Abstract
Mitochondrial dysfunction and stem cell exhaustion are two hallmarks of aging. In the hematopoietic system, aging is linked to imbalanced immune response and reduced regenerative capacity in hematopoietic stem cells (HSCs), as well as an increased predisposition to a spectrum of diseases, including myelodysplastic syndrome and acute myeloid leukemia. Myeloid-biased differentiation and loss of polarity are distinct features of aged HSCs, which generally exhibit enhanced mitochondrial oxidative phosphorylation and increased production of reactive oxygen species (ROS), suggesting a direct role for mitochondria in the degenerative process. Here, we provide an overview of current knowledge of the mitochondrial mechanisms that contribute to age-related phenotypes in HSCs. These include mitochondrial ROS production, alteration/activation of mitochondrial metabolism, the quality control pathway of mitochondria, and inflammation. Greater understanding of the key machineries of HSC aging will allow us to identify new therapeutic targets for preventing, delaying, or even reversing aspects of this process.
1. Introduction
Aging is a time-dependent degenerative process that affects all living organisms. Since the aging population is inexorably growing, there is an imminent need to develop new therapeutic strategies for ameliorating the age-related changes and/or disorders, and first among these is hematopoietic aging. The milestone review paper of Reference [1] has categorized the cellular and molecular hallmarks of this type of aging, which include both mitochondrial dysfunction and stem cell exhaustion, the two main topics of this review.
Mitochondria were described as contributing to aging and degeneration as early as already in the 1920s, when the “rate of living hypothesis” proposed that metabolic rates inversely correlate with organismal lifespan [2]. Many researchers have since described the links between mitochondrial biology and aging [3,4,5,6]. Stem cell exhaustion refers to an impaired functionality of stem cells, which cannot maintain in the tissue in which they reside. In particular, aging of the hematopoietic system displays decreased immune response, declining immuno-competence, increased autoimmunity, diminished stress response, late-onset anemia, reduced regenerative capacity, and increased predisposition to a spectrum of diseases, including myelodysplastic syndrome (MDS) and acute myeloid leukemia (AML) [7,8,9,10].
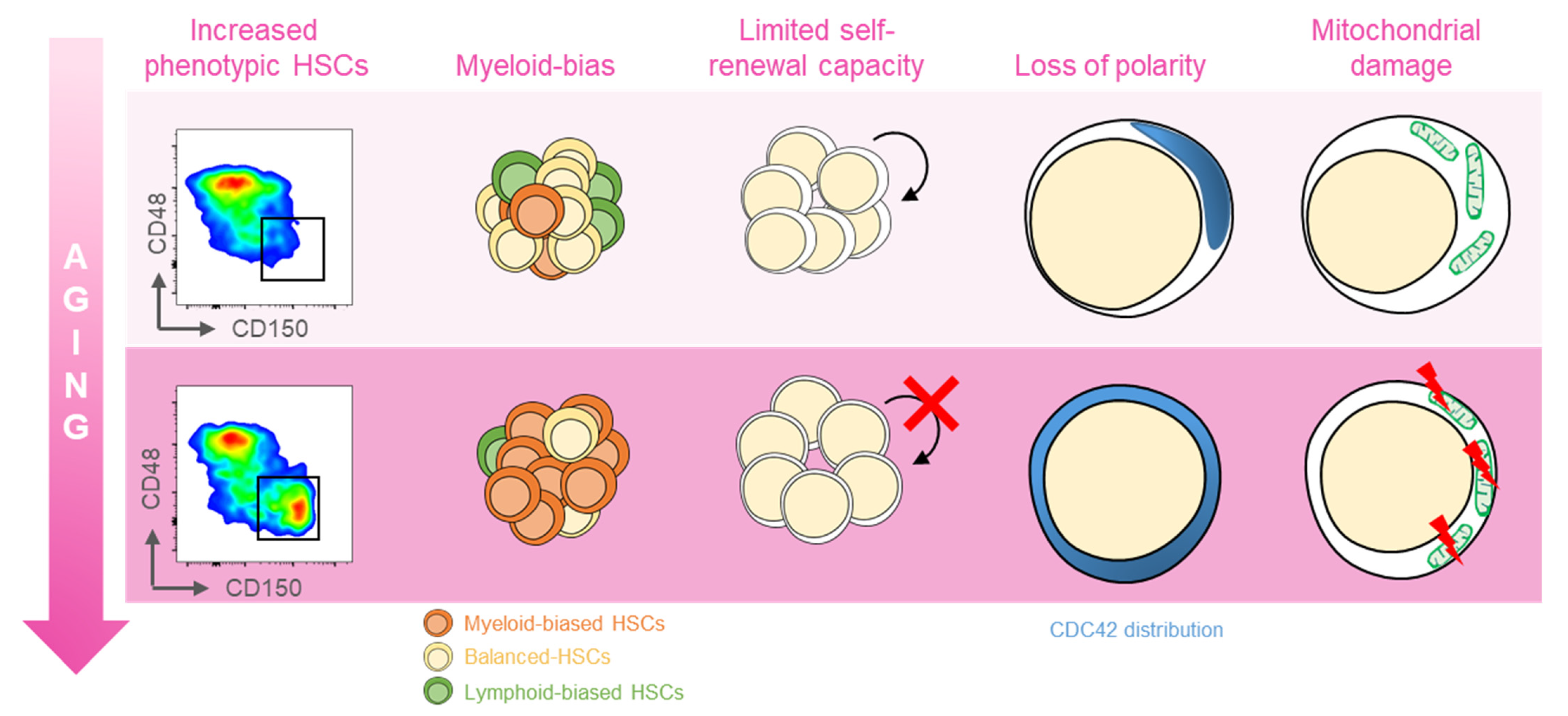
Several distinct features characterize aged hematopoietic stem cells (HSCs) (Figure 1) [11]. For example, it has been shown that phenotypic HSCs in the bone marrow increase in frequency with age, while losing their functionality. Multiple studies have demonstrated a differentiation bias toward the myeloid lineage; the aged murine hematopoietic system is impaired in supporting leukocyte numbers, erythropoiesis, and both B- and T-lymphoid cells in peripheral blood, while the numbers of myeloid cells are increased [12]. These myeloid-biased HSCs express high levels of CD150 (signaling lymphocyte activation molecule, or SLAMF1) and CD41 (integrin alpha 2, or Itga2b) proteins, which have been used to identify the HSC clonal subtypes responsible for hematopoietic aging [13,14]. As expected, stemness decreases in aged HSCs, which show reduced in vivo repopulation capacity, as determined by serial transplantation assays [15], along with a 3-fold lower efficiency in bone marrow homing after transplantation (Figure 1) [16].
Figure 1.
Features of aged HSCs. Upon aging, hematopoietic stem cells (HSCs) acquire phenotypical and functional peculiar properties. Flow cytometry analysis of the murine bone marrow shows an increase in the phenotypic HSCs (defined as c-Kit+Sca-1+Lin−CD135−CD48−CD150+) in old (18 months old, bottom panel) mice (far left). The HSC pool includes balanced HSC, which in equal proportion differentiate in myeloid and lymphoid lineage (yellow, left); with age, the myeloid-biased differentiation prevails at the expense of lymphoid cells (left). The self-renewal capacity typical of HSCs is reduced upon aging (middle). Cytoskeletal polarity detected by CDC42 localization is lost in old HSCs where CDC42 expression is homogeneously distributed (right). Aged HSCs display the mitochondrial damage with the altered metabolism (far right).
Another key feature of aged HSCs is the loss of cell polarity. Young and aged long-term (LT) HSCs exhibit different distribution patterns of CDC42, Tubulin, and AcH4K16, which is believed to be caused by the elevated activity of CDC42, and is linked to age-associated changes in self-renewal and differentiation capacity [12]. Additionally, HSC aging results from cumulative cellular and genomic damage, which leads to permanent cell-cycle arrest, apoptosis, or senescence [15,17,18].
Although historically DNA damage was thought to be the main cause of HSC aging, many new findings have defined an increasing number of biological processes that intrinsically change with age in HSCs. These include epigenetics, chromatin architecture, autophagy, proteostasis, and metabolic changes [19].
