Genome-Wide Comprehensive Analysis of the GASA Gene Family in Populus
Abstract
:1. Introduction
2. Results
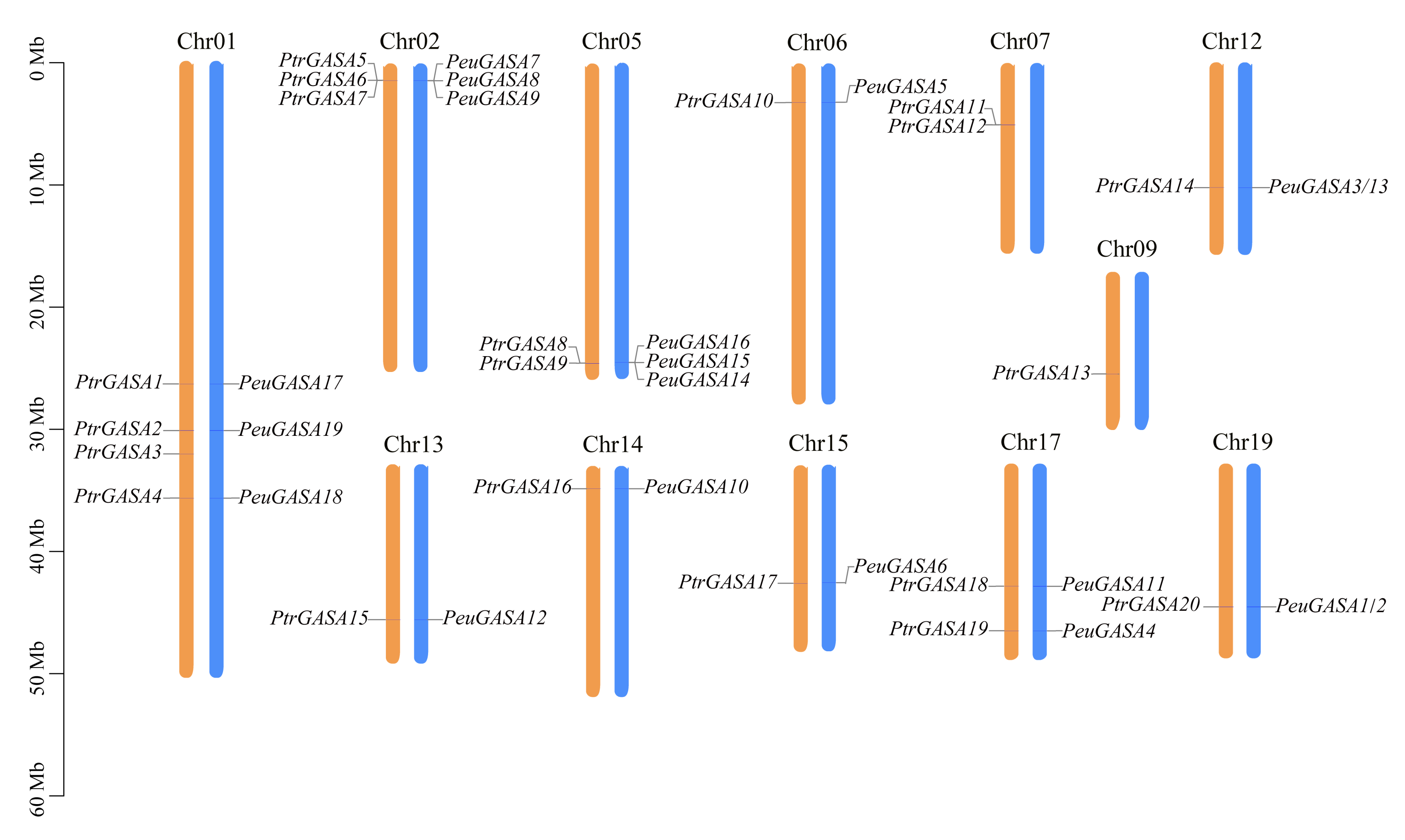
2.1. Genome-Wide Identification of GASA Genes in Populus
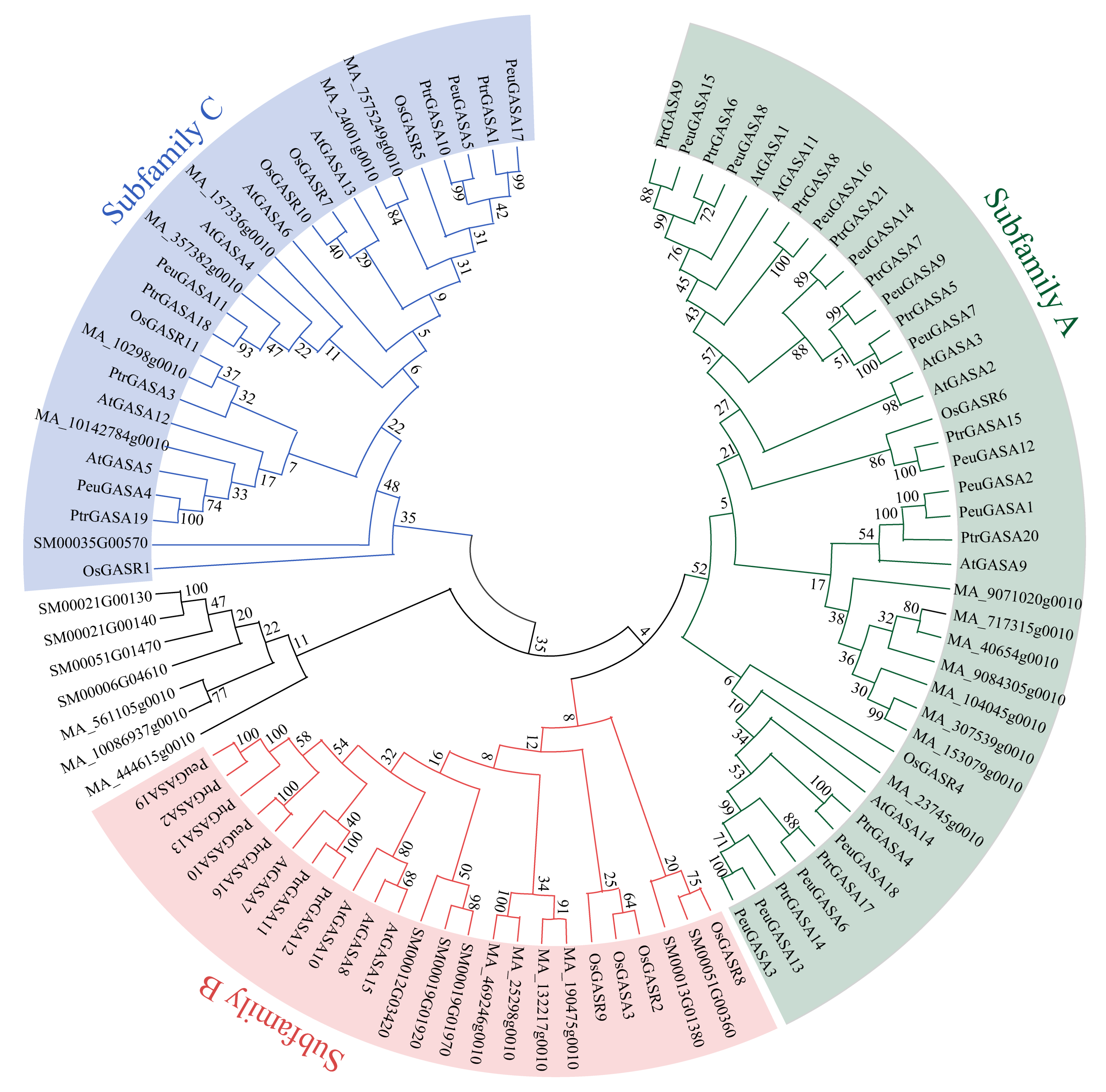
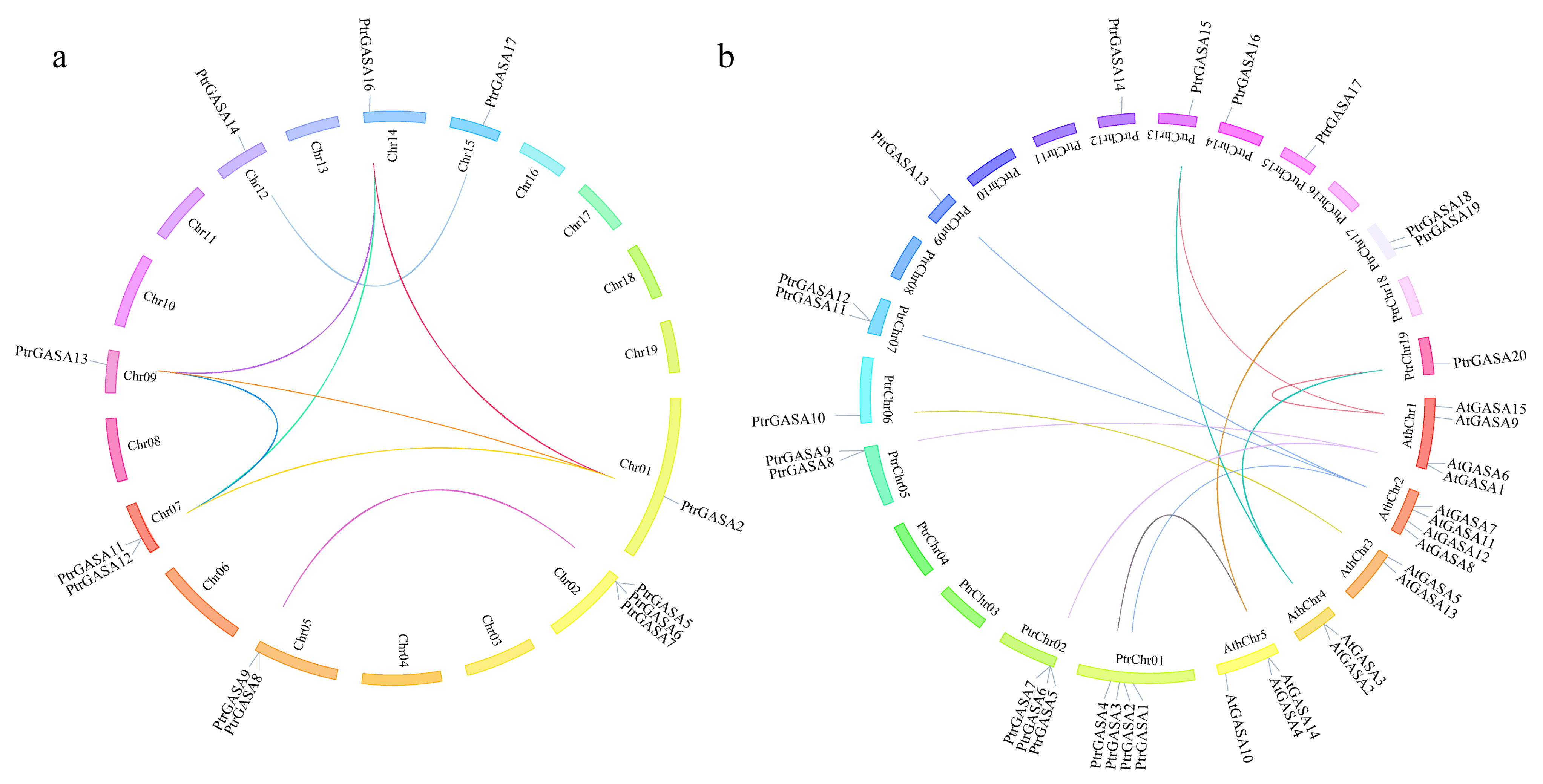
2.2. Evolutionary Relationships Analysis among GASA Domains and Collinearity Analysis among GASA Genes
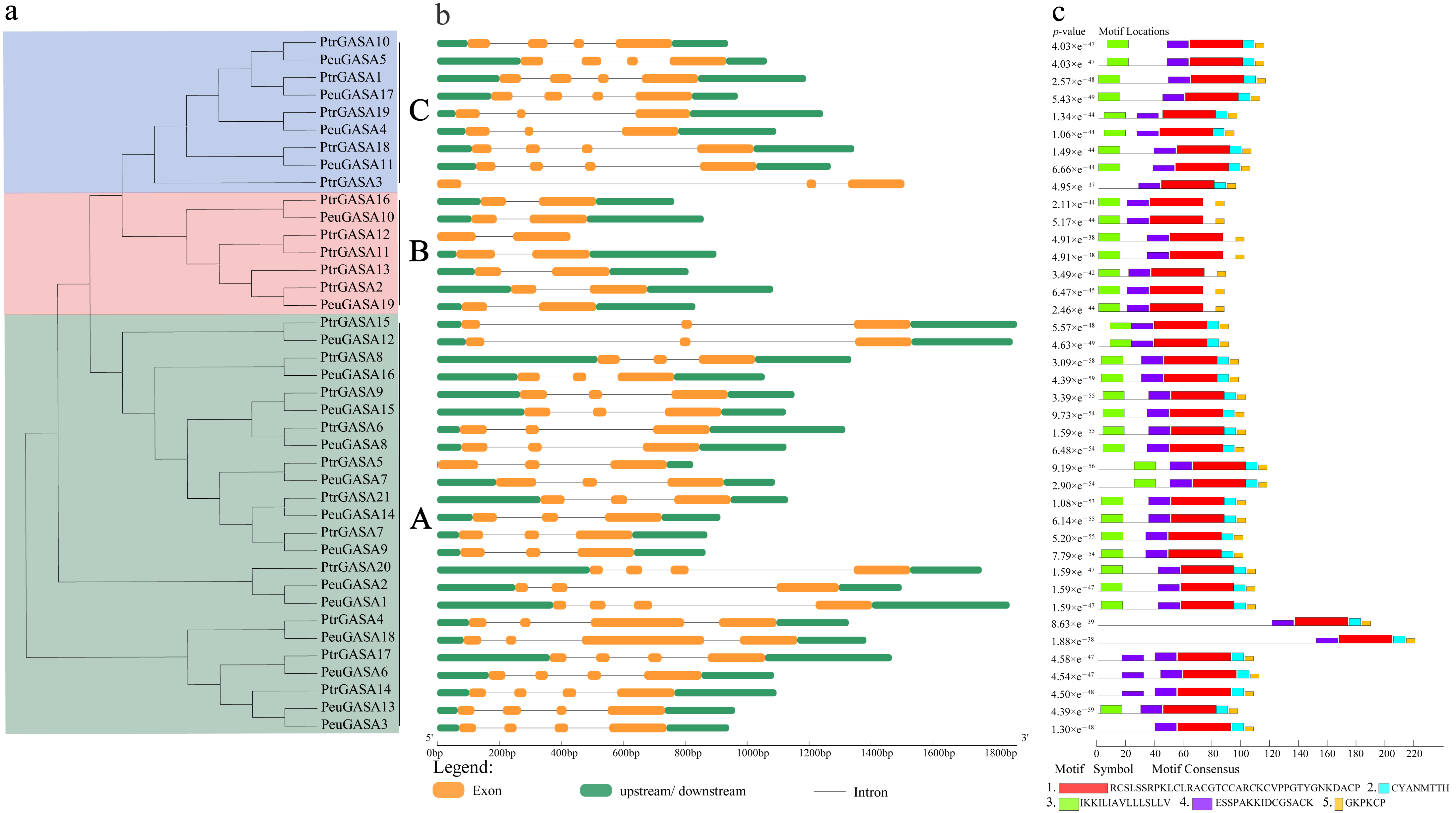
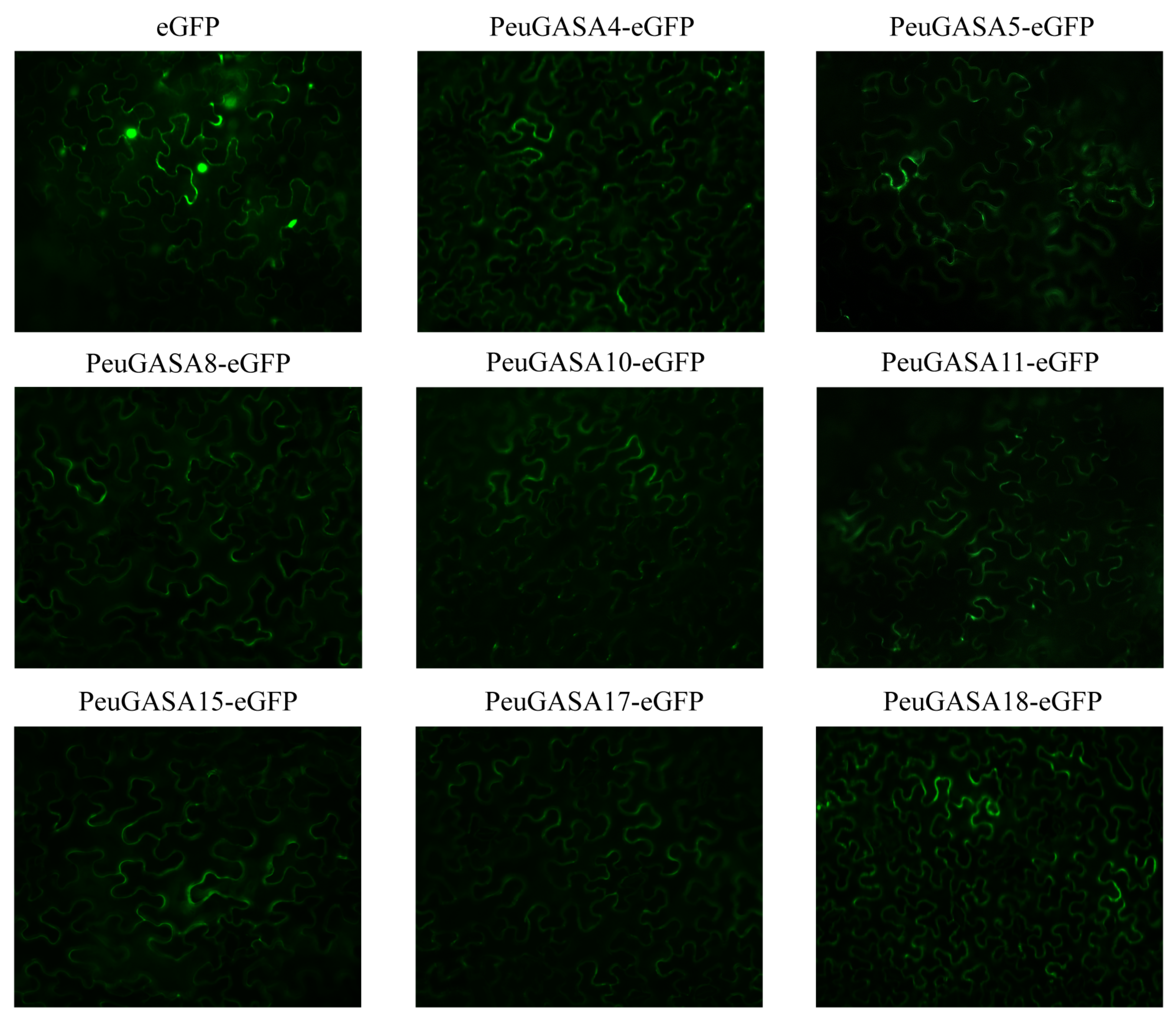
2.3. Gene Structure and Motif Identification, Physicochemical Properties, and Subcellular Localization of GASA Gene Family
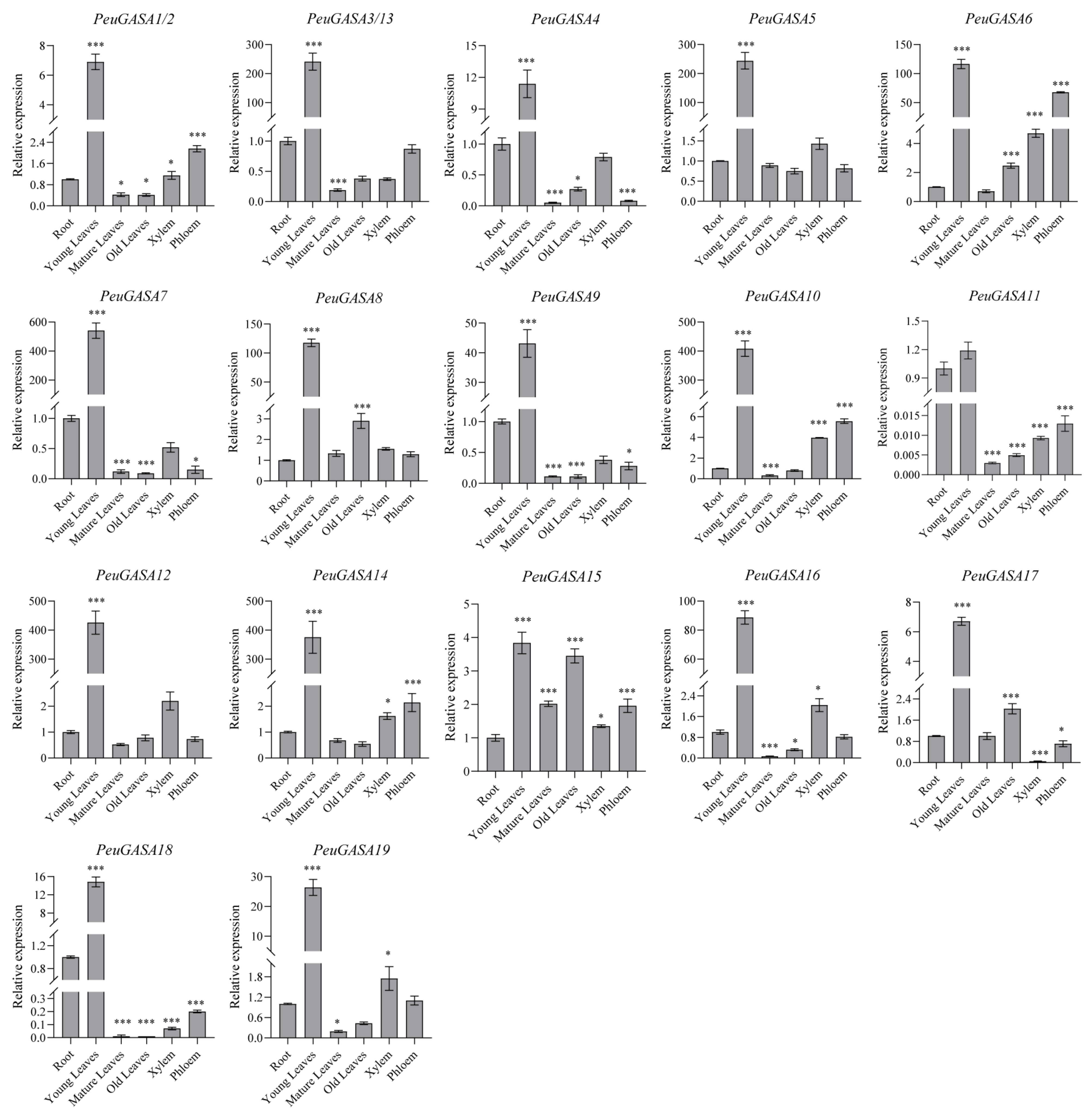
2.4. Expression Patterns of GASA Genes in Different Tissues of Populus
2.5. Responses of GASAs to Exogenous Hormones in P. euphratica Leaves
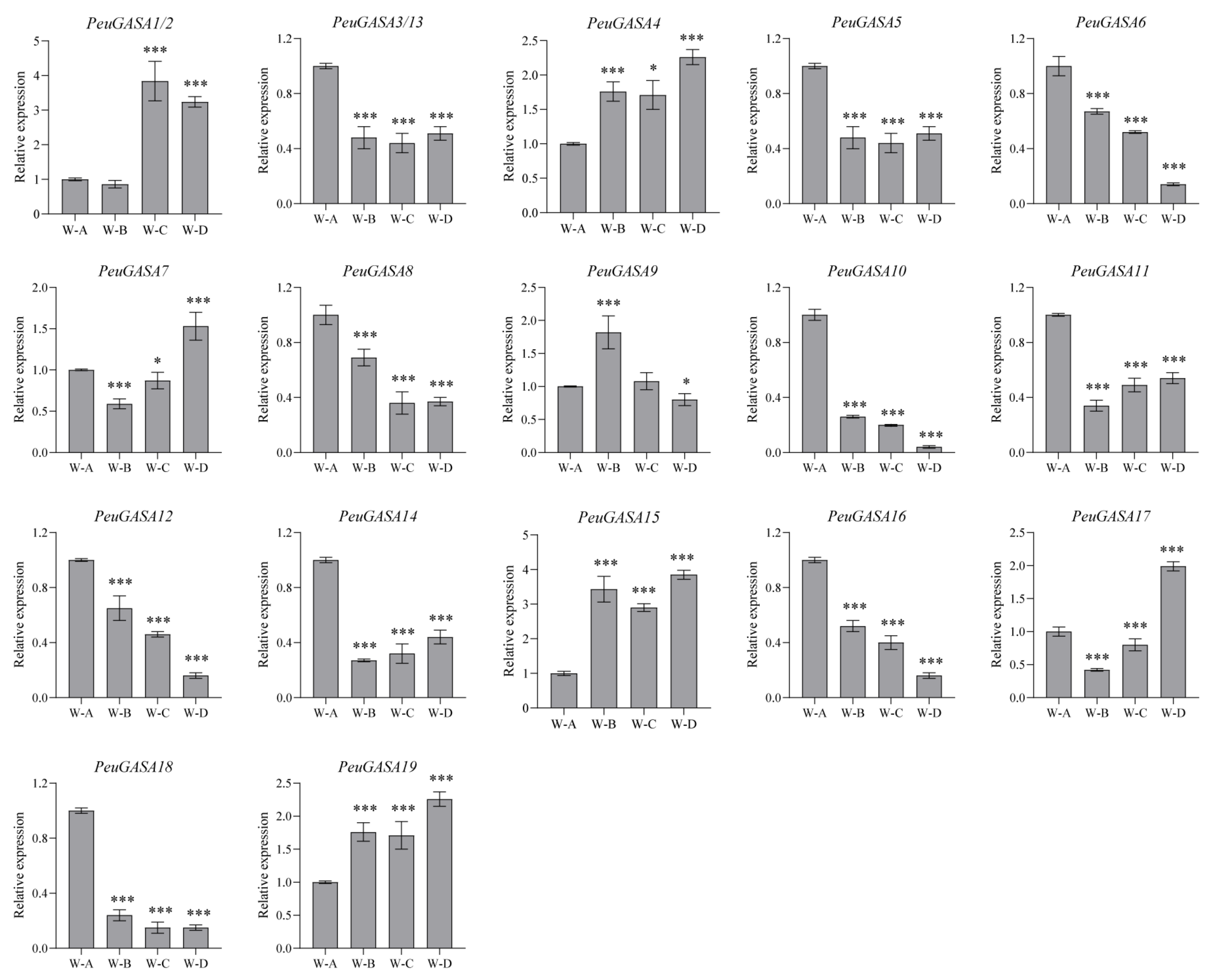
2.6. Response of GASAs to Stresses in Populus Leaves
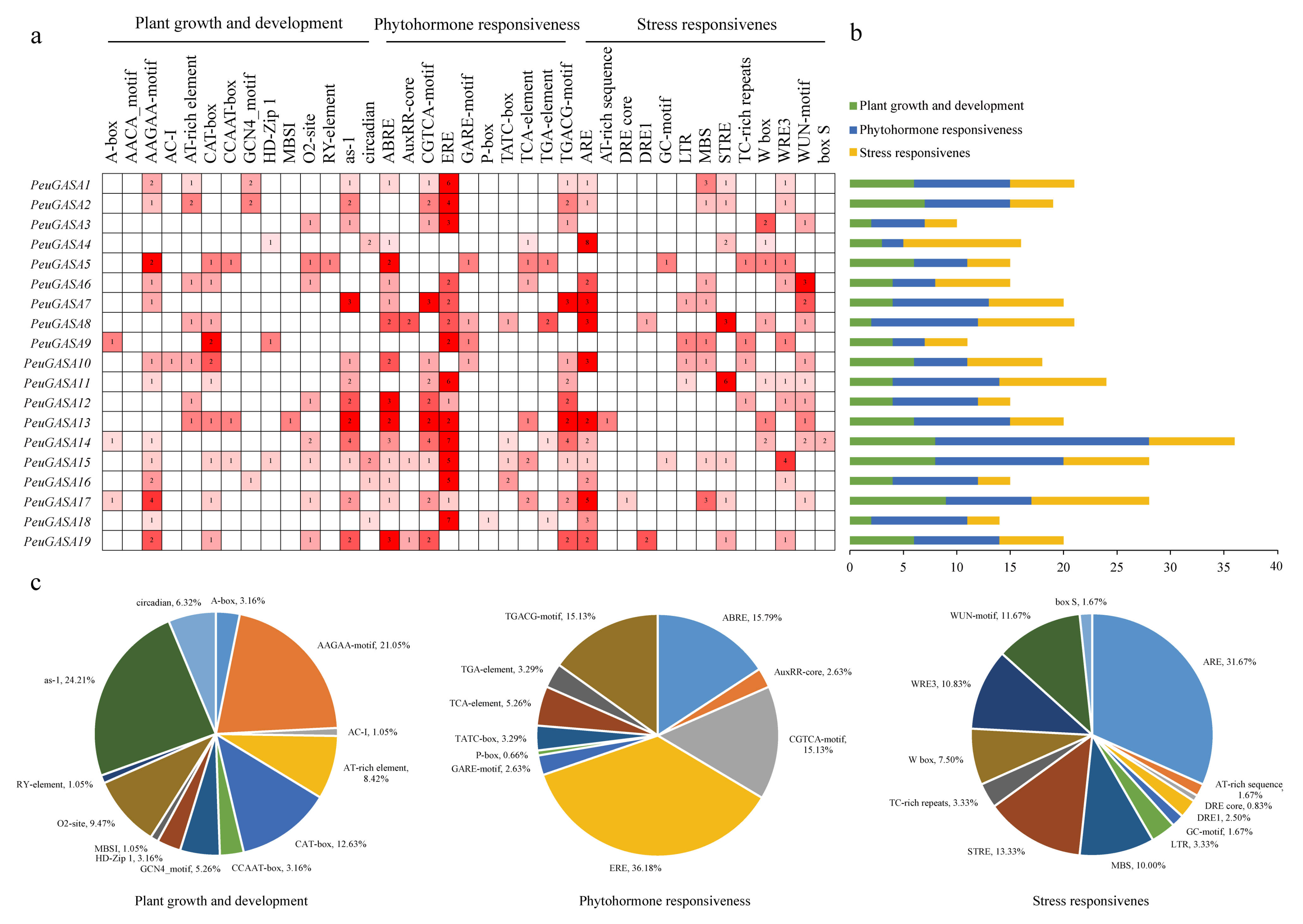
2.7. Identification of cis-Acting Elements in the Promoter Region of GASA Genes
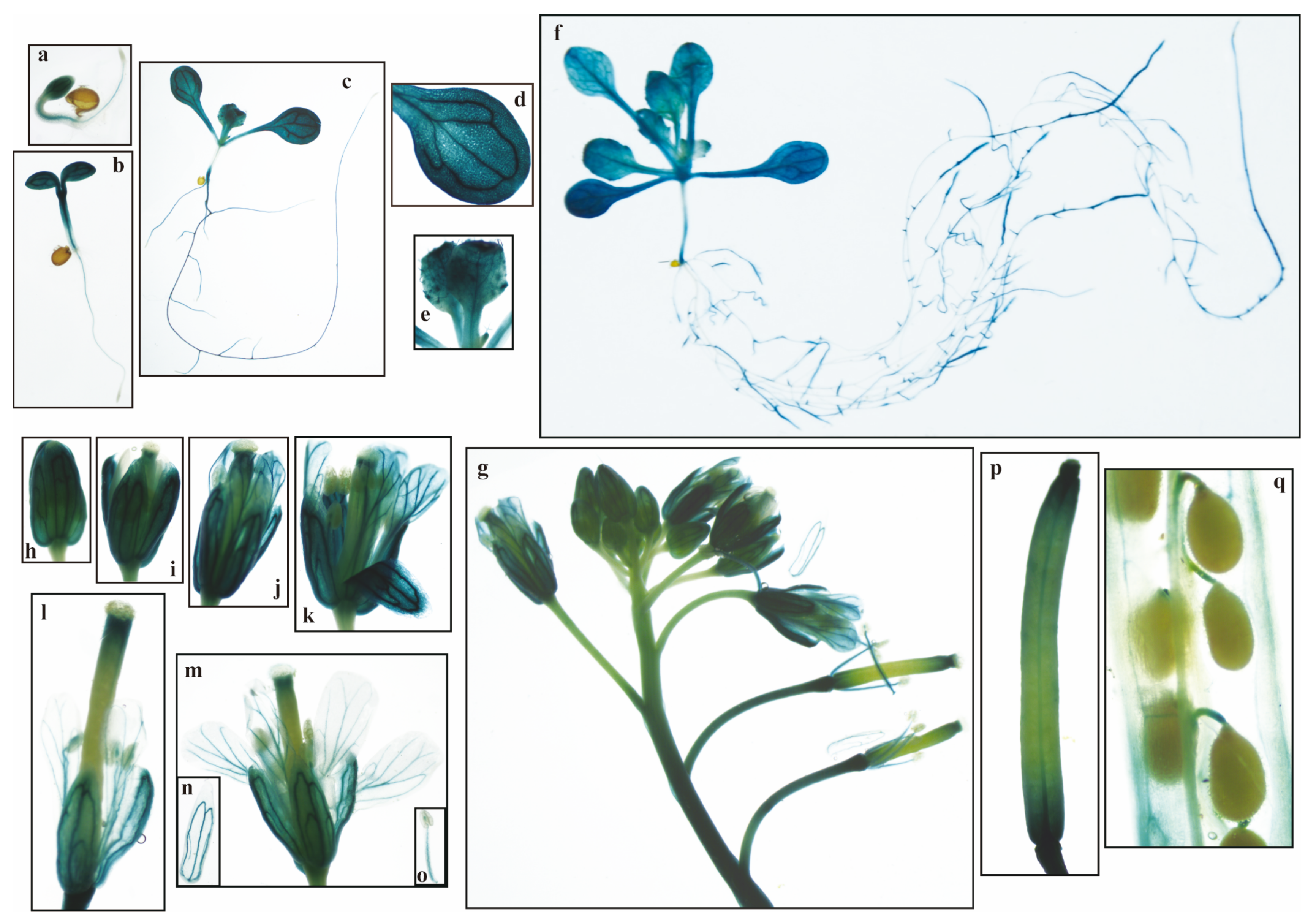
2.8. Analysis of PeuGASA15 Promoter Activity
3. Discussion
4. Materials and Methods
4.1. Identification of GASA Family Genes in P. trichocarpa and P. euphratica
4.2. Physical and Chemical Properties, Subcellular Localization, Multiple Alignments and Collinearity Analysis
4.3. Plant Materials and Sample Collections
4.4. RNA Extraction, cDNA Synthesis, and qRT-PCR Analysis
4.5. Subcellular Localization of PeuGASA Proteins
4.6. Arabidopsis Culture, Construction and Transformation of Vector, and GUS Activity Assay
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Marshall, E.; Costa, L.M.; Gutierrez-Marcos, J. Cysteine-rich peptides (CRPs) mediate diverse aspects of cell-cell communication in plant reproduction and development. J. Exp. Bot. 2011, 62, 1677–1686. [Google Scholar] [CrossRef] [Green Version]
- Matsubayashi, Y. Posttranslationally Modified Small-Peptide Signals in Plants. Annu. Rev. Plant Biol. 2014, 65, 385–413. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Zhang, H.; Jiao, H.; Li, L.; Qiao, X.; Fabrice, M.R.; Wu, J.; Zhang, S. Expansion and evolutionary patterns of cysteine-rich peptides in plants. BMC Genom. 2017, 18, 1–14. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Olsson, V.; Joos, L.; Zhu, S.; Gevaert, K.; Butenko, M.A.; De Smet, I. Look Closely, the Beautiful May Be Small: Precursor-Derived Peptides in Plants. Annu. Rev. Plant Biol. 2019, 70, 153–186. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kereszt, A.; Mergaert, P.; Montiel, J.; Endre, G.; Kondorosi, E. Impact of Plant Peptides on Symbiotic Nodule Development and Functioning. Front. Plant Sci. 2018, 9, 1026. [Google Scholar] [CrossRef]
- Uchi, N.; Fukudome, M.; Nozaki, N.; Suzuki, M.; Osuki, K.-I.; Shigenobu, S.; Uchiumi, T. Antimicrobial Activities of Cysteine-rich Peptides Specific to Bacteriocytes of the Pea Aphid Acyrthosiphon pisum. Microbes Environ. 2019, 34, 155–160. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhong, S.; Liu, M.; Wang, Z.; Huang, Q.; Hou, S.; Xu, Y.-C.; Ge, Z.; Song, Z.; Huang, J.; Qiu, X.; et al. Cysteine-rich peptides promote interspecific genetic isolation in Arabidopsis. Science 2019, 364, eaau9564. [Google Scholar] [CrossRef]
- Wang, L.; Wang, Z.; Xu, Y.; Joo, S.-H.; Kim, S.-K.; Xue, Z.; Xu, Z.; Wang, Z.; Chong, K. OsGSR1is involved in crosstalk between gibberellins and brassinosteroids in rice. Plant J. 2009, 57, 498–510. [Google Scholar] [CrossRef]
- Kumar, A.; Singh, A.; Kumar, P.; Sarkar, A.K. Giberellic Acid-Stimulated Transcript Proteins Evolved through Successive Conjugation of Novel Motifs and Their Subfunctionalization. Plant Physiol. 2019, 180, 998–1012. [Google Scholar] [CrossRef]
