Transglycosylation Activity of Engineered Bifidobacterium Lacto-N-Biosidase Mutants at Donor Subsites for Lacto-N-Tetraose Synthesis
Abstract
1. Introduction
2. Results
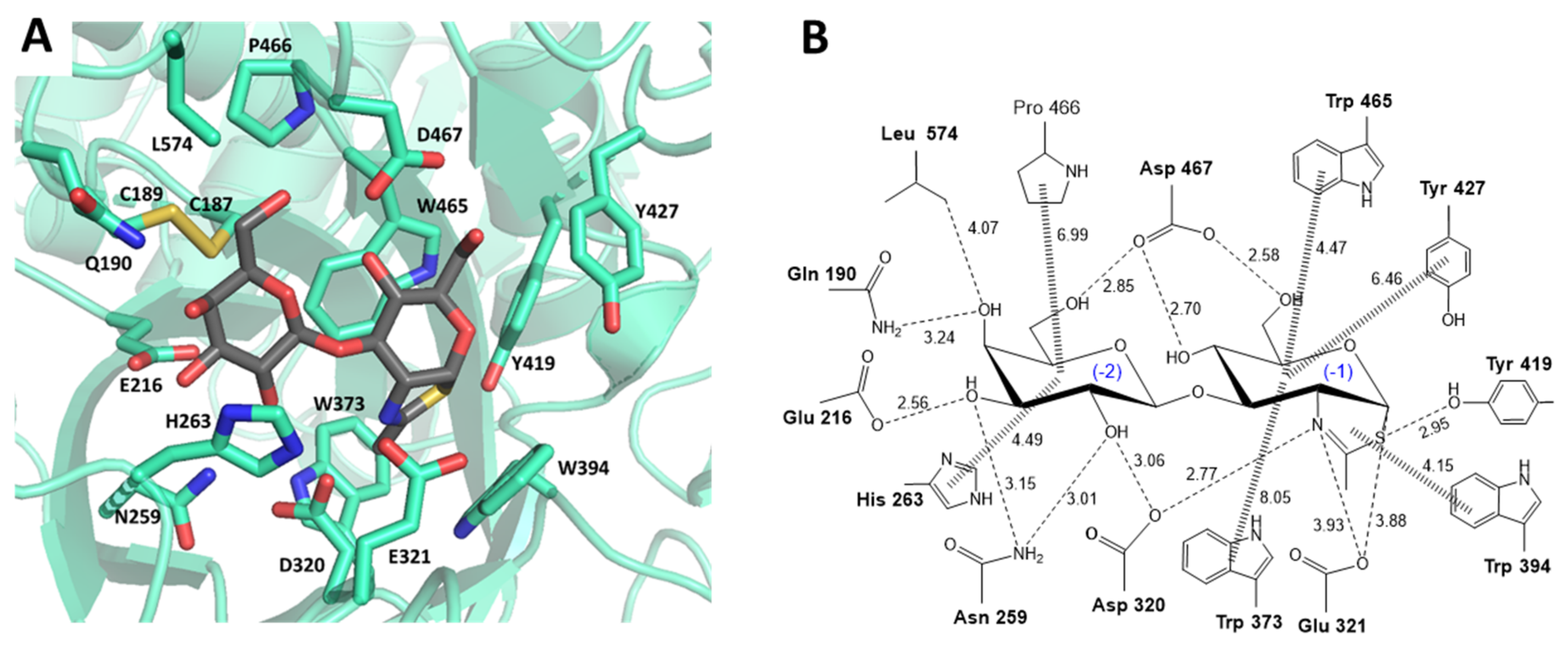
2.1. LnbB Mutants Design
2.2. Hydrolase Activity of LnbB Mutants
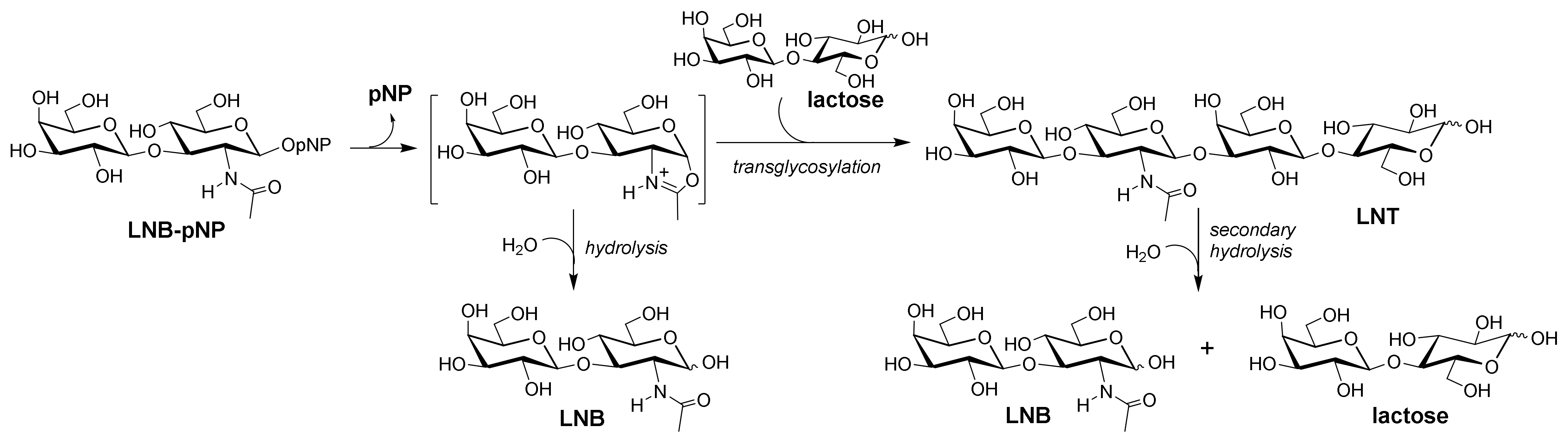
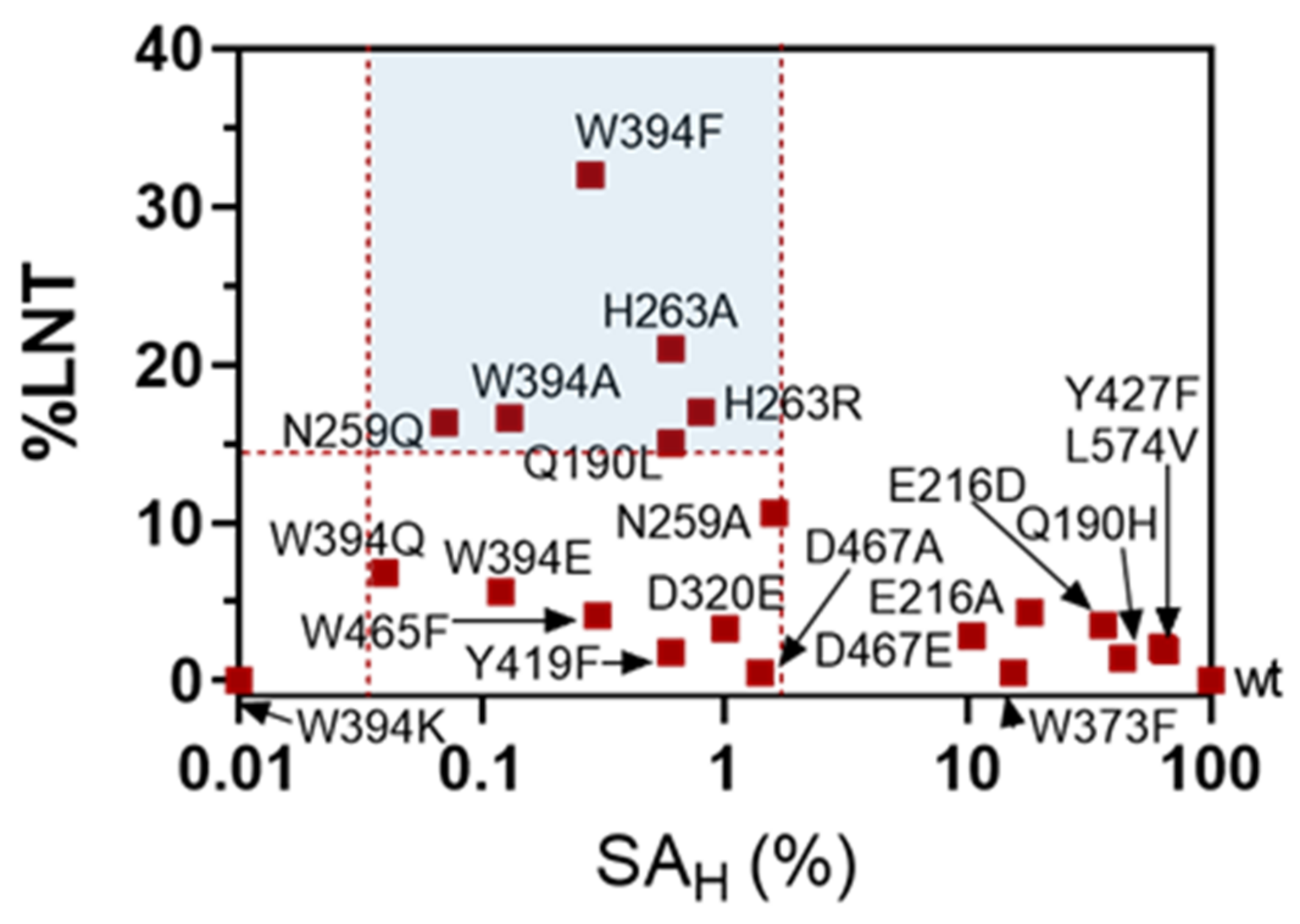
2.3. Formation of LNT by Transglycosylating LnbB Mutants
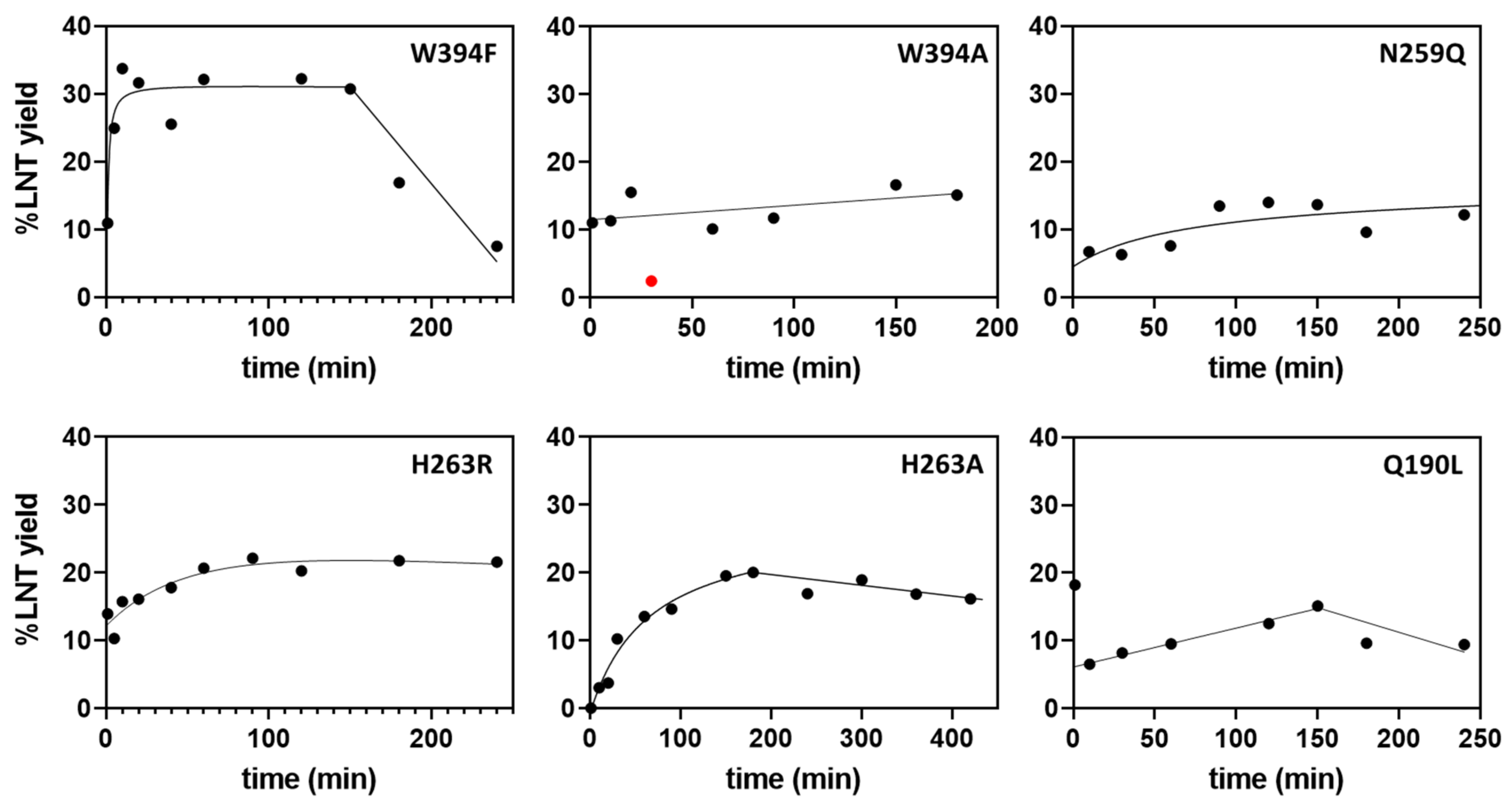
2.4. Transglycosylation Activity of Selected Mutants
2.5. Kinetic Parameters for Donor Substrate in Hydrolysis and Transglycosylation Modes
2.6. Regioselectivity of Tetrasaccharide Formation by LnbB W394F Mutant
3. Discussion
3.1. Assisting Residue
3.2. Hydrophobic Platform
3.3. Catalytic Tyrosine
3.4. Mutations in Subsite -2
4. Materials and Methods
4.1. Substrates
4.2. Mutants Preparation by Site Directed Mutagenesis
4.3. Expression and Purification of Lacto-N-Biosidase Mutants
4.4. Hydrolase Activity Assay
4.5. Transglycosylation Activity Assay
4.6. Enzyme Kinetics
4.7. W394F Thermal Stability
4.8. Regioselectivity of Glycoside Bond Formation by W394A
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Faijes, M.; Castejón-Vilatersana, M.; Val-Cid, C.; Planas, A. Enzymatic and cell factory approaches to the production of human milk oligosaccharides. Biotechnol. Adv. 2019, 37, 667–697. [Google Scholar] [CrossRef]
- Bych, K.; Mikš, M.H.; Johanson, T.; Hederos, M.J.; Vigsnæs, L.K.; Becker, P. Production of HMOs using microbial hosts—From cell engineering to large scale production. Curr. Opin. Biotechnol. 2019, 56, 130–137. [Google Scholar] [CrossRef] [PubMed]