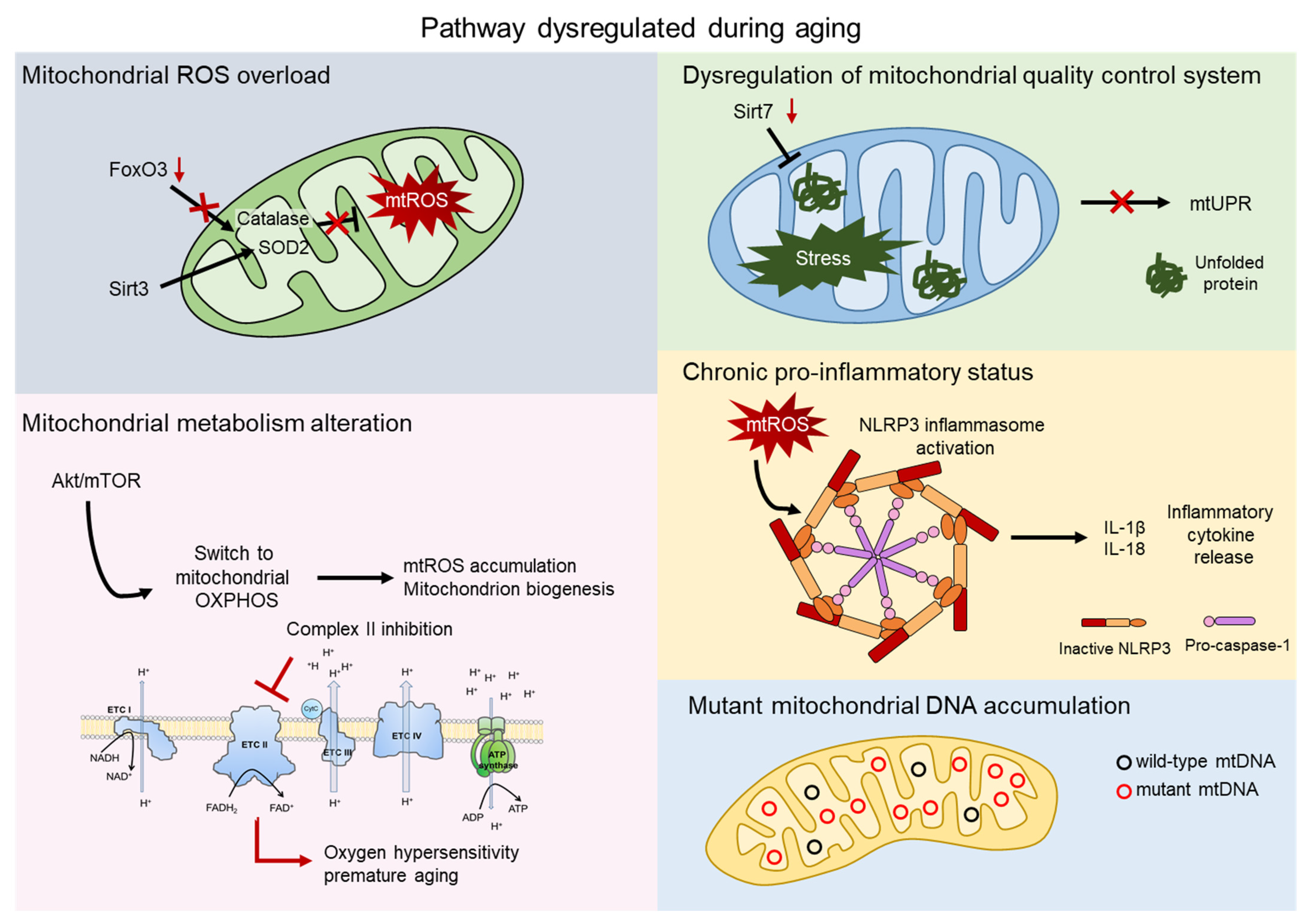
Since aged HSCs generally exhibit enhanced mitochondrial oxidative phosphorylation (OXPHOS) and increased production of reactive oxygen species (ROS), it has often been proposed that mitochondria play a direct role in compromising HSC functions [20,21]. Here, we explore multiple aspects of the impact of the mitochondria on HSC aging. Several mechanisms have been reported to contribute to aging, such as mitochondrial ROS production, alteration in mitochondrial metabolism, and mitochondrial quality control pathways (Figure 2).
Figure 2.
Mitochondrial contribution to aging. Aging process shows dysregulation of several pathway involving the direct contribution of mitochondria. Mitochondrial reactive oxygen species (mtROS) accumulate after the impairment of the major antioxidant defense systems, consist of Superoxide dismutase 2 (SOD2) and Catalase, caused by the aged-related loss of FOXO3 and SIRT3 (grey). AKT/mTOR signaling is a key regulator of mitochondrial metabolism and, upon aging, its imbalance causes mtROS accumulation, mitochondrial biogenesis, and an altered metabolism. Inhibition of complex II of the electron transport chain (ETC) leads to oxygen hypersensitivity and premature aging of HSCs (pink). The mitochondrial unfolded protein response system (mtUPR) is affected by the aged-related loss of SIRT7, leading to accumulation of unfolded protein and mitochondrial stress (green). Accumulation of mtROS activates the NOD-, LRR-, and pyrin domain-containing 3 (NLRP3) inflammasome, which triggers the release of inflammatory cytokines, such as IL-1β and IL-18 (orange). Mitochondrial dysfunctions include accumulation of mtDNA mutations in aged HSCs (blue).
2. Mitochondrial ROS
Since the frequency of HSCs with low levels of ROS decreases with age, ROS generation/accumulation can be considered a distinctive characteristic of aging [22]. Proper levels of ROS are important mediators of various signal transduction pathways. However, increased levels of ROS affect HSCs’s lifespan [20], self-renewal [23,24], and differentiation [25,26]. ROS contribute to HSC aging and senescence, and excessive ROS generation induces apoptotic cell death in HSCs [20,23,27,28]. Increase in ROS levels in adult HSC have similarities with the aging phenotypes, such as myeloid lineage skewing and defective long-term repopulation activity [29,30]. On the other hand, very low levels of intracellular ROS in HSCs are essential to maintaining HSCs quiescence [22]. Mitochondria produce around 90% of cellular ROS, and the impairment of mitochondrial function, for example, by the loss of the Polycomb repressor BMI1, causes a major increase in intracellular ROS [31]. Aging of mitochondria leads to an overload of ROS, which further damage the mitochondria, resulting in perpetual cell cycling [32]. Evidence for this has come from studies of the effects of FOXO transcription factors, key players in the oxidative stress response, on HSC fitness. Genetic variation within the FOXO3 gene is associated with human longevity and aging phenotypes [33]. Mice carrying triple conditional deletions of FOXO1, FOXO3a, and FOXO4 genes in the adult hematopoietic system exhibited myeloid lineage expansion, lymphoid developmental abnormalities, and a decreased long-term repopulation ability in vivo while increased ROS levels [30]. Another study from Dr. Tosho Suda’s group also demonstrated that FOXO3a-deleted HSCs can neither maintain quiescence nor support long-term reconstitution of hematopoiesis in vivo [34]. FOXO3a deficiency increased levels of ROS and downregulated several cyclin-dependent kinases inhibitors, resulting in the exit of HSCs from quiescence. This boosted sensitivity to cell-cycle-specific myelotoxic injury, and loss of self-renewal capacity during aging [34]. FOXO3a knockout HSCs also showed lower expression of mitochondrial Superoxide dismutase 2 (SOD2) and Catalase, two FOXO targets involved in ROS detoxification [35,36]. Further, FOXO3 has been shown to be crucial for the regulation of mitochondrial respiration in HSCs, which, under disrupted conditions, generate more ROS [29]. This strengthens the hypothesis that FOXO3a deficiency causes HSCs cell cycle abnormalities via mitochondrial ROS dysregulation.
Mitochondrial ROS levels and the related signaling pathways, thus, represent a major player in regulating the long-term self-renewal, activation, proliferation, differentiation, and aging of HSCs. Similar roles could also be played by extrinsic factors and the surrounding microenvironment (see Box 1), which can have a direct impact on ROS levels and the signaling pathways regulating HSCs homeostasis [10].
ROS are produced as by-products of mitochondrial respiration; their production is increased, and they are accumulated when mitochondrial respiration is altered. The major ROS source is the mitochondrial electron transport Chain (ETC), which is widely targeted by mitochondrial DNA (mtDNA) mutations [37], as described in detail in the following dedicated paragraphs.
Box 1. Bone Marrow Microenvironment.
Upon aging, drastic changes to the bone marrow
microenvironment may serve as an extrinsic factor that promotes HSC aging.
These changes include higher levels of several niche-derived soluble factors,
such as the pro-inflammatory CC-chemokine ligand 5 (CCL5), osteopontin, and
CXCL12 [38,39,40], as well as niche anatomical remodeling
[41,42].
Pro-inflammatory cytokines increase with age in the bone marrow
microenvironment of both mice and humans, driving myeloid differentiation
[43]. In aged-related
myeloid malignancies, such as myeloid proliferative neoplasms and chronic
myeloid leukemia, serum interleukine (IL)-1β and IL6 levels are elevated [44,45]. Further studies
will clarify whether inflammation is the cause or the consequence of HSC
aging. Anatomical and functional remodeling of the HSC niche accelerates
myeloid-lineage cell expansion during aging. Maryanovich et al. demonstrated
that bone marrow vasculature and its associated stromal cells are remodeled
in elder mice, and that this is associated with progressive deterioration of
the sympathetic nervous system (SNS). SNS neuropathy is an early driver of
niche aging, and loss of SNS by surgical denervation or genetic deletion of
neurotransmitter-targeting β3 adrenergic receptors induces the remodeling of
the HSC niche and leads to premature aging-like changes in HSCs [46].
Another study also
suggested that, during normal aging, increased β2 adrenergic receptors
activity promotes IL6-dependent myeloid differentiation and subsequent
premature HSC aging [47].
3. Mitochondrial Metabolism
Despite the high preference of mitochondria for glycolysis, recent studies have also highlighted the importance of mitochondrial respiration to HSC for proliferation and maintenance [48,49,50]. We have recently shown that HSCs have a relatively high number of mitochondria, which are not completely inactive [51]. Indeed, mitochondrial membrane potential (ΔΨmt) is high in HSCs, although ATP production or intracellular ROS levels are low [52,53]. The higher complex II: complex V ratio gives rise to high ΔΨmt in HSCs due to limited coupling of the electron transport chain (ETC), which supports the idea that mitochondrial complex II is pivotal for both HSC maintenance and the prevention of the aging process [53]. Indeed, inhibition of complex II reduces the in vitro colony-replating capacity of HSCs [53], and genetic mutation of mev-1, a subunit of the succinate dehydrogenase cytochrome b enzyme, which is a component of complex II, leads to oxygen hypersensitivity and premature aging of HSCs [54]. Studies of C. elegans uncovered that the mutated or silenced components of ETC or the ATP synthase can markedly extend [55,56,57] or reduce lifespan [58]. Although these varying experimental results must eventually be resolved, it is clear that imbalances in ETC activity are closely linked to the overall survival of the organism. Interestingly, a recent paper showed that ΔΨmt is a source of heterogeneity in old HSCs, with a prevalent fraction of low ΔΨmt in aged HSCs. Enhancement of ΔΨmt by mitoquinol (Mito-Q), a mitochondrial-targeted coenzyme-Q10 [59], successfully increased ΔΨmt of old HSCs and ameliorated or prevented onset of aging phenotypes [60].