- Shi, L.; Gast, R.T.; Gopalraj, M.; Olszewski, N.E. Characterization of a shoot-specific, GA3- and ABA-Regulated gene from tomato. Plant J. 1992, 2, 153–159. [Google Scholar] [PubMed]
- Almasia, N.I.; Bazzini, A.A.; Hopp, H.E.; Vazquez-Rovere, C. Overexpression of snakin-1 gene enhances resistance to Rhizoctonia solani and Erwinia carotovora in transgenic potato plants. Mol. Plant Pathol. 2008, 9, 329–338. [Google Scholar] [CrossRef]
- Balaji, V.; Smart, C.D. Over-expression of snakin-2 and extensin-like protein genes restricts pathogen invasiveness and enhances tolerance to Clavibacter michiganensis subsp. michiganensis in transgenic tomato (Solanum lycopersicum). Transgenic Res. 2012, 21, 23–37. [Google Scholar] [CrossRef] [Green Version]
- Darqui, F.S.; Radonic, L.M.; Trotz, P.M.; Lopez, N.; Vazquez Rovere, C.; Hopp, H.E.; Lopez Bilbao, M. Potato snakin-1 gene enhances tolerance to Rhizoctonia solani and Sclerotinia sclerotiorum in transgenic lettuce plants. J. Biotechnol. 2018, 283, 62–69. [Google Scholar] [CrossRef] [PubMed]
- Rodriguez-Decuadro, S.; Barraco-Vega, M.; Dans, P.D.; Pandolfi, V.; Benko-Iseppon, A.M.; Cecchetto, G. Antimicrobial and structural insights of a new snakin-like peptide isolated from Peltophorum dubium (Fabaceae). Amino Acids 2018, 50, 1245–1259. [Google Scholar] [CrossRef]
- Fan, S.; Zhang, D.; Zhang, L.; Gao, C.; Xin, M.; Tahir, M.M.; Li, Y.; Ma, J.; Han, M. Comprehensive analysis of GASA family members in the Malus domestica genome: Identification, characterization, and their expressions in response to apple flower induction. BMC Genom. 2017, 18, 827. [Google Scholar] [CrossRef] [PubMed]
- Moyano-Cañete, E.; Bellido, M.L.; García-Caparrós, N.; Medina-Puche, L.; Amil-Ruiz, F.; González-Reyes, J.A.; Caballero, J.L.; Muñoz-Blanco, J.; Blanco-Portales, R. FaGAST2, a Strawberry Ripening-Related Gene, Acts Together with FaGAST1 to Determine Cell Size of the Fruit Receptacle. Plant Cell Physiol. 2013, 54, 218–236. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, S.; Yang, C.; Peng, J.; Sun, S.; Wang, X. GASA5, a regulator of flowering time and stem growth in Arabidopsis thaliana. Plant Mol. Biol. 2009, 69, 745–759. [Google Scholar] [CrossRef] [PubMed]
- Zhong, C.; Xu, H.; Ye, S.; Wang, S.; Li, L.; Zhang, S.; Wang, X. AtGASA6 Serves as an Integrator of Gibberellin-, Abscisic Acid-and Glucose-Signaling during Seed Germination in Arabidopsis. Plant Physiol. 2015, 169, 2288–2303. [Google Scholar] [CrossRef] [Green Version]
- Zimmermann, R.; Sakai, H.; Hochholdinger, F. The Gibberellic Acid Stimulated-Like gene family in maize and its role in lateral root development. Plant Physiol. 2010, 152, 356–365. [Google Scholar] [CrossRef] [Green Version]
- Nahirnak, V.; Almasia, N.I.; Fernandez, P.V.; Hopp, H.E.; Estevez, J.M.; Carrari, F.; Vazquez-Rovere, C. Potato snakin-1 gene silencing affects cell division, primary metabolism, and cell wall composition. Plant Physiol. 2012, 158, 252–263. [Google Scholar] [CrossRef] [Green Version]
- Han, M.; Jin, X.; Yao, W.; Kong, L.; Huang, G.; Tao, Y.; Li, L.; Wang, X.; Wang, Y. A Mini Zinc-Finger Protein (MIF) from Gerbera hybrida Activates the GASA Protein Family Gene, GEG, to Inhibit Ray Petal Elongation. Front. Plant Sci. 2017, 8, 1649. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Huang, G.; Han, M.; Jian, L.; Chen, Y.; Sun, S.; Wang, X.; Wang, Y. An ETHYLENE INSENSITIVE3-LIKE1 Protein Directly Targets the GEG Promoter and Mediates Ethylene-Induced Ray Petal Elongation in Gerbera hybrida. Front. Plant Sci. 2020, 10, 1737. [Google Scholar] [CrossRef] [PubMed]
- Roxrud, I.; Lid, S.E.; Fletcher, J.C.; Schmidt, E.D.; Opsahl-Sorteberg, H.G. GASA4, one of the 14-member Arabidopsis GASA family of small polypeptides, regulates flowering and seed development. Plant Cell Physiol. 2007, 48, 471–483. [Google Scholar] [CrossRef]