- Zeuner, B.; Teze, D.; Muschiol, J.; Meyer, A.S. Synthesis of Human Milk Oligosaccharides: Protein Engineering Strategies for Improved Enzymatic Transglycosylation. Molecules 2019, 24, 2033. [Google Scholar] [CrossRef] [PubMed]
- Chen, X. Human Milk Oligosaccharides (HMOS): Structure, Function, and Enzyme-Catalyzed Synthesis. In Advances in Carbohydrate Chemistry and Biochemistry; Academic Press: Cambridge, MA, USA, 2015; Volume 72, pp. 113–190. ISBN 9780128021415. [Google Scholar]
- Walsh, C.; Lane, J.A.; Van Sinderen, D.; Hickey, R.M. From lab bench to formulated ingredient: Characterization, production, and commercialization of human milk oligosaccharides. J. Funct. Foods 2020, 72, 104052. [Google Scholar] [CrossRef]
- Triantis, V.; Bode, L.; van Neerven, J.R.J. Immunological effects of human milk oligosaccharides. Front. Pediatr. 2018, 6, 190. [Google Scholar] [CrossRef]
- Kunz, C.; Kuntz, S.; Rudloff, S. Bioactivity of Human Milk Oligosaccharides. In Food Oligosaccharides: Production, Analysis and Bioactivity; Moreno, F.J., Sanz, M.L., Eds.; John Wiley & Sons, Ltd.: Chichester, UK, 2014; Volume 9781118426, pp. 5–20. ISBN 9781118817360. [Google Scholar]
- Bode, L.; Contractor, N.; Barile, D.; Pohl, N.; Prudden, A.R.; Boons, G.J.; Jin, Y.S.; Jennewein, S. Overcoming the limited availability of human milk oligosaccharides: Challenges and opportunities for research and application. Nutr. Rev. 2016, 74, 635–644. [Google Scholar] [CrossRef]
- Baumgärtner, F.; Conrad, J.; Sprenger, G.A.; Albermann, C. Synthesis of the Human Milk Oligosaccharide Lacto-N-Tetraose in Metabolically Engineered, Plasmid-Free E. coli. ChemBioChem 2014, 15, 1896–1900. [Google Scholar] [CrossRef]
- Baumgärtner, F.; Sprenger, G.A.; Albermann, C. Enzyme and Microbial Technology Galactose-limited fed-batch cultivation of Escherichia coli for the production of lacto-N-tetraose. Enzym. Microb. Technol. 2015, 75–76, 37–43. [Google Scholar] [CrossRef] [PubMed]