HSCs are mainly dormant but can become highly active on demand, either to maintain hematopoietic homeostasis by replenishing matured/immature hematopoietic cells, or to respond to situations of emergency, such as infection or blood loss [61]. This shift requires a metabolic switch from glycolysis to mitochondrial oxidative phosphorylation, which is precisely regulated by various signaling pathways. The mammalian TOR (mTOR) pathway is a key regulator of cellular and mitochondrial metabolism. mTOR directly controls the mitochondrial oxidative function through a YY1–PGC-1α (peroxisome proliferator-activated receptor gamma coactivator 1-alpha) transcriptional complex [62]. Defects of Tuberous sclerosis complex subunit 1 (TSC1), the major negative regulator for mTOR [63], lead to increased mitochondrion biogenesis and accumulation of ROS. Blockade of ROS activity in vivo restores these HSC defects, demonstrating that the TSC-mTOR pathway controls the quiescence and on-demand functions of HSCs by repressing ROS production [64,65].
HSCs exhibit low AKT/mTOR activity, but, upon stress, upregulation of this pathway drives dormant HSCs toward activation [64,66]. Interestingly, the dysregulation of AKT/mTOR signaling correlates with the aging process in HSCs [67]. Experimental evidence has shown that mTOR activation is involved in HSC aging, as well as that rapamycin treatment restores HSC potential and prolongs the lifespan of mice [68]. mTOR activity is higher in HSCs from elder mice than younger mice, and mTOR activation, through conditional deletion of TSC1 in the HSCs of young mice, mimics the phenotype of HSCs from aged mice; similarly, in older mice, rapamycin restores the self-renewal capacity of HSCs and, importantly, correlates with increased life span [68].
ASXL1 is frequently mutated in age-related clonal hematopoiesis. Its mutation activates the AKT/mTOR pathway, causing aberrant cell cycle progression in the HSC compartment and provoking HSC dysfunction. This is associated with mitochondrial activation, elevated ROS levels, and increased DNA damage, leading to age-associated phenotypes, such as myeloid-biased differentiation, hypocellular bone marrow, and increased frequency, of phenotypic LT-HSCs. Inhibition of the AKT/mTOR pathway can partially rescue these phenotypes, suggesting its involvement in the enhanced aging of the hematopoietic system [69].
A similar phenotype is observed in wild-type p53-induced phosphatase 1 (WIP1), which is highly expressed in HSCs but decreases with age. WIP1-deficient (WIP1−/−) mice exhibit multiple aging-like phenotypes in HSCs, including declines in reconstitution ability and HSC expansion. Mechanistically, their impaired regenerative capacity is due to a p53-mediated differentiation defect, whereas increasing numbers of WIP1−/− HSCs are associated with mTOR-mediated cell cycle progression of HSCs [70]. Notably, experimental results have shown that aged HSCs have higher mTOR [71] activity levels, as well as that its inhibitor rapamycin can restore the self-renewal of aged HSCs, an effect which can be translated to human HSCs [72].
Recent advances have demonstrated that epigenetic, transcriptional, and post-transcriptional mechanisms also control the quiescence of HSCs, which are maintained in a paused state that allows for rapid activation [73]. Mitochondrial activity modifies the epigenetic state of cells affecting their aging process [74]. Citric acid, generated by the tricarboxylic acid (TCA) cycle in the mitochondria, modulates histone acetylation and gene expression through its conversion to acetyl-CoA [74]. Mitochondrial fatty acid oxidation (FAO) also generates acetyl-CoA for histone modification in HSCs [75].
Sirtuins are a family of protein deacetylases, which regulate the mitochondrial metabolic checkpoint in stem cells, and they are key regulators of stem cell aging [21,76]. SIRT3 plays a critical role in the mitochondria, where it deacetylates two critical lysine residues on SOD2 to promote the antioxidative activity. Brown and colleagues have demonstrated that SIRT3 is highly enriched in HSCs, as well as is suppressed with aging [77]. Although SIRT3 has no effect on HSCs maintenance or tissue homeostasis at a young age under homeostatic conditions, it is essential under stress or in old age. Indeed, SIRT3 loss induces HSC quiescence and compromises regenerative capacity in old mice [77].
SIRT1 is a key regulator of HSCs self-renewal and lineage specification under homeostasis. Interestingly, Ghaffari’s group has shown that loss of SIRT1 causes anemia and myeloid expansion at the expense of the lymphoid compartment, overlapping features with aged HSCs. SIRT1 plays a role in HSCs homeostasis by targeting FOXO3, a longevity transcription factor and mitochondrial ROS regulator [78].
Another key regulator of metabolism is nicotinamide adenine dinucleotide (NAD+). Decreased levels of NAD+ are associated with cancer, metabolic disorders, and physiological and accelerated aging processes [79,80,81]. Supplementation of nicotinamide riboside (NR), a NAD+ precursor, significantly improved lifespan and health span in model of aged-related disease, such as ataxia–telangiectasia mutation (ATM), thanks to the improvement of both DNA damage repair and mitophagy [71,82]. Murine models of ATM loss show defects in DNA damage repair associated with mitochondrial dysfunction [83] and loss of hematopoietic stem cell (HSC) potential [23]. NR treatment caused significant alterations in lineage commitment of HSCs with enhanced lymphoid potential [84].
4. Mitochondrial Quality Control
Increased evidence indicates that mitochondrial integrity is disrupted during aging, and this contributes to the pathogenesis of age-related disorders in humans [3,85]. Mitochondria have evolved multiple mechanisms to guarantee mitochondrial quality. For instance, mitochondrial chaperones and proteases prevent the accumulation of misfolded and aggregated proteins by monitoring proteostasis through the mitochondrial unfolded protein stress response (mtUPR) [86], a mechanism that has been shown to be critical for longevity in mammals [87,88]. The mtUPR is a cellular protective program that ensures proteostasis in the mitochondria and is activated by mitochondrial protein folding stress [89]. mtUPR has recently emerged as a regulatory mechanism for adult stem cell maintenance [21,90,91,92]. This protective program is dysregulated during physiological aging, which contributes to the functional deterioration of stem cells, tissue degeneration, and shortened organismal lifespan [21,91]. SIRT7 is an NAD+-dependent H3K18Ac (acetylated lysine 18 of histone H3) deacetylase originally studied for its role in cancer cells [93]. Mohrin et al. have shown that SIRT7 represses the expression of mitochondrial ribosomal proteins in order to regulate mtUPR and reduce mitochondrial protein folding stress [21]. SIRT7 ablation leads to loss of quiescence and aging phenotypes in HSCs, including reduced regenerative capacity and the myeloid-biased differentiation [21].
Stem cells exhibit high levels of autophagy as part of their physiological state [94]. Autophagic activity is necessary for the self-renewal and differentiation capacities of stem cells, particularly HSCs [95]. Autophagy is closely linked with health and longevity, and impaired levels of autophagy in aged HSCs leads to the accumulation of mitochondria, which in turn induces metabolic stress [96,97]. Lysosomal sequestration of mitochondrial enhances the regenerative capacity of HSCs [98]. Overall, autophagy declines in aged stem cells, contributing to loss of quiescence, senescence, and, ultimately, degeneration [99].
Deregulation of other compensatory mitochondrial protective programs, such as mitophagy and mitochondrial dynamics, also affect stem cell maintenance highlighting the importance of mitochondrial integrity [95,96,97,100,101,102]. Indeed, mitochondria are dynamic organelles existing in large tubular and highly dynamic networks that constantly undergo fission and fusion processes, thereby leading to the dilution of non-functional mitochondria [103]. The effects of mitophagy and mitochondrial fusion/fission process on health and lifespan has been particularly demonstrated by using the model organisms C. elegans and D. melanogaster. For instance, the overexpression of the mitochondrial fission protein dynamin-related protein 1 (DRP1) increased the healthy lifespan in flies [104]. The importance of mitochondrial fission on drosophila lifespan was further demonstrated by the observation that lifespan extension caused by the overexpression of p62 was abrogated in DRP1 mutant flies [105]. Lifespan extension in flies was also observed after overexpression of mitophagy key proteins, such as PARKIN and PINK1 [106,107]. These findings are consistent with studies in C. elegans, where mitophagy has been shown to contribute to lifespan regulation [108,109].
5. Inflammation
The natural aging process is associated with activation of the innate immune system, which results in a low-grade chronic pro-inflammatory status, even in the absence of overt diseases [43,110]. Aged-related inflammation (inflammaging) is due to the systemic overabundance of pro-inflammatory cytokines, such as IL-1, tumor necrosis factor (TNF), and IL-6 [111,112]. Several studies focused on HSCs have shown that inflammatory signaling induces the differentiation of the myeloid progenitor cells required to withstand harmful stimuli [112,113,114]. Specifically, IFN-α and IFN-γ activate HSCs entry into the cell cycle and boost the myeloid-biased differentiation of HSCs [115,116], as well as IL-1β, which promotes myeloid lineage biased differentiation of HSCs [117].
Notably, this cytokine network, termed the senescence-associated secretory phenotype (SASP) [118], may be initiated by senescent cells producing IL-1α in the bone marrow microenvironment (Box 1) [119]. HSCs do not solely show mitochondrial defects; for example, CD4+ T cells from elderly people display an elevated number of dysfunctional mitochondria engulfed into autophagosomes compared to cells from young people, suggesting the presence of a defective mitochondrial turnover. These defective mitochondria may be the source of inflammatory stimuli and contribute to the impairment of immune defenses in the elderly [120,121].
Aged HSCs exhibit increased NF-κB activity mediated by RAD21/cohesion, which enhances sensitivity to inflammatory stimuli, higher production of IL-6, and myeloid-biased hematopoietic differentiation [122]. Recently, the Wang group has shown that systemic level of TNF-α, a well-known biomarker of inflammation, increases with age and induces the expression of IL27Ra in HSCs via ERK-ETS1 signaling [123]. The chronic inflammatory process associated with aging leads to dysfunctional differentiation of stem cells, loss of self-renewal capacity, and further promotion of HSC aging [124].
The relevance of mitochondria to the pro-inflammatory response has been largely studied as a source of damage-associated molecular patterns (DAMPs) during cell death [125]. Mitochondrial DAMPs include mtDNA, cytochrome c, ROS, and ATP, which, when released in the cytosol, are recognized by the cell as a red flag of danger and trigger apoptosis or necrosis [125]. Besides pro-apoptotic signals, mitochondria DAMPs are also potent immunostimulators. The exposure to mtDNA triggers a variety of innate immune responses due to its bacterial origin [126], and the modulation of ROS signaling causes the activation of the principal component of innate immunity, the NOD-, LRR-, and pyrin domain-containing 3 (NLRP3) inflammasome [127,128].
