MiR-106b-5p: A Master Regulator of Potential Biomarkers for Breast Cancer Aggressiveness and Prognosis
Abstract
:1. Introduction
2. Results
2.1. BCa Modulates miRNA Expression in 4T1 Allografts and Human Tissue Samples
2.2. Hsa-miR-21-5p and miR-106b-5p Share Several Target Genes
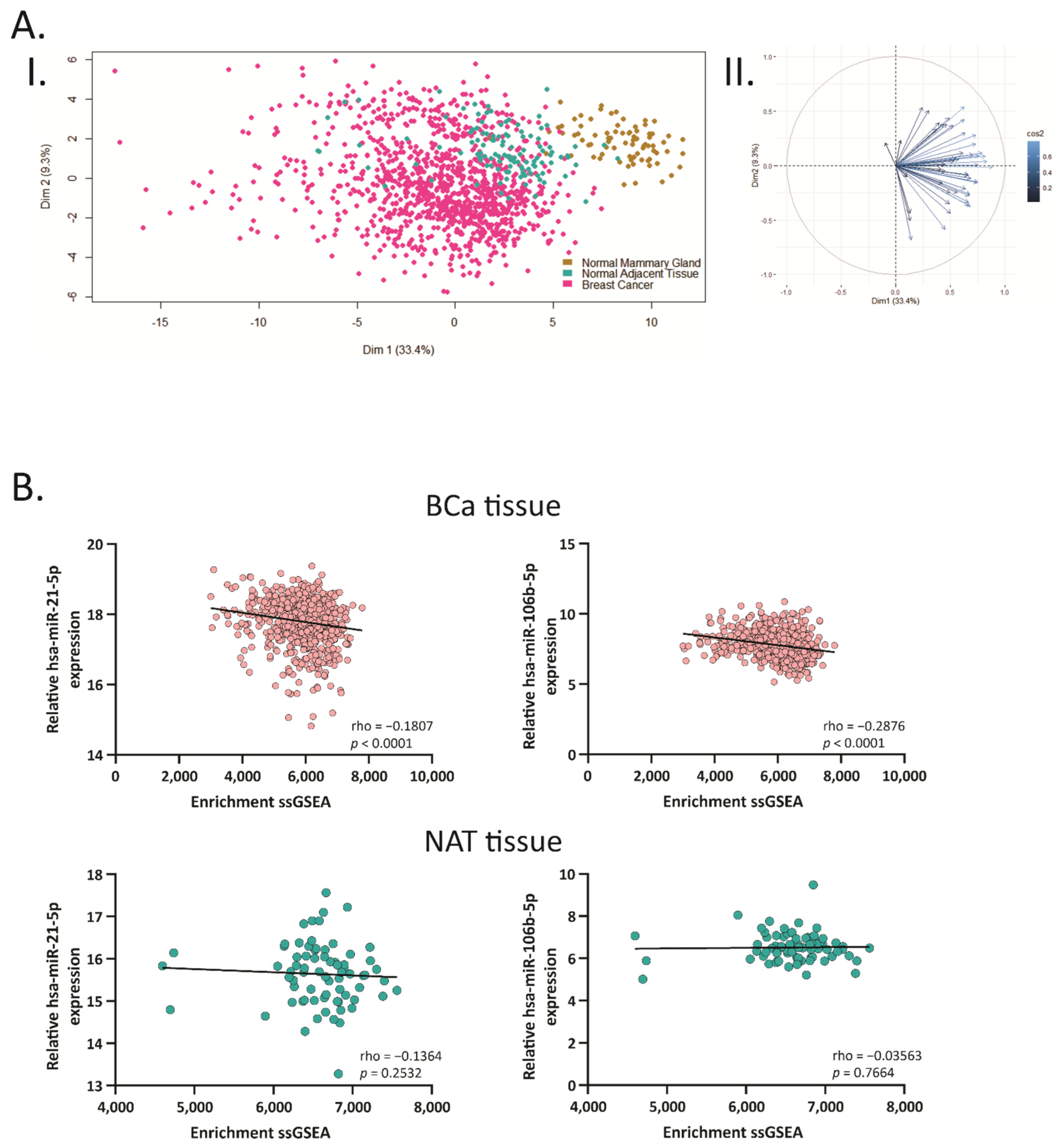
2.3. Hsa-miR-21-5p and miR-106b-5p Negatively Correlate with Their Target Genes
2.4. Hsa-miR-21-5p and miR-106b-5p Modulate Cancer and Metabolic Related Pathways
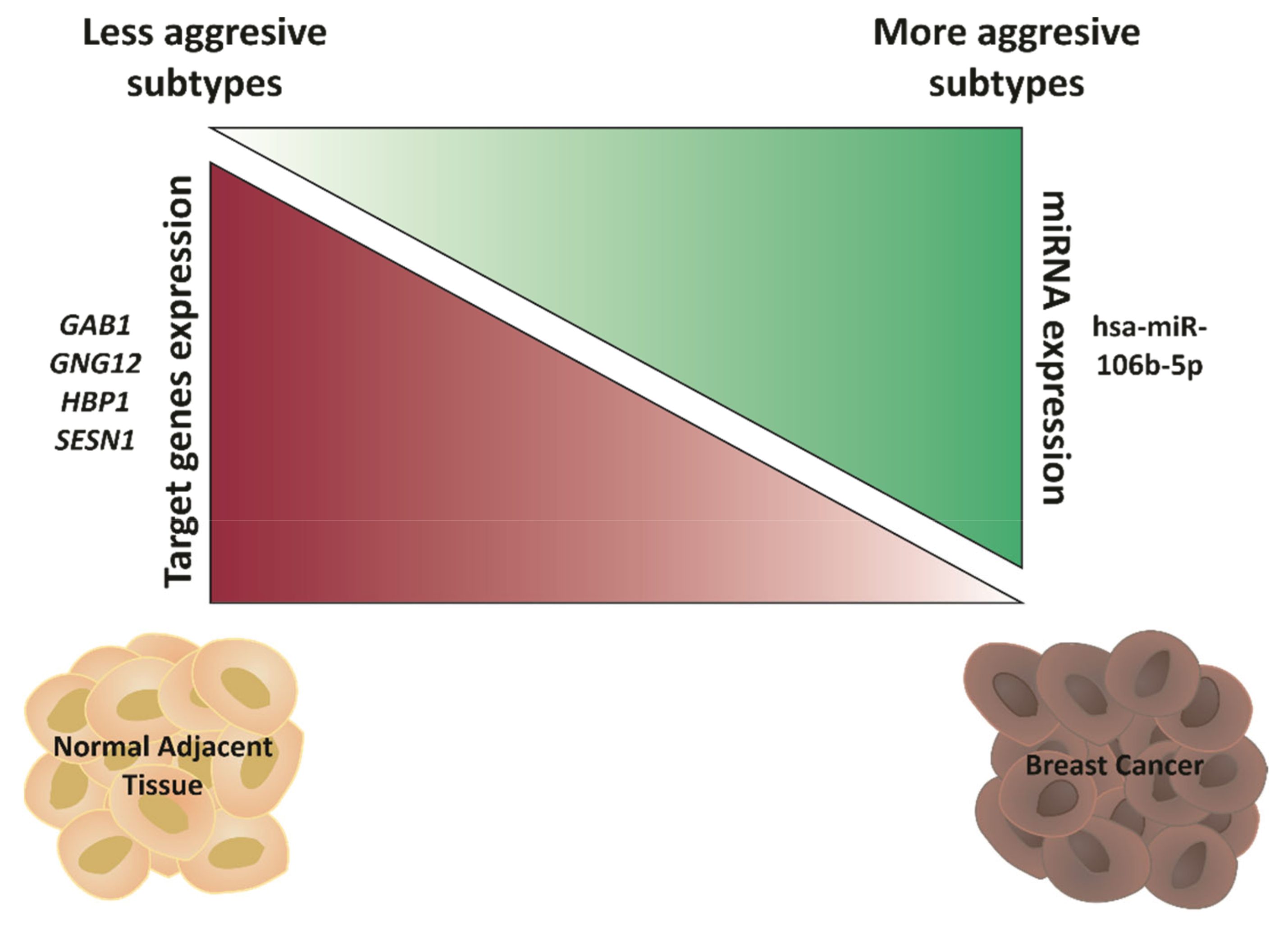
2.5. Hsa-miR-106b-5p and miR-21-5p Are Upregulated in More Aggressive BCa Subtypes and Are Predictors of Worse Overall Survival
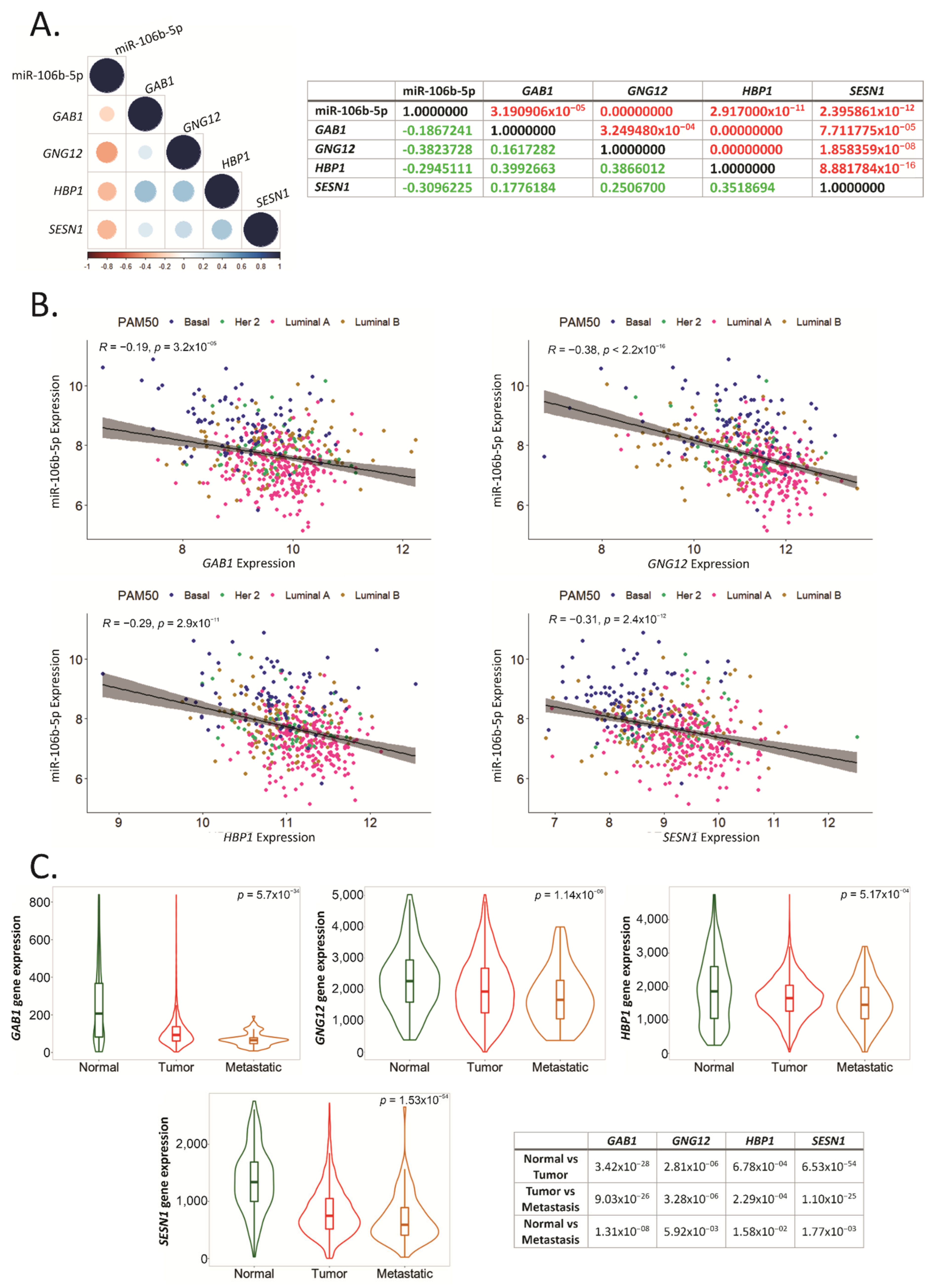
2.6. GAB1, GNG12, HBP1 and SESN1 Are Downregulated in More Aggressive BCa Subtypes and Could Be Used as Prognosis Biomarkers
2.7. GAB1, GNG12, HBP1 and SESN1 Negatively Correlate with hsa-miR-106b-5p in BCa Tissues
3. Discussion
4. Materials and Methods
4.1. Cell Culture
4.2. BCa Allograft Murine Model
4.3. RNA Isolation and RT-qPCR Analysis
4.4. TCGA Dataset Analysis
4.5. Functional Enrichment Analysis
4.6. Principal Component Analysis (PCA)
4.7. Single-Sample Gene-Set Enrichment Analysis
4.8. Correlation Matrix
4.9. Kaplan–Meier Plots
4.10. Normal, Tumor and Metastatic Gene Expression
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Bray, F.; Ferlay, J.; Soerjomataram, I.; Siegel, R.L.; Torre, L.A.; Jemal, A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 2018, 68, 394–424. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Trayes, K.P.; Cokenakes, S.E.H. Breast Cancer Treatment. Am. Fam. Physician 2021, 104, 171–178. [Google Scholar] [PubMed]
- Hortobagyi, G.N.; Edge, S.B.; Giuliano, A. New and Important Changes in the TNM Staging System for Breast Cancer. Am. Soc. Clin. Oncol. Educ. Book 2018, 457–467. [Google Scholar] [CrossRef] [PubMed]
- Yanovich, G.; Agmon, H.; Harel, M.; Sonnenblick, A.; Peretz, T.; Geiger, T. Clinical proteomics of breast cancer reveals a novel layer of breast cancer classification. Cancer Res. 2018, 78, 6001–6010. [Google Scholar] [CrossRef] [Green Version]
- Yin, L.; Duan, J.J.; Bian, X.W.; Yu, S.C. Triple-negative breast cancer molecular subtyping and treatment progress. Breast Cancer Res. 2020, 22, 1–13. [Google Scholar] [CrossRef]
- Ha, M.; Kim, V.N. Regulation of microRNA biogenesis. Nat. Rev. Mol. Cell Biol. 2014, 15, 509–524. [Google Scholar] [CrossRef]