- Craft, K.M.; Townsend, S.D. Synthesis of lacto-N-tetraose. Carbohydr. Res. 2017, 440–441, 43–50. [Google Scholar] [CrossRef] [PubMed]
- Bandara, M.D.; Stine, K.J.; Demchenko, A.V. The chemical synthesis of human milk oligosaccharides: Lacto-N-tetraose (Galβ1→3GlcNAcβ1→3Galβ1→4Glc). Carbohydr. Res. 2019, 486, 107824. [Google Scholar] [CrossRef]
- Blixt, O.; Van Die, I.; Norberg, T.; Van Den Eijnden, D.H. High-level expression of the Neisseria meningitidis lgtA gene in Escherichia coli and characterization of the encoded N-acetylglucosaminyltransferase as a useful catalyst in the synthesis of GlcNAcβ1→3Gal and GalNAcβ1→3Gal linkages. Glycobiology 1999, 9, 1061–1071. [Google Scholar] [CrossRef]
- Liu, X.W.; Xia, C.; Li, L.; Guan, W.Y.; Pettit, N.; Zhang, H.C.; Chen, M.; Wang, P.G. Characterization and synthetic application of a novel β1,3-galactosyltransferase from Escherichia coli O55:H7. Bioorg. Med. Chem. 2009, 17, 4910–4915. [Google Scholar] [CrossRef] [PubMed]
- McArthur, J.B.; Yu, H.; Chen, X. A Bacterial β1-3-Galactosyltransferase Enables Multigram-Scale Synthesis of Human Milk Lacto- N-tetraose (LNT) and its Fucosides. ACS Catal. 2019, 9, 10721–10726. [Google Scholar] [CrossRef]
- Chen, X.; Jin, L.; Jiang, X.; Guo, L.; Gu, G.; Xu, L.; Lu, L.; Wang, F.; Xiao, M. Converting a β-N-acetylhexosaminidase into two trans-β-N-acetylhexosaminidases by domain-targeted mutagenesis. Appl. Microbiol. Biotechnol. 2020, 104, 661–673. [Google Scholar] [CrossRef] [PubMed]
- Murata, T.; Inukai, T.; Suzuki, M.; Yamagishi, M.; Usui, T. Facile enzymatic conversion of lactose into lacto-N-tetraose and lacto-N-neotetraose. Glycoconj. J. 1999, 16, 189–195. [Google Scholar] [CrossRef] [PubMed]
