Isolation and Functional Characterization of a Salt-Responsive Calmodulin-Like Gene MpCML40 from Semi-Mangrove Millettia pinnata
Abstract
1. Introduction
2. Results
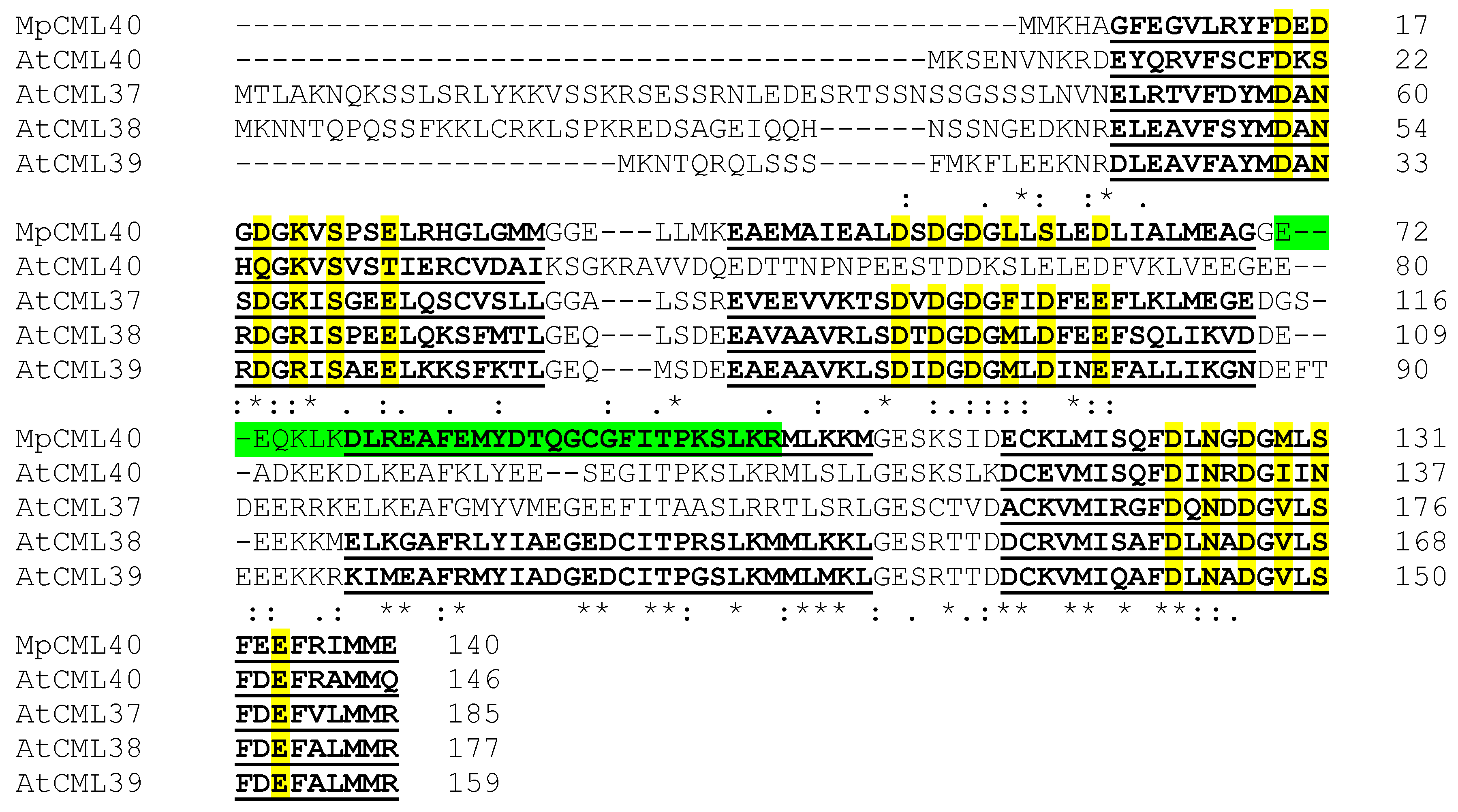
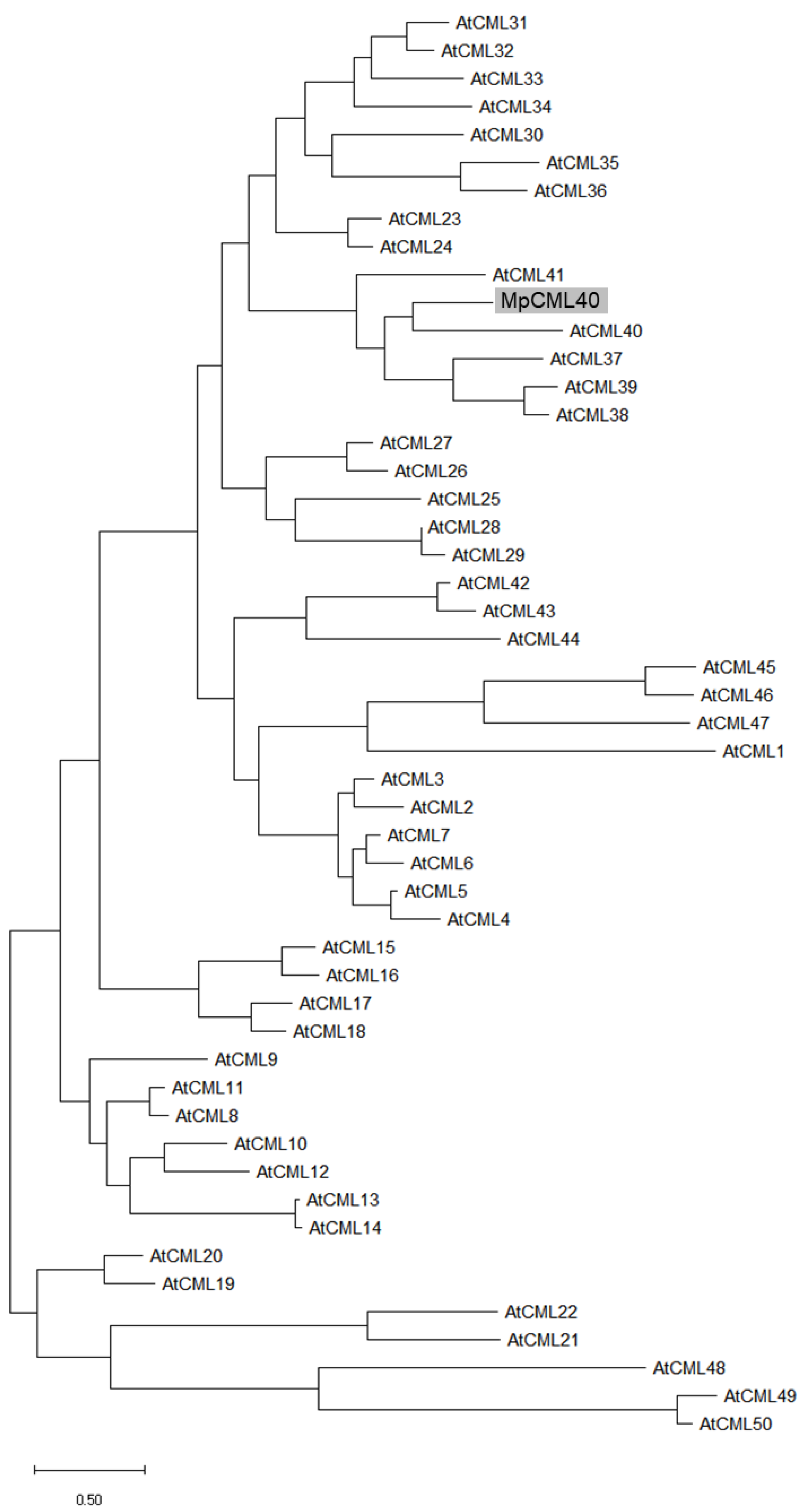
2.1. MpCML40 Is an EF-Hand Motif-Containing Calmodulin-Like Protein
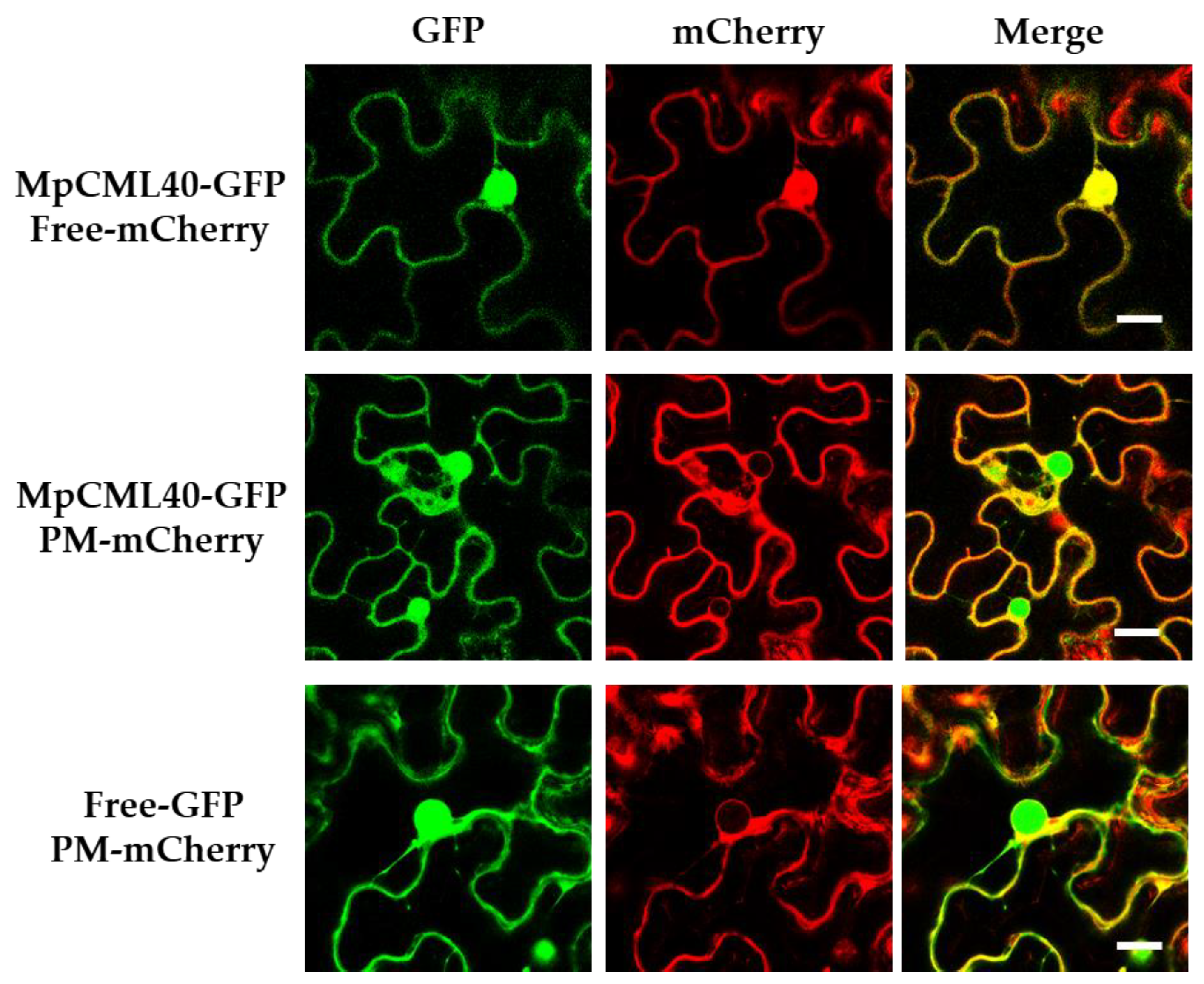
2.2. MpCML40 Localizes at the Plasma Membrane and in the Nucleus
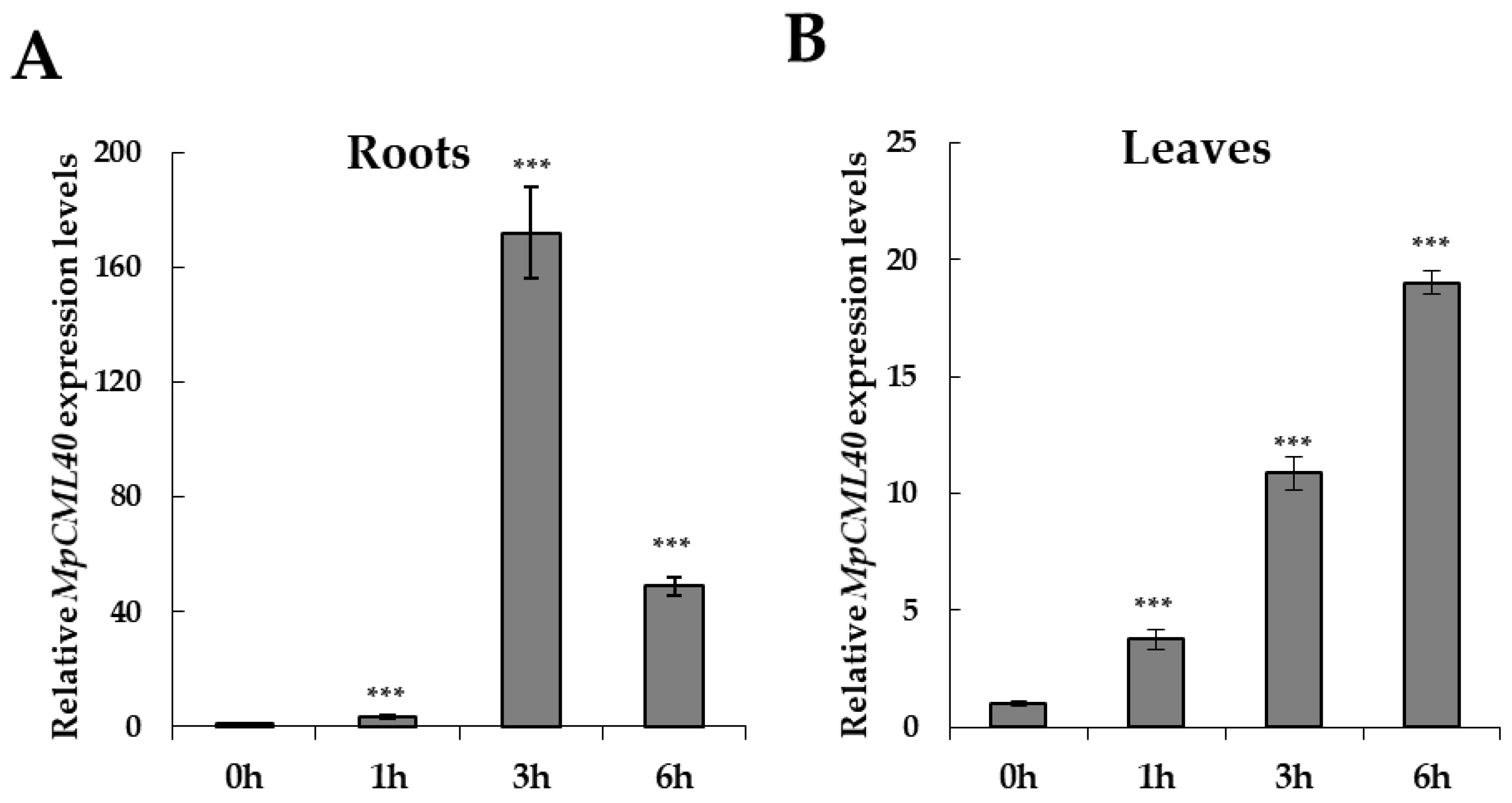
2.3. MpCML40 Gene Is Highly Induced by Salt Stress in Pongamia Roots
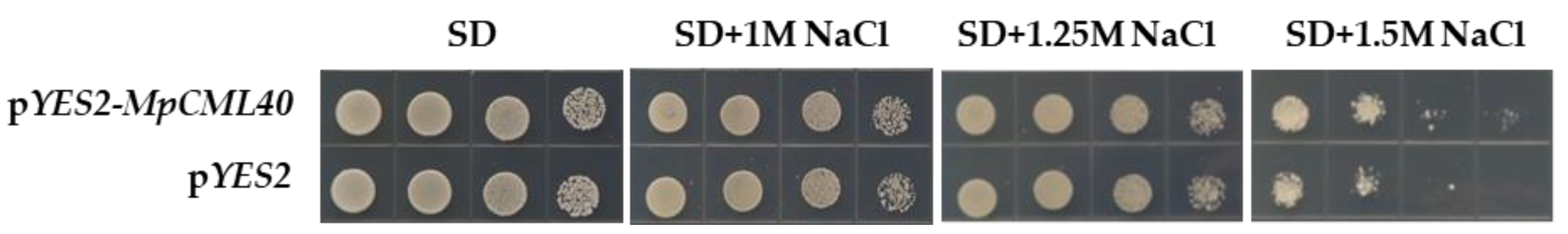
2.4. The pYES22-MpCML40-Transformed Yeast Has Enhanced Salt Tolerance
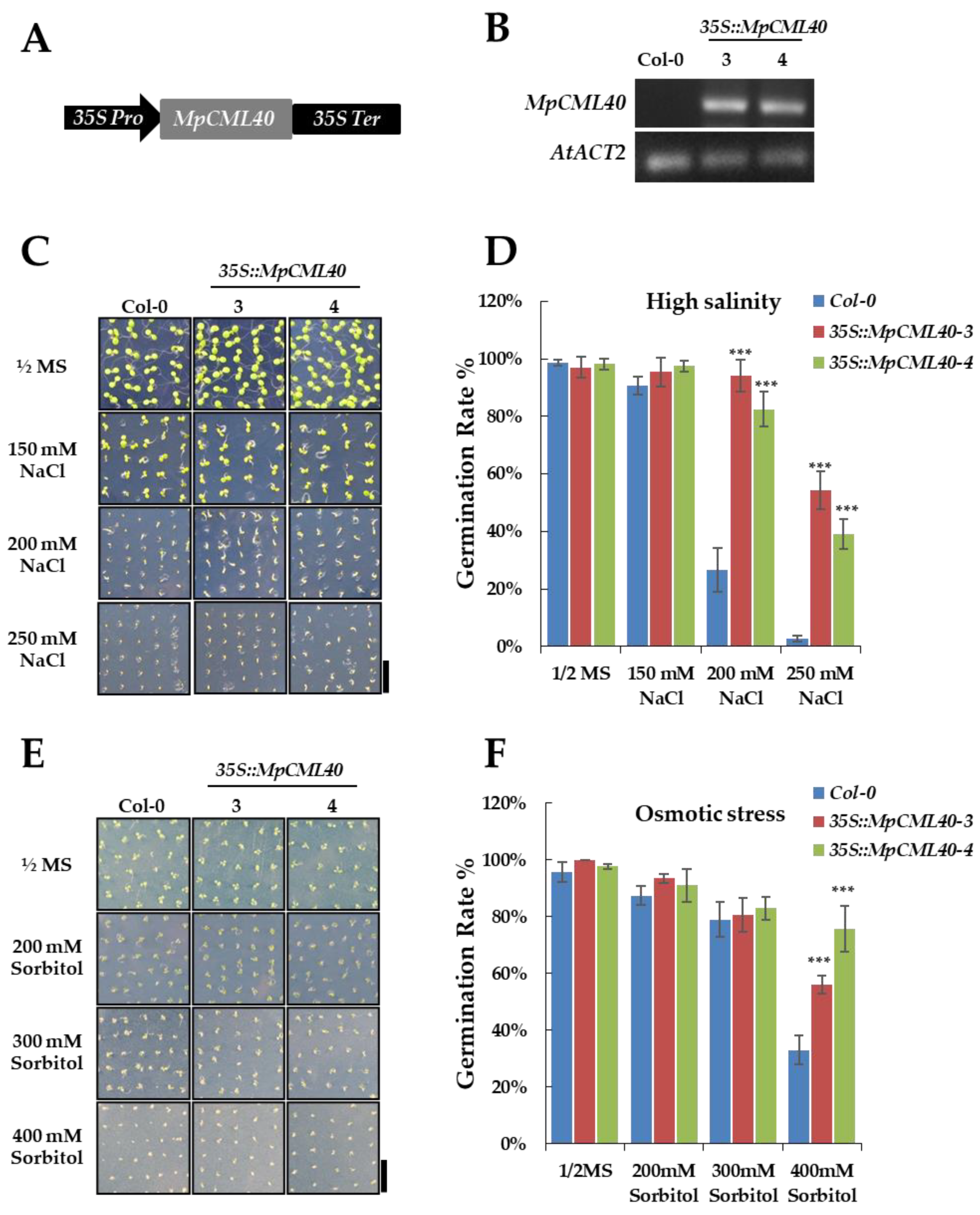
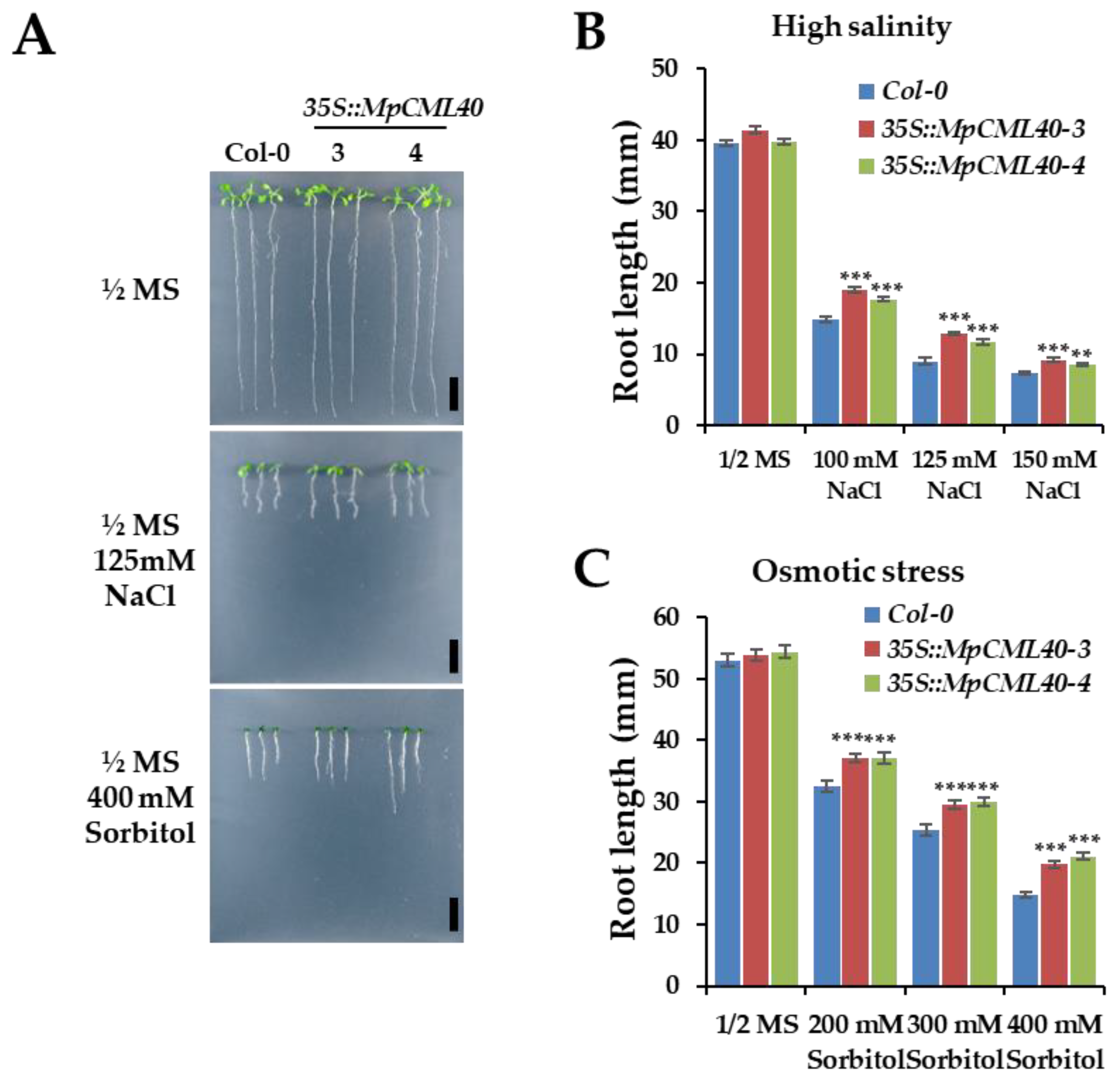
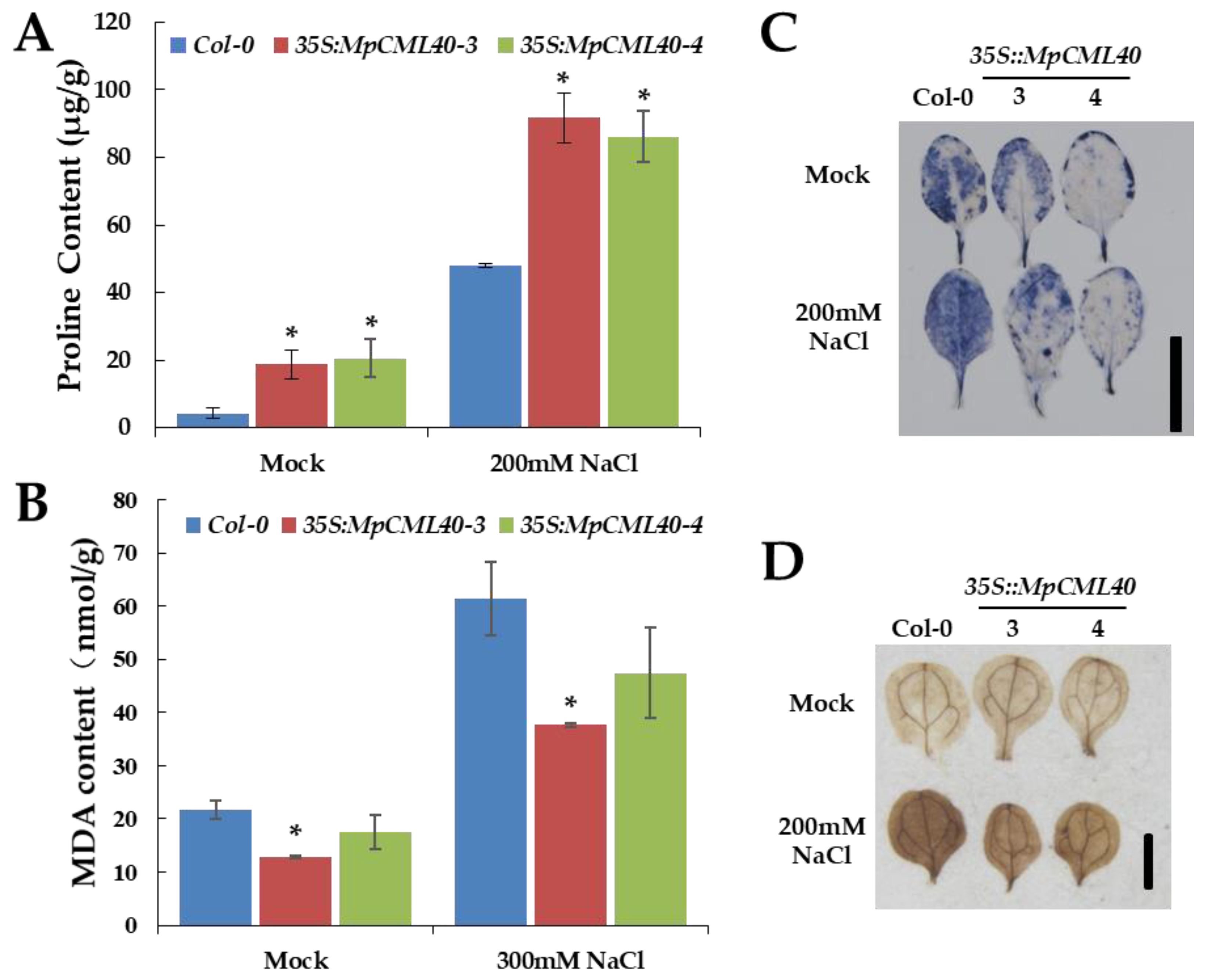
2.5. Heterologous Expression of MpCML40 in Arabidopsis Strongly Enhances Salt and Osmotic Tolerance
3. Discussion
3.1. MpCML40 Is a Salt-Induced CML Gene
3.2. MpCML40 Improves Salt Tolerance in Both Yeast and Arabidopsis
4. Materials and Methods
4.1. Plant Materials and Growth Conditions
4.2. Full-Length cDNA Cloning, Phylogenetic Analysis, and Motif Prediction
4.3. Subcellular Localization
4.4. Quantitative Real-Time PCR
4.5. Yeast Transformation and Growth Assay
4.6. Phenotype Analysis of Wild-Type and 35S::MpCML40 Transgenic Arabidopsis Plants
4.7. Proline Content Measurement
4.8. MDA Content Measurement
4.9. DAB and NBT Staining
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Munns, R.; Tester, M. Mechanisms of salinity tolerance. Annu. Rev. Plant Biol. 2008, 59, 651–681. [Google Scholar] [CrossRef]
- Flowers, T.J. Improving crop salt tolerance. J. Exp. Bot. 2004, 55, 307–319. [Google Scholar] [CrossRef]
- Wang, L.; Mu, M.; Li, X.; Lin, P.; Wang, W. Differentiation between true mangroves and mangrove associates based on leaf traits and salt contents. J. Plant Ecol. 2010, 4, 292–301. [Google Scholar] [CrossRef]
- Choudhury, R.R.; Basak, S.; Ramesh, A.M.; Rangan, L. Nuclear DNA content of Pongamia pinnata L. and genome size stability of in vitro-regenerated plantlets. Protoplasma 2014, 251, 703–709. [Google Scholar] [CrossRef]
