Genophenotypic Factors and Pharmacogenomics in Adverse Drug Reactions
Abstract
1. Introduction
2. ADR-Associated Factors
2.1. Age-Related ADRs
2.2. Sex-Related ADRs
2.3. Ethnic Differences in ADRs
3. Cardiovascular Disorders
4. Cancer
5. Central Nervous System Disorders
6. Immunopharmacogenomics and Cutaneous ADRs
7. COVID-19
8. Future Trends
9. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Osanlou, O.; Pirmohamed, M.; Daly, A.K. Pharmacogenetics of Adverse Drug Reactions. Adv. Pharmacol. 2018, 83, 155–190. [Google Scholar] [CrossRef]
- Ross, C.J.; Carleton, B.; Warn, D.G.; Stenton, S.B.; Rassekh, S.R.; Hayden, M.R. Genotypic approaches to therapy in children: A national active surveillance network (GATC) to study the pharmacogenomics of severe adverse drug reactions in children. Ann. N. Y. Acad. Sci. 2007, 1110, 177–192. [Google Scholar] [CrossRef]
- Eissenberg, J.C.; Aurora, R. Pharmacogenomics: What the Doctor Ordered? Mo. Med. 2019, 116, 217–225. [Google Scholar]
- Becquemont, L. Pharmacogenomics of adverse drug reactions: Practical applications and perspectives. Pharmacogenomics 2009, 10, 961–969. [Google Scholar] [CrossRef]
- Elzagallaai, A.A.; Greff, M.; Rieder, M.J. Adverse Drug Reactions in Children: The Double-Edged Sword of Therapeutics. Clin. Pharmacol. Ther. 2017, 101, 725–735. [Google Scholar] [CrossRef] [PubMed]
- Malki, M.A.; Pearson, E.R. Drug-drug-gene interactions and adverse drug reactions. Pharm. J. 2020, 20, 355–366. [Google Scholar] [CrossRef]
- Pirmohamed, M. Personalized pharmacogenomics: Predicting efficacy and adverse drug reactions. Annu. Rev. Genom. Hum. Genet. 2014, 15, 349–370. [Google Scholar] [CrossRef]
- Aagaard, L.; Hansen, E.H. Information about ADRs explored by pharmacovigilance approaches: A qualitative review of studies on antibiotics, SSRIs and NSAIDs. BMC Clin. Pharmacol. 2009, 9, 4. [Google Scholar] [CrossRef]
- Magro, L.; Moretti, U.; Leone, R. Epidemiology and characteristics of adverse drug reactions caused by drug-drug interactions. Expert Opin. Drug Saf. 2012, 11, 83–94. [Google Scholar] [CrossRef]
- Weinshilboum, R.M.; Wang, L. Pharmacogenomics: Precision Medicine and Drug Response. Mayo Clin. Proc. 2017, 92, 1711–1722. [Google Scholar] [CrossRef]
- Cacabelos, R.; Cacabelos, N.; Carril, J.C. The role of pharmacogenomics in adverse drug reactions. Expert Rev. Clin. Pharmacol. 2019, 12, 407–442. [Google Scholar] [CrossRef]
- Cacabelos, R. Pharmacogenomic of drugs to treat brain disorders. Expert Rev. Prec. Med. Drug Dev. 2020, 5, 181–234. [Google Scholar] [CrossRef]
- Roden, D.M.; McLeod, H.L.; Relling, M.V.; Williams, M.S.; Mensah, G.A.; Peterson, J.F.; van Driest, S.L. Pharmacogenomics. Lancet 2019, 394, 521–532. [Google Scholar] [CrossRef]
- Borroni, R.G. Role of dermatology in pharmacogenomics: Drug-induced skin injury. Pharmacogenomics 2015, 16, 401–412. [Google Scholar] [CrossRef] [PubMed]
- Davies, E.C.; Green, C.F.; Mottram, D.R.; Pirmohamed, M. Adverse drug reactions in hospitals: A narrative review. Curr. Drug Saf. 2007, 2, 79–87. [Google Scholar] [CrossRef] [PubMed]
- Chan, S.L.; Ang, X.; Sani, L.L.; Ng, H.Y.; Winther, M.D.; Liu, J.J.; Brunham, L.R.; Chan, A. Prevalence and characteristics of adverse drug reactions at admission to hospital: A prospective observational study. Br. J. Clin. Pharmacol. 2016, 82, 1636–1646. [Google Scholar] [CrossRef] [PubMed]
- Empey, P.E. Genetic predisposition to adverse drug reactions in the intensive care unit. Crit. Care Med. 2010, 38, S106–S116. [Google Scholar] [CrossRef] [PubMed]
- Alessandrini, M.; Chaudhry, M.; Dodgen, T.M.; Pepper, M.S. Pharmacogenomics and Global Precision Medicine in the Context of Adverse Drug Reactions: Top 10 Opportunities and Challenges for the Next Decade. Omics A J. Integr. Biol. 2016, 20, 593–603. [Google Scholar] [CrossRef] [PubMed]
- Cacabelos, R. (Ed.) World Guide for Drug Use and Pharmacogenomics; EuroEspes Publishing: A Coruña, Spain, 2012. [Google Scholar]
- Kim, G.J.; Lee, S.Y.; Park, J.H.; Ryu, B.Y.; Kim, J.H. Role of Preemptive Genotyping in Preventing Serious Adverse Drug Events in South Korean Patients. Drug Saf. 2017, 40, 65–80. [Google Scholar] [CrossRef] [PubMed]
- Schildcrout, J.S.; Denny, J.C.; Bowton, E.; Gregg, W.; Pulley, J.M.; Basford, M.A.; Cowan, J.D.; Xu, H.; Ramirez, A.H.; Crawford, D.C.; et al. Optimizing drug outcomes through pharmacogenetics: A case for preemptive genotyping. Clin. Pharmacol. Ther. 2012, 92, 235–242. [Google Scholar] [CrossRef]
- Dunnenberger, H.M.; Crews, K.R.; Hoffman, J.M.; Caudle, K.E.; Broeckel, U.; Howard, S.C.; Hunkler, R.J.; Klein, T.E.; Evans, W.E.; Relling, M.V. Preemptive clinical pharmacogenetics implementation: Current programs in five US medical centers. Annu. Rev. Pharmacol. Toxicol. 2015, 55, 89–106. [Google Scholar] [CrossRef]
- Roden, D.M.; van Driest, S.L.; Mosley, J.D.; Wells, Q.S.; Robinson, J.R.; Denny, C.D.; Peterson, J.F. Benefit of Preemptive Pharmacogenetic Information on Clinical Outcome. Clin. Pharmacol. Ther. 2018, 103, 787–794. [Google Scholar] [CrossRef]
- Hong, J.Y.; Kim, J.H. PG-path: Modeling and personalizing pharmacogenomics-based pathways. PLoS ONE 2020, 15, e0230950. [Google Scholar] [CrossRef]
- Wilke, R.A.; Reif, D.M.; Moore, J.H. Combinatorial pharmacogenetics. Nat. Rev. Drug Discov. 2005, 4, 911–918. [Google Scholar] [CrossRef]
- Cacabelos, R.; Tellado, I.; Cacabelos, P. The epigenetic machinery in the life cycle and pharmacoepigenetics. In Pharmacoepigenetics; Cacabelos, R., Ed.; Academic Press/Elsevier: Oxford, UK, 2019; pp. 1–100. [Google Scholar]
- Cacabelos, R.; Carril, J.C.; Sanmartín, A.; Cacabelos, P. Pharmacoepigenetic processors: Epigenetic drugs, Drug resistance, Toxicoepigenetics, and Nutriepigenetics. In Pharmacoepigenetics; Cacabelos, R., Ed.; Academic Press/Elsevier: Oxford, UK, 2019; pp. 191–424. [Google Scholar]
- Song, Y.; Li, C.; Liu, G.; Liu, R.; Chen, Y.; Li, W.; Cao, Z.; Zhao, B.; Lu, C.; Liu, Y. Drug-Metabolizing Cytochrome P450 Enzymes Have Multifarious Influences on Treatment Outcomes. Clin. Pharm. 2021, 60, 585–601. [Google Scholar] [CrossRef]
- Meyer, U.A. Pharmacogenetics and adverse drug reactions. Lancet 2000, 356, 1667–1671. [Google Scholar] [CrossRef]
- Pratt, V.M.; Everts, R.E.; Aggarwal, P.; Beyer, B.N.; Broeckel, U.; Epstein-Baak, R.; Hujsak, P.; Kornreich, R.; Liao, J.; Lorier, R.; et al. Characterization of 137 Genomic DNA Reference Materials for 28 Pharmacogenetic Genes: A GeT-RM Collaborative Project. J. Mol. Diagn. 2016, 18, 109–123. [Google Scholar] [CrossRef]
- Mizzi, C.; Dalabira, E.; Kumuthini, J.; Dzimiri, J.; Balogh, I.; Basal, N.; Bohm, R.; Borg, J.; Borgiani, P.; Bozina, N.; et al. A European Spectrum of Pharmacogenomic Biomarkers: Implications for Clinical Pharmacogenomics. PLoS ONE 2016, 11, e0162866. [Google Scholar] [CrossRef]
- Daly, A.K. Using genome-wide association studies to identify genes important in serious adverse drug reactions. Annu. Rev. Pharmacol. Toxicol. 2012, 52, 21–35. [Google Scholar] [CrossRef]
- La Russa, R.; Fineschi, V.; Di Sanzo, M.; Gatto, V.; Santurro, A.; Martini, G.; Scopetti, M.; Frati, P. Personalized Medicine and Adverse Drug Reactions: The Experience of An Italian Teaching Hospital. Curr. Pharm. Biotechnol. 2017, 18, 274–281. [Google Scholar] [CrossRef]
- Liu, J.; Finkelstein, J. Towards Pharmacogenomics-Driven Medication Risk Assessment in People with Polypharmacy. Stud. Health Technol. Inform. 2018, 247, 880–884. [Google Scholar]
- Ohashi, W.; Tanaka, H. Benefits of pharmacogenomics in drug development-earlier launch of drugs and less adverse events. J. Med. Syst. 2010, 34, 701–707. [Google Scholar] [CrossRef]
- Kaguelidou, F.; Beau-Salinas, F.; Jonville-Bera, A.P.; Jacqz-Aigrain, E. Neonatal adverse drug reactions: An analysis of reports to the French pharmacovigilance database. Br. J. Clin. Pharmacol. 2016, 82, 1058–1068. [Google Scholar] [CrossRef]
- Dotta, A.; Chukhlantseva, N. Ontogeny and drug metabolism in newborns. J. Matern. Fetal. Neonatal. Med. 2012, 25, 83–84. [Google Scholar] [CrossRef]
- Rieder, M. Adverse Drug Reactions Across the Age Continuum: Epidemiology, Diagnostic Challenges, Prevention, and Treatments. J. Clin. Pharmacol. 2018, 58, S36–S47. [Google Scholar] [CrossRef]
- Ramsey, L.B.; Ong, H.H.; Schildcrout, J.S.; Shi, Y.; Tang, L.A.; Hicks, J.K.; elRouby, N.; Cavallari, L.H.; Tuteja, S.; Aquilante, C.L.; et al. Prescribing Prevalence of Medications with Potential Genotype-Guided Dosing in Pediatric Patients. JAMA Netw. Open 2020, 3, e2029411. [Google Scholar] [CrossRef]
- Brown, J.T.; Ramsey, L.B.; van Driest, S.L.; Aka, I.; Colace, S.I. Characterizing Pharmacogenetic Testing Among Children’s Hospitals. Clin. Transl. Sci. 2021, 14, 692–701. [Google Scholar] [CrossRef] [PubMed]
- Loo, T.T.; Ross, C.J.; Sistonen, J.; Visscher, H.; Madadi, P.; Koren, G.; Hayden, M.R.; Carleton, B.C. Pharmacogenomics and active surveillance for serious adverse drug reactions in children. Pharmacogenomics 2010, 11, 1269–1285. [Google Scholar] [CrossRef]
- Castro-Pastrana, L.I.; Ghannadan, R.; Rieder, M.J.; Dahlke, E.; Hayden, M.; Carleton, B. Cutaneous adverse drug reactions in children: An analysis of reports from the Canadian Pharmacogenomics Network for Drug Safety (CPNDS). J. Popul. Ther. Clin. Pharmacol. 2011, 18, e106–e120. [Google Scholar] [PubMed]
- Pussegoda, K.; Ross, C.J.; Visscher, H.; Yazdanpanah, M.; Brooks, B.; Rassekh, S.R.; Zada, Y.F.; Dube, M.P.; Carleton, B.C.; Hayden, M.R.; et al. Replication of TPMT and ABCC3 genetic variants highly associated with cisplatin-induced hearing loss in children. Clin. Pharmacol. Ther. 2013, 94, 243–251. [Google Scholar] [CrossRef] [PubMed]
- Rahawi, S.; Naik, H.; Blake, K.V.; Obeng, A.O.; Wasserman, R.M.; Seki, Y.; Funanage, V.L.; Oishi, K.; Scott, S.A. Knowledge and attitudes on pharmacogenetics among pediatricians. J. Hum. Genet. 2020, 65, 437–444. [Google Scholar] [CrossRef]
- Liko, I.; Lee, Y.M.; Stutzman, D.L.; Blackmer, A.B.; Deininger, K.M.; Reynolds, A.M.; Aquilante, C.L. Providers’ perspectives on the clinical utility of pharmacogenomic testing in pediatric patients. Pharmacogenomics 2021, 22, 263–274. [Google Scholar] [CrossRef]
- Gregornik, D.; Salyakina, D.; Brown, M.; Roiko, S.; Ramos, K. Pediatric pharmacogenomics: Challenges and opportunities: On behalf of the Sanford Children’s Genomic Medicine Consortium. Pharm. J. 2021, 21, 8–19. [Google Scholar] [CrossRef]
- Becker, M.L.; Leeder, J.S. Identifying genomic and developmental causes of adverse drug reactions in children. Pharmacogenomics 2010, 11, 1591–1602. [Google Scholar] [CrossRef]
- Cardelli, M.; Marchegiani, F.; Corsonello, A.; Lattanzio, F.; Provinciali, M. A review of pharmacogenetics of adverse drug reactions in elderly people. Drug Saf. 2012, 35, 3–20. [Google Scholar] [CrossRef]
- Cooper, J.A.; Cadogan, C.A.; Patterson, S.M.; Kerese, N.; Bradley, M.C.; Ryan, C.; Hughes, C.M. Interventions to improve the appropriate use of polypharmacy in older people: A Cochrane systematic review. BMJ Open 2015, 5, e009235. [Google Scholar] [CrossRef]
- Mangin, D.; Bahat, G.; Golomb, B.A.; Mallery, L.H.; Moorhouse, P.; Onder, G.; Petrovi, M.; Garfinkel, D. International Group for Reducing Inappropriate Medication Use & Polypharmacy (IGRIMUP): Position Statement and 10 Recommendations for Action. Drugs Aging 2018, 35, 575–587. [Google Scholar] [CrossRef]
- Finkelstein, J.; Friedman, C.; Hripcsak, G.; Cabrera, M. Pharmacogenetic polymorphism as an independent risk factor for frequent hospitalizations in older adults with polypharmacy: A pilot study. Pharmgenom. Pers. Med. 2016, 9, 107–116. [Google Scholar] [CrossRef][Green Version]
- Sheth, A.R.; Dave, R.B.; Rana, D.; Sheth, D. Comparison of the extent and prevalence of prescription of potentially inappropriate medications prescribed to geriatric age group residing in old-age homes versus those receiving care from tertiary care hospital using Beers criteria. Perspect. Clin. Res. 2020, 11, 144–149. [Google Scholar] [CrossRef]
- Mangoni, A.A. Predicting and detecting adverse drug reactions in old age: Challenges and opportunities. Expert Opin. Drug Metab. Toxicol. 2012, 8, 527–530. [Google Scholar] [CrossRef]
- Lavan, A.H.; Gallagher, P.F.; O’Mahony, D. Methods to reduce prescribing errors in elderly patients with multimorbidity. Clin. Interv. Aging 2016, 11, 857–866. [Google Scholar] [CrossRef]
- Crowley, E.K.; Sallevelt, B.T.G.M.; Huibers, C.J.A.; Murphy, K.D.; Spruit, M.; Shen, Z.; Boland, B.; Spinewine, A.; Dalleur, O.; Moutzouri, E.; et al. Intervention protocol: Optimising therapy to prevent avoidable hospital admission in the multi-morbid elderly (OPERAM): A structured medication review with support of a computerised decision support system. BMC Health Serv. Res. 2020, 20, 220. [Google Scholar] [CrossRef]
- Rankin, A.; Cadogan, C.A.; Patterson, S.M.; Kerse, N.; Cardwell, C.R.; Bradley, M.C.; Ryan, C.; Hughes, C. Interventions to improve the appropriate use of polypharmacy for older people. Cochrane Database Syst. Rev. 2018, 9, CD008165. [Google Scholar] [CrossRef]
- Whitman, A.; Erdeljac, P.; Jones, C.; Pillarella, N.; Nightingale, G. Managing Polypharmacy in Older Adults with Cancer Across Different Healthcare Settings. Drug Healthc. Patient Saf. 2021, 13, 101–116. [Google Scholar] [CrossRef] [PubMed]
- Dücker, C.M.; Brockmöller, J. Genomic Variation and Pharmacokinetics in Old Age: A Quantitative Review of Age- vs. Genotype-Related Differences. Clin. Pharmacol. Ther. 2019, 105, 625–640. [Google Scholar] [CrossRef]
- Beierle, I.; Meibohm, B.; Derendorf, H. Gender differences in pharmacokinetics and pharmacodynamics. Int. J. Clin. Pharmacol. Ther. 1999, 37, 529–547. [Google Scholar]
- Moyer, A.M.; Matey, E.T.; Miller, V.M. Individualized medicine: Sex, hormones, genetics, and adverse drug reactions. Pharmacol. Res. Perspect. 2019, 7, e00541. [Google Scholar] [CrossRef] [PubMed]
- Zucker, I.; Prendergast, B.J. Sex differences in pharmacokinetics predict adverse drug reactions in women. Biol. Sex Differ. 2020, 11, 32. [Google Scholar] [CrossRef]
- Bots, S.H.; Groepenhoff, F.; Eikendal, A.L.M.; Tannenbaum, C.; Rochon, P.A.; Regitz-Zagrosek, V.; Miller, V.M.; Day, D.; Asselbergs, F.W.; den Ruijter, H.M. Adverse Drug Reactions to Guideline-Recommended Heart Failure Drugs in Women: A Systematic Review of the Literature. JACC Heart Fail. 2019, 7, 258–266. [Google Scholar] [CrossRef]
- Brøsen, K. Sex differences in pharmacology. Ugeskr. Laeger 2007, 169, 2408–2411. [Google Scholar]
- Anthony, M.; Berg, M.J. Biologic and molecular mechanisms for sex differences in pharmacokinetics, pharmacodynamics, and pharmacogenetics: Part I. J. Womens Health Gend. Based Med. 2002, 11, 601–615. [Google Scholar] [CrossRef]
- Williams, L.A.; Spector, L.G. Survival Differences Between Males and Females Diagnosed with Childhood Cancer. JNCI Cancer Spectr. 2019, 3, pkz032. [Google Scholar] [CrossRef]
- Duan-Porter, W.; Goldstein, K.M.; McDuffie, J.R.; Hughes, J.M.; Clowse, M.E.B.; Klap, R.S.; Masilamani, V.; Allen, N.M.; Nagi, A.; Fierisch, J.M.; et al. Reporting of Sex Effects by Systematic Reviews on Interventions for Depression, Diabetes, and Chronic Pain. Ann. Intern. Med. 2016, 165, 184–193. [Google Scholar] [CrossRef]
- Momper, J.D.; Misel, M.L.; McKay, D.B. Sex differences in transplantation. Transplant. Rev. 2017, 31, 145–150. [Google Scholar] [CrossRef]
- Ekhart, C.; van Hunsel, F.; Scholl, J.; de Vries, S.; van Puijenbroek, E. Sex Differences in Reported Adverse Drug Reactions of Selective Serotonin Reuptake Inhibitors. Drug Saf. 2018, 41, 677–683. [Google Scholar] [CrossRef]
- Seeman, M.V. The Pharmacodynamics of Antipsychotic Drugs in Women and Men. Front. Psychiatry 2021, 12, 650904. [Google Scholar] [CrossRef]
- Hendriksen, L.C.; van der Linden, P.D.; Lagro-Janssen, A.L.M.; van den Bemt, P.M.L.A.; Siiskonen, S.J.; Teichert, M.; Kuiper, J.G.; Herings, R.M.C.; Stricker, B.H.; Visser, L.E. Sex differences associated with adverse drug reactions resulting in hospital admissions. Biol. Sex Differ. 2021, 12, 34. [Google Scholar] [CrossRef]
- Aboukaoud, M.; Israel, S.; Brautbar, C.; Eyal, S. Genetic Basis of Delayed Hypersensitivity Reactions to Drugs in Jewish and Arab Populations. Pharm. Res. 2018, 35, 211. [Google Scholar] [CrossRef]
- Collins, J.M.; Wang, D. Co-expression of drug metabolizing cytochrome P450 enzymes and estrogen receptor alpha (ESR1) in human liver: Racial differences and the regulatory role of ESR. Drug Metab. Pers. Ther. 2021, 36, 205–214. [Google Scholar] [CrossRef]
- Manikandan, P.; Nagini, S. Cytochrome P450 Structure, Function and Clinical Significance: A Review. Curr. Drug Targets 2018, 19, 38–54. [Google Scholar] [CrossRef]
- Ionova, Y.; Ashenhurst, J.; Zhan, J.; Nhan, H.; Kosinski, C.; Tamraz, B.; Chubb, A. CYP2C19 Allele Frequencies in Over 2.2 Million Direct-to-Consumer Genetics Research Participants and the Potential Implication for Prescriptions in a Large Health System. Clin. Transl. Sci. 2020, 13, 1298–1306. [Google Scholar] [CrossRef]
- Werk, A.N.; Cascorbi, I. Functional gene variants of CYP3A. Clin. Pharmacol. Ther. 2014, 96, 340–348. [Google Scholar] [CrossRef] [PubMed]
- Galaviz-Hernández, C.; Lazalde-Ramos, B.P.; Lares-Assef, I.; Macías-Salas, A.; Ortega-Chavez, A.; Rangel-Villalobos, H.; Sosa-Macías, M. Influence of Genetic Admixture Components on CYP3A5*3 Allele-Associated Hypertension in Amerindian Populations From Northwest Mexico. Front. Pharmacol. 2020, 11, 638. [Google Scholar] [CrossRef] [PubMed]
- Apellániz-Ruiz, M.; Inglada-Pérez, L.; Naranjo, M.E.; Sánchez, L.; Mancikova, V.; Currás-Freixes, M.; de Cubas, A.A.; Comino-Méndez, I.; Triki, S.; Rebai, A.; et al. High frequency and founder effect of the CYP3A4*20 loss-of-function allele in the Spanish population classifies CYP3A4 as a polymorphic enzyme. Pharm. J. 2015, 15, 288–292. [Google Scholar] [CrossRef]
- Hajjej, A.; Almawi, W.Y.; Arnaiz-Villena, A.; Hattab, L.; Hmida, S. The genetic heterogeneity of Arab populations as inferred from HLA genes. PLoS ONE 2018, 13, e0192269. [Google Scholar] [CrossRef]
- Yousef, A.M.; Bulatova, N.R.; Newman, W.; Hakooz, N.; Ismail, S.; Qusa, H.; Zahran, F.; Ababneh, N.A.; Hasan, F.; Zaloom, I.; et al. Allele and genotype frequencies of the polymorphic cytochrome P450 genes (CYP1A1, CYP3A4, CYP3A5, CYP2C9 and CYP2C19) in the Jordanian population. Mol. Biol. Rep. 2012, 39, 9423–9433. [Google Scholar] [CrossRef]
- Gaikovitch, E.A.; Cascorbi, I.; Mrozikiewicz, P.M.; Brockmoller, J.; Frotschl, R.; Kopke, K.; Gerloff, T.; Chernov, J.N.; Roots, I. Polymorphisms of drug-metabolizing enzymes CYP2C9, CYP2C19, CYP2D6, CYP1A1, NAT2 and of P-glycoprotein in a Russian population. Eur. J. Clin. Pharmacol. 2003, 59, 303–312. [Google Scholar] [CrossRef] [PubMed]
- Zand, N.; Tajik, N.; Moghaddam, A.S.; Milanian, I. Genetic polymorphisms of cytochrome P450 enzymes 2C9 and 2C19 in a healthy Iranian population. Clin. Exp. Pharmacol. Physiol. 2007, 34, 102–105. [Google Scholar] [CrossRef] [PubMed]
- Kuzmanovska, M.; Dimishkovska, M.; Maleva Kostovska, I.; Noveski, P.; Stefanovska, E.S.; Plaseka-Karanfilska, D. CYP2D6 allele distribution in Macedonians, Albanians and Romanies in the Republic of Macedonia. Balkan J. Med. Genet. 2016, 18, 49–58. [Google Scholar] [CrossRef]
- Marjani, A.; Gharanjik, A.M. Genetic Polymorphism of CYP2C9 Among Sistani Ethnic Group in Gorgan. Indian J. Clin. Biochem. 2018, 33, 208–213. [Google Scholar] [CrossRef]
- Hadjipanagi, D.; Chrysanthou, S.; Voskarides, K.; Deltas, C. Genetic polymorphisms in warfarin and tacrolimus-related genes VKORC1, CYP2C9 and CYP3A5 in the Greek-Cypriot population. BMC Res. Notes 2014, 7, 123. [Google Scholar] [CrossRef]
- Bozina, N.; Granić, P.; Lalić, Z.; Tramisak, I.; Lovric, M.; Stavljenic-Rukavina, A. Genetic polymorphisms of cytochromes P450: CYP2C9, CYP2C19, and CYP2D6 in Croatian population. Croat. Med. J. 2003, 44, 425–428. [Google Scholar]
- Scordo, M.G.; Caputi, A.P.; D’Arrigo, C.; Fava, G.; Spina, E. Allele and genotype frequencies of CYP2C9, CYP2C19 and CYP2D6 in an Italian population. Pharmacol. Res. 2004, 50, 195–200. [Google Scholar] [CrossRef]
- Petrović, J.; Pešić, V.; Lauschke, V.M. Frequencies of clinically important CYP2C19 and CYP2D6 alleles are graded across Europe. Eur. J. Hum. Genet. 2020, 28, 88–94. [Google Scholar] [CrossRef]
- Scott, S.A.; Edelmann, L.; Kornreich, R.; Erazo, M.; Desnick, R.J. CYP2C9, CYP2C19 and CYP2D6 allele frequencies in the Ashkenazi Jewish population. Pharmacogenomics 2007, 8, 721–730. [Google Scholar] [CrossRef] [PubMed]
- Adedeji, W.A.; Igbinoba, S.I.; Fakeye, T.O.; Odosu, I.A.; Fehintola, F.A.; Ma, Q.; Morse, G.D. Evaluation of CYP2D6 phenotype in the Yoruba Nigerian population. Expert Rev. Clin. Pharmacol. 2017, 10, 1145–1152. [Google Scholar] [CrossRef]
- Lee, S.S.; Kim, K.M.; Thi-Le, H.; Yea, S.S.; Cha, I.J.; Shin, J.G. Genetic polymorphism of CYP2C9 in a Vietnamese Kinh population. Ther. Drug Monit. 2005, 27, 208–210. [Google Scholar] [CrossRef]
- Nguyen, H.H.; Ma, T.T.H.; Vu, N.P.; Bach, Q.T.N.; Vu, T.H.; Nguyen, T.D.; Nong, H.V. Single nucleotide and structural variants of CYP2D6 gene in Kinh Vietnamese population. Medicine 2019, 98, e15891. [Google Scholar] [CrossRef]
- Ding, Y.; Yang, D.; Zhou, L.; He, P.; Yao, J.; Xie, P.; Lin, D.; Sun, D.; Sun, P.; Li, Q.; et al. Cytochrome P450 2C9 (CYP2C9) polymorphisms in Chinese Li population. Int. J. Clin. Exp. Med. 2015, 8, 21024–21033. [Google Scholar]
- He, Y.; Yang, H.; Geng, T.; Feng, T.; Yuan, D.; Kang, L.; Luo, M.; Jin, T. Genetic polymorphisms of pharmacogenomic VIP variants in the lhoba population of southwest China. Int. J. Clin. Exp. Pathol. 2015, 8, 13293–13303. [Google Scholar]
- Koopmans, A.B.; Braakman, M.H.; Vinkers, D.J.; Hoek, H.W.; van Harten, P. Meta-analysis of probability estimates of worldwide variation of CYP2D6 and CYP2C. Transl. Psychiatry 2021, 11, 141. [Google Scholar] [CrossRef]
- Martis, S.; Mei, H.; Vijzelaar, R.; Edelmann, L.; Desnick, R.J.; Scott, S.A. Multi-ethnic cytochrome-P450 copy number profiling: Novel pharmacogenetic alleles and mechanism of copy number variation formation. Pharm. J. 2013, 13, 558–566. [Google Scholar] [CrossRef]
- Naranjo, M.G.; Rodrigues-Soares, F.; Peñas-Lledó, E.M.; Tarazona-Santos, E.; FariñAS, J.; Rodeiro, I.; Terán, E.; Grazina, M.; Moya, G.E.; López-López, M.; et al. Interethnic Variability in CYP2D6, CYP2C9, and CYP2C19 Genes and Predicted Drug Metabolism Phenotypes Among 6060 Ibero- and Native Americans: RIBEF-CEIBA Consortium Report on Population Pharmacogenomics. Omics A J. Integr. Biol. 2018, 22, 575–588. [Google Scholar] [CrossRef]
