Establishment of an Efficient Immortalization Strategy Using HMEJ-Based bTERT Insertion for Bovine Cells
Abstract
:1. Introduction
2. Results
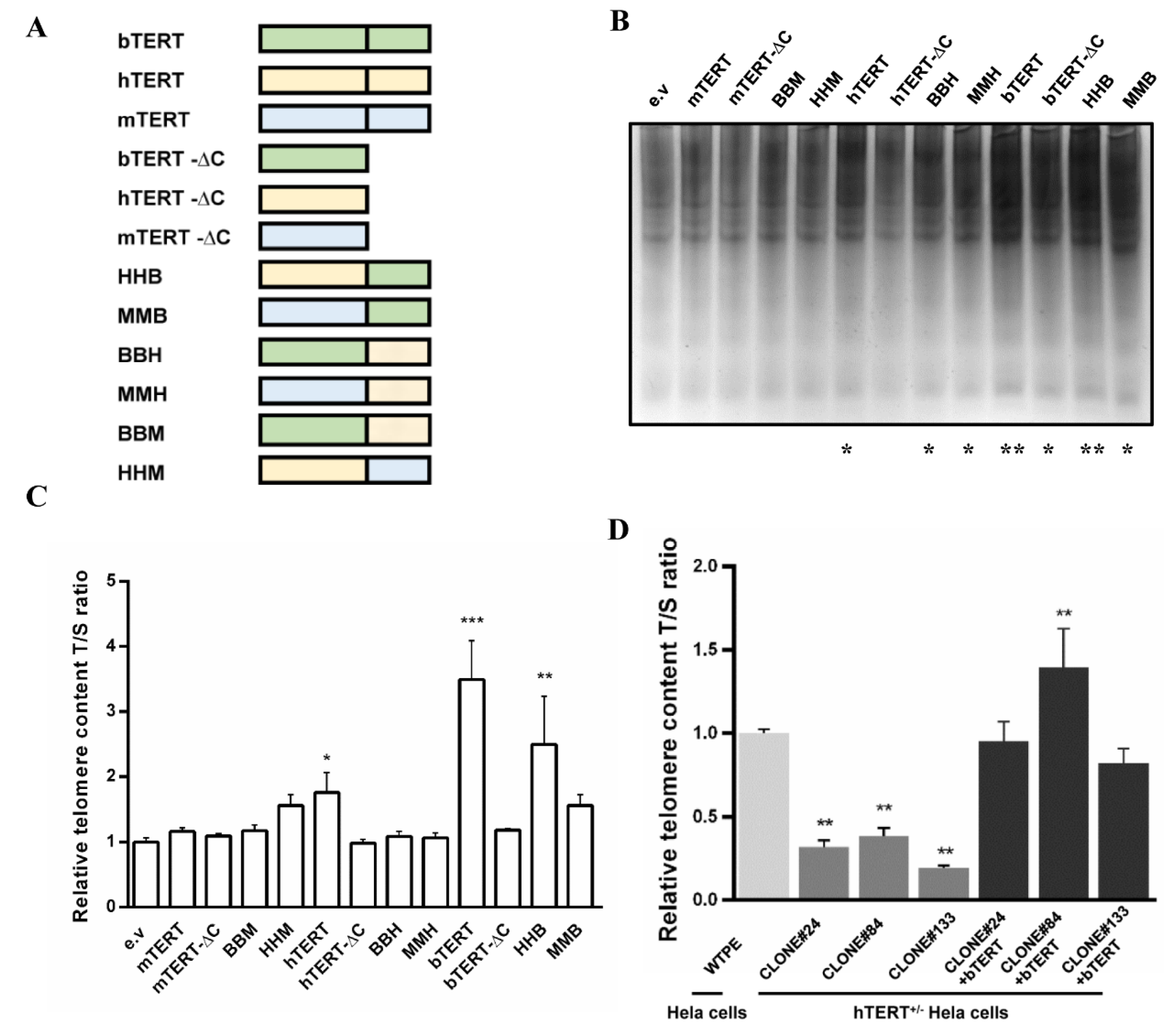
2.1. bTERT Is a Potent Factor for Immortalization
2.2. HMEJ-Based Site-Specific bTERT Insertion Enables Efficient Immortalization of Bovine Cells
2.3. Characteristics of Immortalized Bovine Cells Established via HMEJ-Based bTERT Integration
2.4. bTERT-Driven Immortalization via the CRISPR/Cas9 System Inhibited by the Post-Translational Modification in bBMMs
3. Discussion
4. Materials and Methods
4.1. Cell Isolation and Culture
4.2. Construction of Vectors
4.3. Cell Transfection
4.4. Reverse Transcription PCR and Quantitative RT-PCR
4.5. Western Blot
4.6. Establishment of Immortalized Bovine BFFs and bBMMs Using the HMEJ-Based CRISPR/Cas9-Mediated bTERT Expression at ROSA26 Locus
4.7. RNA Binding Protein Immunoprecipitation Assay
4.8. Telomerase Activity Assay
4.9. Relative Telomere Length Measurement by Quantitative Real-Time PCR
4.10. EdU(5′-ethynyl-2′-deoxyuridine) Incorporation Assay
4.11. Detection of Individual Colonies by PCR
4.12. Off-Target Analyses
4.13. Soft Agar Assay for Colony Formation
4.14. Karyotype Analysis
4.15. Senescence-Associated β-Galactosidase Staining
4.16. Cellular Proliferation Assay
4.17. Cell Cycle Analysis
4.18. FACS Analyses
4.19. Immunofluorescence Staining
4.20. Preparation of Cell Lysates
4.21. Co-Immunoprecipitation
4.22. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
References
- Zhan, K.; Gong, X.; Chen, Y.; Jiang, M.; Yang, T.; Zhao, G. Short-Chain Fatty Acids Regulate the Immune Responses via G Protein-Coupled Receptor 41 in Bovine Rumen Epithelial Cells. Front. Immunol. 2019, 10, 2042. [Google Scholar] [CrossRef]
- Lagrée, A.-C.; Fasani, F.; Rouxel, C.; Pivet, M.; Pourcelot, M.; Fablet, A.; Romey, A.; Caignard, G.; Vitour, D.; Blaise-Boisseau, S.; et al. Bovine Organospecific Microvascular Endothelial Cell Lines as New and Relevant In Vitro Models to Study Viral Infections. Int. J. Mol. Sci. 2020, 21, 5249. [Google Scholar] [CrossRef]
- Mao, R.; Sun, D.; Yang, F.; Tian, H.; Zhu, Z.; Zheng, H.; Liu, X. Establishment and Evaluation of a Stable Bovine Thyroid Cell Line for Investigating Foot-and-Mouth Disease Virus. Front. Microbiol. 2018, 9, 2149. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, J.-X.; Said, A.; Ge, X.-G.; Wang, W.; Zhang, Y.; Jin, T. Development and validation of immortalized bovine mammary epithelial cell line as an in vitro model for the study of mammary gland functions. Cytotechnology 2017, 70, 67–82. [Google Scholar] [CrossRef] [PubMed]
