Non-Cell Autonomous and Epigenetic Mechanisms of Huntington’s Disease
Abstract
1. Introduction
2. Non-Cell Autonomous Cell Death Pathway in HD
2.1. Alteration of Astrocyte Function
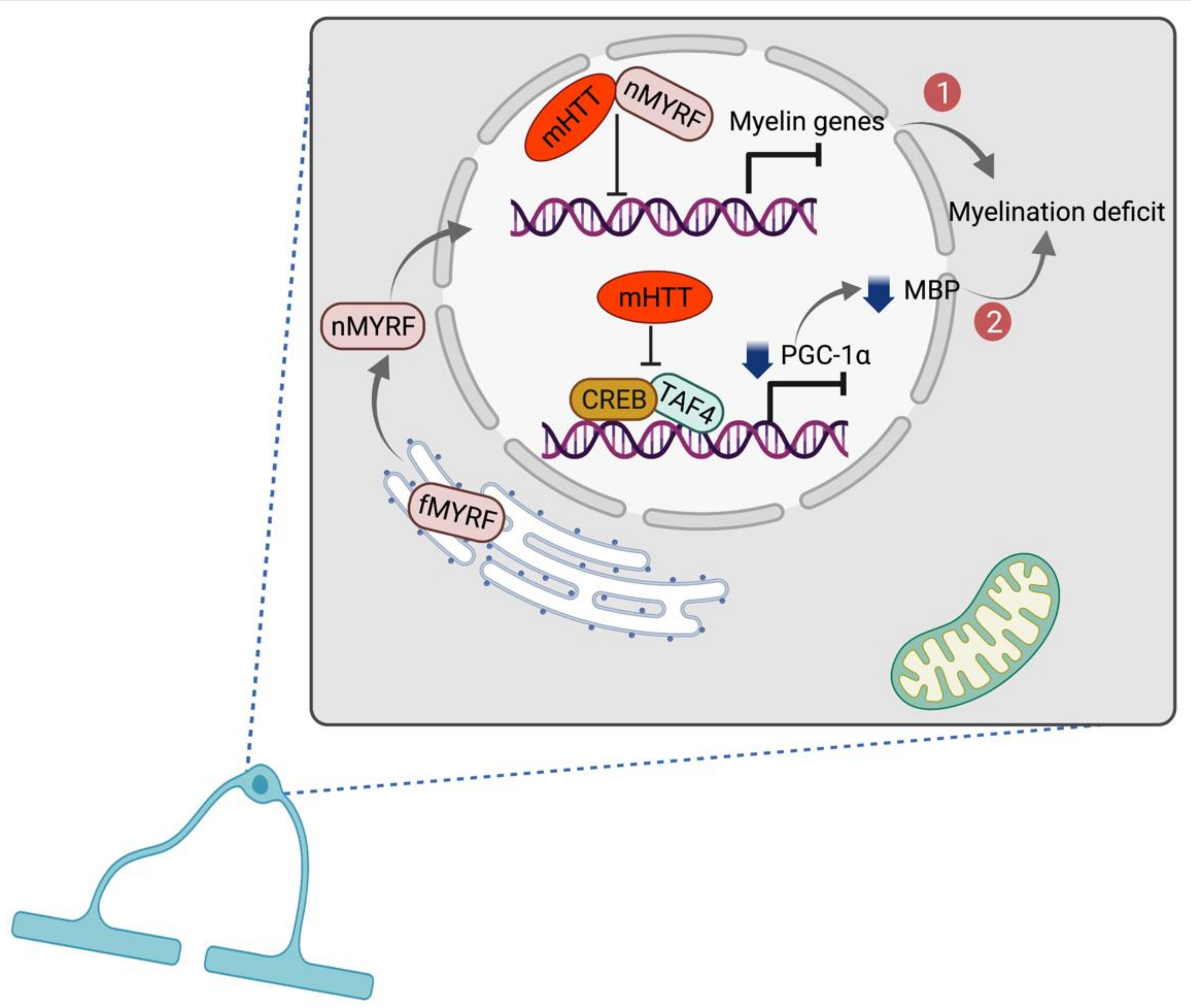
2.2. Alteration of Oligodendrocyte Function
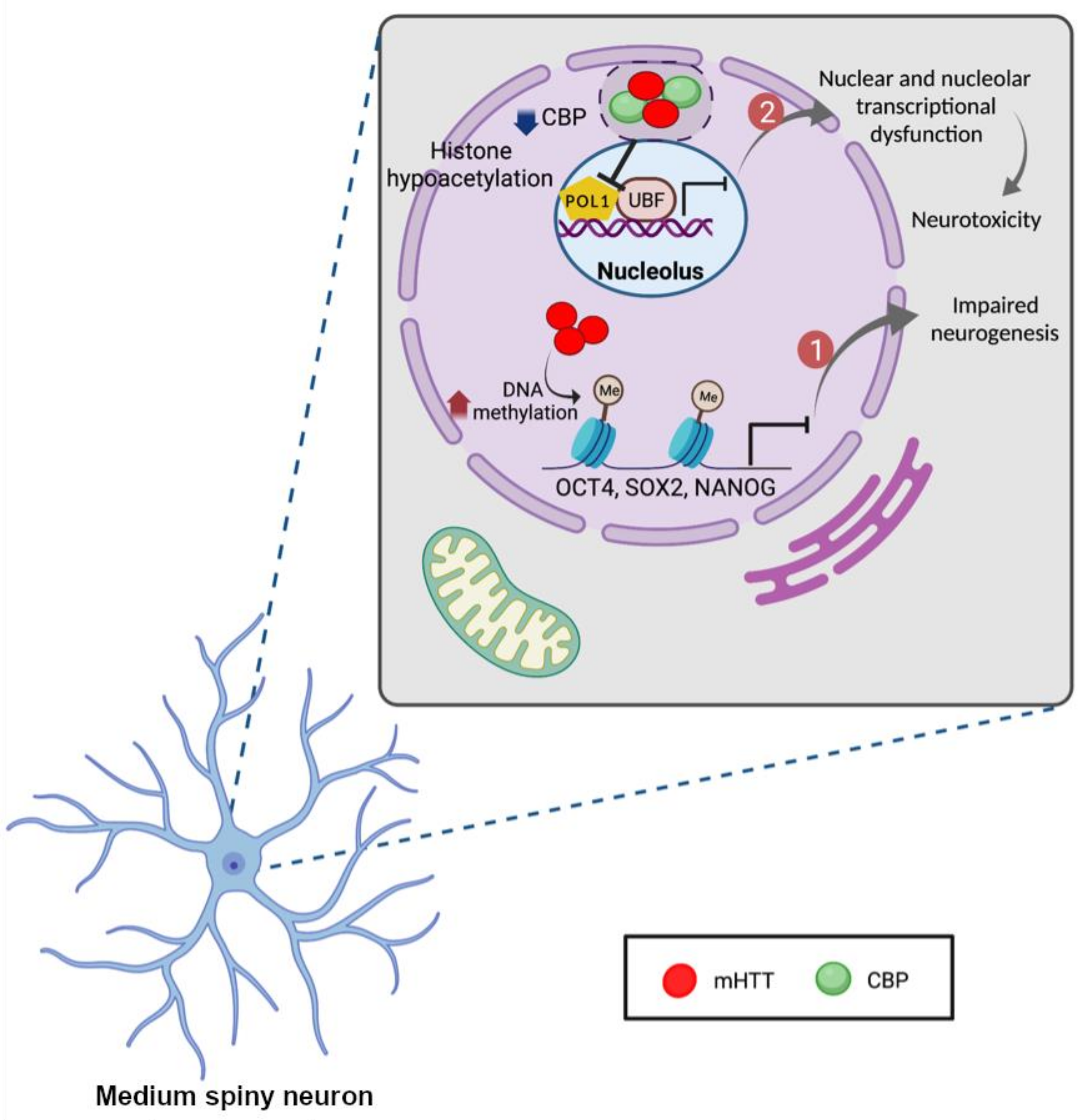
3. The Role of Epigenetic Modifications and Noncoding RNAs in the Pathogenesis of HD
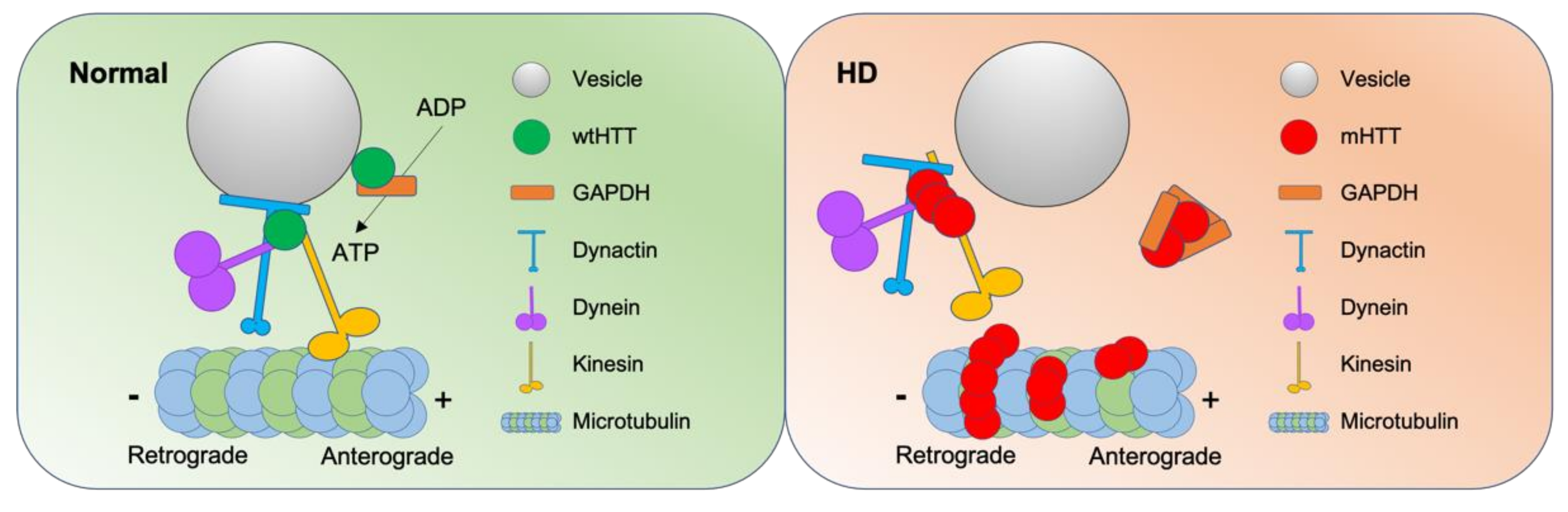
4. Roles of Wild Type HTT (wtHTT) Versus mHTT in Vesicle Trafficking
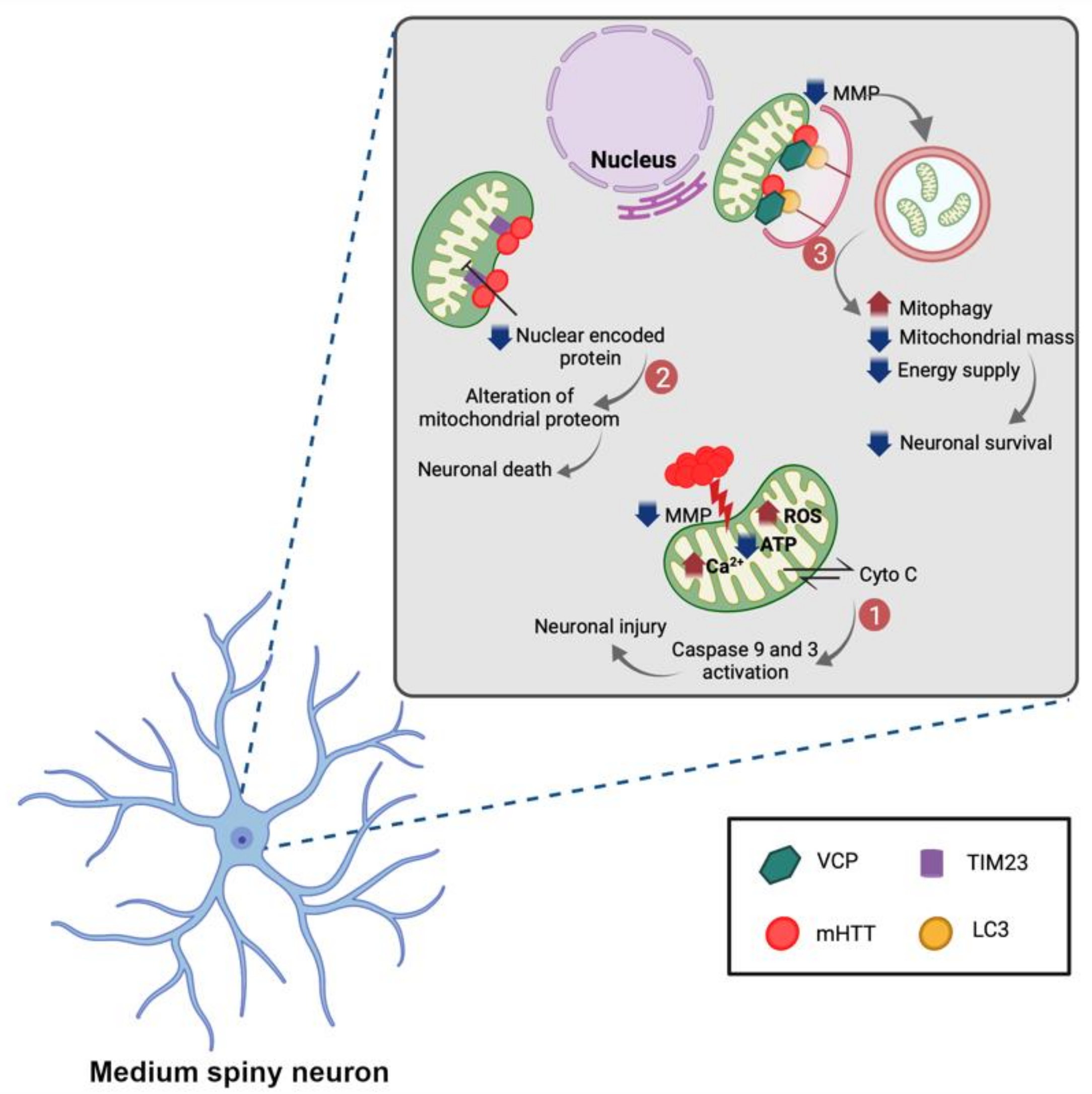
5. Mitochondria Dysfunction in HD
6. Therapeutic Approaches for Huntington’s Disease
7. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Exuzides, A.; Crowell, V.; Reddy, S.R.; Chang, E.; Yohrling, G. Epidemiology of Huntington’s disease (HD) in the US medicare population. Neurology 2020, 94, 670. [Google Scholar]
- The Huntington’s disease collaborative research group. A novel gene containing a trinucleotide repeat that is expanded and unstable on Huntington’s disease chromosomes. Cell 1993, 72, 971–983. [Google Scholar] [CrossRef]
- Di Prospero, N.A.; Fischbeck, K.H. Therapeutics development for triplet repeat expansion diseases. Nat. Rev. Genet. 2005, 6, 756–766. [Google Scholar] [CrossRef] [PubMed]
- Ross, C.A. Polyglutamine pathogenesis: Emergence of unifying mechanisms for Huntington’s disease and related disorders. Neuron 2002, 35, 819–822. [Google Scholar] [CrossRef]
- Cattaneo, E.; Rigamonti, D.; Goffredo, D.; Zuccato, C.; Squitieri, F.; Sipione, S. Loss of normal huntingtin function: New developments in Huntington’s disease research. Trends Neurosci. 2001, 24, 182–188. [Google Scholar] [CrossRef]
- Tomczyk, M.; Glaser, T.; Ulrich, H.; Slominska, E.M.; Smolenski, R.T. Huntingtin protein maintains balanced energetics in mouse cardiomyocytes. Nucleosides Nucleotides Nucleic Acids 2020. accepted. [Google Scholar] [CrossRef]
- Rüb, U.; Seidel, K.; Heinsen, H.; Vonsattel, J.P.; den Dunnen, W.F.; Korf, H.W. Huntington’s disease (HD): The neuropathology of a multisystem neurodegenerative disorder of the human brain. Brain Pathol. 2016, 26, 726–740. [Google Scholar] [CrossRef] [PubMed]
- DiFiglia, M.; Sapp, E.; Chase, K.O.; Davies, S.W.; Bates, G.P.; Vonsattel, J.P.; Aronin, N. Aggregation of Huntingtin in neuronal intranuclear inclusions and dystrophic neurites in brain. Science 1997, 277, 1990–1993. [Google Scholar] [CrossRef] [PubMed]
- Ross, C.A. Intranuclear neuronal inclusions: A common pathogenic mechanism for glutamine-repeat neurodegenerative diseases? Neuron 1997, 19, 1147–1150. [Google Scholar] [CrossRef]
- Kojer, K.; Hering, T.; Bazenet, C.; Weiss, A.; Herrmann, F.; Taanman, J.-W.; Orth, M. Huntingtin aggregates and mitochondrial pathology in skeletal muscle but not heart of late-stage R6/2 mice. J. Huntingt. Dis. 2019, 8, 145–159. [Google Scholar] [CrossRef] [PubMed]
- Creus-Muncunill, J.; Ehrlich, M.E. Cell-autonomous and non-cell-autonomous pathogenic mechanisms in Huntington’s disease: Insights from in vitro and in vivo models. Neurotherapeutics 2019, 16, 957–978. [Google Scholar] [CrossRef]
- Lee, J.; Hwang, Y.J.; Kim, K.Y.; Kowall, N.W.; Ryu, H. Epigenetic mechanisms of neurodegeneration in Huntington’s disease. Neurotherapeutics 2013, 10, 664–676. [Google Scholar] [CrossRef] [PubMed]
- Trushina, E.; Dyer, R.B.; Badger, J.D.; Ure, D.; Eide, L.; Tran, D.D.; Vrieze, B.T.; Legendre-Guillemin, V.; McPherson, P.S.; Mandavilli, B.S. Mutant huntingtin impairs axonal trafficking in mammalian neurons in vivo and in vitro. Mol. Cell. Biol. 2004, 24, 8195–8209. [Google Scholar] [CrossRef] [PubMed]
- Yablonska, S.; Ganesan, V.; Ferrando, L.M.; Kim, J.; Pyzel, A.; Baranova, O.V.; Khattar, N.K.; Larkin, T.M.; Baranov, S.V.; Chen, N.; et al. Mutant huntingtin disrupts mitochondrial proteostasis by interacting with TIM23. Proc. Natl. Acad. Sci. USA 2019, 116, 16593–16602. [Google Scholar] [CrossRef] [PubMed]
- Yamanishi, E.; Hasegawa, K.; Fujita, K.; Ichinose, S.; Yagishita, S.; Murata, M.; Tagawa, K.; Akashi, T.; Eishi, Y.; Okazawa, H. A novel form of necrosis, TRIAD, occurs in human Huntington’s disease. Acta Neuropathol. Commun. 2017, 5, 19. [Google Scholar] [CrossRef] [PubMed]
- Lin, C.-L.; Wang, S.-E.; Hsu, C.-H.; Sheu, S.-J.; Wu, C.-H. Oral treatment with herbal formula B307 alleviates cardiac failure in aging R6/2 mice with Huntington’s disease via suppressing oxidative stress, inflammation, and apoptosis. Clin. Interv. Aging 2015, 10, 1173–1187. [Google Scholar] [PubMed]
- Bassi, S.; Tripathi, T.; Monziani, A.; Di Leva, F.; Biagioli, M. Epigenetics of Huntington’s disease. Adv. Exp. Med. Biol. 2017, 978, 277–299. [Google Scholar] [PubMed]
- Valor, L.M. Understanding histone deacetylation in Huntington’s disease. Oncotarget 2017, 8, 5660–5661. [Google Scholar] [CrossRef] [PubMed]
- Li, S.H.; Li, X.J. Huntingtin-protein interactions and the pathogenesis of Huntington’s disease. Trends Genet. 2004, 20, 146–154. [Google Scholar] [CrossRef]
- Riley, B.E.; Orr, H.T. Polyglutamine neurodegenerative diseases and regulation of transcription: Assembling the puzzle. Genes Dev. 2006, 20, 2183–2192. [Google Scholar] [CrossRef] [PubMed]
- Yamanaka, T.; Miyazaki, H.; Oyama, F.; Kurosawa, M.; Washizu, C.; Doi, H.; Nukina, N. Mutant Huntingtin reduces HSP70 expression through the sequestration of NF-Y transcription factor. Embo. J. 2008, 27, 827–839. [Google Scholar] [CrossRef] [PubMed]
- Zimmer-Bensch, G. Epigenomic remodeling in Huntington’s disease—master or servant? Epigenomes 2020, 4, 15. [Google Scholar] [CrossRef]
- Gray, S.G. Targeting Huntington’s disease through histone deacetylases. Clin. Epigenetics 2011, 2, 257–277. [Google Scholar] [CrossRef] [PubMed]
- Sadri-Vakili, G.; Cha, J.H. Mechanisms of disease: Histone modifications in Huntington’s disease. Nat. Clin. Pr. Neurol. 2006, 2, 330–338. [Google Scholar] [CrossRef]
- Vashishtha, M.; Ng, C.W.; Yildirim, F.; Gipson, T.A.; Kratter, I.H.; Bodai, L.; Song, W.; Lau, A.; Labadorf, A.; Vogel-Ciernia, A.; et al. Targeting H3K4 trimethylation in Huntington disease. Proc. Natl. Acad. Sci. USA 2013, 110, E3027–E3036. [Google Scholar] [CrossRef] [PubMed]
- Jenkins, B.G.; Koroshetz, W.J.; Beal, M.F.; Rosen, B.R. Evidence for impairment of energy metabolism in vivo in Huntington’s disease using localized 1H NMR spectroscopy. Neurology 1993, 43, 2689–2695. [Google Scholar] [CrossRef] [PubMed]
- McBride, H.M.; Neuspiel, M.; Wasiak, S. Mitochondria: More than just a powerhouse. Curr. Biol. 2006, 16, R551–R560. [Google Scholar] [CrossRef]
- Guo, X.; Sun, X.; Hu, D.; Wang, Y.J.; Fujioka, H.; Vyas, R.; Chakrapani, S.; Joshi, A.U.; Luo, Y.; Mochly-Rosen, D.; et al. VCP recruitment to mitochondria causes mitophagy impairment and neurodegeneration in models of Huntington’s disease. Nat. Commun. 2016, 7, 12646. [Google Scholar] [CrossRef] [PubMed]
- Oliveira, J.M. Mitochondrial bioenergetics and dynamics in Huntington’s disease: Tripartite synapses and selective striatal degeneration. J. Bioenerg. Biomembr. 2010, 42, 227–234. [Google Scholar] [CrossRef]
- Yano, H.; Baranov, S.V.; Baranova, O.V.; Kim, J.; Pan, Y.; Yablonska, S.; Carlisle, D.L.; Ferrante, R.J.; Kim, A.H.; Friedlander, R.M. Inhibition of mitochondrial protein import by mutant huntingtin. Nat. Neurosci. 2014, 17, 822–831. [Google Scholar] [CrossRef] [PubMed]
- Faria, A.V.; Ratnanather, J.T.; Tward, D.J.; Lee, D.S.; van den Noort, F.; Wu, D.; Brown, T.; Johnson, H.; Paulsen, J.S.; Ross, C.A.; et al. Linking white matter and deep gray matter alterations in premanifest Huntington disease. Neuroimage Clin. 2016, 11, 450–460. [Google Scholar] [CrossRef] [PubMed]
- Jin, J.; Peng, Q.; Hou, Z.; Jiang, M.; Wang, X.; Langseth, A.J.; Tao, M.; Barker, P.B.; Mori, S.; Bergles, D.E.; et al. Early white matter abnormalities, progressive brain pathology and motor deficits in a novel knock-in mouse model of Huntington’s disease. Hum. Mol. Genet. 2015, 24, 2508–2527. [Google Scholar] [CrossRef] [PubMed]
- Teo, R.T.; Hong, X.; Yu-Taeger, L.; Huang, Y.; Tan, L.J.; Xie, Y.; To, X.V.; Guo, L.; Rajendran, R.; Novati, A.; et al. Structural and molecular myelination deficits occur prior to neuronal loss in the YAC128 and BACHD models of Huntington disease. Hum. Mol. Genet. 2016, 25, 2621–2632. [Google Scholar] [CrossRef] [PubMed]
- Faideau, M.; Kim, J.; Cormier, K.; Gilmore, R.; Welch, M.; Auregan, G.; Dufour, N.; Guillermier, M.; Brouillet, E.; Hantraye, P. In vivo expression of polyglutamine-expanded huntingtin by mouse striatal astrocytes impairs glutamate transport: A correlation with Huntington’s disease subjects. Hum. Mol. Genet. 2010, 19, 3053–3067. [Google Scholar] [CrossRef]
- Khakh, B.S.; Beaumont, V.; Cachope, R.; Munoz-Sanjuan, I.; Goldman, S.A.; Grantyn, R. Unravelling and exploiting astrocyte dysfunction in Huntington’s disease. Trends Neurosci. 2017, 40, 422–437. [Google Scholar] [CrossRef]
- Meunier, C.; Merienne, N.; Jollé, C.; Déglon, N.; Pellerin, L. Astrocytes are key but indirect contributors to the development of the symptomatology and pathophysiology of Huntington’s disease. Glia 2016, 64, 1841–1856. [Google Scholar] [CrossRef] [PubMed]
- Ilieva, H.; Polymenidou, M.; Cleveland, D.W. Non-cell autonomous toxicity in neurodegenerative disorders: ALS and beyond. J. Cell Biol. 2009, 187, 761–772. [Google Scholar] [CrossRef]
- Möller, T. Neuroinflammation in Huntington’s disease. J. Neural. Transm. 2010, 117, 1001–1008. [Google Scholar] [CrossRef] [PubMed]
- Chun, H.; Im, H.; Kang, Y.J.; Kim, Y.; Shin, J.H.; Won, W.; Lim, J.; Ju, Y.; Park, Y.M.; Kim, S. Severe reactive astrocytes precipitate pathological hallmarks of Alzheimer’s disease via H2O2− production. Nat. Neurosci. 2020, 23, 1555–1566. [Google Scholar] [CrossRef]
- Heng, M.Y.; Detloff, P.J.; Wang, P.L.; Tsien, J.Z.; Albin, R.L. In vivo evidence for NMDA receptor-mediated excitotoxicity in a murine genetic model of Huntington disease. J. Neurosci. 2009, 29, 3200–3205. [Google Scholar] [CrossRef]
- Heo, J.Y.; Nam, M.-H.; Yoon, H.H.; Kim, J.; Hwang, Y.J.; Won, W.; Woo, D.H.; Lee, J.A.; Park, H.-J.; Jo, S. Aberrant tonic inhibition of dopaminergic neuronal activity causes motor symptoms in animal models of Parkinson’s disease. Curr. Biol. 2020, 30, 276–291.e9. [Google Scholar] [CrossRef]