- Kaul, S.; Koo, H.L.; Jenkins, J.; Rizzo, M.; Rooney, T.; Tallon, L.J.; Feldblyum, T.; Nierman, W.; Benito, M.I.; Lin, X.; et al. Analysis of the genome sequence of the flowering plant Arabidopsis thaliana. Nature 2000, 408, 796–815. [Google Scholar]
- Wegrzyn, J.L.; Whalen, J.; Kinlaw, C.S.; Harry, D.E.; Puryear, J.; Loopstra, C.A.; Gonzalez-Ibeas, D.; Vasquez-Gross, H.A.; Famula, R.A.; Neale, D.B. Transcriptomic profile of leaf tissue from the leguminous tree, Millettia pinnata. Tree Genet. Genomes 2016, 12, 44. [Google Scholar] [CrossRef]
- Huang, J.; Lu, X.; Yan, H.; Chen, S.; Zhang, W.; Huang, R.; Zheng, Y. Transcriptome characterization and sequencing-based identification of salt-responsive genes in Millettia pinnata, a semi-mangrove plant. DNA Res. 2012, 19, 195–207. [Google Scholar] [CrossRef] [PubMed]
- Sreeharsha, R.V.; Mudalkar, S.; Singha, K.T.; Reddy, A.R. Unravelling molecular mechanisms from floral initiation to lipid biosynthesis in a promising biofuel tree species, Pongamia pinnata using transcriptome analysis. Sci. Rep. 2016, 6, 34315. [Google Scholar] [CrossRef]
- Liu, S.J.; Song, S.H.; Wang, W.Q.; Song, S.Q. De novo assembly and characterization of germinating lettuce seed transcriptome using Illumina paired-end sequencing. Plant Physiol. Biochem. 2015, 96, 154–162. [Google Scholar] [CrossRef]
- Huang, J.; Hao, X.; Jin, Y.; Guo, X.; Shao, Q.; Kumar, K.S.; Ahlawat, Y.K.; Harry, D.E.; Joshi, C.P.; Zheng, Y. Temporal transcriptome profiling of developing seeds reveals a concerted gene regulation in relation to oil accumulation in Pongamia (Millettia pinnata). BMC Plant Biol. 2018, 18, 140. [Google Scholar] [CrossRef] [PubMed]
- Huang, J.Z.; Guo, X.H.; Hao, X.H.; Zhang, W.K.; Chen, S.Y.; Huang, R.F.; Gresshoff, P.M.; Zheng, Y.Z. De novo sequencing and characterization of seed transcriptome of the tree legume Millettia pinnata for gene discovery and SSR marker development. Mol. Breed. 2016, 36, 75. [Google Scholar] [CrossRef]
- Winarto, H.P.; Liew, L.C.; Gresshoff, P.M.; Scott, P.T.; Singh, M.B.; Bhalla, P.L. Isolation and Characterization of Circadian Clock Genes in the Biofuel Plant Pongamia (Millettia pinnata). Bioenergy Res. 2015, 8, 760–774. [Google Scholar] [CrossRef]
