Beta-1,3 Oligoglucans Specifically Bind to Immune Receptor CD28 and May Enhance T Cell Activation
Abstract
1. Introduction
2. Results and Discussion
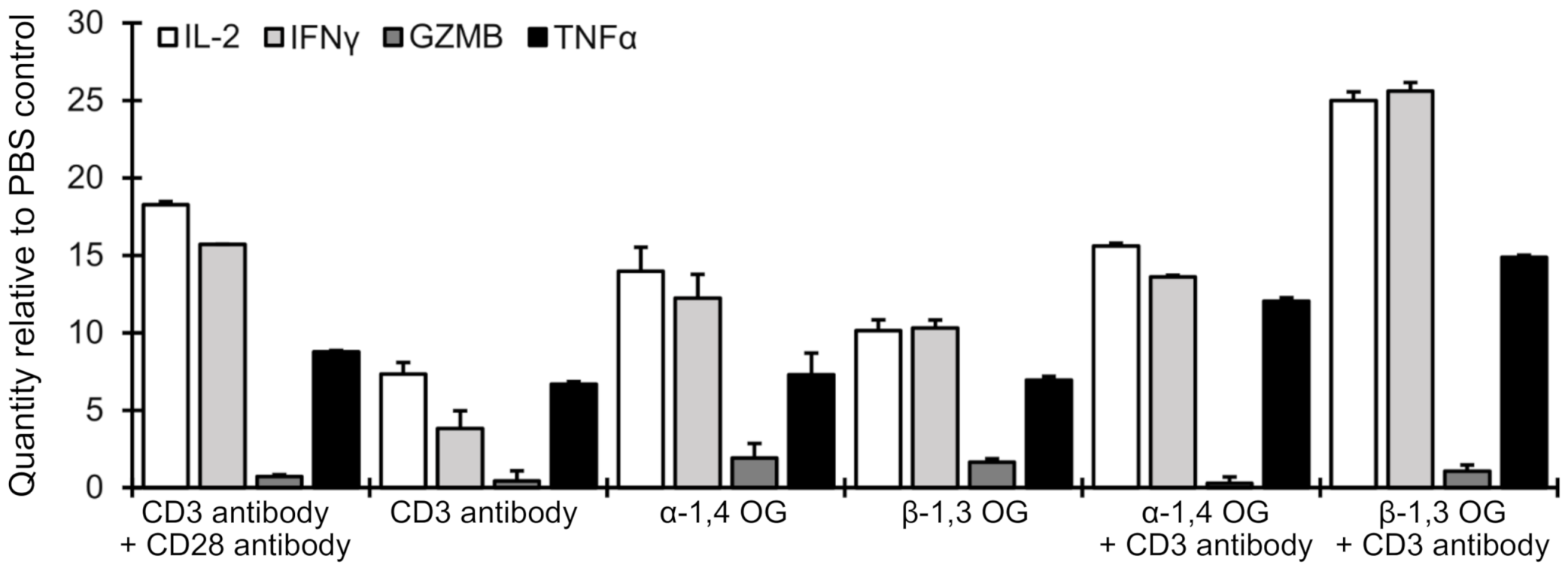
2.1. β-1,3 Oligoglucan Treatment Induced Expression of T Lymphoblasts Activation-Associated Cytokines
