Rapid and Visual Detection of Heterodera schachtii Using Recombinase Polymerase Amplification Combined with Cas12a-Mediated Technology
Abstract
:1. Introduction
2. Results
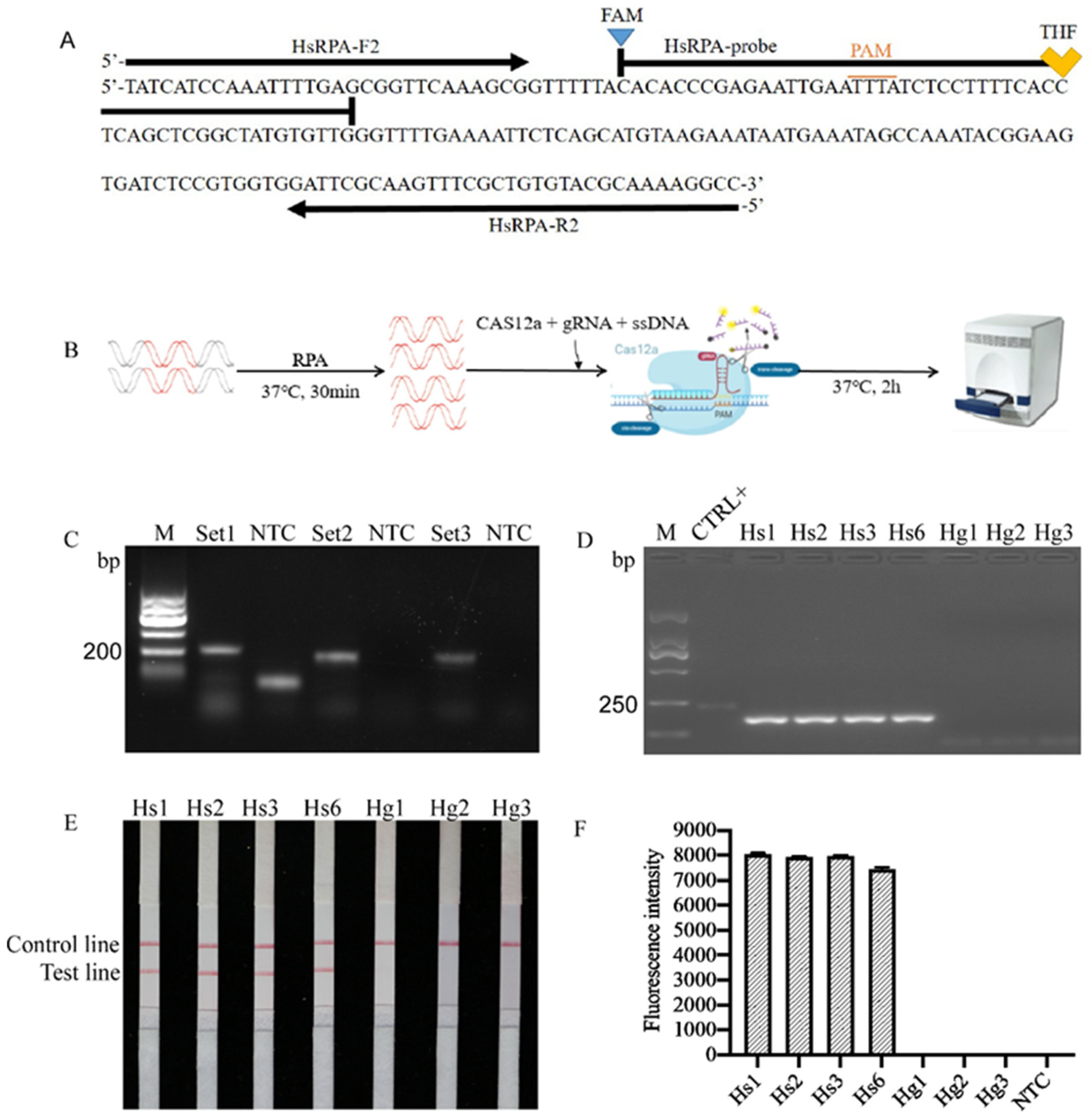
2.1. Primer Design
2.2. Detection and Confirmation of RPA Products
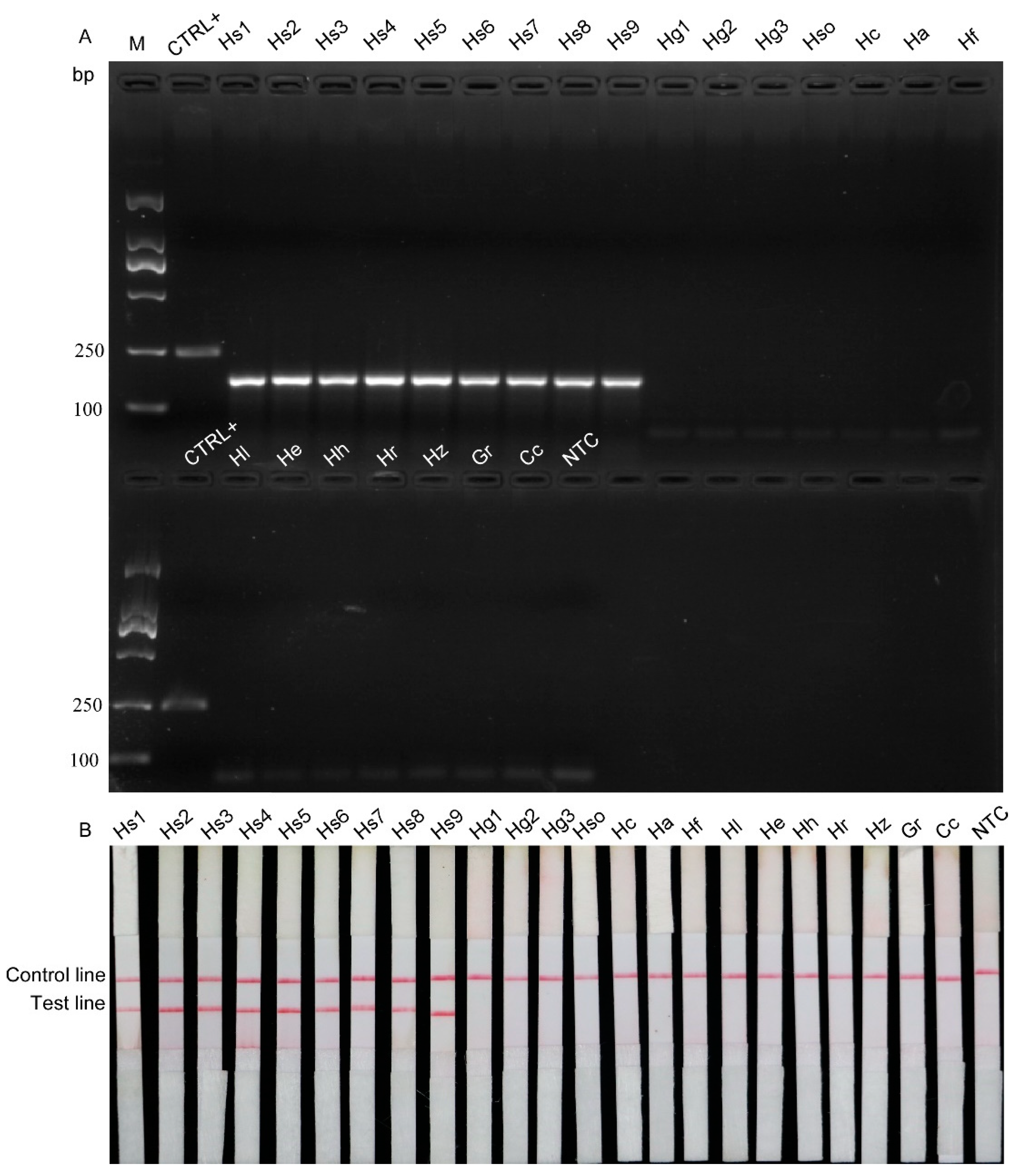
2.3. Specificity Test
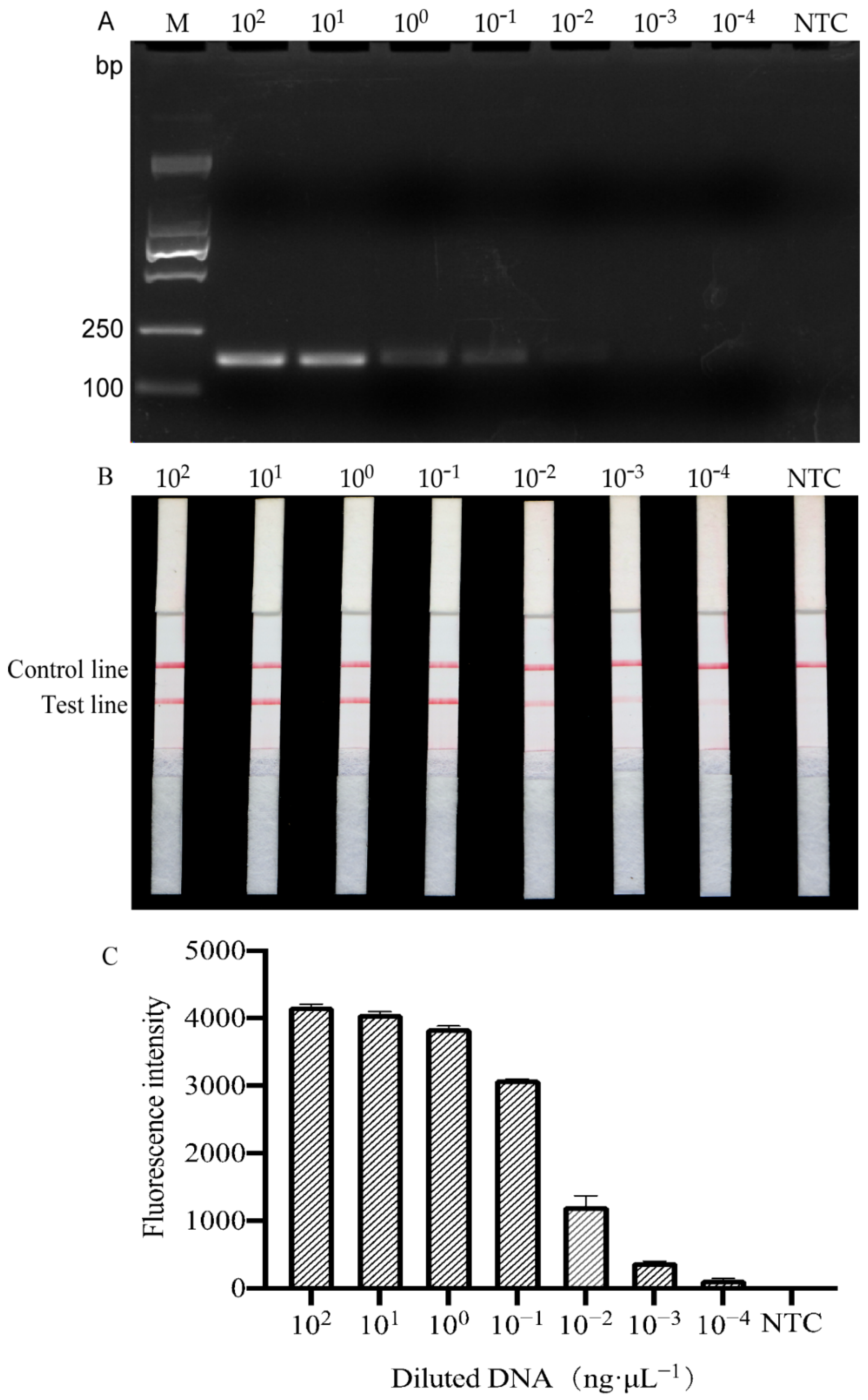
2.4. Sensitivity Test
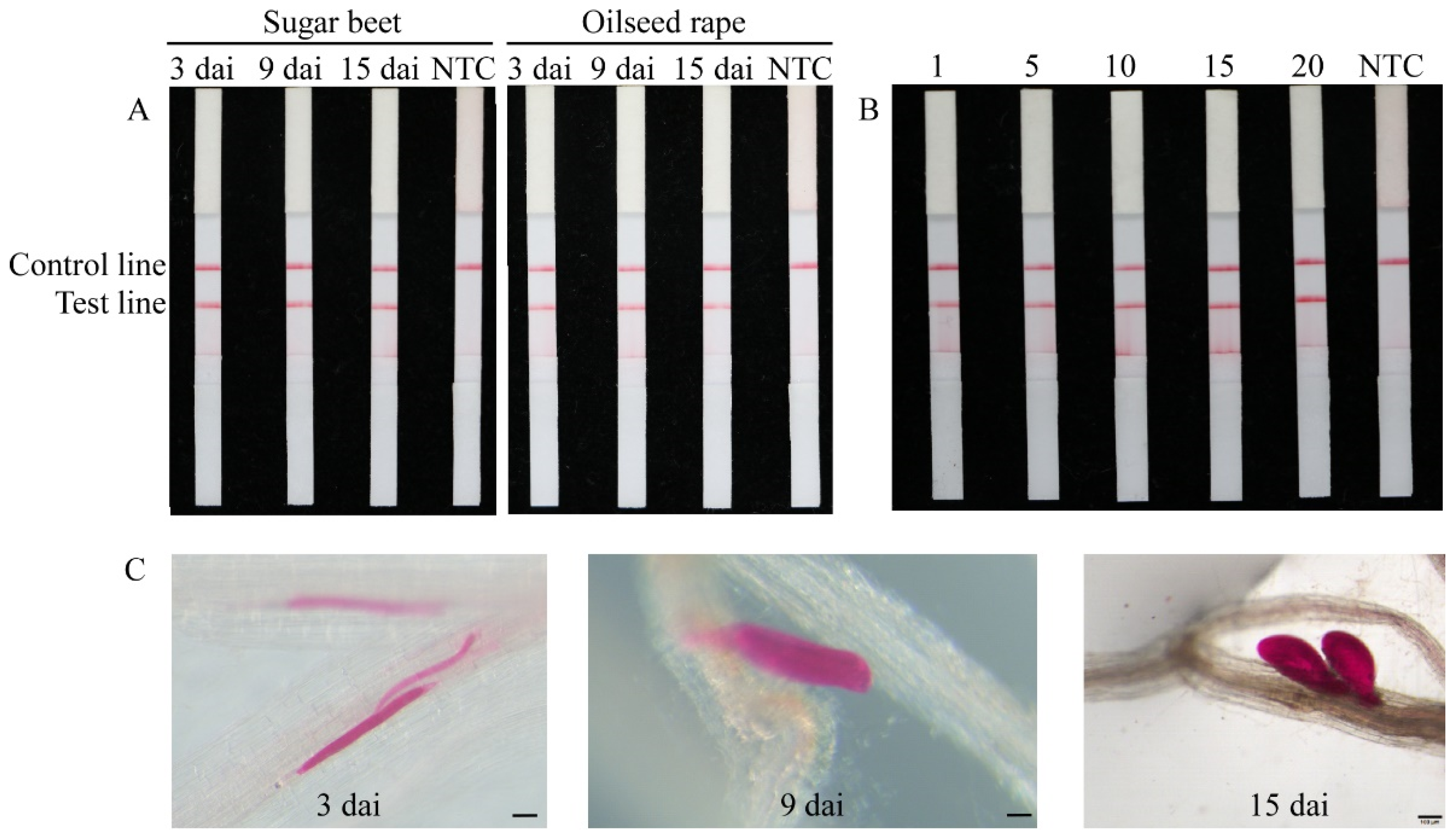
2.5. Direct Detection of Heterodera schachtii in Artificially Inoculated Roots
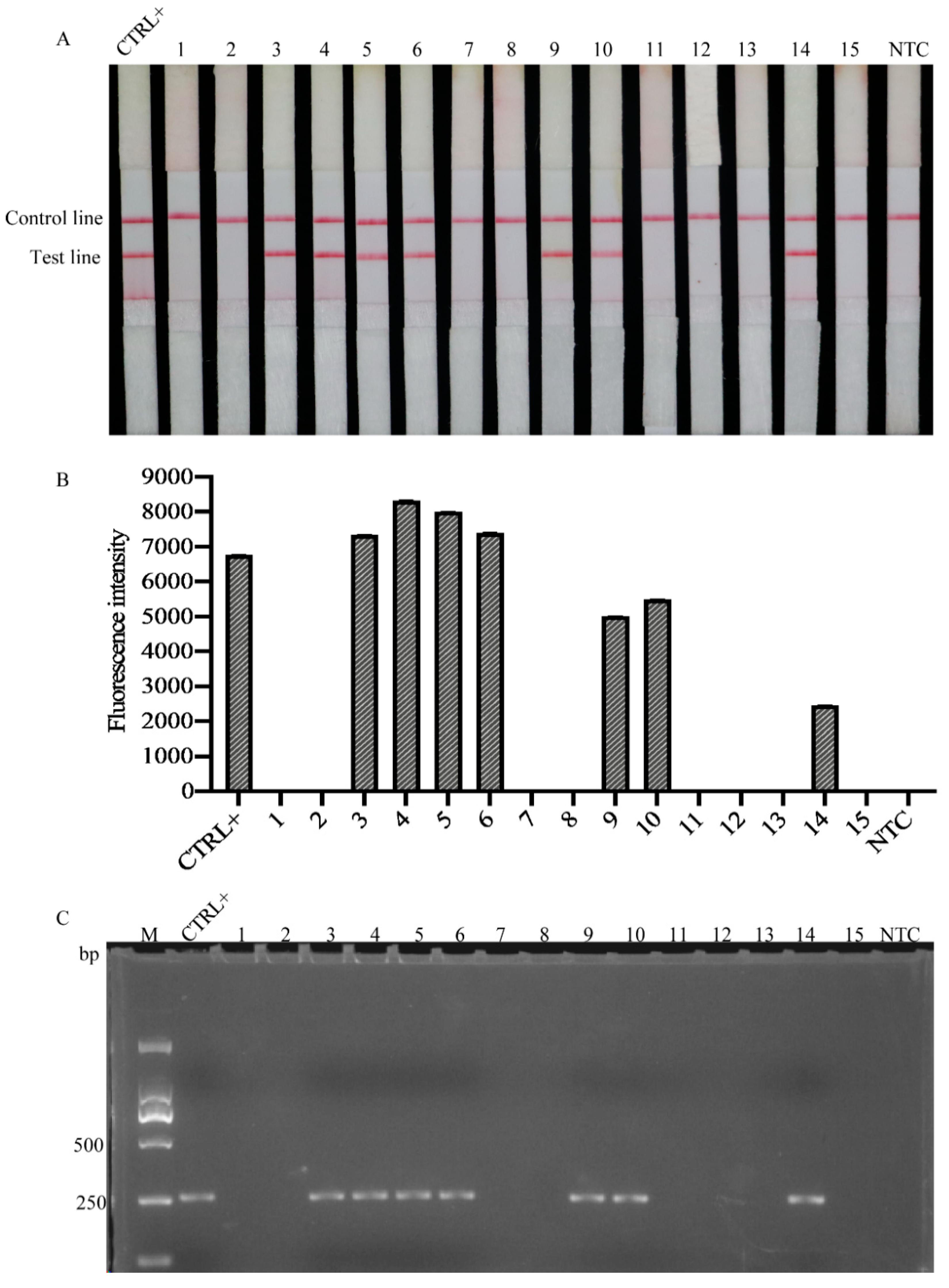
2.6. Detection of Heterodera schachtii in Natural Field Samples
3. Discussion
4. Materials and Methods
4.1. Nematode Populations
4.2. DNA Extraction
4.3. RPA Primer and Probe Design
4.4. RPA Reaction
4.5. Lateral Flow Dipstick (LFD) Assay
4.6. RPA/Cas12a Detection
4.7. RPA Assay Specificity Test
4.8. RPA Assay Sensitivity Test
4.9. Direct Detection of Heterodera schachtii in Inoculated Plant Roots and Soil
4.10. Detection of Heterodera schachtii in Natural Field Roots
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Abad, P.; Gouzy, J.; Aury, J.M. Genome sequence of the metazoan plant-parasitic nematode Meloidogyne incognita. Nat. Biotechnol. 2008, 26, 909–915. [Google Scholar] [CrossRef] [Green Version]
- Jones, J.; Haegeman, A. Top 10 plant-parasitic nematodes in molecular plant pathology. Mol. Plant Pathol. 2013, 14, 946–961. [Google Scholar] [CrossRef] [PubMed]
- Muller, J. The economic importance of Heterodera schachtii in Europe. Helminthologia 1999, 36, 205–213. [Google Scholar]
- Subbotin, S.A.; Waeyenberge, L.; Moens, M. Identification of cyst forming nematodes of the genus Heterodera (Nematoda: Heteroderidae) based on the ribosomal DNA-RFLP. Nematology 2000, 2, 153–164. [Google Scholar] [CrossRef] [Green Version]
- Caswell-Chen, E.P.; Williamson, V.M.; Wu, F.F. Random amplified polymorphic DNA analysis of Heterodera cruciferae and H. schachtii populations. J. Nematol. 1992, 24, 343–351. [Google Scholar] [PubMed]
- Madani, M.; Kyndt, N.T.; Colpaert, S.A.; Subbotin, G.; Moens, M. Polymorphism among sugar beet cyst nematode Heterodera schachtii populations as inferred from AFLP and ITS rDNA gene analyses. Russ. J. Nematol. 2007, 15, 117–128. [Google Scholar]