- Miyazaki, T.; Sato, T.; Furukawa, K.; Ajisaka, K. Enzymatic synthesis of Lacto-N-difucohexaose i which binds to Helicobacter pylori. In Methods in Enzymology; Academic Press: Cambridge, MA, USA, 2010; Volume 480, pp. 511–524. [Google Scholar] [CrossRef]
- Wada, J.; Ando, T.; Kiyohara, M.; Ashida, H.; Kitaoka, M.; Yamaguchi, M.; Kumagai, H.; Katayama, T.; Yamamoto, K. Bifidobacterium bifidum lacto-N-biosidase, a critical enzyme for the degradation of human milk oligosaccharides with a type 1 structure. Appl. Environ. Microbiol. 2008, 74, 3996–4004. [Google Scholar] [CrossRef]
- Schmölzer, K.; Weingarten, M.; Baldenius, K.; Nidetzky, B. Lacto-: N -tetraose synthesis by wild-type and glycosynthase variants of the β-N-hexosaminidase from Bifidobacterium bifidum. Org. Biomol. Chem. 2019, 17, 5661–5665. [Google Scholar] [CrossRef]
- Ito, T.; Katayama, T.; Hattie, M.; Sakurama, H.; Wada, J.; Suzuki, R.; Ashida, H.; Wakagi, T.; Yamamoto, K.; Stubbs, K.A.; et al. Crystal structures of a glycoside hydrolase family 20 lacto-N-biosidase from Bifidobacterium bifidum. J. Biol. Chem. 2013, 288, 11795–11806. [Google Scholar] [CrossRef] [PubMed]
- Val-Cid, C.; Biarnés, X.; Faijes, M.; Planas, A. Structural-functional analysis reveals a specific domain organization in family GH20 hexosaminidases. PLoS ONE 2015, 10, e0128075. [Google Scholar] [CrossRef]
- Lombard, V.; Golaconda-Ramulu, H.; Drula, E.; Coutinho, P.M.; Henrissat, B. The carbohydrate-active enzymes database (CAZy) in 2013. Nucleic Acids Res. 2014, 42, 490–495. [Google Scholar] [CrossRef]