- Ramesh, A.M.; Kesari, V.; Rangan, L. Characterization of a stearoyl-acyl carrier protein desaturase gene from potential biofuel plant, Pongamia pinnata L. Gene 2014, 542, 113–121. [Google Scholar] [CrossRef] [PubMed]
- Moolam, R.A.; Singh, A.; Shelke, R.G.; Scott, P.T.; Gresshoff, P.M.; Rangan, L. Identification of two genes encoding microsomal oleate desaturases (FAD2) from the biodiesel plant Pongamia pinnata L. Trees Struct. Funct. 2016, 30, 1351–1360. [Google Scholar] [CrossRef]
- Wang, H.; Hu, T.; Huang, J.; Lu, X.; Huang, B.; Zheng, Y. The expression of Millettia pinnata chalcone isomerase in Saccharomyces cerevisiae salt-sensitive mutants enhances salt-tolerance. Int. J. Mol. Sci. 2013, 14, 8775–8786. [Google Scholar] [CrossRef]
- Zielinski, R.E. Calmodulin and Calmodulin-Binding Proteins in Plants. Annu. Rev. Plant Physiol. Plant Mol. Biol. 1998, 49, 697–725. [Google Scholar] [CrossRef]
- DeFalco, T.A.; Bender, K.W.; Snedden, W.A. Breaking the code: Ca2+ sensors in plant signalling. Biochem. J. 2010, 425, 27–40. [Google Scholar] [CrossRef] [PubMed]
- Bender, K.W.; Snedden, W.A. Calmodulin-Related Proteins Step Out from the Shadow of Their Namesake. Plant Physiol. 2013, 163, 486–495. [Google Scholar] [CrossRef]
- Beckmann, L.; Edel, K.H.; Batistic, O.; Kudla, J. A calcium sensor—Protein kinase signaling module diversified in plants and is retained in all lineages of Bikonta species. Sci. Rep. 2016, 6, 31645. [Google Scholar] [CrossRef]
- Kretsinger, R.H.; Nockolds, C.E. Carp muscle calcium-binding protein. II. Structure determination and general description. J. Biol. Chem. 1973, 248, 3313–3326. [Google Scholar] [CrossRef]
- Lewit-Bentley, A.; Rety, S. EF-hand calcium-binding proteins. Curr. Opin. Struct. Biol. 2000, 10, 637–643. [Google Scholar] [CrossRef]
- McCormack, E.; Braam, J. Calmodulins and related potential calcium sensors of Arabidopsis. New Phytol. 2003, 159, 585–598. [Google Scholar] [CrossRef]
- Perochon, A.; Aldon, D.; Galaud, J.P.; Ranty, B. Calmodulin and calmodulin-like proteins in plant calcium signaling. Biochimie 2011, 93, 2048–2053. [Google Scholar] [CrossRef]
