Microscopic Interactions of Melatonin, Serotonin and Tryptophan with Zwitterionic Phospholipid Membranes
Abstract
1. Introduction
2. Results and Discussion
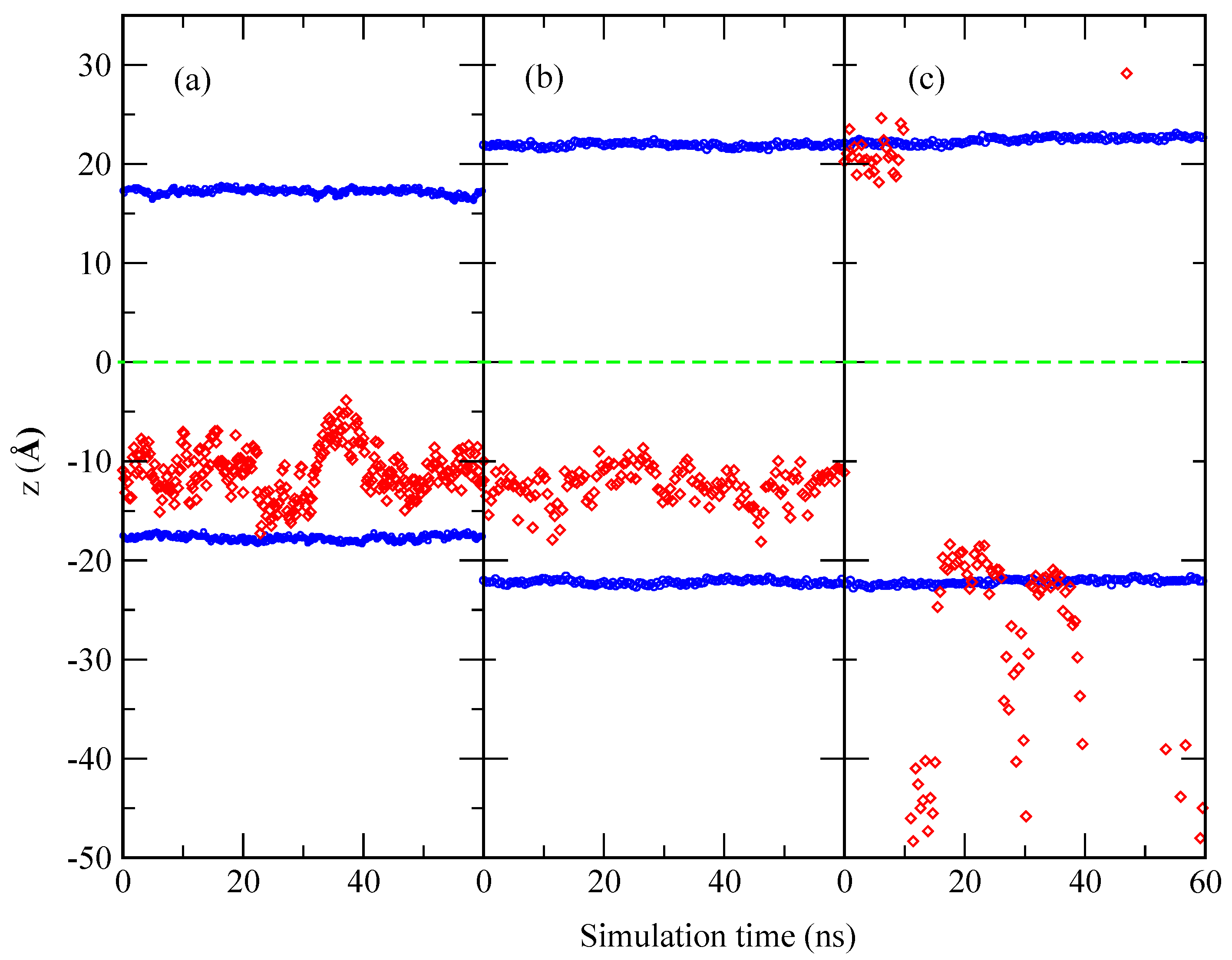
2.1. Structural Properties of the Membranes
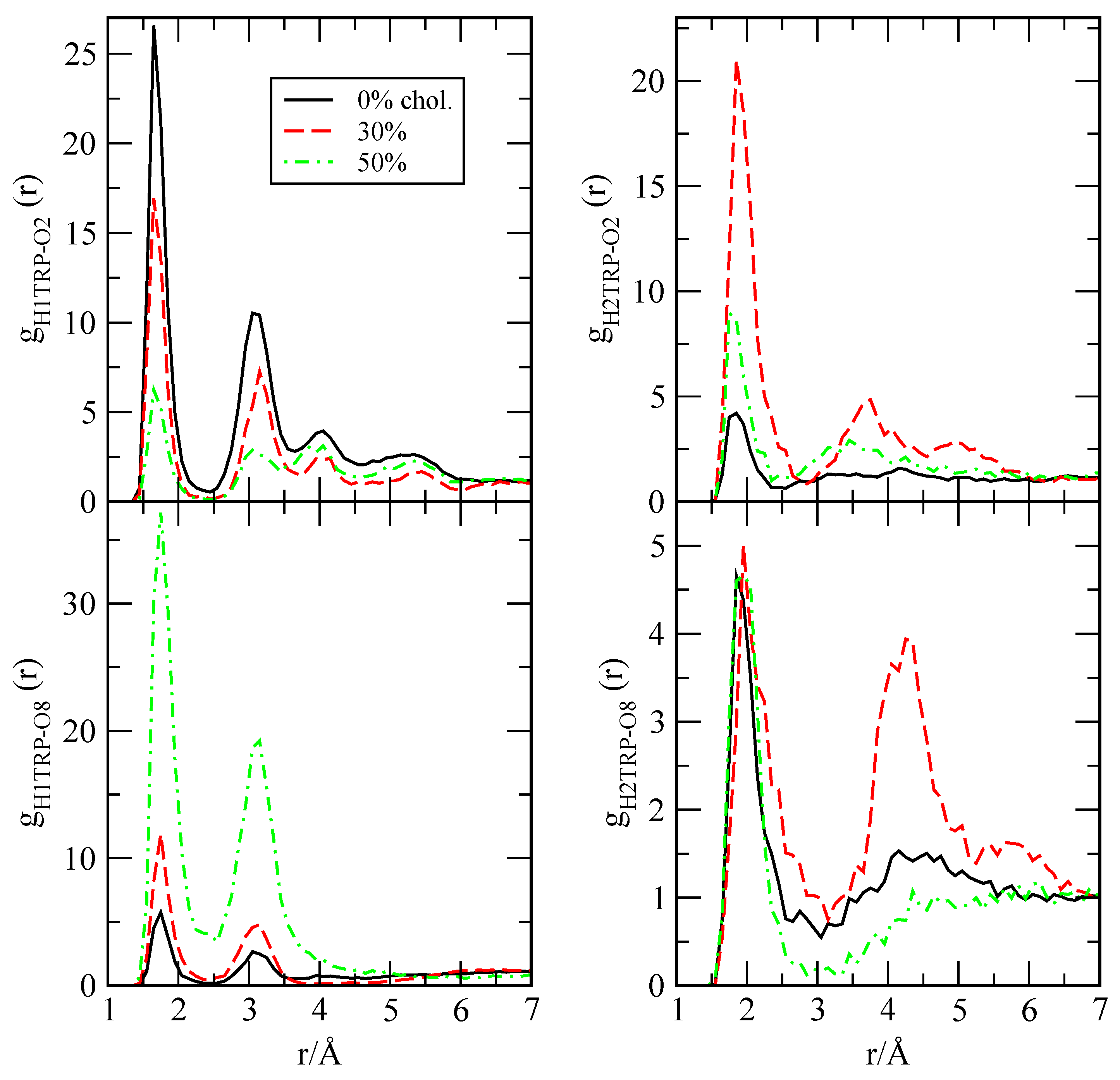
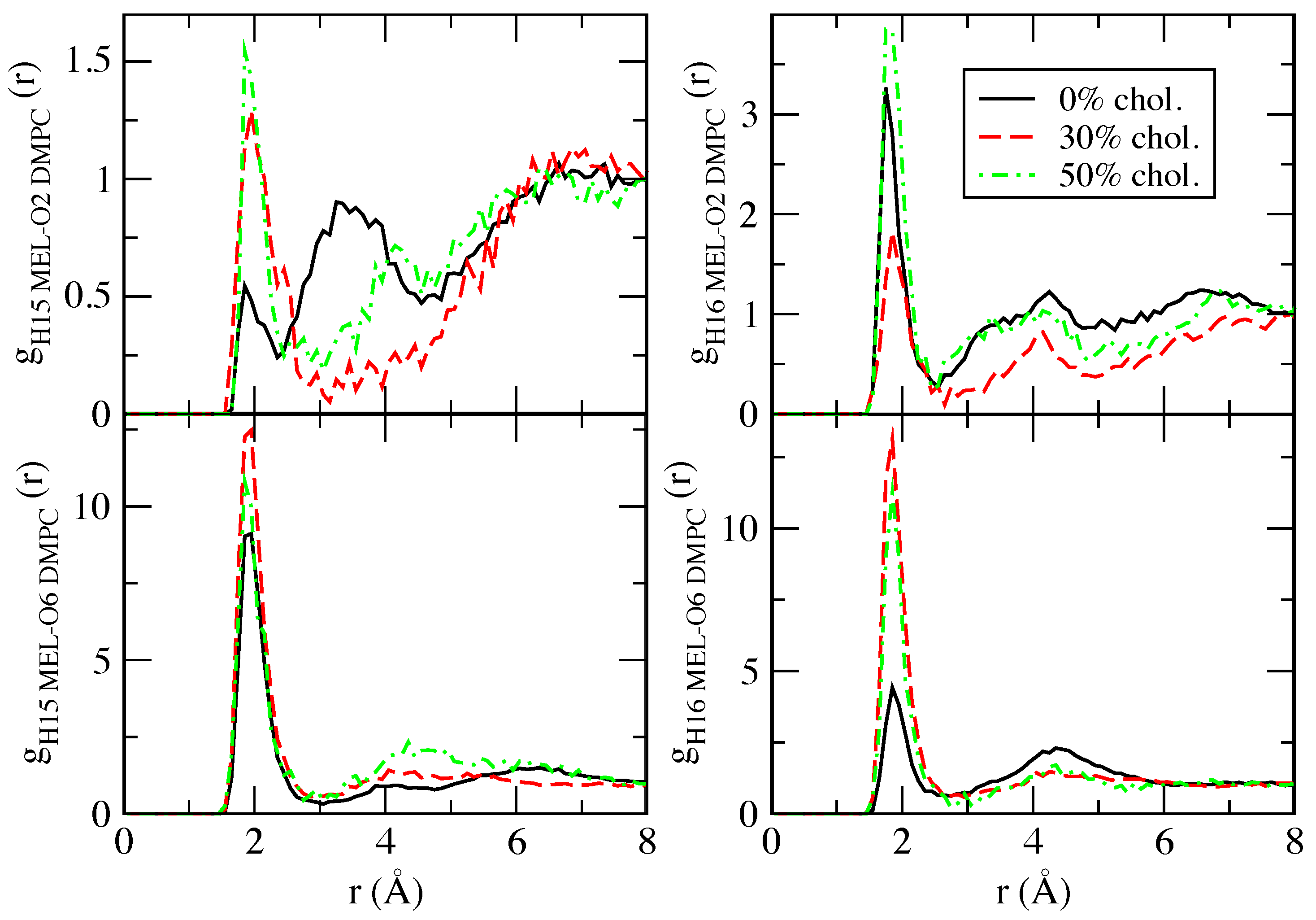
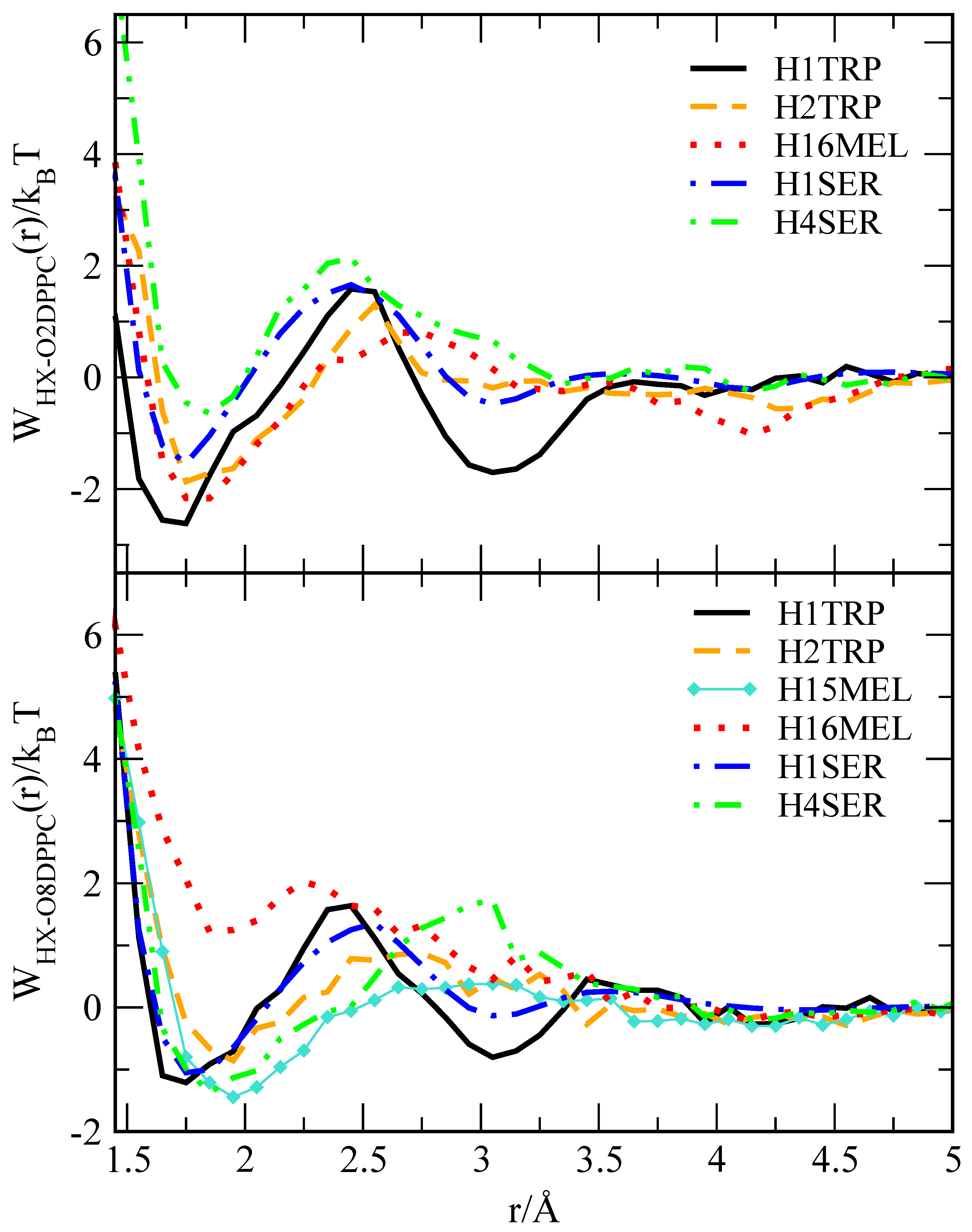
2.2. Preferential Localisations of the Small Molecules at the Interfaces of Phospholipid Membranes: Atomic Radial Distribution Functions
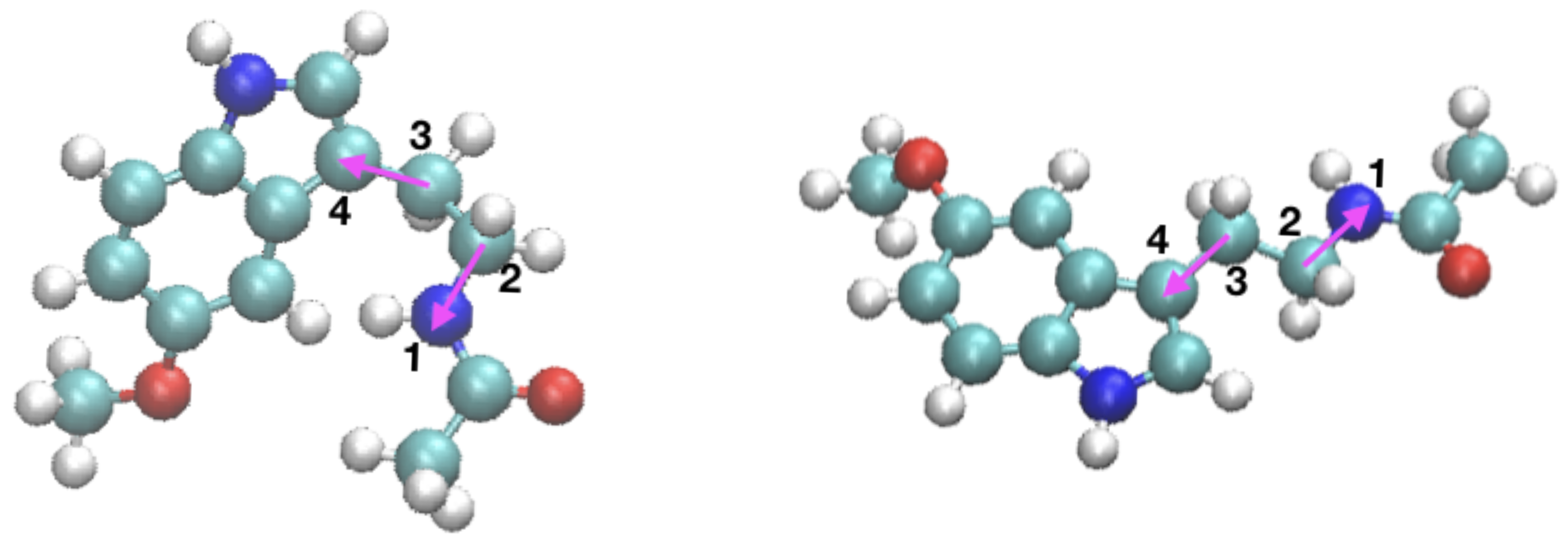
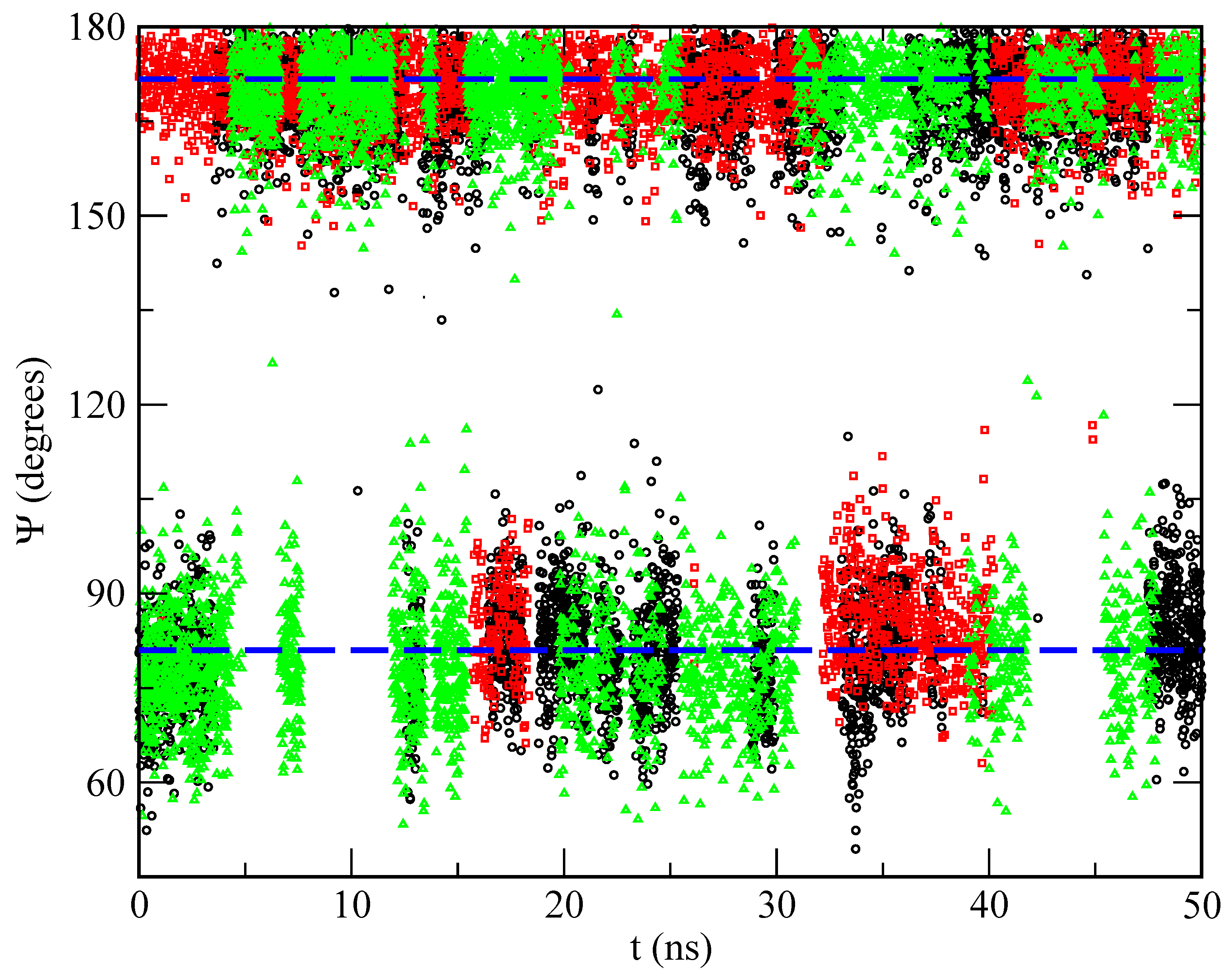
2.3. Orientational Distributions of Melatonin
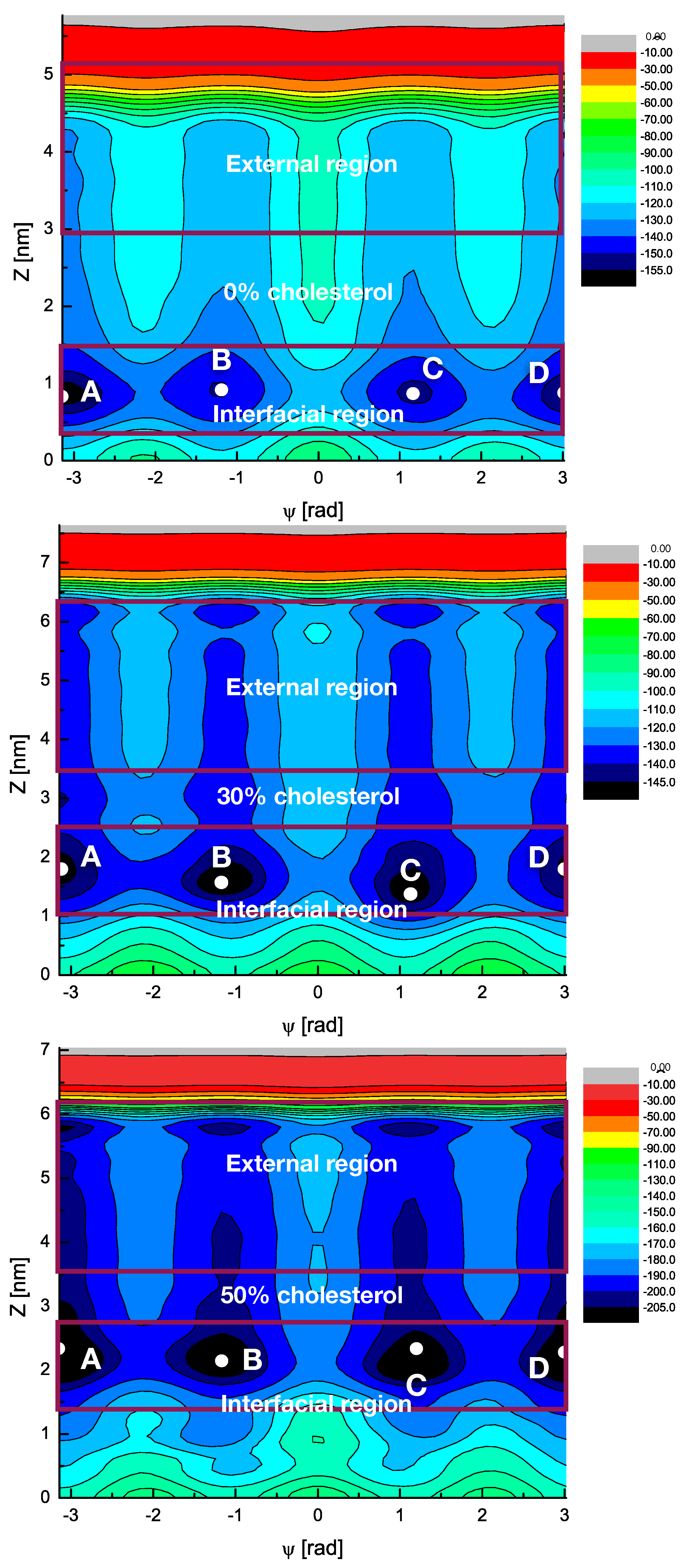
2.4. Free Energy Profiles of Small Molecules and Free Energy Hypersurfaces of Melatonin Binding
2.5. Diffusion Coefficients of Small Molecules: Tryptophan and Melatonin
3. Methods
3.1. Molecular Dynamics
3.2. Well Tempered Metadynamics
4. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| MD | molecular dynamics simulations |
| WTM | Well-tempered metadynamics simulations |
| DMPC | dipalmitoylphosphatidylcholine |
| DPPC | dimyristoylphosphatidylcholine |
| CHOL | cholesterol |
| TRP | tryptophan |
| SER | serotonin |
| MEL | melatonin |
| SI | supporting information |
Appendix A. Supporting Information
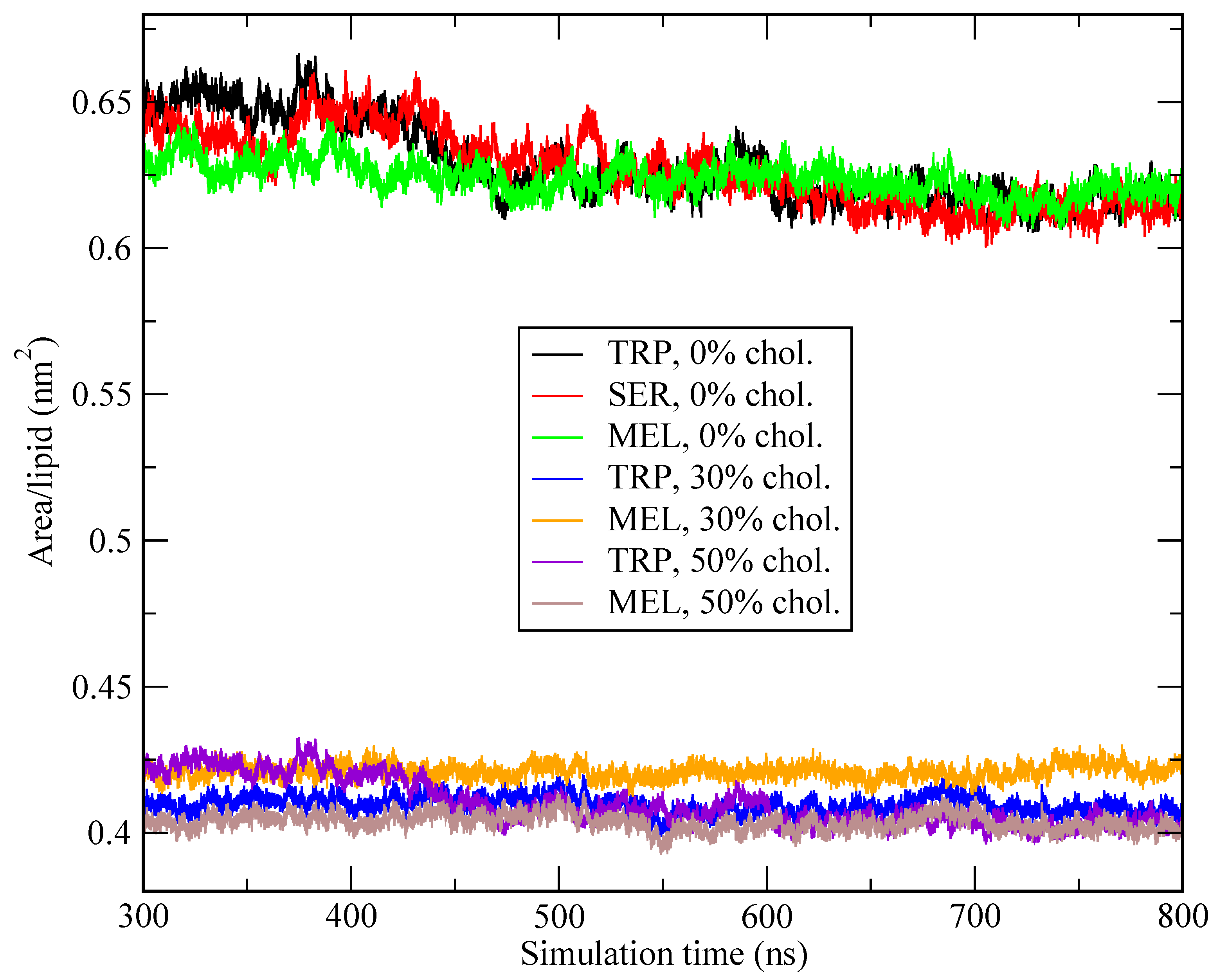
Appendix A.1. Convergence of MD Simulations
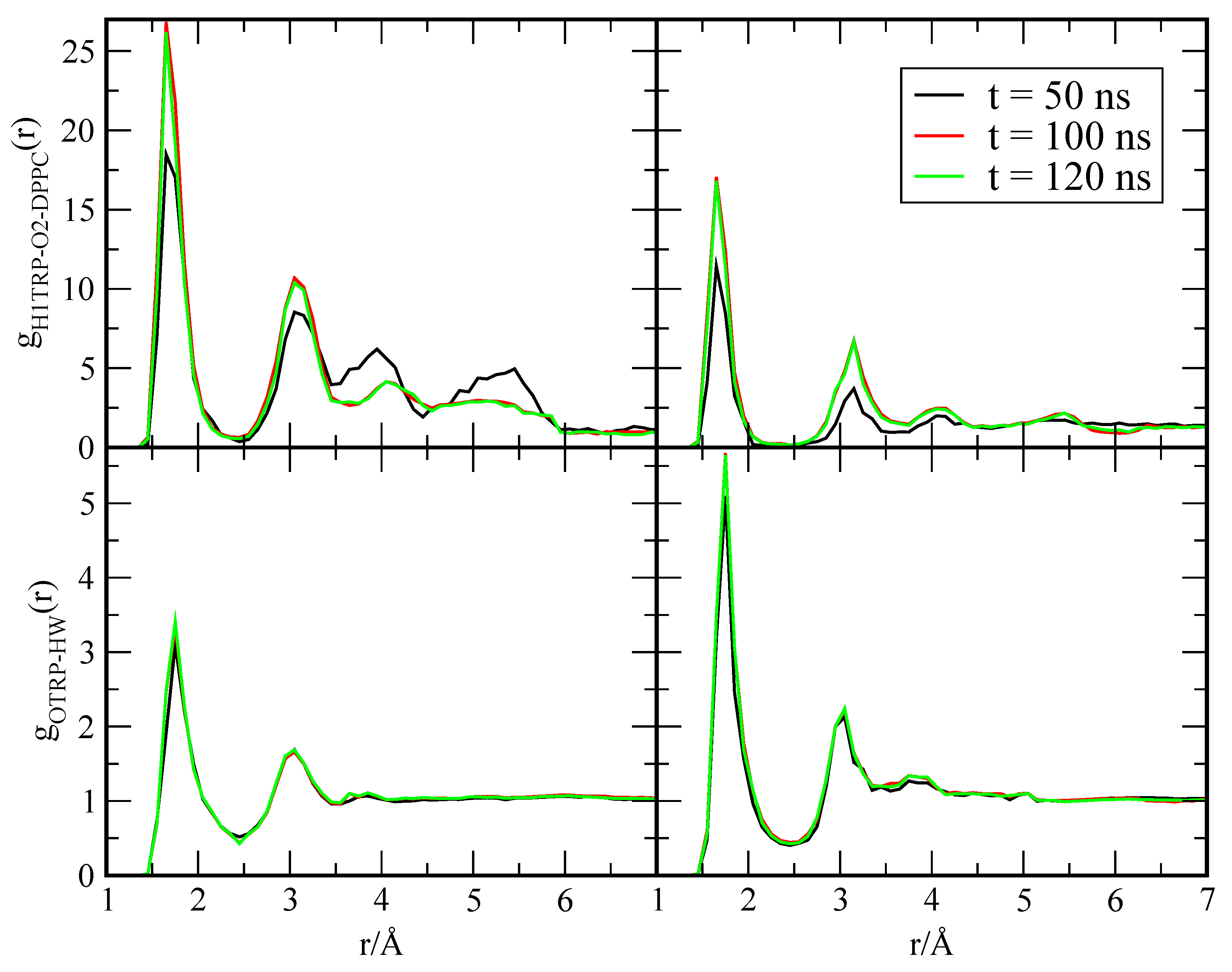
Appendix A.2. Convergence of WTM Simulations
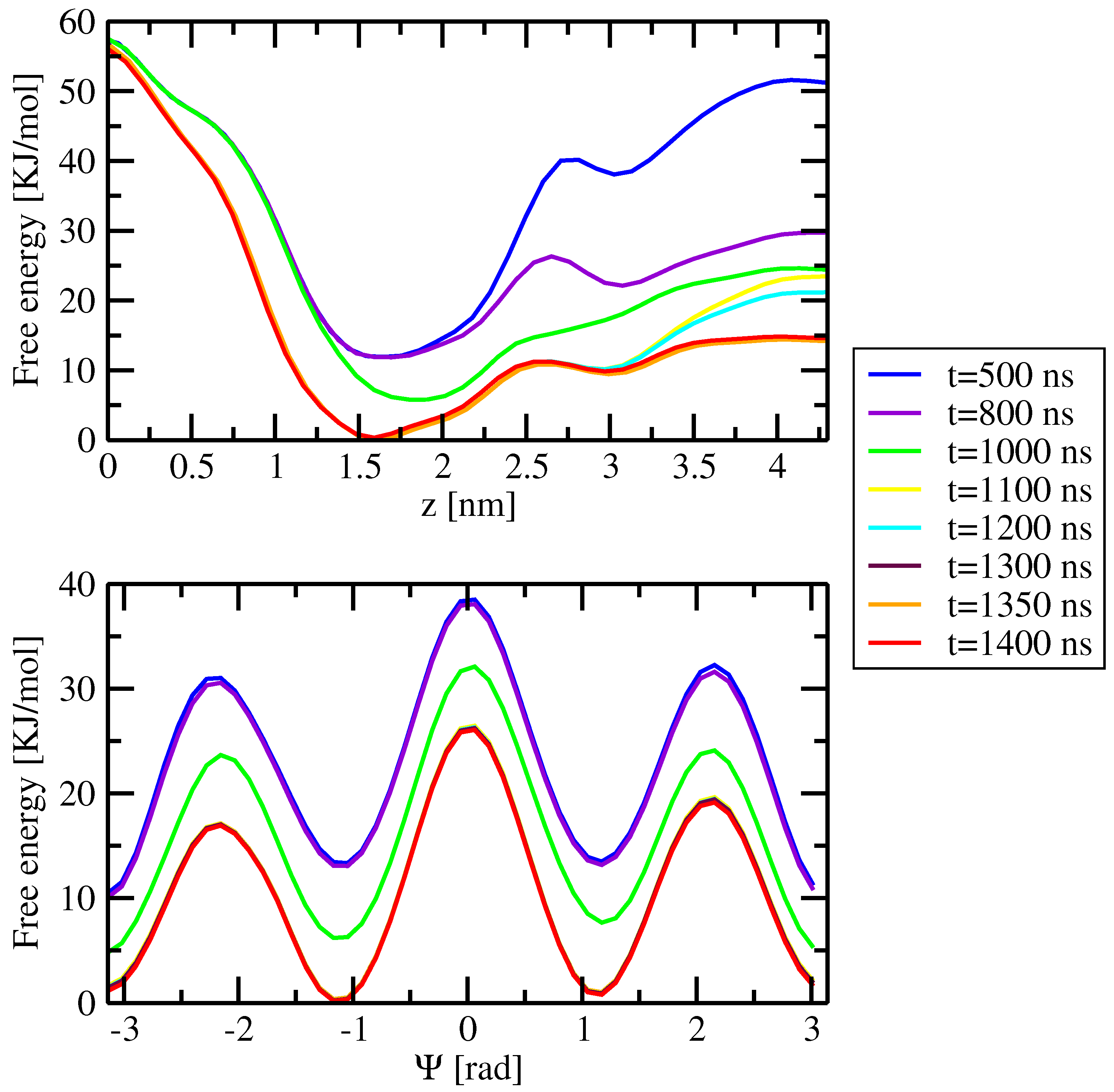
References
- Nagle, J.F.; Tristram-Nagle, S. Structure of lipid bilayers. Biochim. Biophys. Acta (BBA) Rev. Biomembr. 2000, 1469, 159–195. [Google Scholar] [CrossRef]
- Mouritsen, O.G. Life-as a Matter of Fat; Springer-Verlag: Heidelberg, Germany, 2005. [Google Scholar]
- Bassolino-Klimas, D.; Alper, H.E.; Stouch, T.R. Solute diffusion in lipid bilayer membranes: An atomic level study by molecular dynamics simulation. Biochemistry 1993, 32, 12624–12637. [Google Scholar] [CrossRef]
- Berneche, S.; Nina, M.; Roux, B. Molecular dynamics simulation of melittin in a dimyristoylphosphatidylcholine bilayer membrane. Biophys. J. 1998, 75, 1603–1618. [Google Scholar] [CrossRef]
- Högberg, C.J.; Lyubartsev, A.P. A molecular dynamics investigation of the influence of hydration and temperature on structural and dynamical properties of a dimyristoylphosphatidylcholine bilayer. J. Phys. Chem. B 2006, 110, 14326–14336. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.; Calero, C.; Martí, J. Diffusion and spectroscopy of water and lipids in fully hydrated dimyristoylphosphatidylcholine bilayer membranes. J. Chem. Phys. 2014, 140, 03B606_1. [Google Scholar] [CrossRef] [PubMed]
- Mabrey, S.; Sturtevant, J.M. Investigation of phase transitions of lipids and lipid mixtures by sensitivity differential scanning calorimetry. Proc. Natl. Acad. Sci. USA 1976, 73, 3862–3866. [Google Scholar] [CrossRef]
- Almeida, P.F.; Vaz, W.L.; Thompson, T. Lateral diffusion in the liquid phases of dimyristoylphosphatidylcholine/cholesterol lipid bilayers: A free volume analysis. Biochemistry 1992, 31, 6739–6747. [Google Scholar] [CrossRef]
- Clerc, S.G.; Thompson, T.E. Permeability of dimyristoyl phosphatidylcholine/dipalmitoyl phosphatidylcholine bilayer membranes with coexisting gel and liquid-crystalline phases. Biophys. J. 1995, 68, 2333–2341. [Google Scholar] [CrossRef]
- Kučerka, N.; Kiselev, M.A.; Balgavỳ, P. Determination of bilayer thickness and lipid surface area in unilamellar dimyristoylphosphatidylcholine vesicles from small-angle neutron scattering curves: A comparison of evaluation methods. Eur. Biophys. J. 2004, 33, 328–334. [Google Scholar] [CrossRef]
- Edholm, O.; Nagle, J.F. Areas of molecules in membranes consisting of mixtures. Biophys. J. 2005, 89, 1827–1832. [Google Scholar] [CrossRef]
- Vaz, W.L.; Clegg, R.M.; Hallmann, D. Translational diffusion of lipids in liquid crystalline phase phosphatidylcholine multibilayers. A comparison of experiment with theory. Biochemistry 1985, 24, 781–786. [Google Scholar] [CrossRef]