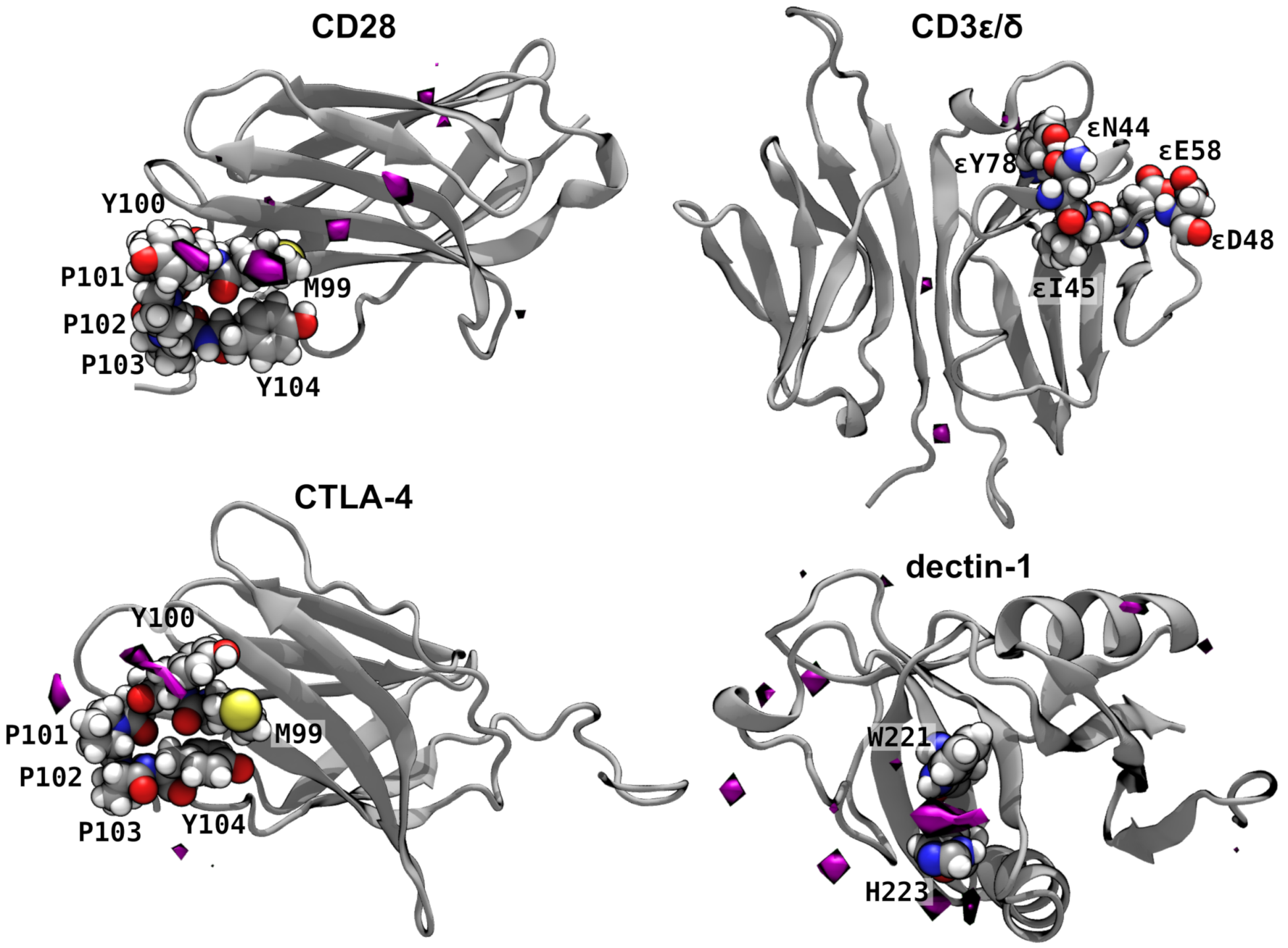
2.2. Interaction of β-1,3 Oligoglucans with Immune Receptors.
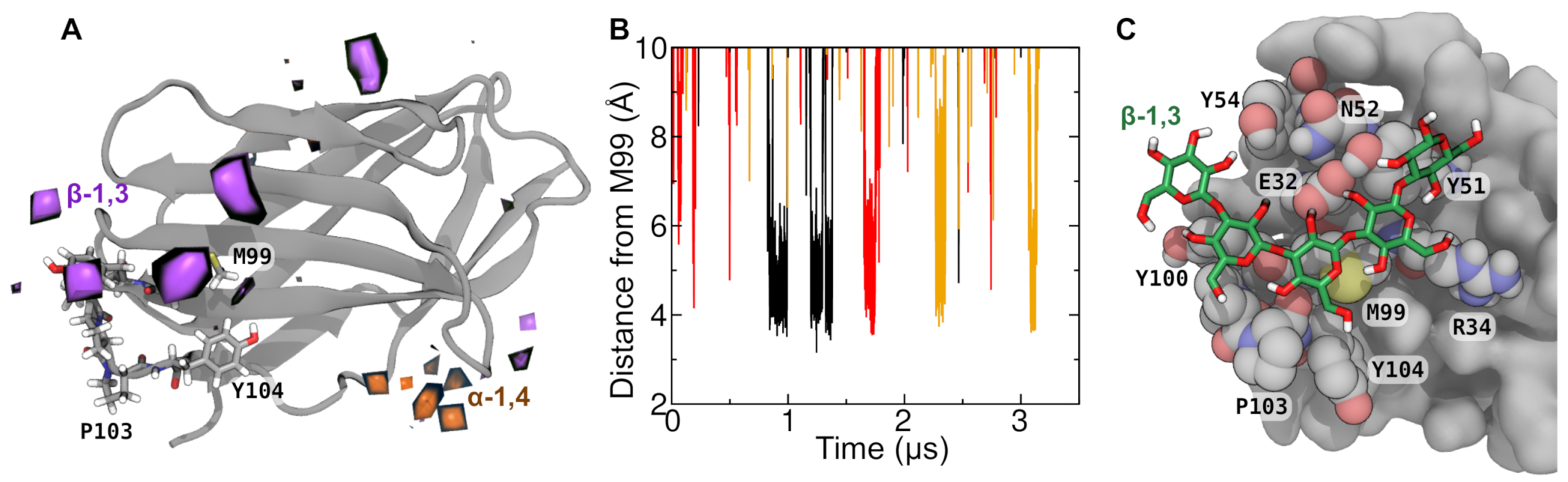
2.3. Specificity of β-1,3 Interaction with CD28.
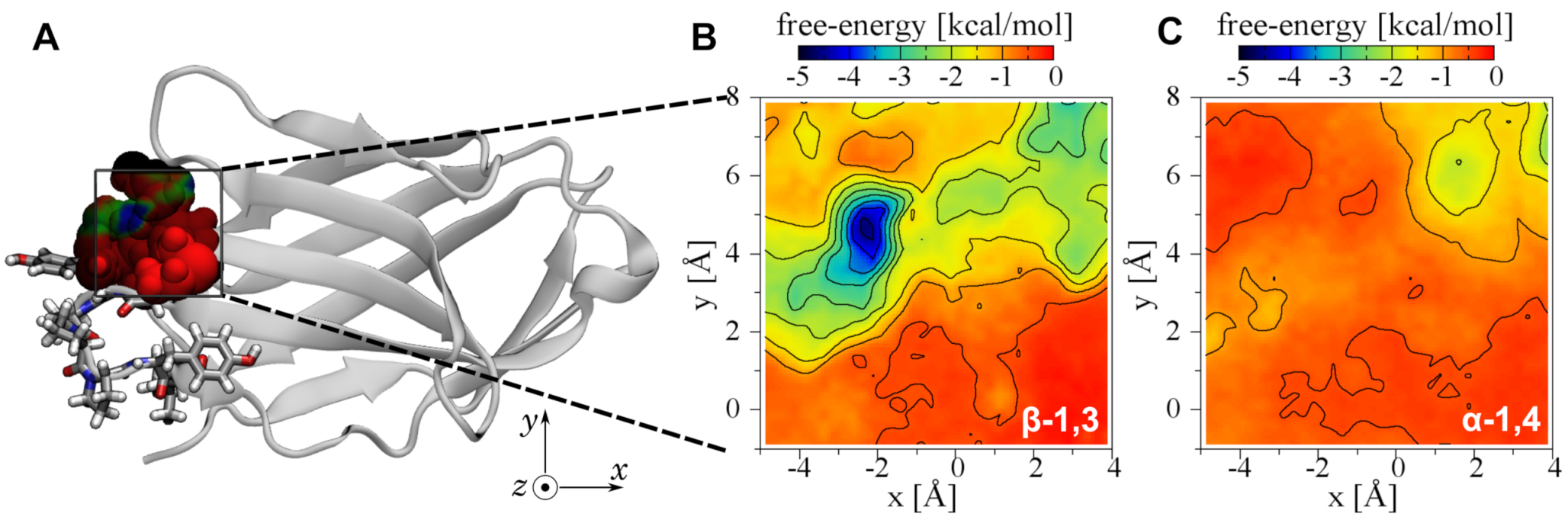
2.4. Three-Dimensional Free Energy Calculations
2.5. Calculation of the Standard Binding Free Energy
3. Conclusions
4. Materials and Methods
4.1. Cell Culture
4.2. Analysis of Cytokine Expression
4.3. Molecular Models
4.4. Molecular Dynamics Simulations
4.5. Unbiased Simulations
4.6. Three-Dimensional Free Energy Calculations
4.7. Calculation of the Standard Binding Free Energy
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Novak, M.; Vetvicka, V. -glucans, history, and the present: Immunomodulatory aspects and mechanisms of action. J. Immunotoxicol. 2008, 5, 47–57. [Google Scholar] [CrossRef]
- McIntosh, M.; Stone, B.; Stanisich, V. Curdlan and other bacterial (1→3)--D-glucans. Appl. Microbiol. Biotechnol. 2005, 68, 163–173. [Google Scholar] [CrossRef] [PubMed]
- Marchessault, R.H.; Deslandes, Y. Fine structure of (1→3)--D-glucans: Curdlan and paramylon. Carbohydr. Res. 1979, 75, 231–242. [Google Scholar] [CrossRef]
- Peat, S.; Whelan, W.; Lawley, H. 141. The structure of laminarin. Part I. The main polymeric linkage. J. Chem. Soc. 1958, 0, 724–728. [Google Scholar] [CrossRef]
- Sasaki, T.; Takasuka, N. Further study of the structure of lentinan, an anti-tumor polysaccharide from Lentinus edodes. Carbohydr. Res. 1976, 47, 99–104. [Google Scholar] [CrossRef]
- Saito, H.; Misaki, A.; Harada, T. A comparison of the structure of curdlan and pachyman. Anal. Biochem. 1968, 32, 1261–1269. [Google Scholar] [CrossRef]
- Karácsonyi, Š.; Kuniak, L. Polysaccharides of Pleurotus ostreatus: Isolation and structure of pleuran, an alkali-insoluble -D-glucan. Carbohydr. Polym. 1994, 24, 107–111. [Google Scholar] [CrossRef]
- Di Carlo, F.J.; Fiore, J.V. On the composition of zymosan. Science 1958, 127, 756–757. [Google Scholar] [CrossRef]
- Bohn, J.A.; BeMiller, J.N. (1→3)--D-glucans as biological response modifiers: A review of structure-functional activity relationships. Carbohydr. Polym. 1995, 28, 3–14. [Google Scholar] [CrossRef]
- Goodridge, H.S.; Wolf, A.J.; Underhill, D.M. -glucan recognition by the innate immune system. Immunol. Rev. 2009, 230, 38–50. [Google Scholar] [CrossRef]