- Trapalis, M.; Li, S.F.; Parish, R.W. The Arabidopsis GASA10 gene encodes a cell wall protein strongly expressed in developing anthers and seeds. Plant Sci. 2017, 260, 71–79. [Google Scholar] [CrossRef]
- Zhang, H.; Yue, M.; Zheng, X.; Gautam, M.; He, S.; Li, L. The Role of Promoter-Associated Histone Acetylation of Haem Oxygenase-1 (HO-1) and Giberellic Acid-Stimulated Like-1 (GSL-1) Genes in Heat-Induced Lateral Root Primordium Inhibition in Maize. Front. Plant Sci. 2018, 9, 1520. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, S.; Wang, X. One new kind of phytohormonal signaling integrator: Up-and-coming GASA family genes. Plant Signal. Behav. 2017, 12, e1226453. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- An, B.; Wang, Q.; Zhang, X.; Zhang, B.; Luo, H.; He, C. Comprehensive transcriptional and functional analyses of HbGASA genes reveal their roles in fungal pathogen resistance in Hevea brasiliensis. Tree Genet. Genomes 2018, 14, 41. [Google Scholar] [CrossRef]
- Cheng, X.; Wang, S.; Xu, D.; Liu, X.; Li, X.; Xiao, W.; Cao, J.; Jiang, H.; Min, X.; Wang, J.; et al. Identification and Analysis of the GASR Gene Family in Common Wheat (Triticum aestivum L.) and Characterization of TaGASR34, a Gene Associated With Seed Dormancy and Germination. Front. Genet. 2019, 10, 980. [Google Scholar] [CrossRef]
- Zhang, S.; Wang, X. Expression pattern of GASA, downstream genes of DELLA, in Arabidopsis. Sci. Bull. 2008, 53, 3839–3846. [Google Scholar] [CrossRef] [Green Version]
- Qiao, K.; Ma, C.; Lv, J.; Zhang, C.; Ma, Q.; Fan, S. Identification, characterization, and expression profiles of the GASA genes in cotton. J. Cotton Res. 2021, 4, 1–16. [Google Scholar] [CrossRef]
- Wang, H.; Wei, T.; Wang, X.; Zhang, L.; Yang, M.; Chen, L.; Song, W.; Wang, C.; Chen, C. Transcriptome Analyses from Mutant Salvia miltiorrhiza Reveals Important Roles for SmGASA4 during Plant Development. Int. J. Mol. Sci. 2018, 19, 2088. [Google Scholar] [CrossRef]
- Sun, S.; Wang, H.; Yu, H.; Zhong, C.; Zhang, X.; Peng, J.; Wang, X. GASA14 regulates leaf expansion and abiotic stress resistance by modulating reactive oxygen species accumulation. J. Exp. Bot. 2013, 64, 1637–1647. [Google Scholar] [CrossRef] [Green Version]
- Iqbal, A.; Wang, T.; Wu, G.; Tang, W.; Zhu, C.; Wang, D.; Li, Y.; Wang, H. Physiological and transcriptome analysis of heteromorphic leaves and hydrophilic roots in response to soil drying in desert Populus euphratica. Sci. Rep. 2017, 7, 1–15. [Google Scholar] [CrossRef] [Green Version]
- Li, B.; Qin, Y.; Duan, H.; Yin, W.; Xia, X. Genome-wide characterization of new and drought stress responsive microRNAs in Populus euphratica. J. Exp. Bot. 2011, 62, 3765–3779. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ottow, E.A.; Polle, A.; Brosché, M.; Kangasjärvi, J.; Dibrov, P.; Zörb, C.; Teichmann, T. Molecular characterization of PeNhaD1: The first member of the NhaD Na+/H+ antiporter family of plant origin. Plant Mol. Biol. 2005, 58, 75–88. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Z.; Chen, Y.; Zhang, J.; Ma, X.; Li, Y.; Li, M.; Wang, D.; Kang, M.; Wu, H.; Yang, Y.; et al. Improved genome assembly provides new insights into genome evolution in a desert poplar (Populus euphratica). Mol. Ecol. Resour. 2020, 20, 781–794. [Google Scholar] [CrossRef] [PubMed]
- Tang, S.; Liang, H.; Yan, D.; Zhao, Y.; Han, X.; Carlson, J.E.; Xia, X.; Yin, W. Populus euphratica: The transcriptomic response to drought stress. Plant Mol. Biol. 2013, 83, 539–557. [Google Scholar] [CrossRef] [PubMed]
- Yan, D.-H.; Fenning, T.; Tang, S.; Xia, X.; Yin, W. Genome-wide transcriptional response of Populus euphratica to long-term drought stress. Plant Sci. 2012, 195, 24–35. [Google Scholar] [CrossRef] [PubMed]
