Alleles of the GRF3-2A Gene in Wheat and Their Agronomic Value
Abstract
1. Introduction
2. Results
2.1. TaGRF3-2A Alleles
2.2. Protein Isoforms
2.3. Phylogenetics of Protein Sequences
2.4. Allele Designation
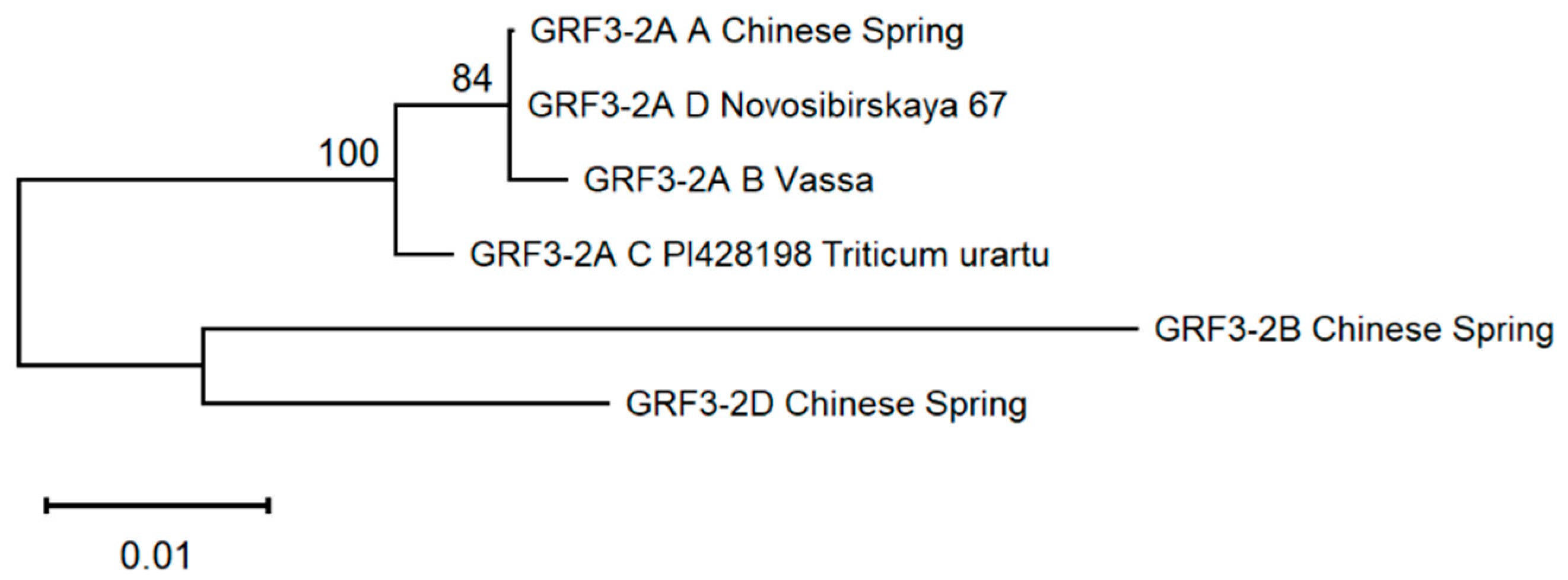
2.5. Allele Phylogenetic Analysis
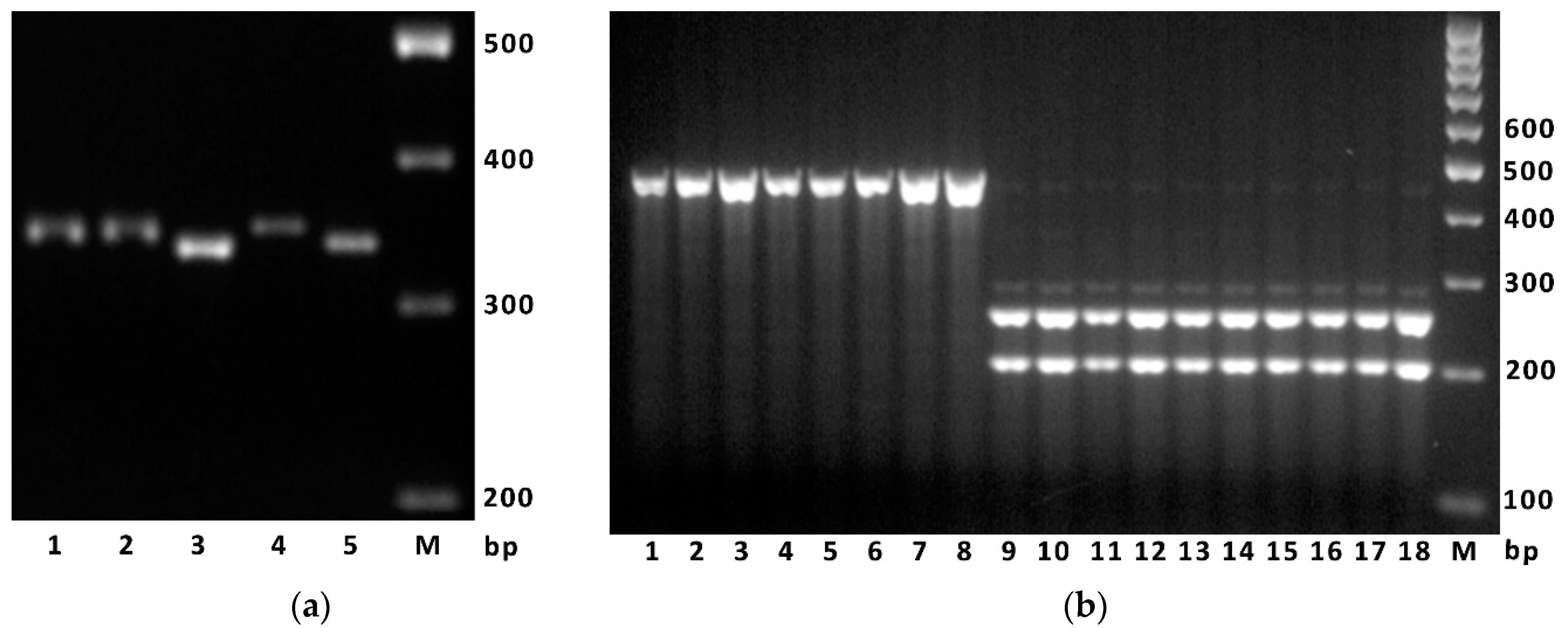
2.6. Molecular Markers and Phenotype
3. Discussion
4. Materials and Methods
4.1. Plant Material and Phenotyping
4.2. Weather Conditions
4.3. DNA Extraction, PCR and Sequencing
4.4. Sequences Obtained from Genome Assemblies
4.5. Sequence Analysis and Phylogenetics
4.6. Pedigrees
4.7. Molecular Markers
4.8. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Huang, W.; He, Y.; Yang, L.; Lu, C.; Zhu, Y.; Sun, C.; Ma, D.; Yin, J. Genome-Wide Analysis of Growth-Regulating Factors (GRFs) in Triticum Aestivum. PeerJ 2021, 9, e10701. [Google Scholar] [CrossRef]
- van der Knaap, E.; Kim, J.H.; Kende, H. A Novel Gibberellin-Induced Gene from Rice and Its Potential Regulatory Role in Stem Growth. Plant. Physiol. 2000, 122, 695–704. [Google Scholar] [CrossRef]
- Kim, J.H.; Choi, D.; Kende, H. The AtGRF Family of Putative Transcription Factors Is Involved in Leaf and Cotyledon Growth in Arabidopsis. Plant. J. 2003, 36, 94–104. [Google Scholar] [CrossRef]
- Liu, J.; Hua, W.; Yang, H.-L.; Zhan, G.-M.; Li, R.-J.; Deng, L.-B.; Wang, X.-F.; Liu, G.-H.; Wang, H.-Z. The BnGRF2 Gene (GRF2-like Gene from Brassica napus) Enhances Seed Oil Production through Regulating Cell Number and Plant Photosynthesis. J. Exp. Bot. 2012, 63, 3727–3740. [Google Scholar] [CrossRef] [PubMed]
- Zan, T.; Zhang, L.; Xie, T.; Li, L. Genome-Wide Identification and Analysis of the Growth-Regulating Factor (GRF) Gene Family and GRF-Interacting Factor Family in Triticum aestivum L. Biochem. Genet. 2020, 58, 705–724. [Google Scholar] [CrossRef]
- Kim, J.H.; Kende, H. A Transcriptional Coactivator, AtGIF1, Is Involved in Regulating Leaf Growth and Morphology in Arabidopsis. Proc. Natl. Acad. Sci. USA 2004, 101, 13374–13379. [Google Scholar] [CrossRef]