- Jung, J.W.; Kim, J.Y.; Park, I.W.; Choi, B.W.; Kang, H.R. Genetic markers of severe cutaneous adverse reactions. Korean J. Intern. Med. 2018, 33, 867–875. [Google Scholar] [CrossRef]
- Chumnumwat, S.; Lu, Z.H.; Sukasem, C.; Winther, M.D.; Capule, F.R.; Hamid, A.A.A.T.A.; Bhandari, B.; Chaikledkaew, U.; Chanhom, N.; Chantarangsu, S.; et al. Southeast Asian Pharmacogenomics Research Network (SEAPharm): Current Status and Perspectives. Public Health Genom. 2019, 22, 132–139. [Google Scholar] [CrossRef]
- Sleder, A.T.; Kalus, J.; Lanfear, D.E. Cardiovascular Pharmacokinetics, Pharmacodynamics, and Pharmacogenomics for the Clinical Practitioner. J. Cardiovasc. Pharmacol. Ther. 2016, 21, 20–26. [Google Scholar] [CrossRef]
- Zhu, Y.; Swanson, K.M.; Rojas, R.L.; Wang, Z.; Sauver, J.L.S.; Visscher, S.L.; Prokop, L.J.; Bielinski, S.J.; Wang, L.; Weinshilboum, R.; et al. Systematic review of the evidence on the cost-effectiveness of pharmacogenomics-guided treatment for cardiovascular diseases. Genet. Med. 2020, 22, 475–486. [Google Scholar] [CrossRef]
- Božina, N.; Vrkić Kirhmajer, M.; Šimičević, L.; Ganoci, L.; Skvrce, N.M.; Domjanović, I.L.; Merćep, I. Use of pharmacogenomics in elderly patients treated for cardiovascular diseases. Croat. Med. J. 2020, 61, 147–158. [Google Scholar] [CrossRef]
- Roden, D.M.; van Driest, S.L.; Wells, Q.S.; Mosley, J.D.; Denny, J.C.; Peterson, J.F. Opportunities and Challenges in Cardiovascular Pharmacogenomics: From Discovery to Implementation. Circ. Res. 2018, 122, 1176–1190. [Google Scholar] [CrossRef]
- Magavern, E.F.; Kaski, J.C.; Turner, R.M.; Janmohamed, A.; Borry, P.; Pirmohamed, M. The Interface of Therapeutics and Genomics in Cardiovascular Medicine. Cardiovasc. Drugs Ther. 2021, 35, 663–676. [Google Scholar] [CrossRef]
- Dong, O.M.; Li, A.; Suzuki, O.; Oni-Orisan, A.; Gonzalez, R.; Stouffer, G.A.; Lee, C.R.; Wiltshire, T. Projected impact of a multigene pharmacogenetic test to optimize medication prescribing in cardiovascular patients. Pharmacogenomics 2018, 19, 771–782. [Google Scholar] [CrossRef]
- Singh, S.; El Rouby, N.; McDonough, C.W.; Gong, Y.; Bailey, K.R.; Boerwinkle, E.; Chapman, A.B.; Gums, J.G.; Turner, S.T.; Cooper-DeHoff, R.M.; et al. Genomic Association Analysis Reveals Variants Associated With Blood Pressure Response to Beta-Blockers in European Americans. Clin. Transl. Sci. 2019, 12, 497–504. [Google Scholar] [CrossRef]
- Xu, J.; Boström, A.E.; Saeed, M.; Dubey, R.K.; Waeber, G.; Vollenweider, P.; Marques-Vidal, P.; Mwinyi, J.; Schiöth, H.B. A genetic variant in the catechol-O-methyl transferase (COMT) gene is related to age-dependent differences in the therapeutic effect of calcium-channel blockers. Medicine 2017, 96, e7029. [Google Scholar] [CrossRef]
- Fontana, V.; Luizon, M.R.; Sandrim, V.C. An update on the pharmacogenetics of treating hypertension. J. Hum. Hypertens. 2015, 29, 283–291. [Google Scholar] [CrossRef]
- Rysz, J.; Franczyk, B.; Rysz-Górzyńska, M.; Gluba-Brózka, A. Pharmacogenomics of Hypertension Treatment. Int. J. Mol. Sci. 2020, 21, 4709. [Google Scholar] [CrossRef]
- Yang, L.; Lu, Y.L.; Wang, H.J.; Zhou, W.Z. Pharmacogenomics study of 620 whole-exome sequencing: Focusing on aspirin application. Zhonghua Er Ke Za Zhi 2016, 54, 332–336. [Google Scholar] [CrossRef]
- El-Sherif, N.; Turitto, G.; Boutjdir, M. Acquired Long QT Syndrome and Electrophysiology of Torsade de Pointes. Arrhythm. Electrophysiol. Rev. 2019, 8, 122–130. [Google Scholar] [CrossRef]
- Niemeijer, M.N.; van den Berg, M.E.; Eijgelsheim, M.; Rijnbeek, P.R.; Stricker, B.H. Pharmacogenetics of Drug-Induced QT Interval Prolongation: An Update. Drug Saf. 2015, 38, 855–867. [Google Scholar] [CrossRef]
- Duan, J.; Tao, J.; Zhai, M.; Li, C.; Zhou, N.; Lv, J.; Wang, L.; Lin, L.; Bai, R. Anticancer drugs-related QTc prolongation, torsade de pointes and sudden death: Current evidence and future research perspectives. Oncotarget 2018, 9, 25738–25749. [Google Scholar] [CrossRef]
- Hu, M.; Tomlinson, B. Pharmacogenomics of lipid-lowering therapies. Pharmacogenomics 2013, 14, 981–995. [Google Scholar] [CrossRef]
- Jiang, J.; Tang, Q.; Feng, J.; Dai, R.; Wang, Y.; Yang, Y.; Tang, X.; Deng, C.; Zeng, H.; Zhao, Y.; et al. Association between SLCO1B1 -521T>C and -388A>G polymorphisms and risk of statin-induced adverse drug reactions: A meta-analysis. Springerplus 2016, 5, 1368. [Google Scholar] [CrossRef] [PubMed]
- Tomlinson, B.; Chan, P.; Liu, Z.M. Statin Responses in Chinese Patients. J. Atheroscler. Thromb. 2018, 25, 199–202. [Google Scholar] [CrossRef]
- Kitzmiller, J.P.; Luzum, J.A.; Dauki, A.; Krauss, R.M.; Medina, M.W. Candidate-Gene Study of Functional Polymorphisms in SLCO1B1 and CYP3A4/5 and the Cholesterol-Lowering Response to Simvastatin. Clin. Transl. Sci. 2017, 10, 172–177. [Google Scholar] [CrossRef]
- Vassy, J.L.; Chun, S.; Advani, S.; Ludin, S.A.; Smith, J.G.; Alligood, E.C. Impact of SLCO1B1 Pharmacogenetic Testing on Patient and Healthcare Outcomes: A Systematic Review. Clin. Pharmacol. Ther. 2019, 106, 360–373. [Google Scholar] [CrossRef]
- House, J.S.; Motsinger-Reif, A.A. Fibrate pharmacogenomics: Expanding past the genome. Pharmacogenomics 2020, 21, 293–306. [Google Scholar] [CrossRef] [PubMed]
- Hertz, D.L.; Rae, J. Pharmacogenetics of cancer drugs. Annu. Rev. Med. 2015, 66, 65–81. [Google Scholar] [CrossRef] [PubMed]
- Cacabelos, R. Pleiotropy and promiscuity in pharmacogenomics for the treatment of Alzheimer’s disease and related risk factors. Future Neurol. 2018, 13, 71–86. [Google Scholar] [CrossRef]
- Miller, C.R.; McLeod, H.L. Pharmacogenomics of cancer chemotherapy-induced toxicity. J. Support. Oncol. 2007, 5, 9–14. [Google Scholar] [PubMed]
- Fatunde, O.A.; Brown, S.A. The Role of CYP450 Drug Metabolism in Precision Cardio-Oncology. Int. J. Mol. Sci. 2020, 21, 604. [Google Scholar] [CrossRef] [PubMed]
- Nichols, D.; Arnold, S.; Weiss, H.L.; Wu, J.; Durbin, E.B.; Miller, R.; Kolesar, J. Pharmacogenomic potential in advanced cancer patients. Am. J. Health Syst. Pharm. 2019, 76, 415–423. [Google Scholar] [CrossRef]
- Udagawa, C.; Zembutsu, H. Pharmacogenetics for severe adverse drug reactions induced by molecular-targeted therapy. Cancer Sci. 2020, 111, 3445–3457. [Google Scholar] [CrossRef]
- Faruque, F.; Noh, H.; Hussain, A.; Neuberger, E.; Onukwugha, E. Economic Value of Pharmacogenetic Testing for Cancer Drugs with Clinically Relevant Drug-Gene Associations: A Systematic Literature Review. J. Manag. Care Spec. Pharm. 2019, 25, 260–271. [Google Scholar] [CrossRef]
- Fleeman, N.; Martin Saborido, C.; Payne, K.; Boland, A.; Dickson, R.; Dundar, Y.; Fernández Santander, A.; Howell, S.; Newman, W.; Oyee, J.; et al. The clinical effectiveness and cost-effectiveness of genotyping for CYP2D6 for the management of women with breast cancer treated with tamoxifen: A systematic review. Health Technol. Assess. 2011, 15, 1–102. [Google Scholar] [CrossRef]
- Wang, T.; Zhou, Y.; Cao, G. Pharmacogenetics of tamoxifen therapy in Asian populations: From genetic polymorphism to clinical outcomes. Eur. J. Clin. Pharmacol. 2021, 77, 1095–1111. [Google Scholar] [CrossRef]
- Scott, E.N.; Hasbullah, J.S.; Carleton, B.C.; Ross, C.J.D. Prevention of adverse drug effects: A pharmacogenomic approach. Curr. Opin. Pediatr. 2020, 32, 646–653. [Google Scholar] [CrossRef]
- Paulík, A.; Nekvindová, J.; Filip, S. Irinotecan toxicity during treatment of metastatic colorectal cancer: Focus on pharmacogenomics and personalized medicine. Tumori 2020, 106, 87–94. [Google Scholar] [CrossRef]
- Martinez-Balibrea, E.; Abad, A.; Martínez-Cardús, A.; Valladares, M.; Navarro, M.; Aranda, E.; Marcuello, E.; Benavides, M.; Massutí, B.; Carrato, A. UGT1A and TYMS genetic variants predict toxicity and response of colorectal cancer patients treated with first-line irinotecan and fluorouracil combination therapy. Br. J. Cancer 2010, 103, 581–589. [Google Scholar] [CrossRef]
- Atasilp, C.; Chansriwong, P.; Sirachainan, E.; Reungwetwattana, T.; Sirilerttakul, S.; Chamnanphon, M.; Puangpetch, A.; Sukasem, C. Effect of drug metabolizing enzymes and transporters in Thai colorectal cancer patients treated with irinotecan-based chemotherapy. Sci. Rep. 2020, 10, 13486. [Google Scholar] [CrossRef]
- Jada, S.R.; Lim, R.; Wong, C.I.; Shu, X.; Lee, S.C.; Zhou, Q.; Goh, B.C.; Chowbay, B. Role of UGT1A1*6, UGT1A1*28 and ABCG2 c.421C>A polymorphisms in irinotecan-induced neutropenia in Asian cancer patients. Cancer Sci. 2007, 98, 1461–1467. [Google Scholar] [CrossRef]
- Zheng, Y.; Deng, Z.; Tang, M.; Xiao, D.; Cai, P. Impact of genetic factors on platinum-induced gastrointestinal toxicity. Mutat. Res. 2020, 786, 108324. [Google Scholar] [CrossRef]
- Li, Y.Q.; Zhang, X.Y.; Chen, J.; Yin, J.Y.; Li, X.P. ATP7B rs9535826 is associated with gastrointestinal toxicity of platinum-based chemotherapy in nonsmall cell lung cancer patients. J. Cancer Res. Ther. 2018, 14, 881–886. [Google Scholar] [CrossRef]
- Kato, M.; Tsurudome, Y.; Kanemitsu, T.; Yasukochi, S.; Kanado, Y.; Ogino, T.; Matsunaga, N.; Koyanagi, S.; Ohdo, S. Diurnal expression of MRP4 in bone marrow cells underlies the dosing-time dependent changes in the oxaliplatin-induced myelotoxicity. Sci. Rep. 2020, 10, 13484. [Google Scholar] [CrossRef]
- Wright, G.E.B.; Amstutz, U.; Drögemöller, B.I.; Shih, J.; Rassekh, S.R.; Hayden, M.R.; Carleton, B.C.; Ross, C.J.D. Pharmacogenomics of Vincristine-Induced Peripheral Neuropathy Implicates Pharmacokinetic and Inherited Neuropathy Genes. Clin. Pharmacol. Ther. 2019, 105, 402–410. [Google Scholar] [CrossRef]
- Zhou, H.; Li, L.; Yang, P.; Zheng, J.E.; Zhou, Y.; Han, Y. Optimal predictor for 6-mercaptopurine intolerance in Chinese children with acute lymphoblastic leukemia: NUDT15, TPMT, or ITPA genetic variants? BMC Cancer 2018, 18, 516. [Google Scholar] [CrossRef] [PubMed]
- Moradveisi, B.; Muwakkit, S.; Zamani, F.; Ghaderi, E.; Mohammadi, E.; Zgheib, N.K. ITPA, TPMT, and NUDT15 Genetic Polymorphisms Predict 6-Mercaptopurine Toxicity in Middle Eastern Children With Acute Lymphoblastic Leukemia. Front. Pharmacol. 2019, 10, 916. [Google Scholar] [CrossRef]
- Kotur, N.; Lazic, J.; Ristivojevic, B.; Stankovic, B.; Gasic, V.; Dokmanovic, L.; Krstovski, N.; Milosevic, G.; Janic, D.; Zukic, B.; et al. Pharmacogenomic Markers of Methotrexate Response in the Consolidation Phase of Pediatric Acute Lymphoblastic Leukemia Treatment. Genes 2020, 11, 468. [Google Scholar] [CrossRef]
- Sini, V.; Botticelli, A.; Lunardi, G.; Gori, S.; Marchetti, P. Pharmacogenetics and aromatase inhibitor induced side effects in breast cancer patients. Pharmacogenomics 2017, 18, 821–830. [Google Scholar] [CrossRef]
- Artigalás, O.; Vanni, T.; Hutz, M.H.; Ashton-Prolla, P.; Schwatz, I.V. Influence of CYP19A1 polymorphisms on the treatment of breast cancer with aromatase inhibitors: A systematic review and meta-analysis. BMC Med. 2015, 13, 139. [Google Scholar] [CrossRef]
- Bojanic, K.; Kuna, L.; Bilic Curcic, I.; Wagner, J.; Smolic, R.; Kralik, K.; Kizivat, T.; Ivanac, G.; Vcev, A.; Wu, G.Y.; et al. Representation of CYP3A4, CYP3A5 and UGT1A4 Polymorphisms within Croatian Breast Cancer Patients’ Population. Int. J. Environ. Res. Public Health 2020, 17, 3692. [Google Scholar] [CrossRef]
- Gal, J.; Milano, G.; Brest, P.; Ebran, N.; Gilhodes, J.; Llorca, L.; Dubot, C.; Romieu, G.; Desmoulins, I.; Brain, E.; et al. VEGF-Related Germinal Polymorphisms May Identify a Subgroup of Breast Cancer Patients with Favorable Outcome under Bevacizumab-Based Therapy-A Message from COMET, a French Unicancer Multicentric Study. Pharmaceuticals 2020, 13, 414. [Google Scholar] [CrossRef]
- Guo, M.; Li, S.; Zhao, X.; Yuan, Y.; Zhang, B.; Guan, Y. Knockdown of Circular RNA Hsa_circ_0000714 Can Regulate RAB17 by Sponging miR-370-3p to Reduce Paclitaxel Resistance of Ovarian Cancer Through CDK6/RB Pathway. OncoTargets Ther. 2020, 13, 13211–13224. [Google Scholar] [CrossRef]
- Zur, R.M.; Roy, L.M.; Ito, S.; Beyene, J.; Carew, C.; Ungar, W.J. Thiopurine S-methyltransferase testing for averting drug toxicity: A meta-analysis of diagnostic test accuracy. Pharm. J. 2016, 16, 305–311. [Google Scholar] [CrossRef]
- Root, A.; Johnson, R.; McGee, A.; Lee, H.J.; Yang, S.; Voora, D. Understanding the state of pharmacogenomic testing for thiopurine methyltransferase within a large health system. Pharmacogenomics 2020, 21, 411–418. [Google Scholar] [CrossRef] [PubMed]
- Zimdahl Kahlin, A.; Helander, S.; Skoglund, K.; Soderkvist, P.; Martensson, L.G.; Appell, M.L. Comprehensive study of thiopurine methyltransferase genotype, phenotype, and genotype-phenotype discrepancies in Sweden. Biochem. Pharmacol. 2019, 164, 263–272. [Google Scholar] [CrossRef]
- Relling, M.V.; Schwab, M.; Whirl-Carrillo, M.; Suarez-Kurtz, G.; Pui, C.H.; Stein, C.M.; Moyer, A.M.; Evans, W.E.; Klein, T.E.; Antilon-Klussmann, F.G.; et al. Clinical Pharmacogenetics Implementation Consortium Guideline for Thiopurine Dosing Based on TPMT and NUDT15 Genotypes: 2018 Update. Clin. Pharmacol. Ther. 2019, 105, 1095–1105. [Google Scholar] [CrossRef]
- Choi, R.; Sohn, I.; Kim, M.J.; Woo, H.I.; Lee, J.W.; Ma, Y.; Yi, E.S.; Koo, H.H.; Lee, S.Y. Pathway genes and metabolites in thiopurine therapy in Korean children with acute lymphoblastic leukaemia. Br. J. Clin. Pharmacol. 2019, 85, 1585–1597. [Google Scholar] [CrossRef]
- D’Alessio, A.; Prete, M.G.; Cammarota, A.; Personeni, N.; Rimassa, L. The Role of Cabozantinib as a Therapeutic Option for Hepatocellular Carcinoma: Current Landscape and Future Challenges. J. Hepatocell. Carcinoma 2021, 8, 177–191. [Google Scholar] [CrossRef]
- van Erp, N.P.; Eechoute, K.; van der Veldt, A.A.; Haanen, J.B.; Reyners, A.K.L.; Mathijssen, R.H.J.; Boven, E.; van der Straaten, T.; Baak-Pablo, R.F.; Wessels, J.A.M.; et al. Pharmacogenetic pathway analysis for determination of sunitinib-induced toxicity. J. Clin. Oncol. 2009, 27, 4406–4412. [Google Scholar] [CrossRef]
- Chu, Y.H.; Li, H.; Tan, H.S.; Koh, V.; Lai, J.; Phyo, W.M.; Choudhury, Y.; Kanesvaran, R.; Chau, N.M.; Toh, C.K.; et al. Association of ABCB1 and FLT3 Polymorphisms with Toxicities and Survival in Asian Patients Receiving Sunitinib for Renal Cell Carcinoma. PLoS ONE 2015, 10, e0134102. [Google Scholar] [CrossRef] [PubMed]
- Sun, F.; Chen, Z.; Yao, P.; Weng, B.; Liu, Z.; Cheng, L. Meta-Analysis of ABCG2 and ABCB1 Polymorphisms With Sunitinib-Induced Toxicity and Efficacy in Renal Cell Carcinoma. Front. Pharmacol. 2021, 12, 641075. [Google Scholar] [CrossRef]
- Zhao, Q.; Zhang, T.; Xiao, X.R.; Huang, J.F.; Wang, Y.; Gonzalez, F.J.; Li, F. Impaired clearance of sunitinib leads to metabolic disorders and hepatotoxicity. Br. J. Pharmacol. 2019, 176, 2162–2178. [Google Scholar] [CrossRef]
- Amaya, G.M.; Durandis, R.; Bourgeois, D.S.; Perkins, J.A.; Abouda, A.A.; Wines, K.J.; Mohamud, M.; Starks, S.A.; Daniels, R.N.; Jackson, K.D. Cytochromes P450 1A2 and 3A4 Catalyze the Metabolic Activation of Sunitinib. Chem. Res. Toxicol. 2018, 31, 570–584. [Google Scholar] [CrossRef]
- Chen, H.; Liu, J.; Zhou, Y.; Hou, Y.; Ma, G.; Cai, W. Association of Hepatic Nuclear Factor 4 Alpha Gene Polymorphisms With Free Imatinib Plasma Levels and Adverse Reactions in Chinese Gastrointestinal Stromal Tumor Patients. Ther. Drug Monit. 2019, 41, 582–590. [Google Scholar] [CrossRef]
- Loganayagam, A.; Arenas Hernandez, M.; Corrigan, A.; Fairbanks, L.; Lewis, C.M.; Harper, P.; Maisey, N.; Ross, P.; Sanderson, J.D.; Marinaki, A.M. Pharmacogenetic variants in the DPYD, TYMS, CDA and MTHFR genes are clinically significant predictors of fluoropyrimidine toxicity. Br. J. Cancer 2013, 108, 2505–2515. [Google Scholar] [CrossRef] [PubMed]
- O’Donnell, P.H.; Trubetskoy, V.; Nurhussein-Patterson, A.; Hall, J.P.; Nath, A.; Huo, D.; Fleming, G.F.; Ingle, J.N.; Abramson, V.G.; Morrow, P.K.; et al. Clinical evaluation of germline polymorphisms associated with capecitabine toxicity in breast cancer: TBCRC. Breast Cancer Res. Treat. 2020, 181, 623–633. [Google Scholar] [CrossRef]
- Aagaard, L.; Hansen, E.H. Adverse drug reactions reported by consumers for nervous system medications in Europe 2007 to. BMC Pharmacol. Toxicol. 2013, 14, 30. [Google Scholar] [CrossRef]
- Cacabelos, R. Epigenetics and pharmacoepigenetics of neurodevelopmental and neuropsychiatric disorders. In Pharmacoepigenetics; Cacabelos, R., Ed.; Academic Press/Elsevier: Oxford, UK, 2019; pp. 609–709. [Google Scholar]
- Kam, H.; Jeong, H. Pharmacogenomic Biomarkers and Their Applications in Psychiatry. Genes 2020, 11, 1445. [Google Scholar] [CrossRef]
- Cacabelos, R.; Torrellas, C.; Teijido, O.; Carril, J.C. Pharmacogenetic considerations in the treatment of Alzheimer’s disease. Pharmacogenomics 2016, 17, 1041–1074. [Google Scholar] [CrossRef]
- Cacabelos, R.; Torrellas, C. Pharmacogenomics of antidepressants. HSOA J. Psych. Dep. Anx. 2015, 1, 001. [Google Scholar] [CrossRef]
- Cacabelos, R. Pharmacogenetic considerations when prescribing cholinesterase inhibitors for the treatment of Alzheimer’s disease. Expert Opin. Drug Metab. Toxicol. 2020, 16, 673–701. [Google Scholar] [CrossRef]
- Cacabelos, R. Pharmacogenomics of Alzheimer’s and Parkinson’s diseases. Neurosci. Lett. 2020, 726, 133807. [Google Scholar] [CrossRef]
- Cacabelos, R. Population-level pharmacogenomics for precision drug development in dementia. Expert Rev. Prec. Med. Drug Dev. 2018, 3, 163–188. [Google Scholar] [CrossRef]
- Cacabelos, R.; Cacabelos, P.; Torrellas, C.; Tellado, I.; Carril, J.C. Pharmacogenomics of Alzheimer’s disease: Novel therapeutic strategies for drug development. Methods Mol. Biol. 2014, 1175, 323–556. [Google Scholar] [CrossRef]
- van Westrhenen, R.; Aitchison, K.J.; Ingelman-Sundberg, M.; Jukic, M.M. Pharmacogenomics of Antidepressant and Antipsychotic Treatment: How Far Have We Got and Where Are We Going? Front. Psychiatry 2020, 11, 94. [Google Scholar] [CrossRef]
- Torrellas, C.; Carril, J.C.; Cacabelos, R. Optimization of antidepressant use with pharmacogenetic strategies. Curr. Genom. 2017, 18, 442–449. [Google Scholar] [CrossRef]
- Marshe, V.S.; Islam, F.; Maciukiewicz, M.; Bousman, C.; Eyre, H.A.; Lavretsky, H.; Mulsant, B.H.; Reynolds, C.F., 3rd; Lenze, E.J.; Muller, D.J. Pharmacogenetic Implications for Antidepressant Pharmacotherapy in Late-Life Depression: A Systematic Review of the Literature for Response, Pharmacokinetics and Adverse Drug Reactions. Am. J. Geriatr. Psychiatry 2020, 28, 609–629. [Google Scholar] [CrossRef]
- Nassan, M.; Nicholson, W.T.; Elliott, M.A.; Rohrer Vitek, C.R.; Black, J.L.; Frye, M.A. Pharmacokinetic Pharmacogenetic Prescribing Guidelines for Antidepressants: A Template for Psychiatric Precision Medicine. Mayo Clin. Proc. 2016, 91, 897–907. [Google Scholar] [CrossRef]
- Ahmed, A.T.; Weinshilboum, R.; Frye, M.A. Benefits of and Barriers to Pharmacogenomics-Guided Treatment for Major Depressive Disorder. Clin. Pharmacol. Ther. 2018, 103, 767–769. [Google Scholar] [CrossRef]
- García-González, J.; Tansey, K.E.; Hauser, J.; Henigsberg, N.; Maier, W.; Mors, O.; Placentino, A.; Rietschel, M.; Souery, D.; Zagar, Y.; et al. Pharmacogenetics of antidepressant response: A polygenic approach. Prog. Neuropsychopharmacol. Biol. Psychiatry 2017, 75, 128–134. [Google Scholar] [CrossRef]
- Shumake, J.; Mallard, T.T.; McGeary, J.E.; Beevers, C.G. Inclusion of genetic variants in an ensemble of gradient boosting decision trees does not improve the prediction of citalopram treatment response. Sci. Rep. 2021, 11, 3780. [Google Scholar] [CrossRef]
- Fabbri, C.; Serretti, A. How to Utilize Clinical and Genetic Information for Personalized Treatment of Major Depressive Disorder: Step by Step Strategic Approach. Clin. Psychopharmacol. Neurosci. 2020, 18, 484–492. [Google Scholar] [CrossRef] [PubMed]
- Fabbri, C.; Tansey, K.E.; Perlis, R.H.; Hauser, J.; Henigsberg, N.; Maier, W.; Mors, O.; Placentino, A.; Rietschel, M.; Souery, D.; et al. Effect of cytochrome CYP2C19 metabolizing activity on antidepressant response and side effects: Meta-analysis of data from genome-wide association studies. Eur. Neuropsychopharmacol. 2018, 28, 945–954. [Google Scholar] [CrossRef] [PubMed]
- Ward, J.; Graham, N.; Strawbridge, R.J.; Ferguson, A.; Jenkins, G.; Chen, W.; Hodgson, K.; Frye, M.; Weinshilboum, R.; Uher, R.; et al. Polygenic risk scores for major depressive disorder and neuroticism as predictors of antidepressant response: Meta-analysis of three treatment cohorts. PLoS ONE 2018, 13, e0203896. [Google Scholar] [CrossRef]
- Li, Q.S.; Tian, C.; Hinds, D.; 23andMe Research Team. Genome-wide association studies of antidepressant class response and treatment-resistant depression. Transl. Psychiatry 2020, 10, 360. [Google Scholar] [CrossRef]
- Le-Niculescu, H.; Roseberry, K.; Gill, S.S.; Levey, D.F.; Phalen, P.L.; Mullen, J.; Williams, A.; Bhairo, S.; Voegtline, T.; Davis, H. Precision medicine for mood disorders: Objective assessment, risk prediction, pharmacogenomics, and repurposed drugs. Mol. Psychiatry 2021, 26, 2776–2804. [Google Scholar] [CrossRef] [PubMed]
- Amare, A.T.; Schubert, K.O.; Baune, B.T. Pharmacogenomics in the treatment of mood disorders: Strategies and Opportunities for personalized psychiatry. EPMA J. 2017, 8, 211–227. [Google Scholar] [CrossRef]
- Bråten, L.S.; Haslemo, T.; Jukic, M.M.; Ingalmen-Sundberg, M.; Molden, E.; Kringen, M.K. Impact of CYP2C19 genotype on sertraline exposure in 1200 Scandinavian patients. Neuropsychopharmacology 2020, 45, 570–576. [Google Scholar] [CrossRef]
- Chang, M.; Tybring, G.; Dahl, M.L.; Lindh, J.D. Impact of cytochrome P450 2C19 polymorphisms on citalopram/escitalopram exposure: A systematic review and meta-analysis. Clin. Pharmacokinet. 2014, 53, 801–811. [Google Scholar] [CrossRef] [PubMed]
- Rossow, K.M.; Aka, I.T.; Maxwell-Horn, A.C.; Roden, D.M.; van Driest, S.L. Pharmacogenetics to Predict Adverse Events Associated With Antidepressants. Pediatrics 2020, 146, e20200957. [Google Scholar] [CrossRef] [PubMed]
- Shishko, I.; Almeida, K.; Silvia, R.J.; Tataronis, G.R. Psychiatric pharmacists’ perception on the use of pharmacogenomic testing in the mental health population. Pharmacogenomics 2015, 16, 949–958. [Google Scholar] [CrossRef]
- Reynolds, K.K.; McNally, B.A.; Linder, M.W. Clinical Utility and Economic Impact of CYP2D6 Genotyping. Clin. Lab. Med. 2016, 36, 525–542. [Google Scholar] [CrossRef]