- Huang, D.; Wu, W.; Abrams, S.R.; Cutler, A.J. The relationship of drought-related gene expression in Arabidopsis thaliana to hormonal and environmental factors. J. Exp. Bot. 2008, 59, 2991–3007. [Google Scholar] [CrossRef] [Green Version]
- Ma, T.; Wang, J.; Zhou, G.; Yue, Z.; Hu, Q.; Chen, Y.; Liu, B.; Qiu, Q.; Wang, Z.; Zhang, J.; et al. Genomic insights into salt adaptation in a desert poplar. Nat. Commun. 2013, 4, 2797. [Google Scholar] [CrossRef]
- Yang, Q.; Han, X.M.; Gu, J.K.; Liu, Y.J.; Yang, M.J.; Zeng, Q.Y. Functional and structural profiles of GST gene family from three Populus species reveal the sequence–function decoupling of orthologous genes. New Phytol. 2018, 221, 1060–1073. [Google Scholar] [CrossRef] [Green Version]
- Tuskan, G.A.; Difazio, S.; Jansson, S.; Bohlmann, J.; Grigoriev, I.; Hellsten, U.; Putnam, N.; Ralph, S.; Rombauts, S.; Salamov, A.; et al. The genome of black cottonwood, Populus trichocarpa (Torr. & Gray). Science 2006, 313, 1596–1604. [Google Scholar]
- Ma, J.; Lu, J.; Xu, J.; Duan, B.; He, X.; Liu, J. Genome-wide Identification of WRKY Genes in the Desert Poplar Populus euphratica and Adaptive Evolution of the Genes in Response to Salt Stress. Evol. Bioinform. Online 2015, 11 (Suppl. 1), 47–55. [Google Scholar] [CrossRef] [PubMed]
- Nahirñak, V.; Rivarola, M.; Gonzalez de Urreta, M.; Paniego, N.; Hopp, H.E.; Almasia, N.I.; Vazquez-Rovere, C. Genome-wide Analysis of the Snakin/GASA Gene Family in Solanum tuberosum cv. Kennebec. Am. J. Potato Res. 2016, 93, 172–188. [Google Scholar] [CrossRef]
- Ben-Nissan, G.; Lee, J.-Y.; Borohov, A.; Weiss, D. GIP, a Petunia hybrida GA-induced cysteine-rich protein: A possible role in shoot elongation and transition to flowering. Plant J. 2004, 37, 229–238. [Google Scholar] [CrossRef]
- Furukawa, T.; Sakaguchi, N.; Shimada, H. Two OsGASR genes, rice GAST homologue genes that are abundant in proliferating tissues, show different expression patterns in developing panicles. Genes Genet. Syst 2006, 81, 171–180. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, K.-L.; Bai, X.; Li, Y.; Cai, H.; Ji, W.; Tang, L.-L.; Wen, Y.-D.; Zhu, Y.-M. GsGASA1 mediated root growth inhibition in response to chronic cold stress is marked by the accumulation of DELLAs. J. Plant Physiol. 2011, 168, 2153–2160. [Google Scholar] [CrossRef] [PubMed]
- Peng, J.; Lai, L.; Wang, X. PRGL: A cell wall proline-rich protein containning GASA domain in Gerbera hybrida. Sci. China Ser. C Life Sci. 2008, 51, 520–525. [Google Scholar] [CrossRef]
- Cannon, S.B.; Mitra, A.; Baumgarten, A.; Young, N.D.; May, G. The roles of segmental and tandem gene duplication in the evolution of large gene families in Arabidopsis thaliana. BMC Plant Biol. 2004, 4, 10. [Google Scholar] [CrossRef] [Green Version]
- Qiao, X.; Li, Q.; Yin, H.; Qi, K.; Li, L.; Wang, R.; Zhang, S.; Paterson, A.H. Gene duplication and evolution in recurring polyploidization–diploidization cycles in plants. Genome Biol. 2019, 20, 1–23. [Google Scholar] [CrossRef] [Green Version]
- Alonso-Peral, M.M.; Li, J.; Li, Y.; Allen, R.S.; Schnippenkoetter, W.; Ohms, S.; White, R.G.; Millar, A.A. The microRNA159-regulated GAMYB-like genes inhibit growth and promote programmed cell death in Arabidopsis. Plant Physiol. 2010, 154, 757–771. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, S.; Shi, Q.; Albrecht, U.; Shatters, R.G., Jr.; Stange, R.; McCollum, G.; Zhang, S.; Fan, C.; Stover, E. Comparative transcriptome analysis during early fruit development between three seedy citrus genotypes and their seedless mutants. Hortic. Res. 2017, 4, 17041. [Google Scholar] [CrossRef]