- Lee, J.; Hyeon, S.J.; Im, H.; Ryu, H.; Kim, Y.; Ryu, H. Astrocytes and microglia as non-cell autonomous players in the pathogenesis of ALS. Exp. Neurobiol. 2016, 25, 233–240. [Google Scholar] [CrossRef]
- Gu, X.; André, V.M.; Cepeda, C.; Li, S.-H.; Li, X.-J.; Levine, M.S.; Yang, X.W. Pathological cell-cell interactions are necessary for striatal pathogenesis in a conditional mouse model of Huntington’s disease. Mol. Neurodegener. 2007, 2, 8. [Google Scholar] [CrossRef] [PubMed]
- Jing, L.; Cheng, S.; Pan, Y.; Liu, Q.; Yang, W.; Li, S.; Li, X.-J. Accumulation of Endogenous Mutant Huntingtin in Astrocytes Exacerbates Neuropathology of Huntington Disease in Mice. Mol. Neurobiol. 2021, 58, 5112–5126. [Google Scholar] [CrossRef]
- Bason, M.; Meister-Broekema, M.; Alberts, N.; Dijkers, P.; Bergink, S.; Sibon, O.C.M.; Kampinga, H.H. Astrocytic expression of the chaperone DNAJB6 results in non-cell autonomous protection in Huntington’s disease. Neurobiol. Dis. 2019, 124, 108–117. [Google Scholar] [CrossRef] [PubMed]
- Benraiss, A.; Mariani, J.N.; Tate, A.; Solly, R.; Capellano, L.; de Mesy Bentley, K.L.; Chandler-Militello, D.; Goldman, S. Huntington Disease Mice Exhibit a TCF7L2-Responsive Suppression of Both Homeostatic and Compensatory Remyelination. SSRN Electron. J. 2021. under review. [Google Scholar] [CrossRef]
- Savage, J.C.; St-Pierre, M.-K.; Carrier, M.; El Hajj, H.; Novak, S.W.; Sanchez, M.G.; Cicchetti, F.; Tremblay, M.-È. Microglial physiological properties and interactions with synapses are altered at presymptomatic stages in a mouse model of Huntington’s disease pathology. J. Neuroinflammation 2020, 17, 98. [Google Scholar] [CrossRef]
- Kim, A.; García-García, E.; Straccia, M.; Comella-Bolla, A.; Miguez, A.; Masana, M.; Alberch, J.; Canals, J.M.; Rodríguez, M.J. Reduced Fractalkine Levels Lead to Striatal Synaptic Plasticity Deficits in Huntington’s Disease. Front. Cell. Neurosci. 2020, 14, 163. [Google Scholar] [CrossRef] [PubMed]
- O’Regan, G.C.; Farag, S.H.; Casey, C.S.; Wood-Kaczmar, A.; Pocock, J.M.; Tabrizi, S.J.; Andre, R. Human Huntington’s disease pluripotent stem cell-derived microglia develop normally but are abnormally hyper-reactive and release elevated levels of reactive oxygen species. J. Neuroinflammation 2021, 18, 94. [Google Scholar] [CrossRef]
- Bolla, A.C.; Valente, T.; Miguez, A.; Brito, V.; Gines, S.; Solà, C.; Straccia, M.; Canals, J.M. CD200 is up-regulated in R6/1 transgenic mouse model of Huntington’s disease. PLoS ONE 2019, 14, e0224901. [Google Scholar]
- Caron, N.S.; Banos, R.; Yanick, C.; Aly, A.E.; Byrne, L.M.; Smith, E.D.; Xie, Y.; Smith, S.E.; Potluri, N.; Black, H.F.; et al. Mutant huntingtin is cleared from the brain via active mechanisms in Huntington disease. J. Neurosci. 2021, 41, 780–796. [Google Scholar] [CrossRef] [PubMed]
- Sameni, S.; Zhang, R.; Digman, M.A. The phasor FLIM method reveals a link between a change in energy metabolism and mHtt protein spread in healthy Mammalian cells when co-cultured with Huntington diseased cells. Methods Appl. Fluoresc. 2021, 9, 015005. [Google Scholar] [CrossRef]
- Singh, A.; Agrawal, N. Deciphering the key mechanisms leading to alteration of lipid metabolism in Drosophila model of Huntington’s disease. Biochim. Et. Biophys. Acta (BBA)-Mol. Basis Dis. 2021, 1867, 166127. [Google Scholar] [CrossRef] [PubMed]
- Sancho, L.; Contreras, M.; Allen, N.J. Glia as sculptors of synaptic plasticity. Neurosci. Res. 2020, 167, 17–29. [Google Scholar] [CrossRef]
- Perea, G.; Navarrete, M.; Araque, A. Tripartite synapses: Astrocytes process and control synaptic information. Trends Neurosci. 2009, 32, 421–431. [Google Scholar] [CrossRef] [PubMed]
- Halassa, M.M.; Fellin, T.; Haydon, P.G. The tripartite synapse: Roles for gliotransmission in health and disease. Trends Mol. Med. 2007, 13, 54–63. [Google Scholar] [CrossRef]
- Brymer, K.J.; Barnes, J.R.; Parsons, M.P. Entering a new era of quantifying glutamate clearance in health and disease. J. Neurosci. Res. 2021, 99, 1598–1617. [Google Scholar] [CrossRef]
- Florence, G.; Pereira, T.; Kurths, J. Extracellular potassium dynamics in the hyperexcitable state of the neuronal ictal activity. Commun. Nonlinear Sci. Numer. Simul. 2012, 17, 4700–4706. [Google Scholar] [CrossRef]
- Radiske, A.; Gonzalez, M.C.; Nôga, D.A.; Rossato, J.I.; Bevilaqua, L.R.M.; Cammarota, M. GluN2B and GluN2A-containing NMDAR are differentially involved in extinction memory destabilization and restabilization during reconsolidation. Sci. Rep. 2021, 11, 186. [Google Scholar] [CrossRef]
- Schmidt, M.E.; Caron, N.S.; Aly, A.E.; Lemarié, F.L.; Dal Cengio, L.; Ko, Y.; Lazic, N.; Anderson, L.; Nguyen, B.; Raymond, L.A. DAPK1 promotes extrasynaptic GluN2B phosphorylation and striatal spine instability in the YAC128 mouse model of Huntington disease. Front. Cell. Neurosci. 2020, 14, 348. [Google Scholar] [CrossRef] [PubMed]
- Miller, B.R.; Bezprozvanny, I. Corticostriatal circuit dysfunction in Huntington’s disease: Intersection of glutamate, dopamine and calcium. Future Neurol. 2010, 5, 735–756. [Google Scholar] [CrossRef]
- Raymond, L.A.; André, V.M.; Cepeda, C.; Gladding, C.M.; Milnerwood, A.J.; Levine, M.S. Pathophysiology of Huntington’s disease: Time-dependent alterations in synaptic and receptor function. Neuroscience 2011, 198, 252–273. [Google Scholar] [CrossRef] [PubMed]
- Kang, R.; Wang, L.; Sanders, S.S.; Zuo, K.; Hayden, M.R.; Raymond, L.A. Altered regulation of striatal neuronal N-methyl-D-aspartate receptor trafficking by palmitoylation in Huntington disease mouse model. Front. Synaptic Neurosci. 2019, 11, 3. [Google Scholar] [CrossRef] [PubMed]
- Marco, S.; Murillo, A.; Pérez-Otaño, I. RNAi-based GluN3A silencing prevents and reverses disease phenotypes induced by mutant huntingtin. Mol. Ther. 2018, 26, 1965–1972. [Google Scholar] [CrossRef]
- Cepeda, C.; Wu, N.; André, V.M.; Cummings, D.M.; Levine, M.S. The corticostriatal pathway in Huntington’s disease. Prog. Neurobiol. 2007, 81, 253–271. [Google Scholar] [CrossRef]
- Bunner, K.D.; Rebec, G.V. Corticostriatal dysfunction in Huntington’s disease: The basics. Front. Hum. Neurosci. 2016, 10, 317. [Google Scholar] [CrossRef]
- Wood, T.E.; Barry, J.; Yang, Z.; Cepeda, C.; Levine, M.S.; Gray, M. Mutant huntingtin reduction in astrocytes slows disease progression in the BACHD conditional Huntington’s disease mouse model. Hum. Mol. Genet. 2019, 28, 487–500. [Google Scholar] [CrossRef]
- Blumenstock, S.; Dudanova, I. Cortical and striatal circuits in Huntington’s disease. Front. Neurosci. 2020, 14, 82. [Google Scholar] [CrossRef]
- Chan, C.S.; Surmeier, D.J. Astrocytes go awry in Huntington’s disease. Nat. Neurosci. 2014, 17, 641–642. [Google Scholar] [CrossRef] [PubMed]
- Ehrlich, M.E. Huntington’s disease and the striatal medium spiny neuron: Cell-autonomous and non-cell-autonomous mechanisms of disease. Neurotherapeutics 2012, 9, 270–284. [Google Scholar] [CrossRef]
- Estrada-Sánchez, A.M.; Rebec, G.V. Corticostriatal dysfunction and glutamate transporter 1 (GLT1) in Huntington’s disease: Interactions between neurons and astrocytes. Basal. Ganglia 2012, 2, 57–66. [Google Scholar] [CrossRef] [PubMed]
- Gray, M. The role of astrocytes in Huntington’s disease. In Pathological Potential of Neuroglia; Springer: Berlin/Heidelberg, Germany, 2014; pp. 213–229. [Google Scholar]
- Gray, M. Astrocytes in Huntington’s disease. Neurogl. Neurodegener. Dis. 2019, 1175, 355–381. [Google Scholar]
- Palpagama, T.H.; Waldvogel, H.J.; Faull, R.L.; Kwakowsky, A. The role of microglia and astrocytes in Huntington’s disease. Front. Mol. Neurosci. 2019, 12, 258. [Google Scholar] [CrossRef] [PubMed]
- Proft, J.; Weiss, N. Rectifying rectifier channels in Huntington disease. Commun. Integr. Biol. 2014, 7, 694–703. [Google Scholar] [CrossRef][Green Version]
- Sánchez, A.M.E.; Mejía-Toiber, J.; Massieu, L. Excitotoxic neuronal death and the pathogenesis of Huntington’s disease. Arch. Med. Res. 2008, 39, 265–276. [Google Scholar] [CrossRef]
- Wilton, D.K.; Stevens, B. The contribution of glial cells to Huntington’s disease pathogenesis. Neurobiol. Dis. 2020, 143, 104963. [Google Scholar] [CrossRef]
- Al-Dalahmah, O.; Sosunov, A.A.; Shaik, A.; Ofori, K.; Liu, Y.; Vonsattel, J.P.; Adorjan, I.; Menon, V.; Goldman, J.E. Single-nucleus RNA-seq identifies Huntington disease astrocyte states. Acta Neuropathol. Commun. 2020, 8, 19. [Google Scholar] [CrossRef]
- Arzberger, T.; Krampfl, K.; Leimgruber, S.; Weindl, A. Changes of NMDA receptor subunit (NR1, NR2B) and glutamate transporter (GLT1) mRNA expression in Huntington’s disease—an in situ hybridization study. J. Neuropathol. Exp. Neurol. 1997, 56, 440–454. [Google Scholar] [CrossRef]
- Petr, G.T.; Schultheis, L.A.; Hussey, K.C.; Sun, Y.; Dubinsky, J.M.; Aoki, C.; Rosenberg, P.A. Decreased expression of GLT-1 in the R6/2 model of Huntington’s disease does not worsen disease progression. Eur. J. Neurosci. 2013, 38, 2477–2490. [Google Scholar] [CrossRef]
- Skotte, N.H.; Andersen, J.V.; Santos, A.; Aldana, B.I.; Willert, C.W.; Nørremølle, A.; Waagepetersen, H.S.; Nielsen, M.L. Integrative characterization of the R6/2 mouse model of Huntington’s disease reveals dysfunctional astrocyte metabolism. Cell Rep. 2018, 23, 2211–2224. [Google Scholar] [CrossRef]
- Cho, I.K.; Yang, B.; Forest, C.; Qian, L.; Chan, A.W. Amelioration of Huntington’s disease phenotype in astrocytes derived from iPSC-derived neural progenitor cells of Huntington’s disease monkeys. PLoS ONE 2019, 14, e0214156. [Google Scholar]
- Dvorzhak, A.; Helassa, N.; Török, K.; Schmitz, D.; Grantyn, R. Single synapse indicators of impaired glutamate clearance derived from fast iGluu imaging of cortical afferents in the striatum of normal and Huntington (Q175) mice. J. Neurosci. 2019, 39, 3970–3982. [Google Scholar] [CrossRef] [PubMed]
- Hassel, B.; Tessler, S.; Faull, R.L.; Emson, P.C. Glutamate uptake is reduced in prefrontal cortex in Huntington’s disease. Neurochem. Res. 2008, 33, 232–237. [Google Scholar] [CrossRef]
- Garcia, M.; Charvin, D.; Caboche, J. Expanded huntingtin activates the C-Jun N terminal kinase/c-Jun pathway prior to aggregate formation in striatal neurons in culture. Neuroscience 2004, 127, 859–870. [Google Scholar] [CrossRef] [PubMed]