- Zeng, H.; Xu, L.; Singh, A.; Wang, H.; Du, L.; Poovaiah, B.W. Involvement of calmodulin and calmodulin-like proteins in plant responses to abiotic stresses. Front. Plant Sci. 2015, 6, 600. [Google Scholar] [CrossRef] [PubMed]
- Braam, J. Regulated expression of the calmodulin-related TCH genes in cultured Arabidopsis cells: Induction by calcium and heat shock. Proc. Natl. Acad. Sci. USA 1992, 89, 3213–3216. [Google Scholar] [CrossRef]
- Vanderbeld, B.; Snedden, W.A. Developmental and stimulus-induced expression patterns of Arabidopsis calmodulin-like genes CML37, CML38 and CML39. Plant Mol. Biol. 2007, 64, 683–697. [Google Scholar] [CrossRef] [PubMed]
- Yamaguchi, T.; Aharon, G.S.; Sottosanto, J.B.; Blumwald, E. Vacuolar Na+/H+ antiporter cation selectivity is regulated by calmodulin from within the vacuole in a Ca2+- and pH-dependent manner. Proc. Natl. Acad. Sci. USA 2005, 102, 16107–16112. [Google Scholar] [CrossRef]
- Magnan, F.; Ranty, B.; Charpenteau, M.; Sotta, B.; Galaud, J.P.; Aldon, D. Mutations in AtCML9, a calmodulin-like protein from Arabidopsis thaliana, alter plant responses to abiotic stress and abscisic acid. Plant J. 2008, 56, 575–589. [Google Scholar] [CrossRef] [PubMed]
- Delk, N.A.; Johnson, K.A.; Chowdhury, N.I.; Braam, J. CML24, regulated in expression by diverse stimuli, encodes a potential Ca2+ sensor that functions in responses to abscisic acid, daylength, and ion stress. Plant Physiol. 2005, 139, 240–253. [Google Scholar] [CrossRef]
- Wu, X.; Qiao, Z.; Liu, H.; Acharya, B.R.; Li, C.; Zhang, W. CML20, an Arabidopsis Calmodulin-like Protein, Negatively Regulates Guard Cell ABA Signaling and Drought Stress Tolerance. Front. Plant Sci. 2017, 8, 824. [Google Scholar] [CrossRef] [PubMed]
- Vadassery, J.; Reichelt, M.; Hause, B.; Gershenzon, J.; Boland, W.; Mithofer, A. CML42-mediated calcium signaling coordinates responses to Spodoptera herbivory and abiotic stresses in Arabidopsis. Plant Physiol. 2012, 159, 1159–1175. [Google Scholar] [CrossRef]
- Xu, G.Y.; Rocha, P.S.; Wang, M.L.; Xu, M.L.; Cui, Y.C.; Li, L.Y.; Zhu, Y.X.; Xia, X. A novel rice calmodulin-like gene, OsMSR2, enhances drought and salt tolerance and increases ABA sensitivity in Arabidopsis. Planta 2011, 234, 47–59. [Google Scholar] [CrossRef] [PubMed]