- Nathan, A.T.; Peterson, E.A.; Chakir, J.; Wills-Karp, M. Innate immune responses of airway epithelium to house dust mite are mediated through -glucan–dependent pathways. J. Allergy Clin. Immun. 2009, 123, 612–618. [Google Scholar] [CrossRef]
- Rop, O.; Mlcek, J.; Jurikova, T. Beta-glucans in higher fungi and their health effects. Nutr. Rev. 2009, 67, 624–631. [Google Scholar] [CrossRef]
- Othman, R.A.; Moghadasian, M.H.; Jones, P.J. Cholesterol-lowering effects of oat -glucan. Nutr. Rev. 2011, 69, 299–309. [Google Scholar] [CrossRef]
- Wang, Y.; Ames, N.P.; Tun, H.M.; Tosh, S.M.; Jones, P.J.; Khafipour, E. High molecular weight barley -glucan alters gut microbiota toward reduced cardiovascular disease risk. Front. Microbiol. 2016, 7, 129. [Google Scholar] [CrossRef]
- Brown, G.D.; Gordon, S. Fungal -glucans and mammalian immunity. Immunity 2003, 19, 311–315. [Google Scholar] [CrossRef]
- Baldassano, S.; Accardi, G.; Vasto, S. Beta-glucans and cancer: The influence of inflammation and gut peptide. Eur. J. Med. Chem. 2017, 142, 486–492. [Google Scholar] [CrossRef] [PubMed]
- Ishiguru, S.; Robben, N.; Burghart, R.; Cote, P.; Greenway, S.; Thakkar, R.; Upreti, D.; Ayaka, N.; Suzuki, K.; Comer, J.; et al. Cell wall membrane fraction of Chlorellasorokiniana enhances host anti-tumor immunity and inhibits colon carcinoma growth in mice. Integr. Cancer Ther. 2020, 19, 1–10. [Google Scholar] [CrossRef]
- Legentil, L.; Paris, F.; Ballet, C.; Trouvelot, S.; Daire, X.; Vetvicka, V.; Ferrières, V. Molecular interactions of -(1→3)-glucans with their receptors. Molecules 2015, 20, 9745–9766. [Google Scholar] [CrossRef] [PubMed]
- Buck, K.; Chen, A.W.; Dickerson, A.; Chain, E. Formation and structure of extracellular glucans produced by Claviceps species. Microbiology 1968, 51, 337–352. [Google Scholar] [CrossRef]
- Kim, Y.W.; Kim, K.H.; Choi, H.J.; Lee, D.S. Anti-diabetic activity of -glucans and their enzymatically hydrolyzed oligosaccharides from Agaricus Blaz. Biotechnol. Lett. 2005, 27, 483–487. [Google Scholar] [CrossRef] [PubMed]
- Ross, G.D.; Cain, J.A.; Myones, B.L.; Newman, S.L.; Lachmann, P.J. Specificity of membrane complement receptor type three (CR3) for ß-glucans. Complement 1987, 4, 61–74. [Google Scholar] [CrossRef]