- Fridrichova, I.; Zmetakova, I. MicroRNAs Contribute to Breast Cancer Invasiveness. Cells 2019, 8, 1361. [Google Scholar] [CrossRef] [Green Version]
- Loh, H.-Y.; Norman, B.P.; Lai, K.-S.; Rahman, N.M.A.N.A.; Alitheen, N.B.M.; Osman, M.A. The Regulatory Role of MicroRNAs in Breast Cancer. Int. J. Mol. Sci. 2019, 20, 4940. [Google Scholar] [CrossRef] [Green Version]
- Ruan, K.; Fang, X.; Ouyang, G. MicroRNAs: Novel regulators in the hallmarks of human cancer. Cancer Lett. 2009, 285, 116–126. [Google Scholar] [CrossRef]
- Bertoli, G.; Cava, C.; Castiglioni, I. Micrornas: New biomarkers for diagnosis, prognosis, therapy prediction and therapeutic tools for breast cancer. Theranostics 2015, 5, 1122–1143. [Google Scholar] [CrossRef]
- Kim, S.Y.; Kim, A.Y.; Lee, H.W.; Son, Y.H.; Lee, G.Y.; Lee, J.-W.; Lee, Y.S.; Kim, J.B. miR-27a is a negative regulator of adipocyte differentiation via suppressing PPARγ expression. Biochem. Biophys. Res. Commun. 2010, 392, 323–328. [Google Scholar] [CrossRef]
- Alarmo, E.L.; Havunen, R.; Häyrynen, S.; Penkki, S.; Ketolainen, J.; Nykter, M.; Kallioniemi, A. Bone morphogenetic protein 4 regulates microRNA expression in breast cancer cell lines in diverse fashion. Genes Chromosom. Cancer 2016, 55, 227–236. [Google Scholar] [CrossRef]
- Kim, Y.C.; Cutler, M.L. Microrna-dependent targeting of rsu1 and the ipp adhesion complex regulates the PTEN/PI3K/AKT signaling pathway in breast cancer cell lines. Int. J. Mol. Sci. 2020, 21, 5458. [Google Scholar] [CrossRef]
- Nurzadeh, M.; Naemi, M.; Sheikh Hasani, S. A comprehensive review on oncogenic miRNAs in breast cancer. J. Genet. 2021, 100. [Google Scholar] [CrossRef]
- Hong, H.C.; Chuang, C.H.; Huang, W.C.; Weng, S.L.; Chen, C.H.; Chang, K.H.; Liao, K.W.; Huang, H. Da A panel of eight microRNAs is a good predictive parameter for triple-negative breast cancer relapse. Theranostics 2020, 10, 8771–8789. [Google Scholar] [CrossRef]
- Goldman, M.J.; Craft, B.; Hastie, M.; Repečka, K.; McDade, F.; Kamath, A.; Banerjee, A.; Luo, Y.; Rogers, D.; Brooks, A.N.; et al. Visualizing and interpreting cancer genomics data via the Xena platform. Nat. Biotechnol. 2020, 38, 675–678. [Google Scholar] [CrossRef]
- Liu, M.; Zhou, S.; Wang, J.; Zhang, Q.; Yang, S.; Feng, J.; Xu, B.; Zhong, S. Identification of genes associated with survival of breast cancer patients. Breast Cancer 2019, 26, 317–325. [Google Scholar] [CrossRef]
