Expression and Functionality of Connexin-Based Channels in Human Liver Cancer Cell Lines
Abstract
1. Introduction
2. Results
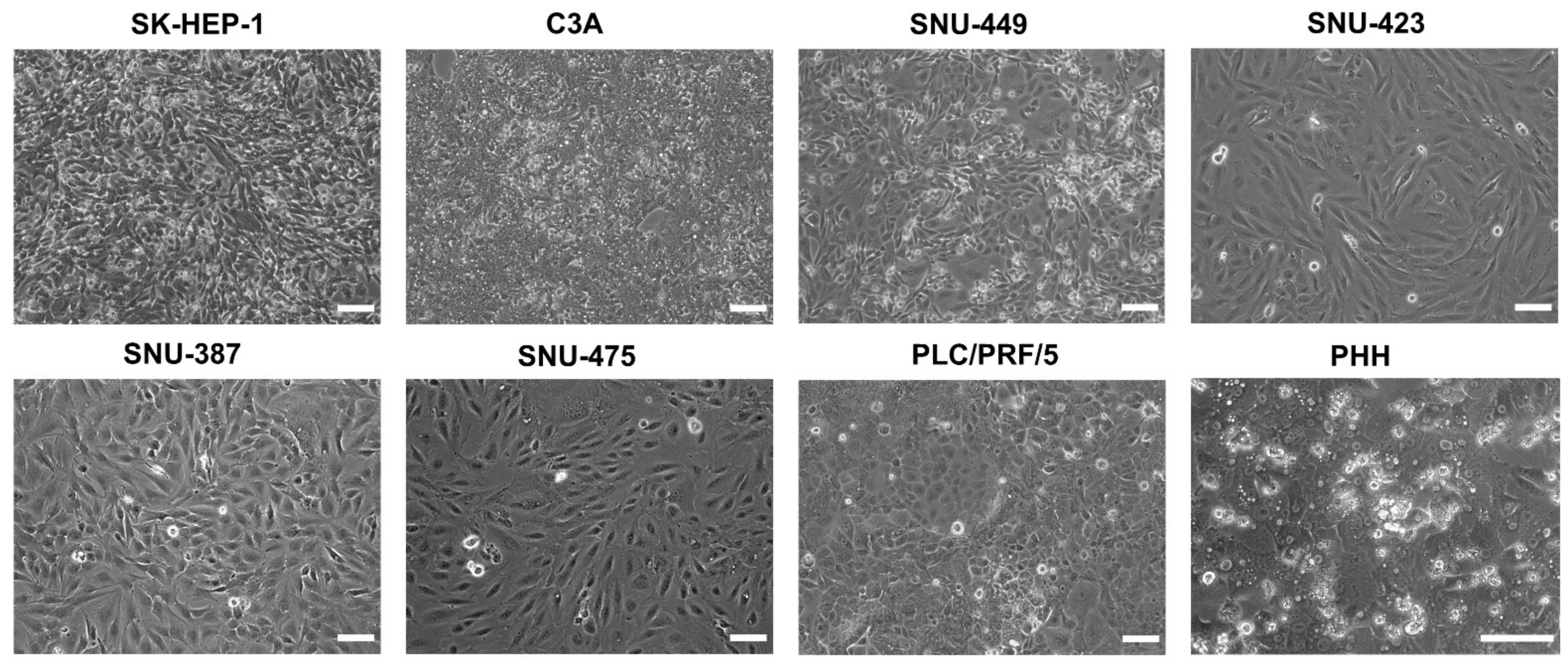
2.1. Selection of Liver Cancer Cell Lines
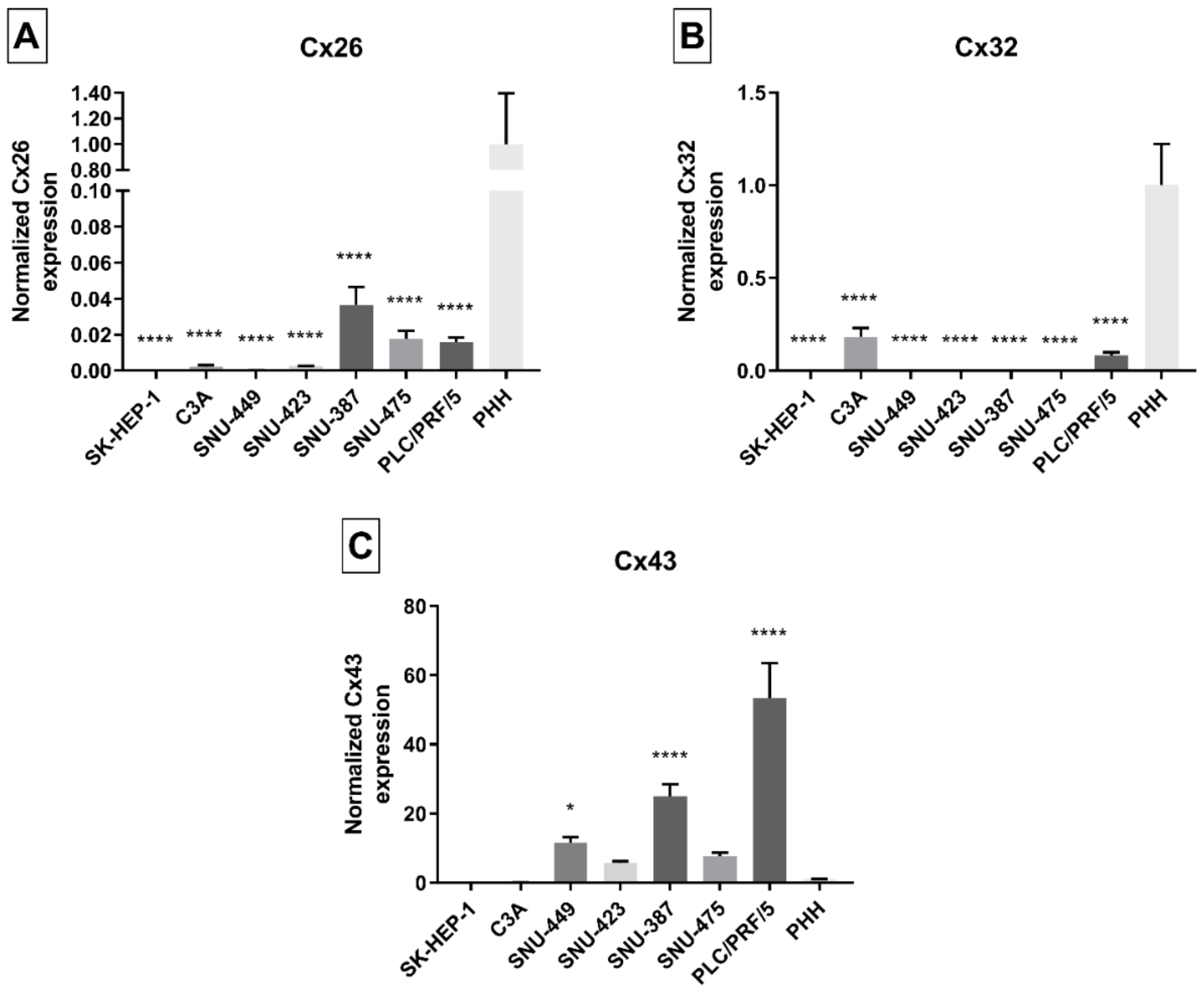
2.2. Connexin Gene Expression in Liver Cancer Cell Lines
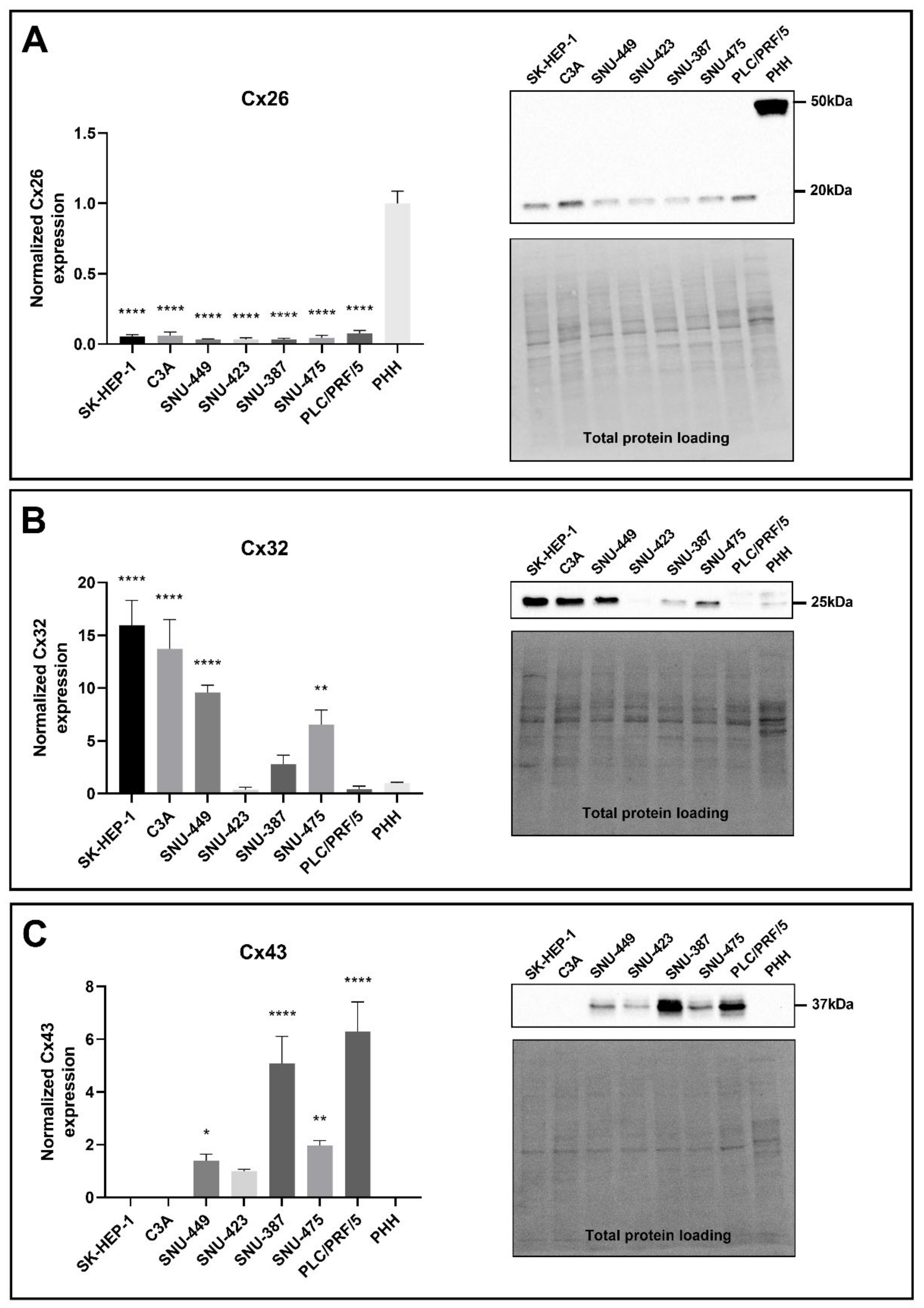
2.3. Connexin Protein Expression in Liver Cancer Cell Lines
2.4. Connexin Protein Subcellular Localization in Liver Cancer Cell Lines
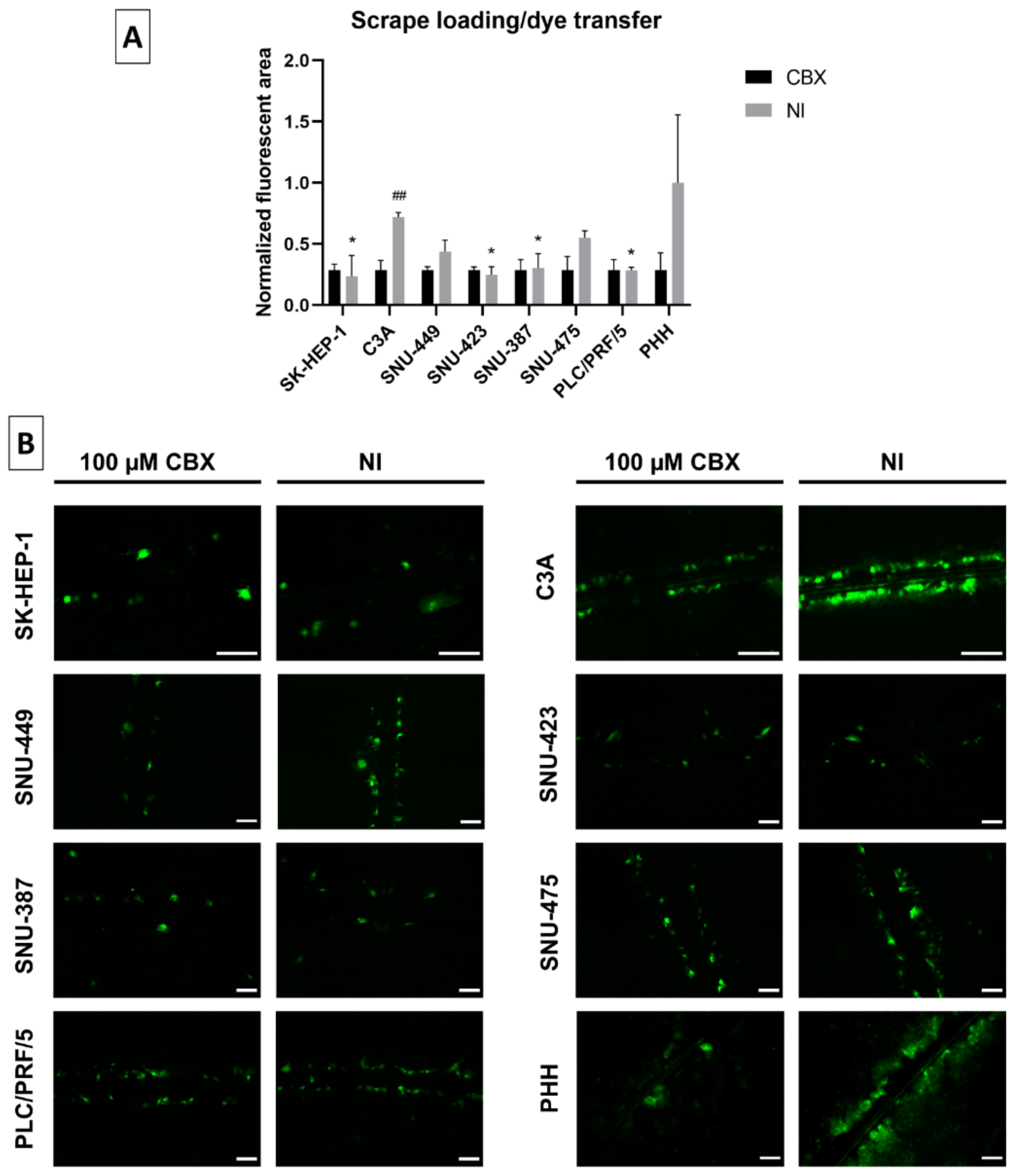
2.5. Gap Junctional Intercellular Communication in Liver Cancer Cell Lines
3. Discussion
4. Materials and Methods
4.1. Reagents
4.2. Liver Cancer Cell Lines and Primary Human Hepatocytes
4.3. Phase-Contrast Microscopy
4.4. Real-Time Quantitative Reverse Transcription Polymerase Chain Reaction Analysis
4.5. Immunoblot Analysis
4.6. Immunocytochemistry Analysis
4.7. Scrape Loading/Dye Transfer Assay
4.8. Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Sung, H.; Ferlay, J.; Siegel, R.L.; Laversanne, M.; Soerjomataram, I.; Jemal, A.; Bray, F. Global Cancer Statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 2021, 71, 209–249. [Google Scholar] [CrossRef] [PubMed]
- Koumbi, L. Dietary factors can protect against liver cancer development. World J. Hepatol. 2017, 9, 119–125. [Google Scholar] [CrossRef]
- Sia, D.; Villanueva, A.; Friedman, S.L.; Llovet, J.M. Liver cancer cell of origin, molecular class, and effects on patient prognosis. Gastroenterology 2017, 152, 745–761. [Google Scholar] [CrossRef]
- Alqahtani, A.; Khan, Z.; Alloghbi, A.; Said Ahmed, T.S.; Ashraf, M.; Hammouda, D.M. Hepatocellular carcinoma: Molecular mechanisms and targeted therapies. Medicina 2019, 55, 526. [Google Scholar] [CrossRef]
- Collins, S.D.; Yuen, G.; Tu, T.; Budzinska, M.A.; Spring, K.; Bryant, K.; Shackel, N.A. In vitro models of the liver: Disease modeling, drug discovery and clinical applications. In Hepatocellular Carcinoma; Tirnitz-Parker, J.E.E., Ed.; Codon Publications: Brisbane, Australia, 2019. [Google Scholar]
- Hirschfield, H.; Bian, C.B.; Higashi, T.; Nakagawa, S.; Zeleke, T.Z.; Nair, V.D.; Fuchs, B.C.; Hoshida, Y. In vitro modeling of hepatocellular carcinoma molecular subtypes for anti-cancer drug assessment. Exp. Mol. Med. 2018, 50, e419. [Google Scholar] [CrossRef]
- Carlessi, R.; Kohn-Gaone, J.; Olynyk, J.K.; Tirnitz-Parker, J.E.E. Mouse models of hepatocellular carcinoma. In Hepatocellular Carcinoma; Tirnitz-Parker, J.E.E., Ed.; Codon Publications: Brisbane, Australia, 2019. [Google Scholar]
- Bagi, C.M.; Andresen, C.J. Models of hepatocellular carcinoma and biomarker strategy. Cancers 2010, 2, 1441–1452. [Google Scholar] [CrossRef] [PubMed]
- Febbraio, M.A.; Reibe, S.; Shalapour, S.; Ooi, G.J.; Watt, M.J.; Karin, M. Preclinical models for studying NASH-driven HCC: How useful are they? Cell Metab. 2019, 29, 18–26. [Google Scholar] [CrossRef]
- Schachtschneider, K.M.; Schwind, R.M.; Darfour-Oduro, K.A.; De, A.K.; Rund, L.A.; Singh, K.; Principe, D.R.; Guzman, G.; Ray, C.E., Jr.; Ozer, H.; et al. A validated, transitional and translational porcine model of hepatocellular carcinoma. Oncotarget 2017, 8, 63620–63634. [Google Scholar] [CrossRef]