- Li, S.; Tian, Y.; Wu, K.; Ye, Y.; Yu, J.; Zhang, J.; Liu, Q.; Hu, M.; Li, H.; Tong, Y.; et al. Modulating Plant Growth–Metabolism Coordination for Sustainable Agriculture. Nature 2018, 560, 595–600. [Google Scholar] [CrossRef]
- Liebsch, D.; Palatnik, J.F. MicroRNA MiR396, GRF Transcription Factors and GIF Co-Regulators: A Conserved Plant Growth Regulatory Module with Potential for Breeding and Biotechnology. Curr. Opin. Plant Biol. 2020, 53, 31–42. [Google Scholar] [CrossRef] [PubMed]
- Beltramino, M.; Debernardi, J.M.; Ferela, A.; Palatnik, J.F. ARF2 Represses Expression of Plant GRF Transcription Factors in a Complementary Mechanism to MicroRNA MiR396. Plant Physiol. 2021, 185, 1798–1812. [Google Scholar] [CrossRef] [PubMed]
- Gao, F.; Wang, K.; Liu, Y.; Chen, Y.; Chen, P.; Shi, Z.; Luo, J.; Jiang, D.; Fan, F.; Zhu, Y.; et al. Blocking MiR396 Increases Rice Yield by Shaping Inflorescence Architecture. Nat. Plants 2016, 2, 15196. [Google Scholar] [CrossRef]
- Che, R.; Tong, H.; Shi, B.; Liu, Y.; Fang, S.; Liu, D.; Xiao, Y.; Hu, B.; Liu, L.; Wang, H.; et al. Control of Grain Size and Rice Yield by GL2-Mediated Brassinosteroid Responses. Nat. Plants 2016, 2, 15195. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.-S.; Mizoi, J.; Kidokoro, S.; Maruyama, K.; Nakajima, J.; Nakashima, K.; Mitsuda, N.; Takiguchi, Y.; Ohme-Takagi, M.; Kondou, Y.; et al. Arabidopsis GROWTH-REGULATING FACTOR7 Functions as a Transcriptional Repressor of Abscisic Acid– and Osmotic Stress–Responsive Genes, Including DREB2A. Plant. Cell 2012, 24, 3393–3405. [Google Scholar] [CrossRef]
- Duan, P.; Ni, S.; Wang, J.; Zhang, B.; Xu, R.; Wang, Y.; Chen, H.; Zhu, X.; Li, Y. Regulation of OsGRF4 by OsmiR396 Controls Grain Size and Yield in Rice. Nat. Plants 2015, 2, 1–5. [Google Scholar] [CrossRef]
- Avni, R.; Oren, L.; Shabtay, G.; Assili, S.; Pozniak, C.; Hale, I.; Ben-David, R.; Peleg, Z.; Distelfeld, A. Genome Based Meta-QTL Analysis of Grain Weight in Tetraploid Wheat Identifies Rare Alleles of GRF4 Associated with Larger Grains. Genes 2018, 9, 636. [Google Scholar] [CrossRef]