- Tong, X.; Ao, Y.; Faas, G.C.; Nwaobi, S.E.; Xu, J.; Haustein, M.D.; Anderson, M.A.; Mody, I.; Olsen, M.L.; Sofroniew, M.V. Astrocyte Kir4. 1 ion channel deficits contribute to neuronal dysfunction in Huntington’s disease model mice. Nat. Neurosci. 2014, 17, 694–703. [Google Scholar] [CrossRef] [PubMed]
- Jiang, R.; Diaz-Castro, B.; Looger, L.L.; Khakh, B.S. Dysfunctional calcium and glutamate signaling in striatal astrocytes from Huntington’s disease model mice. J. Neurosci. 2016, 36, 3453–3470. [Google Scholar] [CrossRef]
- Bazargani, N.; Attwell, D. Astrocyte calcium signaling: The third wave. Nat. Neurosci. 2016, 19, 182–189. [Google Scholar] [CrossRef]
- Garcia, V.J.; Rushton, D.J.; Tom, C.M.; Allen, N.D.; Kemp, P.J.; Svendsen, C.N.; Mattis, V.B. Huntington’s disease patient-derived astrocytes display electrophysiological impairments and reduced neuronal support. Front. Neurosci. 2019, 13, 669. [Google Scholar] [CrossRef]
- Shin, J.-Y.; Fang, Z.-H.; Yu, Z.-X.; Wang, C.-E.; Li, S.-H.; Li, X.-J. Expression of mutant huntingtin in glial cells contributes to neuronal excitotoxicity. J. Cell Biol. 2005, 171, 1001–1012. [Google Scholar] [CrossRef]
- Bankston, A.N.; Mandler, M.D.; Feng, Y. Oligodendroglia and neurotrophic factors in neurodegeneration. Neurosci. Bull. 2013, 29, 216–228. [Google Scholar] [CrossRef]
- Myers, R.H.; Vonsattel, J.P.; Paskevich, P.A.; Kiely, D.K.; Stevens, T.J.; Cupples, L.A.; Richardson, E.P., Jr.; Bird, E.D. Decreased neuronal and increased oligodendroglial densities in Huntington’s disease caudate nucleus. J. Neuropathol. Exp. Neurol. 1991, 50, 729–742. [Google Scholar] [CrossRef] [PubMed]
- Bartzokis, G.; Lu, P.H.; Tishler, T.A.; Fong, S.M.; Oluwadara, B.; Finn, J.P.; Huang, D.; Bordelon, Y.; Mintz, J.; Perlman, S. Myelin breakdown and iron changes in Huntington’s disease: Pathogenesis and treatment implications. Neurochem. Res. 2007, 32, 1655–1664. [Google Scholar] [CrossRef] [PubMed]
- Phillips, O.; Squitieri, F.; Sanchez-Castaneda, C.; Elifani, F.; Caltagirone, C.; Sabatini, U.; Di Paola, M. Deep white matter in Huntington’s disease. PLoS ONE 2014, 9, e109676. [Google Scholar]
- Bardile, C.F.; Garcia-Miralles, M.; Caron, N.S.; Rayan, N.A.; Langley, S.R.; Harmston, N.; Rondelli, A.M.; Teo, R.T.Y.; Waltl, S.; Anderson, L.M.; et al. Intrinsic mutant HTT-mediated defects in oligodendroglia cause myelination deficits and behavioral abnormalities in Huntington disease. Proc. Natl. Acad. Sci. USA 2019, 116, 9622–9627. [Google Scholar] [CrossRef] [PubMed]
- Huang, B.; Wei, W.; Wang, G.; Gaertig, M.A.; Feng, Y.; Wang, W.; Li, X.J.; Li, S. Mutant huntingtin downregulates myelin regulatory factor-mediated myelin gene expression and affects mature oligodendrocytes. Neuron 2015, 85, 1212–1226. [Google Scholar] [CrossRef]
- Wang, N.; Yang, X.W. Huntington disease’s glial progenitor cells hit the pause button in the mouse brain. Cell Stem. Cell 2019, 24, 3–4. [Google Scholar] [CrossRef]
- Cui, L.; Jeong, H.; Borovecki, F.; Parkhurst, C.N.; Tanese, N.; Krainc, D. Transcriptional repression of PGC-1alpha by mutant huntingtin leads to mitochondrial dysfunction and neurodegeneration. Cell 2006, 127, 59–69. [Google Scholar] [CrossRef]
- Johri, A.; Chandra, A.; Beal, M.F. PGC-1α, mitochondrial dysfunction, and Huntington’s disease. Free Radic. Biol. Med. 2013, 62, 37–46. [Google Scholar] [CrossRef]
- Xiang, Z.; Valenza, M.; Cui, L.; Leoni, V.; Jeong, H.K.; Brilli, E.; Zhang, J.; Peng, Q.; Duan, W.; Reeves, S.A.; et al. Peroxisome-proliferator-activated receptor gamma coactivator 1 α contributes to dysmyelination in experimental models of Huntington’s disease. J. Neurosci. 2011, 31, 9544–9553. [Google Scholar] [CrossRef]
- Ryu, H.; Ferrante, R.J. Emerging chemotherapeutic strategies for Huntington’s disease. Expert Opin. Emerg. Drugs 2005, 10, 345–363. [Google Scholar] [CrossRef] [PubMed]
- Allis, C.D.; Jenuwein, T. The molecular hallmarks of epigenetic control. Nat. Rev. Genet. 2016, 17, 487–500. [Google Scholar] [CrossRef]
- Kuehner, J.N.; Bruggeman, E.C.; Wen, Z.; Yao, B. Epigenetic regulations in neuropsychiatric disorders. Front. Genet. 2019, 10, 268. [Google Scholar] [CrossRef] [PubMed]
- Liu, C.; Jiao, C.; Wang, K.; Yuan, N. DNA methylation and psychiatric disorders. Prog Mol. Biol. Transl. Sci. 2018, 157, 175–232. [Google Scholar] [PubMed]
- Grayson, D.R.; Guidotti, A. The dynamics of DNA methylation in Schizophrenia and related psychiatric disorders. Neuropsychopharmacology 2013, 38, 138–166. [Google Scholar] [CrossRef]
- Suzuki, M.M.; Bird, A. DNA methylation landscapes: Provocative insights from epigenomics. Nat. Rev. Genet. 2008, 9, 465–476. [Google Scholar] [CrossRef] [PubMed]
- Ng, C.W.; Yildirim, F.; Yap, Y.S.; Dalin, S.; Matthews, B.J.; Velez, P.J.; Labadorf, A.; Housman, D.E.; Fraenkel, E. Extensive changes in DNA methylation are associated with expression of mutant huntingtin. Proc. Natl. Acad. Sci. USA 2013, 110, 2354–2359. [Google Scholar] [CrossRef]
- Beal, M.F.; Ferrante, R.J. Experimental therapeutics in transgenic mouse models of Huntington’s disease. Nat. Rev. Neurosci. 2004, 5, 373–384. [Google Scholar] [CrossRef]
- Lee, J.; Hwang, Y.J.; Ryu, H.; Kowall, N.W.; Ryu, H. Nucleolar dysfunction in Huntington’s disease. Biochim. Biophys Acta 2014, 1842, 785–790. [Google Scholar] [CrossRef][Green Version]
- Korzus, E.; Rosenfeld, M.G.; Mayford, M. CBP histone acetyltransferase activity is a critical component of memory consolidation. Neuron 2004, 42, 961–972. [Google Scholar] [CrossRef]
- Glajch, K.E.; Sadri-Vakili, G. Epigenetic mechanisms involved in Huntington’s disease pathogenesis. J. Huntingt. Dis. 2015, 4, 1–15. [Google Scholar] [CrossRef]
- Jiang, H.; Poirier, M.A.; Liang, Y.; Pei, Z.; Weiskittel, C.E.; Smith, W.W.; DeFranco, D.B.; Ross, C.A. Depletion of CBP is directly linked with cellular toxicity caused by mutant huntingtin. Neurobiol. Dis. 2006, 23, 543–551. [Google Scholar] [CrossRef] [PubMed]
- McFarland, K.N.; Das, S.; Sun, T.T.; Leyfer, D.; Xia, E.; Sangrey, G.R.; Kuhn, A.; Luthi-Carter, R.; Clark, T.W.; Sadri-Vakili, G.; et al. Genome-wide histone acetylation is altered in a transgenic mouse model of Huntington’s disease. PLoS ONE 2012, 7, e41423. [Google Scholar] [CrossRef] [PubMed]
- Sadri-Vakili, G.; Bouzou, B.; Benn, C.L.; Kim, M.O.; Chawla, P.; Overland, R.P.; Glajch, K.E.; Xia, E.; Qiu, Z.; Hersch, S.M.; et al. Histones associated with downregulated genes are hypo-acetylated in Huntington’s disease models. Hum. Mol. Genet. 2007, 16, 1293–1306. [Google Scholar] [CrossRef]
- Ryu, H.; Lee, J.; Hagerty, S.W.; Soh, B.Y.; McAlpin, S.E.; Cormier, K.A.; Smith, K.M.; Ferrante, R.J. ESET/SETDB1 gene expression and histone H3 (K9) trimethylation in Huntington’s disease. Proc. Natl. Acad. Sci. USA 2006, 103, 19176–19181. [Google Scholar] [CrossRef] [PubMed]
- De Pietri Tonelli, D.; Pulvers, J.N.; Haffner, C.; Murchison, E.P.; Hannon, G.J.; Huttner, W.B. miRNAs are essential for survival and differentiation of newborn neurons but not for expansion of neural progenitors during early neurogenesis in the mouse embryonic neocortex. Development 2008, 135, 3911–3921. [Google Scholar] [CrossRef] [PubMed]
- Dong, X.; Cong, S. MicroRNAs in Huntington’s disease: Diagnostic biomarkers or therapeutic agents? Front. Cell. Neurosci. 2021, 15, 313. [Google Scholar] [CrossRef]
- Juźwik, C.A.; Drake, S.S.; Zhang, Y.; Paradis-Isler, N.; Sylvester, A.; Amar-Zifkin, A.; Douglas, C.; Morquette, B.; Moore, C.S.; Fournier, A.E. microRNA dysregulation in neurodegenerative diseases: A systematic review. Prog. Neurobiol. 2019, 182, 101664. [Google Scholar] [CrossRef] [PubMed]
- Sinha, M.; Ghose, J.; Bhattarcharyya, N.P. Micro RNA -214,-150,-146a and-125b target Huntingtin gene. RNA Biol. 2011, 8, 1005–1021. [Google Scholar] [CrossRef]
- Das, S.; Bhattacharyya, N.P. Heat shock factor 1-regulated miRNAs can target Huntingtin and suppress aggregates of mutant huntingtin. Microrna 2015, 4, 185–193. [Google Scholar] [CrossRef]
- Kozlowska, E.; Krzyzosiak, W.J.; Koscianska, E. Regulation of huntingtin gene expression by miRNA-137, -214, -148a, and their respective isomiRs. Int. J. Mol. Sci. 2013, 14, 16999–17016. [Google Scholar] [CrossRef]
- Bucha, S.; Mukhopadhyay, D.; Bhattacharyya, N.P. Regulation of mitochondrial morphology and cell cycle by microRNA-214 targeting Mitofusin2. Biochem. Biophys Res. Commun. 2015, 465, 797–802. [Google Scholar] [CrossRef]
- Ma, B.; Savas, J.N.; Yu, M.-S.; Culver, B.P.; Chao, M.V.; Tanese, N. Huntingtin mediates dendritic transport of β-actin mRNA in rat neurons. Sci. Rep. 2011, 1, 140. [Google Scholar] [CrossRef] [PubMed]
- Caviston, J.P.; Ross, J.L.; Antony, S.M.; Tokito, M.; Holzbaur, E.L. Huntingtin facilitates dynein/dynactin-mediated vesicle transport. Proc. Natl. Acad. Sci. USA 2007, 104, 10045–10050. [Google Scholar] [CrossRef]
- Caviston, J.P.; Holzbaur, E.L. Huntingtin as an essential integrator of intracellular vesicular trafficking. Trends Cell Biol. 2009, 19, 147–155. [Google Scholar] [CrossRef] [PubMed]
- Zala, D.; Hinckelmann, M.-V.; Yu, H.; Da Cunha, M.M.L.; Liot, G.; Cordelières, F.P.; Marco, S.; Saudou, F. Vesicular glycolysis provides on-board energy for fast axonal transport. Cell 2013, 152, 479–491. [Google Scholar] [CrossRef] [PubMed]
- Burke, J.R.; Enghild, J.J.; Martin, M.E.; Jou, Y.-S.; Myers, R.M.; Roses, A.D.; Vance, J.M.; Strittmatter, W.J. Huntingtin and DRPLA proteins selectively interact with the enzyme GAPDH. Nat. Med. 1996, 2, 347–350. [Google Scholar] [CrossRef]
- Wu, J.; Lin, F.; Qin, Z. Sequestration of glyceraldehyde-3-phosphate dehydrogenase to aggregates formed by mutant huntingtin. Acta Biochim. Et Biophys. Sin. 2007, 39, 885–890. [Google Scholar] [CrossRef][Green Version]
- Chaudhary, R.K.; Patel, K.A.; Patel, M.K.; Joshi, R.H.; Roy, I. Inhibition of aggregation of mutant huntingtin by nucleic acid aptamers in vitro and in a yeast model of Huntington’s disease. Mol. Ther. 2015, 23, 1912–1926. [Google Scholar] [CrossRef] [PubMed]
- Gauthier, L.R.; Charrin, B.C.; Borrell-Pagès, M.; Dompierre, J.P.; Rangone, H.; Cordelières, F.P.; De Mey, J.; MacDonald, M.E.; Leßmann, V.; Humbert, S. Huntingtin controls neurotrophic support and survival of neurons by enhancing BDNF vesicular transport along microtubules. Cell 2004, 118, 127–138. [Google Scholar] [CrossRef]