The use of specific mitochondria ROS scavenger, the mito-TEMPO, inhibit the NLRP3 inflammasome activation, reducing the up-regulation of IL-1β and IL-18 induced by lipopolysaccharide (LPS) [127,129]. Luo et al. recently demonstrated that mitochondrial stress activates the NLRP3 inflammasome in HSCs as part of the key role played by mitochondria in the inflammation process during aging [130]. ROS are a metabolic danger signal and activate an innate immune sensor, the NLRP3 inflammasome. Once triggered, the NLRP3 inflammasome induces pro-inflammatory cytokine secretion and/or caspase 1-dependent cell death [131]. NLRP3 is highly expressed and studied in myeloid cells; however, it is also expressed and functions in HSCs [130]. NLRP3 is a substrate of SIRT2, which inhibits NLRP3 activity through deacetylation [132]. The reduced expression of SIRT2 in aged HSCs enhances the activity of the NLRP3 inflammasome, which increases susceptibility to mitochondrial stress-induced stem cell deterioration. Interestingly, the overexpression of SIRT3 or SIRT7 (described above) reduces caspase 1 activity and improves the function of aged HSCs [130], tightening the link between mitochondria and inflammation.
Finally, patients with high levels of circulating mtDNA (described in the next section) show higher concentrations of IL-6, TNF-a, RANTES, and IL-1 [133].
6. Mitochondrial DNA Mutations
The human mtDNA genome encodes 13 genes, 22 tRNAs, and 2 rRNAs [134]. These 13 genes encode for core subunits of the electron transport chain (ETC) complexes and the ATP synthase. During aging, mtDNA accumulates mutation, causing dysfunction of mitochondria and the respiratory chain [135]. Several papers proposed that mtDNA mutations play a key role in aging [3,136,137]. mtDNA has a much higher mutation rate than nuclear DNA [138,139]. Indeed, the mtDNA spatial proximity to the site of ETC-mediated ROS production make it particularly susceptible to ROS damage. In addition, differently from the nuclear DNA, mtDNA cannot be organized in highly compacted structures by histones, enhancing the probability of coping errors introduced during replication [140]. Interestingly, it has been reported that the aged population has a higher mtDNA copy number in blood [141,142] and exhibits mtDNA heteroplasmy, i.e., the presence of more than one type of organelle genome [143,144]. Of note, individual mtDNA mutations were found in centenarians [145]. One example is represented by A5178C mutation in MT-ND2 gene, which encodes the NADH dehydrogenase 2, a subunit of ETC complex I. This mutation confers protection on mitochondria against oxidative damage contribute to longevity [146].
Conplastic mice strains are a suitable model system for the study of specific mtDNA variations and their influence on ROS and ATP levels upon aging [147]. In this paper, conplastic mouse strains C57BL/6Ntac-mtAKR/J (mtAKR), C57BL/6Ntac-mt129S1SvlmJ (mt129S1), C57BL/6Ntac-mtNOD/LTJ (mtNOD), and the background strain C57BL/6Ntac (B6Ntac) were used to investigated their hematopoietic changes during aging. The three mouse strains harbor specific polymorphisms in the mtDNA affecting complex I, III, and IV of respiratory chain [148]. The presence of mtDNA polymorphisms in these subunits of the respiratory chain decreased intracellular ROS levels and lymphocyte counts during aging [148].
Although somatic mtDNA mutations accumulate in multiple tissues with age [136], its causal role in tissue aging remains to be clarified [149]. The direct effects of mtDNA mutations have been studied through the analysis of a mouse model carrying a proofreading-defective mitochondrial DNA polymerase (POLGAD257A) [150]. The consequent accumulation of mtDNA mutations led to premature aging of the mice, which showed hair loss, weight loss, osteoporosis, anemia, and myeloid lineage skewing with lymphopenia [150,151,152,153,154]. Accumulating mtDNA point mutations destabilized ETC complexes I, III, and IV, leading to respiratory chain deficiency [155]. The POLGAD257A did not show an increase in ROS production [156], but an antioxidant treatment has separately been shown to rescue erythroid differentiation in embryos [151,152]. Chen et al. showed that POLGAD257A mutant animals develop an age-dependent, macrocytic anemia with abnormal erythroid maturation and megaloblastic changes, as well as profound defects, in lymphopoiesis. These abnormalities phenocopy patients with myelodysplastic syndrome (MDS) and refractory anemia, suggesting that abnormalities of mitochondrial function can be involved in the pathogenesis of the diseases [157].
In humans, POLG (DNA Polymerase Subunit Gamma) mutations were not linked to symptoms of premature aging, even though they are one of the most frequent causes of mitochondrial disease [158]. Mice carrying a defect in mitochondrial genome maintenance exonuclease 1 (MGME1) showed mtDNA replication defects and developed a severe multisystemic mitochondrial disorder [159] without signs of premature aging [160]. All things considered, mitochondrial dysfunctions caused by accumulating mtDNA mutations has been shown to cause multiple hematopoietic defects that are typically seen in the elderly, but mtDNA mutations alone may not be responsible for the phenotype associated with aging HSCs [161,162,163].
Dr. Sankaran’s group has recently presented a new fascinating tool for the studying of mtDNA mutation. They developed a high-throughput platform for measuring mtDNA mutation heteroplasmy, along with accessible chromatin states, in thousands of single cells [164,165]. Performing clonal tracing in human hematopoiesis in vivo has the potential to resolve the clonal heterogeneity within malignancies and the aging process.
7. Conclusions and Future Perspective
With a better understanding of the mechanisms of HSC aging, researchers will be able to explore new opportunities to prevent, delay, or even reverse aspects of this process. In this review, we summarize accumulating evidence that support the concept that mitochondrial stress is one of the main drivers of stem cell deterioration with age. Targeting mitochondrial protective pathways could, therefore, allow us to protect stem cells from aging, with important therapeutic implications.
Since the studies of HSC aging have been performed, for the most part, in mouse models, it is crucial to determine whether these findings can be translated to humans. Increasing evidence has revealed a phenomenon termed age-related clonal hematopoiesis (ARCH), describing a clonal expansion of blood cells derived from mutated HSCs in aged humans [166]. Single-cell sequencing of sorted cell populations is able to identify even subclonal mutation inside the stem cell pool, revealing the high complexity of this process [167]. It would be of great interest to determine whether the mitochondria plays a role in the development of human clonal hematopoiesis.
Finally, a better understanding of the molecular properties of the clonal subclass of myeloid-biased HSCs may provide insights into the onset of clinically age-dependent hematological disorders/malignancies derived from stem cells. ARCH can be a precursor to MDS, and MDS is associated with age with a median age at diagnosis of 65–70 years. AML is an aggressive hematological disorder mainly affecting people of advancing age, and 30% of patients with AML are 75 years or older [168]. Notably, we recently revealed a mitochondrial ROS senescence pathway triggered by Nucleophosmin 1 mutant (NPM1c), PML, and TP53, which plays a crucial role in actinomycin D-based therapies in AML [169]. This supports the idea of mitochondria as potential therapeutic target.
Author Contributions
C.M. prepared the original draft. K.I. reviewed and edited the final version. All authors have read and agreed to the published version of the manuscript.
Funding
This research was funded by National Institutes of Health (R01HL148852, R01DK098263, and R01DK115577).
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Acknowledgments
We are grateful to members of the Ito lab and the Einstein Stem Cell Institute for their comments on the topics of hematology and stem cell biology. K.I. is supported by grants from the National Institutes of Health (R01HL148852, R01DK098263, and R01DK115577). K.I. is a Research Scholar of the Leukemia & Lymphoma Society (#1360-19). C.M. is supported by The Einstein Training Program in Stem Cell Research, which is acknowledged from the Empire State Stem Cell Fund through New York State Department of Health Contract (C34874GG).
Conflicts of Interest
The authors declare no conflict of interest.
