase, ecto | 593 | 1214 | 2.05 |
| Ucn | Urocortin | 151 | 283 | 1.87 |
| Itga5 | Integrin alpha 5 | 1404 | 2494 | 1.78 |
| Mean Intensities | ||||
|---|---|---|---|---|
| Gene Symbol | Gene Name | Ctrl | Load | Fold Change |
| Up-regulated | ||||
| Timp1 * | Tissue inhibitor of MMPs 1 | 988 | 2518 | 2.55 |
| Ngf * | Nerve growth factor | 336 | 811 | 2.41 |
| Dusp1 | Dual specificity phosphatase 1 | 1720 | 3590 | 2.09 |
| Ptgs2 | Prostaglandin G/H synthase 2 | 261 | 494 | 1.90 |
| Ucn | Urocortin | 151 | 283 | 1.87 |
| Smo | Smoothened, frizzled class receptor | 189 | 336 | 1.78 |
| Itga5 * | Integrin alpha 5 | 1404 | 2494 | 1.78 |
| Myc | Myc proto-oncogene protein | 263 | 452 | 1.72 |
| Spry2 | Sprouty homolog 2 | 132 | 207 | 1.56 |
| Jun | Jun proto-oncogene | 406 | 625 | 1.54 |
| Sfn | Stratifin, alias 14–3-3 protein sigma | 155 | 237 | 1.53 |
| Gdf5 | Growth differentiation factor 5 | 376 | 567 | 1.51 |
| Down-regulated | ||||
| Bnip3 | BCL2 interacting protein 3 | 1073 | 663 | −1.62 |
| Irs2 | Insulin receptor substrate 2 | 321 | 199 | −1.61 |
| Card14 | Caspase recruitment domain family, member 14 | 134 | 88 | −1.52 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gerstner, M.; Severmann, A.-C.; Chasan, S.; Vortkamp, A.; Richter, W. Heparan Sulfate Deficiency in Cartilage: Enhanced BMP-Sensitivity, Proteoglycan Production and an Anti-Apoptotic Expression Signature after Loading. Int. J. Mol. Sci. 2021, 22, 3726. https://doi.org/10.3390/ijms22073726
Gerstner M, Severmann A-C, Chasan S, Vortkamp A, Richter W. Heparan Sulfate Deficiency in Cartilage: Enhanced BMP-Sensitivity, Proteoglycan Production and an Anti-Apoptotic Expression Signature after Loading. International Journal of Molecular Sciences. 2021; 22(7):3726. https://doi.org/10.3390/ijms22073726
Chicago/Turabian StyleGerstner, Matthias, Ann-Christine Severmann, Safak Chasan, Andrea Vortkamp, and Wiltrud Richter. 2021. "Heparan Sulfate Deficiency in Cartilage: Enhanced BMP-Sensitivity, Proteoglycan Production and an Anti-Apoptotic Expression Signature after Loading" International Journal of Molecular Sciences 22, no. 7: 3726. https://doi.org/10.3390/ijms22073726
APA StyleGerstner, M., Severmann, A.-C., Chasan, S., Vortkamp, A., & Richter, W. (2021). Heparan Sulfate Deficiency in Cartilage: Enhanced BMP-Sensitivity, Proteoglycan Production and an Anti-Apoptotic Expression Signature after Loading. International Journal of Molecular Sciences, 22(7), 3726. https://doi.org/10.3390/ijms22073726