- Twelvetrees, A.E.; Lesept, F.; Holzbaur, E.L.; Kittler, J.T. The adaptor proteins HAP1a and GRIP1 collaborate to activate the kinesin-1 isoform KIF5C. J. Cell Sci. 2019, 132, jcs215822. [Google Scholar] [CrossRef] [PubMed]
- Colin, E.; Zala, D.; Liot, G.; Rangone, H.; Borrell-Pagès, M.; Li, X.J.; Saudou, F.; Humbert, S. Huntingtin phosphorylation acts as a molecular switch for anterograde/retrograde transport in neurons. EMBO J. 2008, 27, 2124–2134. [Google Scholar] [CrossRef]
- Warby, S.C.; Chan, E.Y.; Metzler, M.; Gan, L.; Singaraja, R.R.; Crocker, S.F.; Robertson, H.A.; Hayden, M.R. Huntingtin phosphorylation on serine 421 is significantly reduced in the striatum and by polyglutamine expansion in vivo. Hum. Mol. Genet. 2005, 14, 1569–1577. [Google Scholar] [CrossRef] [PubMed]
- Guo, W.; Dittlau, K.S.; Van Den Bosch, L. Axonal transport defects and neurodegeneration: Molecular mechanisms and therapeutic implications. Semin. Cell Dev. Biol. 2020, 99, 133–150. [Google Scholar] [CrossRef] [PubMed]
- Morfini, G.; Pigino, G.; Brady, S.T. Polyglutamine expansion diseases: Failing to deliver. Trends Mol. Med. 2005, 11, 64–70. [Google Scholar] [CrossRef] [PubMed]
- Her, L.-S.; Goldstein, L.S. Enhanced sensitivity of striatal neurons to axonal transport defects induced by mutant huntingtin. J. Neurosci. 2008, 28, 13662–13672. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Li, S.-H.; Yu, Z.-X.; Shelbourne, P.; Li, X.-J. Huntingtin aggregate-associated axonal degeneration is an early pathological event in Huntington’s disease mice. J. Neurosci. 2001, 21, 8473–8481. [Google Scholar] [CrossRef]
- White, J.A.; Krzystek, T.J.; Hoffmar-Glennon, H.; Thant, C.; Zimmerman, K.; Iacobucci, G.; Vail, J.; Thurston, L.; Rahman, S.; Gunawardena, S. Excess Rab4 rescues synaptic and behavioral dysfunction caused by defective HTT-Rab4 axonal transport in Huntington’s disease. Acta Neuropathol. Commun. 2020, 8, 97. [Google Scholar] [CrossRef]
- Liot, G.; Zala, D.; Pla, P.; Mottet, G.; Piel, M.; Saudou, F. Mutant Huntingtin alters retrograde transport of TrkB receptors in striatal dendrites. J. Neurosci. 2013, 33, 6298–6309. [Google Scholar] [CrossRef]
- Wong, Y.C.; Holzbaur, E.L. The regulation of autophagosome dynamics by huntingtin and HAP1 is disrupted by expression of mutant huntingtin, leading to defective cargo degradation. J. Neurosci. 2014, 34, 1293. [Google Scholar] [CrossRef] [PubMed]
- Gunawardena, S.; Her, L.-S.; Brusch, R.G.; Laymon, R.A.; Niesman, I.R.; Gordesky-Gold, B.; Sintasath, L.; Bonini, N.M.; Goldstein, L.S. Disruption of axonal transport by loss of huntingtin or expression of pathogenic polyQ proteins in Drosophila. Neuron 2003, 40, 25–40. [Google Scholar] [CrossRef]
- Dompierre, J.P.; Godin, J.D.; Charrin, B.C.; Cordelieres, F.P.; King, S.J.; Humbert, S.; Saudou, F. Histone deacetylase 6 inhibition compensates for the transport deficit in Huntington’s disease by increasing tubulin acetylation. J. Neurosci. 2007, 27, 3571–3583. [Google Scholar] [CrossRef] [PubMed]
- Qin, Z.-H.; Wang, Y.; Sapp, E.; Cuiffo, B.; Wanker, E.; Hayden, M.R.; Kegel, K.B.; Aronin, N.; DiFiglia, M. Huntingtin bodies sequester vesicle-associated proteins by a polyproline-dependent interaction. J. Neurosci. 2004, 24, 269–281. [Google Scholar] [CrossRef]
- Liu, Y.F. Expression of polyglutamine-expanded Huntingtin activates the SEK1-JNK pathway and induces apoptosis in a hippocampal neuronal cell line. J. Biol. Chem. 1998, 273, 28873–28877. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Morfini, G.A.; You, Y.-M.; Pollema, S.L.; Kaminska, A.; Liu, K.; Yoshioka, K.; Björkblom, B.; Coffey, E.T.; Bagnato, C.; Han, D. Pathogenic huntingtin inhibits fast axonal transport by activating JNK3 and phosphorylating kinesin. Nat. Neurosci. 2009, 12, 864–871. [Google Scholar] [CrossRef]
- Gelman, A.; Rawet-Slobodkin, M.; Elazar, Z. Huntingtin facilitates selective autophagy. Nat. Cell Biol. 2015, 17, 214–215. [Google Scholar] [CrossRef]
- Rui, Y.N.; Xu, Z.; Patel, B.; Chen, Z.; Chen, D.; Tito, A.; David, G.; Sun, Y.; Stimming, E.F.; Bellen, H.J.; et al. Huntingtin functions as a scaffold for selective macroautophagy. Nat. Cell Biol. 2015, 17, 262–275. [Google Scholar] [CrossRef] [PubMed]
- Ochaba, J.; Lukacsovich, T.; Csikos, G.; Zheng, S.; Margulis, J.; Salazar, L.; Mao, K.; Lau, A.L.; Yeung, S.Y.; Humbert, S.; et al. Potential function for the Huntingtin protein as a scaffold for selective autophagy. Proc. Natl. Acad. Sci. USA 2014, 111, 16889–16894. [Google Scholar] [CrossRef] [PubMed]
- Braatz, E.M.; André, E.A.; Liu, J.P.; Zeitlin, S.O. Characterization of a knock-in mouse model with a huntingtin exon 1 deletion. J. Huntingt. Dis. 2021, 10, 435–454. [Google Scholar] [CrossRef] [PubMed]
- Atwal, R.S.; Truant, R. A stress sensitive ER membrane-association domain in Huntingtin protein defines a potential role for Huntingtin in the regulation of autophagy. Autophagy 2008, 4, 91–93. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Barron, J.C.; Hurley, E.P.; Parsons, M.P. Huntingtin and the synapse. Front. Cell Neurosci. 2021, 15, 225. [Google Scholar] [CrossRef]
- McAdam, R.L.; Morton, A.; Gordon, S.L.; Alterman, J.F.; Khvorova, A.; Cousin, M.A.; Smillie, K.J. Loss of huntingtin function slows synaptic vesicle endocytosis in striatal neurons from the htt(Q140/Q140) mouse model of Huntington’s disease. Neurobiol. Dis. 2020, 134, 104637. [Google Scholar] [CrossRef]
- Bonfanti, S.; Lionetti, M.C.; Fumagalli, M.R.; Chirasani, V.R.; Tiana, G.; Dokholyan, N.V.; Zapperi, S.; La Porta, C.A.M. Molecular mechanisms of heterogeneous oligomerization of huntingtin proteins. Sci. Rep. 2019, 9, 7615. [Google Scholar] [CrossRef]
- Bečanović, K.; Nørremølle, A.; Neal, S.J.; Kay, C.; Collins, J.A.; Arenillas, D.; Lilja, T.; Gaudenzi, G.; Manoharan, S.; Doty, C.N.; et al. A SNP in the HTT promoter alters NF-κB binding and is a bidirectional genetic modifier of Huntington disease. Nat. Neurosci. 2015, 18, 807–816. [Google Scholar] [CrossRef] [PubMed]
- O’Regan, G.C.; Farag, S.H.; Ostroff, G.R.; Tabrizi, S.J.; Andre, R. Wild-type huntingtin regulates human macrophage function. Sci Rep. 2020, 10, 17269. [Google Scholar] [CrossRef]
- Bensalel, J.; Xu, H.; Lu, M.L.; Capobianco, E.; Wei, J. RNA-seq analysis reveals significant transcriptome changes in huntingtin-null human neuroblastoma cells. BMC Med. Genom. 2021, 14, 176. [Google Scholar] [CrossRef] [PubMed]
- Guo, Q.; Huang, B.; Cheng, J.; Seefelder, M.; Engler, T.; Pfeifer, G.; Oeckl, P.; Otto, M.; Moser, F.; Maurer, M. The cryo-electron microscopy structure of huntingtin. Nature 2018, 555, 117–120. [Google Scholar] [CrossRef]
- Kaemmerer, W.F.; Grondin, R.C. The effects of huntingtin-lowering: What do we know so far? Degener Neurol. Neuromuscul. Dis. 2019, 9, 3–17. [Google Scholar] [CrossRef]
- Burtscher, J.; Di Pardo, A.; Maglione, V.; Schwarzer, C.; Squitieri, F. Mitochondrial respiration changes in R6/2 Huntington’s disease model mice during aging in a brain region specific manner. Int. J. Mol. Sci. 2020, 21, 5412. [Google Scholar] [CrossRef]
- Cherubini, M.; Lopez-Molina, L.; Gines, S. Mitochondrial fission in Huntington’s disease mouse striatum disrupts ER-mitochondria contacts leading to disturbances in Ca2+ efflux and reactive oxygen species (ROS) homeostasis. Neurobiol. Dis. 2020, 136, 104741. [Google Scholar] [CrossRef] [PubMed]
- Costa, V.; Giacomello, M.; Hudec, R.; Lopreiato, R.; Ermak, G.; Lim, D.; Malorni, W.; Davies, K.J.; Carafoli, E.; Scorrano, L. Mitochondrial fission and cristae disruption increase the response of cell models of Huntington’s disease to apoptotic stimuli. EMBO Mol. Med. 2010, 2, 490–503. [Google Scholar] [CrossRef]
- Damiano, M.; Galvan, L.; Déglon, N.; Brouillet, E. Mitochondria in Huntington’s disease. Biochim. Biophys Acta 2010, 1802, 52–61. [Google Scholar] [CrossRef] [PubMed]
- Jędrak, P.; Krygier, M.; Tońska, K.; Drozd, M.; Kaliszewska, M.; Bartnik, E.; Sołtan, W.; Sitek, E.J.; Stanisławska-Sachadyn, A.; Limon, J.; et al. Mitochondrial DNA levels in Huntington disease leukocytes and dermal fibroblasts. Metab. Brain Dis. 2017, 32, 1237–1247. [Google Scholar] [CrossRef]
- Kim, J.; Moody, J.P.; Edgerly, C.K.; Bordiuk, O.L.; Cormier, K.; Smith, K.; Beal, M.F.; Ferrante, R.J. Mitochondrial loss, dysfunction and altered dynamics in Huntington’s disease. Hum. Mol. Genet. 2010, 19, 3919–3935. [Google Scholar] [CrossRef] [PubMed]
- Song, W.; Chen, J.; Petrilli, A.; Liot, G.; Klinglmayr, E.; Zhou, Y.; Poquiz, P.; Tjong, J.; Pouladi, M.A.; Hayden, M.R.; et al. Mutant huntingtin binds the mitochondrial fission GTPase dynamin-related protein-1 and increases its enzymatic activity. Nat. Med. 2011, 17, 377–382. [Google Scholar] [CrossRef]
- Jodeiri Farshbaf, M.; Ghaedi, K. Huntington’s disease and mitochondria. Neurotox. Res. 2017, 32, 518–529. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.-Q.; Chen, Q.; Wang, X.; Wang, Q.-C.; Wang, Y.; Cheng, H.-P.; Guo, C.; Sun, Q.; Chen, Q.; Tang, T.-S. Dysregulation of mitochondrial calcium signaling and superoxide flashes cause mitochondrial genomic DNA damage in Huntington disease. J. Biol. Chem. 2013, 288, 3070–3084. [Google Scholar] [CrossRef]
- Elena-Real, C.A.; Díaz-Quintana, A.; González-Arzola, K.; Velázquez-Campoy, A.; Orzáez, M.; López-Rivas, A.; Gil-Caballero, S.; De la Rosa, M.Á.; Díaz-Moreno, I. Cytochrome c speeds up caspase cascade activation by blocking 14-3-3ε-dependent Apaf-1 inhibition. Cell Death Dis. 2018, 9, 365. [Google Scholar] [CrossRef] [PubMed]
- Ryu, H.; Rosas, H.D.; Hersch, S.M.; Ferrante, R.J. The therapeutic role of creatine in Huntington’s disease. Pharm 2005, 108, 193–207. [Google Scholar] [CrossRef]
- Palmer, C.S.; Anderson, A.J.; Stojanovski, D. Mitochondrial protein import dysfunction: Mitochondrial disease, neurodegenerative disease and cancer. FEBS Lett. 2021, 595, 1107–1131. [Google Scholar] [CrossRef] [PubMed]
- Franco-Iborra, S.; Plaza-Zabala, A.; Montpeyo, M.; Sebastian, D.; Vila, M.; Martinez-Vicente, M. Mutant HTT (huntingtin) impairs mitophagy in a cellular model of Huntington disease. Autophagy 2021, 17, 672–689. [Google Scholar] [CrossRef] [PubMed]