- Brown, G.D.; Gordon, S. Immune recognition: A new receptor for -glucans. Nature 2001, 413, 36. [Google Scholar] [CrossRef] [PubMed]
- Sato, M.; Sano, H.; Iwaki, D.; Kudo, K.; Konishi, M.; Takahashi, H.; Takahashi, T.; Imaizumi, H.; Asai, Y.; Kuroki, Y. Direct binding of Toll-like receptor 2 to zymosan, and zymosan-induced NF-κB activation and TNF-α secretion are down-regulated by lung collectin surfactant protein A. J. Immunol. 2003, 171, 417–425. [Google Scholar] [CrossRef] [PubMed]
- Fuller, G.L.; Williams, J.A.; Tomlinson, M.G.; Eble, J.A.; Hanna, S.L.; Pöhlmann, S.; Suzuki-Inoue, K.; Ozaki, Y.; Watson, S.P.; Pearce, A.C. The C-type lectin receptors CLEC-2 and Dectin-1, but not DC-SIGN, signal via a novel YXXL-dependent signaling cascade. J. Biol. Chem. 2007, 282, 12397–12409. [Google Scholar] [CrossRef]
- De Jong, M.A.; Vriend, L.E.; Theelen, B.; Taylor, M.E.; Fluitsma, D.; Boekhout, T.; Geijtenbeek, T.B. C-type lectin Langerin is a -glucan receptor on human Langerhans cells that recognizes opportunistic and pathogenic fungi. Mol. Immunol. 2010, 47, 1216–1225. [Google Scholar] [CrossRef]
- Evans, E.J.; Esnouf, R.M.; Manso-Sancho, R.; Gilbert, R.J.; James, J.R.; Yu, C.; Fennelly, J.A.; Vowles, C.; Hanke, T.; Walse, B.; et al. Crystal structure of a soluble CD28-Fab complex. Nature Immunol. 2005, 6, 271. [Google Scholar] [CrossRef]
- Arnett, K.L.; Harrison, S.C.; Wiley, D.C. Crystal structure of a human CD3-є/δ dimer in complex with a UCHT1 single-chain antibody fragment. Proc. Natl. Acad. Sci. USA 2004, 101, 16268–16273. [Google Scholar] [CrossRef]
- Metzler, W.J.; Bajorath, J.; Fenderson, W.; Shaw, S.Y.; Constantine, K.L.; Naemura, J.; Leytze, G.; Peach, R.J.; Lavoie, T.B.; Mueller, L.; et al. Solution structure of human CTLA-4 and delineation of a CD80/CD86 binding site conserved in CD28. Nat. Struct. Mol. Biol. 1997, 4, 527. [Google Scholar] [CrossRef]
- Chang, T.T.; Kuchroo, V.K.; Sharpe, A.H. Role of the B7-CD28/CTLA-4 pathway in autoimmune disease. Curr. Dir. Autoimmun 2002, 5, 113–130. [Google Scholar]
- Huysamen, C.; Brown, G.D. The fungal pattern recognition receptor, Dectin-1, and the associated cluster of C-type lectin-like receptors. FEMS Microbiol. Lett. 2009, 290, 121–128. [Google Scholar] [CrossRef]
