Tracing CRISPR/Cas12a Mediated Genome Editing Events in Apple Using High-Throughput Genotyping by PCR Capillary Gel Electrophoresis
Abstract
:1. Introduction
2. Results
2.1. Design and Pre-Selection of Efficient MdPDS Specific crRNAs for LbCas12a
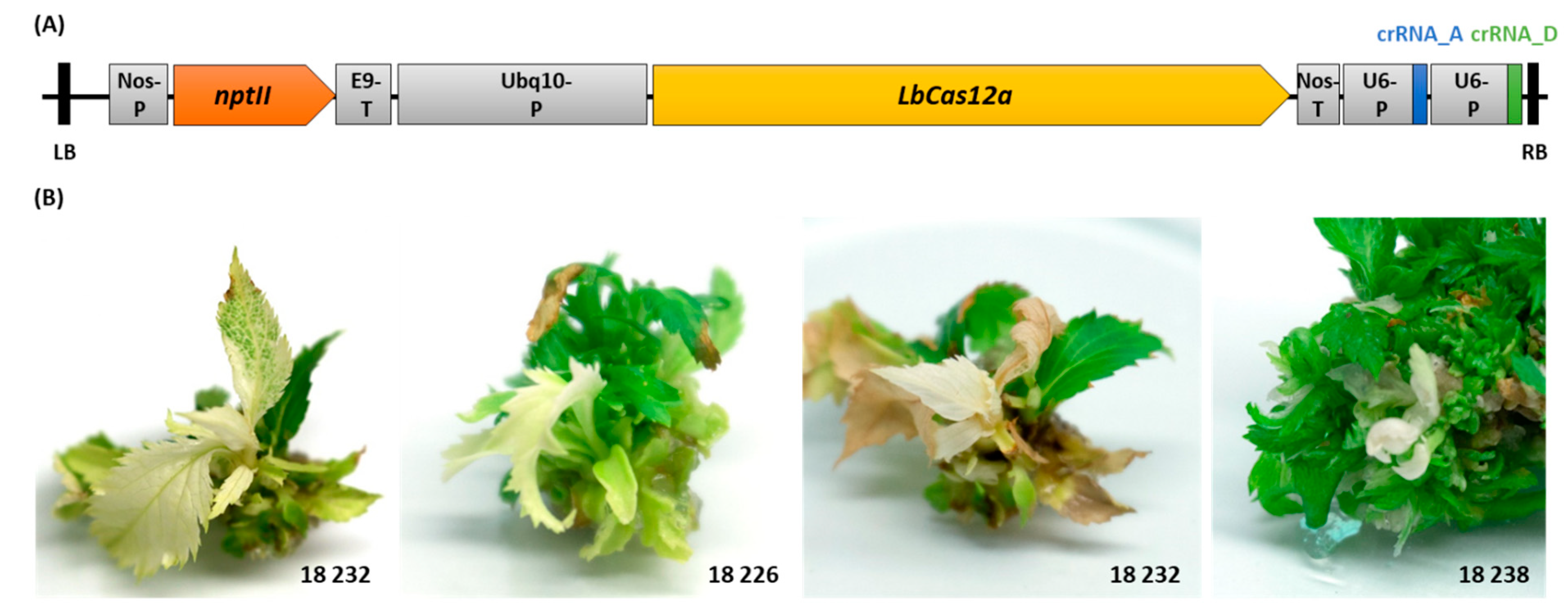
2.2. MdPDS Mutagenesis by T-DNA Transformation of CRISPR/LbCas12a Construct
2.3. High-Throughput Genotyping Indels Using Fluorescent PCR Combined with Capillary Gel Electrophoresis
2.4. Verification of GE Events by Amplicon Deep Sequencing
2.5. Characterization of GE Events in Apple, Introduced by CRISPR/LbCas12a
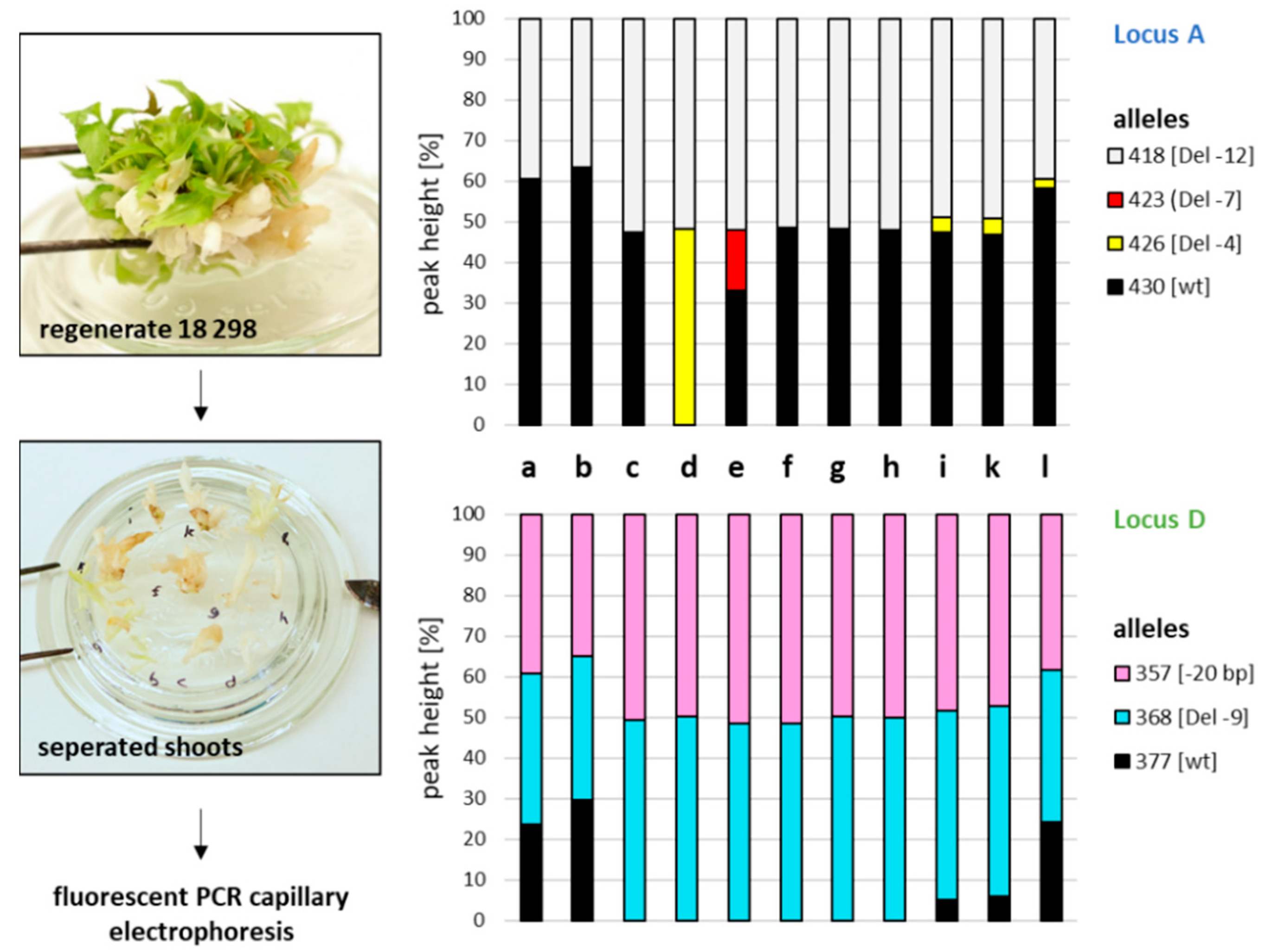
2.6. Tracing GE Events in Chimeric Regenerates
3. Discussion
4. Materials and Methods
4.1. Oligonucleotides and Kits
4.2. Guide RNA Design and Determination of DNA Cleavage Efficiency
4.3. Transformation of the CRISPR/LbCas12a Construct
4.4. Detection of Transgenic Elements by PCR
4.5. Genotyping of Indels by Fluorescent PCR Capillary Gel Electrophoresis
4.6. Amplicon Deep Sequencing of MdPDS Target Sites
4.7. Cloning and Sanger Sequencing
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Jinek, M.; Chylinski, K.; Fonfara, I.; Hauer, M.; Doudna, J.A.; Charpentier, E. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 2012, 337, 816–821. [Google Scholar] [CrossRef]
- Feng, Z.Y.; Zhang, B.T.; Ding, W.N.; Liu, X.D.; Yang, D.L.; Wei, P.L.; Cao, F.Q.; Zhu, S.H.; Zhang, F.; Mao, Y.F.; et al. Efficient genome editing in plants using a CRISPR/Cas system. Cell Res. 2013, 23, 1229–1232. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shan, Q.; Wang, Y.; Li, J.; Zhang, Y.; Chen, K.; Liang, Z.; Zhang, K.; Liu, J.; Xi, J.J.; Qiu, J.L.; et al. Targeted genome modification of crop plants using a CRISPR-Cas system. Nat. Biotechnol. 2013, 31, 686–688. [Google Scholar] [CrossRef]
- Puchta, H. The repair of double-strand breaks in plants: Mechanisms and consequences for genome evolution. J. Exp. Bot. 2005, 56, 1–14. [Google Scholar] [CrossRef] [Green Version]
- Chen, K.; Wang, Y.; Zhang, R.; Zhang, H.; Gao, C. CRISPR/Cas Genome Editing and Precision Plant Breeding in Agriculture. Annu. Rev. Plant Biol. 2019, 70, 667–697. [Google Scholar] [CrossRef]
- Makarova, K.S.; Wolf, Y.I.; Iranzo, J.; Shmakov, S.A.; Alkhnbashi, O.S.; Brouns, S.J.J.; Charpentier, E.; Cheng, D.; Haft, D.H.; Horvath, P.; et al. Evolutionary classification of CRISPR-Cas systems: A burst of class 2 and derived variants. Nat. Rev. Microbiol. 2020, 18, 67–83. [Google Scholar] [CrossRef]