- Dubrovina, A.S.; Aleynova, O.A.; Ogneva, Z.V.; Suprun, A.R.; Ananev, A.A.; Kiselev, K.V. The Effect of Abiotic Stress Conditions on Expression of Calmodulin (CaM) and Calmodulin-Like (CML) Genes in Wild-Growing Grapevine Vitis amurensis. Plants 2019, 8, 602. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.; Sun, X.; Duanmu, H.; Zhu, D.; Yu, Y.; Cao, L.; Liu, A.; Jia, B.; Xiao, J.; Zhu, Y. GsCML27, a Gene Encoding a Calcium-Binding Ef-Hand Protein from Glycine soja, Plays Differential Roles in Plant Responses to Bicarbonate, Salt and Osmotic Stresses. PLoS ONE 2015, 10, e0141888. [Google Scholar] [CrossRef]
- Zhang, X.; Wang, T.; Liu, M.; Sun, W.; Zhang, W.-H. Calmodulin-like gene MtCML40 is involved in salt tolerance by regulating MtHKTs transporters in Medicago truncatula. Environ. Exp. Bot. 2019, 157, 79–90. [Google Scholar] [CrossRef]
- Ma, Q.; Zhou, Q.; Chen, C.; Cui, Q.; Zhao, Y.; Wang, K.; Arkorful, E.; Chen, X.; Sun, K.; Li, X. Isolation and expression analysis of CsCML genes in response to abiotic stresses in the tea plant (Camellia sinensis). Sci. Rep. 2019, 9, 8211. [Google Scholar] [CrossRef]
- Rigden, D.J.; Galperin, M.Y. The DxDxDG motif for calcium binding: Multiple structural contexts and implications for evolution. J. Mol. Biol. 2004, 343, 971–984. [Google Scholar] [CrossRef]
- Zhou, Y.; Xue, S.; Yang, J.J. Calciomics: Integrative studies of Ca2+-binding proteins and their interactomes in biological systems. Metallomics 2013, 5, 29–42. [Google Scholar] [CrossRef] [PubMed]
- Kosugi, S.; Hasebe, M.; Tomita, M.; Yanagawa, H. Systematic identification of cell cycle-dependent yeast nucleocytoplasmic shuttling proteins by prediction of composite motifs. Proc. Natl. Acad. Sci. USA 2009, 106, 10171–10176. [Google Scholar] [CrossRef] [PubMed]
- Kosugi, S.; Hasebe, M.; Matsumura, N.; Takashima, H.; Miyamoto-Sato, E.; Tomita, M.; Yanagawa, H. Six classes of nuclear localization signals specific to different binding grooves of importin alpha. J. Biol. Chem. 2009, 284, 478–485. [Google Scholar] [CrossRef] [PubMed]