- Gumbart, J.C.; Roux, B.; Chipot, C. Standard binding free energies from computer simulations: What is the best strategy? J. Chem. Theory Comput. 2012, 9, 794–802. [Google Scholar] [CrossRef]
- Fu, H.; Cai, W.; Hénin, J.; Roux, B.; Chipot, C. New coarse variables for the accurate determination of standard binding free energies. J. Chem. Theory Comput. 2017, 13, 5173–5178. [Google Scholar] [CrossRef]
- Fu, H.; Gumbart, J.C.; Chen, H.; Shao, X.; Cai, W.; Chipot, C. BFEE: A user-friendly graphical interface facilitating absolute binding free-energy calculations. J. Chem. Inf. Model. 2018, 58, 556–560. [Google Scholar] [CrossRef] [PubMed]
- Adams, E.L.; Rice, P.J.; Graves, B.; Ensley, H.E.; Yu, H.; Brown, G.D.; Gordon, S.; Monteiro, M.A.; Papp-Szabo, E.; Lowman, D.W.; et al. Differential high-affinity interaction of dectin-1 with natural or synthetic glucans is dependent upon primary structure and is influenced by polymer chain length and side-chain branching. J. Pharmacol. Exp. Ther. 2008, 325, 115–123. [Google Scholar] [CrossRef]
- Doi, C.; Egashira, N.; Kawabata, A.; Maurya, D.K.; Ohta, N.; Uppalapati, D.; Ayuzawa, R.; Pickel, L.; Isayama, Y.; Troyer, D.; et al. Angiotensin II type 2 receptor signaling significantly attenuates growth of murine pancreatic carcinoma grafts in syngeneic mice. BMC Cancer 2010, 10, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Law, B.K.; Chytil, A.; Dumont, N.; Hamilton, E.G.; Waltner-Law, M.E.; Aakre, M.E.; Covington, C.; Moses, H.L. Rapamycin potentiates transforming growth factor -induced growth arrest in nontransformed, oncogene-transformed, and human cancer cells. Mol. Cell. Biol. 2002, 22, 8184–8198. [Google Scholar] [CrossRef]
- Kanehira, T.; Tani, T.; Takagi, T.; Nakano, Y.; Howard, E.F.; Tamura, M. Angiotensin II type 2 receptor gene deficiency attenuates susceptibility to tobacco-specific nitrosamine-induced lung tumorigenesis: Involvement of transforming growth factor--dependent cell growth attenuation. Cancer Res. 2005, 65, 7660–7665. [Google Scholar] [CrossRef]
- Schmittgen, T.D.; Livak, K.J. Analyzing real-time PCR data by the comparative CT method. Nat. Protoc. 2008, 3, 1101. [Google Scholar] [CrossRef] [PubMed]