- Shmakov, S.; Smargon, A.; Scott, D.; Cox, D.; Pyzocha, N.; Yan, W.; Abudayyeh, O.O.; Gootenberg, J.S.; Makarova, K.S.; Wolf, Y.I.; et al. Diversity and evolution of class 2 CRISPR-Cas systems. Nat. Rev. Microbiol. 2017, 15, 169–182. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zetsche, B.; Gootenberg, J.S.; Abudayyeh, O.O.; Slaymaker, I.M.; Makarova, K.S.; Essletzbichler, P.; Volz, S.E.; Joung, J.; van der Oost, J.; Regev, A.; et al. Cpf1 is a single RNA-guided endonuclease of a class 2 CRISPR-Cas system. Cell 2015, 163, 759–771. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Alok, A.; Sandhya, D.; Jogam, P.; Rodrigues, V.; Bhati, K.K.; Sharma, H.; Kumar, J. The Rise of the CRISPR/Cpf1 System for Efficient Genome Editing in Plants. Front. Plant Sci. 2020, 11, 264. [Google Scholar] [CrossRef]
- Safari, F.; Zare, K.; Negahdaripour, M.; Barekati-Mowahed, M.; Ghasemi, Y. CRISPR Cpf1 proteins: Structure, function and implications for genome editing. Cell Biosci. 2019, 9, 36. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bernabe-Orts, J.M.; Casas-Rodrigo, I.; Minguet, E.G.; Landolfi, V.; Garcia-Carpintero, V.; Gianoglio, S.; Vazquez-Vilar, M.; Granell, A.; Orzaez, D. Assessment of Cas12a-mediated gene editing efficiency in plants. Plant Biotechnol. J. 2019, 17, 1971–1984. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jia, H.; Orbovic, V.; Wang, N. CRISPR-LbCas12a-mediated modification of citrus. Plant Biotechnol. J. 2019, 17, 1928–1937. [Google Scholar] [CrossRef] [Green Version]
- Kim, H.; Kim, S.T.; Ryu, J.; Kang, B.C.; Kim, J.S.; Kim, S.G. CRISPR/Cpf1-mediated DNA-free plant genome editing. Nat. Commun. 2017, 8, 14406. [Google Scholar] [CrossRef] [Green Version]
- Li, S.; Zhang, X.; Wang, W.; Guo, X.; Wu, Z.; Du, W.; Zhao, Y.; Xia, L. Expanding the Scope of CRISPR/Cpf1-Mediated Genome Editing in Rice. Mol. Plant 2018, 11, 995–998. [Google Scholar] [CrossRef] [Green Version]
- Nishitani, C.; Hirai, N.; Komori, S.; Wada, M.; Okada, K.; Osakabe, K.; Yamamoto, T.; Osakabe, Y. Efficient Genome Editing in Apple Using a CRISPR/Cas9 system. Sci. Rep. 2016, 6, 31481. [Google Scholar] [CrossRef]
- Qin, G.; Gu, H.; Ma, L.; Peng, Y.; Deng, X.W.; Chen, Z.; Qu, L.J. Disruption of phytoene desaturase gene results in albino and dwarf phenotypes in Arabidopsis by impairing chlorophyll, carotenoid, and gibberellin biosynthesis. Cell Res. 2007, 17, 471–482. [Google Scholar] [CrossRef] [Green Version]
- Pompili, V.; Dalla Costa, L.; Piazza, S.; Pindo, M.; Malnoy, M. Reduced fire blight susceptibility in apple cultivars using a high-efficiency CRISPR/Cas9-FLP/FRT-based gene editing system. Plant Biotechnol. J. 2020, 18, 845–858. [Google Scholar] [CrossRef]