- Bousman, C.A.; Bengesser, S.A.; Aitchison, K.J.; Amare, A.T.; Aschauer, H.; Baune, B.; Behroozi Asl, B.; Bishop, J.R.; Burmeister, M.; Chaumette, B.; et al. Review and Consensus on Pharmacogenomic Testing in Psychiatry. Pharmacopsychiatry 2021, 54, 5–17. [Google Scholar] [CrossRef] [PubMed]
- Garon, S.L.; Pavlos, R.K.; White, K.D.; Brown, N.J.; Stone, C.A., Jr.; Phillips, E.J. Pharmacogenomics of off-target adverse drug reactions. Br. J. Clin. Pharmacol. 2017, 83, 1896–1911. [Google Scholar] [CrossRef]
- Redwood, A.J.; Pavlos, R.K.; White, K.D.; Phillips, E.J. HLAs: Key regulators of T-cell-mediated drug hypersensitivity. HLA 2018, 91, 3–16. [Google Scholar] [CrossRef]
- Karnes, J.H.; Miller, M.A.; White, K.D.; Kovinse, K.C.; Pavlos, R.K.; Redwood, A.J.; Peter, J.G.; Lehloenya, R.; Mallal, S.A.; Phillips, E.J. Applications of Immunopharmacogenomics: Predicting, Preventing, and Understanding Immune-Mediated Adverse Drug Reactions. Annu. Rev. Pharmacol. Toxicol. 2019, 59, 463–486. [Google Scholar] [CrossRef]
- Zewde, M.; Kiyotani, K.; Park, J.H.; Fang, H.; Yap, K.L.; Yew, P.Y.; Alachkar, H.; Kato, T.; Mai, T.H.; Ikeda, Y.; et al. The era of immunogenomics/immunopharmacogenomics. J. Hum. Genet. 2018, 63, 865–875. [Google Scholar] [CrossRef]
- Chang, C.J.; Chen, C.B.; Hung, S.I.; Ji, C.; Chung, W.H. Pharmacogenetic Testing for Prevention of Severe Cutaneous Adverse Drug Reactions. Front. Pharmacol. 2020, 11, 969. [Google Scholar] [CrossRef] [PubMed]
- Daneshjou, R.; Huddart, R.; Klein, T.E.; Altman, R.B. Pharmacogenomics in dermatology: Tools for understanding gene-drug associations. Semin. Cutan. Med. Surg. 2019, 38, E19–E24. [Google Scholar] [CrossRef]
- Hasegawa, A.; Abe, R. Recent advances in managing and understanding Stevens-Johnson syndrome and toxic epidermal necrolysis. F1000Research 2020, 9, F1000 Faculty Rev-612. [Google Scholar] [CrossRef] [PubMed]
- Guéant, J.L.; Guéant-Rodriguez, R.M.; Gastin, I.A.; Cornejo-García, J.A.; Viola, M.; Barbaud, A.; Mertes, P.M.; Blanca, M.; Romano, A. Pharmacogenetic determinants of immediate and delayed reactions of drug hypersensitivity. Curr. Pharm. Des. 2008, 14, 2770–2777. [Google Scholar] [CrossRef] [PubMed]
- Tan-Koi, W.C.; Lim, E.S.; Teo, Y.Y. Health regulatory communications of well-established safety-related pharmacogenomics associations in six developed countries: An evaluation of alignment. Pharm. J. 2017, 17, 121–127. [Google Scholar] [CrossRef]
- Jarjour, S.; Barrette, M.; Normand, V.; Rouleau, J.L.; Dubé, M.P.; de Denus, S. Genetic markers associated with cutaneous adverse drug reactions to allopurinol: A systematic review. Pharmacogenomics 2015, 16, 755–767. [Google Scholar] [CrossRef] [PubMed]
- Wu, R.; Cheng, Y.J.; Zhu, L.L.; Yu, L.; Zhao, W.K.; Jia, M.; Wen, C.H.; Long, X.Z.; Tang, T.; He, A.J.; et al. Impact of HLA-B*58:01 allele and allopurinol-induced cutaneous adverse drug reactions: Evidence from 21 pharmacogenetic studies. Oncotarget 2016, 7, 81870–81879. [Google Scholar] [CrossRef]
- Sukasem, C.; Katsila, T.; Tempark, T.; Patrinos, G.P.; Chantratita, W. Drug-Induced Stevens-Johnson Syndrome and Toxic Epidermal Necrolysis Call for Optimum Patient Stratification and Theranostics via Pharmacogenomics. Annu. Rev. Genom. Hum. Genet. 2018, 19, 329–353. [Google Scholar] [CrossRef]
- Sun, B.; Cheng, L.; Xiong, Y.; Hu, L.; Luo, Z.; Zhou, M.; Li, J.; Xie, H.; He, F.; Yuan, X.; et al. PSORS1C1 Hypomethylation Is Associated with Allopurinol-Induced Severe Cutaneous Adverse Reactions during Disease Onset Period: A Multicenter Retrospective Case-Control Clinical Study in Han Chinese. Front. Pharmacol. 2018, 8, 923. [Google Scholar] [CrossRef]
- Manson, L.E.N.; Swen, J.J.; Guchelaar, H.J. Diagnostic Test Criteria for HLA Genotyping to Prevent Drug Hypersensitivity Reactions: A Systematic Review of Actionable HLA Recommendations in CPIC and DPWG Guidelines. Front. Pharmacol. 2020, 11, 567048. [Google Scholar] [CrossRef]
- Hsu, Y.O.; Lu, K.L.; Fu, Y.; Wang, C.W.; Lu, C.W.; Lin, Y.F.; Chang, W.C.; Yeh, K.Y.; Jung, S.I.; Chung, W.H.; et al. The Roles of Immunoregulatory Networks in Severe Drug Hypersensitivity. Front. Immunol. 2021, 12, 597761. [Google Scholar] [CrossRef]
- Fonseca, D.J.; Morel, A.; Llinás-Caballero, K.; Bolívar-Salazar, D.; Laissue, P. Whole-Exome Sequencing in Patients Affected by Stevens-Johnson Syndrome and Toxic Epidermal Necrolysis Reveals New Variants Potentially Contributing to the Phenotype. Pharmgenom. Pers. Med. 2021, 14, 287–299. [Google Scholar] [CrossRef] [PubMed]
- COVID-19 Host Genetics Initiative. Mapping the human genetic architecture of COVID. Nature 2021. [Google Scholar] [CrossRef]
- Stevenson, J.M.; Alexander, G.C.; Palamuttam, N.; Mehta, H.B. Projected Utility of Pharmacogenomic Testing Among Individuals Hospitalized With COVID-19: A Retrospective Multicenter Study in the United States. Clin. Transl. Sci. 2021, 14, 153–162. [Google Scholar] [CrossRef] [PubMed]
- Bickler, S.W.; Cauvi, D.M.; Fisch, K.M.; Prieto, J.M.; Sykes, A.G.; Thangarajah, H.; Lazar, D.A.; Ignacio, R.C.; Gerstmann, D.R.; Ryan, A.F.; et al. Extremes of age are associated with differences in the expression of selected pattern recognition receptor genes and ACE2, the receptor for SARS-CoV-2: Implications for the epidemiology of COVID-19 disease. BMC Med. Genom. 2021, 14, 138. [Google Scholar] [CrossRef]
- Benetti, E.; Tita, R.; Spiga, O.; Ciolfi, A.; Giovanni, B.; Bruselles, A.; Doddato, G.; Giliberti, A.; Marconi, C.; Musacchia, F.; et al. ACE2 gene variants may underlie interindividual variability and susceptibility to COVID-19 in the Italian population. Eur. J. Hum. Genet. 2020, 28, 1602–1614. [Google Scholar] [CrossRef] [PubMed]
- Khayat, A.S.; de Assumpção, P.P.; Meireles Khayat, B.C.; Thomaz Araújo, T.M.; Almeida Batista-Gomes, J.; Carvalho Imbiriba, L.; Ishak, G.; Baraúna de Assumpcao, P.; Cordeiro Moreira, F.; Rodriguez Burbano, R.; et al. ACE2 polymorphisms as potential players in COVID-19 outcome. PLoS ONE 2020, 15, e0243887. [Google Scholar] [CrossRef]
- Cheng, Z.; Zhou, J.; To, K.K.; Chu, H.; Li, C.; Wang, D.; Yang, D.; Zheng, S.; Hao, K.; Bossé, Y.; et al. Identification of TMPRSS2 as a Susceptibility Gene for Severe 2009 Pandemic A(H1N1) Influenza and A(H7N9) Influenza. J. Infect. Dis. 2015, 212, 1214–1221. [Google Scholar] [CrossRef]
- Clinckemalie, L.; Spans, L.; Dubois, V.; Laurent, M.; Helsen, C.; Joniau, S.; Claessens, F. Androgen regulation of the TMPRSS2 gene and the effect of a SNP in an androgen response element. Mol. Endocrinol. 2013, 27, 2028–2040. [Google Scholar] [CrossRef]
- Gemmati, D.; Bramanti, B.; Serino, M.L.; Secchiero, P.; Zauli, G.; Tisato, V. COVID-19 and Individual Genetic Susceptibility/Receptivity: Role of ACE1/ACE2 Genes, Immunity, Inflammation and Coagulation. Might the Double X-chromosome in Females Be Protective against SARS-CoV-2 Compared to the Single X-Chromosome in Males? Int. J. Mol. Sci. 2020, 21, 3474. [Google Scholar] [CrossRef]
- Baratchian, M.; McManus, J.M.; Berk, M.P.; Nakamura, F.; Mukhopadhyay, S.; Xu, W.; Erzurum, S.; Drazba, J.; Peterson, J.; Klein, E.A.; et al. Androgen regulation of pulmonary AR, TMPRSS2 and ACE2 with implications for sex-discordant COVID-19 outcomes. Sci. Rep. 2021, 11, 11130. [Google Scholar] [CrossRef] [PubMed]
- Rossi, A.D.; de Araújo, J.L.F.; de Almeida, T.B.; Ribeiro-Alves, M.; de Almeida Velozo, C.; Maciel de Almeida, J.; de Carvalho Leitao, I.; Natiene Ferreria, S.; da Silva Oliveira, J.; Alves, H.J.; et al. Association between ACE2 and TMPRSS2 nasopharyngeal expression and COVID-19 respiratory distress. Sci. Rep. 2021, 11, 9658. [Google Scholar] [CrossRef]
- Gómez, J.; Albaiceta, G.M.; García-Clemente, M.; Lopez-Larrea, C.; Amado-Rodríguez, L.; Lopez-Alonso, I.; Hermida, T.; Enriquez, A.A.; Herrero, P.; Melón, S.; et al. Angiotensin-converting enzymes (ACE, ACE2) gene variants and COVID-19 outcome. Gene 2020, 762, 145102. [Google Scholar] [CrossRef]
- Lecarpentier, Y.; Vallée, A. The key role of the level of ACE2 gene expression in SARS-CoV-2 infection. Aging 2021, 13, 14552–14556. [Google Scholar] [CrossRef]
- Lambertsen, K.L.; Biber, K.; Finsen, B. Inflammatory cytokines in experimental and human stroke. J. Cereb. Blood Flow. Metab. 2012, 32, 1677–1698. [Google Scholar] [CrossRef]
- Jessurun, N.T.; Drent, M.; van Puijenbroek, E.P.; Bekers, O.; Wijnen, P.A.; Bast, A. Drug-induced interstitial lung disease: Role of pharmacogenetics in predicting cytotoxic mechanisms and risks of side effects. Curr. Opin. Pulm. Med. 2019, 25, 468–477. [Google Scholar] [CrossRef]
- Pellegrino, P.; Falvella, F.S.; Perrone, V.; Carnovale, C.; Brusadelli, T.; Pozzi, M.; Antoniazzi, S.; Cheli, S.; Perrotta, C.; Clementi, E.; et al. The first steps towards the era of personalised vaccinology: Predicting adverse reactions. Pharm. J. 2015, 15, 284–287. [Google Scholar] [CrossRef]
- Collins, S.L.; Carr, D.F.; Pirmohamed, M. Advances in the Pharmacogenomics of Adverse Drug Reactions. Drug Saf. 2016, 39, 15–27. [Google Scholar] [CrossRef]
- Haga, S.B.; Burke, W.; Ginsburg, G.S.; Agans, R. Primary care physicians’ knowledge of and experience with pharmacogenetic testing. Clin. Genet. 2012, 82, 388–394. [Google Scholar] [CrossRef] [PubMed]
- Haga, S.B.; O’Daniel, J.M.; Tindall, G.M.; Mills, R.; Lipkus, I.M.; Agans, R. Survey of genetic counselors and clinical geneticists’ use and attitudes toward pharmacogenetic testing. Clin. Genet. 2012, 82, 115–120. [Google Scholar] [CrossRef]
- Lee, Y.M.; McKillip, R.P.; Borden, B.A.; Klammer, C.E.; Ratain, M.J.; O’Donnell, O.H. Assessment of patient perceptions of genomic testing to inform pharmacogenomic implementation. Pharm. Genom. 2017, 27, 179–189. [Google Scholar] [CrossRef]
- Haddy, C.A.; Ward, H.M.; Angley, M.T.; McKinnon, R.A. Consumers’ views of pharmacogenetics: A qualitative study. Res. Soc. Adm. Pharm. 2010, 6, 221–231. [Google Scholar] [CrossRef]
- Daud, A.N.A.; Bergsma, E.L.; Bergman, J.E.H.; de Walle, H.E.K.; Kerstjens-Frederikse, W.S.; Bijker, B.J.; Hak, E.; Wilffert, B. Knowledge and attitude regarding pharmacogenetics among formerly pregnant women in the Netherlands and their interest in pharmacogenetic research. BMC Pregnancy Childbirth 2017, 17, 120. [Google Scholar] [CrossRef]
- McKillip, R.P.; Borden, B.A.; Galecki, P.; Patricl-Miller, L.; Hall, J.P.; Hussain, S.; Danahey, K.; Suegler, M.; Sorrentino, M.J.; Sacro, Y.; et al. Patient Perceptions of Care as Influenced by a Large Institutional Pharmacogenomic Implementation Program. Clin. Pharmacol. Ther. 2017, 102, 106–114. [Google Scholar] [CrossRef] [PubMed]
- Kichko, K.; Marschal, P.; Flessa, S. Personalized Medicine in the U.S. and Germany: Awareness, Acceptance, Use and Preconditions for the Wide Implementation into the Medical Standard. J. Pers. Med. 2016, 6, 15. [Google Scholar] [CrossRef]
- Kobayashi, E.; Sakurada, T.; Ueda, S.; Satoh, N. Public involvement in pharmacogenomics research: A national survey on patients’ attitudes towards pharmacogenomics research and the willingness to donate DNA samples to a DNA bank in Japan. Cell Tissue Bank. 2011, 12, 71–80. [Google Scholar] [CrossRef]
- Jayasinghe, K.; Quinlan, C.; Mallett, A.J.; Kerr, P.G.; McClaren, B.; Nisselle, A.; Mallawaarachchi, A.; Polkinghorne, K.R.; Patel, C.; Best, S.; et al. Attitudes and Practices of Australian Nephrologists Toward Implementation of Clinical Genomics. Kidney Int. Rep. 2020, 6, 272–283. [Google Scholar] [CrossRef]
- Peterson, J.F.; Field, J.R.; Shi, Y.; Schildcrout, J.S.; Deeny, J.C.; McGregor, T.L.; van Driest, S.L.; Pulley, J.M.; Lubin, I.M.; Laposata, M. Attitudes of clinicians following large-scale pharmacogenomics implementation. Pharm. J. 2016, 16, 393–398. [Google Scholar] [CrossRef]
- Plumpton, C.O.; Roberts, D.; Pirmohamed, M.; Hughes, D.A. A Systematic Review of Economic Evaluations of Pharmacogenetic Testing for Prevention of Adverse Drug Reactions. Pharmacoeconomics 2016, 34, 771–793. [Google Scholar] [CrossRef] [PubMed]
- Walden, L.M.; Brandl, E.J.; Tiwari, A.K.; Cheena, S.; Freeman, N.; Braganza, N.; Kennedy, J.L.; Muller, D.J. Genetic testing for CYP2D6 and CYP2C19 suggests improved outcome for antidepressant and antipsychotic medication. Psychiatry Res. 2019, 279, 111–115. [Google Scholar] [CrossRef] [PubMed]
- Kamenski, G.; Ayazseven, S.; Berndt, A.; Fink, W.; Kamenski, L.; Zehetmayer, S.; Puhringer, H. Clinical Relevance of CYP2D6 Polymorphisms in Patients of an Austrian Medical Practice: A Family Practice-Based Observational Study. Drugs Real. World Outcomes 2020, 7, 63–73. [Google Scholar] [CrossRef] [PubMed]
- Wake, D.T.; Ilbawi, N.; Dunnenberger, H.M.; Hulick, P.J. Pharmacogenomics: Prescribing Precisely. Med. Clin. N. Am. 2019, 103, 977–990. [Google Scholar] [CrossRef]
- Lauschke, V.M.; Ingelman-Sundberg, M. Prediction of drug response and adverse drug reactions: From twin studies to Next Generation Sequencing. Eur. J. Pharm. Sci. 2019, 130, 65–77. [Google Scholar] [CrossRef]
- Young, J.; Bhattacharya, K.; Ramachandran, S.; Lee, A.; Bentley, J.P. Rates of genetic testing in patients prescribed drugs with pharmacogenomic information in FDA-approved labeling. Pharm. J. 2021, 21, 318–325. [Google Scholar] [CrossRef] [PubMed]
- Koutsilieri, S.; Tzioufa, F.; Sismanoglou, D.C.; Patrinos, G.P. Unveiling the guidance heterogeneity for genome-informed drug treatment interventions among regulatory bodies and research consortia. Pharmacol. Res. 2020, 153, 104590. [Google Scholar] [CrossRef]
- Imatoh, T.; Sai, K.; Saito, Y. Pharmacogenomic information in the Warning section of drug labels: A comparison between labels in the United States and those in five other countries/regions. J. Clin. Pharm. Ther. 2018, 43, 493–499. [Google Scholar] [CrossRef]
- Kim, J.A.; Ceccarelli, R.; Lu, C.Y. Pharmacogenomic Biomarkers in US FDA-Approved Drug Labels (2000-2020). J. Pers. Med. 2021, 11, 179. [Google Scholar] [CrossRef]
- Joly, Y.; Koutrikas, G.; Tassé, A.M.; Issa, A.; Carleton, B.; Hayden, M.; Rieder, M.J.; Ramos-Paque, E.; Avard, D. Regulatory approval for new pharmacogenomic tests: A comparative overview. Food Drug Law J. 2011, 66, 1–24. [Google Scholar] [PubMed]
- Tan-Koi, W.C.; Sung, C.; Chong, Y.Y.; Lateef, A.; Pang, S.M.; Vasudevan, A.; Aw, D.; Lui, N.L.; Lee, S.X.; Ren, E.C.; et al. Tailoring of recommendations to reduce serious cutaneous adverse drug reactions: A pharmacogenomics approach. Pharmacogenomics 2017, 18, 881–890. [Google Scholar] [CrossRef]
- Zineh, I.; Pacanowski, M.A. Pharmacogenomics in the assessment of therapeutic risks versus benefits: Inside the United States Food and Drug Administration. Pharmacotherapy 2011, 31, 729–735. [Google Scholar] [CrossRef]
- Lee, Y.M.; Danahey, K.; Knoebel, R.W.; Ratain, M.J.; Maeltzer, D.O.; O’Donnell, P.H. Analysis of comprehensive pharmacogenomic profiling to impact in-hospital prescribing. Pharm. Genom. 2019, 29, 23–30. [Google Scholar] [CrossRef] [PubMed]
- Elliott, L.S.; Henderson, J.C.; Neradilek, M.B.; Moyer, N.A.; Ashcraft, K.C.; Thirumaran, R.K. Clinical impact of pharmacogenetic profiling with a clinical decision support tool in polypharmacy home health patients: A prospective pilot randomized controlled trial. PLoS ONE 2017, 12, e0170905. [Google Scholar] [CrossRef]
- Moyer, A.M.; Caraballo, P.J. The challenges of implementing pharmacogenomic testing in the clinic. Expert Rev. Pharm. Outcomes Res. 2017, 17, 567–577. [Google Scholar] [CrossRef]
- Amara, N.; Blouin-Bougie, J.; Bouthillier, D.; Simard, J. On the readiness of physicians for pharmacogenomics testing: An empirical assessment. Pharm. J. 2018, 18, 308–318. [Google Scholar] [CrossRef]
- O’Donnell, P.H.; Bush, A.; Spitz, J.; Danahey, K.; Saner, D.; Das, S.; Cox, N.J.; Ratain, M.J. The 1200 patients project: Creating a new medical model system for clinical implementation of pharmacogenomics. Clin. Pharmacol. Ther. 2012, 92, 446–449. [Google Scholar] [CrossRef]
- Ramsey, L.B.; Brown, J.T.; Vear, S.I.; Bishop, J.R.; van Driest, S.L. Gene-Based Dose Optimization in Children. Annu. Rev. Pharmacol. Toxicol. 2020, 60, 311–331. [Google Scholar] [CrossRef]
- Caudle, K.E.; Keeling, N.J.; Klein, T.E.; Whirl-Carrillo, M.; Pratt, V.M.; Hoffman, J.M. Standardization can accelerate the adoption of pharmacogenomics: Current status and the path forward. Pharmacogenomics 2018, 19, 847–860. [Google Scholar] [CrossRef]
- Zaid, N.; Limami, Y.; Senhaji, N.; Errafiy, N.; Khalki, L.; Bakri, Y.; Zaid, Y.; Amzazi, S. Coverage rate of ADME genes from commercial sequencing arrays. Medicine 2019, 98, e13975. [Google Scholar] [CrossRef]
- Reisberg, S.; Krebs, K.; Lepamets, M.; Kals, M.; Magi, R.; Metsalu, K.; Lauschke, V.M.; Vilo, J.; Milani, L. Translating genotype data of 44,000 biobank participants into clinical pharmacogenetic recommendations: Challenges and solutions. Genet. Med. 2019, 21, 1345–1354. [Google Scholar] [CrossRef]
- Drozda, K.; Pacanowski, M.A.; Grimstein, C.; Zineh, I. Pharmacogenetic Labeling of FDA-Approved Drugs: A Regulatory Retrospective. JACC Basic. Transl. Sci. 2018, 3, 545–549. [Google Scholar] [CrossRef]
- Skvrce, N.M.; Krivokapić, S.; Božina, N. Implementation of pharmacogenomics in product information. Pharmacogenomics 2020, 21, 443–448. [Google Scholar] [CrossRef]
- Manolopoulos, V.G. Pharmacogenomics and adverse drug reactions in diagnostic and clinical practice. Clin. Chem. Lab. Med. 2007, 45, 801–814. [Google Scholar] [CrossRef]
- Giri, J.; Moyer, A.M.; Bielinski, S.J.; Caraballo, P.J. Concepts Driving Pharmacogenomics Implementation Into Everyday Healthcare. Pharmgenom. Pers. Med. 2019, 12, 305–318. [Google Scholar] [CrossRef] [PubMed]
| Agents Acting on the Renin–Angiotensin System | ||
|---|---|---|
| ACE Inhibitors | ||
| Drug | Properties | Pharmacogenetics |
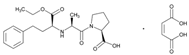 | Name: ENALAPRIL IUPAC Name: L-proline, 1-[N-[1-(ethoxycarbonyl)-3-phenylpropyl]-L-alanyl]-, (S)-, (Z)-2-butenedioate (1:1); 1-[N-[(S)-1-carboxy-3-phenylpropyl]-L-alanyl]-L-proline 1′-ethyl ester, maleate (1:1) Molecular Formula: C20H28N2O5 C4H4O4 Molecular Weight: 492.52 Mechanism: Competitive inhibitor of angiotensin-converting enzyme (ACE). Prevents conversion of angiotensin I to angiotensin II, a potent vasoconstrictor. Results in lower levels of angiotensin II, which causes an increase in plasma renin activity and a reduction in aldosterone secretion. Effect: Treatment of hypertension, symptomatic heart failure, or asymptomatic left ventricular dysfunction. | Mechanistic genes:ACE1, ADRB2, AGTR1, AGT, BDKRB2, NOS3 Metabolic genes Substrate:CYP3A4, CYP3A5 Inhibitor:ACE1 Transporter genes:SLC22A8 Pleiotropic genes:IL6 |
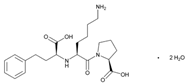 | Name: LISINOPRIL IUPAC Name: L-proline, 1-[N2-(1-carboxy-3-phenylpropyl)-L-lysyl]-, dihydrate, (S)-; 1-[N2-[(S)-1-carboxy-3-phenylpropyl]-L-lysyl]-L-proline dihydrate Molecular Formula: C21H31N3O5 2H2O Molecular Weight: 441.52 Mechanism: Competitive inhibitor of angiotensin-converting enzyme (ACE). Prevents conversion of angiotensin I to angiotensin II, a potent vasoconstrictor. Effect: Treatment of hypertension, either alone or in combination with other antihypertensive agents. Adjunctive therapy in treatment of heart failure. Treatment of acute MI within 24 h in hemodynamically stable patients to improve survival. | Mechanistic genes:ACE1, ADD1, AGTR1, AGT Metabolic genes Substrate:ACE1, ADD1, AGTR1, AGT Inhibitor:ACE1, ACE2 |
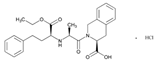 | Name: QUINAPRIL IUPAC Name: 3-Isoquinolinecarboxylic acid, 2-[2-[[1-(ethoxycarbonyl)-3-phenylpropyl]amino]-1-oxopropyl]-1,2,3,4-tetrahydro-, monohydrochloride, [3S-[2[R*(R*)],3R*]]; (S)-2-[(S)-N-[(S)-1-carboxy-3-phenylpropyl]alanyl]-1,2,3,4-tetrahydro-3-isoquinolinecarboxylic acid, 1-ethyl ester, monohydrochloride Molecular Formula: C25H30N2O5 HCl Molecular Weight: 474.98 Mechanism: Treatment of hypertension. Reduction in cardiovascular mortality or non-fatal myocardial infarction in stable coronary artery disease. Effect: Hypertension. Heart failure. | Mechanistic genes: ACE1, AGT, AGTR1, BDKBR2, NR1I21 TGFB1 Metabolic genes Substrate: CYP11B2 Inhibitor: ACE1 |
 | Name: RAMIPRIL IUPAC Name: [1,1′-Biphenyl]-2-carboxylic acid, 4′-[(1,4′-dimethyl-2′-propyl[2,6′-bi-1H-benzimidazol]-1′-yl)methyl]-; 4′-[[4-methyl-6-(1-methyl-2-benzimidazolyl)-2-propyl-1-benzimidazolyl]methyl]-2-biphenylcarboxylic acid Molecular Formula: C23H32N2O5 Molecular Weight: 514.62 Mechanism: A non-peptide AT1 angiotensin II receptor antagonist. This binding prevents angiotensin II from binding to receptor thereby blocking vasoconstriction and aldosterone-secreting effects of angiotensin II. Effect: Treatment of hypertension, alone or in combination with other antihypertensive agents. | Mechanistic genes: ACE1, AGT, AGTR1, BDKRB2, ERAP1, PPARG Metabolic genes Substrate: CYP2C9, CYP11B2, UGT1A1 Inhibitor: ABCB1, ABCG2, CYP2C9, CYP2C19 |
| Angiotensin II Antagonists | ||
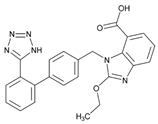 | Name: CANDESARTAN IUPAC Name: 1H-benzimidazole-7-carboxylic acid, 2-ethoxy-1-[[2′-(1H-tetrazol-5-yl)[1,1′-biphenyl]-4-yl]methyl]-; 2-ethoxy-1-[p-(o-1H-tetrazol-5-ylphenyl)benzyl]-7-benzimidazolecarboxylic acid Molecular Formula: C24H20N6O3 Molecular Weight: 440.45 Mechanism: Candesartan is an angiotensin receptor antagonist, blocking vasoconstriction and the aldosterone-secreting effects (reabsorption of sodium and water) of angiotensin II. Effect: Essential hypertension. Heart failure. | Metabolic genes Substrate: CYP1A1, CYP2C9, CYP11B2, UGT1A3, UGT1A5, UGT2B7 Inhibitor: ABCG2, CYP2C8, CYP2C9 Transporter genes:ABCB1, ABCG2 |
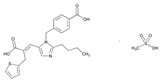 | Name: EPROSARTAN IUPAC Name: 2-Thiophenepropanoic acid, α-[[2-butyl-1-[(4-carboxyphenyl)methyl]-1H-imidazol-5-yl]methylene]-, €-, monomethanesulfonat€(E)-2-butyl-1-(p-carboxybenzyl)-α-2-thenylimidazole-5-acrylic acid, monomethanesulfonate Molecular Formula: C23H24N2O4S CH4O3S Molecular Weight: 520.62 Mechanism: A non-biphenyl, non-tetrazole angiotensin II receptor (AT1) antagonist. Blocks the vasoconstrictor and aldosterone-secreting effects of angiotensin II by selectively blocking the binding of angiotensin II to the AT1 receptor in many tissues, such as vascular smooth muscle and adrenal gland. Does not bind to or block other hormone receptors or ion channels known to be important in cardiovascular regulation. Effect: Used alone or in combination with other classes of antihypertensive agents in the management of hypertension. | Mechanistic genes: ACE1, AGTR1 Metabolic genes Inhibitor: CYP2C9 Inducer: ABCC2 Transporter genes: ABCB1, ABCG2 |