- Zhang, Q.; Li, J.; Middleton, A.; Bhattacharya, S.; Conolly, R.B. Bridging the data gap from in vitro toxicity testing to chemical safety assessment through computational modeling. Front. Public Health 2018, 6, 261. [Google Scholar] [CrossRef]
- Willebrords, J.; Crespo Yanguas, S.; Maes, M.; Decrock, E.; Wang, N.; Leybaert, L.; da Silva, T.C.; Veloso Alves Pereira, I.; Jaeschke, H.; Cogliati, B.; et al. Structure, regulation and function of gap junctions in liver. Cell Commun. Adhes. 2015, 22, 29–37. [Google Scholar] [CrossRef] [PubMed]
- Zefferino, R.; Piccoli, C.; Gioia, S.D.; Capitanio, N.; Conese, M. Gap junction intercellular communication in the carcinogenesis hallmarks: Is this a phenomenon or epiphenomenon? Cells 2019, 8, 896. [Google Scholar] [CrossRef] [PubMed]
- Neveu, M.J.; Hully, J.R.; Babcock, K.L.; Vaughan, J.; Hertzberg, E.L.; Nicholson, B.J.; Paul, D.L.; Pitot, H.C. Proliferation-associated differences in the spatial and temporal expression of gap junction genes in rat liver. Hepatology 1995, 22, 202–212. [Google Scholar]
- Eugenin, E.A.; Gonzalez, H.E.; Sanchez, H.A.; Branes, M.C.; Saez, J.C. Inflammatory conditions induce gap junctional communication between rat Kupffer cells both in vivo and in vitro. Cell Immunol. 2007, 247, 103–110. [Google Scholar] [CrossRef]
- Fischer, R.; Reinehr, R.; Lu, T.P.; Schonicke, A.; Warskulat, U.; Dienes, H.P.; Haussinger, D. Intercellular communication via gap junctions in activated rat hepatic stellate cells. Gastroenterology 2005, 128, 433–448. [Google Scholar] [CrossRef] [PubMed]
- Hernandez-Guerra, M.; Hadjihambi, A.; Jalan, R. Gap junctions in liver disease: Implications for pathogenesis and therapy. J. Hepatol. 2019, 70, 759–772. [Google Scholar] [CrossRef]
- Maes, M.; Crespo Yanguas, S.; Willebrords, J.; Cogliati, B.; Vinken, M. Connexin and pannexin signaling in gastrointestinal and liver disease. Transl. Res. 2015, 166, 332–343. [Google Scholar] [CrossRef]
- Krutovskikh, V.; Mazzoleni, G.; Mironov, N.; Omori, Y.; Aguelon, A.M.; Mesnil, M.; Berger, F.; Partensky, C.; Yamasaki, H. Altered homologous and heterologous gap-junctional intercellular communication in primary human liver tumors associated with aberrant protein localization but not gene mutation of connexin 32. Int. J. Cancer 1994, 56, 87–94. [Google Scholar] [CrossRef] [PubMed]
- Oyamada, M.; Krutovskikh, V.A.; Mesnil, M.; Partensky, C.; Berger, F.; Yamasaki, H. Aberrant expression of gap junction gene in primary human hepatocellular carcinomas: Increased expression of cardiac-type gap junction gene connexin 43. Mol. Carcinog. 1990, 3, 273–278. [Google Scholar] [CrossRef]
- Neveu, M.J.; Hully, J.R.; Babcock, K.L.; Hertzberg, E.L.; Nicholson, B.J.; Paul, D.L.; Pitot, H.C. Multiple mechanisms are responsible for altered expression of gap junction genes during oncogenesis in rat liver. J. Cell Sci. 1994, 107, 83–95. [Google Scholar] [CrossRef] [PubMed]