- Chang, J.T.-H.; Wang, F.; Chapin, W.; Huang, R.S. Identification of MicroRNAs as breast cancer prognosis markers through the cancer genome atlas. PLoS ONE 2016, 11, e0168284. [Google Scholar] [CrossRef] [Green Version]
- Lu, J.; Tan, T.; Zhu, L.; Dong, H.; Xian, R. Hypomethylation Causes MIR21 Overexpression in Tumors. Mol. Ther.-Oncolytics 2020, 18, 47–57. [Google Scholar] [CrossRef]
- Qian, B.; Katsaros, D.; Lu, L.; Preti, M.; Durando, A.; Arisio, R.; Mu, L.; Yu, H. High miR-21 expression in breast cancer associated with poor disease-free survival in early stage disease and high TGF-β1. Breast Cancer Res. Treat. 2009, 117, 131–140. [Google Scholar] [CrossRef]
- Haakensen, V.D.; Nygaard, V.; Greger, L.; Aure, M.R.; Fromm, B.; Bukholm, I.R.K.; Lüders, T.; Chin, S.F.; Git, A.; Caldas, C.; et al. Subtype-specific micro-RNA expression signatures in breast cancer progression. Int. J. Cancer 2016, 139, 1117–1128. [Google Scholar] [CrossRef]
- Yu, X.; Liang, J.; Xu, J.; Li, X.; Xing, S.; Li, H.; Liu, W.; Liu, D.; Xu, J.; Huang, L.; et al. Identification and validation of circulating microRNA signatures for breascancer early detection based on large scale tissue-derived data. J. Breast Cancer 2018, 21, 363–370. [Google Scholar] [CrossRef]
- Pourteimoor, V.; Paryan, M.; Mohammadi-Yeganeh, S. microRNA as a systemic intervention in the specific breast cancer subtypes with C-MYC impacts; introducing subtype-based appraisal tool. J. Cell. Physiol. 2018, 233, 5655–5669. [Google Scholar] [CrossRef]
- Mango, V.L.; Al-Khawari, H.; Dershaw, D.D.; Ashkanani, M.H.; Pennisi, B.; Turner, P.; Thornton, C.; Morris, E.A. Initiating a National Mammographic Screening Program: The Kuwait Experience Training with a US Cancer Center. J. Am. Coll. Radiol. 2019, 16, 202–207. [Google Scholar] [CrossRef]
- Wang, H.; Tan, Z.; Hu, H.; Liu, H.; Wu, T.; Zheng, C.; Wang, X.; Luo, Z.; Wang, J.; Liu, S.; et al. MicroRNA-21 promotes breast cancer proliferation and metastasis by targeting LZTFL1. BMC Cancer 2019, 19. [Google Scholar] [CrossRef] [Green Version]
- Newie, I.; Søkilde, R.; Persson, H.; Jacomasso, T.; Gorbatenko, A.; Borg, Å.; De Hoon, M.; Pedersen, S.F.; Rovira, C. HER2-encoded mir-4728 forms a receptor-independent circuit with miR-21-5p through the non-canonical poly(A) polymerase PAPD5. Sci. Rep. 2016, 6. [Google Scholar] [CrossRef] [Green Version]