- Ismailoglu, I.; Chen, Q.; Popowski, M.; Yang, L.; Gross, S.S.; Brivanlou, A.H. Huntingtin protein is essential for mitochondrial metabolism, bioenergetics and structure in murine embryonic stem cells. Dev. Biol. 2014, 391, 230–240. [Google Scholar] [CrossRef]
- Kim, A.; Lalonde, K.; Truesdell, A.; Gomes Welter, P.; Brocardo, P.S.; Rosenstock, T.R.; Gil-Mohapel, J. New avenues for the treatment of Huntington’s disease. Int. J. Mol. Sci. 2021, 22, 8363. [Google Scholar] [CrossRef]
- Shin, J.W.; Kim, K.-H.; Chao, M.J.; Atwal, R.S.; Gillis, T.; MacDonald, M.E.; Gusella, J.F.; Lee, J.-M. Permanent inactivation of Huntington’s disease mutation by personalized allele-specific CRISPR/Cas9. Hum. Mol. Genet. 2016, 25, 4566–4576. [Google Scholar] [CrossRef] [PubMed]
- Monteys, A.M.; Ebanks, S.A.; Keiser, M.S.; Davidson, B.L. CRISPR/Cas9 editing of the mutant huntingtin allele in vitro and in vivo. Mol. Ther. 2017, 25, 12–23. [Google Scholar] [CrossRef]
- Yang, S.; Chang, R.; Yang, H.; Zhao, T.; Hong, Y.; Kong, H.E.; Sun, X.; Qin, Z.; Jin, P.; Li, S. CRISPR/Cas9-mediated gene editing ameliorates neurotoxicity in mouse model of Huntington’s disease. J. Clin. Investig. 2017, 127, 2719–2724. [Google Scholar] [CrossRef]
- Ekman, F.K.; Ojala, D.S.; Adil, M.M.; Lopez, P.A.; Schaffer, D.V.; Gaj, T. CRISPR-Cas9-mediated genome editing increases lifespan and improves motor deficits in a Huntington’s disease mouse model. Mol. Ther.-Nucleic Acids 2019, 17, 829–839. [Google Scholar] [CrossRef]
- Agustín-Pavón, C.; Mielcarek, M.; Garriga-Canut, M.; Isalan, M. Deimmunization for gene therapy: Host matching of synthetic zinc finger constructs enables long-term mutant Huntingtin repression in mice. Mol. Neurodegener 2016, 11, 64. [Google Scholar] [CrossRef] [PubMed]
- Garriga-Canut, M.; Agustín-Pavón, C.; Herrmann, F.; Sánchez, A.; Dierssen, M.; Fillat, C.; Isalan, M. Synthetic zinc finger repressors reduce mutant huntingtin expression in the brain of R6/2 mice. Proc. Natl. Acad. Sci. USA 2012, 109, E3136–E3145. [Google Scholar] [CrossRef] [PubMed]
- Huang, B.; Schiefer, J.; Sass, C.; Landwehrmeyer, G.B.; Kosinski, C.M.; Kochanek, S. High-capacity adenoviral vector-mediated reduction of huntingtin aggregate load in vitro and in vivo. Hum. Gene Ther. 2007, 18, 303–311. [Google Scholar] [CrossRef] [PubMed]
- Harper, S.Q.; Staber, P.D.; He, X.; Eliason, S.L.; Martins, I.H.; Mao, Q.; Yang, L.; Kotin, R.M.; Paulson, H.L.; Davidson, B.L. RNA interference improves motor and neuropathological abnormalities in a Huntington’s disease mouse model. Proc. Natl. Acad. Sci. USA 2005, 102, 5820–5825. [Google Scholar] [CrossRef]
- Drouet, V.; Perrin, V.; Hassig, R.; Dufour, N.; Auregan, G.; Alves, S.; Bonvento, G.; Brouillet, E.; Luthi-Carter, R.; Hantraye, P. Sustained effects of nonallele-specific Huntingtin silencing. Ann. Neurol. Off. J. Am. Neurol. Assoc. Child. Neurol. Soc. 2009, 65, 276–285. [Google Scholar]
- DiFiglia, M.; Sena-Esteves, M.; Chase, K.; Sapp, E.; Pfister, E.; Sass, M.; Yoder, J.; Reeves, P.; Pandey, R.K.; Rajeev, K.G. Therapeutic silencing of mutant huntingtin with siRNA attenuates striatal and cortical neuropathology and behavioral deficits. Proc. Natl. Acad. Sci. USA 2007, 104, 17204–17209. [Google Scholar] [CrossRef] [PubMed]
- Yu, D.; Pendergraff, H.; Liu, J.; Kordasiewicz, H.B.; Cleveland, D.W.; Swayze, E.E.; Lima, W.F.; Crooke, S.T.; Prakash, T.P.; Corey, D.R. Single-stranded RNAs use RNAi to potently and allele-selectively inhibit mutant huntingtin expression. Cell 2012, 150, 895–908. [Google Scholar] [CrossRef]
- McBride, J.L.; Boudreau, R.L.; Harper, S.Q.; Staber, P.D.; Monteys, A.M.; Martins, I.; Gilmore, B.L.; Burstein, H.; Peluso, R.W.; Polisky, B. Artificial miRNAs mitigate shRNA-mediated toxicity in the brain: Implications for the therapeutic development of RNAi. Proc. Natl. Acad. Sci. USA 2008, 105, 5868–5873. [Google Scholar] [CrossRef]
- Aguiar, S.; van der Gaag, B.; Cortese, F.A.B. RNAi mechanisms in Huntington’s disease therapy: siRNA versus shRNA. Transl. Neurodegener. 2017, 6, 30. [Google Scholar] [CrossRef] [PubMed]
- Keskin, S.; Brouwers, C.C.; Sogorb-Gonzalez, M.; Martier, R.; Depla, J.A.; Vallès, A.; van Deventer, S.J.; Konstantinova, P.; Evers, M.M. AAV5-miHTT lowers huntingtin mRNA and protein without off-target effects in patient-derived neuronal cultures and astrocytes. Mol. Ther.-Methods Clin. Dev. 2019, 15, 275–284. [Google Scholar] [CrossRef]
- Matthes, F.; Massari, S.; Bochicchio, A.; Schorpp, K.; Schilling, J.; Weber, S.; Offermann, N.; Desantis, J.; Wanker, E.; Carloni, P.; et al. Reducing mutant huntingtin protein expression in living cells by a newly identified RNA CAG binder. ACS Chem. Neurosci. 2018, 9, 1399–1408. [Google Scholar] [CrossRef]
- Lopes, C.; Tang, Y.; Anjo, S.I.; Manadas, B.; Onofre, I.; De Almeida, L.P.; Daley, G.Q.; Schlaeger, T.M.; Rego, A.C.C. Mitochondrial and redox modifications in huntington disease induced pluripotent stem cells rescued by CRISPR/Cas9 CAGs targeting. Front. Cell Dev. Biol. 2020, 8, 967. [Google Scholar] [CrossRef]
- Dos Santos, N.T.H.; do Bomfim, F.R.C. Gene editing by CRISPR/CAS9 for treatment of Huntington disease. Int. J. Dev. Res. 2020, 10, 38631–38635. [Google Scholar]
- Jamwal, S.; Elsworth, J.D.; Rahi, V.; Kumar, P. Gene therapy and immunotherapy as promising strategies to combat Huntington’s disease-associated neurodegeneration: Emphasis on recent updates and future perspectives. Expert Rev. Neurother. 2020, 20, 1123–1141. [Google Scholar] [CrossRef] [PubMed]
- Marxreiter, F.; Stemick, J.; Kohl, Z. Huntingtin lowering strategies. Int. J. Mol. Sci. 2020, 21, 2146. [Google Scholar] [CrossRef] [PubMed]
- Leavitt, B.R.; Kordasiewicz, H.B.; Schobel, S.A. Huntingtin-lowering therapies for Huntington disease: A review of the evidence of potential benefits and risks. JAMA Neurol. 2020, 77, 764–772. [Google Scholar] [CrossRef]
- Tabrizi, S.J.; Flower, M.D.; Ross, C.A.; Wild, E.J. Huntington disease: New insights into molecular pathogenesis and therapeutic opportunities. Nat. Rev. Neurol. 2020, 16, 529–546. [Google Scholar] [CrossRef]
- Tabrizi, S.J.; Ghosh, R.; Leavitt, B.R. Huntingtin lowering strategies for disease modification in Huntington’s disease. Neuron 2019, 101, 801–819. [Google Scholar] [CrossRef] [PubMed]
- Estevez-Fraga, C.; Flower, M.D.; Tabrizi, S.J. Therapeutic strategies for Huntington’s disease. Curr. Opin. Neurol. 2020, 33, 508–518. [Google Scholar] [CrossRef]
- Evers, M.M.; Konstantinova, P. AAV5-miHTT gene therapy for Huntington disease: Lowering both huntingtins. Expert Opin. Biol. Ther. 2020, 20, 1121–1124. [Google Scholar] [CrossRef] [PubMed]
- Barker, R.; Fujimaki, M.; Rogers, P.; Rubinsztein, D. Huntingtin-lowering strategies for Huntington’s disease. Expert Opin. Investig. Drugs 2020, 29, 1125–1132. [Google Scholar] [CrossRef] [PubMed]
- Smith, A.V.; Tabrizi, S.J. Therapeutic antisense targeting of huntingtin. DNA Cell Biol. 2020, 39, 154–158. [Google Scholar] [CrossRef]
- Fields, E.; Vaughan, E.; Tripu, D.; Lim, I.; Shrout, K.; Conway, J.; Salib, N.; Lee, Y.; Dhamsania, A.; Jacobsen, M. Gene targeting techniques for Huntington’s disease. Ageing Res. Rev. 2021, 70, 101385. [Google Scholar] [CrossRef]
- Wild, E.J.; Tabrizi, S.J. Therapies targeting DNA and RNA in Huntington’s disease. Lancet Neurol. 2017, 16, 837–847. [Google Scholar] [CrossRef]
- Gardian, G.; Browne, S.E.; Choi, D.-K.; Klivenyi, P.; Gregorio, J.; Kubilus, J.K.; Ryu, H.; Langley, B.; Ratan, R.R.; Ferrante, R.J. Neuroprotective effects of phenylbutyrate in the N171-82Q transgenic mouse model of Huntington’s disease. J. Biol. Chem. 2005, 280, 556–563. [Google Scholar] [CrossRef]
- Ferrante, R.J.; Kubilus, J.K.; Lee, J.; Ryu, H.; Beesen, A.; Zucker, B.; Smith, K.; Kowall, N.W.; Ratan, R.R.; Luthi-Carter, R. Histone deacetylase inhibition by sodium butyrate chemotherapy ameliorates the neurodegenerative phenotype in Huntington’s disease mice. J. Neurosci. 2003, 23, 9418–9427. [Google Scholar] [CrossRef]
- Thomas, E.A.; Coppola, G.; Desplats, P.A.; Tang, B.; Soragni, E.; Burnett, R.; Gao, F.; Fitzgerald, K.M.; Borok, J.F.; Herman, D. The HDAC inhibitor 4b ameliorates the disease phenotype and transcriptional abnormalities in Huntington’s disease transgenic mice. Proc. Natl. Acad. Sci. USA 2008, 105, 15564–15569. [Google Scholar] [CrossRef]
- Jia, H.; Morris, C.D.; Williams, R.M.; Loring, J.F.; Thomas, E.A. HDAC inhibition imparts beneficial transgenerational effects in Huntington’s disease mice via altered DNA and histone methylation. Proc. Natl. Acad. Sci. USA 2015, 112, E56–E64. [Google Scholar] [CrossRef] [PubMed]
- Jia, H.; Pallos, J.; Jacques, V.; Lau, A.; Tang, B.; Cooper, A.; Syed, A.; Purcell, J.; Chen, Y.; Sharma, S. Histone deacetylase (HDAC) inhibitors targeting HDAC3 and HDAC1 ameliorate polyglutamine-elicited phenotypes in model systems of Huntington’s disease. Neurobiol. Dis. 2012, 46, 351–361. [Google Scholar] [CrossRef] [PubMed]
- Siebzehnrübl, F.A.; Raber, K.A.; Urbach, Y.K.; Schulze-Krebs, A.; Canneva, F.; Moceri, S.; Habermeyer, J.; Achoui, D.; Gupta, B.; Steindler, D.A. Early postnatal behavioral, cellular, and molecular changes in models of Huntington disease are reversible by HDAC inhibition. Proc. Natl. Acad. Sci. USA 2018, 115, E8765–E8774. [Google Scholar] [CrossRef]
- Guedes-Dias, P.; de Proença, J.; Soares, T.R.; Leitão-Rocha, A.; Pinho, B.R.; Duchen, M.R.; Oliveira, J.M. HDAC6 inhibition induces mitochondrial fusion, autophagic flux and reduces diffuse mutant huntingtin in striatal neurons. Biochim. Et Biophys. Acta (BBA)-Mol. Basis Dis. 2015, 1852, 2484–2493. [Google Scholar] [CrossRef] [PubMed]
- Ferrante, R.J.; Ryu, H.; Kubilus, J.K.; D’Mello, S.; Sugars, K.L.; Lee, J.; Lu, P.; Smith, K.; Browne, S.; Beal, M.F. Chemotherapy for the brain: The antitumor antibiotic mithramycin prolongs survival in a mouse model of Huntington’s disease. J. Neurosci. 2004, 24, 10335–10342. [Google Scholar] [CrossRef]
- Heiser, V.; Scherzinger, E.; Boeddrich, A.; Nordhoff, E.; Lurz, R.; Schugardt, N.; Lehrach, H.; Wanker, E.E. Inhibition of huntingtin fibrillogenesis by specific antibodies and small molecules: Implications for Huntington’s disease therapy. Proc. Natl. Acad. Sci. USA 2000, 97, 6739–6744. [Google Scholar] [CrossRef] [PubMed]