- Tsukaya, H. Yield Increase: GRFs Provide the Key. Nat. Plants 2015, 2, 1–2. [Google Scholar] [CrossRef] [PubMed]
- Walkowiak, S.; Gao, L.; Monat, C.; Haberer, G.; Kassa, M.T.; Brinton, J.; Ramirez-Gonzalez, R.H.; Kolodziej, M.C.; Delorean, E.; Thambugala, D.; et al. Multiple Wheat Genomes Reveal Global Variation in Modern Breeding. Nature 2020, 588, 277–283. [Google Scholar] [CrossRef]
- Alaux, M.; Rogers, J.; Letellier, T.; Flores, R.; Alfama, F.; Pommier, C.; Mohellibi, N.; Durand, S.; Kimmel, E.; Michotey, C.; et al. Linking the International Wheat Genome Sequencing Consortium Bread Wheat Reference Genome Sequence to Wheat Genetic and Phenomic Data. Genome Biol. 2018, 19, 111. [Google Scholar] [CrossRef] [PubMed]
- Avni, R.; Nave, M.; Barad, O.; Baruch, K.; Twardziok, S.O.; Gundlach, H.; Hale, I.; Mascher, M.; Spannagl, M.; Wiebe, K.; et al. Wild Emmer Genome Architecture and Diversity Elucidate Wheat Evolution and Domestication. Science 2017, 357, 93–97. [Google Scholar] [CrossRef]
- Ling, H.-Q.; Ma, B.; Shi, X.; Liu, H.; Dong, L.; Sun, H.; Cao, Y.; Gao, Q.; Zheng, S.; Li, Y.; et al. Genome Sequence of the Progenitor of Wheat A Subgenome Triticum Urartu. Nature 2018, 557, 424–428. [Google Scholar] [CrossRef]
- den Dunnen, J.T.; Dalgleish, R.; Maglott, D.R.; Hart, R.K.; Greenblatt, M.S.; McGowan-Jordan, J.; Roux, A.-F.; Smith, T.; Antonarakis, S.E.; Taschner, P.E.M. HGVS Recommendations for the Description of Sequence Variants: 2016 Update. Hum. Mutat. 2016, 37, 564–569. [Google Scholar] [CrossRef]
- Chow, C.-N.; Lee, T.-Y.; Hung, Y.-C.; Li, G.-Z.; Tseng, K.-C.; Liu, Y.-H.; Kuo, P.-L.; Zheng, H.-Q.; Chang, W.-C. PlantPAN3.0: A New and Updated Resource for Reconstructing Transcriptional Regulatory Networks from ChIP-Seq Experiments in Plants. Nucleic Acids Res. 2019, 47, D1155–D1163. [Google Scholar] [CrossRef]
- Hong, L.; Ye, C.; Lin, J.; Fu, H.; Wu, X.; Li, Q.Q. Alternative Polyadenylation Is Involved in Auxin-Based Plant Growth and Development. Plant J. 2018, 93, 246–258. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Bhatia, S. A Polymorphic (GA/CT)n- SSR Influences Promoter Activity of Tryptophan Decarboxylase Gene in Catharanthus roseus L. Don. Sci. Rep. 2016, 6, 33280. [Google Scholar] [CrossRef]