References
- Lopez-Otin, C.; Blasco, M.A.; Partridge, L.; Serrano, M.; Kroemer, G. The hallmarks of aging. Cell 2013, 153, 1194–1217. [Google Scholar] [CrossRef] [Green Version]
- Speakman, J.R. Body size, energy metabolism and lifespan. J. Exp. Biol. 2005, 208, 1717–1730. [Google Scholar] [CrossRef] [Green Version]
- Kauppila, T.E.S.; Kauppila, J.H.K.; Larsson, N.G. Mammalian Mitochondria and Aging: An Update. Cell Metab. 2017, 25, 57–71. [Google Scholar] [CrossRef] [Green Version]
- Sebastian, D.; Palacin, M.; Zorzano, A. Mitochondrial Dynamics: Coupling Mitochondrial Fitness with Healthy Aging. Trends Mol. Med. 2017, 23, 201–215. [Google Scholar] [CrossRef] [PubMed]
- Jang, J.Y.; Blum, A.; Liu, J.; Finkel, T. The role of mitochondria in aging. J. Clin. Investig. 2018, 128, 3662–3670. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mu, W.C.; Ohkubo, R.; Widjaja, A.; Chen, D. The mitochondrial metabolic checkpoint in stem cell aging and rejuvenation. Mech. Ageing Dev. 2020, 188, 111254. [Google Scholar] [CrossRef] [PubMed]
- Kiss, T.L.; Sabry, W.; Lazarus, H.M.; Lipton, J.H. Blood and marrow transplantation in elderly acute myeloid leukaemia patients-older certainly is not better. Bone Marrow Transpl. 2007, 40, 405–416. [Google Scholar] [CrossRef]
- Signer, R.A.; Montecino-Rodriguez, E.; Witte, O.N.; McLaughlin, J.; Dorshkind, K. Age-related defects in B lymphopoiesis underlie the myeloid dominance of adult leukemia. Blood 2007, 110, 1831–1839. [Google Scholar] [CrossRef]
- Linton, P.J.; Dorshkind, K. Age-related changes in lymphocyte development and function. Nat. Immunol. 2004, 5, 133–139. [Google Scholar] [CrossRef]
- Ergen, A.V.; Goodell, M.A. Mechanisms of hematopoietic stem cell aging. Exp. Gerontol. 2010, 45, 286–290. [Google Scholar] [CrossRef] [Green Version]
- de Haan, G.; Lazare, S.S. Aging of hematopoietic stem cells. Blood 2018, 131, 479–487. [Google Scholar] [CrossRef] [Green Version]
- Florian, M.C.; Dorr, K.; Niebel, A.; Daria, D.; Schrezenmeier, H.; Rojewski, M.; Filippi, M.D.; Hasenberg, A.; Gunzer, M.; Scharffetter-Kochanek, K.; et al. Cdc42 activity regulates hematopoietic stem cell aging and rejuvenation. Cell Stem Cell 2012, 10, 520–530. [Google Scholar] [CrossRef] [Green Version]
- Beerman, I.; Bhattacharya, D.; Zandi, S.; Sigvardsson, M.; Weissman, I.L.; Bryder, D.; Rossi, D.J. Functionally distinct hematopoietic stem cells modulate hematopoietic lineage potential during aging by a mechanism of clonal expansion. Proc. Natl. Acad. Sci. USA 2010, 107, 5465–5470. [Google Scholar] [CrossRef] [Green Version]
- Gekas, C.; Graf, T. CD41 expression marks myeloid-biased adult hematopoietic stem cells and increases with age. Blood 2013, 121, 4463–4472. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Janzen, V.; Forkert, R.; Fleming, H.E.; Saito, Y.; Waring, M.T.; Dombkowski, D.M.; Cheng, T.; DePinho, R.A.; Sharpless, N.E.; Scadden, D.T. Stem-cell ageing modified by the cyclin-dependent kinase inhibitor p16INK4a. Nature 2006, 443, 421–426. [Google Scholar] [CrossRef] [PubMed]
- Liang, Y.; Van Zant, G.; Szilvassy, S.J. Effects of aging on the homing and engraftment of murine hematopoietic stem and progenitor cells. Blood 2005, 106, 1479–1487. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rossi, D.J.; Jamieson, C.H.; Weissman, I.L. Stems cells and the pathways to aging and cancer. Cell 2008, 132, 681–696. [Google Scholar] [CrossRef] [Green Version]
- Sahin, E.; DePinho, R.A. Axis of ageing: Telomeres, p53 and mitochondria. Nat. Rev. Mol. Cell Biol. 2012, 13, 397–404. [Google Scholar] [CrossRef] [Green Version]
- Mejia-Ramirez, E.; Florian, M.C. Understanding intrinsic hematopoietic stem cell aging. Haematologica 2020, 105, 22–37. [Google Scholar] [CrossRef]
- Ito, K.; Hirao, A.; Arai, F.; Takubo, K.; Matsuoka, S.; Miyamoto, K.; Ohmura, M.; Naka, K.; Hosokawa, K.; Ikeda, Y.; et al. Reactive oxygen species act through p38 MAPK to limit the lifespan of hematopoietic stem cells. Nat. Med. 2006, 12, 446–451. [Google Scholar] [CrossRef]
- Mohrin, M.; Shin, J.; Liu, Y.; Brown, K.; Luo, H.; Xi, Y.; Haynes, C.M.; Chen, D. Stem cell aging. A mitochondrial UPR-mediated metabolic checkpoint regulates hematopoietic stem cell aging. Science 2015, 347, 1374–1377. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jang, Y.Y.; Sharkis, S.J. A low level of reactive oxygen species selects for primitive hematopoietic stem cells that may reside in the low-oxygenic niche. Blood 2007, 110, 3056–3063. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ito, K.; Hirao, A.; Arai, F.; Matsuoka, S.; Takubo, K.; Hamaguchi, I.; Nomiyama, K.; Hosokawa, K.; Sakurada, K.; Nakagata, N.; et al. Regulation of oxidative stress by ATM is required for self-renewal of haematopoietic stem cells. Nature 2004, 431, 997–1002. [Google Scholar] [CrossRef] [PubMed]
- Le, Q.; Yao, W.; Chen, Y.; Yan, B.; Liu, C.; Yuan, M.; Zhou, Y.; Ma, L. GRK6 regulates ROS response and maintains hematopoietic stem cell self-renewal. Cell Death Dis. 2016, 7, e2478. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cao, Y.; Fang, Y.; Cai, J.; Li, X.; Xu, F.; Yuan, N.; Zhang, S.; Wang, J. ROS functions as an upstream trigger for autophagy to drive hematopoietic stem cell differentiation. Hematology 2016, 21, 613–618. [Google Scholar] [CrossRef] [Green Version]
- Maryanovich, M.; Zaltsman, Y.; Ruggiero, A.; Goldman, A.; Shachnai, L.; Zaidman, S.L.; Porat, Z.; Golan, K.; Lapidot, T.; Gross, A. An MTCH2 pathway repressing mitochondria metabolism regulates haematopoietic stem cell fate. Nat. Commun. 2015, 6, 7901. [Google Scholar] [CrossRef]
- Maryanovich, M.; Gross, A. A ROS rheostat for cell fate regulation. Trends Cell Biol. 2013, 23, 129–134. [Google Scholar] [CrossRef]
- Nitta, E.; Yamashita, M.; Hosokawa, K.; Xian, M.; Takubo, K.; Arai, F.; Nakada, S.; Suda, T. Telomerase reverse transcriptase protects ATM-deficient hematopoietic stem cells from ROS-induced apoptosis through a telomere-independent mechanism. Blood 2011, 117, 4169–4180. [Google Scholar] [CrossRef] [Green Version]
- Rimmele, P.; Liang, R.; Bigarella, C.L.; Kocabas, F.; Xie, J.; Serasinghe, M.N.; Chipuk, J.; Sadek, H.; Zhang, C.C.; Ghaffari, S. Mitochondrial metabolism in hematopoietic stem cells requires functional FOXO3. EMBO Rep. 2015, 16, 1164–1176. [Google Scholar] [CrossRef] [Green Version]
- Tothova, Z.; Kollipara, R.; Huntly, B.J.; Lee, B.H.; Castrillon, D.H.; Cullen, D.E.; McDowell, E.P.; Lazo-Kallanian, S.; Williams, I.R.; Sears, C.; et al. FoxOs are critical mediators of hematopoietic stem cell resistance to physiologic oxidative stress. Cell 2007, 128, 325–339. [Google Scholar] [CrossRef] [Green Version]
- Liu, J.; Cao, L.; Chen, J.; Song, S.; Lee, I.H.; Quijano, C.; Liu, H.; Keyvanfar, K.; Chen, H.; Cao, L.Y.; et al. Bmi1 regulates mitochondrial function and the DNA damage response pathway. Nature 2009, 459, 387–392. [Google Scholar] [CrossRef] [PubMed]
- Balaban, R.S.; Nemoto, S.; Finkel, T. Mitochondria, oxidants, and aging. Cell 2005, 120, 483–495. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Willcox, B.J.; Donlon, T.A.; He, Q.; Chen, R.; Grove, J.S.; Yano, K.; Masaki, K.H.; Willcox, D.C.; Rodriguez, B.; Curb, J.D. FOXO3A genotype is strongly associated with human longevity. Proc. Natl. Acad. Sci. USA 2008, 105, 13987–13992. [Google Scholar] [CrossRef] [Green Version]
- Miyamoto, K.; Araki, K.Y.; Naka, K.; Arai, F.; Takubo, K.; Yamazaki, S.; Matsuoka, S.; Miyamoto, T.; Ito, K.; Ohmura, M.; et al. Foxo3a is essential for maintenance of the hematopoietic stem cell pool. Cell Stem Cell 2007, 1, 101–112. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Essers, M.A.; Weijzen, S.; de Vries-Smits, A.M.; Saarloos, I.; de Ruiter, N.D.; Bos, J.L.; Burgering, B.M. FOXO transcription factor activation by oxidative stress mediated by the small GTPase Ral and JNK. EMBO J. 2004, 23, 4802–4812. [Google Scholar] [CrossRef] [Green Version]
- Kops, G.J.; Dansen, T.B.; Polderman, P.E.; Saarloos, I.; Wirtz, K.W.; Coffer, P.J.; Huang, T.T.; Bos, J.L.; Medema, R.H.; Burgering, B.M. Forkhead transcription factor FOXO3a protects quiescent cells from oxidative stress. Nature 2002, 419, 316–321. [Google Scholar] [CrossRef]