- Tews, I.; Terwisscha Van Scheltinga, A.C.; Perrakis, A.; Wilson, K.S.; Dijkstra, B.W. Substrate-assisted catalysis unifies two families of chitinolytic enzymes. J. Am. Chem. Soc. 1997, 119, 7954–7959. [Google Scholar] [CrossRef]
- Mark, B.L.; Vocadlo, D.J.; Knapp, S.; Triggs-Raine, B.L.; Withers, S.G.; James, M.N.G. Crystallographic Evidence for Substrate-assisted Catalysis in a Bacterial β-Hexosaminidase. J. Biol. Chem. 2001, 276, 10330–10337. [Google Scholar] [CrossRef]
- Vocadlo, D.J.; Withers, S.G. Detailed comparative analysis of the catalytic mechanisms of β-N-acetylglucosaminidases from families 3 and 20 of glycoside hydrolases. Biochemistry 2005, 44, 12809–12818. [Google Scholar] [CrossRef]
- Bissaro, B.; Monsan, P.; Fauré, R.; O’Donohue, M.J. Glycosynthesis in a waterworld: New insight into the molecular basis of transglycosylation in retaining glycoside hydrolases. Biochem. J. 2015, 467, 17–35. [Google Scholar] [CrossRef]
- Teze, D.; Jiao, Z.; Wiemann, M.; Gulshan Kazi, Z.; Lupo, R.; Rønne, M.E.; Carlström, G.; Duus, J.Ø.; Sanejouand, Y.-H.; O’Donohue, M.J.; et al. Rational Enzyme Design Without Structural Knowledge: A Sequence-Based Approach for Efficient Generation of Glycosylation Catalysts. ChemRxiv 2020. [Google Scholar] [CrossRef]
- Kapešová, J.; Petrásková, L.; Kulik, N.; Straková, Z.; Bojarová, P.; Markošová, K.; Rebroš, M.; Křen, V.; Slámová, K. Transglycosidase activity of glycosynthase-type mutants of a fungal GH20 β-N-acetylhexosaminidase. Int. J. Biol. Macromol. 2020, 161, 1206–1215. [Google Scholar] [CrossRef] [PubMed]
- Schmölzer, K.; Weingarten, M.; Baldenius, K.; Nidetzky, B. Glycosynthase Principle Transformed into Biocatalytic Process Technology: Lacto-N-triose II Production with Engineered exo-Hexosaminidase. ACS Catal. 2019, 9, 5503–5514. [Google Scholar] [CrossRef]