- Artemova, G.V.; Likhenko, I.E. Historical aspects and main results of scientific research in SIBNIIRS—Branch of the Institute of Cytology and Genetics SO RAN. Lett. Vavilov J. Genet. Breed. 2016, 2, 3. Available online: http://pismavavilov.ru/2016-2-1/ (accessed on 16 November 2021).
- He, F.; Pasam, R.; Shi, F.; Kant, S.; Keeble-Gagnere, G.; Kay, P.; Forrest, K.; Fritz, A.; Hucl, P.; Wiebe, K.; et al. Exome Sequencing Highlights the Role of Wild-Relative Introgression in Shaping the Adaptive Landscape of the Wheat Genome. Nat. Genet. 2019, 51, 896–904. [Google Scholar] [CrossRef]
- Guo, W.; Xin, M.; Wang, Z.; Yao, Y.; Hu, Z.; Song, W.; Yu, K.; Chen, Y.; Wang, X.; Guan, P.; et al. Origin and Adaptation to High Altitude of Tibetan Semi-Wild Wheat. Nat. Commun. 2020, 11, 5085. [Google Scholar] [CrossRef] [PubMed]
- Alvarez, M.A.; Tranquilli, G.; Lewis, S.; Kippes, N.; Dubcovsky, J. Genetic and Physical Mapping of the Earliness per Se Locus Eps-Am1 in Triticum monococcum Identifies EARLY FLOWERING 3 (ELF3) as a Candidate Gene. Funct. Integr. Genom. 2016, 16, 365–382. [Google Scholar] [CrossRef] [PubMed]
- Santra, D.K.; Santra, M.; Allan, R.E.; Campbell, K.G.; Kidwell, K.K. Genetic and Molecular Characterization of Vernalization Genes Vrn-A1, Vrn-B1, and Vrn-D1 in Spring Wheat Germplasm from the Pacific Northwest Region of the U.S.A. Plant. Breed. 2009, 128, 576–584. [Google Scholar] [CrossRef]
- Zanke, C.; Ling, J.; Plieske, J.; Kollers, S.; Ebmeyer, E.; Korzun, V.; Argillier, O.; Stiewe, G.; Hinze, M.; Beier, S.; et al. Genetic Architecture of Main Effect QTL for Heading Date in European Winter Wheat. Front. Plant. Sci. 2014, 5, 217. [Google Scholar] [CrossRef] [PubMed]
- Gao, L.; Zhao, G.; Huang, D.; Jia, J. Candidate Loci Involved in Domestication and Improvement Detected by a Published 90K Wheat SNP Array. Sci. Rep. 2017, 7, 44530. [Google Scholar] [CrossRef]
- KRYMKA. Available online: http://wheatpedigree.net/sort/show/32936 (accessed on 29 July 2021).