- Tocanne, J.F.; Teissié, J. Ionization of phospholipids and phospholipid-supported interfacial lateral diffusion of protons in membrane model systems. Biochim. Biophys. Acta (BBA) Rev. Biomembr. 1990, 1031, 111–142. [Google Scholar] [CrossRef]
- McLaughlin, S.; Murray, D. Plasma membrane phosphoinositide organization by protein electrostatics. Nature 2005, 438, 605–611. [Google Scholar] [CrossRef]
- Ingólfsson, H.I.; Melo, M.N.; Van Eerden, F.J.; Arnarez, C.; Lopez, C.A.; Wassenaar, T.A.; Periole, X.; De Vries, A.H.; Tieleman, D.P.; Marrink, S.J. Lipid organization of the plasma membrane. J. Am. Chem. Soc. 2014, 136, 14554–14559. [Google Scholar] [CrossRef]
- Zhang, Y.; Chen, X.; Gueydan, C.; Han, J. Plasma membrane changes during programmed cell deaths. Cell Res. 2018, 28, 9–21. [Google Scholar] [CrossRef] [PubMed]
- Krapf, D. Compartmentalization of the plasma membrane. Curr. Opin. Cell Biol. 2018, 53, 15–21. [Google Scholar] [CrossRef]
- Zhang, J.; Jin, R.; Jiang, D.; Chen, H.Y. Electrochemiluminescence-based capacitance microscopy for label-free imaging of antigens on the cellular plasma membrane. J. Am. Chem. Soc. 2019, 141, 10294–10299. [Google Scholar] [CrossRef]
- Wyatt, R.; Kupfer, D.; Sjoerdsma, A.; Engelman, K.; Fram, D.; Snyder, F. Effects of L-tryptophan (a natural sedative) on human sleep. Lancet 1970, 296, 842–846. [Google Scholar] [CrossRef]
- Spinweber, C.L. L-tryptophan administered to chronic sleep-onset insomniacs: Late-appearing reduction of sleep latency. Psychopharmacology 1986, 90, 151–155. [Google Scholar] [CrossRef]
- Jouvet, M. Sleep and serotonin: An unfinished story. Neuropsychopharmacology 1999, 21, 24S–27S. [Google Scholar] [PubMed]
- Slominski, A.; Semak, I.; Pisarchik, A.; Sweatman, T.; Szczesniewski, A.; Wortsman, J. Conversion of L-tryptophan to serotonin and melatonin in human melanoma cells. FEBS Lett. 2002, 511, 102–106. [Google Scholar] [CrossRef]
- Wang, L.; Erlandsen, H.; Haavik, J.; Knappskog, P.M.; Stevens, R.C. Three-dimensional structure of human tryptophan hydroxylase and its implications for the biosynthesis of the neurotransmitters serotonin and melatonin. Biochemistry 2002, 41, 12569–12574. [Google Scholar] [CrossRef] [PubMed]
- Paredes, S.D.; Barriga, C.; Reiter, R.J.; Rodríguez, A.B. Assessment of the potential role of tryptophan as the precursor of serotonin and melatonin for the aged sleep-wake cycle and immune function: Streptopelia risoria as a model. Int. J. Tryptophan Res. 2009, 2, S1129. [Google Scholar] [CrossRef]
- Huether, G.; Kochen, W.; Simat, T.J.; Steinhart, H. Tryptophan, Serotonin, and Melatonin: Basic Aspects and Applications; Springer US-Kluwer Academic/Plenum Publishers: New York, NY, USA, 2012; Volume 467. [Google Scholar]
- Yu, H.S. Melatonin in the Eye: Functional Implications; CRC Press: Boca Raton, FL, USA, 1993; Volume 365. [Google Scholar]
- Mockus, S.M.; Vrana, K.E. Advances in the molecular characterization of tryptophan hydroxylase. J. Mol. Neurosci. 1998, 10, 163–179. [Google Scholar] [CrossRef] [PubMed]
- Kema, I.P.; de Vries, E.G.; Muskiet, F.A. Clinical chemistry of serotonin and metabolites. J. Chromatogr. Biomed. Sci. Appl. 2000, 747, 33–48. [Google Scholar] [CrossRef]
- Lerner, A.; Case, J.; Takahashi, Y.; Lee, T.; Mori, W. Isolation of melatonin, a pineal factor that lights melanocytes. J. Am. Chem. Soc. 1958, 80, 2057–2058. [Google Scholar] [CrossRef]
- Mousavi, S.S.; Shohrati, M.; Vahedi, E.; Abdollahpour-Alitappeh, M.; Panahi, Y. Effect of melatonin administration on sleep quality in sulfur mustard exposed patients with sleep disorders. Iran. J. Pharm. Res. 2018, 17, 136. [Google Scholar]
- Savoca, A.; Manca, D. Physiologically-based pharmacokinetic simulations in pharmacotherapy: Selection of the optimal administration route for exogenous melatonin. Admet. Dmpk. 2019, 7, 44–59. [Google Scholar] [CrossRef]
- Kostoglou-Athanassiou, I. Therapeutic applications of melatonin. Ther. Adv. Endocrinol. Metab. 2013, 4, 13–24. [Google Scholar] [CrossRef]
- Cutolo, M.; Sulli, A.; Pizzorni, C.; Secchi, M.E.; Soldano, S.; Seriolo, B.; Straub, R.H.; Otsa, K.; Maestroni, G.J. Circadian rhythms: Glucocorticoids and arthritis. Ann. N. Y. Acad. Sci. 2006, 1069, 289–299. [Google Scholar] [CrossRef]