- Brown, J.; O’Callaghan, C.A.; Marshall, A.S.; Gilbert, R.J.; Siebold, C.; Gordon, S.; Brown, G.D.; Jones, E.Y. Structure of the fungal -glucan-binding immune receptor dectin-1: Implications for function. Prot. Sci. 2007, 16, 1042–1052. [Google Scholar] [CrossRef]
- Jo, S.; Kim, T.; Iyer, V.G.; Im, W. CHARMM-GUI: A web-based graphical user interface for CHARMM. J. Comput. Chem. 2008, 29, 1859–1865. [Google Scholar] [CrossRef]
- Huang, J.; Rauscher, S.; Nawrocki, G.; Ran, T.; Feig, M.; de Groot, B.; Grubmüller, H.; MacKerell, A., Jr. CHARMM36m: An improved force field for folded and intrinsically disordered proteins. Nature Meth. 2017, 14, 71–73. [Google Scholar] [CrossRef]
- Best, R.B.; Zhu, X.; Shim, J.; Lopes, P.E.; Mittal, J.; Feig, M.; MacKerell, A.D., Jr. Optimization of the additive CHARMM all-atom protein force field targeting improved sampling of the backbone ϕ, ψ and side-chain χ1 and χ2 dihedral angles. J. Chem. Theory Comput. 2012, 8, 3257–3273. [Google Scholar] [CrossRef]
- MacKerell, A., Jr.; Feig, M.; Brooks, C., III. Improved treatment of the protein backbone in empirical force fields. J. Am. Chem. Soc. 2004, 126, 698–699. [Google Scholar] [CrossRef] [PubMed]
- MacKerell, A.D., Jr.; Bashford, D.; Bellott, M.; Dunbrack, R.L., Jr.; Evanseck, J.D.; Field, M.J.; Fischer, S.; Gao, J.; Guo, H.; Ha, S.; et al. All–atom empirical potential for molecular modeling and dynamics studies of proteins. J. Phys. Chem. B 1998, 102, 3586–3616. [Google Scholar] [CrossRef]
- Guvench, O.; Greene, S.N.; Kamath, G.; Brady, J.W.; Venable, R.M.; Pastor, R.W.; Mackerell, A.D. Additive empirical force field for hexopyranose monosaccharides. J. Comput. Chem. 2008, 29, 2543–2564. [Google Scholar] [CrossRef]
- Guvench, O.; Hatcher, E.; Venable, R.M.; Pastor, R.W.; MacKerell, A.D., Jr. CHARMM additive all-atom force field for glycosidic linkages between hexopyranoses. J. Chem. Theory Comput. 2009, 5, 2353–2370. [Google Scholar] [CrossRef]
- Miyamoto, S.; Kollman, P.A. SETTLE: An analytical version of the SHAKE and RATTLE algorithm for rigid water models. J. Comput. Chem. 1992, 13, 952–962. [Google Scholar] [CrossRef]
- Andersen, H.C. RATTLE: A “velocity” version of the SHAKE algorithm for molecular dynamics calculations. J. Chem. Phys. 1983, 52, 24–34. [Google Scholar] [CrossRef]
- Hopkins, C.W.; Le Grand, S.; Walker, R.C.; Roitberg, A.E. Long-time-step molecular dynamics through hydrogen mass repartitioning. J. Chem. Theory Comput. 2015, 11, 1864–1874. [Google Scholar] [CrossRef] [PubMed]