- Charrier, A.; Vergne, E.; Dousset, N.; Richer, A.; Petiteau, A.; Chevreau, E. Efficient Targeted Mutagenesis in Apple and First Time Edition of Pear Using the CRISPR-Cas9 System. Front. Plant Sci. 2019, 10, 40. [Google Scholar] [CrossRef] [Green Version]
- Zhou, H.; Bai, S.; Wang, N.; Sun, X.; Zhang, Y.; Zhu, J.; Dong, C. CRISPR/Cas9-Mediated Mutagenesis of MdCNGC2 in Apple Callus and VIGS-Mediated Silencing of MdCNGC2 in Fruits Improve Resistance to Botryosphaeria dothidea. Front. Plant Sci. 2020, 11, 575477. [Google Scholar] [CrossRef] [PubMed]
- Malnoy, M.; Viola, R.; Jung, M.H.; Koo, O.J.; Kim, S.; Kim, J.S.; Velasco, R.; Kanchiswamy, C.N. DNA-Free Genetically Edited Grapevine and Apple Protoplast Using CRISPR/Cas9 Ribonucleoproteins. Front. Plant Sci. 2016, 7, 1904. [Google Scholar] [CrossRef] [PubMed]
- Osakabe, Y.; Liang, Z.C.; Ren, C.; Nishitani, C.; Osakabe, K.; Wada, M.; Komori, S.; Malnoy, M.; Velasco, R.; Poli, M.; et al. CRISPR-Cas9-mediated genome editing in apple and grapevine. Nat. Protoc. 2018, 13, 2844–2863. [Google Scholar] [CrossRef]
- Malabarba, J.; Chevreau, E.; Dousset, N.; Veillet, F.; Moizan, J.; Vergne, E. New Strategies to Overcome Present CRISPR/Cas9 Limitations in Apple and Pear: Efficient Dechimerization and Base Editing. Int. J. Mol. Sci. 2020, 22, 319. [Google Scholar] [CrossRef]
- Zhang, Y.; Zhou, P.; Bozorov, T.A.; Zhang, D. Application of CRISPR/Cas9 technology in wild apple (Malus sieverii) for paired sites gene editing. Plant Methods 2021, 17, 79. [Google Scholar] [CrossRef] [PubMed]
- Zischewski, J.; Fischer, R.; Bortesi, L. Detection of on-target and off-target mutations generated by CRISPR/Cas9 and other sequence-specific nucleases. Biotechnol. Adv. 2017, 35, 95–104. [Google Scholar] [CrossRef] [PubMed]
- Vouillot, L.; Thelie, A.; Pollet, N. Comparison of T7E1 and surveyor mismatch cleavage assays to detect mutations triggered by engineered nucleases. G3 Genes Genomes Genet. 2015, 5, 407–415. [Google Scholar] [CrossRef] [Green Version]
- Thomas, H.R.; Percival, S.M.; Yoder, B.K.; Parant, J.M. High-throughput genome editing and phenotyping facilitated by high resolution melting curve analysis. PLoS ONE 2014, 9, e114632. [Google Scholar] [CrossRef] [PubMed]
- Zhu, X.; Xu, Y.; Yu, S.; Lu, L.; Ding, M.; Cheng, J.; Song, G.; Gao, X.; Yao, L.; Fan, D.; et al. An efficient genotyping method for genome-modified animals and human cells generated with CRISPR/Cas9 system. Sci. Rep. 2014, 4, 6420. [Google Scholar] [CrossRef] [PubMed]