- Park, C.B.; Larsson, N.G. Mitochondrial DNA mutations in disease and aging. J. Cell Biol. 2011, 193, 809–818. [Google Scholar] [CrossRef] [Green Version]
- Guidi, N.; Sacma, M.; Standker, L.; Soller, K.; Marka, G.; Eiwen, K.; Weiss, J.M.; Kirchhoff, F.; Weil, T.; Cancelas, J.A.; et al. Osteopontin attenuates aging-associated phenotypes of hematopoietic stem cells. EMBO J. 2017, 36, 840–853. [Google Scholar] [CrossRef] [Green Version]
- Ergen, A.V.; Boles, N.C.; Goodell, M.A. Rantes/Ccl5 influences hematopoietic stem cell subtypes and causes myeloid skewing. Blood 2012, 119, 2500–2509. [Google Scholar] [CrossRef] [Green Version]
- Tuljapurkar, S.R.; McGuire, T.R.; Brusnahan, S.K.; Jackson, J.D.; Garvin, K.L.; Kessinger, M.A.; Lane, J.T.; BJ, O.K.; Sharp, J.G. Changes in human bone marrow fat content associated with changes in hematopoietic stem cell numbers and cytokine levels with aging. J. Anat. 2011, 219, 574–581. [Google Scholar] [CrossRef]
- Kusumbe, A.P.; Ramasamy, S.K.; Itkin, T.; Mae, M.A.; Langen, U.H.; Betsholtz, C.; Lapidot, T.; Adams, R.H. Age-dependent modulation of vascular niches for haematopoietic stem cells. Nature 2016, 532, 380–384. [Google Scholar] [CrossRef] [Green Version]
- Poulos, M.G.; Ramalingam, P.; Gutkin, M.C.; Llanos, P.; Gilleran, K.; Rabbany, S.Y.; Butler, J.M. Endothelial transplantation rejuvenates aged hematopoietic stem cell function. J. Clin. Investig. 2017, 127, 4163–4178. [Google Scholar] [CrossRef] [Green Version]
- Kovtonyuk, L.V.; Fritsch, K.; Feng, X.; Manz, M.G.; Takizawa, H. Inflamm-Aging of Hematopoiesis, Hematopoietic Stem Cells, and the Bone Marrow Microenvironment. Front. Immunol. 2016, 7, 502. [Google Scholar] [CrossRef] [Green Version]
- Arranz, L.; Sanchez-Aguilera, A.; Martin-Perez, D.; Isern, J.; Langa, X.; Tzankov, A.; Lundberg, P.; Muntion, S.; Tzeng, Y.S.; Lai, D.M.; et al. Neuropathy of haematopoietic stem cell niche is essential for myeloproliferative neoplasms. Nature 2014, 512, 78–81. [Google Scholar] [CrossRef]
- Reynaud, D.; Pietras, E.; Barry-Holson, K.; Mir, A.; Binnewies, M.; Jeanne, M.; Sala-Torra, O.; Radich, J.P.; Passegue, E. IL-6 controls leukemic multipotent progenitor cell fate and contributes to chronic myelogenous leukemia development. Cancer Cell 2011, 20, 661–673. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Maryanovich, M.; Zahalka, A.H.; Pierce, H.; Pinho, S.; Nakahara, F.; Asada, N.; Wei, Q.; Wang, X.; Ciero, P.; Xu, J.; et al. Adrenergic nerve degeneration in bone marrow drives aging of the hematopoietic stem cell niche. Nat. Med. 2018, 24, 782–791. [Google Scholar] [CrossRef] [PubMed]
- Ho, Y.H.; Del Toro, R.; Rivera-Torres, J.; Rak, J.; Korn, C.; Garcia-Garcia, A.; Macias, D.; Gonzalez-Gomez, C.; Del Monte, A.; Wittner, M.; et al. Remodeling of Bone Marrow Hematopoietic Stem Cell Niches Promotes Myeloid Cell Expansion during Premature or Physiological Aging. Cell Stem Cell 2019, 25, 407–418. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bejarano-Garcia, J.A.; Millan-Ucles, A.; Rosado, I.V.; Sanchez-Abarca, L.I.; Caballero-Velazquez, T.; Duran-Galvan, M.J.; Perez-Simon, J.A.; Piruat, J.I. Sensitivity of hematopoietic stem cells to mitochondrial dysfunction by SdhD gene deletion. Cell Death Dis. 2016, 7, e2516. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Anso, E.; Weinberg, S.E.; Diebold, L.P.; Thompson, B.J.; Malinge, S.; Schumacker, P.T.; Liu, X.; Zhang, Y.; Shao, Z.; Steadman, M.; et al. The mitochondrial respiratory chain is essential for haematopoietic stem cell function. Nat. Cell Biol. 2017, 19, 614–625. [Google Scholar] [CrossRef]
- Umemoto, T.; Hashimoto, M.; Matsumura, T.; Nakamura-Ishizu, A.; Suda, T. Ca2+-mitochondria axis drives cell division in hematopoietic stem cells. J. Exp. Med. 2018, 215, 2097–2113. [Google Scholar] [CrossRef]
- Bonora, M.; Ito, K.; Morganti, C.; Pinton, P.; Ito, K. Membrane-potential compensation reveals mitochondrial volume expansion during HSC commitment. Exp. Hematol. 2018, 68, 30–37.e1. [Google Scholar] [CrossRef] [PubMed]
- Morganti, C.; Bonora, M.; Ito, K. Improving the Accuracy of Flow Cytometric Assessment of Mitochondrial Membrane Potential in Hematopoietic Stem and Progenitor Cells Through the Inhibition of Efflux Pumps. J. Vis. Exp. 2019, 149. [Google Scholar] [CrossRef] [PubMed]
- Morganti, C.; Bonora, M.; Ito, K.; Ito, K. Electron transport chain complex II sustains high mitochondrial membrane potential in hematopoietic stem and progenitor cells. Stem Cell Res. 2019, 40, 101573. [Google Scholar] [CrossRef] [PubMed]
- Ishii, N.; Fujii, M.; Hartman, P.S.; Tsuda, M.; Yasuda, K.; Senoo-Matsuda, N.; Yanase, S.; Ayusawa, D.; Suzuki, K. A mutation in succinate dehydrogenase cytochrome b causes oxidative stress and ageing in nematodes. Nature 1998, 394, 694–697. [Google Scholar] [CrossRef] [PubMed]
- Dillin, A.; Hsu, A.L.; Arantes-Oliveira, N.; Lehrer-Graiwer, J.; Hsin, H.; Fraser, A.G.; Kamath, R.S.; Ahringer, J.; Kenyon, C. Rates of behavior and aging specified by mitochondrial function during development. Science 2002, 298, 2398–2401. [Google Scholar] [CrossRef] [PubMed]
- Feng, J.; Bussiere, F.; Hekimi, S. Mitochondrial electron transport is a key determinant of life span in Caenorhabditis elegans. Dev. Cell 2001, 1, 633–644. [Google Scholar] [CrossRef] [Green Version]
- Lee, S.S.; Lee, R.Y.; Fraser, A.G.; Kamath, R.S.; Ahringer, J.; Ruvkun, G. A systematic RNAi screen identifies a critical role for mitochondria in C. elegans longevity. Nat. Genet. 2003, 33, 40–48. [Google Scholar] [CrossRef] [PubMed]
- Kayser, E.B.; Sedensky, M.M.; Morgan, P.G. The effects of complex I function and oxidative damage on lifespan and anesthetic sensitivity in Caenorhabditis elegans. Mech. Ageing Dev. 2004, 125, 455–464. [Google Scholar] [CrossRef] [PubMed]
- Murphy, M.P.; Smith, R.A. Targeting antioxidants to mitochondria by conjugation to lipophilic cations. Annu. Rev. Pharmacol. Toxicol. 2007, 47, 629–656. [Google Scholar] [CrossRef]
- Mansell, E.; Sigurdsson, V.; Deltcheva, E.; Brown, J.; James, C.; Miharada, K.; Soneji, S.; Larsson, J.; Enver, T. Mitochondrial Potentiation Ameliorates Age-Related Heterogeneity in Hematopoietic Stem Cell Function. Cell Stem Cell 2021, 28, 241–256. [Google Scholar] [CrossRef]
- Wilson, A.; Laurenti, E.; Oser, G.; van der Wath, R.C.; Blanco-Bose, W.; Jaworski, M.; Offner, S.; Dunant, C.F.; Eshkind, L.; Bockamp, E.; et al. Hematopoietic stem cells reversibly switch from dormancy to self-renewal during homeostasis and repair. Cell 2008, 135, 1118–1129. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cunningham, J.T.; Rodgers, J.T.; Arlow, D.H.; Vazquez, F.; Mootha, V.K.; Puigserver, P. mTOR controls mitochondrial oxidative function through a YY1-PGC-1alpha transcriptional complex. Nature 2007, 450, 736–740. [Google Scholar] [CrossRef]
- Inoki, K.; Corradetti, M.N.; Guan, K.L. Dysregulation of the TSC-mTOR pathway in human disease. Nat. Genet. 2005, 37, 19–24. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.; Liu, Y.; Liu, R.; Ikenoue, T.; Guan, K.L.; Liu, Y.; Zheng, P. TSC-mTOR maintains quiescence and function of hematopoietic stem cells by repressing mitochondrial biogenesis and reactive oxygen species. J. Exp. Med. 2008, 205, 2397–2408. [Google Scholar] [CrossRef] [Green Version]
- Chen, C.; Liu, Y.; Liu, Y.; Zheng, P. The axis of mTOR-mitochondria-ROS and stemness of the hematopoietic stem cells. Cell Cycle 2009, 8, 1158–1160. [Google Scholar] [CrossRef] [Green Version]
- Hirao, A.; Hoshii, T. Mechanistic/mammalian target protein of rapamycin signaling in hematopoietic stem cells and leukemia. Cancer Sci. 2013, 104, 977–982. [Google Scholar] [CrossRef] [Green Version]
- Wu, F.; Chen, Z.; Liu, J.; Hou, Y. The Akt-mTOR network at the interface of hematopoietic stem cell homeostasis. Exp. Hematol. 2021. online ahead of print. [Google Scholar] [CrossRef]
- Chen, C.; Liu, Y.; Liu, Y.; Zheng, P. mTOR regulation and therapeutic rejuvenation of aging hematopoietic stem cells. Sci. Signal 2009, 2, ra75. [Google Scholar] [CrossRef] [Green Version]