- Forrest, C.M.; Mackay, G.M.; Stoy, N.; Stone, T.W.; Darlington, L.G. Inflammatory status and kynurenine metabolism in rheumatoid arthritis treated with melatonin. Br. J. Clin. Pharmacol. 2007, 64, 517–526. [Google Scholar] [CrossRef]
- Bang, J.; Chang, H.W.; Jung, H.R.; Cho, C.H.; Hur, J.A.; Lee, S.I.; Choi, T.H.; Kim, S.H.; Ha, E. Melatonin attenuates clock gene Cryptochrome1, which may aggravates mouse anti-type II collagen antibody-induced arthritis. Rheumatol. Int. 2012, 32, 379–385. [Google Scholar] [CrossRef]
- Huang, C.C.; Chiou, C.H.; Liu, S.C.; Hu, S.L.; Su, C.M.; Tsai, C.H.; Tang, C.H. Melatonin attenuates TNF-α and IL-1β expression in synovial fibroblasts and diminishes cartilage degradation: Implications for the treatment of rheumatoid arthritis. J. Pineal Res. 2019, 66, e12560. [Google Scholar] [CrossRef]
- Rusanova, I.; Martínez-Ruiz, L.; Florido, J.; Rodríguez-Santana, C.; Guerra-Librero, A.; Acuña-Castroviejo, D.; Escames, G. Protective effects of melatonin on the skin: Future perspectives. Int. J. Mol. Sci. 2019, 20, 4948. [Google Scholar] [CrossRef]
- Slominski, A.T.; Zmijewski, M.A.; Semak, I.; Kim, T.K.; Janjetovic, Z.; Slominski, R.M.; Zmijewski, J.W. Melatonin, mitochondria, and the skin. Cell. Mol. Life Sci. 2017, 74, 3913–3925. [Google Scholar] [CrossRef]
- Slominski, A.T.; Hardeland, R.; Zmijewski, M.A.; Slominski, R.M.; Reiter, R.J.; Paus, R. Melatonin: A cutaneous perspective on its production, metabolism, and functions. J. Investig. Dermatol. 2018, 138, 490–499. [Google Scholar] [CrossRef] [PubMed]
- Hussain, S.A.R. Effect of melatonin on cholesterol absorption in rats. J. Pineal Res. 2007, 42, 267–271. [Google Scholar] [CrossRef] [PubMed]
- Costa, E.; Lopes, R.; Lamy-Freund, M.T. Permeability of pure lipid bilayers to melatonin. J. Pineal Res. 1995, 19, 123–126. [Google Scholar] [CrossRef]
- Bongiorno, D.; Ceraulo, L.; Ferrugia, M.; Filizzola, F.; Ruggirello, A.; Liveri, V.T. Localization and interactions of melatonin in dry cholesterol/lecithin mixed reversed micelles used as cell membrane models. J. Pineal Res. 2005, 38, 292–298. [Google Scholar] [CrossRef]
- Acuña-Castroviejo, D.; Escames, G.; Macías, M.; Muñoz Hoyos, A.; Carballo Molina, A.; Arauzo, M.; Montes, R.; Vives, F. Minireview: Cell protective role of melatonin in the brain. J. Pineal Res. 1995, 19, 57–63. [Google Scholar] [CrossRef]
- Maestroni, G.J. The immunotherapeutic potential of melatonin. Expert Opin. Investig. Drugs 2001, 10, 467–476. [Google Scholar] [CrossRef] [PubMed]
- Dies, H.; Toppozini, L.; Rheinstädter, M.C. The interaction between amyloid-β peptides and anionic lipid membranes containing cholesterol and melatonin. PLoS ONE 2014, 9, e99124. [Google Scholar] [CrossRef] [PubMed]
- Zhang, R.; Wang, X.; Ni, L.; Di, X.; Ma, B.; Niu, S.; Liu, C.; Reiter, R.J. COVID-19: Melatonin as a potential adjuvant treatment. Life Sci. 2020, 250, 117583. [Google Scholar] [CrossRef] [PubMed]
- Pandi-Perumal, S.R.; Srinivasan, V.; Maestroni, G.; Cardinali, D.; Poeggeler, B.; Hardeland, R. Melatonin: Nature’s most versatile biological signal? FEBS J. 2006, 273, 2813–2838. [Google Scholar] [CrossRef] [PubMed]
- Hardeland, R.; Cardinali, D.P.; Srinivasan, V.; Spence, D.W.; Brown, G.M.; Pandi-Perumal, S.R. Melatonin—A pleiotropic, orchestrating regulator molecule. Prog. Neurobiol. 2011, 93, 350–384. [Google Scholar] [CrossRef]
- Tan, D.X.; Manchester, L.C.; Esteban-Zubero, E.; Zhou, Z.; Reiter, R.J. Melatonin as a potent and inducible endogenous antioxidant: Synthesis and metabolism. Molecules 2015, 20, 18886–18906. [Google Scholar] [CrossRef]
- Severcan, F.; Sahin, I.; Kazancı, N. Melatonin strongly interacts with zwitterionic model membranes—Evidence from Fourier transform infrared spectroscopy and differential scanning calorimetry. Biochim. Biophys. Acta (BBA) Biomembr. 2005, 1668, 215–222. [Google Scholar] [CrossRef]
- Dies, H.; Cheung, B.; Tang, J.; Rheinstädter, M.C. The organization of melatonin in lipid membranes. Biochim. Biophys. Acta (BBA) Biomembr. 2015, 1848, 1032–1040. [Google Scholar] [CrossRef]