- Barthel, R.; Feng, J.; Piedrahita, J.A.; McMurray, D.N.; Templeton, J.W.; Adams, L.G. Stable transfection of the bovine NRAMP1 gene into murine RAW264.7 cells: Effect on Brucella abortus survival. Infect. Immun. 2001, 69, 3110–3119. [Google Scholar] [CrossRef] [Green Version]
- Lin, X.; Wei, M.; Song, F.; Xue, D.I.; Wang, Y. N-acetylcysteine (NAC) Attenuating Apoptosis and Autophagy in RAW264.7 Cells in Response to Incubation with Mycolic Acid from Bovine Mycobacterium tuberculosis Complex. Pol. J. Microbiol. 2020, 69, 223–229. [Google Scholar] [CrossRef] [PubMed]
- Stepanenko, A.A.; Kavsan, V.M. Immortalization and malignant transformation of eukaryotic cells. Tsitol. Genet. 2012, 46, 36–75. [Google Scholar] [CrossRef] [PubMed]
- Katwal, P.; Thomas, M.; Uprety, T.; Hildreth, M.B.; Kaushik, R.S. Development and biochemical and immunological characterization of early passage and immortalized bovine intestinal epithelial cell lines from the ileum of a young calf. Cytotechnology 2019, 71, 127–148. [Google Scholar] [CrossRef] [Green Version]
- Chiou, H.C.; Vasu, S.; Liu, C.Y.; Cisneros, I.; Jones, M.B.; Zmuda, J.F. Scalable transient protein expression. Methods Mol. Biol. 2014, 1104, 35–55. [Google Scholar]
- Tenenbaum, L.; Lehtonen, E.; Monahan, P.E. Evaluation of risks related to the use of adeno-associated virus-based vectors. Curr. Gene Ther. 2003, 3, 545–565. [Google Scholar] [CrossRef] [Green Version]
- Wade-Martins, R.; Frampton, J.; James, M.R. Long-term stability of large insert genomic DNA episomal shuttle vectors in human cells. Nucleic Acids Res. 1999, 27, 1674–1682. [Google Scholar] [CrossRef] [Green Version]
- Kafri, T.; Blomer, U.; Peterson, D.A.; Gage, F.H.; Verma, I.M. Sustained expression of genes delivered directly into liver and muscle by lentiviral vectors. Nat. Genet. 1997, 17, 314–317. [Google Scholar] [CrossRef]
- Middleton, T.; Sugden, B. Retention of plasmid DNA in mammalian cells is enhanced by binding of the Epstein-Barr virus replication protein EBNA1. J. Virol. 1994, 68, 4067–4071. [Google Scholar] [CrossRef] [Green Version]
- Niu, D.; Wei, H.J.; Lin, L.; George, H.; Wang, T.; Lee, I.H.; Zhao, H.Y.; Wang, Y.; Kan, Y.; Shrock, E.; et al. Inactivation of porcine endogenous retrovirus in pigs using CRISPR-Cas9. Science 2017, 357, 1303–1307. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Carlson, D.F.; Lancto, C.A.; Zang, B.; Kim, E.S.; Walton, M.; Oldeschulte, D.; Seabury, C.; Sonstegard, T.S.; Fahrenkrug, S.C. Production of hornless dairy cattle from genome-edited cell lines. Nat. Biotechnol. 2016, 34, 479–481. [Google Scholar] [CrossRef] [PubMed]