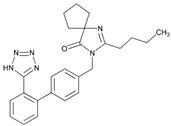 | Name: IRBESARTAN IUPAC Name: 1,3-Diazaspiro[4.4]non-1-en-4-one, 2-butyl-3-[[2′-(1H-tetrazol-5-yl)[1,1′-biphenyl]-4-yl]methyl]-; 2-butyl-3-[p-(o-1H-tetrazol-5-ylphenyl)benzyl]-1,3-diazaspiro[4.4]non-1-en-4-one Molecular Formula: C25H28N6O Molecular Weight: 428.53 Mechanism: Irbesartan binds to AT1 angiotensin II receptor. This binding prevents angiotensin II from binding to receptor, thereby blocking the vasoconstriction and aldosterone-secreting effects of angiotensin II. Effect: Treatment of hypertension alone or in combination with other antihypertensives. Treatment of diabetic nephropathy in type 2 diabetes mellitus (non-insulin-dependent, NIDDM) and hypertension. | Mechanistic genes: ADRA1A, AGTR1, APOB, BDKRB2, ERAP1, EDN1, NPPA, AGT, APOE, LDLR, NOS3, TGFB1 Metabolic genes Substrate: CYP2C9, CYP3A4, CYP3A5, CYP11B2 Inhibitor: CYP1A2, CYP2C8, CYP2C9, CYP2D6, CYP3A4, CYP3A5 Transporter genes: ABCB1, ABCG2 |
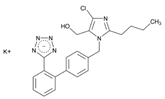 | Name: LOSARTAN IUPAC Name: 1H-Imidazole-5-methanol, 2-butyl-4-chloro-1-[[2′-(1H-tetrazol-5-yl)[1,1′-biphenyl]-4-yl]methyl]-, monopotassium salt; 2-butyl-4-chloro-1-[p-(o-1H-tetrazol-5-ylphenyl)benzyl]imidazole-5-methanol, monopotassium salt Molecular Formula: C22H22ClKN6O Molecular Weight: 461.00 Mechanism: As a selective and competitive non-peptide angiotensin II receptor antagonist, losartan blocks vasoconstrictor and aldosterone-secreting effects of angiotensin II. Losartan increases urinary flow rate and in addition to being natriuretic and kaliuretic, increases excretion of chloride, magnesium, uric acid, calcium, and phosphate. Effect: Treatment of hypertension. Treatment of diabetic nephropathy in type 2 diabetes mellitus (non-insulin-dependent) and history of hypertension. Stroke risk reduction in hypertension and left ventricular hypertrophy. | Mechanistic genes:ACE1, ADD1, AGT, AGTR1, AGTR2, BDKRB2, EDN1, FOS, MMP2, NOS3, PDGFRB, REN, TGFB1 Metabolic genes Substrate:CYP1A2, CYP2C8, CYP2C9, CYP2C19, CYP2D6, CYP3A4, CYP3A5, UGT1A1, UGT1A3, UGT1A10, UGT2B7, UGT2B17 Inhibitor:CYP1A2, CYP2C8, CYP2C9, CYP2C19, CYP3A4, CYP3A5, CYP11B2 Transporter genes:ABCB1, ABCG2 Pleiotropic genes:TNF |
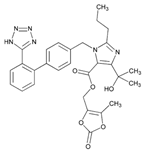 | Name: OLMESARTAN IUPAC Name: 1H-Imidazole-5-carboxylic acid, 4-(1-hydroxy-1-methylethyl)-2-propyl-1-[[2′-(1H-tetrazol-5-yl) [1,1′-biphenyl]-4-yl]methyl]-, (5-methyl-2-oxo-1,3-dioxol-4-yl) methyl ester Molecular Formula: C29H30N6O6 Molecular Weight: 558.59 Mechanism: Blocks vasoconstrictor and aldosterone-secreting effects of angiotensin II. Interacts reversibly at AT1 and AT2 receptors and has slow dissociation kinetics (has greater affinity for AT1 receptor). Olmesartan increases urinary flow rate and, besides being natriuretic and kaliuretic, increases excretion of chloride, magnesium, uric acid, calcium, and phosphate. Effect: Hypertension. | Mechanistic genes:ACE2, AGTR1, EDN1, TGFB1 Metabolic genes Substrate:CMBL, CYP2C9 Inducer:ABCC2 Transporter genes:ABCB1, ABCC2, ABCG2, SLC22A8, SLCO1A2, SLCO1B1 Pleiotropic genes:APOE |
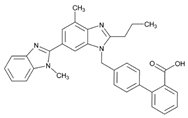 | Name: TELMISARTAN IUPAC Name: [1,1′-Biphenyl]-2-carboxylic acid, 4′-[(1,4′-dimethyl-2′-propyl[2,6′-bi-1H-benzimidazol]-1′-yl)methyl]-; 4′-[[4-methyl-6-(1-methyl-2-benzimidazolyl)-2-propyl-1-benzimidazolyl]methyl]-2-biphenylcarboxylic acid Molecular Formula: C33H30N4O2 Molecular Weight: 514.62 Mechanism: A non-peptide AT1 angiotensin II receptor antagonist. This binding prevents angiotensin II from binding to receptor thereby blocking vasoconstriction and aldosterone-secreting effects of angiotensin II. Effect: Treatment of hypertension, alone or in combination with other antihypertensive agents. | Mechanistic genes:ACE1, AGT, AGTR1, BDKRB2, ERAP1, PPARG Metabolic genes Substrate:CYP2C9, CYP11B2, UGT1A1 Inhibitor:ABCB1, ABCG2, CYP2C9, CYP2C19 |
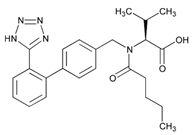 | Name: VALSARTAN IUPAC Name: L-valine, N-(1-oxopentyl)-N-[[2′-(1H-tetrazol-5-yl)[1,1′-biphenyl]-4-yl]methyl]-; N-[p-(o-1H-tetrazol-5-ylphenyl)benzyl]-N-valeryl-L-valine Molecular Formula: C24H29N5O3 Molecular Weight: 435.52 Mechanism: Displaces angiotensin II from AT1 receptor and produces its blood pressure-lowering effects by antagonizing AT1-induced vasoconstriction, aldosterone release, catecholamine release, arginine vasopressin release, water intake, and hypertrophic responses. Effect: Treatment of essential hypertension (alone or in combination with other antihypertensive agents). Reduction in cardiovascular mortality in left ventricular dysfunction postmyocardial infarction. Treatment of heart failure. | Mechanistic genes:ACE1, AGT, AGTR1, AGT2R1, BDKRB2, ERAP1, GNB3, STAT3, TGFB1 Metabolic genes Substrate:CYP2C9, CYP2C19, CYP2D6, CYP3A4, CYP3A5, CYP11B2 Inhibitor:CYP2C9 Transporter genes:ABCC2, SLCO1B1, SLCO1B3 |
| Other Agents acting on the Renin–Angiotensin system | ||
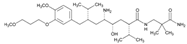 | Name: ALISKIREN IUPAC Name: Benzeneoctanamide, δ-amino-N-(3-amino-2,2-dimethyl-3-oxopropyl)-γ-hydroxy-4-methoxy-3-(3-methoxypropoxy)-α,ζ-bis(1-methylethyl)-, (αS, γS, δS, ζS)-; (2) (2S,4S,5S,7S)-5-amino-N-(2-carbamoyl-2-methylpropyl)-4-hydroxy-2-isopropyl-7-[4-methoxy-3-(3-methoxypropoxy)benzyl]-8-methylnonamide Molecular Formula: C30H53N3O6 Molecular Weight: 551.76 Mechanism: Blocks conversion of angiotensinogen to angiotensin I. Effect: Treatment of hypertension. | Mechanistic genes:REN Metabolic genes Substrate:CYP3A4, CYP3A5 Inhibitor:CYP3A4, CYP3A5, REN Transporter genes:ABCB1 Mechanistic genes:ACE1, ACE2, ADD1, ADRB2, AGT, AGTR1, MTHFR, MTR |
| Antihypertensives | ||
| Drug | Properties | Pharmacogenetics |
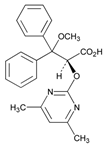 | Name: AMBRISENTAN IUPAC Name: (+)-(2S)-2-[(4,6-dimethylpyrimidin-2-yl)oxy]-3-methoxy-3,3-diphenylpropanoic acid Molecular Formula: C22H22N2O4 Molecular Weight: 378.42 Mechanism: Blocks endothelin receptor ETA and ETB on vascular endothelium and smooth muscle. Effect: Treatment of pulmonary artery hypertension. | Mechanistic genes:EDN1, EDNRA, NOS3 Metabolic genes Substrate:CYP2C9, CYP2C19, CYP3A4, CYP3A5, GSTs, UGT1A3, UGT1A9, UGT2B7 Transporter genes:ABCB1, SLCO1A2 Pleiotropic genes:IL1B, IL6 |
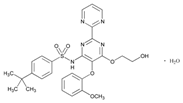 | Name: BOSENTAN IUPAC Name: Benzenesulfonamide, 4-(1,1-dimethylethyl)-N-[6-(2-hydroxyethoxy)-5-(2-methoxyphenoxy)[2,2′-bipyrimidin]-4-yl]-, monohydrate; (2) p-tert-butyl-N-[6-(2-hydroxyethoxy)-5-(o-methoxyphenoxy)-2-(2-pyrimidinyl)-4-pyrimidinyl]benzenesulfonamide monohydrate Molecular Formula: C27H29N5O6S.H2O Molecular Weight: 569.63 Mechanism: Acts as a competitive antagonist and blocks endothelin receptors on vascular endothelium and smooth muscle. Stimulation of endothelin receptors is associated with vasoconstriction and proliferation. Although bosentan blocks both ETA and ETB receptors, the affinity is slightly higher for ETA. Effect: Adjunctive therapy for the treatment of pulmonary arterial hypertension (WHO group I), in patients with WHO class III or IV symptoms. | Metabolic genes Substrate:ACVRL1, BMPR2, EDNRA, EDNRB, TGFBR1 Inhibitor:CYP2B6, CYP2C9, CYP3A4, CYP3A5 Inducer:CYP2C9, CYP2C19, CYP3A4, CYP3A5 Transporter genes:ABCB1, ABCB11, SLCO1B1, SLCO1B3 Pleiotropic genes:TNF |
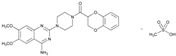 | Name: DOXAZOSIN IUPAC Name: Piperazine, 1-(4-amino-6,7-dimethoxy-2-quinazolinyl)-4-[(2,3-dihydro-1,4-benzodioxin-2-yl)carbonyl]-, monomethanesulfonate; 1-(4-amino-6,7-dimethoxy-2-quinazolinyl)-4-(1,4-benzodioxan-2-ylcarbonyl)piperazine monomethanesulfonate Molecular Formula: C23H25N5O5.CH4O3S Molecular Weight: 547.58 Mechanism: Doxazosin is a quinazoline-derivative postsynaptic α1-adrenergic blocking agent. It reduces peripheral vascular resistance and blood pressure as a result of its vasodilating effects. The drug produces both arterial and venous dilation. Effects of doxazosin on the cardiovascular system are mediated by the drug’s activity at α1-receptor sites on vascular smooth muscle. Because of the prevalence of α receptors on the prostate capsule, prostate adenoma, and the bladder trigone and the relative absence of these receptors on the bladder body, α-blockers decrease urinary outflow resistance in men. Doxazosin may improve to a limited extent the serum lipid profile and can reduce blood glucose and serum insulin concentrations. The drug does not appear to affect plasma renin activity appreciably. Effect: Treatment of hypertension alone or in conjunction with diuretics, ACE inhibitors, β-blockers, or calcium antagonists. Treatment of urinary outflow obstruction and/or obstructive and irritative symptoms associated with BPH; can be used in combination with finasteride. | Mechanistic genes:ACE1, ADD1, ADRA1A Metabolic genes Substrate:CYP2C19, CYP2D6, CYP3A4, CYP3A5 Transporter genes:ABCB1 |
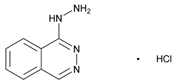 | Name: HYDRALAZINE IUPAC Name: Phthalazine, 1-hydrazino-, monohydrochloride; 1-hydrazinophthalazine monohydrochloride Molecular Formula: C8H8N4 HCl Molecular Weight: 196.64 Mechanism: Direct vasodilation of arterioles (with little effect on veins) with decreased systemic resistance. Effect: Management of moderate-to-severe hypertension, congestive heart failure, hypertension secondary to pre-eclampsia/eclampsia. Treatment of primary pulmonary hypertension. | Mechanistic genes:AGPAT2, AGT, AKR1C4, CHRNA1, COL1A1, ESR1, GSTP1, HBB, HFE, HIF1A, MAOA, MGMT, NR3C1, PDGFRB Metabolic genes Substrate:NAT2 Inhibitor:CEL, CYP3A4, CYP3A5 Transporter genes:SLC6A2, SLC12A3, SLC22A16 Pleiotropic genes:APC, HLA-A, HLA-B, IL6, IL10, TNF, TP53 |
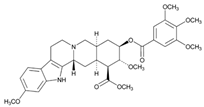 | Name: RESERPINE IUPAC Name: Yohimban-16-carboxylic acid, 11,17-dimethoxy-18-[(3,4,5-trimethoxybenzoyl)oxy]-, methyl ester, (3β,16β,17α,18β,20α)-; methyl 18β-hydroxy-11,17α-dimethoxy-3β,20α-yohimban-16β-carboxylate 3,4,5-trimethoxybenzoate (ester) Molecular Formula: C33H40N2O9 Molecular Weight: 608.68 Mechanism: Reduces blood pressure via depletion of sympathetic biogenic amines (norepinephrine and dopamine). This also commonly results in sedative effects. Effect: Management of mild-to-moderate hypertension. Treatment of agitated psychotic states (schizophrenia). | Mechanistic genes:COMT, ERBB2, LDLR, MAOA, MAOB, NR1I2 Metabolic genes Substrate:CYP1A1, CYP3A4, CYP3A5, CYP7A1, UGT1A1 Inhibitor:ABCB1, ABCG2 Inducer:ABCB1 Transporter genes:ABCB11, SLC18A2 |
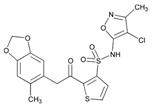 | Name: SITAXENTAN IUPAC Name: N-(4-chloro-3-methyl-5-isoxazolyl)-2-[[4,5-(methylenedioxy)-o-toly]acetyl]-3-thiophenesulfonamide Molecular Formula: C18H15ClN2O6S2 Molecular Weight: 454.90 Mechanism: A selective antagonist of A subtype of endothelin-1 receptors (ETA) located in pulmonary smooth muscle. Stimulation of these receptors by endogenous endothelin-1 causes vasoconstriction. Sitaxsentan exhibits 6500-fold greater selectivity for ETA over the ETB subtype; the latter predominates on vascular endothelial cells. Thus, preferential antagonism of ETA reduces vasoconstriction, without compromising vasodilatory/antiproliferative actions mediated through endothelin-1 binding to ETB subtype. Effect: Primary pulmonary arterial hypertension or pulmonary hypertension secondary to connective tissue disease. | Mechanistic genes:EDNRA, EDNRB Metabolic genes Substrate:CYP2C9, CYP3A4, CYP3A5 Inhibitor:CYP2C9, CYP2C19, CYP3A4, CYP3A5 Transporter genes:ABCB1 |
| Beta Blocking Agents | ||
| Drug | Properties | Pharmacogenetics |
 | Name: ATENOLOL IUPAC Name: Benzeneacetamide, 4-[2-hydroxy-3-[(1-methylethyl)amino]propoxy]-; 2-[p-[2-hydroxy-3-(isopropylamino)propoxy]phenyl]acetamide Molecular Formula: C14H22N2O3 Molecular Weight: 266.34 Mechanism: Competitively blocks response to β-adrenergic stimulation, selectively blocks β1 receptors with little or no effect on β2 receptors except at high doses. Effect: Treatment of hypertension. Management of angina pectoris, postmyocardial infarction. | Mechanistic genes:ACE1, ACE2, ADRB1, ADRB2, AGT, APOB, BDKRB2, EDN1, ERAP1, GNAS, GNB3, GRK5, LDLR Metabolic genes Substrate:CYP2C9 |
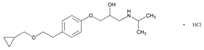 | Name: BETAXOLOL IUPAC Name: 2-Propanol, 1-[4-[2-(cyclopropylmethoxy)ethyl]phenoxy]-3-[(1-methylethyl)amino]-, hydrochloride, (±)-; (2)(±)-1-[p-[2-(cyclopropylmethoxy)ethyl]phenoxy]-3-(isopropylamino)-2-propanol hydrochloride Molecular Formula: C18H29NO3 HCl Molecular Weight: 343.89 Mechanism: Competitively blocks β1 receptors, with little or no effect on β2 receptors (bronchial and vascular smooth muscle; only at high doses). No intrinsic sympathomimetic activity and little or no membrane-stabilizing effect (local anesthetic) on the heart. Reduces blood pressure by decreasing cardiac output, decreasing sympathetic outflow from the CNS, and/or suppressing renin release. Reduces intraocular pressure by reducing the production of aqueous humor (may block endogenous catecholamine-stimulated increases in cyclic adenosine monophosphate concentrations within the ciliary processes and subsequent formation of aqueous humor). Effect: Reduction in elevated intraocular pressure in chronic open-angle glaucoma and ocular hypertension (has been used effectively in glaucoma following laser trabeculoplasty). Reduction in systemic hypertension. Initial management of hypertension in heart failure, postmyocardial infarction, high coronary disease risk, and/or diabetes mellitus. | Mechanistic genes:ADRB1, ADRB2, AGT, BDKRB2, GNAS Metabolic genes Substrate:CYP1A2, CYP2D6 Inhibitor:CYP2D6 |
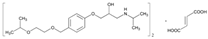 | Name: BISOPROLOL IUPAC Name: 2-Propanol, 1-[4-[[2-(1-methylethoxy)ethoxy]methyl]phenoxy]-3-[(1-methylethyl)amino]-, (±)-, I-2-butenedioate (2:1); (2)(±)-1-[[α-(2-Isopropoxyethoxy)-p-tolyl]oxy]-3-(isopropylamino)-2-propanol fumarate (2:1) Molecular Formula: (C18H31NO4)2 C4H4O4 Molecular Weight: 766.96 Mechanism: Selective inhibitor of β1-adrenergic receptors (competitively blocks β1 receptors in myocardium), with little or no effect on β2 receptors at doses ≤20 mg (at high doses may block β2-adrenergic receptors within the bronchial and vascular smooth muscle). Decreases resting and exercise-stimulated heart rate and cardiac output, decreases isoproterenol-induced tachycardia, prolongs sinus node recovery time, refractory period of the AV node, and AV nodal conduction (with rapid atrial stimulation). No intrinsic sympathomimetic activity or membrane-stabilizing effect on the heart. Reduces blood pressure by decreasing cardiac output, decreasing sympathetic outflow from the CNS, and/or suppressing renin release. Effect: Treatment of hypertension, alone or in combination with other agents. Management of mild to moderately severe heart failure of ischemic or cardiomyopathic origin in conjunction with other agents (do not use in patients with acutely decompensated heart failure requiring I.V. inotropic therapy, those with substantial fluid retention requiring intensive diuresis, and those who require hospitalization for heart failure). | Mechanistic genes:ACE1, ADRB1, AGT, BDKRB2, GNAS Metabolic genes Substrate:CYP2D6, CYP3A4, CYP3A5 |
 | Name: CARTEOLOL IUPAC Name: 2(1H)-quinolinone, 5-[3-[(1,1-dimethylethyl)amino]-2-hydroxypropoxy]-3,4-dihydro-, monohydrochloride; 5-[3-(tert-Butylamino)-2-hydroxypropoxy]-3,4-dihydrocarbostyril monohydrochloride Molecular Formula: C16H24N2O3 HCl Molecular Weight: 328.83 Mechanism: Blocks both β1 and β2 receptors and has mild intrinsic sympathomimetic activity. Reduces intraocular pressure by decreasing aqueous humor production. Has negative inotropic and chronotropic effects and can significantly slow AV nodal conduction. Effect: Chronic open-angle glaucoma and intraocular hypertension. | Mechanistic genes:ADRB1, ADRB2 Metabolic genes Substrate:CYP2D6, CYP3A4, CYP3A5 |
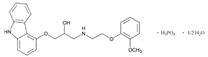 | Name: CARVEDILOL IUPAC Name: 2-Propanol, 1-(9H-carbazol-4-yloxy)-3-[[2-(2-methoxyphenoxy)ethyl]amino]-, phosphate (salt), hydrate (2:2:1) Molecular Formula: C24H26N2O4 H3O4P ½H2O Molecular Weight: 513.48 Mechanism: Non-selective β-adrenergic blocker with α-adrenergic blocking activity. Available as a racemic mixture. Does not possess intrinsic sympathomimetic activity. Has calcium channel blocking activity at higher dose (30-fold the normal dose). In hypertensive patients, reduces cardiac output, exercise- or β-agonist-induced tachycardia, reflex orthostatic tachycardia, vasodilatation, peripheral vascular resistance (especially in standing position), renal vascular resistance, plasma renin activity, and increases levels of atrial natriuretic peptide. In congestive heart failure, decreases systemic blood pressure, pulmonary capillary wedge pressure, pulmonary artery pressure, heart rate, systemic vascular resistance, right arterial pressure, and increases stroke volume index and left ventricular ejection fraction. Effect: Mild-to-severe heart failure of ischemic or cardiomyopathic origin (usually in addition to standard therapy). Left ventricular dysfunction following myocardial infarction (clinically stable with LVEF ≤40%). Management of hypertension. | Mechanistic genes:ADRA1A, ADRB1, ADRB2, GRK5, MMP2, NOX1 Metabolic genes Substrate:CYP1A2, CYP2C9, CYP2C19, CYP2D6, CYP2E1, CYP3A4, CYP3A5, UGT1A1, UGT1A4, UGT1A6, UGT2B7 Inhibitor:ABCB1 Transporter genes:ABCB1 |
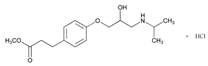 | Name: ESMOLOL IUPAC Name: Benzenepropanoic acid, 4-[2-hydroxy-3-[(1-methylethyl)amino]propoxy]-, methyl ester, hydrochloride, (±)-; (±)-methyl p-[2-hydroxy-3-(isopropylamino)propoxy]hydrocinnamate hydrochloride Molecular Formula: C16H25NO4 HCl Molecular Weight: 331.83 Mechanism: A short-acting β1-selective adrenergic blocking agent. Competitively blocks response to β1-adrenergic stimulation with little or no effect on β2 receptors except at high doses, no intrinsic sympathomimetic activity, no membrane-stabilizing activity. Effect: Used in management of supraventricular tachyarrhythmias (e.g., atrial flutter and/or fibrillation, and sinus tachycardia). Used to prevent or treat increases in blood pressure associated with surgical events, including hypertensive crises (i.e., emergencies and urgencies). Treatment of non-compensatory sinus tachycardia. | Mechanistic genes:ADRB1 Metabolic genes Substrate:CYP2D6 |
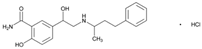 | Name: LABETALOL IUPAC Name: Benzamide, 2-hydroxy-5-[1-hydroxy-2-[(1-methyl-3-phenylpropyl)amino]ethyl]-, monohydrochloride; 5-[1-hydroxy-2-[(1-methyl-3-phenylpropyl)amino]ethyl]salicylamide monohydrochloride Molecular Formula: C19H24N2O3 HCl Molecular Weight: 364.87 Mechanism: Blocks α-, β1-, and β2-adrenergic receptor sites. Elevated renins reduced. Effect: Treatment of mild to severe hypertension. Used intravenously for hypertensive emergencies. | Mechanistic genes:ADRA, ADRB1, ADRB2 Metabolic genes Substrate:UGT1A1, UGT2B7 |
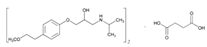 | Name: METOPROLOL IUPAC Name: 2-Propanol, 1-[4-(2-methoxyethyl)phenoxy]-3-[(1-methylethyl)amino]-, (±)-, butanedioate (2:1); (±)-1-(isopropylamino)-3-[p-(2-methoxyethyl)phenoxy]-2-propanol succinate (2:1) Molecular Formula: (C15H25NO3)2 C4H6O4 Molecular Weight: 652.82 Mechanism: Competitively blocks β1 receptors, with little or no effect on β2 receptors at doses <100 mg. Effect: Treatment of angina pectoris, hypertension, or hemodynamically stable acute myocardial infarction. Treatment of angina pectoris or hypertension. To reduce mortality/hospitalization in heart failure patients already receiving ACEIs, diuretics, and/or digoxin. | Mechanistic genes:ADRB1, ADRA2C, ACE, GRK5, KCNH2 Metabolic genes Substrate:CYP2C19, CYP2D6 Inhibitor:CYP2D6 Transporter genes:ABCB1 |
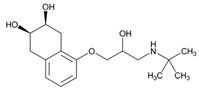 | Name: NADOLOL IUPAC Name: 2,3-Naphthalenediol, 5-[3-[(1,1-dimethylethyl)amino]-2-hydroxypropoxy]-1,2,3,4-tetrahydro-, cis-; 1-(tert-butylamino)-3-[(5,6,7,8-tetrahydro-cis-6,7-dihydroxy-1-naphthyl)oxy]-2-propanol Molecular Formula: C17H27NO4 Molecular Weight: 309.40 Mechanism: Competitively blocks response to β1- and β2-adrenergic stimulation. Effect: Treatment of hypertension and angina pectoris. Prophylaxis of migraine headaches. | Mechanistic genes:ADRB1, ADRB2, ADRB3 Transporter genes:ABCB1 Pleiotropic genes:IL10, IL12B |
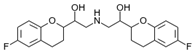 | Name: NEBIVOLOL IUPAC Name: 2-H-1-benzopyran-2-methanol, α,α’-[iminobis(methylene)bis[6-fluoro-3,4-dihydro-; α,α’-(iminodimethylene)bis[6-fluoro-2-chromanmethanol] Molecular Formula: C22H25F2NO4 Molecular Weight: 405.44 Mechanism: Highly-selective inhibitor of β1-adrenergic receptors. Nebivolol, unlike other β-blockers, also produces an endothelium-derived nitric oxide-dependent vasodilation resulting in a reduction in systemic vascular resistance. Effect: Treatment of hypertension, alone or in combination with other agents. | Mechanistic genes:ADRB1 Metabolic genes Substrate:CYP2D6, UGTs Inducer:NOS3 |
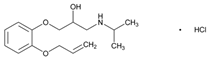 | Name: OXPRENOLOL IUPAC Name: 2-Propanol, 1-(o-allyloxyphenoxy)-3-isopropylamino-, hydrochloride Molecular Formula: C15H23NO3 HCl Molecular Weight: 301.81 Mechanism: A competitive and non-selective antagonist of β-adrenergic receptors. Antagonizes catecholamine-induced tachycardia, thus decreasing cardiac output. Inhibits renin release by kidneys and inhibits vasomotor centers. Effect: Mild or moderate hypertension. | Mechanistic genes:ADRB1, ADRB2, ADRB3 Metabolic genes Inhibitor:CYP2D6 |
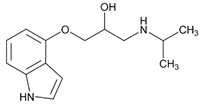 | Name: PINDOLOL IUPAC Name: 2-Propanol, 1-(1H-indol-4-yloxy)-3-(1-methylethyl)amino-; 1-(indol-4-yloxy)-3-(isopropylamino)-2-propanol Molecular Formula: C14H20N2O2 Molecular Weight: 248.32 Mechanism: Blocks both β1 and β2 receptors and has mild intrinsic sympathomimetic activity. Has negative inotropic and chronotropic effects and can significantly slow AV nodal conduction. Augmentative action of antidepressants thought to be mediated via serotonin 1A autoreceptor antagonism. Effect: Treatment of hypertension, alone or in combination with other agents. | Mechanistic genes:ADRB1, ADRB2, ADRB3, GRK5, HTR1A, HTR1B Metabolic genes Substrate:CYP2D6 Inhibitor:CYP2D6 |