- Calvano Filho, C.M.C.; Calvano-Mendes, D.C.; Carvalho, K.C.; Maciel, G.A.; Ricci, M.D.; Torres, A.P.; Filassi, J.R.; Baracat, E.C. Triple-negative and luminal A breast tumors: Differential expression of miR-18a-5p, miR-17-5p, and miR-20a-5p. Tumor Biol. 2014, 35, 7733–7741. [Google Scholar] [CrossRef]
- Moi, L.; Braaten, T.; Al-Shibli, K.; Lund, E.; Busund, L.T.R. Differential expression of the miR-17-92 cluster and miR-17 family in breast cancer according to tumor type; Results from the Norwegian Women and Cancer (NOWAC) study. J. Transl. Med. 2019, 17, 1–20. [Google Scholar] [CrossRef] [Green Version]
- De Rinaldis, E.; Gazinska, P.; Mera, A.; Modrusan, Z.; Fedorowicz, G.M.; Burford, B.; Gillett, C.; Marra, P.; Grigoriadis, A.; Dornan, D.; et al. Integrated genomic analysis of triple-negative breast cancers reveals novel microRNAs associated with clinical and molecular phenotypes and sheds light on the pathways they control. BMC Genom. 2013, 14, 1. [Google Scholar] [CrossRef] [Green Version]
- Zheng, R.; Pan, L.; Gao, J.; Ye, X.; Chen, L.; Zhang, X.; Tang, W.; Zheng, W. Prognostic value of miR-106b expression in breast cancer patients. J. Surg. Res. 2015, 195, 158–165. [Google Scholar] [CrossRef]
- Guarnieri, A.L.; Towers, C.G.; Drasin, D.J.; Oliphant, M.U.J.; Andrysik, Z.; Hotz, T.J.; Vartuli, R.L.; Linklater, E.S.; Pandey, A.; Khanal, S.; et al. The miR-106b-25 cluster mediates breast tumor initiation through activation of NOTCH1 via direct repression of NEDD4L. Oncogene 2018, 37, 3879–3893. [Google Scholar] [CrossRef] [PubMed]
- Tan, W.; Liang, G.; Xie, X.; Jiang, W.; Tan, L.; Sanders, A.J.; Liu, Z.; Ling, Y.; Zhong, W.; Tian, Z.; et al. Incorporating MicroRNA into Molecular Phenotypes of Circulating Tumor Cells Enhances the Prognostic Accuracy for Patients with Metastatic Breast Cancer. Oncologist 2019, 24. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chen, K.-H.E.; Bustamante, K.; Nguyen, V.; Walker, A.M. Involvement of miR-106b in tumorigenic actions of both prolactin and estradiol. Oncotarget 2017, 8, 36368–36382. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, Z.; Li, T.E.; Chen, M.; Pan, J.J.; Shen, K.W. miR-106b-5p contributes to the lung metastasis of breast cancer via targeting CNN1 and regulating Rho/ROCK1 pathway. Aging 2020, 12, 1867–1887. [Google Scholar] [CrossRef] [PubMed]