- Tanaka, M.; Machida, Y.; Niu, S.; Ikeda, T.; Jana, N.R.; Doi, H.; Kurosawa, M.; Nekooki, M.; Nukina, N. Trehalose alleviates polyglutamine-mediated pathology in a mouse model of Huntington disease. Nat. Med. 2004, 10, 148–154. [Google Scholar] [CrossRef] [PubMed]
- Lee, C.Y.D.; Wang, N.; Shen, K.; Stricos, M.; Langfelder, P.; Cheon, K.H.; Cortés, E.P.; Vinters, H.V.; Vonsattel, J.P.; Wexler, N.S.; et al. Disease-related Huntingtin seeding activities in cerebrospinal fluids of Huntington’s disease patients. Sci. Rep. 2020, 10, 20295. [Google Scholar] [CrossRef] [PubMed]
- Xu, X.; Ng, B.; Sim, B.; Radulescu, C.I.; Yusof, N.A.B.M.; Goh, W.I.; Lin, S.; Lim, J.S.Y.; Cha, Y.; Kusko, R.; et al. pS421 huntingtin modulates mitochondrial phenotypes and confers neuroprotection in an HD hiPSC model. Cell Death Dis. 2020, 11, 809. [Google Scholar] [CrossRef] [PubMed]
- Kratter, I.H.; Zahed, H.; Lau, A.; Tsvetkov, A.S.; Daub, A.C.; Weiberth, K.F.; Gu, X.; Saudou, F.; Humbert, S.; Yang, X.W.; et al. Serine 421 regulates mutant huntingtin toxicity and clearance in mice. J. Clin. Investig. 2016, 126, 3585–3597. [Google Scholar] [CrossRef] [PubMed]
- Weber, J.J.; Kloock, S.J.; Nagel, M.; Ortiz-Rios, M.M.; Hofmann, J.; Riess, O.; Nguyen, H.P. Calpastatin ablation aggravates the molecular phenotype in cell and animal models of Huntington disease. Neuropharmacology 2018, 133, 94–106. [Google Scholar] [CrossRef]
- Kim, M.; Lee, H.; LaForet, G.; McIntyre, C.; Martin, E.J.; Chang, P.; Kim, T.W.; Williams, M.; Reddy, P.; Tagle, D. Mutant huntingtin expression in clonal striatal cells: Dissociation of inclusion formation and neuronal survival by caspase inhibition. J. Neurosci. 1999, 19, 964–973. [Google Scholar] [CrossRef] [PubMed]
- Rieux, M.; Alpaugh, M.; Sciacca, G.; Saint-Pierre, M.; Masnata, M.; Denis, H.L.; Lévesque, S.A.; Herrmann, F.; Bazenet, C.; Garneau, A.P. Shedding a new light on Huntington’s disease: How blood can both propagate and ameliorate disease pathology. Mol. Psychiatry 2020, 26, 5441–5463. [Google Scholar] [CrossRef] [PubMed]
- Ferrante, R.J.; Andreassen, O.A.; Jenkins, B.G.; Dedeoglu, A.; Kuemmerle, S.; Kubilus, J.K.; Kaddurah-Daouk, R.; Hersch, S.M.; Beal, M.F. Neuroprotective effects of creatine in a transgenic mouse model of Huntington’s disease. J. Neurosci. 2000, 20, 4389–4397. [Google Scholar] [CrossRef]
- Koroshetz, W.J.; Jenkins, B.G.; Rosen, B.R.; Beal, M.F. Energy metabolism defects in Huntington’s disease and effects of coenzyme Q10. Ann. Neurol. Off. J. Am. Neurol. Assoc. Child. Neurol. Soc. 1997, 41, 160–165. [Google Scholar] [CrossRef]
- Investigators, H.S.G.P.C. Safety and tolerability of high-dosage coenzyme Q10 in Huntington’s disease and healthy subjects. Mov. Disord. 2010, 25, 1924–1928. [Google Scholar]
- Essa, M.M.; Moghadas, M.; Ba-Omar, T.; Qoronfleh, M.W.; Guillemin, G.J.; Manivasagam, T.; Justin-Thenmozhi, A.; Ray, B.; Bhat, A.; Chidambaram, S.B. Protective effects of antioxidants in Huntington’s disease: An extensive review. Neurotox. Res. 2019, 35, 739–774. [Google Scholar] [CrossRef]
- McMeekin, L.J.; Fox, S.N.; Boas, S.M.; Cowell, R.M. Dysregulation of PGC-1α-dependent transcriptional programs in neurological and developmental disorders: Therapeutic challenges and opportunities. Cells 2021, 10, 352. [Google Scholar] [CrossRef]
- Lopes, C.; Ribeiro, M.; Duarte, A.I.; Humbert, S.; Saudou, F.; De Almeida, L.P.; Hayden, M.; Rego, A.C. IGF-1 intranasal administration rescues Huntington’s disease phenotypes in YAC128 mice. Mol. Neurobiol. 2014, 49, 1126–1142. [Google Scholar] [CrossRef] [PubMed]
- Martin, D.D.; Ladha, S.; Ehrnhoefer, D.E.; Hayden, M.R. Autophagy in Huntington disease and huntingtin in autophagy. Trends Neurosci. 2015, 38, 26–35. [Google Scholar] [CrossRef] [PubMed]
- Lo, C.H.; Pandey, N.K.; Lim, C.K.-W.; Ding, Z.; Tao, M.; Thomas, D.D.; Langen, R.; Sachs, J.N. Discovery of small molecule inhibitors of huntingtin exon 1 aggregation by FRET-Based high-throughput screening in living cells. ACS Chem. Neurosci. 2020, 11, 2286–2295. [Google Scholar] [CrossRef] [PubMed]
- Gies, E.; Wilde, I.; Winget, J.M.; Brack, M.; Rotblat, B.; Novoa, C.A.; Balgi, A.D.; Sorensen, P.H.; Roberge, M.; Mayor, T. Niclosamide prevents the formation of large ubiquitin-containing aggregates caused by proteasome inhibition. PLoS ONE 2010, 5, e14410. [Google Scholar] [CrossRef]
- Tang, T.-S.; Slow, E.; Lupu, V.; Stavrovskaya, I.G.; Sugimori, M.; Llinás, R.; Kristal, B.S.; Hayden, M.R.; Bezprozvanny, I. Disturbed Ca2+ signaling and apoptosis of medium spiny neurons in Huntington’s disease. Proc. Natl. Acad. Sci. USA 2005, 102, 2602–2607. [Google Scholar] [CrossRef]
- Farina, F.; Lambert, E.; Commeau, L.; Lejeune, F.-X.; Roudier, N.; Fonte, C.; Parker, J.A.; Boddaert, J.; Verny, M.; Baulieu, E.-E. The stress response factor daf-16/FOXO is required for multiple compound families to prolong the function of neurons with Huntington’s disease. Sci. Rep. 2017, 7, 4014. [Google Scholar] [CrossRef]
- Ehrnhoefer, D.E.; Skotte, N.H.; Reinshagen, J.; Qiu, X.; Windshügel, B.; Jaishankar, P.; Ladha, S.; Petina, O.; Khankischpur, M.; Nguyen, Y.T.N.; et al. Activation of caspase-6 is promoted by a mutant huntingtin fragment and blocked by an allosteric inhibitor compound. Cell Chem. Biol. 2019, 26, 1295–1305.e1296. [Google Scholar] [CrossRef]
- Wei, H.; Qin, Z.-H.; Senatorov, V.; Wei, W.; Wang, Y.; Qian, Y.; Chuang, D.-M. Lithium suppresses excitotoxicity-induced striatal lesions in a rat model of Huntington’s disease. Neuroscience 2001, 106, 603–612. [Google Scholar] [CrossRef]
- Ellrichmann, G.; Blusch, A.; Fatoba, O.; Brunner, J.; Reick, C.; Hayardeny, L.; Hayden, M.; Sehr, D.; Winklhofer, K.F.; Saft, C.; et al. Laquinimod treatment in the R6/2 mouse model. Sci. Rep. 2017, 7, 4947. [Google Scholar] [CrossRef]
- Megahed, M.; El-Azab, M.F.; El Sayed, M.I.; Moustafa, Y. Huntington’s disease and NMDA receptors; a new arena for therapeutic development. Rec. Pharm. Biomed. Sci. 2018, 2, 1–13. [Google Scholar] [CrossRef]
- Zhu, S.; Zhang, Y.; Bai, G.; Li, H. Necrostatin-1 ameliorates symptoms in R6/2 transgenic mouse model of Huntington’s disease. Cell Death Dis. 2011, 2, e115. [Google Scholar] [CrossRef] [PubMed]
- Beconi, M.G.; Howland, D.; Park, L.; Lyons, K.; Giuliano, J.; Dominguez, C.; Munoz-Sanjuan, I.; Pacifici, R. Pharmacokinetics of memantine in rats and mice. PLoS Curr. 2011, 3, RRN1291. [Google Scholar] [CrossRef] [PubMed]
- Reilmann, R.; Ross, C.; Testa, C.; Frank, S.; Evers, M.; de Haan, M.; Valles-Sanchez, A.; Konstantinova, P.; van Deventer, S.; Higgins, J. Translation of AMT-130 preclinical data to inform the design of the first FDA-approved human AAV gene therapy clinical trial in adults with early manifest Huntington’s disease (4531). Neurology 2020, 94, 4531. [Google Scholar]
- Tabrizi, S.J.; Leavitt, B.R.; Landwehrmeyer, G.B.; Wild, E.J.; Saft, C.; Barker, R.A.; Blair, N.F.; Craufurd, D.; Priller, J.; Rickards, H. Targeting huntingtin expression in patients with Huntington’s disease. New Engl. J. Med. 2019, 380, 2307–2316. [Google Scholar] [CrossRef] [PubMed]
- Kordasiewicz, H.B.; Stanek, L.M.; Wancewicz, E.V.; Mazur, C.; McAlonis, M.M.; Pytel, K.A.; Artates, J.W.; Weiss, A.; Cheng, S.H.; Shihabuddin, L.S. Sustained therapeutic reversal of Huntington’s disease by transient repression of huntingtin synthesis. Neuron 2012, 74, 1031–1044. [Google Scholar] [CrossRef]
- Karpuj, M.V.; Becher, M.W.; Springer, J.E.; Chabas, D.; Youssef, S.; Pedotti, R.; Mitchell, D.; Steinman, L. Prolonged survival and decreased abnormal movements in transgenic model of Huntington disease, with administration of the transglutaminase inhibitor cystamine. Nat. Med. 2002, 8, 143–149. [Google Scholar] [CrossRef]
- Dedeoglu, A.; Kubilus, J.K.; Jeitner, T.M.; Matson, S.A.; Bogdanov, M.; Kowall, N.W.; Matson, W.R.; Cooper, A.J.; Ratan, R.R.; Beal, M.F. Therapeutic effects of cystamine in a murine model of Huntington’s disease. J. Neurosci. 2002, 22, 8942–8950. [Google Scholar] [CrossRef]
- Sanchez, I.; Mahlke, C.; Yuan, J. Pivotal role of oligomerization in expanded polyglutamine neurodegenerative disorders. Nature 2003, 421, 373–379. [Google Scholar] [CrossRef]
- Chen, M.; Ona, V.O.; Li, M.; Ferrante, R.J.; Fink, K.B.; Zhu, S.; Bian, J.; Guo, L.; Farrell, L.A.; Hersch, S.M. Minocycline inhibits caspase-1 and caspase-3 expression and delays mortality in a transgenic mouse model of Huntington disease. Nat. Med. 2000, 6, 797–801. [Google Scholar] [CrossRef]
- Rea, S.; Della-Morte, D.; Pacifici, F.; Capuani, B.; Pastore, D.; Coppola, A.; Arriga, R.; Andreadi, A.; Donadel, G.; Di Daniele, N. Insulin and exendin-4 reduced mutated Huntingtin accumulation in neuronal cells. Front. Pharmacol. 2020, 11, 779. [Google Scholar] [CrossRef]
- Di Pardo, A.; Maglione, V.; Alpaugh, M.; Horkey, M.; Atwal, R.S.; Sassone, J.; Ciammola, A.; Steffan, J.S.; Fouad, K.; Truant, R. Ganglioside GM1 induces phosphorylation of mutant huntingtin and restores normal motor behavior in Huntington disease mice. Proc. Natl. Acad. Sci. USA 2012, 109, 3528–3533. [Google Scholar] [CrossRef] [PubMed]
- Ermak, G.; Hench, K.J.; Chang, K.T.; Sachdev, S.; Davies, K.J. Regulator of calcineurin (RCAN1-1L) is deficient in Huntington disease and protective against mutant huntingtin toxicity in vitro. J. Biol. Chem. 2009, 284, 11845–11853. [Google Scholar] [CrossRef]
- Rangone, H.; Poizat, G.; Troncoso, J.; Ross, C.A.; MacDonald, M.E.; Saudou, F.; Humbert, S. The serum-and glucocorticoid-induced kinase SGK inhibits mutant huntingtin-induced toxicity by phosphorylating serine 421 of huntingtin. Eur. J. Neurosci. 2004, 19, 273–279. [Google Scholar] [CrossRef]
- Dickey, A.S.; Pineda, V.V.; Tsunemi, T.; Liu, P.P.; Miranda, H.C.; Gilmore-Hall, S.K.; Lomas, N.; Sampat, K.R.; Buttgereit, A.; Torres, M.-J.M. PPAR-δ is repressed in Huntington’s disease, is required for normal neuronal function and can be targeted therapeutically. Nat. Med. 2016, 22, 37–45. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.J.; Li, J. Caspase blockade induces RIP3-mediated programmed necrosis in Toll-like receptor-activated microglia. Cell Death Dis. 2013, 4, e716. [Google Scholar] [CrossRef] [PubMed]