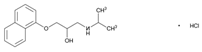 | Name: PROPRANOLOL IUPAC Name: 2-Propanol, 1-[(1-methylethyl)amino]-3-(1-naphthalenyloxy)-, hydrochloride, (±)-; (±)-1-(isopropylamino)-3-(1-naphthyloxy)-2-propanol hydrochloride Molecular Formula: C16H21NO2 HCl Molecular Weight: 295.80 Mechanism: Competitively blocks response to β1- and β2-adrenergic stimulation, which results in decreases in heart rate, myocardial contractility, blood pressure, and myocardial oxygen demand. Reduces portal pressure by producing splanchnic vasoconstriction (β2-effect) thereby reducing portal blood flow. Effect: Management of hypertension. Angina pectoris. Pheochromocytoma. Essential tremor. Supraventricular arrhythmias (such as atrial fibrillation and flutter, AV nodal re-entrant tachycardias), ventricular tachycardias (catecholamine-induced arrhythmias, digoxin toxicity). Prevention of myocardial infarction. Migraine headache prophylaxis. Symptomatic treatment of hypertrophic subaortic stenosis (hypertrophic obstructive cardiomyopathy). | Mechanistic genes:ADRB1, ADRB2, ADRB3, ALOX5, CFTR, COMT, FOS, GNAS, GRK5, HTR1B, HTR3B, KCNE2; KCNH2, KCNQ1, NPY, PPARGC1A, PTGS2 Metabolic genes Substrate:CYP1A1, CYP1A2, CYP2C9, CYP2C19, CYP2D6, CYP3A4, CYP3A5, UGT2B7 Inhibitor:ABCB1, CYP1A2, CYP2D6 Transporter genes:ABCB1 Pleiotropic genes:IL1B, TNF |
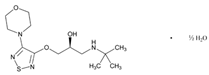 | Name: TIMOLOL IUPAC Name: 2-Propanol, 1-[(1,1-dimethylethyl)amino]-3-[[4-(4-morpholinyl)-1,2,5-thiadiazol-3-yl]oxy]-, hemihydrate, (S)-; (S)-1-(tert-butylamino)-3-[(4-morpholino-1,2,5-thiadiazol-3-yl)oxy]-2-propanol hemihydrate Molecular Formula: C13H24N4O3S ½H2O Molecular Weight: 325.43 Mechanism: Blocks both β1- and β2-adrenergic receptors, reduces intraocular pressure by reducing aqueous humor production or possibly outflow. Reduces blood pressure by blocking adrenergic receptors and decreasing sympathetic outflow, produces negative chronotropic and inotropic activity through unknown mechanism. Effect: Treatment of elevated intraocular pressure such as glaucoma or ocular hypertension. Treatment of hypertension and angina, to reduce mortality following myocardial infarction. Prophylaxis of migraine. | Mechanistic genes:ADRB1, GNAS Metabolic genes Substrate:CYP2C19, CYP2D6 Inhibitor:CYP2D6 |
| Calcium-Channel Blockers | ||
| Drug | Properties | Pharmacogenetics |
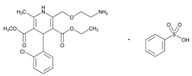 | Name: AMLODIPINE IUPAC Name: 3,5-Pyridinedicarboxylic acid, 2-[(2-aminoethoxy)methyl]-4-(2-chlorophenyl)-1,4-dihydro-6-methyl-, 3-ethyl 5-methyl ester, (±)-, monobenzenesulfonate; 3-ethyl 5-methyl (±)-2-[(2-aminoethoxy)methyl]-4-(o-chlorophenyl)-1,4-dihydro-6-methyl-3,5-pyridinedicarboxylate, monobenzenesulfonate Molecular Formula: C20H25ClN2O5 C6H6O3S Molecular Weight: 567.05 Mechanism: Inhibits calcium ion from entering the voltage-sensitive channels of vascular smooth muscle and myocardium during depolarization, producing a relaxation of coronary vascular smooth muscle and coronary vasodilation. Increases myocardial oxygen delivery in vasospastic angina. Effect: Treatment of hypertension, symptomatic chronic stable angina, vasospastic (Prinzmetal’s) angina. Prevention of hospitalization due to angina with documented coronary artery disease. | Mechanistic genes:ADD1, AGT, CACNs, NPPA Metabolic genes Substrate:CYP3A4, CYP3A5 Inhibitor:ABCB1, CYP1A1, CYP1A2, CYP2A6, CYP2B6, CYP2C8, CYP2C9, CYP2D6, CYP3A4, CYP3A5 Transporter genes:ABCB1 |
 | Name: DILTIAZEM IUPAC Name: 1,5-Benzothiazepin-4(5H)-one, 3-(acetyloxy)-5-[2-(dimethylamino)ethyl]-2,3-dihydro-2-(4-methoxyphenyl)-, monohydrochloride, (+)-cis-; (+)-5-[2-(dimethylamino)ethyl]-cis-2,3-dihydro-3-hydroxy-2-(p-methoxyphenyl)-1,5-benzothiazepin-4(5H)-one acetate (ester) monohydrochloride Molecular Formula: C22H26N2O4S HCl Molecular Weight: 450.98 Mechanism: Inhibits calcium ions from entering the “slow channels” or select voltage-sensitive areas of vascular smooth muscle and myocardium during depolarization, producing a relaxation of coronary vascular smooth muscle and coronary vasodilation. Increases myocardial oxygen delivery in vasospastic angina. Effect: Essential hypertension, chronic stable angina or angina from coronary artery spasm. Temporary control of rapid ventricular rate in atrial fibrillation or flutter. Management of supraventricular tachyarrhythmias, including rapid conversion to sinus rhythm of paroxysmal supraventricular tachycardias (e.g., those associated with Wolff–Parkinson–White or Lown–Ganong–Levine syndrome). | Mechanistic genes:CACNA Metabolic genes Substrate:CYB5s, CYP2C8, CYP2C9, CYP2D6, CYP3A4, CYP3A5 Inhibitor:ABCB1, CYB5s, CYP2C9, CYP2D6, CYP3A4, CYP3A5 Transporter genes:ABCB1, SLCO1B1 Pleiotropic genes:IL12B |
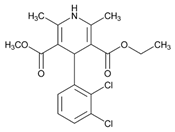 | Name: FELODIPINE IUPAC Name: 3,5-Pyridinedicarboxylic acid 4-(2,3-dichlorophenyl)-1,4-dihydro-2,6-dimethyl-, ethyl methyl ester, (±)-; (±)-ethyl methyl 4-(2,3-dichlorophenyl)-1,4-dihydro-2,6-dimethyl-3,5-pyridinedicarboxylate Molecular Formula: C18H19Cl2NO4 Molecular Weight: 384.25 Mechanism: Inhibits calcium ions from entering the “slow channels” or select voltage-sensitive areas of vascular smooth muscle and myocardium during depolarization, producing a relaxation of coronary vascular smooth muscle and coronary vasodilation. Increases myocardial oxygen delivery in vasospastic angina. Effect: Treatment of hypertension. | Mechanistic genes:CACNA1C, NR1I2 Metabolic genes Substrate:CYP3A4, CYP3A5 Inhibitor:ABCB1, CYP2C8, CYP2C9, CYP2D6, CYP3A4, CYP3A5 |
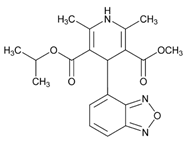 | Name: ISRADIPINE IUPAC Name: 3,5-Pyridinedicarboxylic acid, 4-(4-benzofurazanyl)-1,4-dihydro-2,6-dimethyl-, methyl 1-methylethyl ester, (±)-; isopropyl methyl (±)-4-(4-benzofurazanyl)-1,4-dihydro-2,6-dimethyl-3,5-pyridinedicarboxylate Molecular Formula: C19H21N3O5 Molecular Weight: 371.39 Mechanism: Inhibits transmembrane influx of extracellular calcium ions across membranes of myocardial cells and vascular smooth muscle cells, without changing serum calcium concentrations. Increases myocardial oxygen delivery in vasospastic angina. Effect: Management of hypertension (alone or in combination with other classes of antihypertensive agents). | Mechanistic genes:CACNA1C, NR1I2 Metabolic genes Substrate:CYP3A4, CYP3A5 Inhibitor:CYP3A4, CYP3A5 |
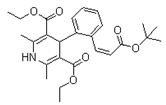 | Name: LACIDIPINE IUPAC Name: 3,5-Pyridinedicarboxylic acid, 4-[2-[3-(1,1-dimethylethoxy)-3-oxo-1-propenyl]phenyl]-1,4-dihydro-2,6-dimethyl-, diethyl ester, I-; 4-I(E)-2-carboxyvinyl]-phenyl]-1,4-dihydro-2,6-dimethyl-3,5-pyridinedicarboxylic acid, 4-tert-butyl diethyl ester Molecular Formula: C26H33NO6 Molecular Weight: 455.54 Mechanism: A specific and potent calcium antagonist with predominant selectivity for calcium channels in vascular smooth muscle. Its main action is to dilate peripheral arterioles, reducing peripheral vascular resistance and lowering blood pressure. Effect: Indicated for treatment of hypertension either alone or in combination with other antihypertensive agents, including β-adrenoceptor antagonists, diuretics, and ACEIs. | Mechanistic genes:CACN Metabolic genes Substrate: CYP3A4, CYP35 Inhibitor: CYP3A4, CYP35 |
 | Name: NICARDIPINE IUPAC Name: 3,5-Pyridinedicarboxylic acid, 1,4-dihydro-2,6-dimethyl-4-(3-nitrophenyl)-, methyl 2-[methyl(phenylmethyl)amino]ethyl ester, monohydrochloride; 2-(benzylmethylamino)ethyl methyl 1,4-dihydro-2,6-dimethyl-4-(m-nitrophenyl)-3,5-pyridinedicarboxylate monohydrochloride Molecular Formula: C26H29N3O6 HCl Molecular Weight: 515.99 Mechanism: Inhibits calcium ions from entering “slow channels” or select voltage-sensitive areas of vascular smooth muscle and myocardium during depolarization, producing relaxation of coronary vascular smooth muscle and coronary vasodilation. Increases myocardial oxygen delivery in vasospastic angina. Effect: Chronic stable angina. Management of hypertension. | Mechanistic genes:CACNA1C, CACNA1D, CACNA2D1, CACNB2 Metabolic genes Substrate:CYP1A1, CYP1A2, CYP2B6, CYP2C8, CYP2C9, CYP2D6, CYP2E1, CYP3A4, CYP3A5 Inhibitor:ABCB1, CYP1A1, CYP2A6, CYP2C8, CYP2C9, CYP2C19, CYP2D6, CYP3A4, CYP3A5 Inducer:CYP1A2 Transporter genes:ABCB1 |
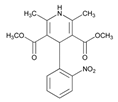 | Name: NIFEDIPINE IUPAC Name: 3,5-Pyridinedicarboxylic acid, 1,4-dihydro-2,6-dimethyl-4-(2-nitrophenyl)-, dimethyl ester; dimethyl 1,4-dihydro-2,6-dimethyl-4-(o-nitrophenyl)-3,5-pyridinedicarboxylate Molecular Formula: C17H18N2O6 Molecular Weight: 346.33 Mechanism: Inhibits calcium ions from entering “slow channels” or select voltage-sensitive areas of vascular smooth muscle and myocardium during depolarization, producing relaxation of coronary vascular smooth muscle and coronary vasodilation. Increases myocardial oxygen delivery in vasospastic angina. Effect: Angina and hypertension. Pulmonary hypertension. | Mechanistic genes:ACE1, ACE2, CACNA1C, DRD2, FOS, MMP2, SCN5A Metabolic genes Substrate:CYP2D6, CYP2C8, CYP11B2, CYP3A4, CYP3A5, POR Inhibitor:ABCB1, CYP1A2, CYP2C9, CYP2D6, CYP2E1, CYP3A4, CYP3A5 Transporter genes:ABCB1, ABCC2, ABCC3, SLC14A2 Pleiotropic genes:TP53 |
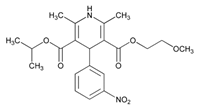 | Name: NIMODIPINE IUPAC Name: 3,5-Pyridinedicarboxylic acid, 1,4-dihydro-2,6-dimethyl-4-(3-nitrophenyl)-, 2-methoxyethyl 1-methylethyl ester; isopropyl 2-methoxyethyl 1,4-dihydro-2,6-dimethyl-4-(m-nitrophenyl)-3,5-pyridinedicarboxylate Molecular Formula: C21H26N2O7 Molecular Weight: 418.44 Mechanism: Animal studies indicate that nimodipine has greater effect on cerebral arterials than other arterials; this increased specificity may be due to increased lipophilicity and cerebral distribution of the drug as compared to nifedipine. Inhibits calcium ions from entering “slow channels” or select voltage-sensitive areas of vascular smooth muscle and myocardium during depolarization. Effect: Spasm following subarachnoid hemorrhage from ruptured intracranial aneurysms regardless of patient’s postictus neurological condition. | Mechanistic genes:CACNA1C, DRD2 Metabolic genes Substrate:CYP3A4, CYP35 Pleiotropic genes:APP |
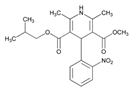 | Name: NISOLDIPINE IUPAC Name: 3,5-Pyridinedicarboxylic acid, 1,4-dihydro-2,6-dimethyl-4-(2-nitrophenyl)-, methyl 2-methylpropyl ester, (±)-; (±)-isobutyl methyl 1,4-dihydro-2,6-dimethyl-4-(o-nitrophenyl)-3,5-pyridinedicarboxylate Molecular Formula: C20H24N2O6 Molecular Weight: 388.41 Mechanism: As a dihydropyridine calcium-channel blocker, structurally similar to nifedipine, nisoldipine impedes movement of calcium ions into vascular smooth muscle and cardiac muscle. Dihydropyridines are potent vasodilators and not as likely to suppress cardiac contractility and slow cardiac conduction as other calcium antagonists such as verapamil and diltiazem. Nisoldipine is 5–10-fold as potent vasodilator as nifedipine. Effect: Management of hypertension, alone or in combination with other antihypertensive agents. | Mechanistic genes:CACNA1C Metabolic genes Substrate:CYP34, CYP3A5 Inhibitor:ABCB1, CYP1A2, CYP3A4, CYP3A5 |
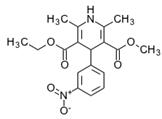 | Name: NITRENDIPINE IUPAC Name: 3,5-Pyridinedicarboxylic acid, 1,4-dihydro-2,6-dimethyl-4-(3-nitrophenyl)-, ethyl methyl ester, (±)-; (±)-ethyl methyl 1,4dihydro-2,6-dimethyl-4-(m-nitrophenyl)-3,5-pyridinedicarboxylate Molecular Formula: C18H20N2O6 Molecular Weight: 360.36 Mechanism: Dihydropyridine calcium channel-blocking agent with actions similar to nifedipine. Effect: Management of hypertension. | Mechanistic genes:CACNA1C, CACNG1 Metabolic genes Substrate:CYP3A4, CYP3A5 Inhibitor:ABCB1, CYP3A4, CYP3A5 Transporter genes:ABCG2 |
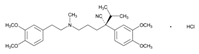 | Name: VERAPAMIL IUPAC Name: Benzeneacetonitrile, α-[3-[[2-(3,4-dimethoxyphenyl)ethyl]methylamino]propyl]-3,4-dimethoxy-α-(1-methylethyl)-, monohydrochloride, (±)-; (±)-5-[(3,4-dimethoxyphenethyl)methylamino]-2-(3,4-dimethoxyphenyl)-2-isopropylvaleronitrile monohydrochloride Molecular Formula: C27H38N2O4 HCl Molecular Weight: 491.06 Mechanism: Inhibits calcium ions from entering “slow channels” or select voltage-sensitive areas of vascular smooth muscle and myocardium during depolarization. Produces relaxation of coronary vascular smooth muscle and coronary vasodilation. Increases myocardial oxygen delivery in vasospastic angina. Slows automaticity and conduction of the AV node. Effect: Orally for treatment of angina pectoris (vasospastic, chronic stable, and unstable) and hypertension. I.V. for supraventricular tachyarrhythmias (PSVT, atrial fibrillation, and atrial flutter). | Mechanistic genes:ADRB1, ADRB2, CACNA1C, CACNs, CFTR, KCNMB1, LDLR, NOS1AP, RET, TGFB1 Metabolic genes Substrate:CYP1A2, CYP2B6, CYP2C8, CYP2C9, CYP2C18, CYP2C19, CYP2E1, CYP2J2, CYP3A4, CYP3A5, CYP3A7, SOD2 Inhibitor:ABCB1, ABCC1, ABCC2, CYP1A2, CYP2C8, CYP2C9, CYP2D6, CYP3A4, CYP3A5, CYP3A7 Inducer:ABCB1 Transporter genes:ABCB1, ABCC3, SLCO1B1 Pleiotropic genes:TNF |
| Cardiotonic Agents | ||
| Drug | Properties | Pharmacogenetics |
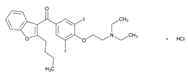 | Name: AMIODARONE IUPAC Name: Methanone, (2-butyl-3-benzofuranyl)[4-[2-(diethylamino)ethoxy]-3,5-diiodophenyl]-; 2-butyl-3-benzofuranyl 4-[2-(diethylamino)ethoxy]-3,5-diiodophenyl ketone Molecular Formula: C25H29I2NO3 Molecular Weight: 645.31 Mechanism: Inhibits adrenergic stimulation (α- and β-blocking properties), affects sodium, potassium and calcium channels, and prolongs the action potential and refractory period in myocardial tissue. Decreases AV conduction and sinus node function. Effect: Management of recurrent ventricular fibrillation or hemodynamically-unstable ventricular tachycardia. | Mechanistic genes:ABL1, ACOX1, ADRA2A, ADRB1, ADRB2, CHRM2, FABP1, FMO1, FOS, ICAM1, KCNE1, KCNE2, KCNH2, KCNJ11, KCNQ1, PSEN1, SCN5A Metabolic genes Substrate:CYP1A1, CYP1A2, CYP2C8, CYP2C9, CYP2C19, CYP2D6, CYP2J2, CYP3A4, CYP3A5 Inhibitor:ABCB1, CYP1A2, CYP2A6, CYP2B6, CYP2C8, CYP2C9, CYP2C19, CYP2D6, CYP3A4, CYP3A5 Inducer:CYP1A2 Transporter genes:ABCB1, ABCC6, ABCC8, SLC5A5 |
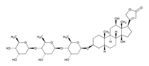 | Name: DIGOXIN IUPAC Name: Card-20(22)-enolide, 3-[(O-2,6-dideoxy-β-D-ribo-hexopyranosyl-(1→4)-O-2,6-dideoxy-β-D-ribo-hexopyranosyl-(1→4)-2,6-dideoxy-β-D-ribo-hexopyranosyl)oxy]-12,14-dihydroxy-, (3β,5β,12β)-; 3β-[(O-2,6-dideoxy-β-D-ribo-hexopyranosyl-(1→4)-O-2,6-dideoxy-β-D-ribo-hexopyranosyl-(1→4)-2,6-dideoxy-β-D-ribo-hexopyranosyl)oxy]-12β,14-dihydroxy-5β-card-20(22)-enolide Molecular Formula: C41H64O14 Molecular Weight: 780.94 Mechanism: Digoxin is a cardiac glycoside with positive inotropic effects. In congestive heart failure it inhibits the sodium/potassium ATPase pump which acts to increase the intracellular sodium-calcium exchange to increase intracellular calcium, leading to increased contractility. In supraventricular arrhythmias: Direct suppression of the AV node conduction to increase effective refractory period and decrease conduction velocity; positive inotropic effect, enhanced vagal tone, and decreased ventricular rate to fast atrial arrhythmias. Atrial fibrillation may decrease sensitivity and increase tolerance to higher serum digoxin concentrations. Effect: Digitalization and maintenance therapy. Used principally in the prophylactic management and treatment of congestive heart failure and to control the ventricular rate in supraventricular tachyarrhythmias (e.g., atrial fibrillation or flutter). Used to improve left ventricular function in cardiogenic shock and atrial fibrillation or flutter with rapid ventricular rate. May be useful, especially in conjunction with a β-adrenergic blocking agent, in the treatment of angina pectoris when cardiomegaly and congestive heart failure are present. | Mechanistic genes:ATP1A1 Metabolic genes Substrate:CYP3A4, CYP3A5 Transporter genes:ABCB1, ABCB11, ABCG2, SLCO1B3 |
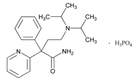 | Name: DISOPYRAMIDE IUPAC Name: 2-Pyridineacetamide, α-[2-[bis(1-methylethyl)amino]ethyl]-α-phenyl-, (±)-, phosphate (1:1); (±)-α-[2-(diisopropylamino)ethyl]-α-phenyl-2-pyridineacetamide phosphate (1:1) Molecular Formula: C21H29N3O H3PO4 Molecular Weight: 437.47 Mechanism: Decreases myocardial excitability and conduction velocity. Reduces disparity in refractory period between normal and infarcted myocardium. Possesses anticholinergic, peripheral vasoconstrictive, and negative inotropic effects. Effect: Suppression and prevention of unifocal and multifocal ventricular premature complexes, coupled ventricular premature complexes and/or paroxysmal ventricular tachycardia in primary arrhythmias or arrhythmias secondary to coronary artery disease. | Mechanistic genes:ADRB1, ADRB2, CHRM2, KCNE1, KCNE2, KCNH2, KCNJ11, KCNQ1 Metabolic genes Substrate:CYP2D6, CYP3A4, CYP3A5 Inhibitor:CYP1A1, CYP1A2, CYP2C19, CYP3A4, CYP3A5 Transporter genes:ABCC8 |
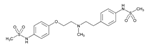 | Name: DOFETILIDE IUPAC Name: Methanesulfonamide, N-[4-[2-[methyl[2-[4-[(methylsulfonyl)amino]phenoxy]ethyl]amino]ethyl]phenyl]-; β-[(p-methanesulfonamidophenethyl)methylamino]methanesulfono-p-phenetidide Molecular Formula: C19H27N3O5S2 Molecular Weight: 441.56 Mechanism: Class III antiarrhythmic agent. Blockade of the cardiac ion channel carrying the rapid component of the delayed rectifier potassium current. It increases the monophasic action potential duration due to delayed repolarization. The increase in the QT interval is a function of prolongation of both effective and functional refractory periods in the His–Purkinje system and the ventricles. Effect: Used for the maintenance of normal sinus rhythm in patients with atrial fibrillation/flutter of more than 1 week duration who have been converted to normal sinus rhythm. Additionally used for the conversion of atrial fibrillation and atrial flutter to normal sinus rhythm. | Mechanistic genes:ADRA2A, ADRB1, CHRM2, KCNE1, KCNE2, KCNH2, KCNJ11, KCNQ1 Metabolic genes Substrate:CYP2D6, CYP3A4, CYP3A5 Transporter genes:ABCC8 |
 | Name: DRONEDARONE IUPAC Name: N-[2-butyl-3-[4-[3-(dibutylamino)propoxy]benzoyl]-1-benzofuran-5-yl] methanesulfonamide, hydrochloride Molecular Formula: C31H44N2O5S HCl Molecular Weight: 593.22 Mechanism: Dronedarone has antiarrhythmic properties belonging to all four antiarrhythmic (Vaughan-Williams) classes, but the contribution of each of these activities to the clinical effect is unknown. Inhibits sodium (INa) and potassium (Ikr, IkS, Ik1, and Ik-ACh) channels resulting in prolongation of the action potential and refractory period in myocardial tissue without reverse-use-dependent effects. Decreases AV conduction and sinus node function through inhibition of calcium (ICa-L) channels and β1-receptor blocking activity. Similar to amiodarone, dronedarone also inhibits α1-receptor-mediated increases in blood pressure. Effect: Indicated to reduce the risk of hospitalization related to paroxysmal or persistent atrial fibrillation (AF) or atrial flutter (AFl) in patients with a recent episode of AF/AFl and associated cardiovascular risk factors (e.g., age >70 years, hypertension, diabetes, prior cerebrovascular accident, left atrial diameter ≥50 mm or left ventricular ejection fraction <40%), who are in normal sinus rhythm or will be cardioverted. | Mechanistic genes:KCNA5, KCNE2, KCNH2 Metabolic genes Substrate:CYP3A4, CYP3A5 Inhibitor:ABCB1, CYP2C9, CYP2D6, CYP3A4, CYP3A5 |
 | Name: FLECAINIDE IUPAC Name: Benzamide, N-(2-piperidinylmethyl)-2,5-bis(2,2,2-trifluoroethoxy)-, monoacetate Molecular Formula: C17H20F6N2O3 C2H4O2 Molecular Weight: 474.39 Mechanism: Flecainide is a local anesthetic-type class Ic antiarrhythmic agent. Slows conduction in cardiac tissue by altering transport of ions across cell membranes. Causes slight prolongation of refractory periods. Decreases rate of rise of action potential without affecting duration. Increases electrical stimulation threshold of ventricle, the His–Purkinje system. Possesses local anesthetic and moderate negative inotropic effects. Effect: Prevention and suppression of documented life-threatening ventricular arrhythmias (e.g., sustained ventricular tachycardia). Control of symptomatic, disabling supraventricular tachycardias in patients without structural heart disease where other agents fail. | Mechanistic genes:ADRA2A, CHRM2, FABP1, KCNE1, KCNE2, KCNH2, KCNJ11, KCNQ1, SCN5A Metabolic genes Substrate:CYP1A2, CYP2D6 Inhibitor:CYP2D6 Transporter genes:ABCC8 |
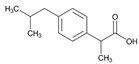 | Name: IBUPROFEN IUPAC Name: Benzeneacetic acid, α-methyl-4-(2-methylpropyl), (±)-; (±)-p-isobutylhydratropic acid; (3)(±)-2-(p-isobutylphenyl)propionic acid Molecular Formula: C13H18O2 Molecular Weight: 206.28 Mechanism: Inhibits prostaglandin synthesis by decreasing activity of enzyme cyclooxygenase (COX/PTGS), which results in decreased formation of prostaglandin precursors. Effect: Inflammatory diseases and rheumatoid disorders including juvenile rheumatoid arthritis, mild-to-moderate pain, fever, dysmenorrhea. Ibuprofen lysine is for use in premature infants weighing between 500 and 1500 g and who are ≤32 weeks gestational age to induce closure of clinically significant patent ductus arteriosus (PDA) when usual treatments are ineffective. Management of pain and swelling. | Mechanistic genes:ACSL1, AGT, BCAR1, CCND1, CNR1, ERBB4, FOS, ICAM1, IFNG, IGF1, LTA, MMP3, NOS3, NQO1, NR3C1, PPARA, PPARG, PTGER3, PTGES, PTGIS, PTGS1, PTGS2, RB1, REN, SCARB1, SNCA, TBX21, TBXAS1, VCAM1, VEGFA Metabolic genes Substrate:CYP1A2, CYP2B6, CYP2C8, CYP2C9, CYP2C19, CYP2D6, CYP2E1, CYP3A4, CYP3A5, CYP19A1, GSTT1, SOD2, UGT1A1, UGT1A3, UGT1A7, UGT1A9, UGT1A10, UGT2B7 Inhibitor:ACACA, CYP2C9, DDC, PTGS2, SLC5A8, SULT1A1, UGT2B15 Inducer:CYP19A1 Transporter genes:SLC15A1, SLC22A1, SLC22A6, SLC22A7, SLC22A8 Pleiotropic genes:APP, IL1B, IL1RN, IL4, IL6, IL10, TNF |
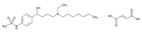 | Name: IBUTILIDE IUPAC Name: Methanesulfonamide, N-[4-[4-(ethylheptylamino)-1-hydroxybutyl]phenyl]-, (±)-, (E)-2-butenedioate (2:1); (±)-4′-[4-(ethylheptylamino)-1-hydroxybutyl]methanesulfonanilide fumarate (2:1) Molecular Formula: (C20H36N2O3S)2 C4H4O4 Molecular Weight: 885.23 Mechanism: Prolongs action potential duration and effective refractory period in both atrial and ventricular cardiac tissue. Delays repolarization by activating a slow, predominantly sodium, inward current. Produces dose-related prolongation of QT interval, thought to be associated with antiarrhythmic activity. Negligible effects on heart rate, cardiac contractility, or blood pressure. Lacks β-adrenergic-blocking activity. Effect: Used for rapid conversion of recent-onset atrial flutter or fibrillation to sinus rhythm. | Mechanistic genes:ADRA2A, CACNA1C, CHRM2, KCNE1, KCNE2, KCNH2, KCNJ11, KCNQ1 Metabolic genes Substrate:CYP2D6 Transporter genes:ABCB1, ABCC8 |