- Moens, M.; Subbotin, S.; Maafi, Z.T. Molecular identification of cyst-forming nematodes (Heteroderidae) from Iran and a phylogeny based on ITS-rDNA sequences. Nematology 2003, 5, 99–111. [Google Scholar]
- Amiri, S.; Subbotin, S.A.; Moens, M. Identification of the beet cyst nematode Heterodera schachtii by PCR. Eur. J. Plant Pathol. 2002, 108, 497–506. [Google Scholar] [CrossRef]
- Madani, M.; Subbotin, S.A.; Moens, M. Quantitative detection of the potato cyst nematode, Globodera pallida, and the beet cyst nematode, Heterodera schachtii, using Real-Time PCR with SYBR green I dye. Mol. Cell. Probes 2005, 19, 81–86. [Google Scholar] [CrossRef]
- Baidoo, R.; Yan, G.P. Developing a real-time PCR assay for direct identification and quantification of soybean cyst nematode, Heterodera glycines, in soil and its discrimination from sugar beet cyst nematode, Heterodera schachtii. Plant Dis. 2021. [Google Scholar] [CrossRef] [PubMed]
- Dhama, K.; Karthik, K.; Chakraborty, S.; Tiwari, R.; Kapoor, S.; Kumar, A. Loop-mediated isothermal amplification of DNA (lamp): A new diagnostic tool lights the world of diagnosis of animal and human pathogens: A review. Pak. J. Biol. 2014, 17, 151–166. [Google Scholar] [CrossRef] [Green Version]
- Notomi, T.; Okayama, H.; Masubuchi, H. Loop-mediated isothermal amplification of DNA. Nucleic Acids Res. 2000, 28, e63. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Piepenburg, O.; Williams, C.H.; Stemple, D.L. DNA Detection Using Recombination Proteins. PLoS Biol. 2006, 4, 1115–1121. [Google Scholar] [CrossRef]
- Lobato, I.M.; O’Sullivan, C.K. Recombinase polymerase amplification: Basics, applications and recent advances. TrAC Trend. Anal. Chem. 2018, 98, 19–35. [Google Scholar] [CrossRef]
- Londoño, M.A.; Harmon, C.L.; Polston, J.E. Evaluation of recombinase polymerase amplification for detection of begomoviruses by plant diagnostic clinics. Virol. J. 2016, 13, 48. [Google Scholar] [CrossRef] [Green Version]
- Song, Z.; Yang, X.; Zhang, X.; Luan, M.; Mei, S. Rapid and visual detection of Meloidogyne Hapla using recombinase polymerase amplification combined with a lateral flow dipstick (RPA-LFD) assay. Plant Dis. 2020, 7, 448–453. [Google Scholar]