- Ran, F.A.; Hsu, P.D.; Lin, C.Y.; Gootenberg, J.S.; Konermann, S.; Trevino, A.E.; Scott, D.A.; Inoue, A.; Matoba, S.; Zhang, Y.; et al. Double nicking by RNA-guided CRISPR Cas9 for enhanced genome editing specificity. Cell 2013, 154, 1380–1389. [Google Scholar] [CrossRef] [Green Version]
- Yu, C.; Zhang, Y.; Yao, S.; Wei, Y. A PCR based protocol for detecting indel mutations induced by TALENs and CRISPR/Cas9 in zebrafish. PLoS ONE 2014, 9, e98282. [Google Scholar] [CrossRef] [Green Version]
- Bauer, D.E.; Canver, M.C.; Orkin, S.H. Generation of genomic deletions in mammalian cell lines via CRISPR/Cas9. J. Vis. Exp. 2015, e52118. [Google Scholar] [CrossRef]
- Kim, J.M.; Kim, D.; Kim, S.; Kim, J.S. Genotyping with CRISPR-Cas-derived RNA-guided endonucleases. Nat. Commun. 2014, 5, 3157. [Google Scholar] [CrossRef] [PubMed]
- Ramlee, M.K.; Yan, T.D.; Cheung, A.M.S.; Chuah, C.T.H.; Li, S. High-throughput genotyping of CRISPR/Cas9-mediated mutants using fluorescent PCR-capillary gel electrophoresis. Sci. Rep. 2015, 5, 15587. [Google Scholar] [CrossRef]
- Brinkman, E.K.; Chen, T.; Amendola, M.; van Steensel, B. Easy quantitative assessment of genome editing by sequence trace decomposition. Nucleic Acids Res. 2014, 42, e168. [Google Scholar] [CrossRef]
- Guell, M.; Yang, L.H.; Church, G.M. Genome editing assessment using CRISPR Genome Analyzer (CRISPR-GA). Bioinformatics 2014, 30, 2968–2970. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hendel, A.; Kildebeck, E.J.; Fine, E.J.; Clark, J.T.; Punjya, N.; Sebastiano, V.; Bao, G.; Porteus, M.H. Quantifying Genome-Editing Outcomes at Endogenous Loci with SMRT Sequencing. Cell Rep. 2014, 7, 293–305. [Google Scholar] [CrossRef] [Green Version]
- Anders, C.; Niewoehner, O.; Duerst, A.; Jinek, M. Structural basis of PAM-dependent target DNA recognition by the Cas9 endonuclease. Nature 2014, 513, 569–573. [Google Scholar] [CrossRef]
- Wolter, F.; Puchta, H. In planta gene targeting can be enhanced by the use of CRISPR/Cas12a. Plant J. 2019, 100, 1083–1094. [Google Scholar] [CrossRef] [Green Version]
- Moreno-Mateos, M.A.; Fernandez, J.P.; Rouet, R.; Vejnar, C.E.; Lane, M.A.; Mis, E.; Khokha, M.K.; Doudna, J.A.; Giraldez, A.J. CRISPR-Cpf1 mediates efficient homology-directed repair and temperature-controlled genome editing. Nat. Commun. 2017, 8, 2024. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Schindele, P.; Puchta, H. Engineering CRISPR/LbCas12a for highly efficient, temperature-tolerant plant gene editing. Plant Biotechnol. J. 2020, 18, 1118–1120. [Google Scholar] [CrossRef] [Green Version]