 | Name: INDOMETHACIN IUPAC Name: 1H-indole-3-acetic acid, 1-(4-chlorobenzoyl)-5-methoxy-2-methyl-; 1-(p-chlorobenzoyl)-5-methoxy-2-methylindole-3-acetic acid Molecular Formula: C19H16ClNO4 Molecular Weight: 357.79 Mechanism: Inhibits cyclooxygenase-1 (COX-1/PTGS1) and COX-2/PTGS2. Exhibits anti-inflammatory, analgesic, and antipyretic activity. Permits closure of ductus arteriosus in premature neonates by inhibiting prostaglandin synthesis. Effect: Symptomatic treatment of osteoarthritis, rheumatoid arthritis, and ankylosing spondylitis. Symptomatic relief of acute gout and acute painful shoulder (i.e., bursitis and/or tendinitis). Treatment of patent ductus arteriosus in premature neonates. | Mechanistic genes:AGTR1, BCAR1, CAT, CBR1, CBS, CCND1, CDK2, CDK4, COL1A1, DIO2, EDN1, EGFR, FOS, G6PD, GNAS, ICAM1, IFNG, LEP, LPL, LTA, MAPT, MMP2, MMP3, NOS3, NPPA, NPY, PARK2, PDGFRA, PDGFRB, PPARA, PPARD, PPARG, PTGER2, PTGES, PTGIS, PTGS1, PTGS2, RB1, TBXAS1, TGFB1, TGFBR1, THBD, TLR, UCP2, VEGFA, XDH Metabolic genes Substrate:CYP1A1, CYP1A2, CYP1B1, CYP2B6, CYP2C8, CYP2C9, CYP2C19, CYP2D6, CYP2E1, CYP3A4, CYP3A5, CYP7A1, CYP19A1, CYP27A1, GSTM1, GSTT1, SOD2, UGT1A1, UGT1A3, UGT1A4, UGT1A7, UGT1A9, UGT1A10, UGT2B7 Inhibitor: CYP2C9, CYP2C19, PTGS1, PTGS2, SLC22A7, SULT1A1 Transporter genes:ABCB1, ABCC1, ABCC2, ABCC3, ABCC6, ABCG2, SLC10A1, SLC19A1, SLC22A1, SLC22A2, SLC22A6, SLC22A7, SLC22A8 Pleiotropic genes:APC, APP, IL1B, IL1RN, IL2, IL4, IL6, IL10, IL12B, TNF, TNFRSF1A, TP53 |
 | Name: ISOSORBIDE DINITRATE IUPAC Name: D-glucitol, 1,4:3,6-dianhydro-, dinitrate Molecular Formula: C6H8N2O8 Molecular Weight: 236.14 Mechanism: Stimulation of intracellular cyclic GMP results in vascular smooth muscle relaxation of both arterial and venous vasculature. Effect: Acute relief of angina pectoris, prophylactic management in situations likely to provoke angina attacks, and long-term prophylactic management of angina pectoris. Additionally used to relieve pain, dysphagia, and spasm in esophageal spasm with gastroesophageal reflux. | Mechanistic genes:ALDH3A1, EDN1, NOS3 Metabolic genes Substrate:CYP3A4, CYP3A5, CYP11B2 |
 | Name: ISOSORBIDE MONONITRATE IUPAC Name: D-glucitol, 1,4:3,6-dianhydro-, 5-nitrate; 1,4:3,6-dianhydro-D-glucitol 5-nitrate Molecular Formula: C6H9NO6 Molecular Weight: 191.14 Mechanism: Decreases preload as measured by pulmonary capillary wedge pressure and left ventricular end diastolic volume and pressure. This effect improves congestive symptoms in heart failure and improves myocardial perfusion gradient in coronary artery disease. Effect: Long-acting metabolite of vasodilator isosorbide dinitrate used for prophylactic treatment of angina pectoris. | Mechanistic genes:NOS3 Metabolic genes Substrate:CYP3A4, CYP3A5, CYP11B2 |
 | Name: IVABRADINE IUPAC Name: 3-[3-[[[(7S)-3,4-dimethoxybicyclo[4.2.0]octa-1,3,4,5-tetrahydro-7,8-dimethoxy-2H-3-benzazepin-2-one Molecular Formula: C27H36N2O5 Molecular Weight: 468.59 Mechanism: Ivabradine is a pure heart rate-lowering agent, acting by selective and specific inhibition of the cardiac pacemaker If current that controls the spontaneous diastolic depolarization in the sinus node and regulates heart rate. The cardiac effects are specific to the sinus node with no effect on intra-atrial, atrioventricular or intraventricular conduction times, nor on myocardial contractility or ventricular repolarization. Effect: Symptomatic treatment of chronic stable angina pectoris in patients with normal sinus rhythm, who have a contraindication or intolerance for β-blockers. | Mechanistic genes:HCN1, HCN4 Metabolic genes Substrate:CYP3A4, CYP3A5 |
 | Name: LIDOCAINE IUPAC Name: Acetamide, 2-(diethylamino)-N-(2,6-dimethylphenyl)-; (2) 2-(diethylamino)-2′,6′-acetoxylidide Molecular Formula: C14H22N2O Molecular Weight: 234.34 Mechanism: Suppresses automaticity of conduction tissue, by increasing electrical stimulation threshold of ventricle, the His–Purkinje system, and spontaneous depolarization of ventricles during diastole by direct action on tissues. Effect: Local anesthetic and acute treatment of ventricular arrhythmias (such as from myocardial infarction or cardiac manipulation). | Mechanistic genes:CHRM2, FOS, KCNE2, KCNH2, KCNJ11, KCNQ1, SCN5A, TRPV1, VEGFA Metabolic genes Substrate:CYP1A2, CYP2A6, CYP2B6, CYP2C9, CYP2D6, CYP3A4, CYP3A5 Inhibitor:ABCB1, CYP1A2, CYP2D6, CYP3A4, CYP3A5 Transporter genes:ABCC8, SLC22A16 Pleiotropic genes:IL6 |
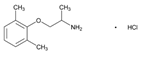 | Name: MEXILETINE IUPAC Name: 2-Propanamine, 1-(2,6-dimethylphenoxy)-, hydrochloride, (±)-; (±)-1-methyl-2-(2,6-xylyloxy)ethylamine hydrochloride Molecular Formula: C11H17NO HCl Molecular Weight: 215.72 Mechanism: Inhibits inward sodium current, decreases rate of rise of phase 0, increases effective refractory period/action potential duration ratio. Effect: Management of serious ventricular arrhythmias. Suppression of premature ventricular contractions. | Mechanistic genes:CHRM2, KCNE1, KCNE2, KCNH2, KCNJ11, KCNQ1, SCN5A Metabolic genes Substrate:CYP1A2, CYP2D6, CYP3A4, CYP3A5 Inhibitor:CYP1A2 Transporter genes:ABCC8 |
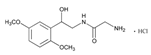 | Name: MIDODRINE IUPAC Name: Acetamide, 2-amino-N-[2-(2,5-dimethoxyphenyl)-2-hydroxyethyl]-, monohydrochloride, (±)-; (±)-2-amino-N-(β-hydroxy-2,5-dimethoxyphenethyl)acetamide monohydrochloride Molecular Formula: C12H18N2O4 HCl Molecular Weight: 290.74 Mechanism: Midodrine forms an active metabolite, desglymidodrine, an α1 agonist. This agent increases arteriolar and venous tone resulting in a rise in standing, sitting, and supine systolic and diastolic blood pressure in orthostatic hypotension. Effect: Orphan drug: Treatment of symptomatic orthostatic hypotension. | Mechanistic genes:ADRA1A, ADRA1B Metabolic genes Substrate:CYP2D6 |
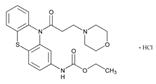 | Name: MORICIZINE IUPAC Name: Carbamic acid, [10-[3-(4-morpholinyl)-l-oxopropyl]-10H-phenothiazin-2yl]-, ethyl ester, hydrochloride; ethyl 10-(3-morpholinopropionyl)phenothiazine-2-carbamate, hydrochloride Molecular Formula: C22H25N3O4S HCl Molecular Weight: 463.98 Mechanism: Reduces fast inward current carried by sodium ions, shortens phase I and phase II repolarization, resulting in decreased action potential duration and effective refractory period. Effect: Treatment of ventricular tachycardia and life-threatening ventricular arrhythmias. | Mechanistic genes:SCN5A Metabolic genes Substrate:CYP3A4, CYP3A5 Inducer:CYP1A2, CYP3A4, CYP3A5 |
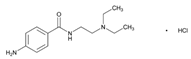 | Name: PROCAINAMIDE IUPAC Name: Benzamide, 4-amino-N-[2-(diethylamino)ethyl]-, monohydrochloride; p-amino-N-[2-(diethylamino)ethyl]benzamide monohydrochloride Molecular Formula: C13H21N3O HCl Molecular Weight: 271.79 Mechanism: Decreases myocardial excitability and conduction velocity and may depress myocardial contractility, by increasing electrical stimulation threshold of ventricle, the His–Purkinje system and through direct cardiac effects. Effect: Treatment of ventricular tachycardia, premature ventricular contractions, paroxysmal atrial tachycardia, and atrial fibrillation. Prevention of recurrence of ventricular tachycardia, paroxysmal supraventricular tachycardia, atrial fibrillation or flutter. | Mechanistic genes:CHRM2, KCNE1, KCNE2, KCNQ1, KCNH2, KCNJ11, SCN5A Metabolic genes Substrate:CYP2D6, CYP3A4, CYP3A5, NAT2 Transporter genes:ABCC8, SLC22A16 |
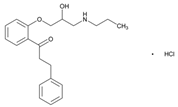 | Name: PROPAFENONE IUPAC Name: 1-Propanone, 1-[2-[2-hydroxy-3-(propylamino)propoxy]phenyl]-3-phenyl-, hydrochloride; 2′-[2-hydroxy-3-(propylamino)propoxy]-3-phenylpropiophenone hydrochloride Molecular Formula: C21H27NO3 HCl Molecular Weight: 377.90 Mechanism: Possesses local anesthetic properties, blocks fast inward sodium current, and slows rate of increase of action potential. Prolongs effective refractory period, reduces spontaneous automaticity and exhibits some β-blockade activity. Effect: Treatment of life-threatening ventricular arrhythmias. Maintenance of normal sinus rhythm in symptomatic atrial fibrillation. | Mechanistic genes:ADRB, CHRM2, KCNE1, KCNE2, KCNH2, KCNJ11, KCNQ1 Metabolic genes Substrate:CYP1A1, CYP1A2, CYP2D6, CYP3A4, CYP3A5, UGTs Transporter genes:ABCC8 |
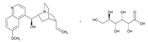 | Name: QUINIDINE IUPAC Name: (8R,9S)-6′-Methoxycinchonan-9-ol Molecular Formula: C20H24N2O2 Molecular Weight: 324.42 Mechanism: Depresses phase 0 of action potential. Decreases myocardial excitability and conduction velocity, and myocardial contractility by decreasing sodium influx during depolarization and potassium efflux in repolarization. Additionally reduces calcium transport across cell membrane. Decreases conduction velocity in atria, ventricles, and the His–Purkinje system, and may decrease or cause no change in conduction velocity through the AV node. May suppress atrial fibrillation or flutter. May produce sinus tachycardia via its anticholinergic effects. Has direct negative inotropic effect, but therapeutic plasma concentrations of drug do not usually depress contractility in normal heart. May reduce peripheral resistance and blood pressure by blockade of α-adrenergic receptors and by effects on myocardial contractility. Acts principally as intraerythrocytic schizonticide (has little effect on sporozoites or pre-erythrocytic parasites). Gametocidal against Plasmodium vivax and P. malariae, but not P. falciparum. Effect: Prophylaxis after cardioversion of atrial fibrillation and/or flutter to maintain normal sinus rhythm. Prevent recurrence of paroxysmal supraventricular tachycardia, paroxysmal AV junctional rhythm, paroxysmal ventricular tachycardia, paroxysmal atrial fibrillation, and atrial or ventricular premature contractions. Has activity against Plasmodium falciparum malaria. | Mechanistic genes:CHRM2, CHRNA1, FGB, G6PD, KCNE1, KCNE2, KCNH2, KCNJ11, KCNQ1, LIPC, SCN5A, TNFRSF1A Metabolic genes Substrate:CYP1A2, CYP2C9, CYP2D6, CYP2E1, CYP3A4, CYP3A5, GSTM1, GSTP1 Inhibitor:ABCB1, CYP2C9, CYP2D6, CYP3A4, CYP3A5 Transporter genes:ABCB1, ABCC1, ABCC8 |
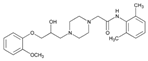 | Name: RANOLAZINE IUPAC Name: 1-Piperazineacetamide, N-(2,6-dimethylphenyl)-4-[2-hydroxy-3-(2-methoxyphenoxy)propyl]-; (±)-N-(2,6-dimethylphenyl)-4-[2-hydroxy-3-(2-methoxyphenoxy)propyl]-1-piperazineacetamide Molecular Formula: C24H33N3O4 Molecular Weight: 427.54 Mechanism: Ranolazine exerts antianginal and anti-ischemic effects without changing hemodynamic parameters (heart rate or blood pressure). At therapeutic levels, ranolazine inhibits late phase of inward sodium channel (late INa) in ischemic cardiac myocytes during cardiac repolarization reducing intracellular sodium concentrations and thereby reducing calcium influx via Na+-Ca2+ exchange. Decreased intracellular calcium reduces ventricular tension and myocardial oxygen consumption. It is thought that ranolazine produces myocardial relaxation and reduces anginal symptoms through this mechanism although this is uncertain. At higher concentrations, ranolazine inhibits rapid delayed rectifier potassium current (IKr) thus prolonging ventricular action potential duration and subsequent prolongation of QT interval. Effect: Treatment of chronic angina. | Mechanistic genes:SCN5A Metabolic genes Substrate:CYP2D6, CYP3A4, CYP3A5 Inhibitor:ABCB1, CYP2D6, CYP3A4, CYP3A5 Transporter genes:ABCB1 |
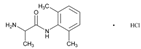 | Name: TOCAINIDE IUPAC Name: Propanamide, 2-amino-N-(2,6-dimethylphenyl)-, hydrochloride, (±)-; (±)-amino-2′,6′-propionoxylidide hydrochloride Molecular Formula: C11H16N2O.HCl Molecular Weight: 228.72 Mechanism: Suppresses automaticity of conduction tissue, by increasing electrical stimulation threshold of ventricle, the His–Purkinje system, and spontaneous depolarization of ventricles during diastole by direct action on tissues. Blocks both initiation and conduction of nerve impulses by decreasing permeability to sodium ions of neuronal membrane, which results in inhibition of depolarization with resultant blockade of conduction. Effect: Suppression and prevention of symptomatic life-threatening ventricular arrhythmias. | Mechanistic genes:CHRM2, KCNE1, KCNE2, KCNH2, KCNJ11, KCNQ1, SCN5A Inhibitor:CYP1A2 Transporter genes:ABCC8 |
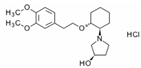 | Name: VERNAKALANT IUPAC Name: (3R)-1-[(1R,2R)-2-[2-(3,4-dimethoxyphenyl)ethoxy]cyclo34enzazepineolidin-3-ol hydrochloride Molecular Formula: C20H31NO4 HCl Molecular Weight: 385.20 Mechanism: Vernakalant is an antiarrhythmic medicine that acts preferentially in the atria to prolong atrial refractoriness and to rate-dependently slow impulse conduction. These antifibrillatory actions on refractoriness and conduction are thought to suppress re-entry, and are potentiated in the atria during atrial fibrillation. The relative selectivity of vernakalant on atrial vs. ventricular refractoriness is postulated to result from the block of currents that are expressed in the atria, but not in the ventricles, as well as the unique electrophysiologic condition of the fibrillating atria. However, blockade of cationic currents, including hERG channels and cardiac voltage-dependent sodium channels present in the ventricles, has been documented. Effect: Rapid conversion of recent onset atrial fibrillation to sinus rhythm in adults (for non-surgery patients, atrial fibrillation ≤7 days duration; and for post-cardiac surgery patients, atrial fibrillation ≤3 days duration). | Mechanistic genes:KCNA5, KCNH2 Metabolic genes Substrate:CYP2D6, UGTs |
| Diuretics | ||
| Drug | Properties | Pharmacogenetics |
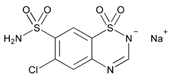 | Name: CHLOROTHIAZIDE IUPAC Name: 2H-1,2,4-benzothiadiazine-7-sulfonamide, 6-chloro-, 1,1-dioxide, monosodium salt; 6-chloro-2H-1,2,4-benzothiadiazine-7-sulfonamide, 1,1-dioxide, monosodium salt Molecular Formula: C7H5ClN3NaO4S2 Molecular Weight: 317.71 Mechanism: Primary site of diuretic action appears to be the cortical diluting segment of the nephron. Enhances excretion of sodium, chloride, and water by interfering with the transport of sodium ions across the renal tubular epithelium. Enhances urinary excretion of potassium secondary to increased amount of sodium at distal tubular site of sodium–potassium exchange. Increases urinary bicarbonate excretion (although to a lesser extent than chloride excretion) but change in urinary pH is usually minimal (diuretic efficacy not affected by acid–base balance of patient). Increases calcium urinary excretion from a decrease in extracellular fluid volume, although calcium reabsorption in the nephron may be increased (also, slight or intermittent elevations in serum calcium concentration). Hypotensive activity in hypertensive patients (unknown mechanism; potential direct arteriolar dilation). Effect: Mild-to-moderate hypertension. Edema. | Mechanistic genes:ADD1, ADRB1, ADRB2, ACE1, AGT, GNB3, NOS3, SCNN1G, WNK1 Metabolic genes Substrate:CYP11B2 |
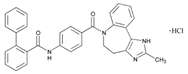 | Name: CONIVAPTAN IUPAC Name: [1,1′-Biphenyl]-2-carboxamide, N-[4-[(4,5-dihydro-2-methylimidazo[4,535enzazepineazepin-6(1H)-yl)carbonyl]phenyl]-, monohydrochloride; 4′’-[(4,5-dihydro-2-methylimidazo[4,535enzazepineazepin-6(1H)-yl)carbonyl]-2-biphenylcarboxanilide monohydrochloride Molecular Formula: C32H26N4O2 HCl Molecular Weight: 535.04 Mechanism: Conivaptan is an arginine vasopressin (AVP) receptor antagonist with affinity for AVP receptor subtypes V1A and V2. The antidiuretic action of AVP is mediated through activation of the V2 receptor, which functions to regulate water and electrolyte balance at the level of the collecting ducts in the kidney. Antagonism of the V2 receptor by conivaptan promotes the excretion of free water (without loss of serum electrolytes) resulting in net fluid loss, increased urine output, decreased urine osmolality, and subsequent restoration of normal serum sodium levels. Blockade of vascular V1A receptors may cause splanchnic vasodilation, and thus hypotension or variceal bleeding in patients with cirrhosis (especially those with portal hypertension). Effect: Euvolemic and hypervolemic hyponatremia in hospitalized patients. | Mechanistic genes:AVPR1A Metabolic genes Substrate:CYP3A4, CYP3A5 Inhibitor:CYP3A4, CYP3A5 |
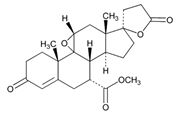 | Name: EPLERENONE IUPAC Name: Pregn-4-ene-7,21-dicarboxylic acid, 9,11-epoxy-17-hydroxy-3-oxo-, γ-lactone, methyl ester, (7α,11α,17α)-; 9,11α-epoxy-17-hydroxy-3-oxo-17α-pregn-4-ene-7α,21-dicarboxylic acid, γ-lactone, methyl ester Molecular Formula: C24H30O6 Molecular Weight: 414.49 Mechanism: A relatively selective competitive mineralocorticoid (aldosterone) receptor antagonist. Binds selectively to mineralocorticoid receptors and has low (less than 1%) affinity for glucocorticoid, progesterone, and androgen receptors. It is a competitive antagonist of aldosterone at mineralocorticoid receptors in the kidney, myocardium, salivary gland, GI tract, brain, and vasculature, and has been shown to inhibit the physiologic effects of aldosterone in these organs. Some of the antihypertensive effects of eplerenone may be related to restoration of endothelial function by increasing the release of nitric oxide, which results in vasodilation. Has been shown to produce sustained increases in plasma renin and serum aldosterone concentrations, reflecting the inhibition of the negative feedback of aldosterone on renin secretion. Appears to have cardioprotective effects in congestive heart failure and left ventricular dysfunction following MI. The cardioprotective action mechanism appears to be related more to the ability of the drug to competitively inhibit the pathophysiologic effects of aldosterone on the myocardium than to its hypotensive effects. Eplerenone reduces coronary vascular inflammation, risk of subsequent development of interstitial myocardial and coronary perivascular fibrosis, cardiac hypertrophy, and/or ventricular remodeling. Effect: To reduce the risk of mortality following acute MI in clinically stable patients with left ventricular dysfunction (i.e., LVEF 40% or less) who have demonstrated clinical evidence of CHF. Used orally in the management of hypertension. May be used as monotherapy or in combination with other classes of antihypertensive agents (e.g., ACEIs, angiotensin II receptor antagonists, calcium-channel blocking agents, β-adrenergic blocking agents, and thiazide diuretics). | Mechanistic genes:NOS3, NPPA, NR3C2 Metabolic genes Substrate:CYP3A4, CYP3A5, CYP11B2 |
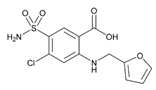 | Name: FUROSEMIDE IUPAC Name: Benzoic acid, 5-(aminosulfonyl)-4-chloro-2-[(2-furanylmethyl)amino]-; 4-chloro-N-furfuryl-5-sulfamoylanthranilic acid Molecular Formula: C12H11ClN2O5S Molecular Weight: 330.74 Mechanism: Inhibits reabsorption of sodium and chloride in ascending loop of Henle and distal renal tubule, interfering with chloride-binding cotransport system, thus causing increased excretion of water, sodium, chloride, magnesium, and calcium. Effect: Management of edema associated with CHF, nephrotic syndrome, and hepatic cirrhosis. I.V. furosemide may also be used as an adjunct in treatment of acute pulmonary edema. | Mechanistic genes:COL1A1, FOS, GABRA6, IFNA1, IGF1, KDR, LTA, MMP2, NOS3, PDGFRA, PDGFRB, PTGER4, PTGS1, PTGS2, REN, SCN1B, SCNN1B, SCNN1G, TGFB1, TNFRSF1A, TNFRSF1B, VCAM1, VEGFA Metabolic genes Substrate:UGT1A1, UGT1A3, UGT1A7, UGT1A10 Transporter genes:ABCC2, ABCC3, ABCC4, SLC12A1, SLC12A3, SLC22A6, SLC22A7 Pleiotropic genes:IL6, IL10, TNF |
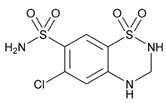 | Name: HYDROCHLOROTHIAZIDE IUPAC Name: 2H-1,2,4-benzothiadiazine-7-sulfonamide, 6-chloro-3,4-dihydro-, 1,1-dioxide; 6-chloro-3,4-dihydro-2H-1,2,4-benzothiadiazine-7-sulfonamide 1,1-dioxide Molecular Formula: C7H8ClN3O4S2 Molecular Weight: 297.74 Mechanism: Inhibits sodium reabsorption in distal tubules causing increased excretion of sodium and water as well as potassium and hydrogen ions. Effect: Management of mild-to-moderate hypertension. Treatment of edema in congestive heart failure and nephrotic syndrome. | Mechanistic genes:ACE1, ACE2, ADD1, ADRB1, ADRB2, AGT, GNB3, GRIA3, NOS3, PTGS2, REN, SCNN1G, WNK1 Metabolic genes Substrate:CYP11B2 Transporter genes:ABCC4, SLC22A6 |
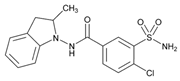 | Name: INDAPAMIDE IUPAC Name: Benzamide, 3-(aminosulfonyl)-4-chloro-N-(2,3-dihydro-2-methyl-1H-indol-1-yl)-; 4-chloro-N-(2-methyl-1-indolinyl)-3-sulfamoylbenzamide Molecular Formula: C16H16ClN3O3S Molecular Weight: 365.83 Mechanism: Diuretic effect localized at proximal segment of distal tubule of nephron. Enhances sodium, chloride, and water excretion by interfering with transport of sodium ions across renal tubular epithelium. Effect: Management of hypertension, or of edema associated with CHF and nephrotic syndrome. | Mechanistic genes:KCNE1, KCNQ1 Metabolic genes Substrate:CYP3A4, CYP3A5 |
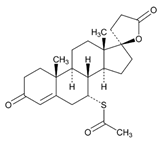 | Name: SPIRONOLACTONE IUPAC Name: Pregn-4-ene-21-carboxylic acid, 7-(acetylthio)-17-hydroxy-3-oxo-, γ-lactone, (7α,17α)-; 17-hydroxy-7α-mercapto-3-oxo-17α-pregn-4-ene-21-carboxylic acid, γ-lactone acetate Molecular Formula: C24H32O4S Molecular Weight: 416.57 Mechanism: Synthetic steroid mineralocorticoid receptor antagonist (aldosterone antagonist). Exhibits magnesium- and potassium-sparing, natriuretic, diuretic, and hypotensive effects by competitively inhibiting physiologic effects of adrenocorticortical hormone aldosterone on distal renal tubules, myocardium, and vasculature. Does not generally cause potassium depletion or affect glucose metabolism or uric acid excretion. Androgen and progesterone receptor antagonist. Effect: Edema associated with excessive aldosterone excretion. Hypertension. Primary hyperaldosteronism. Hypokalemia. Cirrhosis accompanied by edema or ascites. Nephritic syndrome. Severe heart failure. | Mechanistic genes:ACE1, AR, NR3C2, SCNN1G Metabolic genes Substrate:CYP2C8, CYP3A4, CYP3A5, CYP7A1, UGT1A1, UGT1A6 Inhibitor:CYP11B2 Transporter genes:ABCB1, ABCB11, ABCC2, ABCC3 |