- Ju, Y.L.; Lin, Y.; Yang, G.G.; Wu, H.P.; Pan, Y.M. Development of recombinase polymerase amplification assay for rapid detection of Meloidogyne incognita, M. javanica, M. arenaria, and M. enterolobii. Eur. J. Plant Pathol. 2019, 155, 1155–1163. [Google Scholar] [CrossRef]
- Chi, Y.K.; Zhao, W.; Ye, M.D.; Ali, F.; Wang, T.; Qi, R.D. Evaluation of recombinase polymerase amplification assay for detecting Meloidogyne javanica. Plant Dis. 2020, 104, 801–807. [Google Scholar] [CrossRef]
- Subbotin, S.A. Recombinase polymerase amplification assay for rapid detection of the root-knot nematode Meloidogyne enterolobii. Nematology 2019, 21, 243–251. [Google Scholar] [CrossRef] [Green Version]
- Cha, D.J.; Kim, D.S.; Lee, S.K.; Han, H.R. A new on-site detection method for Bursaphelenchus xylophilus in infected pine trees. For. Pathol. 2019, 49, e12503. [Google Scholar] [CrossRef]
- Cha, D.; Kim, D.; Choi, W.; Park, S.; Han, H. Point-of-care diagnostic (POCD) method for detecting Bursaphelenchus xylophilus in pinewood using recombinase polymerase amplification (RPA) with the portable optical isothermal device (POID). PLoS ONE 2020, 15, e0227476. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chen, J.S.; Ma, E.; Harrington, L.B.; Da Costa, M.; Tian, X.; Palefsky, J.M. Crispr-cas12a target binding unleashes indiscriminate single-stranded dnase activity. Science 2018, 360, 436–439. [Google Scholar] [CrossRef] [Green Version]
- Gootenberg, J.S.; Abudayyeh, O.O.; Kellner, M.J.; Joung, J.; Collins, J.J.; Zhang, F. Multiplexed and portable nucleic acid detection platform with cas13, cas12a, and csm6. Science 2018, 360, 439–444. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zetsche, B.; Gootenberg, J.; Abudayyeh, O.; Slaymaker, I.; Makarova, K.; Essletzbichler, P. Cpf1 is a single rna-guided endonuclease of a class 2 crispr-cas system. Cell 2015, 163, 759–771. [Google Scholar] [CrossRef] [Green Version]
- Hua, L.; Jwa, B.; Hza, B.; Xl, C.; Jiang, W.; Yu, W.C. RPA-Cas12a-FS: A frontline nucleic acid rapid detection system for food safety based on CRISPER-Cas12a combined with recombinase polymerase amplification. Food Chem. 2020, 334, 127608. [Google Scholar]