- Lee, K.; Zhang, Y.; Kleinstiver, B.P.; Guo, J.A.; Aryee, M.J.; Miller, J.; Malzahn, A.; Zarecor, S.; Lawrence-Dill, C.J.; Joung, J.K.; et al. Activities and specificities of CRISPR/Cas9 and Cas12a nucleases for targeted mutagenesis in maize. Plant Biotechnol. J. 2019, 17, 362–372. [Google Scholar] [CrossRef] [Green Version]
- Malzahn, A.A.; Tang, X.; Lee, K.; Ren, Q.; Sretenovic, S.; Zhang, Y.; Chen, H.; Kang, M.; Bao, Y.; Zheng, X.; et al. Application of CRISPR-Cas12a temperature sensitivity for improved genome editing in rice, maize, and Arabidopsis. BMC Biol. 2019, 17, 9. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, Y.; Ren, Q.; Tang, X.; Liu, S.; Malzahn, A.A.; Zhou, J.; Wang, J.; Yin, D.; Pan, C.; Yuan, M.; et al. Expanding the scope of plant genome engineering with Cas12a orthologs and highly multiplexable editing systems. Nat. Commun. 2021, 12, 1944. [Google Scholar] [CrossRef] [PubMed]
- Nishitani, C.; Osakabe, K.; Osakabe, Y. Genome Editing in Apple. In The Apple Genome; Compendium of Plant Genomes; Korban, S.S., Ed.; Springer: Cham, Switzerland, 2021; pp. 213–225. [Google Scholar]
- Duan, H.Y.; Ding, X.S.; Song, J.Y.; Duan, Z.K.; Zhou, Y.Q.; Zhou, C.N. Effects of Kanamycin on Growth and Development of Arabidopsis Thaliana Seedling, Cotyledon and Leaf. Pak. J. Bot. 2009, 41, 1611–1618. [Google Scholar]
- Zhang, B.H.; Liu, F.; Liu, Z.H.; Wang, H.M.; Yao, C.B. Effects of kanamycin on tissue culture and somatic embryogenesis in cotton. Plant Growth Regul. 2001, 33, 137–149. [Google Scholar] [CrossRef]
- Kirik, A.; Salomon, S.; Puchta, H. Species-specific double-strand break repair and genome evolution in plants. Eur. Mol. Biol. Organ. J. 2000, 19, 5562–5566. [Google Scholar] [CrossRef] [Green Version]
- Fan, D.; Liu, T.; Li, C.; Jiao, B.; Li, S.; Hou, Y.; Luo, K. Efficient CRISPR/Cas9-mediated Targeted Mutagenesis in Populus in the First Generation. Sci. Rep. 2015, 5, 12217. [Google Scholar] [CrossRef]
- Wang, Z.; Wang, S.; Li, D.; Zhang, Q.; Li, L.; Zhong, C.; Liu, Y.; Huang, H. Optimized paired-sgRNA/Cas9 cloning and expression cassette triggers high-efficiency multiplex genome editing in kiwifruit. Plant Biotechnol. J. 2018, 16, 1424–1433. [Google Scholar] [CrossRef] [Green Version]
- Wang, X.; Tu, M.; Wang, D.; Liu, J.; Li, Y.; Li, Z.; Wang, Y.; Wang, X. CRISPR/Cas9-mediated efficient targeted mutagenesis in grape in the first generation. Plant Biotechnol. J. 2018, 16, 844–855. [Google Scholar] [CrossRef] [Green Version]
- Abuqaoud, H.; Skirvin, R.M.; Chevreau, E. Invitro Separation of Chimeral Pears into Their Component Genotypes. Euphytica 1990, 48, 189–196. [Google Scholar] [CrossRef]