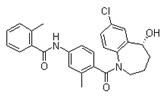 | Name: TOLVAPTAN IUPAC Name: N-[4-(7-chloro-5-hydroxy-2,3,4,5-tetrahydro-1-benzazepine-1-carbonyl)-3-methylphenyl]-2-methylbenzamide; (±)-4′-[(7-chloro-2,3,4,5-tetrahydro-5-hydroxy-1H-1-benzazepin-1-yl)carbonyl]-o-tolu-m-toluidide Molecular Formula: C26H25ClN2O3 Molecular Weight: 448.94 Mechanism: Tolvaptan is a selective vasopressin V2-receptor antagonist with an affinity for the V2 receptor that is 1.8-fold that of native arginine vasopressin (AVP). Tolvaptan affinity for the V2 receptor is 29-fold greater than for the V1a receptor. Antagonism of the V2 receptor by tolvaptan promotes the excretion of free water (without loss of serum electrolytes) resulting in net fluid loss, increased urine output, decreased urine osmolality, and subsequent restoration of normal serum sodium levels. Tolvaptan metabolites have no or weak antagonist activity for human V2 receptors compared with tolvaptan. Effect: Treatment of clinically significant hypervolemic or euvolemic hyponatremia (associated with heart failure, cirrhosis or SIADH) with either a serum sodium <125 mEq/L or less marked hyponatremia that is symptomatic and resistant to fluid restriction. | Mechanistic genes:AVPR2, PKD1, PKD2 Metabolic genes Substrate:ABCB1, CYP3A4, CYP3A5 Inhibitor:ABCB1 |
 | Name: TORSEMIDE IUPAC Name: 3-Pyridinesulfonamide, N-[[(1-methylethyl)amino]carbonyl]-4-[(3-methylphenyl)amino]-; 1-isopropyl-3-[(4-m-toluidino-3-pyridyl)sulfonyl]urea Molecular Formula: C16H20N4O3S Molecular Weight: 348.42 Mechanism: Inhibits reabsorption of sodium and chloride in ascending loop of Henle and distal renal tubule, interfering with chloride-binding cotransport system, thus causing increased excretion of water, sodium, chloride, magnesium, and calcium. Effect: Management of edema associated with congestive heart failure and hepatic or renal disease. Used alone or in combination with antihypertensives in treatment of hypertension. I.V. form indicated when rapid onset is desired. | Mechanistic genes:ADD1, SCNN1G Metabolic genes Substrate:CYP2C8, CYP2C9, CYP11B2 Inhibitor:CYP2C19 Transporter genes:SLC12A1, SLC12A3, SLCO1B1 |
 | Name: TRIAMTERENE IUPAC Name: 2,4,7-Pteridinetriamine, 6-phenyl-; (2) 2,4,7-triamino-6-phenylpteridine Molecular Formula: C12H11N7 Molecular Weight: 253.26 Mechanism: Interferes with potassium/sodium exchange (active transport) in distal tubule, cortical collecting tubule and collecting duct by inhibiting sodium, potassium-ATPase. Decreases calcium excretion. Increases magnesium loss. Effect: Alone or in combination with other diuretics in treatment of edema and hypertension. Decreases potassium excretion caused by kaliuretic diuretics. | Mechanistic genes:SCNN1A, SCNN1B, SCNN1D, SCNN1G Metabolic genes Substrate:CYP1A2, CYP2A6, CYP2D6, CYP3A4, CYP3A5 Inducer:CYP1A2 |
| Lipid-Modifying Agents | ||
|---|---|---|
| Drug | Properties | Pharmacogenetics |
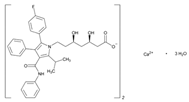 | Name: ATORVASTATIN IUPAC Name: 1H-Pyrrole-1-heptanoic acid, 2-(4-fluorophenyl)-β,δ-dihydroxy-5-(1-methylethyl)-3-phenyl-4-[(phenylamino)carbonyl]-, calcium salt (2:1), [R-(R*,R*)]-; calcium (βR,δR)-2-(p-fluorophenyl)-β,δ-dihydroxy-5-isopropyl-3-phenyl-4-(phenylcarbamoyl)pyrrole-1-heptanoate (1:2) Molecular Formula: C66H68CaF2N4O10 Molecular Weight: 1155.34 Mechanism: Inhibits 3-hydroxy-3-methylglutaryl coenzyme A (HMG-CoA) reductase, resulting in a compensatory increase in the expression of LDL receptors on hepatocyte membranes and a stimulation of LDL catabolism. Effect: Treatment of dyslipidemias or primary prevention of cardiovascular disease (atherosclerotic). | Mechanistic genes: APOA5, APOB, APOC3, ACE1, APOA1, APOE, CETP, CRP, ESR1, FGB, GNB3, HTR3B, ITGB3, LDLR, LIPC, MMP3, MTTP, NOS3, PON1, USP5 Metabolic genes Substrate: CYP2C8, CYP2C9, CYP3A4, CYP3A5, CYP7A1, CYP11B2, UGT1A1, UGT1A3 Inhibitor: ABCB1, CYP2B6, CYP2C9, CYP2C19, CYP2D6, CYP3A4, CYP3A5, HMGCR Inducer: CYP2B6 Transporter genes: ABCA1, ABCB1, ABCB11, ABCC1, ABCC2, ABCC3, ABCG2, SLC16As, SLCO1B1, SLCO1B3 Pleiotropic genes: IL6, IL10, TNF |
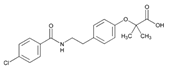 | Name: BEZAFIBRATE IUPAC Name: Propanoic acid, 2-[4-[2-[(4-chlorobenzoyl)amino]ethyl]phenoxy]-2-methyl-; 2-[p-[2-(p-chlorobenzamido)ethyl]phenoxy]-2-methylpropionic acid Molecular Formula: C19H20ClNO4 Molecular Weight: 361.82 Mechanism: Mechanism not established. May increase VLDL catabolism by increasing lipoprotein and hepatic triglyceride lipase activities. May decrease triglyceride biosynthesis by inhibition of acetyl-CoA carboxylase. May decrease cholesterol biosynthesis by inhibition of 3-hydroxy-3-methyglutaryl-coenzyme A reductase. Effect: Adjunct to diet and other therapeutic measures for treatment of type IIa and IIb mixed hyperlipidemia, to regulate lipid and apoprotein levels (reduce serum triglycerides, LDL-C and apolipoprotein B, increase HDL-C and apolipoprotein A). Treatment of adult patients with high to very high triglyceride levels (hypertriglyceridemia)(Fredrickson classification type IV and V hyperlipidemias) who are at high risk of complications from their dyslipidemia. | Mechanistic genes:ACSL1, ACOX1, APOA5, APOB, APOC3, APOE, CETP, LDLR, LIPC, MGMT, PPARA, SCARB1 Metabolic genes Substrate:CYP1A1, CYP3A4, CYP3A5, CYP7A1, UGT1A1 Inhibitor:ACACA, CYP2C8, HMGCR Inducer:CYP3A4, CYP3A5, UGT1A1 Transporter genes:ABCB11 |
| Name: COLESTIPOL IUPAC Name: Colestipol hydrochloride; copolymer of diethylenetriamine and 1-chloro-2,3-epoxypropane, hydrochloride Molecular Formula: N.A. Molecular Weight: N.A. Mechanism: Binds to bile acids in the intestine and forms a non-absorbable complex. Thus, bile acids are partially removed from enterohepatic circulation and conversion of cholesterol to bile acids in the liver is increased. This enhanced demand for cholesterol in liver cells causes a compensatory increase in hepatic uptake (and thus systemic clearance) of circulating LDL-C. Serum triglyceride concentrations may remain unchanged or increase slightly (5–10%). Antilipemic effects are additive when used with lovastatin or niacin. Effect: Primary hypercholesterolemia. Arteriolosclerosis. Pruritus associated with elevated levels of bile acids. To decrease plasma half-life of digoxin in toxicity. | Mechanistic genes:APOE, LIPC Metabolic genes Substrate:CYP7A1 | |
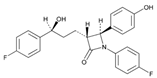 | Name: EZETIMIBE IUPAC Name: 2-Azetidinone, 1-(4-fluorophenyl)-3-[3-(4-fluorophenyl)-3-hydroxypropyl]-4-(4-hydroxyphenyl)-, [3R-[3α(S*),4β]]-; (2)(3R,4S)-1-(p-fluorophenyl)-3-[(3S)-3-(p-fluorophenyl)-3-hydroxypropyl]-4-(p-hydroxyphenyl)-2-azetidinone Molecular Formula: C24H21F2NO3 Molecular Weight: 409.43 Mechanism: Inhibits absorption of cholesterol at the brush border of the small intestine via the sterol transporter, Niemann–Pick C1-Like 1 (NPC1L1). This leads to a decreased delivery of cholesterol to the liver, reduction in hepatic cholesterol stores and increased clearance of cholesterol from the blood. Decreases total cholesterol, LDL-C, APOB, and triglycerides while increasing HDL-C. Effect: Used in combination with dietary therapy for treatment of primary hypercholesterolemia (as monotherapy or in combination with HMG-CoA reductase inhibitors). Homozygous sitosterolemia. Homozygous familial hypercholesterolemia (in combination with atorvastatin or simvastatin). Mixed hyperlipidemia (in combination with fenofibrate). | Mechanistic genes:APOE, APOA1, APOA5, APOB, APOC3, CETP, NPC1, NPC1L1, SCARB1 Metabolic genes Substrate:CYP7A1, UGT1A1, UGT1A3, UGT2B7, UGT2B15 Transporter genes:ABCB1, ABCA1, ABCC2, SLCO1B1, SLCO1B3 |
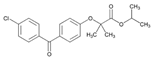 | Name: FENOFIBRATE IUPAC Name: Isopropyl 2-[p-(p-chlorobenzoyl)phenoxy]-2-methylpropionate Molecular Formula: C20H21ClO4 Molecular Weight: 360.83 Mechanism: Fenofibric acid is believed to increase VLDL catabolism by enhancing the synthesis of lipoprotein lipase. As a result of a decrease in VLDL levels, total plasma triglycerides are reduced by 30%–60%. Modest increase in HDL occurs in some hypertriglyceridemic patients. Effect: Adjunct to dietary therapy to decrease elevated serum total and LDL-C, triglyceride, and apolipoprotein B (APOB) concentrations, and to increase HDL-C concentrations in the management of primary hypercholesterolemia and mixed dyslipidemia, including heterozygous familial hypercholesterolemia and other causes of hypercholesterolemia. Additive antilipemic effects when used concomitantly with other antilipemic agents (e.g., colesevelam and ezetimibe). Adjunct to dietary therapy in management of hypertriglyceridemia. | Mechanistic genes:ACE1, ACOX1, APOA1, APOA5, APOB, APOE, FABP1, FABP2, LPL, PPARA, SCARB1 Metabolic genes Substrate:CYP2C8, CYP3A4, CYP3A5, UGT1A1 Inhibitor:CYP2A6, CYP2C8, CYP2C9, CYP2C19, CYP4F2 Inducer:CYP2C8, CYP3A4, CYP3A5, UGT1A1 Transporter genes:ABCA1, ABCB1 Pleiotropic genes:APP, IL6 |
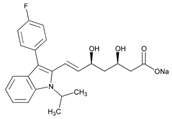 | Name: FLUVASTATIN IUPAC Name: 6-Heptenoic acid, 7-[3-(4-fluorophenyl)-1-(1-methylethyl)-1H-indol-2-yl]-3,5-dihydroxy-, monosodium salt, [R*,S*-(E)]-(±)-; sodium (±)-(3R*,5S*,6E)-7-[3-(p-fluorophenyl)-1-isopropylindol-2-yl]-3,5-dihydroxy-6-heptenoate Molecular Formula: C24H25FNNaO4 Molecular Weight: 433.45 Mechanism: Acts by competitively inhibiting 3-hydroxyl-3-methylglutaryl-coenzyme A (HMG-CoA) reductase (HMGCR), the enzyme that catalyzes reduction in HMG-CoA to mevalonate. HDL is increased while total, LDL and VLDL cholesterols, apolipoprotein B, and plasma triglycerides are decreased. Effect: Used as a component of multiple risk factor intervention in patients at risk for atherosclerosis vascular disease due to hypercholesterolemia. | Mechanistic genes:ACE1, APOA1, APOA5, APOB, APOC3, APOE, CETP, LDLR, LIPC, LPL, NOS3, NR1I2, NR1I3, PPARD, PON1, USP5 Metabolic genes Substrate:CYP1A1, CYP2B6, CYP2C8, CYP2C9, CYP2D6, CYP3A4, CYP3A5, CYP7A1, UGT1A3 Inhibitor:CYP1A2, CYP2C8, CYP2C9, CYP2C19, CYP2D6, CYP3A4, CYP3A5, HMGCR Transporter genes:ABCA1, ABCB1, ABCB11, ABCC2, ABCG2, SLC15A1, SLC22A8, SLCO1B1, SLCO1B3, SLCO2B1 |
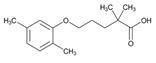 | Name: GEMFIBROZIL IUPAC Name: Pentanoic acid, 5-(2,5-dimethylphenoxy)-2,2-dimethyl-; 2,2-dimethyl-5-(2,5-xylyloxy)valeric acid Molecular Formula: C15H22O3 Molecular Weight: 250.33 Mechanism: Gemfibrozil can inhibit lipolysis and decrease subsequent hepatic fatty acid uptake as well as inhibit hepatic secretion of VLDL. Together, these actions decrease serum VLDL levels. Increases HDL-C. Effect: Gemfibrozil is used to reduce risk of developing coronary heart disease (CHD) in patients with type IIb hyperlipoproteinemia without clinical evidence of CHD (primary prevention) who have inadequate response to dietary management, weight loss, exercise, and drugs known to reduce LDL-C and increase HDL-C and who have low HDL-C concentrations in addition to elevated LDL-C and triglycerides. In addition, gemfibrozil is used for treatment of hypertriglyceridemia in types IV and V hyperlipidemia for patients at greater risk for pancreatitis and who have not responded to dietary intervention. | Mechanistic genes:ACE1, ACOX1, AHR, APOE, CES2, CETP, CFTR, LPL, MMP3, NR1I2, PPARA, SCARB1 Metabolic genes Substrate:CYP2D6, CYP2C8, CYP3A4, CYP3A5, CYP7A1, UGT1A1, UGT1A3 Inhibitor:CYP1A2, CYP2C8, CYP2C9, CYP2C19, CYP3A4, CYP3A5 Inducer:CYP2C8, UGT1A1 Transporter genes:ABCB1, SLCO1B1, SLCO1B2 Pleiotropic genes:IL1B, IL12B, TNF |
 | Name: LOVASTATIN IUPAC Name: Butanoic acid, 2-methyl-, 1,2,3,7,8,8a-hexahydro-3,7-dimethyl-8-[2-(tetrahydro-4-hydroxy-6-oxo-2H-pyran-2-yl)ethyl]-1-naphthalenyl ester, [1S-[1α(R*),3α,7β,8β(2S*,4S*),8aβ]]-; (S)-2-methylbutyric acid, 8-ester with (4R,6R)-6-[2-[(1S,2S,6R,8S,8aR)-1,2,6,7,8,8a-hexahydro-8-hydroxy-2,6-dimethyl-1-naphthyl]ethyl]tetrahydro-4-hydroxy-2H-pyran-2-one Molecular Formula: C24H36O5 Molecular Weight: 404.54 Mechanism: Acts by competitively inhibiting 3-hydroxyl-3-methylglutaryl-coenzyme A (HMG-CoA) reductase, enzyme which catalyzes rate-limiting step in cholesterol biosynthesis. Effect: Adjunct to dietary therapy to decrease elevated serum total and LDL-C concentrations in primary hypercholesterolemia. Primary prevention of coronary artery disease (patients without symptomatic disease with average to moderately elevated total and LDL-C and below average HDL-C). Slow progression of coronary atherosclerosis in coronary heart disease. Adjunct to dietary therapy in adolescent patients (10–17 years of age, females >1 year postmenarche) with heterozygous familial hypercholesterolemia having LDL>189 mg/dL, or LDL>160 mg/dL with positive family history of premature cardiovascular disease, or LDL>160 mg/dL with presence of at least two other CVD risk factors. | Mechanistic genes:APOA1, APOA5, APOB, APOC3, CETP, KCNH2, LDLR, LIPC, LPL, USP5 Metabolic genes Substrate:CYP2C8, CYP3A4, CYP3A5, UGT1A3 Inhibitor:ABCB1, CYP2C9, CYP2C19, CYP2D6, CYP3A4, CYP3A5, HMGCR Inducer:CYP2B6, CYP7A1 Transporter genes:ABCA1, ABCB1, ABCB11, ABCC2, ABCG2, SLCO1B1, SLCO1B3 Pleiotropic genes:TP53 |
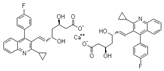 | Name: PITAVASTATIN IUPAC Name: Calcium (E,3R,5S)-7-[2-cyclopropyl-4-(4-fluorophenyl)quinolin-3-yl]-3,5-dihydroxyhept-6-enoate; (+)monocalcium bis{(3R, 5S, 6E)-7-[2-cyclopropyl-4-(4-fluorophenyl)-3-quinolyl]-3,5-dihydroxy-6-heptenoate} Molecular Formula: C50H46CaF2N2O8 Molecular Weight: 880.98 Mechanism: Pitavastatin competitively inhibits HMG-CoA reductase, which is a rate-determining enzyme involved with biosynthesis of cholesterol, in a manner of competition with the substrate so that it inhibits cholesterol synthesis in the liver. As a result, the expression of LDL receptors followed by the uptake of LDL from blood to liver is accelerated and then the plasma total cholesterol (TC) decreases. Furthermore, sustained inhibition of cholesterol synthesis in the liver decreases levels of very low-density lipoproteins. Effect: Adjunct to dietary therapy to reduce elevations in TC, LDL-C, apolipoprotein B (Apo B), and triglycerides (TG), and to increase low HDL-C in patients with primary hyperlipidemia and mixed dyslipidemia. | Metabolic genes Substrate:CYP2C8, CYP2C9, CYP3A4, CYP3A5, UGT1A3, UGT2B7 Inhibitor:HMGCR Transporter genes:ABCB1, ABCG2, SLCO1B1 |
 | Name: PRAVASTATIN IUPAC Name: 1-Naphthaleneheptanoic acid, 1,2,6,7,8,8a-hexahydro-β,δ,6-trihydroxy-2-methyl-8-(2-methyl-1-oxobutoxy)-, monosodium salt, [1S-[1α(βS*,δS*),2α,6α,8β(R*),8aα]]-; sodium (+)-(βR,δR,1S,2S,6S,8S,8aR)-1,2,6,7,8,8a-hexahydro-β,δ,6,8-tetrahydroxy-2-methyl-1-naphthaleneheptanoate, 8-[(2S)-2-methylbutyrate] Molecular Formula: C23H35NaO7 Molecular Weight: 446.51 Mechanism: A competitive inhibitor of 3-hydroxy-3-methylglutaryl coenzyme A (HMG-CoA) reductase. Effect: Primary prevention of coronary events (reduction in cardiovascular morbidity (myocardial infarction, coronary revascularization procedures) and mortality). Secondary prevention of cardiovascular events in established coronary heart disease (slowing of progression of coronary atherosclerosis, reduction in cardiovascular morbidity (myocardial infarction, coronary vascular procedures) and reduction in mortality, reduction in risk of stroke and transient ischemic attacks). Hyperlipidemias (reduction in elevations in total cholesterol, LDL-C, apolipoprotein B, and triglycerides). Heterozygous familial hypercholesterolemia. | Mechanistic genes:ACE1, ALDH1A1, APOA1, APOA5, APOB, APOC3, APOE, CBS, CETP, CRP, FGB, HMGCR, HTR3B, ITGB3, LDLR, LEP, LIPC, LPL, MMP2, MMP3, MTHFR, NOS3, PON1, USP5 Metabolic genes Substrate:CYP1A1, CYP1A2, CYP2C8, CYP2E1, CYP3A4, CYP3A5, CYP7A1, UGT1A3 Inhibitor:ABCB1, CYP2C9, CYP2C19, CYP2D6, CYP3A4, CYP3A5, HMGCR Inducer:CYP2B6 Transporter genes:ABCA1, ABCB1, ABCB11, ABCC2, ABCG2, SLC22A8, SLCO1A2, SLCO1B1, SLCO1B3, SLCO2B1 Pleiotropic genes:IL1B, IL6, IL10, TP53 |
 | Name: PROBUCOL IUPAC Name: Phenol, 4,4′-[(1-methylethylidene)bis(thio)]bis[2,6-bis(1,1-dimethylethyl)-; acetone bis(3,5-di-tert-butyl-4-hydroxyphenyl) mercaptole Molecular Formula: C31H48O2S2 Molecular Weight: 516.84 Mechanism: Increases fecal loss of bile acid-bound low-density lipoprotein cholesterol, decreases synthesis of cholesterol and inhibits enteral cholesterol absorption. Effect: Adjunct to dietary therapy to decrease elevated serum total and LDL-C concentrations in primary hypercholesterolemia. | Mechanistic genes:APOB, APOC3, KCNH2, SCARB1 Metabolic genes Substrate:CYP2D6, CYP3A4, CYP3A5 Pleiotropic genes:TNF |
 | Name: ROSUVASTATIN IUPAC Name: 6-Heptenoic acid, 7-[4-(4-fluorophenyl)-6-(1-methylethyl)-2-[methyl(methylsulfonyl)amino]-5-pyrimidinyl]-3,5-dihydroxy-, calcium salt (2:1), (3R,5S,6E)-; [S-[R*,S*-(E)]]-7-[4-(4-fluorophenyl)-6-(1-methylethyl)-2-[methyl(methylsulfonyl)amino]-5-pyrimidinyl]-3,5-dihydroxy-6-heptenoic acid, calcium salt (2:1) Molecular Formula: [C22H27FN3O6S]2Ca Molecular Weight: 1001.14 Mechanism: Inhibitor of 3-hydroxy-3-methylglutaryl coenzyme A (HMG-CoA) reductase, rate-limiting enzyme in cholesterol synthesis. This results in compensatory increase in expression of LDL receptors on hepatocyte membranes and stimulation of LDL catabolism. Effect: Used with dietary therapy for hyperlipidemias to reduce elevations in total cholesterol (TC), LDL-C, apolipoprotein B, non-HDL-C, and triglycerides in primary hypercholesterolemia (elevations of 1 or more components present in Fredrickson type IIa, IIb, and IV hyperlipidemias). Treatment of primary dysbetalipoproteinemia (Fredrickson type III hyperlipidemia). Treatment of homozygous familial hypercholesterolemia. To slow progression of atherosclerosis as adjunct to TC- and LDL-C-lowering diet. | Mechanistic genes:ACE1, APOA1, APOA5, APOB, APOC3, APOE, CETP, FGB, ITGB3, LDLR, LIPC, LPL, NOS3, TCF20, USP5 Metabolic genes Substrate:CYP2C8, CYP2C9, CYP2C19, CYP2D6, CYP3A4, CYP3A5, CYP7A1, UGT1A3 Inhibitor:CETP, HMGCR, SLCO1B1 Inducer:CYP2B6, CYP2C9, CYP3A4, CYP3A5 Transporter genes:ABCA1, ABCB1, ABCB11, ABCC1, ABCC4, ABCG2, SLC10A1, SLCO1A2, SLCO1B1, SLCO1B3, SLCO2B1, SLC22A8 |
 | Name: SIMVASTATIN IUPAC Name: Butanoic acid, 2,2-dimethyl-, 1,2,3,7,8,8a-hexahydro-3,7-dimethyl-8-[2-(tetrahydro-4-hydroxy-6-oxo-2H-pyran-2-yl)ethyl]-1-naphthalenyl ester, [1S-[1α,3α,7β,8β(2S*,4S*),8aβ]]-; 2,2-dimethylbutyric acid, 8-ester with (4R,6R)-6-[2-[(1S,2S,6R,8S,8aR)-1,2,6,7,8,8a-hexahydro-8-hydroxy-2,6-dimethyl-1-naphthyl]ethyl]tetrahydro-4-hydroxy-2H-pyran-2-one Molecular Formula: C25H38O5 Molecular Weight: 418.57 Mechanism: Prodrug requiring hydrolysis in vivo for activity. Inhibits HMG-CoA reductase, causing subsequent reduction in hepatic cholesterol synthesis. Reduces serum concentrations of total cholesterol, LDL-C, Apo B, and triglycerides. Statins may slow progression and/or induce regression of atherosclerosis in coronary and/or carotid arteries, modulate blood pressure in hypercholesterolemic patients with hypertension, and possess anti-inflammatory activity. Effect: Secondary prevention of cardiovascular events in hypercholesterolemic patients with established coronary heart disease or at high risk for coronary heart disease. Hyperlipidemias (primary hypercholesterolemia, homozygous familial hypercholesterolemia, heterozygous familial hypercholesterolemia). | Mechanistic genes:APOA1, APOA5, APOB, APOC3, APOE, CETP, F2, FGB, GNB3, HMGCR, HTR3B, LDLR, LIPC, LPL, NOS3, PRNP, USP5, VCAM1 Metabolic genes Substrate:CYP2C8, CYP2C9, CYP2C19, CYP3A4, CYP3A5, CYP7A1, POR, UGT1A3 Inhibitor:CYP2C8, CYP2C9, CYP2C19, CYP2D6, CYP3A4, CYP3A5, HMGCR Inducer:CYP2B6 Transporter genes:ABCA1, ABCB1, ABCB11, ABCC2, ABCC3, ABCG2, SLCO1B1, SLCO1B3 Pleiotropic genes:IL6, TNF |
| Antithrombotic Drugs | ||
|---|---|---|
| Vitamin K Antagonists | ||
| Drug | Properties | Pharmacogenetics |
 | Name: Dicoumarol IUPAC Name: 4-hydroxy-3-[(4-hydroxy-2-oxo-2H-chromen-3-yl)methyl]-2H-chromen-2-one. Molecular Formula: C19H12O6 Molecular Weight: 336.295 Da. Mechanism: Inhibits vitamin K reductase, depletes vitamin KH2, cofactor for vitamin K-dependent protein-carboxylation, limits gamma-carboxylation, and activates vitamin K-dependent coagulant proteins. Inhibits synthesis of vitamin K-dependent coagulation factors II, VII, IX, and X and anticoagulant proteins C and S. Depresses vitamin K-dependent coagulation factors II, VII, and X, lowers prothrombin levels and the amount of fibrin-bound thrombin, reducing thrombogenicity. Effect: Antithrombotic agents. Vitamin K antagonists. | Mechanistic genes:CRYZ, F2, F7, F9, F10, NQO1, PROC, PROS1, VKORC1 Metabolic genes Substrate:CYP2C9 Inhibitor:CYP2C6, CYP2C11 Transporter genes:ALB |
 | Name: Phenindione IUPAC Name: 2-phenyl-2,3-dihydro-1H-indene-1,3-dione. Molecular Formula: C15H10O2 Molecular Weight: 222.24 Da. Mechanism: Similar mode of action as Dicoumarol. Effect: Antithrombotic agents. Vitamin K antagonists. | Mechanistic genes:ANXA5, F2, F7, F9, F10, PROC, PROS1, VKORC1 |
 | Name:Warfarin IUPAC Name, (1) 2H-1-benzopyran-2-one, 4-hydroxy-3-(3-oxo-1-phenylbutyl)-, sodium salt, (2) 3-(α-acetonylbenzyl)-4-hydroxycoumarin sodium salt Molecular Formula: C19H15NaO4 Molecular Weight: 330.31 Da Mechanism: Competitively inhibits subunit-1 of multi-unit VKOR complex, depleting vitamin K reserves. Antithrombogenic effects occur after functional coagulation factors IX and X are diminished. Phytonadione (vitamin K1) reverses anticoagulant effect. Slightly affects platelet-rich arterial thrombi-adherence to abnormal vessel wall. Effect: Antithrombotic agents, Anticoagulants, Coumarin Derivatives. Vitamin K Antagonist | Mechanistic genes: F2, F5, F7, F9, F10, NR1I2, PROC, VKORC1 Metabolic genes Substrate: CALU, CYP1A2, CYP2C8, CYP2C9, CYP2C18, CYP2C19, CYP3A4, CYP3A5, EPHX1, GGCX Inhibitor: CYP2C9, CYP2C19, VKORC1 Inducer: CYP2C9, CYP3A4 Transporter genes: ABCB1, ALB, ORM1 Pleiotropic genes: APOE |
 | Name: Phenprocoumon IUPAC Name: 4-hydroxy-3-(1-phenylpropyl)-2H-chromen-2-one. Molecular Formula: C18H16O3 Molecular Weight: 280.32 Da. Mechanism: as per Dicoumarol Effect: Antithrombotic agents. Vitamin K antagonists. | Mechanistic genes:F2, F7, F9, F10, PROC, PROS1, VKORC1 Metabolic genes Substrate:CYP2C8, CYP2C9, CYP3A4 Transporter genes:ALB, ORM1 |
 | Name:Acenocoumarol IUPAC Name, (1) 2H-1-benzopyran-2-one, 4-hydroxy-3-[1-(4-nitrophenyl)-3-oxobutyl]-, (2) 3-(α-acetonyl-p-nitrobenzyl)-4-hydroxycoumarin Molecular Formula: C19H15NO6 Molecular Weight: 353.33 Da Mechanism: Interferes with hepatic synthesis of vitamin K-dependent coagulation factors II, VII, IX, X. Effect: Antithrombotic agents. Vitamin K antagonists. | Mechanistic genes:CALU, F2, F7, F9, F10, VKORC1 Metabolic genes Substrate:CYP1A2, CYP2C9, CYP2C18, CYP2C19, CYP3A4 Transporter genes:ABCB1, ALB, ORM Pleiotropic gens: APOE |