- Rashid, A.; Ahmed, M.; Tin, M.; Norhan, H.; Magdy, M.M. Efficient, rapid, and sensitive detection of plant RNA viruses with one-pot RT-RPA–CRISPR/Cas12a assay. Front. Microbiol. 2020, 11, 610872. [Google Scholar]
- Ren, M.; Mei, H.; Zhou, J.; Zhou, M.; Zhao, L. Early diagnosis of rabies virus infection by RPA-CRISPR techniques in a rat model. Arch. Virol. Suppl. 2021, 166, 1083–1092. [Google Scholar] [CrossRef]
- Hou, T.; Zeng, W.; Yang, M.; Chen, W.; Xu, T. Development and evaluation of a rapid CRISPR-based diagnostic for COVID-19. PLoS Pathog. 2020, 16, e1008705. [Google Scholar] [CrossRef] [PubMed]
- Kang, H.X.; Peng, Y.; Hua, K.Y.; Deng, Y.F. Rapid detection of wheat blast pathogen Magnaporthe Oryzae Triticum pathotype using genome-specific primers and cas12a-mediated technology. Engineering 2021, 7, 1326–1335. [Google Scholar] [CrossRef]
- Samuelian, S.; Kleine, M.; Ruyter-Spira, C.P.; Klein-Lankhorst, R.M.; Jung, C. Cloning and functional analyses of a gene from sugar beet up-regulated upon cyst nematode infection. Plant Mol. Biol. 2004, 54, 147–156. [Google Scholar] [CrossRef]
- Cui, J.K.; Erginbas-Orakci, G.; Peng, H.; Huang, W.K.; Liu, S.M.; Qiao, F.; Elekcioglu, H.; Imren, M.; Dababat, A.A.; Peng, D.L. First report of sugar beet nematode, Heterodera schachtii Schmidt, 1871 (Nemata: Heteroderidae) in sugar beet growing areas of Sanliurfa, Turkey. Turk. J. Entomol. 2016, 40, 303–314. [Google Scholar] [CrossRef]
- Ou, S.Q.; Peng, D.L.; Liu, X.M.; Li, Y.; Moens, M. Identification of Heterodera glycines using PCR with sequence characterised amplified region (SCAR) primers. Nematology 2008, 10, 397–403. [Google Scholar]
- Zhen, H.Y.; Peng, H.; Kong, L.A.; Hong, B.Y.; Zhu, G.L.; Wang, R.H.; Peng, D.L.; Wen, Y.H. Heterodera sojae, a new cyst nematode record in China and its parasitism to Legume crops. Sci. Agric. Sin. 2018, 51, 2913–2924. (In Chinese) [Google Scholar]
- Holgado, R. Cereal Cyst Nematodes Heterodera spp. In Norway: Studies on Occurrence, Species, Pathotypes and Antagonists; Agricultural University of Norway: Aas, Norway, 2003; Available online: https://www.plantwise.org/KnowledgeBank/pestalert?pan=20043055141./ (accessed on 1 August 2021).