- Gong, C.; Qu, S.; Liu, B.; Pan, S.; Jiao, Y.; Nie, Y.; Su, F.; Liu, Q.; Song, E. MiR-106b expression determines the proliferation paradox of TGF-β in breast cancer cells. Oncogene 2015, 34, 84–93. [Google Scholar] [CrossRef] [PubMed]
- Schrijver, W.A.M.E.; van Diest, P.J.; Moelans, C.B. Unravelling site-specific breast cancer metastasis: A microRNA expression profiling study. Oncotarget 2017, 8, 3111–3123. [Google Scholar] [CrossRef] [PubMed]
- Lee, C.M.; Chung, I.Y.; Park, Y.; Yun, K.W.; Jo, H.G.; Park, H.J.; Lee, H.J.; Lee, S.B.; Kim, H.J.; Ko, B.S.; et al. The impact of androgen receptor and histone deacetylase 1 expression on the prognosis of ductal carcinoma in situ. J. Breast Cancer 2020, 23, 610–621. [Google Scholar] [CrossRef]
- Sanli, T.; Linher-Melville, K.; Tsakiridis, T.; Singh, G. Sestrin2 modulates AMPK subunit expression and its response to ionizing radiation in breast cancer cells. PLoS ONE 2012, 7. [Google Scholar] [CrossRef]
- Cordani, M.; Butera, G.; Dando, I.; Torrens-Mas, M.; Butturini, E.; Pacchiana, R.; Oppici, E.; Cavallini, C.; Gasperini, S.; Tamassia, N.; et al. Mutant p53 blocks SESN1/AMPK/PGC-1α/UCP2 axis increasing mitochondrial O2−· production in cancer cells. Br. J. Cancer 2018, 119, 994–1008. [Google Scholar] [CrossRef]
- Cordani, M.; Oppici, E.; Dando, I.; Butturini, E.; Dalla Pozza, E.; Nadal-Serrano, M.; Oliver, J.; Roca, P.; Mariotto, S.; Cellini, B.; et al. Mutant p53 proteins counteract autophagic mechanism sensitizing cancer cells to mTOR inhibition. Mol. Oncol. 2016, 10, 1008–1029. [Google Scholar] [CrossRef] [Green Version]
- Veeraraghavan, J.; Tan, Y.; Cao, X.X.; Kim, J.A.; Wang, X.; Chamness, G.C.; Maiti, S.N.; Cooper, L.J.N.; Edwards, D.P.; Contreras, A.; et al. Recurrent ESR1-CCDC170 rearrangements in an aggressive subset of oestrogen receptor-positive breast cancers. Nat. Commun. 2014, 5. [Google Scholar] [CrossRef] [Green Version]
- Ortiz-Padilla, C.; Gallego-Ortega, D.; Browne, B.C.; Hochgräfe, F.; Caldon, C.E.; Lyons, R.J.; Croucher, D.R.; Rickwood, D.; Ormandy, C.J.; Brummer, T.; et al. Functional characterization of cancer-associated Gab1 mutations. Oncogene 2013, 32, 2696–2702. [Google Scholar] [CrossRef] [Green Version]
- Mouradian, M.; Kikawa, K.D.; Johnson, E.D.; Beck, K.L.; Pardini, R.S. Key roles for GRB2-associated-binding protein 1, phosphatidylinositol-3-kinase, cyclooxygenase 2, prostaglandin E2 and transforming growth factor alpha in linoleic acid-induced upregulation of lung and breast cancer cell growth. Prostaglandins Leukot. Essent. Fat. Acids 2014, 90, 105–115. [Google Scholar] [CrossRef] [Green Version]