- Wilgenbus, K.K.; Kirkpatrick, C.J.; Knuechel, R.; Willecke, K.; Traub, O. Expression of Cx26, Cx32 and Cx43 gap junction proteins in normal and neoplastic human tissues. Int. J. Cancer 1992, 51, 522–529. [Google Scholar] [CrossRef] [PubMed]
- Sheen, I.S.; Jeng, K.S.; Wang, P.C.; Shih, S.C.; Chang, W.H.; Wang, H.Y.; Chen, C.C.; Shyung, L.R. Are gap junction gene connexins 26, 32 and 43 of prognostic values in hepatocellular carcinoma? A prospective study. World J. Gastroenterol. 2004, 10, 2785–2790. [Google Scholar] [CrossRef]
- Krutovskikh, V.A.; Oyamada, M.; Yamasaki, H. Sequential changes of gap-junctional intercellular communications during multistage rat liver carcinogenesis: Direct measurement of communication in vivo. Carcinogenesis 1991, 12, 1701–1706. [Google Scholar] [CrossRef]
- Fukuyama, K.; Asagiri, M.; Sugimoto, M.; Tsushima, H.; Seo, S.; Taura, K.; Uemoto, S.; Iwaisako, K. Gene expression profiles of liver cancer cell lines reveal two hepatocyte-like and fibroblast-like clusters. PLoS ONE 2021, 16, e0245939. [Google Scholar] [CrossRef] [PubMed]
- Eun, J.R.; Jung, Y.J.; Zhang, Y.; Zhang, Y.; Tschudy-Seney, B.; Ramsamooj, R.; Wan, Y.J.; Theise, N.D.; Zern, M.A.; Duan, Y. Hepatoma SK Hep-1 cells exhibit characteristics of oncogenic mesenchymal stem cells with highly metastatic capacity. PLoS ONE 2014, 9, e110744. [Google Scholar] [CrossRef]
- Nelson, L.J.; Morgan, K.; Treskes, P.; Samuel, K.; Henderson, C.J.; LeBled, C.; Homer, N.; Grant, M.H.; Hayes, P.C.; Plevris, J.N. Human hepatic HepaRG cells maintain an organotypic phenotype with high intrinsic CYP450 activity/metabolism and significantly outperform standard HepG2/C3A cells for pharmaceutical and therapeutic applications. Basic Clin. Pharmacol. 2017, 120, 30–37. [Google Scholar] [CrossRef] [PubMed]
- Caruso, S.; Calatayud, A.L.; Pilet, J.; La Bella, T.; Rekik, S.; Imbeaud, S.; Letouze, E.; Meunier, L.; Bayard, Q.; Rohr-Udilova, N.; et al. Analysis of liver cancer cell lines identifies agents with likely efficacy against hepatocellular carcinoma and markers of response. Gastroenterology 2019, 157, 760–776. [Google Scholar] [CrossRef]
- Bartlett, D.C.; Hodson, J.; Bhogal, R.H.; Youster, J.; Newsome, P.N. Combined use of N-acetylcysteine and Liberase improves the viability and metabolic function of human hepatocytes isolated from human liver. Cytotherapy 2014, 16, 800–809. [Google Scholar] [CrossRef]
- Montalbano, M.; Rastellini, C.; Wang, X.; Corsello, T.; Eltorky, M.A.; Vento, R.; Cicalese, L. Transformation of primary human hepatocytes in hepatocellular carcinoma. Int. J. Oncol. 2016, 48, 1205–1217. [Google Scholar] [CrossRef]
- Maes, M.; Decrock, E.; Cogliati, B.; Oliveira, A.G.; Marques, P.E.; Dagli, M.L.; Menezes, G.B.; Mennecier, G.; Leybaert, L.; Vanhaecke, T.; et al. Connexin and pannexin (hemi)channels in the liver. Front. Physiol. 2014, 4, 405. [Google Scholar] [CrossRef] [PubMed]
- Cooreman, A.; Van Campenhout, R.; Ballet, S.; Annaert, P.; Van Den Bossche, B.; Colle, I.; Cogliati, B.; Vinken, M. Connexin and pannexin (hemi)channels: Emerging targets in the treatment of liver disease. Hepatology 2019, 69, 1317–1323. [Google Scholar] [CrossRef]