- Peng, D.L.; Ye, W.X.; Peng, H.; Gu, X.C. First report of the cyst nematode (Heterodera filipjevi) on wheat in Henan Province, China. Plant Dis. 2010, 94, 1262. [Google Scholar] [CrossRef]
- Franklin, M.T. Heterodera latipons n. sp. a cereal cyst nematode from the Mediterranean region. Nematologic 1969, 15, 535–542. [Google Scholar] [CrossRef] [Green Version]
- Ding, Z.; Namphueng, J.; He, X.F.; Peng, D.L.; Huang, W.K. First report of the cyst nematode (Heterodera elachista) on Rice in Hunan Province, China. Plant Dis. 2012, 96, 151. [Google Scholar] [CrossRef]
- Wu, H.Y.; Qiu, Z.Q.; Mo, A.S.; Li, J.Q.; Peng, D.L. First report of Heterodera zeae on maize in China. Plant Dis. 2017, 101, 1330. [Google Scholar] [CrossRef]
- Ge, J.J.; Cao, A.X.; Chen, H.J.; Marice, M. Real time PCR assay of the golden nematode, Globodera rostochiensis with TaqMan probe. Plant Prot. 2009, 35, 105–109. (In Chinese) [Google Scholar]
- Duan, Y.X.; Wang, D.; Chen, L.J. First report of the Cactus Cyst Nematode, Cactodera cacti, on Cactus in Northern China. Plant Dis. 2012, 96, 1385. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.G.; Ni, T.; Kim, J.Y.; Chun, E.M.; Kern, A.; Park, J.K. Phytogeography of the highly invasive sugar beet nematode, Heterodera schachtii (Schmidt, 1871), based on microsatellites. Evol. Appl. 2019, 12, 324–336. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Subbotin, S.A.; Mundoocampo, M.; Baldwin, J.G. Systematics of Cyst Nematodes (Nematoda: Heteroderinae), Volume 8B; Plant Pathology; Brill: Leiden, The Netherlands; Academic Press: Cambridge, MA, USA, 2010; pp. 35–42. [Google Scholar]
- Peng, H.; Liu, H.; Gao, L.; Jiang, R.; Li, G.K.; Gao, H.F.; Wu, W.; Wang, J.; Zhang, Y.; Huang, W.K.; et al. Identification of Heterodera schachtii on sugar beet in Xinjiang Uygur Autonomous Region of China. J. Integr. Agric. 2021. [Google Scholar] [CrossRef]
- Li, T.T.; Wang, J.L.; Zhang, N.Z.; Li, W.H.; Yan, H.B.; Li, L. Rapid and visual detection of Trichinella spp. using a lateral flow strip-based recombinase polymerase amplification (lf-rpa) assay. Front. Cell. Infect. Microbiol. 2019, 9, 1. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Radice, A.D.; Powers, T.O.; Sandall, L.J. Comparisons of Mitochondrial DNA from the Sibling Species Heterodera glycines and H. schachtii. J. Nematol. 1988, 20, 443–450. [Google Scholar]