- Miao, X. Recent advances in the development of new transgenic animal technology. Cell Mol. Life Sci. 2013, 70, 815–828. [Google Scholar] [CrossRef]
- Zhao, Z.; Fowle, H.; Valentine, H.; Liu, Z.; Tan, Y.; Pei, J.; Badal, S.; Testa, J.R.; Grana, X. Immortalization of human primary prostate epithelial cells via CRISPR inactivation of the CDKN2A locus and expression of telomerase. Prostate Cancer Prostatic Dis. 2021, 24, 233–243. [Google Scholar] [CrossRef] [PubMed]
- Hu, X.; Li, L.; Yu, X.; Zhang, R.; Yan, S.; Zeng, Z.; Shu, Y.; Zhao, C.; Wu, X.; Lei, J.; et al. CRISPR/Cas9-mediated reversibly immortalized mouse bone marrow stromal stem cells (BMSCs) retain multipotent features of mesenchymal stem cells (MSCs). Oncotarget 2017, 8, 111847–111865. [Google Scholar] [CrossRef]
- Takenouchi, T.; Kitani, H.; Suzuki, S.; Nakai, M.; Fuchimoto, D.I.; Tsukimoto, M.; Shinkai, H.; Sato, M.; Uenishi, H. Immortalization and Characterization of Porcine Macrophages That Had Been Transduced with Lentiviral Vectors Encoding the SV40 Large T Antigen and Porcine Telomerase Reverse Transcriptase. Front. Vet. Sci. 2017, 4, 132. [Google Scholar] [CrossRef] [Green Version]
- Wang, W.; Zhang, T.; Wu, C.; Wang, S.; Wang, Y.; Li, H.; Wang, N. Immortalization of chicken preadipocytes by retroviral transduction of chicken TERT and TR. PLoS ONE 2017, 12, e0177348. [Google Scholar] [CrossRef]
- Wang, J.; Feng, H.; Huang, X.Q.; Xiang, H.; Mao, Y.W.; Liu, J.P.; Yan, Q.; Liu, W.B.; Liu, Y.; Deng, M.; et al. Human telomerase reverse transcriptase immortalizes bovine lens epithelial cells and suppresses differentiation through regulation of the ERK signaling pathway. J. Biol. Chem. 2005, 280, 22776–22787. [Google Scholar] [CrossRef] [Green Version]
- Jia, W.; Wang, S.; Horner, J.W.; Wang, N.; Wang, H.; Gunther, E.J.; DePinho, R.A.; Zhu, J. A BAC transgenic reporter recapitulates in vivo regulation of human telomerase reverse transcriptase in development and tumorigenesis. FASEB J. 2011, 25, 979–989. [Google Scholar] [CrossRef] [Green Version]
- Wright, W.E.; Piatyszek, M.A.; Rainey, W.E.; Byrd, W.; Shay, J.W. Telomerase activity in human germline and embryonic tissues and cells. Dev. Genet. 1996, 18, 173–179. [Google Scholar] [CrossRef]
- Middleman, E.J.; Choi, J.; Venteicher, A.S.; Cheung, P.; Artandi, S.E. Regulation of cellular immortalization and steady-state levels of the telomerase reverse transcriptase through its carboxy-terminal domain. Mol. Cell. Biol. 2006, 26, 2146–2159. [Google Scholar] [CrossRef] [Green Version]