- de Jesus, A.J.; Allen, T.W. The role of tryptophan side chains in membrane protein anchoring and hydrophobic mismatch. Biochim. Biophys. Acta (BBA) Biomembr. 2013, 1828, 864–876. [Google Scholar] [CrossRef]
- Costa, E.J.; Shida, C.S.; Biaggi, M.H.; Ito, A.S.; Lamy-Freund, M.T. How melatonin interacts with lipid bilayers: A study by fluorescence and ESR spectroscopies. FEBS Lett. 1997, 416, 103–106. [Google Scholar] [CrossRef]
- Yau, W.M.; Wimley, W.C.; Gawrisch, K.; White, S.H. The preference of tryptophan for membrane interfaces. Biochemistry 1998, 37, 14713–14718. [Google Scholar] [CrossRef]
- MacCallum, J.L.; Bennett, W.D.; Tieleman, D.P. Distribution of amino acids in a lipid bilayer from computer simulations. Biophys. J. 2008, 94, 3393–3404. [Google Scholar] [CrossRef]
- Yu, H.; Dickson, E.J.; Jung, S.R.; Koh, D.S.; Hille, B. High membrane permeability for melatonin. J. Gen. Physiol. 2016, 147, 63–76. [Google Scholar] [CrossRef]
- Hevia, D.; Sainz, R.M.; Blanco, D.; Quirós, I.; Tan, D.X.; Rodríguez, C.; Mayo, J.C. Melatonin uptake in prostate cancer cells: Intracellular transport versus simple passive diffusion. J. Pineal Res. 2008, 45, 247–257. [Google Scholar] [CrossRef]
- Hevia, D.; González-Menéndez, P.; Quiros-González, I.; Miar, A.; Rodríguez-García, A.; Tan, D.X.; Reiter, R.J.; Mayo, J.C.; Sainz, R.M. Melatonin uptake through glucose transporters: A new target for melatonin inhibition of cancer. J. Pineal Res. 2015, 58, 234–250. [Google Scholar] [CrossRef] [PubMed]
- Andersen, L.P.H.; Gögenur, I.; Rosenberg, J.; Reiter, R.J. The safety of melatonin in humans. Clin. Drug Investig. 2016, 36, 169–175. [Google Scholar] [CrossRef]
- Drolle, E.; Kučerka, N.; Hoopes, M.; Choi, Y.; Katsaras, J.; Karttunen, M.; Leonenko, Z. Effect of melatonin and cholesterol on the structure of DOPC and DPPC membranes. Biochim. Biophys. Acta (BBA) Biomembr. 2013, 1828, 2247–2254. [Google Scholar] [CrossRef] [PubMed]
- Choi, Y.; Attwood, S.J.; Hoopes, M.I.; Drolle, E.; Karttunen, M.; Leonenko, Z. Melatonin directly interacts with cholesterol and alleviates cholesterol effects in dipalmitoylphosphatidylcholine monolayers. Soft Matter 2014, 10, 206–213. [Google Scholar] [CrossRef]
- Park, K.R.; Kim, E.C.; Hong, J.T.; Yun, H.M. Dysregulation of 5-hydroxytryptamine 6 receptor accelerates maturation of bone-resorbing osteoclasts and induces bone loss. Theranostics 2018, 8, 3087. [Google Scholar] [CrossRef]
- Rapport, M.M.; Green, A.A.; Page, I.H. Crystalline serotonin. Science 1948, 108, 329–330. [Google Scholar] [CrossRef] [PubMed]
- Berger, M.; Gray, J.A.; Roth, B.L. The expanded biology of serotonin. Annu. Rev. Med. 2009, 60, 355–366. [Google Scholar] [CrossRef] [PubMed]
- Daubert, E.A.; Condron, B.G. Serotonin: A regulator of neuronal morphology and circuitry. Trends Neurosci. 2010, 33, 424–434. [Google Scholar] [CrossRef] [PubMed]
- Hartmann, E. Effects of L-tryptophan on sleepiness and on sleep. J. Psychiatr. Res. 1982, 17, 107–113. [Google Scholar] [CrossRef]
- Schneider-Helmert, D.; Spinweber, C.L. Evaluation of L-tryptophan for treatment of insomnia: A review. Psychopharmacology 1986, 89, 1–7. [Google Scholar] [CrossRef]
- Frenkel, D.; Smit, B. Understanding Molecular Simulation: From Algorithms to Applications; Academic Press: San Diego, CA, USA, 2001; Volume 1. [Google Scholar]
- Martí, J.; Diaz, B. Efficient recursive Adams–Bashforth methods in molecular dynamics simulations of N-body systems interacting through pairwise potentials. Mol. Simul. 2020, 46, 1248–1254. [Google Scholar] [CrossRef]
- Martí, J.; Padró, J.; Guardia, E. Computer simulation of molecular motions in liquids: Infrared spectra of water and heavy water. Mol. Simul. 1993, 11, 321–336. [Google Scholar] [CrossRef]
- Martí, J.; Gordillo, M. Microscopic dynamics of confined supercritical water. Chem. Phys. Lett. 2002, 354, 227–232. [Google Scholar] [CrossRef]
- Nagy, G.; Gordillo, M.; Guàrdia, E.; Martí, J. Liquid water confined in carbon nanochannels at high temperatures. J. Phys. Chem. B 2007, 111, 12524–12530. [Google Scholar] [CrossRef]