- CRIMEAN. Available online: http://wheatpedigree.net/sort/show/14818 (accessed on 29 July 2021).
- Moon, D. Introduction of Russian wheat in the Great Plains of the United States of America. In American Yearbook 2011; Ves′ Mir: Moscow, Russia, 2011; pp. 161–176. [Google Scholar]
- Competitive Variety Testing Methodologies. Available online: https://gossortrf.ru/en/metodiki_ksi/ (accessed on 6 September 2021).
- Doyle, P.J. DNA Protocols for Plants. In Molecular Techniques in Taxonomy; Hewitt, G.M., Johnston, A.W.B., Young, J.P.W., Eds.; NATO ASI Series; Springer: Berlin/Heidelberg, Germany, 1991; ISBN 978-3-642-83964-1. [Google Scholar]
- Ye, J.; Coulouris, G.; Zaretskaya, I.; Cutcutache, I.; Rozen, S.; Madden, T.L. Primer-BLAST: A Tool to Design Target-Specific Primers for Polymerase Chain Reaction. BMC Bioinform. 2012, 13, 134. [Google Scholar] [CrossRef] [PubMed]
- Bazhenov, M.S.; Chernook, A.G.; Goncharov, N.P.; Chikida, N.N.; Belousova, M.K.; Karlov, G.I.; Divashuk, M.G. The Allelic Diversity of the Gibberellin Signaling Pathway Genes in Aegilops tauschii Coss. Plants 2020, 9, 1696. [Google Scholar] [CrossRef] [PubMed]
- MikhailBazhenov Consensus: A Program for Making Consensus Sequence from CAP3 Output. Available online: https://github.com/MikhailBazhenov/Consensus (accessed on 27 September 2021).
- Fruzangohar, M.; Kalashyan, E.; Kalambettu, P.; Ens, J.; Wiebe, K.; Pozniak, C.J.; Tricker, P.J.; Baumann, U. Novel Informatic Tools to Support Functional Annotation of the Durum Wheat Genome. Front. Plant. Sci. 2019, 10, 1244. [Google Scholar] [CrossRef] [PubMed]
- Camacho, C.; Coulouris, G.; Avagyan, V.; Ma, N.; Papadopoulos, J.; Bealer, K.; Madden, T.L. BLAST+: Architecture and Applications. BMC Bioinform. 2009, 10, 421. [Google Scholar] [CrossRef] [PubMed]
- MikhailBazhenov Fastfinder: A Program for Extracting Some Part of a Sequence from a Genome-Large FASTA File. Available online: https://github.com/MikhailBazhenov/fastfinder (accessed on 27 September 2021).
- Edgar, R.C. MUSCLE: Multiple Sequence Alignment with High Accuracy and High Throughput. Nucleic Acids Res. 2004, 32, 1792–1797. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X: Molecular Evolutionary Genetics Analysis across Computing Platforms. Mol. Biol. Evol. 2018, 35, 1547–1549. [Google Scholar] [CrossRef]
- NICHOLAS, K.B. GeneDoc: Analysis and Visualization of Genetic Variation. EMBNEW News 1997, 4, 14. [Google Scholar]
- Marchler-Bauer, A.; Derbyshire, M.K.; Gonzales, N.R.; Lu, S.; Chitsaz, F.; Geer, L.Y.; Geer, R.C.; He, J.; Gwadz, M.; Hurwitz, D.I.; et al. CDD: NCBI′s Conserved Domain Database. Nucleic Acids Res. 2015, 43, D222226. [Google Scholar] [CrossRef]
- Choi, Y.; Chan, A.P. PROVEAN Web Server: A Tool to Predict the Functional Effect of Amino Acid Substitutions and Indels. Bioinformatics 2015, 31, 2745–2747. [Google Scholar] [CrossRef]
- Hasegawa, M.; Kishino, H.; Yano, T. Dating of the Human-Ape Splitting by a Molecular Clock of Mitochondrial DNA. J. Mol. Evol. 1985, 22, 160–174. [Google Scholar] [CrossRef]
- Jones, D.T.; Taylor, W.R.; Thornton, J.M. The Rapid Generation of Mutation Data Matrices from Protein Sequences. Comput. Appl. Biosci. Cabios 1992, 8, 275–282. [Google Scholar] [CrossRef] [PubMed]
- Genetic Resources Information System for Wheat and Triticale. Available online: http://wheatpedigree.net/ (accessed on 16 August 2021).