- Huard, S.; Moriarty, T.J.; Autexier, C. The C terminus of the human telomerase reverse transcriptase is a determinant of enzyme processivity. Nucleic Acids Res. 2003, 31, 4059–4070. [Google Scholar] [CrossRef] [Green Version]
- Wen, L.; Zhao, C.; Song, J.; Ma, L.; Ruan, J.; Xia, X.; Chen, Y.E.; Zhang, J.; Ma, P.X.; Xu, J. CRISPR/Cas9-Mediated TERT Disruption in Cancer Cells. Int. J. Mol. Sci. 2020, 21, 653. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chiruvella, K.K.; Liang, Z.; Wilson, T.E. Repair of Double-Strand Breaks by End Joining. Cold Spring Harb. Perspect. Biol. 2013, 5, a012757. [Google Scholar] [CrossRef] [PubMed]
- Symington, L.S.; Gautier, J. Double-strand break end resection and repair pathway choice. Annu Rev. Genet. 2011, 45, 247–271. [Google Scholar] [CrossRef]
- Deriano, L.; Roth, D.B. Modernizing the nonhomologous end-joining repertoire: Alternative and classical NHEJ share the stage. Annu. Rev. Genet. 2013, 47, 433–455. [Google Scholar] [CrossRef]
- Jha, K.K.; Banga, S.; Palejwala, V.; Ozer, H.L. SV40-Mediated immortalization. Exp. Cell Res. 1998, 245, 1–7. [Google Scholar] [CrossRef]
- Yan, T.; Ooi, W.F.; Qamra, A.; Cheung, A.; Ma, D.; Sundaram, G.M.; Xu, C.; Xing, M.; Poon, L.; Wang, J.; et al. HoxC5 and miR-615–3p target newly evolved genomic regions to repress hTERT and inhibit tumorigenesis. Nat. Commun. 2018, 9, 100. [Google Scholar] [CrossRef]
- Jie, M.M.; Chang, X.; Zeng, S.; Liu, C.; Liao, G.B.; Wu, Y.R.; Liu, C.H.; Hu, C.J.; Yang, S.M.; Li, X.Z. Diverse regulatory manners of human telomerase reverse transcriptase. Cell Commun. Signal. 2019, 17, 63. [Google Scholar] [CrossRef] [Green Version]
- Chang, J.T.; Lu, Y.C.; Chen, Y.J.; Tseng, C.P.; Chen, Y.L.; Fang, C.W.; Cheng, A.J. hTERT phosphorylation by PKC is essential for telomerase holoprotein integrity and enzyme activity in head neck cancer cells. Br. J. Cancer 2006, 94, 870–878. [Google Scholar] [CrossRef]
- Li, H.; Zhao, L.; Yang, Z.; Funder, J.W.; Liu, J.P. Telomerase is controlled by protein kinase Calpha in human breast cancer cells. J. Biol. Chem. 1998, 273, 33436–33442. [Google Scholar] [CrossRef] [Green Version]
- Kang, S.S.; Kwon, T.; Kwon, D.Y.; Do, S.I. Akt protein kinase enhances human telomerase activity through phosphorylation of telomerase reverse transcriptase subunit. J. Biol. Chem. 1999, 274, 13085–13090. [Google Scholar] [CrossRef] [Green Version]
- Chung, J.; Khadka, P.; Chung, I.K. Nuclear import of hTERT requires a bipartite nuclear localization signal and Akt-mediated phosphorylation. J. Cell Sci. 2012, 125, 2684–2697. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jeong, S.A.; Kim, K.; Lee, J.H.; Cha, J.S.; Khadka, P.; Cho, H.S.; Chung, I.K. Akt-mediated phosphorylation increases the binding affinity of hTERT for importin alpha to promote nuclear translocation. J. Cell Sci. 2015, 128, 2287–2301. [Google Scholar] [CrossRef] [Green Version]