- Gensler, M.; Buschbeck, M.; Ullrich, A. Negative Regulation of HER2 Signaling by the PEST-type Protein-tyrosine Phosphatase BDP1. J. Biol. Chem. 2004, 279, 12110–12116. [Google Scholar] [CrossRef] [Green Version]
- Wang, X.; Peng, J.; Yang, Z.; Zhou, P.J.; An, N.; Wei, L.; Zhu, H.H.; Lu, J.; Fang, Y.X.; Gao, W.Q. Elevated expression of Gab1 promotes breast cancer metastasis by dissociating the PAR complex. J. Exp. Clin. Cancer Res. 2019, 38. [Google Scholar] [CrossRef] [Green Version]
- Escamilla-Powers, J.R.; Daniel, C.J.; Farrell, A.; Taylor, K.; Zhang, X.; Byers, S.; Sears, R. The tumor suppressor protein HBP1 is a novel c-Myc-binding protein that negatively regulates c-Myc transcriptional activity. J. Biol. Chem. 2010, 285, 4847–4858. [Google Scholar] [CrossRef] [Green Version]
- Paulson, K.E.; Rieger-Christ, K.; McDevitt, M.A.; Kuperwasser, C.; Kim, J.; Unanue, V.E.; Zhang, X.; Hu, M.; Ruthazer, R.; Berasi, S.P.; et al. Alterations of the HBP1 transcriptional repressor are associated with invasive breast cancer. Cancer Res. 2007, 67, 6136–6145. [Google Scholar] [CrossRef] [Green Version]
- Kim, J.; Zhang, X.; Rieger-Christ, K.M.; Summerhayes, I.C.; Wazer, D.E.; Paulson, K.E.; Yee, A.S. Suppression of Wnt signaling by the green tea compound (-)-epigallocatechin 3-gallate (EGCG) in invasive breast cancer cells: Requirement of the transcriptional repressor HBP1. J. Biol. Chem. 2006, 281, 10865–10875. [Google Scholar] [CrossRef] [Green Version]
- Li, H.; Bian, C.; Liao, L.; Li, J.; Zhao, R.C. miR-17-5p promotes human breast cancer cell migration and invasion through suppression of HBP1. Breast Cancer Res. Treat. 2011, 126, 565–575. [Google Scholar] [CrossRef]
- Bollaert, E.; de Rocca Serra, A.; Demoulin, J.B. The HMG box transcription factor HBP1: A cell cycle inhibitor at the crossroads of cancer signaling pathways. Cell. Mol. Life Sci. 2019, 76, 1529–1539. [Google Scholar] [CrossRef]