- State Commission for Selection Achievements Test and Protection. Available online: https://gossortrf.ru/en/ (accessed on 16 August 2021).
- Kroupin, P.Y.; Chernook, A.G.; Bazhenov, M.S.; Karlov, G.I.; Goncharov, N.P.; Chikida, N.N.; Divashuk, M.G. Allele Mining of TaGRF-2D Gene 5′-UTR in Triticum aestivum and Aegilops tauschii Genotypes. PLoS ONE 2020, 15, e0231704. [Google Scholar] [CrossRef] [PubMed]
- R Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2020. [Google Scholar]



| Isoform. | Differences in Protein Sequence | PROVEAN Score | Representative Accessions * |
|---|---|---|---|
| A | - | - | Chinese Spring |
| B | Gln165His | −2.601 ** | Stan, Vassa, Vid |
| C | Gln176_Ala177delinsHisGly | −1.224 | PI428198 (T. urartu) |
| Asp243Glu | 0.524 | ||
| D | Gln42_Gln44del | 0.821 | Novosibirskaya 67 |
| Primer Sequence, 5′→3′ | Tm, °C | Expected Product Size, bp |
|---|---|---|
| GRF-2A-1F: AAATTGAAGGCTAGACAATCGGC GRF-2A-1R: CCTTTTACTCCTACTTGCCTGGT | 60 | 1179 |
| GRF-2A-2F: CAAACGAACTTGACGGTACAGAT GRF-2A-2R: CACATGAGGATGAGGCTTCTTGA | 60 | 1188 |
| GRF-2A-3F: AGATTTCAGGTGTACTCGACCTC GRF-2A-3R: AGCATGCAGAAGATAAAAACGGC | 60 | 1185 |
| GRF-2A-4F: GCTCAGCTGCACATGGATAATG GRF-2A-4R: CGAGTCAGATTTGCAGCATAGTG | 60 | 1119 |
| GRF-2A-5F: TGCAGCAACAATTGCTCGTATAG GRF-2A-5R: CACCCCCACCCCTAAGATAGATA | 60 | 1250 |
Publisher′s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bazhenov, M.S.; Chernook, A.G.; Bespalova, L.A.; Gritsay, T.I.; Polevikova, N.A.; Karlov, G.I.; Nazarova, L.A.; Divashuk, M.G. Alleles of the GRF3-2A Gene in Wheat and Their Agronomic Value. Int. J. Mol. Sci. 2021, 22, 12376. https://doi.org/10.3390/ijms222212376
Bazhenov MS, Chernook AG, Bespalova LA, Gritsay TI, Polevikova NA, Karlov GI, Nazarova LA, Divashuk MG. Alleles of the GRF3-2A Gene in Wheat and Their Agronomic Value. International Journal of Molecular Sciences. 2021; 22(22):12376. https://doi.org/10.3390/ijms222212376
Chicago/Turabian StyleBazhenov, Mikhail S., Anastasiya G. Chernook, Ludmila A. Bespalova, Tatiana I. Gritsay, Nadezhda A. Polevikova, Gennady I. Karlov, Lubov A. Nazarova, and Mikhail G. Divashuk. 2021. "Alleles of the GRF3-2A Gene in Wheat and Their Agronomic Value" International Journal of Molecular Sciences 22, no. 22: 12376. https://doi.org/10.3390/ijms222212376
APA StyleBazhenov, M. S., Chernook, A. G., Bespalova, L. A., Gritsay, T. I., Polevikova, N. A., Karlov, G. I., Nazarova, L. A., & Divashuk, M. G. (2021). Alleles of the GRF3-2A Gene in Wheat and Their Agronomic Value. International Journal of Molecular Sciences, 22(22), 12376. https://doi.org/10.3390/ijms222212376