- Buchner, N.; Zschauer, T.C.; Lukosz, M.; Altschmied, J.; Haendeler, J. Downregulation of mitochondrial telomerase reverse transcriptase induced by H2O2 is Src kinase dependent. Exp. Gerontol. 2010, 45, 558–562. [Google Scholar] [CrossRef] [PubMed]
- Jakob, S.; Schroeder, P.; Lukosz, M.; Buchner, N.; Spyridopoulos, I.; Altschmied, J.; Haendeler, J. Nuclear protein tyrosine phosphatase Shp-2 is one important negative regulator of nuclear export of telomerase reverse transcriptase. J. Biol. Chem. 2008, 283, 33155–33161. [Google Scholar] [CrossRef] [Green Version]
- Cheung, D.H.; Kung, H.F.; Huang, J.J.; Shaw, P.C. PinX1 is involved in telomerase recruitment and regulates telomerase function by mediating its localization. FEBS Lett. 2012, 586, 3166–3171. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhou, X.Z.; Lu, K.P. The Pin2/TRF1-interacting protein PinX1 is a potent telomerase inhibitor. Cell 2001, 107, 347–359. [Google Scholar] [CrossRef] [Green Version]
- Yuan, K.; Li, N.; Jiang, K.; Zhu, T.; Huo, Y.; Wang, C.; Lu, J.; Shaw, A.; Thomas, K.; Zhang, J.; et al. PinX1 is a novel microtubule-binding protein essential for accurate chromosome segregation. J. Biol. Chem. 2009, 284, 23072–23082. [Google Scholar] [CrossRef] [Green Version]
- Shore, D.; Bianchi, A. Telomere length regulation: Coupling DNA end processing to feedback regulation of telomerase. EMBO J. 2009, 28, 2309–2322. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, F.; Podell, E.R.; Zaug, A.J.; Yang, Y.; Baciu, P.; Cech, T.R.; Lei, M. The POT1-TPP1 telomere complex is a telomerase processivity factor. Nature 2007, 445, 506–510. [Google Scholar] [CrossRef]
- Giardini, M.A.; Segatto, M.; da Silva, M.S.; Nunes, V.S.; Cano, M.I. Telomere and telomerase biology. Prog. Mol. Biol. Transl. Sci. 2014, 125, 1–40. [Google Scholar] [PubMed]
- Liu, K.; Hodes, R.J.; Weng, N. Cutting edge: Telomerase activation in human T lymphocytes does not require increase in telomerase reverse transcriptase (hTERT) protein but is associated with hTERT phosphorylation and nuclear translocation. J. Immunol. 2001, 166, 4826–4830. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kim, N.W.; Piatyszek, M.A.; Prowse, K.R.; Harley, C.B.; West, M.D.; Ho, P.L.; Coviello, G.M.; Wright, W.E.; Weinrich, S.L.; Shay, J.W. Specific association of human telomerase activity with immortal cells and cancer. Science 1994, 266, 2011–2015. [Google Scholar] [CrossRef] [PubMed]