- Gao, X.; Wang, L.; Zhang, H.; Zhu, B.; Lv, G.; Xiao, J. Transcriptome analysis and identification of genes associated with floral transition and fruit development in rabbiteye blueberry (Vaccinium ashei). PLoS ONE 2021, 16, e0259119. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Guo, W.; Liu, B.; Zhang, Y.; Song, X.; Cheng, Y.; Zhang, L.; Zhang, T. Molecular mechanisms of fiber differential development between G. barbadense and G. hirsutum revealed by genetical genomics. PLoS ONE 2012, 7, e30056. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lin, M.; Pang, C.; Fan, S.; Song, M.; Wei, H.; Yu, S. Global analysis of the Gossypium hirsutum L. Transcriptome during leaf senescence by RNA-Seq. BMC Plant Biol. 2015, 15, 43. [Google Scholar] [CrossRef] [Green Version]
- Tobar, M.; Fiore, N.; Perez-Donoso, A.G.; Leon, R.; Rosales, I.M.; Gambardella, M. Divergent molecular and growth responses of young “Cabernet Sauvignon” (Vitis vinifera) plants to simple and mixed infections with Grapevine rupestris stem pitting-associated virus. Hortic. Res. 2020, 7, 1–14. [Google Scholar] [CrossRef] [Green Version]
- Dias, P.M.; Brunel-Muguet, S.; Durr, C.; Huguet, T.; Demilly, D.; Wagner, M.H.; Teulat-Merah, B. QTL analysis of seed germination and pre-emergence growth at extreme temperatures in Medicago truncatula. Theor. Appl. Genet. 2011, 122, 429–444. [Google Scholar] [CrossRef] [Green Version]
- Bouquin, T.; Meier, C.; Foster, R.; Nielsen, M.E.; Mundy, J. Control of Specific Gene Expression by Gibberellin and Brassinosteroid. Plant Physiol. 2001, 127, 450–458. [Google Scholar] [CrossRef]
- Paponov, I.A.; Paponov, M.; Teale, W.; Menges, M.; Chakrabortee, S.; Murray, J.A.; Palme, K. Comprehensive transcriptome analysis of auxin responses in Arabidopsis. Mol. Plant 2008, 1, 321–337. [Google Scholar] [CrossRef]
- Ceserani, T.; Trofka, A.; Gandotra, N.; Nelson, T. VH1/BRL2 receptor-like kinase interacts with vascular-specific adaptor proteins VIT and VIK to influence leaf venation. Plant J. 2009, 57, 1000–1014. [Google Scholar] [CrossRef] [Green Version]
- Lakhssassi, N.; Doblas, V.G.; Rosado, A.; del Valle, A.E.; Pose, D.; Jimenez, A.J.; Castillo, A.G.; Valpuesta, V.; Borsani, O.; Botella, M.A. The Arabidopsis tetratricopeptide thioredoxin-like gene family is required for osmotic stress tolerance and male sporogenesis. Plant Physiol. 2012, 158, 1252–1266. [Google Scholar] [CrossRef] [Green Version]
- Amorim-Silva, V.; Garcia-Moreno, A.; Castillo, A.G.; Lakhssassi, N.; Esteban Del Valle, A.; Perez-Sancho, J.; Li, Y.; Pose, D.; Perez-Rodriguez, J.; Lin, J.; et al. TTL Proteins Scaffold Brassinosteroid Signaling Components at the Plasma Membrane to Optimize Signal Transduction in Arabidopsis. Plant Cell 2019, 31, 1807–1828. [Google Scholar] [CrossRef] [Green Version]
- Rosado, A.; Schapire, A.L.; Bressan, R.A.; Harfouche, A.L.; Hasegawa, P.M.; Valpuesta, V.; Botella, M.A. The Arabidopsis tetratricopeptide repeat-containing protein TTL1 is required for osmotic stress responses and abscisic acid sensitivity. Plant Physiol. 2006, 142, 1113–1126. [Google Scholar] [CrossRef] [Green Version]
- Cuadrado-Pedetti, M.B.; Rauschert, I.; Sainz, M.M.; Amorim-Silva, V.; Botella, M.A.; Borsani, O.; Sotelo-Silveira, M. The Arabidopsis TETRATRICOPEPTIDE THIOREDOXIN-LIKE 1 Gene Is Involved in Anisotropic Root Growth during Osmotic Stress Adaptation. Genes 2021, 12, 236. [Google Scholar] [CrossRef] [PubMed]
- Imtiaz, M.; Yang, Y.; Liu, R.; Xu, Y.; Khan, M.A.; Wei, Q.; Gao, J.; Hong, B. Identification and functional characterization of the BBX24 promoter and gene from chrysanthemum in Arabidopsis. Plant Mol. Biol. 2015, 89, 1–19. [Google Scholar] [CrossRef] [PubMed]