- Rodriguez, J.; Laria, D.; Guardia, E.; Martí, J. Dynamics of water nanodroplets and aqueous protons in non-ionic reverse micelles. Phys. Chem. Chem. Phys. 2009, 11, 1484–1490. [Google Scholar] [CrossRef][Green Version]
- Sala, J.; Guardia, E.; Martí, J. Specific ion effects in aqueous eletrolyte solutions confined within graphene sheets at the nanometric scale. Phys. Chem. Chem. Phys. 2012, 14, 10799–10808. [Google Scholar] [CrossRef]
- Calero, C.; Gordillo, M.; Martí, J. Size effects on water adsorbed on hydrophobic probes at the nanometric scale. J. Chem. Phys. 2013, 138, 214702. [Google Scholar] [CrossRef]
- Rodriguez, J.; Elola, M.D.; Martí, J.; Laria, D. Surface behavior of aprotic mixtures: Dimethyl sulfoxide/acetonitrile. J. Phys. Chem. C 2017, 121, 14618–14627. [Google Scholar] [CrossRef]
- MacKerell, A.D., Jr.; Banavali, N.K. All-atom empirical force field for nucleic acids: II. Application to molecular dynamics simulations of DNA and RNA in solution. J. Comput. Chem. 2000, 21, 105–120. [Google Scholar] [CrossRef]
- Ponomarev, S.Y.; Thayer, K.M.; Beveridge, D.L. Ion motions in molecular dynamics simulations on DNA. Proc. Natl. Acad. Sci. USA 2004, 101, 14771–14775. [Google Scholar] [CrossRef] [PubMed]
- Johnson, R.R.; Johnson, A.C.; Klein, M.L. Probing the structure of DNA- carbon nanotube hybrids with molecular dynamics. Nano Lett. 2008, 8, 69–75. [Google Scholar] [CrossRef] [PubMed]
- Van der Ploeg, P.; Berendsen, H. Molecular dynamics simulation of a bilayer membrane. J. Chem. Phys. 1982, 76, 3271–3276. [Google Scholar] [CrossRef]
- Egberts, E.; Marrink, S.J.; Berendsen, H.J. Molecular dynamics simulation of a phospholipid membrane. Eur. Biophys. J. 1994, 22, 423–436. [Google Scholar] [CrossRef]
- Berkowitz, M.L. Detailed molecular dynamics simulations of model biological membranes containing cholesterol. Biochim. Biophys. Acta (BBA) Biomembr. 2009, 1788, 86–96. [Google Scholar] [CrossRef] [PubMed]
- Gedeon, P.C.; Indarte, M.; Surratt, C.K.; Madura, J.D. Molecular dynamics of leucine and dopamine transporter proteins in a model cell membrane lipid bilayer. Proteins Struct. Funct. Bioinform. 2010, 78, 797–811. [Google Scholar] [CrossRef]
- Venable, R.M.; Kramer, A.; Pastor, R.W. Molecular dynamics simulations of membrane permeability. Chem. Rev. 2019, 119, 5954–5997. [Google Scholar] [CrossRef] [PubMed]
- Senn, H.M.; Thiel, W. QM/MM Methods for Biological Systems. In Atomistic Approaches in Modern Biology; Springer-Verlag: Berlin/Heidelberg, Germany, 2006; pp. 173–290. [Google Scholar]
- Martí, J.; Csajka, F.S.; Chandler, D. Stochastic transition pathways in the aqueous sodium chloride dissociation process. Chem. Phys. Lett. 2000, 328, 169–176. [Google Scholar] [CrossRef]
- Martí, J.; Csajka, F. The aqueous solvation of sodium chloride: A Monte Carlo transition path sampling study. J. Chem. Phys. 2000, 113, 1154–1161. [Google Scholar] [CrossRef]
- Geissler, P.L.; Dellago, C.; Chandler, D.; Hutter, J.; Parrinello, M. Autoionization in liquid water. Science 2001, 291, 2121–2124. [Google Scholar] [CrossRef]
- Martí, J. Transition path sampling study of the local molecular structure in the aqueous solvation of sodium chloride. Mol. Simul. 2001, 27, 169–185. [Google Scholar] [CrossRef]
- Dellago, C.; Bolhuis, P.G.; Geissler, P.L. Transition path sampling. Adv. Chem. Phys. 2002, 123, 1–78. [Google Scholar]
- Martí, J.; Csajka, F.S. Transition path sampling study of flip-flop transitions in model lipid bilayer membranes. Phys. Rev. E 2004, 69, 061918. [Google Scholar] [CrossRef] [PubMed]
- Dellago, C.; Bolhuis, P.G. Transition Path Sampling simulations of biological systems. In Atomistic Approaches in Modern Biology; Springer: New York, NY, USA, 2006; pp. 291–317. [Google Scholar]
- Henin, J.; Fiorin, G.; Chipot, C.; Klein, M.L. Exploring multidimensional free energy landscapes using time-dependent biases on collective variables. J. Chem. Theory Comput. 2009, 6, 35–47. [Google Scholar] [CrossRef]
- Mezei, M. Adaptive umbrella sampling: Self-consistent determination of the non-Boltzmann bias. J. Comput. Phys. 1987, 68, 237–248. [Google Scholar] [CrossRef]