- Reichert, S.; Froy, H.; Boner, W.; Burg, T.M.; Daunt, F.; Gillespie, R.; Griffiths, K.; Lewis, S.; Phillips, R.A.; Nussey, D.H.; et al. Telomere length measurement by qPCR in birds is affected by storage method of blood samples. Oecologia 2017, 184, 341–350. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cawthon, R.M. Telomere measurement by quantitative PCR. Nucleic Acids Res. 2002, 30, e47. [Google Scholar] [CrossRef]



| Construct | Enzymatic Activity | Telomere Length |
|---|---|---|
| bTERT | ++++ | ++++ |
| hTERT | +++ | +++ |
| mTERT | − | − |
| bTERT-△C | ++ | − |
| hTERT-△C | − | − |
| mTERT-△C | − | − |
| HHB | +++ | +++ |
| MMB | ++ | + |
| BBH | + | − |
| MMH | + | − |
| BBM | − | − |
| HHM | − | − |
| Immortalization Strategy | Passage 0 | Passage 5 | Passage 15 | Passage 35 | Passage 50 | Passage 80 | |
|---|---|---|---|---|---|---|---|
| Transient transfection system | hTERT | 26 | 1 | 0 | 0 | 0 | 0 |
| 25 | 0 | 0 | 0 | 0 | 0 | ||
| 16 | 0 | 0 | 0 | 0 | 0 | ||
| bTERT | 25 | 2 | 1 | 0 | 0 | 0 | |
| 21 | 1 | 3 | 0 | 0 | 0 | ||
| 30 | 3 | 2 | 0 | 0 | 0 | ||
| Lentivirus- mediated delivery system | hTERT | 6 | 2 | 1 | 1 | 0 | 0 |
| 9 | 1 | 2 | 1 | 1 | 0 | ||
| 9 | 3 | 1 | o | 2 | 0 | ||
| bTERT | 1 | 3 | 2 | 4 | 1 | 1 | |
| 3 | 3 | 4 | 1 | 3 | 0 | ||
| 4 | 2 | 0 | 2 | 2 | 2 | ||
| CRISPR/Cas9 system | hTERT | 2 | 4 | 3 | 1 | 2 | 1 |
| 3 | 2 | 1 | 2 | 1 | 2 | ||
| 2 | 2 | 3 | 4 | 3 | 1 | ||
| bTERT | 0 | 1 | 2 | 3 | 3 | 4 | |
| 1 | 0 | 1 | 3 | 2 | 3 | ||
| 0 | 0 | 3 | 4 | 2 | 7 | ||
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhang, Z.; Han, Z.; Guo, Y.; Liu, X.; Gao, Y.; Zhang, Y. Establishment of an Efficient Immortalization Strategy Using HMEJ-Based bTERT Insertion for Bovine Cells. Int. J. Mol. Sci. 2021, 22, 12540. https://doi.org/10.3390/ijms222212540
Zhang Z, Han Z, Guo Y, Liu X, Gao Y, Zhang Y. Establishment of an Efficient Immortalization Strategy Using HMEJ-Based bTERT Insertion for Bovine Cells. International Journal of Molecular Sciences. 2021; 22(22):12540. https://doi.org/10.3390/ijms222212540
Chicago/Turabian StyleZhang, Zihan, Zhuo Han, Ying Guo, Xin Liu, Yuanpeng Gao, and Yong Zhang. 2021. "Establishment of an Efficient Immortalization Strategy Using HMEJ-Based bTERT Insertion for Bovine Cells" International Journal of Molecular Sciences 22, no. 22: 12540. https://doi.org/10.3390/ijms222212540
APA StyleZhang, Z., Han, Z., Guo, Y., Liu, X., Gao, Y., & Zhang, Y. (2021). Establishment of an Efficient Immortalization Strategy Using HMEJ-Based bTERT Insertion for Bovine Cells. International Journal of Molecular Sciences, 22(22), 12540. https://doi.org/10.3390/ijms222212540