- Morigaki, R.; Goto, S. Striatal vulnerability in Huntington’s disease: Neuroprotection versus neurotoxicity. Brain Sci. 2017, 7, 63. [Google Scholar] [CrossRef] [PubMed]
- Casella, C.; Lipp, I.; Rosser, A.; Jones, D.K.; Metzler-Baddeley, C. A critical review of white matter changes in huntington’s disease. Mov. Disord. 2020, 35, 1302–1311. [Google Scholar] [CrossRef] [PubMed]
- Franklin, G.L.; Camargo, C.H.F.; Meira, A.T.; Lima, N.S.; Teive, H.A. The role of the cerebellum in Huntington’s disease: A systematic review. Cerebellum 2021, 20, 254–265. [Google Scholar] [CrossRef]
- Reiner, A.; Deng, Y.P. Disrupted striatal neuron inputs and outputs in Huntington’s disease. CNS Neurosci. Ther. 2018, 24, 250–280. [Google Scholar] [CrossRef]
- Yang, H.-M.; Yang, S.; Huang, S.-S.; Tang, B.-S.; Guo, J.-F. Microglial Activation in the Pathogenesis of Huntington’s Disease. Front. Aging Neurosci. 2017, 9, 193. [Google Scholar] [CrossRef] [PubMed]
- Cepeda, C.; Levine, M.S. Synaptic dysfunction in Huntington’s disease: Lessons from genetic animal models. Neuroscience 2020, 1–21. [Google Scholar] [CrossRef] [PubMed]
- Tang, B.L. Unconventional secretion and intercellular transfer of mutant huntingtin. Cells 2018, 7, 59. [Google Scholar] [CrossRef]
- Wanker, E.E.; Ast, A.; Schindler, F.; Trepte, P.; Schnoegl, S. The pathobiology of perturbed mutant huntingtin protein–protein interactions in Huntington’s disease. J. Neurochem. 2019, 151, 507–519. [Google Scholar] [CrossRef]
- Pandey, M.; Rajamma, U. Huntington’s disease: The coming of age. J. Genet. 2018, 97, 649–664. [Google Scholar] [CrossRef]
- Tellone, E.; Galtieri, A.; Ficarra, S. Reviewing biochemical implications of normal and mutated Huntingtin in Huntington’s disease. Curr. Med. Chem. 2020, 27, 5137–5158. [Google Scholar] [CrossRef]
- Ehrnhoefer, D.E.; Sutton, L.; Hayden, M.R. Small changes, big impact: Posttranslational modifications and function of huntingtin in Huntington disease. Neurosci. 2011, 17, 475–492. [Google Scholar] [CrossRef]
- Soares, T.R.; Reis, S.D.; Pinho, B.R.; Duchen, M.R.; Oliveira, J.M.A. Targeting the proteostasis network in Huntington’s disease. Ageing Res. Rev. 2019, 49, 92–103. [Google Scholar] [CrossRef] [PubMed]
- Rui, Y.-N.; Xu, Z.; Patel, B.; Cuervo, A.M.; Zhang, S. HTT/Huntingtin in selective autophagy and Huntington disease: A foe or a friend within? Autophagy 2015, 11, 858–860. [Google Scholar] [CrossRef]
- Valionyte, E.; Yang, Y.; Roberts, S.L.; Kelly, J.; Lu, B.; Luo, S. Lowering mutant huntingtin levels and toxicity: Autophagy-endolysosome pathways in Huntington’s disease. J. Mol. Biol. 2020, 432, 2673–2691. [Google Scholar] [CrossRef]
- Paul, B.D.; Snyder, S.H. Impaired redox signaling in Huntington’s disease: Therapeutic implications. Front. Mol. Neurosci. 2019, 12, 68. [Google Scholar] [CrossRef] [PubMed]
- Dubinsky, J.M. Towards an understanding of energy impairment in Huntington’s disease brain. J. Huntingt. Dis. 2017, 6, 267–302. [Google Scholar] [CrossRef] [PubMed]
- Singh, A.; Agrawal, N. Metabolism in Huntington’s disease: A major contributor to pathology. Metab. Brain Dis. 2021, 1–15. [Google Scholar] [CrossRef]
- Romo, L.; Mohn, E.S.; Aronin, N. A fresh look at Huntingtin mRNA processing in Huntington’s disease. J. Huntingt. Dis. 2018, 7, 101–108. [Google Scholar] [CrossRef]
- Heinz, A.; Nabariya, D.K.; Krauss, S. Huntingtin and its role in mechanisms of RNA-mediated toxicity. Toxins 2021, 13, 487. [Google Scholar] [CrossRef]
- Valadão, P.A.C.; Santos, K.B.S.; E Vieira, T.H.F.; E Cordeiro, T.M.; Teixeira, A.L.; Guatimosim, C.; de Miranda, A.S. Inflammation in Huntington’s disease: A few new twists on an old tale. J. Neuroimmunol. 2020, 348, 577380. [Google Scholar] [CrossRef]
- Mattis, V.B.; Svendsen, C.N. Modeling Huntington’s disease with patient-derived neurons. Brain Res. 2017, 1656, 76–87. [Google Scholar] [CrossRef]
- Monk, R.; Connor, B. Cell Reprogramming to Model Huntington’s Disease: A Comprehensive Review. Cells 2021, 10, 1565. [Google Scholar] [CrossRef] [PubMed]
- Kandasamy, M.; Aigner, L. Reactive Neuroblastosis in Huntington’s Disease: A Putative Therapeutic Target for Striatal Regeneration in the Adult Brain. Front. Cell. Neurosci. 2018, 12, 37. [Google Scholar] [CrossRef] [PubMed]
- Connor, B. Concise review: The use of stem cells for understanding and treating Huntington’s disease. Stem Cells 2018, 36, 146–160. [Google Scholar] [CrossRef] [PubMed]
- Colpo, G.D.; Stimming, E.F.; Teixeira, A.L. Stem cells in animal models of Huntington disease: A systematic review. Mol. Cell. Neurosci. 2019, 95, 43–50. [Google Scholar] [CrossRef]
- Gorantla, V.R.; Bhat, A.; Ghosh, A.R.; Bolla, S.R.; Bhojaraj, S.; Mohan, S.K.; Veeraraghavan, V.P.; Chidambaram, S.B.; Essa, M.M.; Qoronfleh, M.W. Stem cells therapy: A ray of hope for Huntington disease. Int. J. Nutr. Pharmacol. Neurol. Dis. 2021, 11, 95. [Google Scholar]
- Albin, R.L.; Kordower, J.H. A. Failed Future. Mov. Disord. 2020, 35, 1299–1301. [Google Scholar] [CrossRef] [PubMed]
- Rosser, A.E.; Busse, M.; Aron Badin, R.; Canals, J.M.; Wheelock, V.; Perrier, A.L.; Gray, W.; Thompson, L.; Goldman, S. Cell therapy for Huntington’s disease: Learning from failure. Mov. Disord. 2021, 36, 787–788. [Google Scholar] [CrossRef]
- Csobonyeiova, M.; Polak, S.; Danisovic, L. Recent overview of the use of iPSCs Huntington’s disease modeling and therapy. Int. J. Mol. Sci. 2020, 21, 2239. [Google Scholar] [CrossRef] [PubMed]
- Srinageshwar, B.; Petersen, R.B.; Dunbar, G.L.; Rossignol, J. Prion-like mechanisms in neurodegenerative disease: Implications for Huntington’s disease therapy. Stem Cells Transl. Med. 2020, 9, 559–566. [Google Scholar] [CrossRef] [PubMed]
- Bachoud-Lévi, A.C.; Massart, R.; Rosser, A. Cell therapy in Huntington’s disease: Taking stock of past studies to move the field forward. Stem Cells 2021, 39, 144–155. [Google Scholar]
- Choi, K.-A.; Choi, Y.; Hong, S. Stem cell transplantation for Huntington’s diseases. Methods 2018, 133, 104–112. [Google Scholar] [CrossRef]
- Wojtecki, L.; Groiss, S.J.; Hartmann, C.J.; Elben, S.; Omlor, S.; Schnitzler, A.; Vesper, J. Deep brain stimulation in Huntington’s disease—preliminary evidence on pathophysiology, efficacy and safety. Brain Sci. 2016, 6, 38. [Google Scholar] [CrossRef]
- Pini, L.; Jacquemot, C.; Cagnin, A.; Meneghello, F.; Semenza, C.; Mantini, D.; Vallesi, A. Aberrant brain network connectivity in presymptomatic and manifest Huntington’s disease: A systematic review. Hum. Brain Mapp. 2020, 41, 256–269. [Google Scholar] [CrossRef]
- Cardoso, F. Chapter fifty-Nonmotor symptoms in Huntington disease. In International Review of Neurobiology; Chaudhuri, K.R., Titova, N., Eds.; Academic Press: Cambridge, MA, 2017; Volume 134, pp. 1397–1408. [Google Scholar]
- Patil, R.S.; Vyas, S.G.; Quazi, W.T.; Tembhurnikar, H.J.; Milmile, P.S.; Umekar, M.J. The gut microbiome in Huntington disease: A review. GSC Biol. Pharm. Sci. 2021, 15, 317–326. [Google Scholar] [CrossRef]
- Schultz, J.L.; Nopoulos, P.C.; Gonzalez-Alegre, P. Human Immunodeficiency Virus Infection in Huntington’s Disease is Associated with an Earlier Age of Symptom Onset. J. Huntingt. Dis. 2018, 7, 163–166. [Google Scholar] [CrossRef] [PubMed]
- McCusker, E.A.; Loy, C.T. Huntington disease: The complexities of making and disclosing a clinical diagnosis after premanifest genetic testing. Tremor Other Hyperkinet Mov. 2017, 7, 467. [Google Scholar] [CrossRef]
- Przybyl, L.; Wozna-Wysocka, M.; Kozlowska, E.; Fiszer, A. What, when and how to measure—peripheral biomarkers in therapy of Huntington’s disease. Int. J. Mol. Sci. 2021, 22, 1561. [Google Scholar] [CrossRef] [PubMed]
- Schobel, S.A.; Palermo, G.; Auinger, P.; Long, J.D.; Ma, S.; Khwaja, O.S.; Trundell, D.; Cudkowicz, M.; Hersch, S.; Sampaio, C.; et al. Motor, cognitive, and functional declines contribute to a single progressive factor in early HD. Neurology 2017, 89, 2495–2502. [Google Scholar] [CrossRef]
- Claassen, D.O.; Iyer, R.G.; Shah-Manek, B.; DiBonaventura, M.; Abler, V.; Sung, V.W. Tetrabenazine treatment patterns and outcomes for chorea associated with Huntington disease: A retrospective chart review. J. Huntingt. Dis. 2018, 7, 345–353. [Google Scholar] [CrossRef] [PubMed]
- Quinn, L.; Kegelmeyer, D.; Kloos, A.; Rao, A.K.; Busse, M.; Fritz, N.E. Clinical recommendations to guide physical therapy practice for Huntington disease. Neurology 2020, 94, 217–228. [Google Scholar] [CrossRef]
- Zarotti, N.; Dale, M.; Eccles, F.; Simpson, J. Psychological Interventions for People with Huntington’s Disease: A Call to Arms. J. Huntingt. Dis. 2020, 9, 231–243. [Google Scholar] [CrossRef]
- Eccles, F.J.R.; Craufurd, D.; Smith, A.; Davies, R.; Glenny, K.; Homberger, M.; Peeren, S.; Rogers, D.; Rose, L.; Skitt, Z.; et al. A feasibility investigation of mindfulness-based cognitive therapy for people with Huntington’s disease. Pilot Feasibility Stud. 2020, 6, 90. [Google Scholar] [CrossRef]
- Rossi, G.; Oh, J.C. Management of agitation in Huntington’s disease: A review of the literature. Cureus 2020, 12, e9748. [Google Scholar] [CrossRef] [PubMed]
- Liu, Q.; Cheng, S.; Yang, H.; Zhu, L.; Pan, Y.; Jing, L.; Tang, B.; Li, S.; Li, X.-J. Loss of Hap1 selectively promotes striatal degeneration in Huntington disease mice. Proc. Natl. Acad. Sci. USA 2020, 117, 20265–20273. [Google Scholar] [CrossRef] [PubMed]
- Yang, H.; Yang, S.; Jing, L.; Huang, L.; Chen, L.; Zhao, X.; Yang, W.; Pan, Y.; Yin, P.; Qin, Z.S.; et al. Truncation of mutant huntingtin in knock-in mice demonstrates exon1 huntingtin is a key pathogenic form. Nat. Commun. 2020, 11, 2582. [Google Scholar] [CrossRef]
- Sharma, M.; Subramaniam, S. Rhes travels from cell to cell and transports Huntington disease protein via TNT-like protrusion. J. Cell Biol. 2019, 218, 1972–1993. [Google Scholar] [CrossRef] [PubMed]
- Sharma, M.; Rajendrarao, S.; Shahani, N.; Ramírez-Jarquín, U.N.; Subramaniam, S. Cyclic GMP-AMP synthase promotes the inflammatory and autophagy responses in Huntington disease. Proc. Natl. Acad. Sci. USA 2020, 117, 15989–15999. [Google Scholar] [CrossRef] [PubMed]