- Petree, C.; Varshney, G.K. MultiFRAGing: Rapid and Simultaneous Genotyping of Multiple Alleles in a Single Reaction. Sci. Rep. 2020, 10, 3172. [Google Scholar] [CrossRef] [Green Version]
- Park, J.; Bae, S.; Kim, J.S. Cas-Designer: A web-based tool for choice of CRISPR-Cas9 target sites. Bioinformatics 2015, 31, 4014–4016. [Google Scholar] [CrossRef] [Green Version]
- Bae, S.; Park, J.; Kim, J.S. Cas-OFFinder: A fast and versatile algorithm that searches for potential off-target sites of Cas9 RNA-guided endonucleases. Bioinformatics 2014, 30, 1473–1475. [Google Scholar] [CrossRef] [Green Version]
- Hood, E.E.; Gelvin, S.B.; Melchers, L.S.; Hoekema, A. New Agrobacterium Helper Plasmids for Gene-Transfer to Plants. Transgen. Res. 1993, 2, 208–218. [Google Scholar] [CrossRef]
- Flachowsky, H.; Peil, A.; Sopanen, T.; Elo, A.; Hanke, V. Overexpression of BpMADS4 from silver birch (Betula pendula Roth.) induces early-flowering in apple (Malus x domestica Borkh.). Plant Breed. 2007, 126, 137–145. [Google Scholar] [CrossRef]
- Boudichevskaia, A.; Flachowsky, H.; Dunemann, F. Identification and molecular analysis of candidate genes homologous to HcrVf genes for scab resistance in apple. Plant Breed. 2009, 128, 84–91. [Google Scholar] [CrossRef]
- Afgan, E.; Baker, D.; Batut, B.; van den Beek, M.; Bouvier, D.; Cech, M.; Chilton, J.; Clements, D.; Coraor, N.; Gruning, B.A.; et al. The Galaxy platform for accessible, reproducible and collaborative biomedical analyses: 2018 update. Nucleic Acids Res. 2018, 46, W537–W544. [Google Scholar] [CrossRef] [PubMed] [Green Version]




| Material | Locus A | Locus D | |||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Deep Sequencing | Fragment Length Analysis | Deep Sequencing | Fragment Length Analysis | ||||||||||||||||
| Regenerate | Shoot | Reads analyzed | Type | Diff. from wt | [%] Reads | Allele size | Diff. to wt | Peak Height | Sum Peak Heights | [%] Peak Height | Reads analyzed | Type | Diff. from wt | [%] Reads | Allele size | Diff. from wt | Peak Height | Sum Peak Heights | [%] Peak Height |
| Gala | 163,807 | wt | 0 | 100.0 | 430 | 0 | 13,617 | 13,617 | 100.0 | 151,045 | wt | 0 | 100.0 | 377 | 0 | 21,545 | 21,545 | 100.0 | |
| 18 219 | a | 197,114 | Del | −84 | 62.9 | 346 | −84 | 11,069 | 18,988 | 58.3 | 167,120 | Del | −12 | 51.0 | 365 | −12 | 13,582 | 26,958 | 50.4 |
| wt | 0 | 35.1 | 430 | 0 | 7480 | 39.4 | Del | −2 | 49.0 | 375 | −2 | 13,376 | 49.6 | ||||||
| Del | −4 | 2.1 | 426 | −4 | 439 | 2.3 | |||||||||||||
| 18 234 | 1 | 215,296 | Del | −1 | 38.6 | 429 | −1 | 6854 | 17,093 | 40.1 | 136,560 | wt | 0 | 43.8 | 377 | 0 | 9734 | 23,160 | 42.0 |