- Rosenberg, E.; Spray, D.C.; Reid, L.M. Transcriptional and posttranscriptional control of connexin mRNAs in periportal and pericentral rat hepatocytes. Eur. J. Cell Biol. 1992, 59, 21–26. [Google Scholar]
- Berthoud, V.M.; Iwanij, V.; Garcia, A.M.; Saez, J.C. Connexins and glucagon receptors during development of rat hepatic acinus. Am. J. Physiol. 1992, 263, 650–658. [Google Scholar] [CrossRef] [PubMed]
- Greenwel, P.; Rubin, J.; Schwartz, M.; Hertzberg, E.L.; Rojkind, M. Liver fat-storing cell clones obtained from a Cci4-cirrhotic rat are heterogeneous with regard to proliferation, expression of extracellular-matrix components, interleukin-6, and connexin-43. Lab. Investig. 1993, 69, 210–216. [Google Scholar]
- Saez, J.C. Intercellular gap junctional communication is required for an optimal metabolic response of the functional units of liver. Hepatology 1997, 25, 775–776. [Google Scholar] [CrossRef] [PubMed]
- Gingalewski, C.; Wang, K.; Clemens, M.G.; De Maio, A. Posttranscriptional regulation of connexin 32 expression in liver during acute inflammation. J. Cell Physiol. 1996, 166, 461–467. [Google Scholar] [CrossRef]
- Yang, Y.; Zhu, J.; Zhang, N.; Zhao, Y.; Li, W.Y.; Zhao, F.Y.; Ou, Y.R.; Qin, S.K.; Wu, Q. Impaired gap junctions in human hepatocellular carcinoma limit intrinsic oxaliplatin chemosensitivity: A key role of connexin 26. Int. J. Oncol. 2016, 48, 703–713. [Google Scholar] [CrossRef] [PubMed]
- Kawasaki, Y.; Kubomoto, A.; Yamasaki, H. Control of intracellular localization and function of Cx43 by SEMA3F. J. Membr. Biol. 2007, 217, 53–61. [Google Scholar] [CrossRef]
- Yang, Y.; Qin, S.K.; Wu, Q.; Wang, Z.S.; Zheng, R.S.; Tong, X.H.; Liu, H.; Tao, L.; He, X.D. Connexin-dependent gap junction enhancement is involved in the synergistic effect of sorafenib and all-trans retinoic acid on HCC growth inhibition. Oncol. Rep. 2014, 31, 540–550. [Google Scholar] [CrossRef]
- Ambrosi, C.; Boassa, D.; Pranskevich, J.; Smock, A.; Oshima, A.; Xu, J.; Nicholson, B.J.; Sosinsky, G.E. Analysis of four connexin26 mutant gap junctions and hemichannels reveals variations in hexamer stability. Biophys. J. 2010, 98, 1809–1819. [Google Scholar] [CrossRef]
- Degasperi, A.; Birtwistle, M.R.; Volinsky, N.; Rauch, J.; Kolch, W.; Kholodenko, B.N. Evaluating strategies to normalise biological replicates of Western blot data. PLoS ONE 2014, 9, e87293. [Google Scholar] [CrossRef]
- Spray, D.C.; Bai, S.; Burk, R.D.; Saez, J.C. Regulation and function of liver gap junctions and their genes. Prog. Liver Dis. 1994, 12, 1–18. [Google Scholar]
- Maes, M.; McGill, M.R.; da Silva, T.C.; Abels, C.; Lebofsky, M.; Maria Monteiro de Araujo, C.; Tiburcio, T.; Veloso Alves Pereira, I.; Willebrords, J.; Crespo Yanguas, S.; et al. Involvement of connexin43 in acetaminophen-induced liver injury. Biochim. Biophys. Acta 2016, 1862, 1111–1121. [Google Scholar] [CrossRef]
- Traub, O.; Look, J.; Paul, D.; Willecke, K. Cyclic adenosine monophosphate stimulates biosynthesis and phosphorylation of the 26 kDa gap junction protein in cultured mouse hepatocytes. Eur. J. Cell Biol. 1987, 43, 48–54. [Google Scholar]