- Bartels, C.; Karplus, M. Multidimensional adaptive umbrella sampling: Applications to main chain and side chain peptide conformations. J. Comput. Chem. 1997, 18, 1450–1462. [Google Scholar] [CrossRef]
- Calero, C.; Martí, J.; Guàrdia, E.; Masia, M. Characterization of the methane–graphene hydrophobic interaction in aqueous solution from ab initio simulations. J. Chem. Theory Comput. 2013, 9, 5070–5075. [Google Scholar] [CrossRef] [PubMed]
- Trzesniak, D.; Kunz, A.P.E.; van Gunsteren, W.F. A comparison of methods to compute the potential of mean force. ChemPhysChem 2007, 8, 162–169. [Google Scholar] [CrossRef] [PubMed]
- Lu, H.; Martí, J. Binding free energies of small-molecules in phospholipid membranes: Aminoacids, serotonin and melatonin. Chem. Phys. Lett. 2018, 712, 190–195. [Google Scholar] [CrossRef]
- Barducci, A.; Bonomi, M.; Parrinello, M. Metadynamics. Wiley Interdiscip. Rev. Comput. Mol. Sci. 2011, 1, 826–843. [Google Scholar] [CrossRef]
- Bussi, G.; Gervasio, F.L.; Laio, A.; Parrinello, M. Free-energy landscape for β hairpin folding from combined parallel tempering and metadynamics. J. Am. Chem. Soc. 2006, 128, 13435–13441. [Google Scholar] [CrossRef] [PubMed]
- Deighan, M.; Bonomi, M.; Pfaendtner, J. Efficient simulation of explicitly solvated proteins in the well-tempered ensemble. J. Chem. Theory Comput. 2012, 8, 2189–2192. [Google Scholar] [CrossRef]
- Palmer, J.C.; Car, R.; Debenedetti, P.G. The liquid–liquid transition in supercooled ST2 water: A comparison between umbrella sampling and well-tempered metadynamics. Faraday Discuss. 2013, 167, 77–94. [Google Scholar] [CrossRef]
- Haldar, S.; Kührová, P.; Banáš, P.; Spiwok, V.; Sponer, J.; Hobza, P.; Otyepka, M. Insights into stability and folding of GNRA and UNCG tetraloops revealed by microsecond molecular dynamics and well-tempered metadynamics. J. Chem. Theory Comput. 2015, 11, 3866–3877. [Google Scholar] [CrossRef] [PubMed]
- Martí, J. Free-energy surfaces of ionic adsorption in cholesterol-free and cholesterol-rich phospholipid membranes. Mol. Simul. 2018, 44, 1136–1146. [Google Scholar] [CrossRef]
- Poger, D.; Mark, A.E. On the validation of molecular dynamics simulations of saturated and cis-monounsaturated phosphatidylcholine lipid bilayers: A comparison with experiment. J. Chem. Theory Comput. 2010, 6, 325–336. [Google Scholar] [CrossRef]
- Pandey, P.R.; Roy, S. Headgroup mediated water insertion into the DPPC bilayer: A molecular dynamics study. J. Phys. Chem. B 2011, 115, 3155–3163. [Google Scholar] [CrossRef]
- Lu, H.; Martí, J. Effects of cholesterol on the binding of the precursor neurotransmitter tryptophan to zwitterionic membranes. J. Chem. Phys. 2018, 149, 164906. [Google Scholar] [CrossRef] [PubMed]
- Lu, H.; Martí, J. Binding and dynamics of melatonin at the interface of phosphatidylcholine-cholesterol membranes. PLoS ONE 2019, 14, e0224624. [Google Scholar] [CrossRef]
- Petrache, H.I.; Dodd, S.W.; Brown, M.F. Area per lipid and acyl length distributions in fluid phosphatidylcholines determined by 2H NMR spectroscopy. Biophys. J. 2000, 79, 3172–3192. [Google Scholar] [CrossRef]
- Kučerka, N.; Nieh, M.P.; Katsaras, J. Fluid phase lipid areas and bilayer thicknesses of commonly used phosphatidylcholines as a function of temperature. Biochim. Biophys. Acta (BBA) Biomembr. 2011, 1808, 2761–2771. [Google Scholar] [CrossRef]
- Chiu, S.; Jakobsson, E.; Mashl, R.J.; Scott, H.L. Cholesterol-induced modifications in lipid bilayers: A simulation study. Biophys. J. 2002, 83, 1842–1853. [Google Scholar] [CrossRef]
- Hofsäß, C.; Lindahl, E.; Edholm, O. Molecular dynamics simulations of phospholipid bilayers with cholesterol. Biophys. J. 2003, 84, 2192–2206. [Google Scholar] [CrossRef]
- Wang, Y.; Gkeka, P.; Fuchs, J.E.; Liedl, K.R.; Cournia, Z. DPPC-cholesterol phase diagram using coarse-grained Molecular Dynamics simulations. Biochim. Biophys. Acta (BBA) Biomembr. 2016, 1858, 2846–2857. [Google Scholar] [CrossRef]
- Armstrong, C.L.; Barrett, M.A.; Hiess, A.; Salditt, T.; Katsaras, J.; Shi, A.C.; Rheinstädter, M.C. Effect of cholesterol on the lateral nanoscale dynamics of fluid membranes. Eur. Biophys. J. 2012, 41, 901–913. [Google Scholar] [CrossRef]