 | Name: Ethyl biscoumacetate IUPAC Name: ethyl 2,2-bis(4-hydroxy-2-oxochromen-3-yl)acetate. Molecular Formula: C22H16O8 Molecular Weight: 408.4 Da. Mechanism: Anticoagulant, mode of action similar to that of warfarin. Effect: Antithrombotic agents. Vitamin K antagonists. | Mechanistic genes:F2, F7, F9, F10, VKORC1 Metabolic genes Substrate:CYP3A4 Inhibitor:GLUL |
| Heparins | ||
 | Name: Heparin IUPAC Name: (2S,3S,4R,5R,6R)-3-({(2R,3R,4R,5S,6R)-3-(acetylamino)-4,5-dihydroxy-6-[(sulfooxy)methyl]tetrahydro-2H-pyran-2-yl}oxy)-6-{[(2S,3S,4S,5R,6S)-6-{[(2R,3S,4S,5R)-2-carboxy-4,6-dihydroxy-5-(sulfooxy)tetrahydro-2H-pyran-3-yl]oxy}-2-hydroxy-4-(sulfomethyl)-5-(sulfooxy)tetrahydro-2H-pyran-3-yl]oxy}-4,5-dihydroxytetrahydro-2H-pyran-2-carboxylic acid. Molecular Formula: C26H41NO34S4. Molecular Weight: 1039.85 Da Mechanism: Potentiates antithrombin III activity; inactivates thrombin, coagulation factors IX, X, XI, XII, and plasmin; prevents conversion of fibrinogen to fibrin. Effect: Antithrombotic agents. Heparin group. | Mechanistic genes:F9, F10, F11, F12, FCGR2A, FCGR3A, FGF1, FGF19, FGF2, FGF4, FGFR1, FGFR2, FGFR4, HGF, ITGB3, LIPA, PF4, PROC, SELP, SERPINA5, SERPINC1, VWF Metabolic genes Substrate:HPSE Transporter genes:ABCC1, SERPINA7 Pleiotropic genes:APP |
 | Name: Enoxaparin Mechanism: Enhances the inhibition rate of clotting proteases by antithrombin III, impairing normal hemostasis; strongly inhibits factor Xa. Effect: Antithrombotic agents. Heparin group. | Mechanistic genes:ACE, F2, F5, F10, FCGR3A, IL1RN, ITGB3, MPO, SERPINC1, THBD Metabolic genes Substrate:HPSE Transporter genes:SERPINA7 |
| Platelet aggregation inhibitors | ||
 | Name:Clopidogrel IUPAC Name: (1) Thieno[3,2-c]pyridine-5(4H)-acetic acid, α-(2-chlorophenyl)-6,7-dihydro-, methyl ester, (S)-, sulfate (1:1), (2) Methyl (+)-(S)-α-(o-chlorophenyl)-6,7-dihydrothieno[3,2-c]pyridine-5(4H)-acetate, sulfate (1:1) Molecular Formula: C16H16ClNO2S Molecular Weight: 419.90 Da Mechanism: Platelet inhibitor; irreversibly binds to P2Y12 ADP receptors on platelets preventing ADP binding to same receptors, activating the glycoprotein GPIIb/IIIa complex, and reducing platelet aggregation. Inhibits ADP-mediated release of platelet dense granule (e.g., ADP, Ca2+, and serotonin) and α-granule (e.g., fibrinogen and thrombospondin) contents that augment platelet aggregation. Effect: Antithrombotic agents. Platelet aggregation inhibitors excl. heparin. | Mechanistic genes:ITGA2B, ITGB3, P2RY12 Metabolic genes Substrate:CES1, CYP1A2, CYP2B6, CYP2C9, CYP2C19, CYP3A4, CYP3A5 Inhibitor:CYP2B6, CYP2C8, CYP2C9, CYP2C19 Transporter genes:ABCB1, SLC22A1, SLC22A2 |
 | Name:Ticlopidine IUPAC Name, (1) Thieno[3,2-c]pyridine, 5-[(2-chlorophenyl)methyl]-4,5,6,7-tetrahydro-, hydrochloride, (2) 5-(o-chlorobenzyl)-4,5,6,7-tetrahydrothieno-[3,2-c]pyridine hydrochloride Molecular Formula: C14H14ClNS Molecular Weight: 300.25 Da Mechanism: Irreversibly blocks P2Y12 receptors as an active metabolite, preventing GPIIb/IIIa receptor complex activation, reducing platelet aggregation. Effect: Antithrombotic agents. Platelet aggregation inhibitors excl. heparin. | Mechanistic genes:ITGB3, P2RY12 Metabolic genes Substrate:CYP2B6, CYP2C19, CYP2D6, CYP3A4, MPO Inhibitor:CYP1A2, CYP2B6, CYP2C8, CYP2C9, CYP2C19, CYP2D6, CYP2E1, CYP3A4 Pleiotropic genes:HLA-B |
 | Name:Acetylsalicylic acid IUPAC Name: Benzoic acid, 2-(acetyloxy)-, (2) salicylic acid acetate Molecular Formula: C9H8O4 Molecular Weight: 180.16 Da Mechanism: Inhibits prostaglandin synthesis; acts on the preoptic area of the anterior hypothalamus to reduce fever. Blocks prostaglandin synthetase activity, preventing thromboxane A2 formation. Effect: Antithrombotic agents. Platelet aggregation inhibitors excl. heparin. | Mechanistic genes:AKR1C1, CASP1, CASP3, CCNA2, CCND1, EDNRA, GP1BA, GP6, HSPA5, IKBKB, MAP2K4; MYC, NFKBIA, PCNA, PRKAs, PTGER1, PTGER2, PTGER3, PTGER4, PTGES, PTGIR, PTGS1, PTGS2, RPS6KA3, TBX21, TBXA2R, TNFAIP6, TP53 Metabolic genes Substrate:ACSM1, CYP2C9, CYP3A4, GLYAT, NAT2, UGT1A1, UGT1A10, UGT1A3, UGT1A6, UGT1A7, UGT1A9, UGT2B4, UGT2B7 Inhibitor:CYP19A1, PTGS1, PTGS2 Inducer:CYP2C19, CYP2E1 Transporter genes, ABCB1, SLC22A6, SLC22A8 |
 | Name:Dipyridamole IUPAC Name: Ethanol, 2,2′,2′’,2′’’-[(4,8-di-1-piperidinylpyrimido[5,4-d]pyrimidine-2,6-diyl)dinitrilo]tetrakis-, (2) 2,2′,2′’,2′’-[(4,8-sipiperidinopyrimido[5,4-d]pyrimidine-2,6-diyl)dinitrilo]tetraethanol Molecular Formula: C24H40N8O4 Molecular Weight: 504.63 Da Mechanism: Non-nitrate coronary vasodilator; inhibits adenosine deaminase and phosphodiesterase activity, inducing accumulation of adenosine, adenine nucleotides, and cAMP which inhibit platelet aggregation, causing vasodilation. May stimulate prostacyclin or PGD2 activity. Effect: Antithrombotic agents. Platelet aggregation inhibitors excl. heparin. | Mechanistic genes:ADA, ORM1, PDE10A, PDE4A, PDE5A, PTGDR2, RCAN1 Transporter genes:ABCB1, ABCB11, ABCC4, ABCC5, SLCO1B1, SLCO1B3, SLCO2B1 |
 | Name:Abciximab IUPAC Name: (1) Immunoglobulin G1, anti-(human integrin αIIbβ3) Fab fragment (human-mouse monoclonal c7E3 clone p7E3VHhCγ1 γ1-chain), disulfide with human-mouse monoclonal c7E3 clone p7E3VκhCκ κ-chain. Molecular Formula: C22H27ClN4O3S Molecular Weight: 462.993 Da Mechanism: Monoclonal antiglycoprotein IIb/IIIa receptor antibody; GPIIb/IIIa is the major platelet surface receptor in platelet aggregation. Blocks vitronectin receptor-mediated cell adhesion, and Mac-1 receptor on monocytes and neutrophils, inhibiting adhesion to monocyte cells. Effect: Antithrombotic agents. Platelet aggregation inhibitors excl. heparin. Glycoprotein IIb/IIIa Inhibitor. | Mechanistic genes:FCGR2A, FCGR2B, ITGA2B, ITGB3, P2RY1, VTN Metabolic genes Substrate:CYP1A2, CYP2C19 |
 | Name:Eptifibatide IUPAC Name: (1) N6-amidino-N2-(3-mercaptopropionyl)-L-lysylglycyl-L-α-aspartyl-L-tryptophyl-L-prolyl-L-cysteinamide, cyclic (1–6)-disulfide. Molecular Formula: C35H49N11O9S2 Molecular Weight: 831.96 Da Mechanism: Blocks the platelet GPIIb/IIIa receptor, reversibly inhibiting platelet aggregation, preventing thrombosis. Effect: Antithrombotic agents, platelet aggregation inhibitors, and glycoprotein IIb/IIIa Inhibitors. | Mechanistic genes:IL6, ITGA2B, ITGB3, P2RY1, TBXAS1 Metabolic genes Substrate:CYP1A2, CYP2C19 |
 | Name: Triflusal IUPAC Name: 2-(acetyloxy)-4-(trifluoromethyl)benzoic acid. Molecular Formula: C10H7F3O4 Molecular Weight: 248.15 Da Mechanism: irreversible COX-1 inhibitor in platelets; spares the arachidonic acid pathway in endothelial cells; favors nitric oxide production. Effect: Antithrombotic agents. Platelet aggregation inhibitors excl. heparin. | Mechanistic genes:NFKB1, NOS2, PDE10A, PTGS1 Metabolic genes Substrate:CYP2C8 Transporter genes:ALB |
 | Name: Beraprost IUPAC Name: sodium 4-[(2S,3R,4R,6S)-4-hydroxy-3-[(1E,3S)-3-hydroxy-4-methyloct-1-en-6-yn-1-yl]-7-oxatricyclo[6.4.0.0^{2,6}]dodeca-1(8),9,11-trien-9-yl]butanoate. Molecular Formula: C24H29NaO5 Molecular Weight: 420.47 Da Mechanism: Binds prostacyclin membrane receptors, inhibiting Ca2+ release from intracellular stores, promoting vasodilation. Effect: Antithrombotic agents. Platelet aggregation inhibitors excl. heparin. | Mechanistic genes:PTGIR Metabolic genes Substrate:CYP2C8 |
 | Name: Treprostinil IUPAC Name: 2-{[(1R,2R,3aS,9aS)-2-hydroxy-1-[(3S)-3-hydroxyoctyl]-1H,2H,3H,3aH,4H,9H,9aH-cyclopenta[b]naphthalen-5-yl]oxy}acetic acid. Molecular Formula: C23H34O5 Molecular Weight: 390.51 Da Mechanism: Prostacyclin vasodilator; binds to the prostacyclin receptor inducing vasodilation of pulmonary and systemic arterial vascular beds, inhibits platelet aggregation and inflammatory cytokine production. Effect: Antithrombotic agents. Platelet aggregation inhibitors excl. heparin. | Mechanistic genes:P2RY12, PPARD, PTGIR Metabolic genes Substrate:CYP2C9 |
 | Name,Prasugrel IUPAC Name: (1) 5-[(1RS)-2-cyclopropyl-1-(2-fluorophenyl)-2-oxoethyl]-4,5,6,7-tetrahydrothieno[3,2-c]pyridin-2-yl acetate hydrochloride. (2) 2-[2-(Acetyloxy)-6,7-dihydrothieno[3,2-c]pyridin-5(4H)-yl]-1-cyclopropyl-2-(2-fluorophenyl)ethanone hydrochloride Molecular Formula: C20H21ClFNO3S Molecular Weight: 409.90 Da Mechanism: P2Y12 platelet inhibitor; impairs ADP-mediated activation of the GPIIb/IIIa complex. Effect: Antithrombotic agents. Platelet aggregation inhibitors excl. heparin. | Mechanistic genes:P2RY12 Metabolic genes Substrate:CES1, CES2, CYP2B6, CYP2C9, CYP2C19, CYP2D6, CYP3A4, GSTs, POR Inhibitor:CYP2B6, CYP2C9, CYP2C19, CYP2D6, CYP3A4, CYP3A5 Transporter genes:ABCB1, ALB |
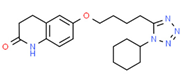 | Name:Cilostazol IUPAC Name: (1) 2(1H)-quinolinone, 6-[4-(1-cyclohexyl-1H-tetrazol-5-yl)butoxy]-3,4-dihydro-, (2) 6-[4-(1-cyclohexyl-1H-tetrazol-5-yl)butoxy]-3,4-dihydrocarbostyril Molecular Formula: C20H27N5O2 Molecular Weight: 369.46 Da Mechanism: Antiplatelet agent and vasodilator; inhibits PDE3 activation, increasing cAMP concentrations in platelets and blood vessels and mediating arterial vasodilation and inhibition of platelet aggregation. Reduces plasma triglyceride, but increases high-density lipoprotein cholesterol, levels. Effect: Antithrombotic agents. Platelet aggregation inhibitors excl. heparin. Phosphodiesterase Enzyme Inhibitor. | Mechanistic genes:PDE3A Metabolic genes Substrate:CYP1A2, CYP1B1, CYP2C8, CYP2C19, CYP2D6, CYP3A4, CYP3A5, CYP3A7 Transporter genes:ABCB1 |
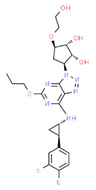 | Name:Ticagrelor IUPAC Name: (1S,2S,3R,5S)-3-[7-[[(1R,2S)-2-(3,4-difluorophenyl)cyclopropyl]amino]-5-propylsulfanyltriazolo[4,5-d]pyrimidin-3-yl]-5-(2-hydroxyethoxy)cyclopentane-1,2-diol Molecular Formula: C23H28F2N6O4S Molecular Weight: 522.57 Da Mechanism: P2Y12 platelet inhibitor; couples with Gαi2 and other Gi proteins to inhibit adenylyl cyclase. Activates PI3K, Akt, Rap1b, and K+ channels, mediating hemostasis and platelet aggregation. P2Y12 receptor blockade reduces development of occlusive thromboses, risk of MI and ischemic stroke. Effect: Antithrombotic agents. Platelet aggregation inhibitors excl. heparin. Selective adenosine diphosphate (ADP) receptor antagonist. | Mechanistic genes:P2RY12 Metabolic genes Substrate:CYP2C19, CYP3A4, CYP3A5, UGTs Inhibitor:ABCB1, CYP1A2, CYP2C9, CYP3A4 Inducer:CYP2B6, CYP2C9 Transporter genes:ABCB1, ALB |
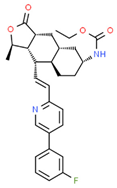 | Name: Vorapaxar IUPAC Name: ethyl N-[(1R,3aR,4aR,6R,8aR,9S,9aS)-9-[(1E)-2-[5-(3-fluorophenyl)pyridin-2-yl]ethenyl]-1-methyl-3-oxo-dodecahydronaphtho[2,3-c]furan-6-yl]carbamate. Molecular Formula: C29H33FN2O4 Molecular Weight: 492.58 Da Mechanism: Reversible PAR-1 antagonist, inhibits thrombin- and TRAP-induced platelet aggregation. Reduces thrombotic cardiovascular events in patients with a history of MI or PAD. Effect: Antithrombotic agents. Platelet aggregation inhibitors excl. heparin. | Mechanistic genes:F2R Metabolic genes Substrate:CYP2J2, CYP3A4 Transporter genes:ABCB1 |
 | Name: Selexipag IUPAC Name: 2-{4-[(5,6-diphenylpyrazin-2-yl)(propan-2-yl)amino]butoxy}-N-methanesulfonylacetamide. Molecular Formula: C26H32N4O4S Molecular Weight: 496.63 Da Mechanism: Selective PGI2 receptor agonist. Potent vasodilator with antiproliferative, anti-inflammatory, and antithrombotic effects. Effect: Antithrombotic agents. Platelet aggregation inhibitors excl. heparin. | Mechanistic genes:PTGIR Metabolic genes Substrate:CES1, CYP2C8, CYP3A4 Transporter genes:ABCB1, SLCO1B1, SLCO1B3 |
| Direct thrombin inhibitors | ||
 | Name: Desirudin Molecular Formula: C287H440N80O110S6 Molecular Weight: 6963.52 Da Mechanism: Direct, highly selective thrombin inhibitor. Reversibly binds to the active thrombin site of free and clot-associated thrombin. Inhibits fibrin formation, activation of coagulation factors V, VII, and XIII, and thrombin-induced platelet aggregation. Effect: Antithrombotic agents. Direct thrombin inhibitors. | Mechanistic genes:F2, F5, F7, F13A1 Metabolic genes Substrate:CPA1 |
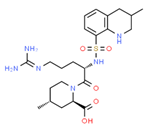 | Name:Argatroban IUPAC Name: (1) (2R,4R)-1-{N5-(Diaminomethylene)-N2-[(3-methyl-1,2,3,4-tetrahydro-8-quinolinyl)sulfonyl]-L-ornithyl}-4-methyl-2-piperidinecarboxylic acid Molecular Formula: C23H36N6O5S Molecular Weight: 508.634 Da Mechanism: Direct thrombin inhibitor; Reversibly binds to the active thrombin site of free and clot-associated thrombin. Inhibits fibrin formation, activation of coagulation factors V, VIII, and XIII, protein C, and platelet aggregation. Effect: Antithrombotic agents. Direct thrombin inhibitors. | Mechanistic genes:F2, F5, F8, F13, PROC Metabolic genes Substrate:CYP3A4, CYP3A5 |
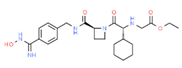 | Name: Ximelagatran IUPAC Name: ethyl 2-{[(1R)-1-cyclohexyl-2-[(2S)-2-[({4-[(Z)-N’-hydroxycarbamimidoyl]phenyl}methyl)carbamoyl]azetidin-1-yl]-2-oxoethyl]amino}acetate. Molecular Formula: C24H35N5O5 Molecular Weight: 473.56 Da Mechanism: Bioconverted to the active moiety, melagatran, which inhibits clot-bound thrombin. Effect: Antithrombotic agents. Direct thrombin inhibitors. | Mechanistic genes:F2 Metabolic genes Substrate:CYP2C9 |
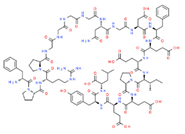 | Name:Bivalirudin IUPAC Name: 1) L-Leucine, D-phenylalanyl-L-prolyl-L-arginyl-L-prolylglycylglycylglycylglycyl-L-asparaginylglycyl-L-α-aspartyl-L-phenylalanyl-L-α-glutamyl-L-α-glutamyl-L-isoleucyl-L-prolyl-L-α-glutamyl-L-α-glutamyl-L-tyrosyl-, (2) D-phenylalanyl-L-prolyl-L-arginyl-L-prolylglycylglycylglycylglycyl-L-asparaginylglycyl-L-α-aspartyl-L-phenylalanyl-L-α-glutamyl-L-α-glutamyl-L-isoleucyl-L-prolyl-L-α-glutamyl-L-α-glutamyl-L-tyrosyl-L-leucine Molecular Formula: C98H138N24O33 Molecular Weight: 2180.29 Da Mechanism: Reversible direct thrombin inhibitor for heparin-induced thrombocytopenia. Inhibits thrombin by binding to its catalytic and anion-binding exosite, preventing thrombin-mediated cleavage of fibrinogen to fibrin, activation of factors V, VIII, and XIII, conversion of fibrinogen to fibrin, and platelet activation and aggregation. Effect: Antithrombotic agents. Direct thrombin inhibitors. | Mechanistic genes:F2, F5, F8, F13, FGA Metabolic genes Inhibitor:MPO |
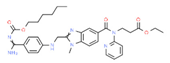 | Name:Dabigatran IUPAC Name: Ethyl 3-[[[2-[[[4-[[[(hexyloxy)carbonyl]amino]iminomethyl]phenyl]amino]methyl]-1-methyl-1H-benzimidazol-5-yl]carbonyl](pyridin-2-yl)amino]propanoate Molecular Formula: C34H41N7O5 Molecular Weight: 627.73 Da Mechanism: Inhibits coagulation by preventing thrombin-mediated effects, including cleavage of fibrinogen to fibrin monomers, activation of factors V, VIII, XI and XIII, and inhibition of thrombin-induced platelet aggregation. Effect: Antithrombotic agents. Direct thrombin inhibitors. | Mechanistic genes:F2, F5, F8, F11, F13, FGA Metabolic genes Substrate:CES1 Transporter genes:ABCB1 |
| Direct factor Xa inhibitors | ||
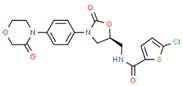 | Name: Rivaroxaban IUPAC Name: 5-chloro-N-{[(5S)-2-oxo-3-[4-(3-oxomorpholin-4-yl)phenyl]-1,3-oxazolidin-5-yl]methyl}thiophene-2-carboxamide. Molecular Formula: C19H18ClN3O5S Molecular Weight: 435.88 Da Mechanism: Anticoagulant, irreversibly inhibits free and clot bound factor Xa; treating DVT and PE. Effect: Antithrombotic agents. Direct Factor Xa inhibitors. | Mechanistic genes:F2, F10 Metabolic genes Substrate:CYP2J2, CYP3A4, CYP3A5 Transporter genes:ABCB1, ABCG2 |
 | Name: Apixaban IUPAC Name: 1-(4-methoxyphenyl)-7-oxo-6-[4-(2-oxopiperidin-1-yl)phenyl]-1H,4H,5H,6H,7H-pyrazolo[3,4-c]pyridine-3-carboxamide. Molecular Formula: C25H25N5O4 Molecular Weight: 459.50 Da Mechanism: Inhibits factor Xa, independent of antithrombin III. Inhibits prothrombin, preventing thrombus formation. Effect: Antithrombotic agents. Direct Factor Xa inhibitors. | Mechanistic genes:F2, F5, F10 Metabolic genes Substrate:CYP2C8, CYP2C19, CYP2C9, CYP1A2, CYP2J2, CYP3A4, CYP3A5 Inhibitor:CYP2C19 Transporter genes:ABCB1, ABCG2 |
 | Name: Edoxaban IUPAC Name: N’-(5-chloropyridin-2-yl)-N-[(1S,2R,4S)-4-(dimethylcarbamoyl)-2-{5-methyl-4H,5H,6H,7H-[1,3]thiazolo[5,4-c]pyridine-2-amido}cyclohexyl]ethanediamide. Molecular Formula: C24H30ClN7O4S Molecular Weight: 548.06 Da Mechanism: Selective Factor Xa inhibitor. Effect: Antithrombotic agents. Direct Factor Xa inhibitors. | Mechanistic genes:F10 Transporter genes:ABCB1 |
 | Name: Betrixaban IUPAC Name: N-(5-chloropyridin-2-yl)-2-[4-(N,N-dimethylcarbamimidoyl)benzamido]-5-methoxybenzamide. Molecular Formula: C23H22ClN5O3 Molecular Weight: 451.91 Da Mechanism: Cofactor-independent direct inhibitor of free and prothrombinase-bound Factor Xa. Effect: Antithrombotic agents. Direct Factor Xa inhibitors. | Mechanistic genes:F2, F10 Transporter genes:ABCB1, KCNH2 |
| ABCB1 | ABCC2 | CYP2C9 | CYP2C19 | CYP2D6 | CYP3A4 | CYP3A5 | NAT2 | SLCO1B1 | |||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| ANTIVIRALS | |||||||||||||||||||||||||||
| CHLOROQUINE |  |  |  | ||||||||||||||||||||||||
| HYDROXYCHLOROQUINE |  |  |  | ||||||||||||||||||||||||
| LOPINAVIR |  |  |  |  |  |  |  |  | |||||||||||||||||||
| REMSEDIVIR |  |  |  |  |  |  | |||||||||||||||||||||
| RITONAVIR |  |  |  |  |  |  |  |  | |||||||||||||||||||
| ANTIPYRETICS | |||||||||||||||||||||||||||
| PARACETAMOL |  |  |  |  |  |  |  |  |  |  | |||||||||||||||||
| CORTICOSTEROIDS FOR SYSTEMIC USE | |||||||||||||||||||||||||||
| ALDOSTERONE |  |  |  |  | |||||||||||||||||||||||
| BECLOMETASONE |  |  |  | ||||||||||||||||||||||||
| BETAMETASONE |  |  |  | ||||||||||||||||||||||||
| BUDESONIDE |  |  |  |  |  |  |  | ||||||||||||||||||||
| CORTISONE |  |  |  | ||||||||||||||||||||||||
| CICLESONIDE |  |  |  |  | |||||||||||||||||||||||
| DEFLAZACORT |  |  | |||||||||||||||||||||||||
| DEXAMETASONE |  |  |  |  |  |  |  |  |  |  |  |  |  | ||||||||||||||
| FLUNISOLIDE |  |  |  |  | |||||||||||||||||||||||
| FLUTICASONE |  |  |  |  | |||||||||||||||||||||||
| HYDROCORTISONE |  |  |  |  | |||||||||||||||||||||||
| METHYLPRENISOLONE |  |  |  |  |  | ||||||||||||||||||||||
| MOMETASONE |  |  |  | ||||||||||||||||||||||||
| PREDNISOLONE |  |  |  |  |  |  |  |  |  | ||||||||||||||||||
| RIMEXOLONE |  | ||||||||||||||||||||||||||
| TIXOCORTOL |  | ||||||||||||||||||||||||||
| TRIAMCINOLONE |  | ||||||||||||||||||||||||||
| NON-STEROIDAL ANTIINFLAMMATORY PRODUCTS | |||||||||||||||||||||||||||
| ACECLOFENAC |  | ||||||||||||||||||||||||||
| BENZYDAMINE |  |  |  |  | |||||||||||||||||||||||
| BENOXAPROFEN | |||||||||||||||||||||||||||
| CELECOXIB |  |  |  |  |  |  | |||||||||||||||||||||
| DEXIBUPROFEN |  | ||||||||||||||||||||||||||
| DICLOFENAC |  |  |  |  |  |  |  |  |  |  |  |  |  |  | |||||||||||||
| ETODOLAC |  |  |  |  | |||||||||||||||||||||||
| ETORICOXIB |  |  |  |  |  |  |  |  |  | ||||||||||||||||||
| FENOPROFEN |  | ||||||||||||||||||||||||||
| FLURBIPROFEN |  |  |  |  |  | ||||||||||||||||||||||
| IBUPROFEN |  |  |  |  |  |  |  |  | |||||||||||||||||||
| INDOMETHACIN |  |  |  |  |  |  |  |  |  |  |  |  |  | ||||||||||||||
| INDOPROFEN | |||||||||||||||||||||||||||
| KETOPROFEN |  |  | |||||||||||||||||||||||||
| KETOROLAC | |||||||||||||||||||||||||||
| LORNOXICAM |  |  |  |  | |||||||||||||||||||||||
| LUMIRACOXIB |  |  |  |  | |||||||||||||||||||||||
| MEFENAMIC ACID |  |  | |||||||||||||||||||||||||
| MELOXICAM |  |  |  |  |  | ||||||||||||||||||||||
| NABUMETONE |  |  |  |  |  | ||||||||||||||||||||||
| NAPROXEN |  |  | |||||||||||||||||||||||||
| NIFLUMIC ACID | |||||||||||||||||||||||||||
| NIMESULIDE |  | ||||||||||||||||||||||||||
| OXAPROZIN |  | ||||||||||||||||||||||||||
| PENICILLAMINE |  | ||||||||||||||||||||||||||
| PIROXICAM |  |  |  |  | |||||||||||||||||||||||
| ROFECOXIB |  | ||||||||||||||||||||||||||
| SULINDAC |  |  |  |  | |||||||||||||||||||||||
| SUPROFEN |  | ||||||||||||||||||||||||||
| TENOXICAM |  |  |  |  | |||||||||||||||||||||||
| VALDECOXIB |  |  |  | ||||||||||||||||||||||||
| ANTIHYPERTENSIVE AGENTS | |||||||||||||||||||||||||||
| AMLODIPINE |  |  |  |  |  |  |  |  | |||||||||||||||||||
| ATENOLOL |  | ||||||||||||||||||||||||||
| BISOPROLOL |  |  |  |  | |||||||||||||||||||||||
| CANDESARTAN |  |  |  | ||||||||||||||||||||||||
| CARVEDILOL |  |  |  |  |  |  |  | ||||||||||||||||||||
| ENALAPRIL |  |  | |||||||||||||||||||||||||
| HYDROCHLOROTIAZIDE | |||||||||||||||||||||||||||
| INDAPAMIDE |  |  | |||||||||||||||||||||||||
| IRBESARTAN |  |  |  |  |  |  |  |  | |||||||||||||||||||
| ISOSORBIDE |  |  | |||||||||||||||||||||||||
| LERCANIDIPINE | |||||||||||||||||||||||||||
| LISINOPRIL |  |  |  | ||||||||||||||||||||||||
| LOSARTAN |  |  |  |  |  |  |  |  |  | ||||||||||||||||||
| METOPROLOL |  |  |  |  |  | ||||||||||||||||||||||
| NEBIVOLOL |  |  | |||||||||||||||||||||||||
| NIFEDIPINE |  |  |  |  |  |  |  |  |  |  | |||||||||||||||||
| OLMERSARTAN |  |  |  |  |  | ||||||||||||||||||||||
| PERINDOPRIL | |||||||||||||||||||||||||||
| RAMIPRIL | |||||||||||||||||||||||||||
| TELMISARTAN |  |  |  |  |  |  |  | ||||||||||||||||||||
| VALSARTAN |  |  |  | ||||||||||||||||||||||||
| DIURETICS | |||||||||||||||||||||||||||
| AMILORIDE |  |  | |||||||||||||||||||||||||
| CONIVAPTAN |  |  |  |  | |||||||||||||||||||||||
| FUROSEMIDE |  | ||||||||||||||||||||||||||
| INDAPAMIDE |  |  | |||||||||||||||||||||||||
| MONTELUKAST |  |  |  |  |  | ||||||||||||||||||||||
| TERBUTALINE | |||||||||||||||||||||||||||
| TOLVAPTAN |  |  |  |  | |||||||||||||||||||||||
| TORASEMIDE |  |  | |||||||||||||||||||||||||
| TRIAMTERENE |  |  | |||||||||||||||||||||||||
 Substrate;
Substrate;  Inhibitor;
Inhibitor;  Inducer.
Inducer.Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Cacabelos, R.; Naidoo, V.; Corzo, L.; Cacabelos, N.; Carril, J.C. Genophenotypic Factors and Pharmacogenomics in Adverse Drug Reactions. Int. J. Mol. Sci. 2021, 22, 13302. https://doi.org/10.3390/ijms222413302
Cacabelos R, Naidoo V, Corzo L, Cacabelos N, Carril JC. Genophenotypic Factors and Pharmacogenomics in Adverse Drug Reactions. International Journal of Molecular Sciences. 2021; 22(24):13302. https://doi.org/10.3390/ijms222413302
Chicago/Turabian StyleCacabelos, Ramón, Vinogran Naidoo, Lola Corzo, Natalia Cacabelos, and Juan C. Carril. 2021. "Genophenotypic Factors and Pharmacogenomics in Adverse Drug Reactions" International Journal of Molecular Sciences 22, no. 24: 13302. https://doi.org/10.3390/ijms222413302
APA StyleCacabelos, R., Naidoo, V., Corzo, L., Cacabelos, N., & Carril, J. C. (2021). Genophenotypic Factors and Pharmacogenomics in Adverse Drug Reactions. International Journal of Molecular Sciences, 22(24), 13302. https://doi.org/10.3390/ijms222413302