- Chu, F.F.; Doyle, D. Turnover of plasma membrane proteins in rat hepatoma cells and primary cultures of rat hepatocytes. J. Biol. Chem. 1985, 260, 3097–3107. [Google Scholar] [CrossRef]
- Traub, O.; Druge, P.M.; Willecke, K. Degradation and resynthesis of gap junction protein in plasma membranes of regenerating liver after partial hepatectomy or cholestasis. Proc. Natl. Acad. Sci. USA 1983, 80, 755–759. [Google Scholar] [CrossRef]
- Yano, T.; Hernandez-Blazquez, F.J.; Omori, Y.; Yamasaki, H. Reduction of malignant phenotype of HEPG2 cell is associated with the expression of connexin 26 but not connexin 32. Carcinogenesis 2001, 22, 1593–1600. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.; Yao, J.H.; Du, Q.Y.; Zhou, Y.C.; Yao, T.J.; Wu, Q.; Liu, J.; Ou, Y.R. Connexin 32 downregulation is critical for chemoresistance in oxaliplatin-resistant HCC cells associated with EMT. Cancer Manag. Res. 2019, 11, 5133–5146. [Google Scholar] [CrossRef]
- Ye, Z.C.; Oberheim, N.; Kettenmann, H.; Ransom, B.R. Pharmacological “cross-inhibition” of connexin hemichannels and swelling activated anion channels. Glia 2009, 57, 258–269. [Google Scholar] [CrossRef] [PubMed]
- Raju, K.L.; Augustine, D.; Rao, R.S.; SV, S.; Haragannavar, V.C.; Nambiar, S.; Prasad, K.; Awan, K.H.; Patil, S. Biomarkers in tumorigenesis using cancer cell lines: A systematic review. Asian Pac. J. Cancer Prev. 2017, 18, 2329–2337. [Google Scholar] [CrossRef]
- Schalper, K.A.; Carvajal-Hausdorf, D.; Oyarzo, M.P. Possible role of hemichannels in cancer. Front. Physiol. 2014, 5, 237. [Google Scholar] [CrossRef] [PubMed]
- Xiang, Y.; Wang, Q.; Guo, Y.; Ge, H.; Fu, Y.; Wang, X.; Tao, L. Cx32 exerts anti-apoptotic and pro-tumor effects via the epidermal growth factor receptor pathway in hepatocellular carcinoma. J. Exp. Clin. Cancer Res. 2019, 38, 145. [Google Scholar] [CrossRef]
- Neveu, M.J.; Hully, J.R.; Paul, D.L.; Pitot, H.C. Reversible alteration in the expression of the gap junctional protein connexin 32 during tumor promotion in rat liver and its role during cell proliferation. Cancer Commun. 1990, 2, 21–31. [Google Scholar] [CrossRef]
- Krutovskikh, V.A.; Mesnil, M.; Mazzoleni, G.; Yamasaki, H. Inhibition of rat liver gap junction intercellular communication by tumor-promoting agents in vivo. Association with aberrant localization of connexin proteins. Lab. Invest. 1995, 72, 571–577. [Google Scholar] [PubMed]
- Zhang, D.; Kaneda, M.; Nakahama, K.; Arii, S.; Morita, I. Connexin 43 expression promotes malignancy of HuH7 hepatocellular carcinoma cells via the inhibition of cell-cell communication. Cancer Lett. 2007, 252, 208–215. [Google Scholar] [CrossRef]
- Ogawa, K.; Pitchakarn, P.; Suzuki, S.; Chewonarin, T.; Tang, M.; Takahashi, S.; Naiki-Ito, A.; Sato, S.; Takahashi, S.; Asamoto, M.; et al. Silencing of connexin 43 suppresses invasion, migration and lung metastasis of rat hepatocellular carcinoma cells. Cancer Sci. 2012, 103, 860–867. [Google Scholar] [CrossRef]
- Elaut, G.; Henkens, T.; Papeleu, P.; Snykers, S.; Vinken, M.; Vanhaecke, T.; Rogiers, V. Molecular mechanisms underlying the dedifferentiation process of isolated hepatocytes and their cultures. Curr. Drug Metab. 2006, 7, 629–660. [Google Scholar] [CrossRef]