- Yang, J.; Martí, J.; Calero, C. Pair interactions among ternary DPPC/POPC/cholesterol mixtures in liquid-ordered and liquid-disordered phases. Soft Matter 2016, 12, 4557–4561. [Google Scholar] [CrossRef]
- Abankwa, D.; Gorfe, A.A.; Inder, K.; Hancock, J.F. Ras membrane orientation and nanodomain localization generate isoform diversity. Proc. Natl. Acad. Sci. USA 2010, 107, 1130–1135. [Google Scholar] [CrossRef]
- Kapoor, S.; Triola, G.; Vetter, I.R.; Erlkamp, M.; Waldmann, H.; Winter, R. Revealing conformational substates of lipidated N-Ras protein by pressure modulation. Proc. Natl. Acad. Sci. USA 2012, 109, 460–465. [Google Scholar] [CrossRef] [PubMed]
- Mazhab-Jafari, M.T.; Marshall, C.B.; Smith, M.J.; Gasmi-Seabrook, G.M.; Stathopulos, P.B.; Inagaki, F.; Kay, L.E.; Neel, B.G.; Ikura, M. Oncogenic and RASopathy-associated K-RAS mutations relieve membrane-dependent occlusion of the effector-binding site. Proc. Natl. Acad. Sci. USA 2015, 112, 6625–6630. [Google Scholar] [CrossRef] [PubMed]
- Prakash, P.; Zhou, Y.; Liang, H.; Hancock, J.F.; Gorfe, A.A. Oncogenic K-Ras binds to an anionic membrane in two distinct orientations: A molecular dynamics analysis. Biophys. J. 2016, 110, 1125–1138. [Google Scholar] [CrossRef]
- Prakash, P.; Litwin, D.; Liang, H.; Sarkar-Banerjee, S.; Dolino, D.; Zhou, Y.; Hancock, J.F.; Jayaraman, V.; Gorfe, A.A. Dynamics of membrane-bound G12V-KRAS from simulations and single-molecule FRET in native nanodiscs. Biophys. J. 2019, 116, 179–183. [Google Scholar] [CrossRef]
- Laio, A.; Parrinello, M. Escaping free-energy minima. Proc. Natl. Acad. Sci. USA 2002, 99, 12562–12566. [Google Scholar] [CrossRef]
- Barducci, A.; Bussi, G.; Parrinello, M. Well-tempered metadynamics: A smoothly converging and tunable free-energy method. Phys. Rev. Lett. 2008, 100, 020603. [Google Scholar] [CrossRef]
- Chandler, D. Introduction to Modern Statistical Mechanics; Oxford University Press: Oxford, UK, 1987; Volume 40. [Google Scholar]
- Peters, G.H.; Werge, M.; Elf-Lind, M.N.; Madsen, J.J.; Velardez, G.F.; Westh, P. Interaction of neurotransmitters with a phospholipid bilayer: A molecular dynamics study. Chem. Phys. Lipids 2014, 184, 7–17. [Google Scholar] [CrossRef]
- Jämbeck, J.P.; Lyubartsev, A.P. Exploring the free energy landscape of solutes embedded in lipid bilayers. J. Phys. Chem. Lett. 2013, 4, 1781–1787. [Google Scholar] [CrossRef]
- Florio, G.M.; Christie, R.A.; Jordan, K.D.; Zwier, T.S. Conformational preferences of jet-cooled melatonin: Probing trans-and cis-amide regions of the potential energy surface. J. Am. Chem. Soc. 2002, 124, 10236–10247. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Gallagher, E.; Jorgensen, C.; Troendle, E.P.; Hu, D.; Searson, P.C.; Ulmschneider, M.B. An experimentally validated approach to calculate the blood-brain barrier permeability of small molecules. Sci. Rep. 2019, 9, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Jo, S.; Kim, T.; Iyer, V.G.; Im, W. CHARMM-GUI: A web-based graphical user interface for CHARMM. J. Comput. Chem. 2008, 29, 1859–1865. [Google Scholar] [CrossRef]
- Klauda, J.B.; Venable, R.M.; Freites, J.A.; O’Connor, J.W.; Tobias, D.J.; Mondragon-Ramirez, C.; Vorobyov, I.; MacKerell, A.D., Jr.; Pastor, R.W. Update of the CHARMM all-atom additive force field for lipids: Validation on six lipid types. J. Phys. Chem. B 2010, 114, 7830–7843. [Google Scholar] [CrossRef]
- Lim, J.B.; Rogaski, B.; Klauda, J.B. Update of the cholesterol force field parameters in CHARMM. J. Phys. Chem. B 2012, 116, 203–210. [Google Scholar] [CrossRef] [PubMed]
- Essmann, U.; Perera, L.; Berkowitz, M.L.; Darden, T.; Lee, H.; Pedersen, L.G. A smooth particle mesh Ewald method. J. Chem. Phys. 1995, 103, 8577–8593. [Google Scholar] [CrossRef]
- Ytreberg, F.M.; Swendsen, R.H.; Zuckerman, D.M. Comparison of free energy methods for molecular systems. J. Chem. Phys. 2006, 125, 184114. [Google Scholar] [CrossRef]
- Huber, T.; Torda, A.E.; Van Gunsteren, W.F. Local elevation: A method for improving the searching properties of molecular dynamics simulation. J. Comput. Aided Mol. Des. 1994, 8, 695–708. [Google Scholar] [CrossRef]