| Del | −4 | 33.4 | 426 | −4 | 5267 | 30.8 | Del | −8 | 27.8 | 369 | −8 | 6586 | 28.4 | ||||||
| wt | 0 | 25.6 | 430 | 0 | 4552 | 26.6 | Del | −10 | 16.2 | 367 | −10 | 3769 | 16.3 | ||||||
| Del | −8 | 2.4 | 422 | −8 | 420 | 2.5 | Del | −13 | 7.2 | 364 | −13 | 1657 | 7.2 | ||||||
| Del | −17 | 1.9 | 360 | −17 | 499 | 2.2 | |||||||||||||
| Del | −16 | 1.7 | 361 | −16 | 406 | 1.8 | |||||||||||||
| Del | −9 | 1.4 | n.d. | ||||||||||||||||
| 358 | −19 | 509 | 2.2 | ||||||||||||||||
| 18 238 | a | 202,366 | Del a | −4 | 80.4 | 426 | −4 | 5580 | 6604 | 84.5 | 125,784 | wt | 0 | 79.9 | 377 | 0 | 7954 | 9010 | 88.3 |
| wt | 0 | 14.8 | 430 | 0 | 1024 | 15.5 | Del | −16 | 11.6 | 361 | −16 | 1056 | 11.7 | ||||||
| Del | −6 | 2.8 | n.d. | Del g | −6 | 3.6 | n.d. | ||||||||||||
| Del b | −5 | 1.9 | n.d. | Del | −9 | 2.7 | n.d. | ||||||||||||
| Del h | −13 | 2.2 | n.d. | ||||||||||||||||
| b | 183,884 | Del a | −4 | 78.2 | 426 | −4 | 11,714 | 15,395 | 76.1 | 143,519 | Del i | −18 | 23.1 | 359 | −18 | 4320 | 19,405 | 22.3 | |
| wt | 0 | 19.1 | 430 | 0 | 3149 | 20.5 | Del k | −12 | 22.4 | 365 | −12 | 4453 | 22.9 | ||||||
| Del | −10 | 2.7 | 420 | −10 | 532 | 3.5 | wt | 0 | 19.5 | 377 | 0 | 3917 | 20.2 | ||||||
| Del | −14 | 16.8 | 363 | −14 | 3507 | 18.1 | |||||||||||||
| Del | −20 | 16.2 | 357 | −20 | 3208 | 16.5 | |||||||||||||
| Del h | −13 | 2.0 | n.d. | ||||||||||||||||
| c | 172,597 | Del b | −5 | 40.4 | 425 | −5 | 1545 | 3904 | 39.6 | 155,911 | Del k | −12 | 39.8 | 365 | −12 | 3956 | 5,335 | 74.2 | |
| Del a | −4 | 37.2 | 426 | −4 | 1300 | 33.3 | Del | −12 | 38.9 | ||||||||||
| wt | 0 | 22.4 | 430 | 0 | 1059 | 27.1 | wt | 0 | 19.5 | 377 | 0 | 910 | 17.1 | ||||||
| Del | −8 | 1.8 | 369 | −8 | 150 | 2.8 | |||||||||||||
| 370 | −7 | 173 | 3.2 | ||||||||||||||||
| 371 | −6 | 146 | 2.7 | ||||||||||||||||
| d | 192,845 | Del a | −4 | 63.2 | 426 | −4 | 6797 | 10,654 | 63.8 | 126,005 | Del | −14 | 44.8 | 363 | −14 | 5807 | 13,844 | 41.9 | |
| wt | 0 | 36.8 | 430 | 0 | 3857 | 36.2 | Del | −4 | 30.6 | 373 | −4 | 4289 | 31.0 | ||||||
| Del i | −18 | 13.2 | 359 | −18 | 1857 | 13.4 | |||||||||||||
| wt | 0 | 7.7 | 377 | 0 | 1010 | 7.3 | |||||||||||||
| Del | −19 | 2.0 | 358 | −19 | 500 | 3.6 | |||||||||||||
| Del g | −6 | 1.7 | 371 | −6 | 381 | 2.8 | |||||||||||||
| 18 268 | 201,374 | wt | 0 | 93.6 | 430 | 0 | 15,210 | 16,634 | 91.4 | 207,734 | wt | 0 | 92.2 | 377 | 0 | 23,847 | 26,738 | 89.2 | |
| Del | −10 | 4.2 | 420 | −10 | 919 | 5.5 | Del | −38 | 2.2 | 339 | −38 | 743 | 2.8 | ||||||
| Del | −14 | 2.2 | 416 | −14 | 505 | 3.0 | Del | −15 | 2.0 | 362 | −15 | 754 | 2.8 | ||||||