- Farré, P.L.; Scalise, G.D.; Duca, R.B.; Dalton, G.N.; Massillo, C.; Porretti, J.; Graña, K.; Gardner, K.; De Luca, P.; De Siervi, A. CTBP1 and metabolic syndrome induce an mRNA and miRNA expression profile critical for breast cancer progression and metastasis. Oncotarget 2018, 9, 13848–13858. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dalton, G.N.; Massillo, C.; Scalise, G.D.; Duca, R.; Porretti, J.; Farré, P.L.; Gardner, K.; Paez, A.; Gueron, G.; De Luca, P.; et al. CTBP1 depletion on prostate tumors deregulates miRNA/mRNA expression and impairs cancer progression in metabolic syndrome mice. Cell Death Dis. 2019, 10, 299. [Google Scholar] [CrossRef] [Green Version]
- Porretti, J.; Dalton, G.N.; Massillo, C.; Scalise, G.D.; Farré, P.L.; Elble, R.; Gerez, E.N.; Accialini, P.; Cabanillas, A.M.; Gardner, K.; et al. CLCA2 epigenetic regulation by CTBP1, HDACs, ZEB1, EP300 and miR-196b-5p impacts prostate cancer cell adhesion and EMT in metabolic syndrome disease. Int. J. Cancer 2018, 143, 897–906. [Google Scholar] [CrossRef] [Green Version]
- Chen, C.; Ridzon, D.A.; Broomer, A.J.; Zhou, Z.; Lee, D.H.; Nguyen, J.T.; Barbisin, M.; Xu, N.L.; Mahuvakar, V.R.; Andersen, M.R.; et al. Real-time quantification of microRNAs by stem-loop RT-PCR. Nucleic Acids Res. 2005, 33, e179. [Google Scholar] [CrossRef]
- Karagkouni, D.; Paraskevopoulou, M.D.; Chatzopoulos, S.; Vlachos, I.S.; Tastsoglou, S.; Kanellos, I.; Papadimitriou, D.; Kavakiotis, I.; Maniou, S.; Skoufos, G.; et al. DIANA-TarBase v8: A decade-long collection of experimentally supported miRNA-gene interactions. Nucleic Acids Res. 2018, 46, D239–D245. [Google Scholar] [CrossRef] [Green Version]
- Lánczky, A.; Nagy, Á.; Bottai, G.; Munkácsy, G.; Szabó, A.; Santarpia, L.; Győrffy, B. miRpower: A web-tool to validate survival-associated miRNAs utilizing expression data from 2178 breast cancer patients. Breast Cancer Res. Treat. 2016, 160, 439–446. [Google Scholar] [CrossRef]
- Győrffy, B. Survival analysis across the entire transcriptome identifies biomarkers with the highest prognostic power in breast cancer. Comput. Struct. Biotechnol. J. 2021, 19, 4101–4109. [Google Scholar] [CrossRef]
- Bartha, Á.; Győrffy, B. TNMplot.com: A Web Tool for the Comparison of Gene Expression in Normal, Tumor and Metastatic Tissues. Int. J. Mol. Sci. 2021, 22, 2622. [Google Scholar] [CrossRef]





| Primer | Sequence (5′–3′) | Tann (°C) |
|---|---|---|
| RT-Stem-loop-Rv | TGGTGCAGGGTCCGAGGTATT | – |
| RT-mmu-miR-21a-5p-STEM | GTCTCCTCTGGTGCAGGGTCCGAGGTATTCGCACCAGAGGAGACTCAACA | – |
| RT-mmu-miR-21a-5p Fw | CGGGGGGTAGCTTATCAGACTG | 65 |
| RT-mmu-miR-106b-5p-STEM | GTCTCCTCTGGTGCAGGGTCCGAGGTATTCGCACCAGAGGAGACATCTGC | – |