- Fujino, T.; Goyama, S.; Sugiura, Y.; Inoue, D.; Asada, S.; Yamasaki, S.; Matsumoto, A.; Yamaguchi, K.; Isobe, Y.; Tsuchiya, A.; et al. Mutant ASXL1 induces age-related expansion of phenotypic hematopoietic stem cells through activation of Akt/mTOR pathway. Nat. Commun. 2021, 12, 1826. [Google Scholar] [CrossRef] [PubMed]
- Chen, Z.; Yi, W.; Morita, Y.; Wang, H.; Cong, Y.; Liu, J.P.; Xiao, Z.; Rudolph, K.L.; Cheng, T.; Ju, Z. Wip1 deficiency impairs haematopoietic stem cell function via p53 and mTORC1 pathways. Nat. Commun. 2015, 6, 6808. [Google Scholar] [CrossRef] [PubMed]
- Fang, E.F.; Kassahun, H.; Croteau, D.L.; Scheibye-Knudsen, M.; Marosi, K.; Lu, H.; Shamanna, R.A.; Kalyanasundaram, S.; Bollineni, R.C.; Wilson, M.A.; et al. NAD+ Replenishment Improves Lifespan and Healthspan in Ataxia Telangiectasia Models via Mitophagy and DNA Repair. Cell Metab. 2016, 24, 566–581. [Google Scholar] [CrossRef] [Green Version]
- Huang, J.; Nguyen-McCarty, M.; Hexner, E.O.; Danet-Desnoyers, G.; Klein, P.S. Maintenance of hematopoietic stem cells through regulation of Wnt and mTOR pathways. Nat. Med. 2012, 18, 1778–1785. [Google Scholar] [CrossRef] [Green Version]
- Cheung, T.H.; Rando, T.A. Molecular regulation of stem cell quiescence. Nat. Rev. Mol. Cell Biol. 2013, 14, 329–340. [Google Scholar] [CrossRef]
- Reid, M.A.; Dai, Z.; Locasale, J.W. The impact of cellular metabolism on chromatin dynamics and epigenetics. Nat. Cell Biol. 2017, 19, 1298–1306. [Google Scholar] [CrossRef]
- Ito, K.; Suda, T. Metabolic requirements for the maintenance of self-renewing stem cells. Nat. Rev. Mol. Cell Biol. 2014, 15, 243–256. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Qiu, X.; Brown, K.; Hirschey, M.D.; Verdin, E.; Chen, D. Calorie restriction reduces oxidative stress by SIRT3-mediated SOD2 activation. Cell Metab. 2010, 12, 662–667. [Google Scholar] [CrossRef] [Green Version]
- Brown, K.; Xie, S.; Qiu, X.; Mohrin, M.; Shin, J.; Liu, Y.; Zhang, D.; Scadden, D.T.; Chen, D. SIRT3 reverses aging-associated degeneration. Cell Rep. 2013, 3, 319–327. [Google Scholar] [CrossRef] [Green Version]
- Rimmele, P.; Bigarella, C.L.; Liang, R.; Izac, B.; Dieguez-Gonzalez, R.; Barbet, G.; Donovan, M.; Brugnara, C.; Blander, J.M.; Sinclair, D.A.; et al. Aging-like phenotype and defective lineage specification in SIRT1-deleted hematopoietic stem and progenitor cells. Stem Cell Rep. 2014, 3, 44–59. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fang, E.F.; Lautrup, S.; Hou, Y.; Demarest, T.G.; Croteau, D.L.; Mattson, M.P.; Bohr, V.A. NAD+ in Aging: Molecular Mechanisms and Translational Implications. Trends Mol. Med. 2017, 23, 899–916. [Google Scholar] [CrossRef] [PubMed]
- Fang, E.F.; Scheibye-Knudsen, M.; Brace, L.E.; Kassahun, H.; SenGupta, T.; Nilsen, H.; Mitchell, J.R.; Croteau, D.L.; Bohr, V.A. Defective mitophagy in XPA via PARP-1 hyperactivation and NAD+/SIRT1 reduction. Cell 2014, 157, 882–896. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Das, A.; Huang, G.X.; Bonkowski, M.S.; Longchamp, A.; Li, C.; Schultz, M.B.; Kim, L.J.; Osborne, B.; Joshi, S.; Lu, Y.; et al. Impairment of an Endothelial NAD+-H2S Signaling Network Is a Reversible Cause of Vascular Aging. Cell 2018, 173, 74–89.e20. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fang, E.F.; Bohr, V.A. NAD+: The convergence of DNA repair and mitophagy. Autophagy 2017, 13, 442–443. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Valentin-Vega, Y.A.; Maclean, K.H.; Tait-Mulder, J.; Milasta, S.; Steeves, M.; Dorsey, F.C.; Cleveland, J.L.; Green, D.R.; Kastan, M.B. Mitochondrial dysfunction in ataxia-telangiectasia. Blood 2012, 119, 1490–1500. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zong, L.; Tanaka-Yano, M.; Park, B.; Yanai, H.; Turhan, F.T.; Croteau, D.L.; Tian, J.; Fang, E.F.; Bohr, V.A.; Beerman, I. NAD+ augmentation with nicotinamide riboside improves lymphoid potential of Atm−/− and old mice HSCs. NPJ Aging Mech. Dis. 2021, 7, 25. [Google Scholar] [CrossRef]
- Youle, R.J. Mitochondria-Striking a balance between host and endosymbiont. Science 2019, 365, eaaw9855. [Google Scholar] [CrossRef] [PubMed]
- Melber, A.; Haynes, C.M. UPR(mt) regulation and output: A stress response mediated by mitochondrial-nuclear communication. Cell Res. 2018, 28, 281–295. [Google Scholar] [CrossRef] [PubMed]
- Houtkooper, R.H.; Mouchiroud, L.; Ryu, D.; Moullan, N.; Katsyuba, E.; Knott, G.; Williams, R.W.; Auwerx, J. Mitonuclear protein imbalance as a conserved longevity mechanism. Nature 2013, 497, 451–457. [Google Scholar] [CrossRef] [Green Version]
- Mouchiroud, L.; Houtkooper, R.H.; Moullan, N.; Katsyuba, E.; Ryu, D.; Canto, C.; Mottis, A.; Jo, Y.S.; Viswanathan, M.; Schoonjans, K.; et al. The NAD+/Sirtuin Pathway Modulates Longevity through Activation of Mitochondrial UPR and FOXO Signaling. Cell 2013, 154, 430–441. [Google Scholar] [CrossRef] [Green Version]
- Haynes, C.M.; Fiorese, C.J.; Lin, Y.F. Evaluating and responding to mitochondrial dysfunction: The mitochondrial unfolded-protein response and beyond. Trends Cell Biol. 2013, 23, 311–318. [Google Scholar] [CrossRef] [Green Version]
- Berger, E.; Rath, E.; Yuan, D.; Waldschmitt, N.; Khaloian, S.; Allgauer, M.; Staszewski, O.; Lobner, E.M.; Schottl, T.; Giesbertz, P.; et al. Mitochondrial function controls intestinal epithelial stemness and proliferation. Nat. Commun. 2016, 7, 13171. [Google Scholar] [CrossRef] [Green Version]
- Zhang, H.; Ryu, D.; Wu, Y.; Gariani, K.; Wang, X.; Luan, P.; D’Amico, D.; Ropelle, E.R.; Lutolf, M.P.; Aebersold, R.; et al. NAD+ repletion improves mitochondrial and stem cell function and enhances life span in mice. Science 2016, 352, 1436–1443. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mohrin, M.; Widjaja, A.; Liu, Y.; Luo, H.; Chen, D. The mitochondrial unfolded protein response is activated upon hematopoietic stem cell exit from quiescence. Aging Cell 2018, 17, e12756. [Google Scholar] [CrossRef] [PubMed]
- Barber, M.F.; Michishita-Kioi, E.; Xi, Y.; Tasselli, L.; Kioi, M.; Moqtaderi, Z.; Tennen, R.I.; Paredes, S.; Young, N.L.; Chen, K.; et al. SIRT7 links H3K18 deacetylation to maintenance of oncogenic transformation. Nature 2012, 487, 114–118. [Google Scholar] [CrossRef] [PubMed]
- Salemi, S.; Yousefi, S.; Constantinescu, M.A.; Fey, M.F.; Simon, H.U. Autophagy is required for self-renewal and differentiation of adult human stem cells. Cell Res. 2012, 22, 432–435. [Google Scholar] [CrossRef]
- Ito, K.; Turcotte, R.; Cui, J.; Zimmerman, S.E.; Pinho, S.; Mizoguchi, T.; Arai, F.; Runnels, J.M.; Alt, C.; Teruya-Feldstein, J.; et al. Self-renewal of a purified Tie2+ hematopoietic stem cell population relies on mitochondrial clearance. Science 2016, 354, 1156–1160. [Google Scholar] [CrossRef] [Green Version]
- Ho, T.T.; Warr, M.R.; Adelman, E.R.; Lansinger, O.M.; Flach, J.; Verovskaya, E.V.; Figueroa, M.E.; Passegue, E. Autophagy maintains the metabolism and function of young and old stem cells. Nature 2017, 543, 205–210. [Google Scholar] [CrossRef] [Green Version]
- Vannini, N.; Girotra, M.; Naveiras, O.; Nikitin, G.; Campos, V.; Giger, S.; Roch, A.; Auwerx, J.; Lutolf, M.P. Specification of haematopoietic stem cell fate via modulation of mitochondrial activity. Nat. Commun. 2016, 7, 13125. [Google Scholar] [CrossRef]
- Liang, R.; Arif, T.; Kalmykova, S.; Kasianov, A.; Lin, M.; Menon, V.; Qiu, J.; Bernitz, J.M.; Moore, K.; Lin, F.; et al. Restraining Lysosomal Activity Preserves Hematopoietic Stem Cell Quiescence and Potency. Cell Stem Cell 2020, 26, 359–376. [Google Scholar] [CrossRef]
- Garcia-Prat, L.; Martinez-Vicente, M.; Perdiguero, E.; Ortet, L.; Rodriguez-Ubreva, J.; Rebollo, E.; Ruiz-Bonilla, V.; Gutarra, S.; Ballestar, E.; Serrano, A.L.; et al. Autophagy maintains stemness by preventing senescence. Nature 2016, 529, 37–42. [Google Scholar] [CrossRef] [PubMed]