- Ruzic, L.; Bolivar, J.M.; Nidetzky, B. Glycosynthase reaction meets the flow: Continuous synthesis of lacto-N-triose II by engineered β-hexosaminidase immobilized on solid support. Biotechnol. Bioeng. 2020, 117, 1597–1602. [Google Scholar] [CrossRef] [PubMed]
- Muschiol, J.; Vuillemin, M.; Meyer, A.S.; Zeuner, B. β-N-Acetylhexosaminidases for Carbohydrate Synthesis via Trans-Glycosylation. Catalysts 2020, 20, 365. [Google Scholar] [CrossRef]
- Slámová, K.; Krejzová, J.; Marhol, P.; Kalachova, L.; Kulik, N.; Pelantová, H.; Cvačka, J.; Křen, V. Synthesis of derivatized chitooligomers using transglycosidases engineered from the fungal GH20 β-N-acetylhexosaminidase. Adv. Synth. Catal. 2015, 357, 1941–1950. [Google Scholar] [CrossRef]
- Studier, F.W. Protein production by auto-induction in high density shaking cultures. Protein Expr. Purif. 2005, 41, 207–234. [Google Scholar] [CrossRef] [PubMed]

| RESIDUE (Subsite) | MUTANT | SAH 1 (s−1) | %H 2 (mut/wt) | % LNT 3 Yield (max) | t (min) 4 Max Yield | t Interval 5 (min) |
|---|---|---|---|---|---|---|
| WT | WT | 24 ± 1.8 | 100 | n.d.6 | ||
| Trp373 (-1) | W373F | 3.7 ± 0.2 | 15.4 | 0.5 | 30 | 30–120 |
| Trp394 (-1) | W394F | 0.07 ± 0.01 | 0.28 | 32 | 105 | 60–150 |
| W394A | 0.030 ± 0.001 | 0.13 | 16.6 | 150 | 90–180 | |
| W394K | <0.01 | <0.04 | n.d. | |||
| W394E | 0.031 ± 0.002 | 0.12 | 5.6 | 180 | 20–360 | |
| W394Q | 0.011 ± 0.001 | 0.04 | 6.8 | 1440 | 1434–45 | |
| Tyr419 (-1) | Y419F | 0.13 ± 0.09 | 0.6 | 1.8 | 1 | 1–10 |
| Tyr427 (-1) | Y427F | 15.4 ± 0.8 | 64 | 2.1 | 1 | 1–3 |
| Trp465 (-1) | W465F | 0.06 ± 0.03 | 0.3 | 4.1 | 300 | 1–360 |
| Glu321 *(-1) | E321A | 0.80 ± 0.08 | 3 | - | ||
| Asp320 ** (-1/-2) | D320A | <0.01 | <0.04 | |||