- Lucas, W.J.; Groover, A.; Lichtenberger, R.; Furuta, K.; Yadav, S.R.; Helariutta, Y.; He, X.Q.; Fukuda, H.; Kang, J.; Brady, S.M.; et al. The plant vascular system: Evolution, development and functions. J. Integr. Plant Biol. 2013, 55, 294–388. [Google Scholar] [CrossRef] [PubMed]
- Ray, S.; Agarwal, P.; Arora, R.; Kapoor, S.; Tyagi, A.K. Expression analysis of calcium-dependent protein kinase gene family during reproductive development and abiotic stress conditions in rice (Oryza sativa L. ssp. indica). Mol. Genet. Genom. 2007, 278, 493–505. [Google Scholar] [CrossRef]
- Ljung, K.; Hull, A.K.; Celenza, J.; Yamada, M.; Estelle, M.; Normanly, J.; Sandberg, G. Sites and regulation of auxin biosynthesis in Arabidopsis roots. Plant Cell 2005, 17, 1090–1104. [Google Scholar] [CrossRef] [PubMed]
- Powers, S.K.; Strader, L.C. Regulation of auxin transcriptional responses. Dev. Dyn. 2020, 249, 483–495. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yamaguchi, N.; Winter, C.M.; Wu, M.F.; Kanno, Y.; Yamaguchi, A.; Seo, M.; Wagner, D. Gibberellin acts positively then negatively to control onset of flower formation in Arabidopsis. Science 2014, 344, 638–641. [Google Scholar] [CrossRef]
- Rizza, A.; Jones, A.M. The makings of a gradient: Spatiotemporal distribution of gibberellins in plant development. Curr. Opin. Plant Biol. 2019, 47, 9–15. [Google Scholar] [CrossRef]
- Tamura, K.; Stecher, G.; Peterson, D.; Filipski, A.; Kumar, S. MEGA6: Molecular Evolutionary Genetics Analysis version 6.0. Mol. Biol. Evol. 2013, 30, 2725–2729. [Google Scholar] [CrossRef] [Green Version]
- Yu, Z.; Zhao, C.; Zhang, G.; Teixeira da Silva, J.A.; Duan, J. Genome-Wide Identification and Expression Profile of TPS Gene Family in Dendrobium officinale and the Role of DoTPS10 in Linalool Biosynthesis. Int. J. Mol. Sci. 2020, 21, 5419. [Google Scholar] [CrossRef] [PubMed]
- Huang, W.; Hu, B.; Liu, J.; Zhou, Y.; Liu, S. Identification and Characterization of Tonoplast Sugar Transporter (TST) Gene Family in Cucumber. Hortic. Plant J. 2020, 6, 145–157. [Google Scholar] [CrossRef]
- Chao, J.; Huang, Z.; Yang, S.; Deng, X.; Tian, W. Genome-wide identification and expression analysis of the phosphatase 2A family in rubber tree (Hevea brasiliensis). PLoS ONE 2020, 15, e0228219. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, Y.; Tang, H.; Debarry, J.D.; Tan, X.; Li, J.; Wang, X.; Lee, T.H.; Jin, H.; Marler, B.; Guo, H.; et al. MCScanX: A toolkit for detection and evolutionary analysis of gene synteny and collinearity. Nucleic Acids Res. 2012, 40, e49. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chen, C.; Chen, H.; Zhang, Y.; Thomas, H.R.; Frank, M.H.; He, Y.; Xia, R. TBtools: An Integrative Toolkit Developed for Interactive Analyses of Big Biological Data. Mol. Plant 2020, 13, 1194–1202. [Google Scholar] [CrossRef]
- Wang, D.; Gao, Z.; Du, P.; Xiao, W.; Tan, Q.; Chen, X.; Li, L.; Gao, D. Expression of ABA Metabolism-Related Genes Suggests Similarities and Differences Between Seed Dormancy and Bud Dormancy of Peach (Prunus persica). Front Plant Sci. 2015, 6, 1248. [Google Scholar] [CrossRef]
- Zhang, H.; Hong, Y.; Huang, L.; Liu, S.; Tian, L.; Dai, Y.; Cao, Z.; Huang, L.; Li, D.; Song, F. Virus-Induced Gene Silencing-Based Functional Analyses Revealed the Involvement of Several Putative Trehalose-6-Phosphate Synthase/Phosphatase Genes in Disease Resistance against Botrytis cinerea and Pseudomonas syringae pv. tomato DC3000 in Tomato. Front Plant Sci. 2016, 7, 1176. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.L.; Li, L.; Tang, S.; Yuan, C.; Tian, Q.; Su, Y.; Li, H.G.; Zhao, L.; Yin, W.; Zhao, R.; et al. Evaluation of Appropriate Reference Genes for Reverse Transcription-Quantitative PCR Studies in Different Tissues of a Desert Poplar via Comparision of Different Algorithms. Int. J. Mol. Sci. 2015, 16, 20468–20491. [Google Scholar] [CrossRef] [Green Version]