- Oyamada, M.; Oyamada, Y.; Takamatsu, T. Regulation of connexin expression. Biochim. Biophys. Acta 2005, 1719, 6–23. [Google Scholar] [CrossRef] [PubMed]
- Mitchell, J.A.; Lye, S.J. Differential activation of the connexin 43 promoter by dimers of activator protein-1 transcription factors in myometrial cells. Endocrinology 2005, 146, 2048–2054. [Google Scholar] [CrossRef] [PubMed]
- Shen, Y.M.; Li, Y.X.; Ma, X.F.; Wan, Q.H.; Jiang, Z.M.; Liu, Y.X.; Zhang, D.Y.; Liu, X.Z.; Wu, W.H. Connexin 43 SUMOylation improves gap junction functions between liver cancer stem cells and enhances their sensitivity to HSVtk/GCV. Int. J. Oncol. 2018, 52, 872–880. [Google Scholar] [CrossRef]
- Tork, O.M.; Khaleel, E.F.; Abdelmaqsoud, O.M. Altered cell to cell communication, autophagy and mitochondrial dysfunction in a model of hepatocellular carcinoma: Potential protective effects of curcumin and stem cell therapy. Asian Pac. J. Cancer Prev. 2015, 16, 8271–8279. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Piechocki, M.P.; Burk, R.D.; Ruch, R.J. Regulation of connexin32 and connexin43 gene expression by DNA methylation in rat liver cells. Carcinogenesis 1999, 20, 401–406. [Google Scholar] [CrossRef] [PubMed]
- Tai, Y.; Gao, J.H.; Zhao, C.; Tong, H.; Zheng, S.P.; Huang, Z.Y.; Liu, R.; Tang, C.W.; Li, J. SK-Hep1: Not hepatocellular carcinoma cells but a cell model for liver sinusoidal endothelial cells. Int. J. Clin. Exp. Pathol. 2018, 11, 2931–2938. [Google Scholar] [PubMed]
- Hermans, M.M.; Kortekaas, P.; Jongsma, H.J.; Rook, M.B. pH sensitivity of the cardiac gap junction proteins, Connexin 45 and 43. Pflugers Arch. 1995, 431, 138–140. [Google Scholar] [CrossRef] [PubMed]
- Moreno, A.P.; Saez, J.C.; Fishman, G.I.; Spray, D.C. Human Connexin43 gap junction channels—Regulation of unitary conductances by phosphorylation. Circ. Res. 1994, 74, 1050–1057. [Google Scholar] [CrossRef]
- Moreno, A.P.; Laing, J.G.; Beyer, E.C.; Spray, D.C. Properties of gap junction channels formed of connexin-45 endogenously expressed in human hepatoma (Skhep1) cells. Am. J. Physiol. Cell Physiol. 1995, 268, C356–C365. [Google Scholar] [CrossRef] [PubMed]
- Tsujiuchi, T.; Shimizu, K.; Itsuzaki, Y.; Onishi, M.; Sugata, E.; Fujii, H.; Honoki, K. CpG site hypermethylation of E-cadherin and Connexin26 genes in hepatocellular carcinomas induced by a choline-deficient L-Amino Acid-defined diet in rats. Mol. Carcinog. 2007, 46, 269–274. [Google Scholar] [CrossRef]
- Hokaiwado, N.; Asamoto, M.; Futakuchi, M.; Ogawa, K.; Takahashi, S.; Shirai, T. Both early and late stages of hepatocarcinogenesis are enhanced in cx32 dominant negative mutant transgenic rats with disrupted gap junctional intercellular communication. J. Membr. Biol. 2007, 218, 101–106. [Google Scholar] [CrossRef]
- Kato, H.; Naiki-Ito, A.; Naiki, T.; Suzuki, S.; Yamashita, Y.; Sato, S.; Sagawa, H.; Kato, A.; Kuno, T.; Takahashi, S. Connexin 32 dysfunction promotes ethanol-related hepatocarcinogenesis via activation of Dusp1-Erk axis. Oncotarget 2016, 7, 2009–2021. [Google Scholar] [CrossRef]