| D320E | 0.24 ± 0.02 | 1.0 | 3.3 | 90 | 60–120 | |
| Gln190 (-2) | Q190L | 0.14 ± 0.02 | 0.6 | 15 | 150 | 120–180 |
| Q190H | 10 ± 1 | 43.3 | 1.4 | 240 | 1–240 | |
| Glu216 (-2) | E216A | 4.3 ± 0.2 | 18.0 | 4.3 | 5 | 1–10 |
| E216D | 8.7 ± 0.5 | 36 | 3.5 | 10 | 1–60 | |
| Asn259 (-2) | N259A | 0.4 ± 0.1 | 1.6 | 10.6 | 20 | 10–30 |
| N259Q | 0.017 ± 0.002 | 0.07 | 16.3 | 360 | 90–360 | |
| His263 (-2) | H263R | 0.20 ± 0.01 | 0.8 | 17 | 90 | 40–240 |
| H263A | 0.14 ± 0.03 | 0.6 | 21 | 240 | 90–360 | |
| Asp467 (-1/-2) | D467A | 0.34 ± 0.03 | 1.4 | 0.5 | 30 | 30–120 |
| D467E | 2.5 ± 0.6 | 10.4 | 2.8 | 5 | 3–30 | |
| Leu574 (-2) | L574V | 15 ± 3 | 65 | 1.9 | 1 | 1–5 |
| MUTANT | SAH 1 (s−1) | % H 2 (mut/wt) | % LNT 3 Yield (max) |
|---|---|---|---|
| W394F_Y419F | 0.014 ± 0.009 | 0.06 | 4.7 |
| W394F_N259A | 0.0060 ± 0.0008 | 0.02 | 2.9 |
| W394F_N259Q | 0.0053 ± 0.0003 | 0.02 | n.d |
| W394F_H263A | 0.044 ± 0.003 | 0.2 | 5.2 |
| H263R_Y419F | <0.01 | <0.04 | 0.7 |
| H263A_Y419F | 0.05 ± 0.005 | 0.2 | 1 |
| Q190L_Y419F | 0.17 ± 0.01 | 0.7 | 9.7 |
| MUTANT | SAH 1 (s−1) | SAT 2 (s−1) | App.T/H Ratio 3 |
|---|---|---|---|
| wt | 27.7 ± 1.54 | 27.96 ± 2.63 | 1.0 |
| W394F | 0.069 ± 0.003 | 0.098 ± 0.0001 | 1.4 |
| W394A | 0.024 ± 0.004 | 0.018 ± 0.002 | 0.8 |
| H263R | 0.169 ± 0.003 | 0.22 ± 0.02 | 1.3 |
| H263A | 0.135 ± 0.001 | 0.13 ± 0.02 | 0.9 |
| N259Q | 0.017 ± 0.001 | 0.023 ± 0.005 | 1.3 |
| Q190L | 0.135 ± 0.003 | 0.122 ± 0.004 | 0.9 |
| MUTANT | Hydrolysis Mode | Transglycosylation Mode | |||||
|---|---|---|---|---|---|---|---|
| kcat (s−1) | KM (μM) | kcat/KM (M−1 · s−1) | kcat (s−1) | KM (μM) | kcat/KM (M−1 · s−1) | T/H (kcat/KM) 1 | |
| WT | 80 ± 8 | 520 ± 120 | 1.53·105 | 90 ± 13 | 750 ± 200 | 1.20·105 | 0.8 |