- Wang, H.L.; Zhang, Y.; Wang, T.; Yang, Q.; Yang, Y.; Li, Z.; Li, B.; Wen, X.; Li, W.; Yin, W.; et al. An alternative splicing variant of PtRD26 delays leaf senescence by regulating multiple NAC transcription factors in Populus. Plant Cell 2021, 33, 1594–1614. [Google Scholar] [CrossRef]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]
- Zhao, R.; Sun, H.L.; Mei, C.; Wang, X.J.; Yan, L.; Liu, R.; Zhang, X.F.; Wang, X.F.; Zhang, D.P. The Arabidopsis Ca(2+) -dependent protein kinase CPK12 negatively regulates abscisic acid signaling in seed germination and post-germination growth. New Phytol. 2011, 192, 61–73. [Google Scholar] [CrossRef] [PubMed]
- Li, X. Infiltration of Nicotiana benthamiana Protocol for Transient Expression via Agrobacterium. Bio-Protocol 2011, 1, e95. [Google Scholar] [CrossRef] [Green Version]
- Clough, S.J.; Bent, A.F. Floral dip: A simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J. 1998, 16, 735–743. [Google Scholar] [CrossRef] [Green Version]


| (a) Identification in the Protein Database Reported by Ma et al. | |||||
|---|---|---|---|---|---|
| Protein ID | Transcript ID | Gene | Gene Description | CDS (bp) | Peptide (aa) |
| XP_011001772.1 | XM_011003470.1 | LOC105108948 | gibberellin-regulated protein 9-like | 333 | 110 |
| XP_011015007.1 | XM_011016705.1 | LOC105118698 | gibberellin-regulated protein 9-like | 333 | 110 |
| XP_011016723.1 | XM_011018421.1 | LOC105120240 | snakin-2-like | 330 | 109 |
| XP_011021371.1 | XM_011023069.1 | LOC105123460 | protein RSI-1 | 288 | 95 |
| XP_011027306.1 | XM_011029004.1 | LOC105127637 | protein GAST1-like | 351 | 116 |
| XP_011028622.1 /XP_011028623.1 | XM_011030320.1 /XM_011030321.1 | LOC105128593 | snakin-2-like | 342 | 113 |
| XP_011029010.1 | XM_011030708.1 | LOC105128863 | gibberellin-regulated protein 11-like | 357 | 118 |
| XP_011029011.1 | XM_011030709.1 | LOC105128865 | gibberellin-regulated protein 1-like | 309 | 102 |
| XP_011029012.1 | XM_011030710.1 | LOC105128866 | gibberellin-regulated protein 1-like | 309 | 102 |
| XP_011029030.1 | XM_011030728.1 | LOC105128880 | peamaclein-like | 267 | 88 |
| XP_011035751.1 | XM_011037449.1 | LOC105133442 | gibberellin-regulated protein 4 | 321 | 106 |
| XP_011038137.1 | XM_011039835.1 | LOC105135117 | gibberellin-regulated protein 2 | 276 | 91 |
| XP_011044353.1 | XM_011046051.1 | LOC105139569 | snakin-2-like | 330 | 109 |
| XP_011045095.1 | XM_011046793.1 | LOC105140101 | gibberellin-regulated protein 1-like | 312 | 103 |
| XP_011045096.1 | XM_011046794.1 | LOC105140102 | gibberellin-regulated protein 1-like | 309 | 102 |
| XP_011045097.1 | XM_011046795.1 | LOC105140103 | gibberellin-regulated protein 11-like | 297 | 98 |
| XP_011045379.1 | XM_011047077.1 | LOC105140300 | protein GAST1-like | 342 | 113 |
| XP_011047278.1 | XM_011048976.1 | LOC105141680 | gibberellin-regulated protein 14-like | 669 | 222 |
| (b) Identification in the Protein Database Reported by Zhang et al. | |||||
| Protein | mRNA | Gene | CDS (bp) | Peptide (aa) | BLAST |
| GWHPAAYU001024 | GWHTAAYU001024 | GWHGAAYU001024 | 477 | 158 | (XP_011047278.1) |
| GWHPAAYU001570 | GWHTAAYU001570 | GWHGAAYU001570 | 267 | 88 | No |
| GWHPAAYU001972 | GWHTAAYU001972 | GWHGAAYU001972 | 342 | 113 | XP_011045379.1 |
| GWHPAAYU008043 | GWHTAAYU008043 | GWHGAAYU008043 | 321 | 106 | XP_011035751.1 |
| GWHPAAYU008049 | GWHTAAYU008049 | GWHGAAYU008049 | 321 | 106 | XP_011035751.1 |
| GWHPAAYU008363 | GWHTAAYU008363 | GWHGAAYU008363 | 288 | 95 | XP_011021371.1 |
| GWHPAAYU012087 | GWHTAAYU012087 | GWHGAAYU012087 | 342 | 113 | XP_011028622.1 /XP_011028623.1 |
| GWHPAAYU013554 | GWHTAAYU013554 | GWHGAAYU013554 | 330 | 109 | XP_011016723.1 |