- Darden, T.A.; York, D.M.; Pedersen, L.G. Particle mesh Ewald: An NlogN method for Ewald sums in large systems. J. Chem. Phys. 1993, 98, 10089–10092. [Google Scholar] [CrossRef]
- Feller, S.E.; Zhang, Y.H.; Pastor, R.W.; Brooks, B.R. Constant pressure molecular dynamics simulations — The Langevin piston method. J. Chem. Phys. 1995, 103, 4613–4621. [Google Scholar] [CrossRef]
- Luo, Y.; Roux, B. Simulation of osmotic pressure in concentrated aqueous salt solutions. J. Phys. Chem. Lett. 2009, 1, 183–189. [Google Scholar] [CrossRef]
- Venable, R.M.; Luo, Y.; Gawrisch, K.; Roux, B.; Pastor, R.W. Simulations of anionic lipid membranes: Development of interaction-specific ion parameters and validation using NMR data. J. Phys. Chem. B 2013, 117, 10183–10192. [Google Scholar] [CrossRef] [PubMed]
- Martínez, L.; Andrade, R.; Birgin, E.G.; Martínez, J.M. Packmol: A package for building initial configurations for molecular dynamics simulations. J. Comput. Chem. 2009, 30, 2157–2164. [Google Scholar] [CrossRef] [PubMed]
- Darve, E.; Pohorille, A. Calculating free energies using average force. J. Chem. Phys. 2001, 115, 9169–9183. [Google Scholar] [CrossRef]
- Comer, J.; Gumbart, J.C.; Hénin, J.; Lelièvre, T.; Pohorille, A.; Chipot, C. The adaptive biasing force method: Everything you always wanted to know but were afraid to ask. J. Phys. Chem. B 2015, 119, 1129–1151. [Google Scholar] [CrossRef] [PubMed]
- Humphrey, W.; Dalke, A.; Schulten, K. VMD – Visual Molecular Dynamics. J. Mol. Graph. 1996, 14, 33–38. [Google Scholar] [CrossRef]
- Rastelli, G.; Rio, A.D.; Degliesposti, G.; Sgobba, M. Fast and accurate predictions of binding free energies using MM-PBSA and MM-GBSA. J. Comput. Chem. 2010, 31, 797–810. [Google Scholar] [CrossRef]
- Poblete, H.; Miranda-Carvajal, I.; Comer, J. Determinants of alanine dipeptide conformational equilibria on graphene and hydroxylated derivatives. J. Phys. Chem. B 2017, 121, 3895–3907. [Google Scholar] [CrossRef]
| Primer | Sequence | Size | |
|---|---|---|---|
| Human | Forward (5–3) | ATGAGACAGCAACCATTGTAGAATTT | 87 bp |
| IL-2 | Reverse (5–3) | CACTTAATTATCAAGTCAGTGTTGAGATGA | |
| Human | Forward (5–3) | GCCAGAATGCTGCAGGACTT | 63 bp |