- Kosugi, S.; Hasebe, M.; Entani, T.; Takayama, S.; Tomita, M.; Yanagawa, H. Design of peptide inhibitors for the importin alpha/beta nuclear import pathway by activity-based profiling. Chem. Biol. 2008, 15, 940–949. [Google Scholar] [CrossRef]
- Boonburapong, B.; Buaboocha, T. Genome-wide identification and analyses of the rice calmodulin and related potential calcium sensor proteins. BMC Plant Biol. 2007, 7, 4. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Sun, Q.; Yu, S.; Guo, Z. Calmodulin-Like (CML) Gene Family in Medicago truncatula: Genome-Wide Identification, Characterization and Expression Analysis. Int. J. Mol. Sci. 2020, 21, 7142. [Google Scholar] [CrossRef] [PubMed]
- Munir, S.; Khan, M.R.; Song, J.; Munir, S.; Zhang, Y.; Ye, Z.; Wang, T. Genome-wide identification, characterization and expression analysis of calmodulin-like (CML) proteins in tomato (Solanum lycopersicum). Plant Physiol. Biochem. 2016, 102, 167–179. [Google Scholar] [CrossRef] [PubMed]
- Shi, J.; Du, X. Identification, characterization and expression analysis of calmodulin and calmodulin-like proteins in Solanum pennellii. Sci. Rep. 2020, 10, 7474. [Google Scholar] [CrossRef]
- Zhang, Q.; Liu, X.; Liu, X.; Wang, J.; Yu, J.; Hu, D.; Hao, Y. Genome-Wide Identification, Characterization, and Expression Analysis of Calmodulin-Like Proteins (CMLs) in Apple. Hortic. Plant J. 2017, 3, 219–231. [Google Scholar] [CrossRef]
- Zhu, X.; Dunand, C.; Snedden, W.; Galaud, J.P. CaM and CML emergence in the green lineage. Trends Plant Sci. 2015, 20, 483–489. [Google Scholar] [CrossRef]
- Marriboina, S.; Sengupta, D.; Kumar, S.; Reddy, A.R. Physiological and molecular insights into the high salinity tolerance of Pongamia pinnata (L.) pierre, a potential biofuel tree species. Plant Sci. 2017, 258, 102–111. [Google Scholar] [CrossRef]
- Mishra, A.; Tanna, B. Halophytes: Potential resources for salt stress tolerance genes and promoters. Front. Plant Sci. 2017, 8, 829. [Google Scholar] [CrossRef]
- Wang, W.; Vinocur, B.; Altman, A. Plant responses to drought, salinity and extreme temperatures: Towards genetic engineering for stress tolerance. Planta 2003, 218, 1–14. [Google Scholar] [CrossRef]