- Lee, H.; Noh, J.-Y.; Oh, Y.; Kim, Y.; Chang, J.-W.; Chung, C.-W.; Lee, S.-T.; Kim, M.; Ryu, H.; Jung, Y.-K. IRE1 plays an essential role in ER stress-mediated aggregation of mutant huntingtin via the inhibition of autophagy flux. Hum. Mol. Genet. 2011, 21, 101–114. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.; Kosaras, B.; Del Signore, S.J.; Cormier, K.; McKee, A.; Ratan, R.R.; Kowall, N.W.; Ryu, H. Modulation of lipid peroxidation and mitochondrial function improves neuropathology in Huntington’s disease mice. Acta Neuropathol. 2011, 121, 487–498. [Google Scholar] [CrossRef] [PubMed]
- Jeon, G.S.; Kim, K.Y.; Hwang, Y.J.; Jung, M.-K.; An, S.; Ouchi, M.; Ouchi, T.; Kowall, N.; Lee, J.; Ryu, H. Deregulation of BRCA1 leads to impaired spatiotemporal dynamics of γ-H2AX and DNA damage responses in Huntington’s disease. Mol. Neurobiol. 2012, 45, 550–563. [Google Scholar] [CrossRef]
- Hyeon, S.J.; Park, J.; Yoo, J.; Kim, S.-H.; Hwang, Y.J.; Kim, S.-C.; Liu, T.; Shim, H.S.; Kim, Y.; Cho, Y.; et al. Dysfunction of X-linked inhibitor of apoptosis protein (XIAP) triggers neuropathological processes via altered p53 activity in Huntington’s disease. Prog. Neurobiol. 2021, 204, 102110. [Google Scholar] [CrossRef] [PubMed]
- Mario Isas, J.; Pandey, N.K.; Xu, H.; Teranishi, K.; Okada, A.K.; Fultz, E.K.; Rawat, A.; Applebaum, A.; Meier, F.; Chen, J.; et al. Huntingtin fibrils with different toxicity, structure, and seeding potential can be interconverted. Nat. Commun. 2021, 12, 4272. [Google Scholar] [CrossRef] [PubMed]
- Monteiro, O.; Chen, C.; Bingham, R.; Argyrou, A.; Buxton, R.; Jönsson, C.P.; Jones, E.; Bridges, A.; Gatfield, K.; Krauß, S. Pharmacological disruption of the MID1/α4 interaction reduces mutant Huntingtin levels in primary neuronal cultures. Neurosci. Lett. 2018, 673, 44–50. [Google Scholar] [CrossRef]
- Miyazaki, H.; Yamanaka, T.; Oyama, F.; Kino, Y.; Kurosawa, M.; Yamada-Kurosawa, M.; Yamano, R.; Shimogori, T.; Hattori, N.; Nukina, N. FACS-array–based cell purification yields a specific transcriptome of striatal medium spiny neurons in a murine Huntington disease model. J. Biol. Chem. 2020, 295, 9768–9785. [Google Scholar] [CrossRef]
- Critchley, B.J.; Isalan, M.; Mielcarek, M. Neuro-cardio mechanisms in Huntington’s disease and other neurodegenerative disorders. Front. Physiol. 2018, 9, 559. [Google Scholar] [CrossRef] [PubMed]
- Hsu, Y.T.; Chang, Y.G.; Chern, Y. Insights into GABA(A)ergic system alteration in Huntington’s disease. Open Biol. 2018, 8, 180165. [Google Scholar] [CrossRef] [PubMed]
- Burgold, J.; Schulz-Trieglaff, E.K.; Voelkl, K.; Gutiérrez-Ángel, S.; Bader, J.M.; Hosp, F.; Mann, M.; Arzberger, T.; Klein, R.; Liebscher, S.; et al. Cortical circuit alterations precede motor impairments in Huntington’s disease mice. Sci. Rep. 2019, 9, 6634. [Google Scholar] [CrossRef] [PubMed]
- Raj, A.; Powell, F. Network model of pathology spread recapitulates neurodegeneration and selective vulnerability in Huntington’s Disease. NeuroImage 2021, 235, 118008. [Google Scholar] [CrossRef] [PubMed]
- Lebouc, M.; Richard, Q.; Garret, M.; Baufreton, J. Striatal circuit development and its alterations in Huntington’s disease. Neurobiol. Dis. 2020, 145, 105076. [Google Scholar] [CrossRef] [PubMed]
- Federspiel, J.D.; Greco, T.M.; Lum, K.K.; Cristea, I.M. Hdac4 interactions in Huntington’s disease viewed through the prism of multiomics. Mol. Cell Proteom. 2019, 18, S92–S113. [Google Scholar] [CrossRef] [PubMed]
- Cheng, J.; Liu, H.P.; Lin, W.Y.; Tsai, F.J. Identification of contributing genes of Huntington’s disease by machine learning. BMC Med. Genom. 2020, 13, 176. [Google Scholar] [CrossRef]
- Seefelder, M.; Kochanek, S. A meta-analysis of transcriptomic profiles of Huntington’s disease patients. PLoS ONE 2021, 16, e0253037. [Google Scholar]
- Imamura, T.; Fujita, K.; Tagawa, K.; Ikura, T.; Chen, X.; Homma, H.; Tamura, T.; Mao, Y.; Taniguchi, J.B.; Motoki, K.; et al. Identification of hepta-histidine as a candidate drug for Huntington’s disease by in silico-in vitro- in vivo-integrated screens of chemical libraries. Sci. Rep. 2016, 6, 33861. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Panwar, S.; Sharma, M.K.; Sharma, M.K. Genes to drug: An in-silico approach to design a drug for Huntington disease (HD) in Homo sapiens. Int. J. Comput. Biol. Drug Des. 2021, 14, 190–201. [Google Scholar] [CrossRef]
- Kohli, H.; Kumar, P.; Ambasta, R.K. In silico designing of putative peptides for targeting pathological protein Htt in Huntington’s disease. Heliyon 2021, 7, e06088. [Google Scholar] [CrossRef] [PubMed]
- Sundaram, J.R.; Wu, Y.; Lee, I.C.; George, S.E.; Hota, M.; Ghosh, S.; Kesavapany, S.; Ahmed, M.; Tan, E.-K.; Shenolikar, S. PromISR-6, a guanabenz analogue, improves cellular survival in an experimental model of Huntington’s disease. ACS Chem. Neurosci. 2019, 10, 3575–3589. [Google Scholar] [CrossRef] [PubMed]
- Deepa, S.; Rymbai, E.; Praveen, T.; Saravanan, J. Neuroprotective effects of farnesol on motor and cognitive impairment against 3-nitropropionic acid-induced Huntington’s disease. Thai J. Pharm. Sci. 2021, 45, 16–23. [Google Scholar]
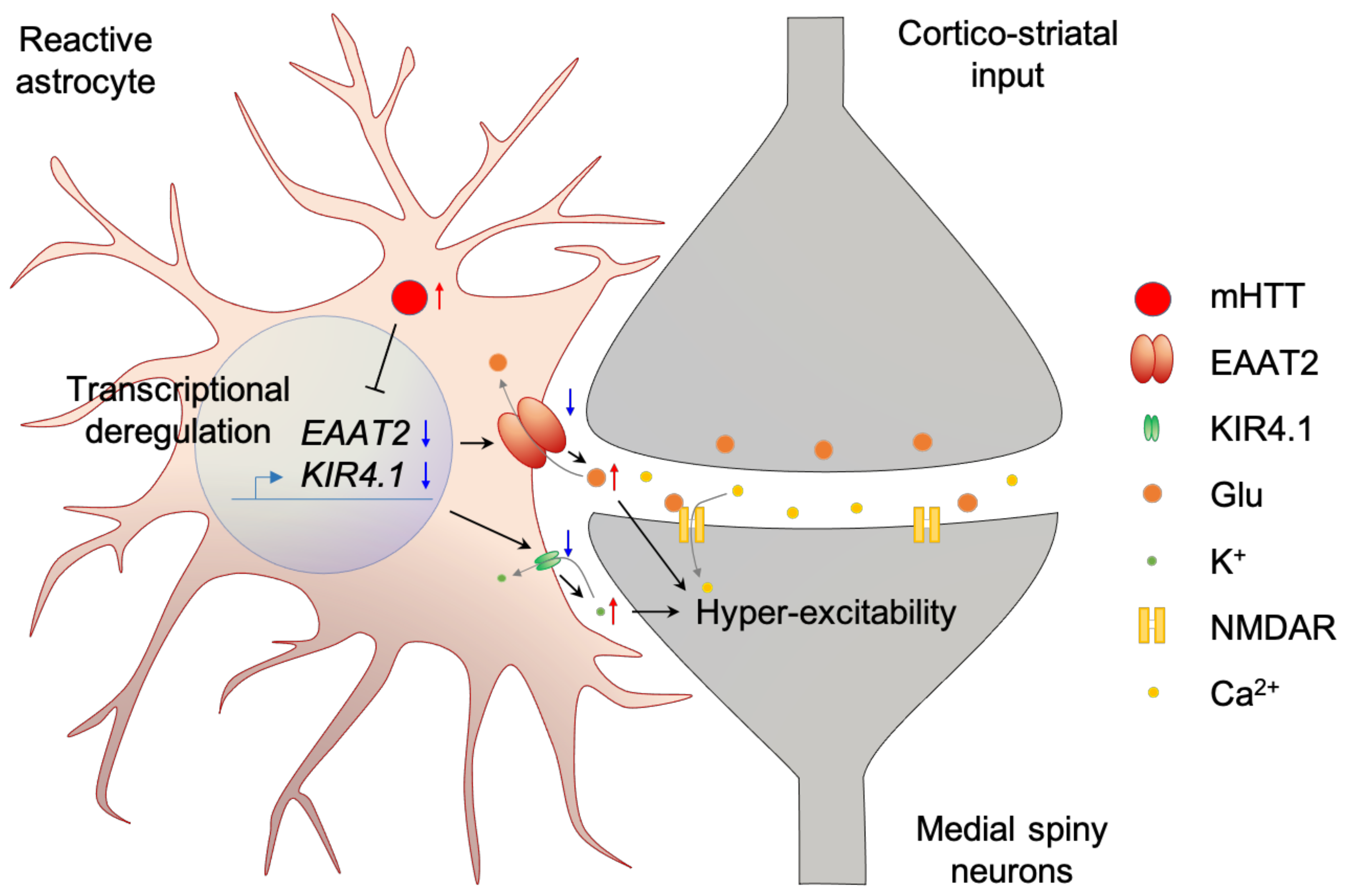
| HD Pathology | Specimen | Brain Region/Cell Type | Experimental Method | Reference |
|---|---|---|---|---|
| Less EAAT2 mRNA | Postmortem | Cingulate cortex | RNA sequencing | [78] |
| Neostriatum | In situ hybridization | [79] | ||
| Less EAAT2 protein | Postmortem | Striatum | Immunohistochemistry | [34] |
| Striatum, cortex | Western blot | [80] | ||
| Mouse; R6/2 | Striatum, cortex, hippocampus, midbrain | Quantitative proteomics | [81] | |
| Less glutamate uptake | Postmortem | Prefrontal cortex | Glutamate uptake assay | [84] |
| Cell; astrocyte | differentiated from Q77 monkey iPSC | Glutamate uptake assay | [82] | |
| Mouse; Q175 | Single corticostriatal synapse | Imaging assay with glutamate sensor | [83] | |
| Less Kir4.1 mRNA | Postmortem | cingulate cortex | RNA sequencing | [78] |
| Less Kir4.1 protein Higher extracellular K+More excitable | Mouse; R6/2 | Striatal MSN and astrocyte | qPCR, IHC, Western blot, Virus microinjection, Electrophysiology | [86] |
| Altered Ca2+ signal | Mouse; R6/2 | Striatal astrocyte | Virus microinjectionElectrophysiology | [87] |
| More excitotoxicity | Cell; neuron and astrocyte | Co-culture of HD neurons and astrocytes from human iPSC | Cell count after glutamate exposure | [89] |
| Co-culture of wild type neurons with mHTT infected glia from rat primary culture | [90] |
| Target | Strategy | Mode of Action | Disease Model | Clinical Trial & NCTno. | References |
|---|---|---|---|---|---|
| mHTT gene | CRISPR/Cas9 | Excise mHTT DNA selectively | Cell iPS | [174] | |
| Mouse BacHD | [175] | ||||
| Mouse HD140Q | [176] | ||||
| Mouse R6/2 | [177] | ||||
| KRAB-ZFPs | Inhibition of translation or transcriptdegradation | Mouse R6/1,2 | [178,179] | ||
| shRNA | Mouse R6/2 | [180] | |||
| Mouse N171-82Q | [181,182] | ||||
| siRNA | Mouse HTT injected | [183] | |||
| Mouse Hdh-150Q | [184] | ||||
| miRNA | Mouse HD140Q | Phase I/II [235], NCT04120493 | [185] | ||
| Antisense nucleotide | Bind to HTT mRNA | Mouse BACHD | Phase II, NCT02519036 | [236,237] | |
| Transcriptional dysregulation | Phenylbutyrate | Inactivate histone deacetylase | Mouse N171-82Q | Phase II, NCT00212316 | [202] |
| Sodium butyrate | Mouse R6/2 | [203] | |||
| HDACi 4b | Mouse N171–82Q | [204,205,206] | |||
| HDACi LBH589 | Transgenic Rodent HD Models | [207] | |||
| Tubastatin A | Cell primary neuron | [208] | |||
| CKD-504 | Phase I, NCT03713892 | ||||
| Mithramycin | Increase H3K9 | Mouse R6/2 | [209] | ||
| mHTT aggregation | Cystamine | Suppress mHTT crosslinking | Mouse R6/2 | [238,239] | |
| Congo red | Bind and inhibit polyglutamineoligomerization | Mouse R6/2 | [240] | ||
| ChrysamineG, Direct fast yellow | [210] | ||||
| Trehalose | [211] | ||||
| mHTT fragmentation | Minocycline | Inhibit caspase | Mouse R6/2 | Phase III, NCT00277355 | [241] |
| Z-VAD-FMK | Cell X57 | [216] | |||
| mHTT lowering | Blood transfusion | Remove circulating mHTT | Mouse zQ175 | [217] | |
| mHTT post-modification | Insulin, exendin-4 | Increase mHTT Phosphorylation | Cell SH-SY5Y | [242] | |
| GM1 | Mouse YAC128 | [243] | |||
| RCAN1-1L | Cell ST14A | [244] | |||
| SGK | Cell primary neuron | [245] | |||
| Transactivation | KD3010 | Increased PPARδ transactivation | Mouse pCAGGS-loxP-STOP-loxP | [246] | |
| Mitochondrial dysfunction | Creatine | Inactivate mitochondrial permeability transition | Mouse R6/2 | Phase II, NCT00026988 | [218] |
| Coenzyme Q10 | Enhance electron transport | Phase II, NCT00920699 | [219,220] | ||
| PGC-1α | Upregulate mitochondrial gene | [222] | |||
| Metabolism | rhIGF-1 | increase glucose uptake | Mouse YAC128 Mouse R6/2 | [223] | |
| Autophagy | Niclosamide | Inhibit mTOR | Cell HEK293, N2a | [225] | |
| MAP4343, 17βE2, Isoquercitrin | Regulate stress response | C.elegans HD mutants | [228] | ||
| Apoptosis | Z-VAD-FMK, Z-DEVD-FMK, Z-LEHD-FMK | Inhibit caspase | Cell primary neuron | [227] | |
| PG3d | Cell COS-7 | [229] | |||
| Lithium chloride | Rat QA injected | [230] | |||
| Excitotoxicity | Memantine | Inhibit NMDA receptor | Phase IV, NCT00652457; Phase II, NCT00652457 | ||
| Necrostatin-1 | Inhibit RIP1 kinase | Mouse R6/2 | [233,247] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kim, C.; Yousefian-Jazi, A.; Choi, S.-H.; Chang, I.; Lee, J.; Ryu, H. Non-Cell Autonomous and Epigenetic Mechanisms of Huntington’s Disease. Int. J. Mol. Sci. 2021, 22, 12499. https://doi.org/10.3390/ijms222212499
Kim C, Yousefian-Jazi A, Choi S-H, Chang I, Lee J, Ryu H. Non-Cell Autonomous and Epigenetic Mechanisms of Huntington’s Disease. International Journal of Molecular Sciences. 2021; 22(22):12499. https://doi.org/10.3390/ijms222212499
Chicago/Turabian StyleKim, Chaebin, Ali Yousefian-Jazi, Seung-Hye Choi, Inyoung Chang, Junghee Lee, and Hoon Ryu. 2021. "Non-Cell Autonomous and Epigenetic Mechanisms of Huntington’s Disease" International Journal of Molecular Sciences 22, no. 22: 12499. https://doi.org/10.3390/ijms222212499
APA StyleKim, C., Yousefian-Jazi, A., Choi, S.-H., Chang, I., Lee, J., & Ryu, H. (2021). Non-Cell Autonomous and Epigenetic Mechanisms of Huntington’s Disease. International Journal of Molecular Sciences, 22(22), 12499. https://doi.org/10.3390/ijms222212499