| GWHPAAYU016490 | GWHTAAYU016490 | GWHGAAYU016490 | 309 | 102 | XP_011029012.1 |
| GWHPAAYU016491 | GWHTAAYU016491 | GWHGAAYU016491 | 309 | 102 | XP_011029011.1 |
| GWHPAAYU025413 | GWHTAAYU025413 | GWHGAAYU025413 | 351 | 116 | XP_011027306.1 |
| GWHPAAYU029870 | GWHTAAYU029870 | GWHGAAYU029870 | 309 | 102 | XP_011045096.1 |
| GWHPAAYU029871 | GWHTAAYU029871 | GWHGAAYU029871 | 312 | 103 | XP_011045095.1 |
| GWHPAAYU031162 | GWHTAAYU031162 | GWHGAAYU031162 | 276 | 91 | XP_011038137.1 |
| Gene 1 | Gene 2 | Duplication | E-Value | Ka | Ks | Ka/Ks | Selection Pressure |
|---|---|---|---|---|---|---|---|
| PtrGASA2 | PtrGASA11 | WGD | 5E-16 | 0.306574081 | 1.511177265 | 0.202871025 | Purifying selection |
| PtrGASA2 | PtrGASA13 | WGD | 1E-41 | 0.024592367 | 0.245409683 | 0.100209441 | Purifying selection |
| PtrGASA2 | PtrGASA16 | WGD | 3E-34 | 0.189006113 | 0.854755332 | 0.221123058 | Purifying selection |
| PtrGASA11 | PtrGASA13 | WGD | 8E-14 | 0.314370832 | 1.291132145 | 0.24348463 | Purifying selection |
| PtrGASA11 | PtrGASA16 | WGD | 2E-14 | 0.2812637 | 1.166421628 | 0.241133818 | Purifying selection |
| PtrGASAd13 | PtrGASA16 | WGD | 6E-34 | 0.175785289 | 1.055787598 | 0.166496831 | Purifying selection |
| PtrGASA7 | PtrGASA9 | WGD | 1E-33 | 0.189204442 | 0.883649795 | 0.214116998 | Purifying selection |
| PtrGASA14 | PtrGASA17 | WGD | 4E-68 | 0.041106177 | 0.225565616 | 0.182236008 | Purifying selection |
| PtrGASA5 | PtrGASA6 | TD | 4E-31 | 0.257142422 | 0.939998969 | 0.273556068 | Purifying selection |
| PtrGASA6 | PtrGASA7 | TD | 1E-34 | 0.162561762 | 0.966817469 | 0.16814111 | Purifying selection |
| PtrGASA8 | PtrGASA9 | TD | 7E-34 | 0.320382084 | 1.907862372 | 0.167927251 | Purifying selection |
| PtrGASA11 | PtrGASA12 | TD | 4E-30 | 0 | 0 | NaN | - |
| PeuGASA6 | PeuGASA13 | WGD | 6E-49 | 0.057332466 | 0.184523724 | 0.31070512 | Purifying selection |
| PeuGASA9 | PeuGASA15 | WGD | 2E-39 | 0.097292607 | 0.362138829 | 0.268661075 | Purifying selection |
| PeuGASA10 | PeuGASA19 | WGD | 2E-34 | 0.195458562 | 0.965443142 | 0.202454762 | Purifying selection |
| PeuGASA7 | PeuGASA8 | TD | 2E-32 | 0.204373366 | 0.898115959 | 0.227557883 | Purifying selection |
| PeuGASA8 | PeuGASA9 | TD | 6E-30 | 0.185223217 | 0.962509794 | 0.192437747 | Purifying selection |
| PeuGASA14 | PeuGASA15 | TD | 1E-44 | 0.253240964 | 0.898681597 | 0.281791644 | Purifying selection |
| PeuGASA15 | PeuGASA16 | TD | 3E-35 | 0.297194132 | 1.782104232 | 0.166765853 | Purifying selection |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Han, S.; Jiao, Z.; Niu, M.-X.; Yu, X.; Huang, M.; Liu, C.; Wang, H.-L.; Zhou, Y.; Mao, W.; Wang, X.; et al. Genome-Wide Comprehensive Analysis of the GASA Gene Family in Populus. Int. J. Mol. Sci. 2021, 22, 12336. https://doi.org/10.3390/ijms222212336
Han S, Jiao Z, Niu M-X, Yu X, Huang M, Liu C, Wang H-L, Zhou Y, Mao W, Wang X, et al. Genome-Wide Comprehensive Analysis of the GASA Gene Family in Populus. International Journal of Molecular Sciences. 2021; 22(22):12336. https://doi.org/10.3390/ijms222212336
Chicago/Turabian StyleHan, Shuo, Zhiyin Jiao, Meng-Xue Niu, Xiao Yu, Mengbo Huang, Chao Liu, Hou-Ling Wang, Yangyan Zhou, Wei Mao, Xiaofei Wang, and et al. 2021. "Genome-Wide Comprehensive Analysis of the GASA Gene Family in Populus" International Journal of Molecular Sciences 22, no. 22: 12336. https://doi.org/10.3390/ijms222212336
APA StyleHan, S., Jiao, Z., Niu, M.-X., Yu, X., Huang, M., Liu, C., Wang, H.-L., Zhou, Y., Mao, W., Wang, X., Yin, W., & Xia, X. (2021). Genome-Wide Comprehensive Analysis of the GASA Gene Family in Populus. International Journal of Molecular Sciences, 22(22), 12336. https://doi.org/10.3390/ijms222212336