- Mahajan, S.; Tuteja, N. Cold, salinity and drought stresses: An overview. Arch. Biochem. Biophys. 2005, 444, 139–158. [Google Scholar] [CrossRef] [PubMed]
- Liang, W.; Ma, X.; Wan, P.; Liu, L. Plant salt-tolerance mechanism: A review. Biochem. Biophys. Res. Commun. 2018, 495, 286–291. [Google Scholar] [CrossRef] [PubMed]
- Jones, D.T.; Taylor, W.R.; Thornton, J.M. The rapid generation of mutation data matrices from protein sequences. Comput. Appl. Biosci. 1992, 8, 275–282. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X: Molecular Evolutionary Genetics Analysis across Computing Platforms. Mol. Biol. Evol. 2018, 35, 1547–1549. [Google Scholar] [CrossRef] [PubMed]
- Madeira, F.; Park, Y.M.; Lee, J.; Buso, N.; Gur, T.; Madhusoodanan, N.; Basutkar, P.; Tivey, A.R.N.; Potter, S.C.; Finn, R.D.; et al. The EMBL-EBI search and sequence analysis tools APIs in 2019. Nucleic Acids Res. 2019, 47, W636–W641. [Google Scholar] [CrossRef]
- Letunic, I.; Khedkar, S.; Bork, P. SMART: Recent updates, new developments and status in 2020. Nucleic Acids Res. 2020, 49, D458–D460. [Google Scholar] [CrossRef]
- Letunic, I.; Bork, P. 20 years of the SMART protein domain annotation resource. Nucleic Acids Res. 2018, 46, D493–D496. [Google Scholar] [CrossRef]


| 1 | CCACAACATA TAATAACAAC TCAATTTTCC ATTTGCATAC AAGTTACATT TTCTCCTTCT |
| 61 | TATTCTTCTT GTTATTGTGT ACATTAAAGA TTTGAACAAA TTACACTACA CCTTTAAGAT |
| 121 | AAGGAGCATT GTAACATGAT GAAGCATGCG GGTTTTGAGG GTGTTCTTCG ATATTTTGAT |
| 181 | GAAGATGGGG ATGGAAAGGT TTCACCTTCA GAGTTAAGGC ATGGATTGGG AATGATGGGT |
| 241 | GGGGAGCTTT TGATGAAAGA AGCAGAGATG GCAATTGAGG CACTGGATTC TGATGGTGAT |
| 301 | GGGTTGTTGA GTTTGGAGGA TTTGATTGCT TTGATGGAAG CAGGGGGAGA GGAACAAAAG |
| 361 | TTGAAGGATT TGAGAGAAGC TTTTGAGATG TATGACACTC AAGGGTGCGG ATTTATAACC |
| 421 | CCAAAGAGCT TGAAGAGGAT GCTTAAGAAG ATGGGAGAGT CCAAGTCCAT TGATGAATGC |
| 481 | AAATTGATGA TTAGTCAATT TGATTTGAAT GGGGATGGGA TGCTTAGCTT TGAAGAATTC |
| 541 | AGAATTATGA TGGAGTGAGG CCAGTATATT TGTTGATGAT ATTGTTTAGT TTGTTTGTTT |
| 601 | GTTTGGGAGG AAGAGGGGTA TAGTTAAGTG GATTTGATTT ATTTGTTTGC AGTTTGCACA |