| TNFα | Reverse (5–3) | GGCCTAAGGTCCACTTGTGTCA | |
| Human | Forward (5–3) | AGGGAAGCGAAAAAGGAGTCA | 64 bp |
| IFN | Reverse (5–3) | GGACAACCATTACTGGGATGCT | |
| Human | Forward (5–3) | TGCAGGAAGATCGAAAGTGCG | 180 bp |
| GZMB | Reverse (5–3) | GAGGCATGCCATTGTTTCGTC | |
| 18S | Forward (5–3) | GAGGTTCGAAGACGATCAGA | 315 bp |
| Reverse (5–3) | TCGCTCCACCAACTAAGAAC |
| Stage | System | Free-Energy | Free Energy | Sim. Time |
|---|---|---|---|---|
| Term | (kcal/mol) | (ns) | ||
| 1 | protein–ligand | 200 | ||
| 2 | protein–ligand | 200 | ||
| 3 | protein–ligand | 170 | ||
| 4 | protein–ligand | 160 | ||
| 5 | protein–ligand | 120 | ||
| 6 | protein–ligand | 100 | ||
| 7 | protein–ligand | 1190 | ||
| 8 | ligand only | 240 | ||
| 9 | ligand only | 0 | ||
| total | – | 2380 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Comer, J.; Bassette, M.; Burghart, R.; Loyd, M.; Ishiguro, S.; Azhagiya Singam, E.R.; Vergara-Jaque, A.; Nakashima, A.; Suzuki, K.; Geisbrecht, B.V.; et al. Beta-1,3 Oligoglucans Specifically Bind to Immune Receptor CD28 and May Enhance T Cell Activation. Int. J. Mol. Sci. 2021, 22, 3124. https://doi.org/10.3390/ijms22063124
Comer J, Bassette M, Burghart R, Loyd M, Ishiguro S, Azhagiya Singam ER, Vergara-Jaque A, Nakashima A, Suzuki K, Geisbrecht BV, et al. Beta-1,3 Oligoglucans Specifically Bind to Immune Receptor CD28 and May Enhance T Cell Activation. International Journal of Molecular Sciences. 2021; 22(6):3124. https://doi.org/10.3390/ijms22063124
Chicago/Turabian StyleComer, Jeffrey, Molly Bassette, Riley Burghart, Mayme Loyd, Susumu Ishiguro, Ettayapuram Ramaprasad Azhagiya Singam, Ariela Vergara-Jaque, Ayaka Nakashima, Kengo Suzuki, Brian V. Geisbrecht, and et al. 2021. "Beta-1,3 Oligoglucans Specifically Bind to Immune Receptor CD28 and May Enhance T Cell Activation" International Journal of Molecular Sciences 22, no. 6: 3124. https://doi.org/10.3390/ijms22063124
APA StyleComer, J., Bassette, M., Burghart, R., Loyd, M., Ishiguro, S., Azhagiya Singam, E. R., Vergara-Jaque, A., Nakashima, A., Suzuki, K., Geisbrecht, B. V., & Tamura, M. (2021). Beta-1,3 Oligoglucans Specifically Bind to Immune Receptor CD28 and May Enhance T Cell Activation. International Journal of Molecular Sciences, 22(6), 3124. https://doi.org/10.3390/ijms22063124