- Grubmüller, H. Predicting slow structural transitions in macromolecular systems: Conformational flooding. Phys. Rev. E 1995, 52, 2893. [Google Scholar] [CrossRef]
- Bonomi, M.; Parrinello, M. Enhanced sampling in the well-tempered ensemble. Phys. Rev. Lett. 2010, 104, 190601. [Google Scholar] [CrossRef] [PubMed]
- Bonomi, M.; Branduardi, D.; Bussi, G.; Camilloni, C.; Provasi, D.; Raiteri, P.; Donadio, D.; Marinelli, F.; Pietrucci, F.; Broglia, R.A.; et al. PLUMED: A portable plugin for free-energy calculations with molecular dynamics. Comput. Phys. Commun. 2009, 180, 1961–1972. [Google Scholar] [CrossRef]
- Tribello, G.A.; Bonomi, M.; Branduardi, D.; Camilloni, C.; Bussi, G. PLUMED 2: New feathers for an old bird. Comput. Phys. Commun. 2014, 185, 604–613. [Google Scholar] [CrossRef]
- Martí, J.; Lu, H. Molecular dynamics of di-palmitoyl-phosphatidyl-choline biomembranes in ionic solution: Adsorption of the precursor neurotransmitter tryptophan. Procedia Comput. Sci. 2017, 108, 1242–1250. [Google Scholar] [CrossRef][Green Version]
- Liu, H.; Zhang, H.; Jin, B. Fluorescence of tryptophan in aqueous solution. Spectrochim. Acta Part Mol. Biomol. Spectrosc. 2013, 106, 54–59. [Google Scholar] [CrossRef]
- Lu, H.; Martí, J. Long-lasting Salt Bridges Provide the Anchoring Mechanism of Oncogenic Kirsten Rat Sarcoma Proteins at Cell Membranes. J. Phys. Chem. Lett. 2020, 11, 9938–9945. [Google Scholar] [CrossRef]
| Small Molecule and Cholesterol Percentage | Phospholipid Species | A (nm) | z (nm) |
|---|---|---|---|
| TRP-0% | DPPC | 0.614 (0.008) | 3.97 (0.05) |
| TRP-30% | DPPC | 0.408 (0.002) | 4.89 (0.04) |
| TRP-50% | DPPC | 0.401 (0.002) | 4. 78 (0.03) |
| SER-0% | DPPC | 0.613 (0.015) | 3.83 (0.05) |
| MEL-0% | DMPC | 0.618 (0.005) | 3.49 (0.06) |
| MEL-30% | DMPC | 0.421 (0.007) | 4.43 (0.03) |
| MEL-50% | DMPC | 0.402 (0.008) | 4.47 (0.03) |
| Probe (Active Site) | O2-DPPC | O8-DPPC |
|---|---|---|
| H1 TRP | 11.29 | 7.53 |
| H2 TRP | 8.02 | 4.18 |
| H1 SER | 7.95 | 6.53 |
| H4 SER | 7.45 | 7.87 |
| H15 MEL | - | 4.85 |
| H16 MEL | 8.03 | 1.97 |
| Cholesterol Percentage | Folded-Extended | Interface-Bulk | Interface to Internal Regions |
|---|---|---|---|
| 0 % | 18.8 | 25.3 | 40.2 |
| 30 % | 19.7 | 14.1 | 50.7 |
| 50 % | 17.6 | 9.1 | 55.5 |
| Small Molecule | 0% CHOL | 30% CHOL | 50% CHOL |
|---|---|---|---|
| TRP | 3.48(0.80) | 2.91(0.35) | 14.0(0.2) |
| MEL | 1.1(0.4) | 3.9(0.6) | 4.1(0.9) |
| Phospholipids | Small Molecule | Waters | Total Length (ns) | Temperature (K) | Ion Pairs |
|---|---|---|---|---|---|
| 204 DPPC | TRP | 4962 | 800 | 323.15 | 17 Na + 17 Cl |
| 204 DPPC | SER | 4962 | 800 | 323.15 | 17 Na + 17 Cl |
| 204 DMPC | MEL | 10250 | 800 | 303.15 | 21 Na + 21 Cl |
| Parameter | 0% | 30% | 50% |
|---|---|---|---|
| Gaussian width of CV1 [nm] | 0.30 | 0.30 | 0.25 |
| Gaussian width of CV2 [degrees] | 20 | 20 | 20 |
| Starting (Gaussian) hill [kJ/mol] | 1.0 | 1.0 | 1.0 |
| Deposition stride [ps] | 1 | 1 | 1 |
| Bias factor | 10 | 10 | 20 |
| Simulation time [ns] | 1100 | 1400 | 1400 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Martí, J.; Lu, H. Microscopic Interactions of Melatonin, Serotonin and Tryptophan with Zwitterionic Phospholipid Membranes. Int. J. Mol. Sci. 2021, 22, 2842. https://doi.org/10.3390/ijms22062842
Martí J, Lu H. Microscopic Interactions of Melatonin, Serotonin and Tryptophan with Zwitterionic Phospholipid Membranes. International Journal of Molecular Sciences. 2021; 22(6):2842. https://doi.org/10.3390/ijms22062842
Chicago/Turabian StyleMartí, Jordi, and Huixia Lu. 2021. "Microscopic Interactions of Melatonin, Serotonin and Tryptophan with Zwitterionic Phospholipid Membranes" International Journal of Molecular Sciences 22, no. 6: 2842. https://doi.org/10.3390/ijms22062842
APA StyleMartí, J., & Lu, H. (2021). Microscopic Interactions of Melatonin, Serotonin and Tryptophan with Zwitterionic Phospholipid Membranes. International Journal of Molecular Sciences, 22(6), 2842. https://doi.org/10.3390/ijms22062842