- Zhao, B.; Zhao, W.; Wang, Y.; Xu, Y.; Xu, J.; Tang, K.; Zhang, S.; Yin, Z.; Wu, Q.; Wang, X. Connexin32 regulates hepatoma cell metastasis and proliferation via the p53 and Akt pathways. Oncotarget 2015, 6, 10116–10133. [Google Scholar] [CrossRef]
- Shoda, T.; Mitsumori, K.; Onodera, H.; Toyoda, E.; Uneyama, C.; Takada, K.; Hirose, H. Liver tumor-promoting effect of beta-naphthoflavone, a strong CYP 1A1/2 inducer, and the relationship between CYP 1A1/2 induction and Cx32 decrease in its hepatocarcinogenesis in the rat. Toxicol. Pathol. 2000, 28, 540–547. [Google Scholar] [CrossRef]
- Shoda, T.; Mitsumori, K.; Onodera, H.; Toyoda, K.; Uneyama, C.; Imazawa, T.; Hirose, M. The relationship between decrease in Cx32 and induction of P450 isozymes in the early phase of clofibrate hepatocarcinogenesis in the rat. Arch. Toxicol. 1999, 73, 373–380. [Google Scholar] [CrossRef] [PubMed]
- Yu, M.; Zou, Q.; Wu, X.; Han, G.; Tong, X. Connexin 32 affects doxorubicin resistance in hepatocellular carcinoma cells mediated by Src/FAK signaling pathway. Biomed. Pharmacother. 2017, 95, 1844–1852. [Google Scholar] [CrossRef] [PubMed]
- Willebrords, J.; Cogliati, B.; Pereira, I.V.A.; da Silva, T.C.; Crespo Yanguas, S.; Maes, M.; Govoni, V.M.; Lima, A.; Felisbino, D.A.; Decrock, E.; et al. Inhibition of connexin hemichannels alleviates non-alcoholic steatohepatitis in mice. Sci. Rep. 2017, 7, 8268. [Google Scholar] [CrossRef] [PubMed]
- De Maio, A.; Gingalewski, C.; Theodorakis, N.G.; Clemens, M.G. Interruption of hepatic gap junctional communication in the rat during inflammation induced by bacterial lipopolysaccharide. Shock 2000, 14, 53–59. [Google Scholar] [CrossRef]
- Theodorakis, N.G.; De Maio, A. Cx32 mRNA in rat liver: Effects of inflammation on poly(A) tail distribution and mRNA degradation. Am. J. Physiol. 1999, 276, 1249–1257. [Google Scholar] [CrossRef]
- Crespo Yanguas, S.; Willebrords, J.; Maes, M.; da Silva, T.C.; Veloso Alves Pereira, I.; Cogliati, B.; Zaidan Dagli, M.L.; Vinken, M. Connexins and pannexins in liver damage. EXCLI J. 2016, 15, 177–186. [Google Scholar] [CrossRef]
- Van Rijen, H.V.; van Veen, T.A.; Hermans, M.M.; Jongsma, H.J. Human connexin40 gap junction channels are modulated by cAMP. Cardiovasc. Res. 2000, 45, 941–951. [Google Scholar] [CrossRef]
- Pfaffl, M.W. A new mathematical model for relative quantification in real-time RT-PCR. Nucleic Acids Res. 2001, 29, e45. [Google Scholar] [CrossRef]
- Cooreman, A.; Van Campenhout, R.; Crespo Yanguas, S.; Gijbels, E.; Leroy, K.; Pieters, A.; Tabernilla, A.; Van Brantegem, P.; Annaert, P.; Cogliati, B.; et al. Cholestasis differentially affects liver connexins. Int. J. Mol. Sci. 2020, 21, 6534. [Google Scholar] [CrossRef]



| Cell Line | Morphology | Disease | Age | Gender | Ethnicity | HBV | Medium |
|---|---|---|---|---|---|---|---|
| SK-HEP-1 | Epithelial | Adenocarcinoma | 52 | Male | Caucasian | Not detected | EMEM + FBS |
| C3A | Epithelial | HCC | 15 | Male | Caucasian | Not detected | EMEM + FBS |
| SNU-449 | Epithelial | Grade II-III/IV, HCC | 52 | Male | Asian | DNA detected | RPMI-1640 + heat-inactivated FBS |