- Cao, J.; Yao, Y.; Fan, K.; Tan, G.; Xiang, W.; Xia, X.; Zhang, L. Harnessing a previously unidentified capability of bacterial allosteric transcription factors for sensing diverse small molecules in vitro. Sci. Adv. 2018, 4, eaau4602. [Google Scholar] [CrossRef] [Green Version]
- Chai, Z.; Ma, W.; Fu, F.; Lang, Y.; Wang, W.; Tong, G.; Li, X. A SYBR Greenbased real-time RT-PCR assay for simple and rapid detection and differentiation of highly pathogenic and classical type 2 porcine reproductive and respiratory syndrome virus circulating in China. Arch. Virol. 2013, 158, 407–415. [Google Scholar] [CrossRef] [PubMed]
- Isabel, S.; Saifuddin, F.A.; Rybarski, J.R.; Finkelstein, I.J.; Rick, R. Kinetic basis for DNA target specificity of crispr-cas12a. Mol. Cell 2018, 71, 816–824. [Google Scholar]
- Subbotin, S.A.; Burbridge, J. Sensitive, accurate and rapid detection of the northern root-knot nematode, Meloidogyne hapla, using recombinase polymerase amplification assays. Plants 2021, 10, 336. [Google Scholar] [CrossRef] [PubMed]
- Tefft, P.M.; Bone, L.W. Zinc-mediated hatching of eggs of soybean cyst nematode, Heterodera glycines. J. Chem. Ecol. 1984, 10, 361–372. [Google Scholar] [CrossRef]
- Byrd, D.W., Jr.; Kirkpatrick, T.; Barker, K.R. An improved technique for clearing and staining plant tissues for detection of nematodes. J. Nematol. 1983, 15, 142–143. [Google Scholar]

| Primers | Sequences | Purpose | References |
|---|---|---|---|
| HsRPA-F1 | 5′-GAGAATTGAATTTATCTCCTTTTCACCTCAGCTC-3′ | RPA amplification | This study |
| HsRPA-R1 | 5′-CACAGAGACAACACGAAGGAGCGAGCTCAATG-3′ | ||
| HsRPA-F2 | 5′-TATCATCCAAATTTTGAGCGGTTCAAAGCG-3′ | ||
| HsRPA-R2 | 5′-GGCCTTTTGCGTACACAGCGAAACTTGCGAATC-3′ | ||
| HsRPA-F3 | 5′-CAACAACTACCACCACAATTACCGAACCACCG-3′ | ||
| HsRPA-R3 | 5′-CAACTCCTTCAATGATTTGTCTAAGAATTCTC-3′ | ||
| ssDNA | 5′-FAMTTTTT-3′BHQ1 | Cas12a degradation | |
| Hs-probe | 5′-FAMCACCCGAGAATTGAATTTATCTCCTTTTCAC(THF)TCAGCTCGGCTATGTGTTG-3′ | ||
| Hs-gRNA | 5′-AAUUUCUACUGUUGUAGAUGAAUUUAUCUCCUUUUCACCUCA-3′ | ||
| SHF6 | 5′-GTTCTTACGTTACTTCCA-3′ | PCR amplification | [8] |