| 661 | TGTATAAATA ACTCCTTTTG TGCTTTGCAA TACTTTTGAC AATTGATTAA CTGTTAGATT |
| 721 | TCTCCCAAGT TCCTACATAA AAAATTATTC AAATTTTCTT AATGGGAGTT GTATTATGAC |
| 781 | TATTATCATG GTTAAATATA TTTTTTATTT CGTTCCAAAA AAAAAAAAAA AAAAAAAAAA |
| Primer | Primer Sequence (5′ → 3′) |
|---|---|
| MpActin.RtF | AGAGCAGTTCTTCAGTTGAG |
| MpActin.RtR | TCCTCCAATCCAGACACTAT |
| MpCML40.RtF | GCACTGGATTCTGATGGTGATGGG |
| MpCML40.RtR | GCTCTTTGGGGTTATAAATCCGCA |
| MpCML40.3’GSP | TGGCAATTGAGGCACTGGATTCTGATGG |
| MpCML40.3’NGSP | ATGGAAGCAGGGGGAGAGGAACAA |
| MpCML40.5’GSP | TCCGCACCCTTGAGTGTCATACATCTCA |
| MpCML40.5’NGSP | CCTCCAAACTCAACAACCCATCACCATC |
| MpCML40.yeastF | CGCGGATCCATGATGAAGCATGCGGGTTTTGAGG |
| MpCML40.yeastR | CCGCTCGAGTCACTCCATCATAATTCTGAA |
| MpCML40.plantF | TATGGCGCGCCCATGATGAAGCATGCGGGTTTTGA |
| MpCML40.plantR | CTCCATCATAATTCTGAATTCTT |
| AtACT2.RtF | GACCTTTAACTCTCCCGCTATG |
| AtACT2.RtR | GAGACACACCATCACCAGAAT |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhang, Y.; Huang, J.; Hou, Q.; Liu, Y.; Wang, J.; Deng, S. Isolation and Functional Characterization of a Salt-Responsive Calmodulin-Like Gene MpCML40 from Semi-Mangrove Millettia pinnata. Int. J. Mol. Sci. 2021, 22, 3475. https://doi.org/10.3390/ijms22073475
Zhang Y, Huang J, Hou Q, Liu Y, Wang J, Deng S. Isolation and Functional Characterization of a Salt-Responsive Calmodulin-Like Gene MpCML40 from Semi-Mangrove Millettia pinnata. International Journal of Molecular Sciences. 2021; 22(7):3475. https://doi.org/10.3390/ijms22073475
Chicago/Turabian StyleZhang, Yi, Jianzi Huang, Qiongzhao Hou, Yujuan Liu, Jun Wang, and Shulin Deng. 2021. "Isolation and Functional Characterization of a Salt-Responsive Calmodulin-Like Gene MpCML40 from Semi-Mangrove Millettia pinnata" International Journal of Molecular Sciences 22, no. 7: 3475. https://doi.org/10.3390/ijms22073475
APA StyleZhang, Y., Huang, J., Hou, Q., Liu, Y., Wang, J., & Deng, S. (2021). Isolation and Functional Characterization of a Salt-Responsive Calmodulin-Like Gene MpCML40 from Semi-Mangrove Millettia pinnata. International Journal of Molecular Sciences, 22(7), 3475. https://doi.org/10.3390/ijms22073475