| Del | −5 | 1.8 | 372 | −5 | 737 | 2.8 | |||||||||||||
| Del | −11 | 1.8 | 366 | −11 | 657 | 2.5 | |||||||||||||
| 18 298 | c | 201,668 | Del c | −12 | 43.6 | 418 | −12 | 7547 | 14,362 | 52.5 | 182,157 | Del m | −20 | 50.4 | 357 | −20 | 10,278 | 20,324 | 50.6 |
| wt | 0 | 37.2 | 430 | 0 | 6815 | 47.5 | Del o | −9 | 49.6 | 368 | −9 | 10,046 | 49.4 | ||||||
| Art e | −165 | 17.0 | n.d. | ||||||||||||||||
| Art f | −172 | 2.3 | n.d. | ||||||||||||||||
| d | 204,297 | Del c | −12 | 52.0 | 418 | −12 | 9693 | 18,744 | 51.7 | 182,047 | Del m | −20 | 51.6 | 357 | −20 | 12,387 | 24,866 | 49.8 | |
| Del d | −4 | 48.0 | 426 | −4 | 9051 | 48.3 | Del o | −9 | 48.4 | 368 | −9 | 12,479 | 50.2 | ||||||
| e | 210,743 | Del c | −12 | 50.8 | 418 | −12 | 4822 | 9280 | 52.0 | 188,583 | Del m | −20 | 52.7 | 357 | −20 | 7393 | 14,335 | 51.6 | |
| wt | wt | 28.5 | 430 | 0 | 3066 | 33.0 | Del o | −9 | 47.3 | 368 | −9 | 6942 | 48.4 | ||||||
| Del | −7 | 10.3 | 423 | −7 | 1392 | 15.0 | |||||||||||||
| Art e | −165 | 8.2 | n.d. | ||||||||||||||||
| Art f | −172 | 2.3 | n.d. | ||||||||||||||||
| i | 221,032 | Del c | −12 | 49.7 | 418 | −12 | 9484 | 19,358 | 49.0 | 197,422 | Del m | −20 | 48.9 | 357 | −20 | 13,184 | 27,323 | 48.3 | |
| wt | 0 | 45.1 | 430 | 0 | 9175 | 47.4 | Del o | −9 | 46.2 | 368 | −9 | 12,774 | 46.8 | ||||||
| Del d | −4 | 3.1 | 426 | −4 | 699 | 3.6 | wt | 0 | 4.9 | 377 | 0 | 1365 | 5.0 | ||||||
| Art e | −165 | 2.0 | n.d. | ||||||||||||||||
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Schröpfer, S.; Flachowsky, H. Tracing CRISPR/Cas12a Mediated Genome Editing Events in Apple Using High-Throughput Genotyping by PCR Capillary Gel Electrophoresis. Int. J. Mol. Sci. 2021, 22, 12611. https://doi.org/10.3390/ijms222212611
Schröpfer S, Flachowsky H. Tracing CRISPR/Cas12a Mediated Genome Editing Events in Apple Using High-Throughput Genotyping by PCR Capillary Gel Electrophoresis. International Journal of Molecular Sciences. 2021; 22(22):12611. https://doi.org/10.3390/ijms222212611
Chicago/Turabian StyleSchröpfer, Susan, and Henryk Flachowsky. 2021. "Tracing CRISPR/Cas12a Mediated Genome Editing Events in Apple Using High-Throughput Genotyping by PCR Capillary Gel Electrophoresis" International Journal of Molecular Sciences 22, no. 22: 12611. https://doi.org/10.3390/ijms222212611
APA StyleSchröpfer, S., & Flachowsky, H. (2021). Tracing CRISPR/Cas12a Mediated Genome Editing Events in Apple Using High-Throughput Genotyping by PCR Capillary Gel Electrophoresis. International Journal of Molecular Sciences, 22(22), 12611. https://doi.org/10.3390/ijms222212611