| rDNA2 | 5′-TTTCACTCGCCGTTACTAAGG-3′ |
| Species Code | Species | Population Origin | Host | Sampling Date | References |
|---|---|---|---|---|---|
| Hs1 | H. schachtii | Germany | Sugar beet | - a | [30] |
| Hs2 | H. schachtii | Belgium | Sugar beet | - a | [9] |
| Hs3 | H. schachtii | Bozova, anliurfa Province, Turkey | Sugar beet | - | [31] |
| Hs4 | H. schachtii | Karakprü anliurfa Province,Turkey | Sugar beet | - | [31] |
| Hs5 | H. schachtii | Siverek, anliurfa Province, Turkey | Sugar beet | - | [31] |
| Hs6 | H. schachtii | Xinyuan county Xinjiang, China | Sugar beet | Sept. 2019 | This study |
| Hs7 | H. schachtii | Xinyuan county Xinjiang, China | Sugar beet | Sept. 2019 | This study |
| Hs8 | H. schachtii | Xinyuan county Xinjiang, China | Sugar beet | Sept. 2019 | This study |
| Hs9 | H. schachtii | Emin county, Xinjiang, China | Sugar beet | Sept. 2019 | This study |
| Hg1 | H. glycines | Langfang, Hebei, China | Soybean | Oct. 2019 | [32] |
| Hg2 | H. glycines | Heilongjiang, China | Soybean | Oct. 2019 | [32] |
| Hg3 | H. glycines | Japan | - | - b | This study |
| Hso | H. sojae | Jiangxi, China | Soybean | Sept. 2017 | [33] |
| Hc | H. cruciferae | Hubei, China | Oilseed rape | Sept. 2017 | This study |
| Ha | H. avenae | Norway | Wheat | Sept. 2016 | [34] |
| Hf | H. filipjevi | Tangyin. Henan, China | Wheat | July 2019 | [35] |
| Hl | H. latinpones | Belgium | Wheat | - a | [36] |
| He | H. elachista | Hunan, China | Rice | Dec. 2016 | [37] |
| Hh | H. humuli | Guizhou, China | Weeds | July 2018 | This study |
| Hr | H. ripae | Guizhou, China | Weeds | July 2018 | This study |
| Hz | H. zeae | Laibin, Guangxi, China | Maize | Mar. 2018 | [38] |
| Gr | Globodera rostochiensis | Belgium | Potato | - a | [39] |
| Cc | Cactoderacacti | Liaoning, China | Weeds | July 2018 | [40] |
| Samples | Host | Location | Cyst Nematode Density a | Detection Results | ||
|---|---|---|---|---|---|---|
| RPA/LFD (Detections/Trials) | RPA/CRISPR (Detections/Trial) | PCR | ||||
| 1 | Sugar beet | Huocheng County, Xinjiang, China | 0 | 0/3 | 0/3 | − |
| 2 | Zhaosu County, Xinjiang, China | 0 | 0/3 | 0/3 | − | |
| 3 | Xinyuan county-1, Xinjiang, China | 22 ± 7 | 3/3 | 3/3 | + | |