| RT-mmu-miR-106b-5p Fw | GCGGCGGTAAAGTGCTGACAG | 67 |
| RT-mmu-miR-125b-5p-STEM | GTCTCCTCTGGTGCAGGGTCCGAGGTATTCGCACCAGAGGAGACTCACAA | – |
| RT-mmu-miR-125b-5p Fw | CCGCCTCCCTGAGACCCTAAC | 65 |
| RT-mmu-miR-221-3p-STEM | GTCTCCTCTGGTGCAGGGTCCGAGGTATTCGCACCAGAGGAGACGAAACC | – |
| RT-mmu-miR-221-3p Fw | GGCGGAGCTACATTGTCTGCTG | 65 |
| RT-mmu-miR-138-5p-STEM | GTCTCCTCTGGTGCAGGGTCCGAGGTATTCGCACCAGAGGAGACCGGCCT | – |
| RT-mmu-miR-138-5p Fw | GGCGGAGCTGGTGTTGTGAATC | 67 |
| RT-mmu-miR-143-3p-STEM | GTCTCCTCTGGTGCAGGGTCCGAGGTATTCGCACCAGAGGAGACGAGCTA | – |
| RT-mmu-miR-143-3p Fw | GGGCGGTGAGATGAAGCACTG | 67 |
| RT-mmu-miR-146a-5p-STEM | GTCTCCTCTGGTGCAGGGTCCGAGGTATTCGCACCAGAGGAGACAACCCA | – |
| RT-mmu-miR-146a-5p Fw | CGGGCGGTGAGAACTGAATTCC | 65 |
| RT-mmu-miR-205-5p-STEM | GTCTCCTCTGGTGCAGGGTCCGAGGTATTCGCACCAGAGGAGACCAGACT | – |
| RT-mmu-miR-205-5p Fw | CGCGTCCTTCATTCCACCGG | 65 |
| RT-mmu-miR-191-5p-STEM | GTCTCCTCTGGTGCAGGGTCCGAGGTATTCGCACCAGAGGAGACCAGCTG | – |
| RT-mmu-miR-191-5p Fw | GCGGCAACGGAATCCCAAAAG | 70 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Farré, P.L.; Duca, R.B.; Massillo, C.; Dalton, G.N.; Graña, K.D.; Gardner, K.; Lacunza, E.; De Siervi, A. MiR-106b-5p: A Master Regulator of Potential Biomarkers for Breast Cancer Aggressiveness and Prognosis. Int. J. Mol. Sci. 2021, 22, 11135. https://doi.org/10.3390/ijms222011135
Farré PL, Duca RB, Massillo C, Dalton GN, Graña KD, Gardner K, Lacunza E, De Siervi A. MiR-106b-5p: A Master Regulator of Potential Biomarkers for Breast Cancer Aggressiveness and Prognosis. International Journal of Molecular Sciences. 2021; 22(20):11135. https://doi.org/10.3390/ijms222011135
Chicago/Turabian StyleFarré, Paula Lucía, Rocío Belén Duca, Cintia Massillo, Guillermo Nicolás Dalton, Karen Daniela Graña, Kevin Gardner, Ezequiel Lacunza, and Adriana De Siervi. 2021. "MiR-106b-5p: A Master Regulator of Potential Biomarkers for Breast Cancer Aggressiveness and Prognosis" International Journal of Molecular Sciences 22, no. 20: 11135. https://doi.org/10.3390/ijms222011135
APA StyleFarré, P. L., Duca, R. B., Massillo, C., Dalton, G. N., Graña, K. D., Gardner, K., Lacunza, E., & De Siervi, A. (2021). MiR-106b-5p: A Master Regulator of Potential Biomarkers for Breast Cancer Aggressiveness and Prognosis. International Journal of Molecular Sciences, 22(20), 11135. https://doi.org/10.3390/ijms222011135