| SNU-423 | Epithelial | Grade III/IV, pleomorphic HCC | 40 | Male | Asian | DNA detected | RPMI-1640 + heat-inactivated FBS |
| SNU-387 | Epithelial | Grade IV/V, pleomorphic HCC | 41 | Female | Asian | DNA detected | RPMI-1640 + FBS |
| SNU-475 | Epithelial | Grade II-IV/V, HCC | 43 | Male | Asian | DNA detected | RPMI-1640 + heat-inactivated FBS |
| PLC/PRF/5 | Epithelial | HCC | 24 | Male | African | Secreted HbsAg | EMEM + FBS |
| PHH | Epithelial | Liver metastasis of colorectal adenocarcinoma | 74 | Female | Caucasian | Not detected | CHRM + CHPM |
| Gene Symbol | Assay ID | Accession Number | Assay Location | Amplicon Size (Base Pairs) | Exon Boundary |
|---|---|---|---|---|---|
| GJA1 (Cx43) | Hs00748445-s1 | NM_000165.4 | 1031 | 142 | 2 |
| GJB1 (Cx32) | Hs00939759-s1 | NM_000166.5 NM_001097642.2 | 1547 1496 | 63 | 2 |
| GJB2 (Cx26) | Hs00269615-s1 | NM_004004.5 | 715 | 123 | 2 |
| ACTB | Hs01060665-g1 | NM_001101.3 | 208 | 63 | 2–3 |
| B2M | Hs00187842-m1 | NM_004048.2 | 134 | 64 | 1–2 |
| GAPDH | Hs02786624-g1 | NM_001256799.2 NM_001289745.1 NM_001289746.1 NM_002046.5 | 870 928 822 836 | 157 | 7 8 7 8 |
| HMBS | Hs00609296-g1 | NM_000190.3 NM_001024382.1 NM_001258208.1 NM_001258209.1 | 1070 972 950 1041 | 69 | 13–14 13–14 12–13 12–13 |
| UBC | Hs01871556-s1 | M26880.1 | 2173 | 135 | – |
| Target | Product Code | Dilution Primary Antibody | Supplier | |
|---|---|---|---|---|
| Immunoblot | Immunocytochemistry | |||
| Cx26 | 51–2800 | 1:250 | 1:250 | Invitrogen, Waltham, MA, USA |
| Cx32 | C3470 | 1:600 | 1:500 | Sigma-Aldrich, St. Louis, MO, USA |
| Cx43 | C6219 | 1:4000 | 1:1000 | Sigma-Aldrich, St. Louis, MO, USA |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Leroy, K.; Silva Costa, C.J.; Pieters, A.; dos Santos Rodrigues, B.; Van Campenhout, R.; Cooreman, A.; Tabernilla, A.; Cogliati, B.; Vinken, M. Expression and Functionality of Connexin-Based Channels in Human Liver Cancer Cell Lines. Int. J. Mol. Sci. 2021, 22, 12187. https://doi.org/10.3390/ijms222212187
Leroy K, Silva Costa CJ, Pieters A, dos Santos Rodrigues B, Van Campenhout R, Cooreman A, Tabernilla A, Cogliati B, Vinken M. Expression and Functionality of Connexin-Based Channels in Human Liver Cancer Cell Lines. International Journal of Molecular Sciences. 2021; 22(22):12187. https://doi.org/10.3390/ijms222212187
Chicago/Turabian StyleLeroy, Kaat, Cícero Júlio Silva Costa, Alanah Pieters, Bruna dos Santos Rodrigues, Raf Van Campenhout, Axelle Cooreman, Andrés Tabernilla, Bruno Cogliati, and Mathieu Vinken. 2021. "Expression and Functionality of Connexin-Based Channels in Human Liver Cancer Cell Lines" International Journal of Molecular Sciences 22, no. 22: 12187. https://doi.org/10.3390/ijms222212187
APA StyleLeroy, K., Silva Costa, C. J., Pieters, A., dos Santos Rodrigues, B., Van Campenhout, R., Cooreman, A., Tabernilla, A., Cogliati, B., & Vinken, M. (2021). Expression and Functionality of Connexin-Based Channels in Human Liver Cancer Cell Lines. International Journal of Molecular Sciences, 22(22), 12187. https://doi.org/10.3390/ijms222212187