| 4 | Xinyuan county-2, Xinjiang, China | 46 ± 4 | 3/3 | 3/3 | + | |
| 5 | Xinyuan county-3, Xinjiang, China | 38 ± 5 | 3/3 | 3/3 | + | |
| 6 | Xinyuan county-4, Xinjiang, China | 20 ± 5 | 3/3 | 3/3 | + | |
| 7 | Gongliu county, Xinjiang, China | 0 | 0/3 | 0/3 | − | |
| 8 | Gongliu county, Xinjiang, China | 0 | 0/3 | 0/3 | − | |
| 9 | Emin county, Xinjiang, China | 11 ± 4 | 3/3 | 3/3 | + | |
| 10 | Emin county, Xinjiang, China | 19 ± 4 | 3/3 | 3/3 | + | |
| 11 | Keyouqian Banner, Inner Mongolia, China | 0 | 0/3 | 0/3 | − | |
| 12 | Urad Front Banner, Inner Mongolia, China | 0 | 0/3 | 0/3 | − | |
| 13 | Linxi county, Inner Mongolia, China | 0 | 0/3 | 0/3 | − | |
| 14 | Zhangbei county, Hebei province, China | 5 ± 2 | 3/3 | 3/3 | + | |
| 15 | Kangbao county, Hebei province, China | 0 | 0/3 | 0/3 | − | |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yao, K.; Peng, D.; Jiang, C.; Zhao, W.; Li, G.; Huang, W.; Kong, L.; Gao, H.; Zheng, J.; Peng, H. Rapid and Visual Detection of Heterodera schachtii Using Recombinase Polymerase Amplification Combined with Cas12a-Mediated Technology. Int. J. Mol. Sci. 2021, 22, 12577. https://doi.org/10.3390/ijms222212577
Yao K, Peng D, Jiang C, Zhao W, Li G, Huang W, Kong L, Gao H, Zheng J, Peng H. Rapid and Visual Detection of Heterodera schachtii Using Recombinase Polymerase Amplification Combined with Cas12a-Mediated Technology. International Journal of Molecular Sciences. 2021; 22(22):12577. https://doi.org/10.3390/ijms222212577
Chicago/Turabian StyleYao, Ke, Deliang Peng, Chen Jiang, Wei Zhao, Guangkuo Li, Wenkun Huang, Lingan Kong, Haifeng Gao, Jingwu Zheng, and Huan Peng. 2021. "Rapid and Visual Detection of Heterodera schachtii Using Recombinase Polymerase Amplification Combined with Cas12a-Mediated Technology" International Journal of Molecular Sciences 22, no. 22: 12577. https://doi.org/10.3390/ijms222212577
APA StyleYao, K., Peng, D., Jiang, C., Zhao, W., Li, G., Huang, W., Kong, L., Gao, H., Zheng, J., & Peng, H. (2021). Rapid and Visual Detection of Heterodera schachtii Using Recombinase Polymerase Amplification Combined with Cas12a-Mediated Technology. International Journal of Molecular Sciences, 22(22), 12577. https://doi.org/10.3390/ijms222212577