| W394F | 0.186 ± 0.008 | 260 ± 30 | 7.18·102 | 0.26 ± 0.02 | 240 ± 52 | 1.09·103 | 1.5 |
| W394A | 0.041 ± 0.003 | 87 ± 22 | 4.58·102 | 0.06 ± 0.01 | 530 ± 190 | 1.19·102 | 0.3 |
| H263R | 1.0 ± 0.2 | 780 ± 310 | 1.29·103 | 1.0 ± 0.2 | 720 ± 300 | 1.44·103 | 1.1 |
| H263A | 0.67 ± 0.03 | 670 ± 60 | 9.82·102 | 1.08 ± 0.18 | 1280 ± 320 | 8.93·102 | 0.9 |
| N259Q | 0.034 ± 0.002 | 235 ± 42 | 1.45·102 | 0.07 ± 0.01 | 490 ± 150 | 1.50·102 | 1.0 |
| Q190L | 0.201 ± 0.009 | 34 ± 7 | 5.93·103 | 0.41 ± 0.02 | 78 ± 14 | 5.24·103 | 0.9 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Castejón-Vilatersana, M.; Faijes, M.; Planas, A. Transglycosylation Activity of Engineered Bifidobacterium Lacto-N-Biosidase Mutants at Donor Subsites for Lacto-N-Tetraose Synthesis. Int. J. Mol. Sci. 2021, 22, 3230. https://doi.org/10.3390/ijms22063230
Castejón-Vilatersana M, Faijes M, Planas A. Transglycosylation Activity of Engineered Bifidobacterium Lacto-N-Biosidase Mutants at Donor Subsites for Lacto-N-Tetraose Synthesis. International Journal of Molecular Sciences. 2021; 22(6):3230. https://doi.org/10.3390/ijms22063230
Chicago/Turabian StyleCastejón-Vilatersana, Mireia, Magda Faijes, and Antoni Planas. 2021. "Transglycosylation Activity of Engineered Bifidobacterium Lacto-N-Biosidase Mutants at Donor Subsites for Lacto-N-Tetraose Synthesis" International Journal of Molecular Sciences 22, no. 6: 3230. https://doi.org/10.3390/ijms22063230
APA StyleCastejón-Vilatersana, M., Faijes, M., & Planas, A. (2021). Transglycosylation Activity of Engineered Bifidobacterium Lacto-N-Biosidase Mutants at Donor Subsites for Lacto-N-Tetraose Synthesis. International Journal of Molecular Sciences, 22(6), 3230. https://doi.org/10.3390/ijms22063230