- Luchsinger, L.L.; de Almeida, M.J.; Corrigan, D.J.; Mumau, M.; Snoeck, H.W. Mitofusin 2 maintains haematopoietic stem cells with extensive lymphoid potential. Nature 2016, 529, 528–531. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Morganti, C.; Missiroli, S.; Lebiedzinska-Arciszewska, M.; Ferroni, L.; Morganti, L.; Perrone, M.; Ramaccini, D.; Occhionorelli, S.; Zavan, B.; Wieckowski, M.R.; et al. Regulation of PKCbeta levels and autophagy by PML is essential for high-glucose-dependent mesenchymal stem cell adipogenesis. Int. J. Obes. 2019, 43, 963–973. [Google Scholar] [CrossRef] [PubMed]
- Morganti, C.; Bonora, M.; Marchi, S.; Ferroni, L.; Gardin, C.; Wieckowski, M.R.; Giorgi, C.; Pinton, P.; Zavan, B. Citrate Mediates Crosstalk between Mitochondria and the Nucleus to Promote Human Mesenchymal Stem Cell In Vitro Osteogenesis. Cells 2020, 9, 1034. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Youle, R.J.; van der Bliek, A.M. Mitochondrial fission, fusion, and stress. Science 2012, 337, 1062–1065. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rana, A.; Oliveira, M.P.; Khamoui, A.V.; Aparicio, R.; Rera, M.; Rossiter, H.B.; Walker, D.W. Promoting Drp1-mediated mitochondrial fission in midlife prolongs healthy lifespan of Drosophila melanogaster. Nat. Commun. 2017, 8, 448. [Google Scholar] [CrossRef]
- Aparicio, R.; Rana, A.; Walker, D.W. Upregulation of the Autophagy Adaptor p62/SQSTM1 Prolongs Health and Lifespan in Middle-Aged Drosophila. Cell Rep. 2019, 28, 1029–1040.e5. [Google Scholar] [CrossRef] [Green Version]
- Todd, A.M.; Staveley, B.E. Expression of Pink1 with alpha-synuclein in the dopaminergic neurons of Drosophila leads to increases in both lifespan and healthspan. Genet. Mol. Res. 2012, 11, 1497–1502. [Google Scholar] [CrossRef] [Green Version]
- Rana, A.; Rera, M.; Walker, D.W. Parkin overexpression during aging reduces proteotoxicity, alters mitochondrial dynamics, and extends lifespan. Proc. Natl. Acad. Sci. USA 2013, 110, 8638–8643. [Google Scholar] [CrossRef] [Green Version]
- Palikaras, K.; Lionaki, E.; Tavernarakis, N. Coordination of mitophagy and mitochondrial biogenesis during ageing in C. elegans. Nature 2015, 521, 525–528. [Google Scholar] [CrossRef]
- Schiavi, A.; Maglioni, S.; Palikaras, K.; Shaik, A.; Strappazzon, F.; Brinkmann, V.; Torgovnick, A.; Castelein, N.; De Henau, S.; Braeckman, B.P.; et al. Iron-Starvation-Induced Mitophagy Mediates Lifespan Extension upon Mitochondrial Stress in C. elegans. Curr. Biol. 2015, 25, 1810–1822. [Google Scholar] [CrossRef] [Green Version]
- Franceschi, C.; Campisi, J. Chronic inflammation (inflammaging) and its potential contribution to age-associated diseases. J. Gerontol. Ser. A Biol. Sci. Med. Sci. 2014, 69, S4–S9. [Google Scholar] [CrossRef]
- Franceschi, C.; Capri, M.; Monti, D.; Giunta, S.; Olivieri, F.; Sevini, F.; Panourgia, M.P.; Invidia, L.; Celani, L.; Scurti, M.; et al. Inflammaging and anti-inflammaging: A systemic perspective on aging and longevity emerged from studies in humans. Mech. Ageing Dev. 2007, 128, 92–105. [Google Scholar] [CrossRef]
- Pietras, E.M. Inflammation: A key regulator of hematopoietic stem cell fate in health and disease. Blood 2017, 130, 1693–1698. [Google Scholar] [CrossRef] [Green Version]
- Manz, M.G.; Boettcher, S. Emergency granulopoiesis. Nat. Rev. Immunol. 2014, 14, 302–314. [Google Scholar] [CrossRef]
- Zhao, J.L.; Ma, C.; O’Connell, R.M.; Mehta, A.; DiLoreto, R.; Heath, J.R.; Baltimore, D. Conversion of danger signals into cytokine signals by hematopoietic stem and progenitor cells for regulation of stress-induced hematopoiesis. Cell Stem Cell 2014, 14, 445–459. [Google Scholar] [CrossRef] [Green Version]
- Essers, M.A.; Offner, S.; Blanco-Bose, W.E.; Waibler, Z.; Kalinke, U.; Duchosal, M.A.; Trumpp, A. IFNalpha activates dormant haematopoietic stem cells In Vivo. Nature 2009, 458, 904–908. [Google Scholar] [CrossRef]
- Baldridge, M.T.; King, K.Y.; Boles, N.C.; Weksberg, D.C.; Goodell, M.A. Quiescent haematopoietic stem cells are activated by IFN-gamma in response to chronic infection. Nature 2010, 465, 793–797. [Google Scholar] [CrossRef] [PubMed]
- Pietras, E.M.; Mirantes-Barbeito, C.; Fong, S.; Loeffler, D.; Kovtonyuk, L.V.; Zhang, S.; Lakshminarasimhan, R.; Chin, C.P.; Techner, J.M.; Will, B.; et al. Chronic interleukin-1 exposure drives haematopoietic stem cells towards precocious myeloid differentiation at the expense of self-renewal. Nat. Cell Biol. 2016, 18, 607–618. [Google Scholar] [CrossRef] [PubMed]
- Coppe, J.P.; Desprez, P.Y.; Krtolica, A.; Campisi, J. The senescence-associated secretory phenotype: The dark side of tumor suppression. Annu. Rev. Pathol. 2010, 5, 99–118. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bousounis, P.; Bergo, V.; Trompouki, E. Inflammation, Aging and Hematopoiesis: A Complex Relationship. Cells 2021, 10, 1386. [Google Scholar] [CrossRef]
- Bektas, A.; Schurman, S.H.; Gonzalez-Freire, M.; Dunn, C.A.; Singh, A.K.; Macian, F.; Cuervo, A.M.; Sen, R.; Ferrucci, L. Age-associated changes in human CD4+ T cells point to mitochondrial dysfunction consequent to impaired autophagy. Aging 2019, 11, 9234–9263. [Google Scholar] [CrossRef]
- Callender, L.A.; Carroll, E.C.; Bober, E.A.; Akbar, A.N.; Solito, E.; Henson, S.M. Mitochondrial mass governs the extent of human T cell senescence. Aging Cell 2020, 19, e13067. [Google Scholar] [CrossRef] [Green Version]
- Chen, Z.; Amro, E.M.; Becker, F.; Holzer, M.; Rasa, S.M.M.; Njeru, S.N.; Han, B.; Di Sanzo, S.; Chen, Y.; Tang, D.; et al. Cohesin-mediated NF-kappaB signaling limits hematopoietic stem cell self-renewal in aging and inflammation. J. Exp. Med. 2019, 216, 152–175. [Google Scholar] [CrossRef]
- He, H.; Xu, P.; Zhang, X.; Liao, M.; Dong, Q.; Cong, T.; Tang, B.; Yang, X.; Ye, M.; Chang, Y.; et al. Aging-induced IL27Ra signaling impairs hematopoietic stem cells. Blood 2020, 136, 183–198. [Google Scholar] [CrossRef] [PubMed]
- Jones, D.L.; Rando, T.A. Emerging models and paradigms for stem cell ageing. Nat. Cell Biol. 2011, 13, 506–512. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Krysko, D.V.; Agostinis, P.; Krysko, O.; Garg, A.D.; Bachert, C.; Lambrecht, B.N.; Vandenabeele, P. Emerging role of damage-associated molecular patterns derived from mitochondria in inflammation. Trends Immunol. 2011, 32, 157–164. [Google Scholar] [CrossRef] [PubMed]
- Collins, L.V.; Hajizadeh, S.; Holme, E.; Jonsson, I.M.; Tarkowski, A. Endogenously oxidized mitochondrial DNA induces in vivo and in vitro inflammatory responses. J. Leukoc. Biol. 2004, 75, 995–1000. [Google Scholar] [CrossRef] [PubMed]
- Zhou, R.; Yazdi, A.S.; Menu, P.; Tschopp, J. A role for mitochondria in NLRP3 inflammasome activation. Nature 2011, 469, 221–225. [Google Scholar] [CrossRef] [PubMed]
- Zitvogel, L.; Kepp, O.; Galluzzi, L.; Kroemer, G. Inflammasomes in carcinogenesis and anticancer immune responses. Nat. Immunol. 2012, 13, 343–351. [Google Scholar] [CrossRef]
- Alfonso-Loeches, S.; Urena-Peralta, J.R.; Morillo-Bargues, M.J.; Oliver-De La Cruz, J.; Guerri, C. Role of mitochondria ROS generation in ethanol-induced NLRP3 inflammasome activation and cell death in astroglial cells. Front. Cell Neurosci. 2014, 8, 216. [Google Scholar] [CrossRef]
- Luo, H.; Mu, W.C.; Karki, R.; Chiang, H.H.; Mohrin, M.; Shin, J.J.; Ohkubo, R.; Ito, K.; Kanneganti, T.D.; Chen, D. Mitochondrial Stress-Initiated Aberrant Activation of the NLRP3 Inflammasome Regulates the Functional Deterioration of Hematopoietic Stem Cell Aging. Cell Rep. 2019, 26, 945–954. [Google Scholar] [CrossRef] [Green Version]
- Guo, H.; Callaway, J.B.; Ting, J.P. Inflammasomes: Mechanism of action, role in disease, and therapeutics. Nat. Med. 2015, 21, 677–687. [Google Scholar] [CrossRef] [Green Version]
- He, M.; Chiang, H.H.; Luo, H.; Zheng, Z.; Qiao, Q.; Wang, L.; Tan, M.; Ohkubo, R.; Mu, W.C.; Zhao, S.; et al. An Acetylation Switch of the NLRP3 Inflammasome Regulates Aging-Associated Chronic Inflammation and Insulin Resistance. Cell Metab. 2020, 31, 580–591. [Google Scholar] [CrossRef]
- Pinti, M.; Cevenini, E.; Nasi, M.; De Biasi, S.; Salvioli, S.; Monti, D.; Benatti, S.; Gibellini, L.; Cotichini, R.; Stazi, M.A.; et al. Circulating mitochondrial DNA increases with age and is a familiar trait: Implications for “inflamm-aging”. Eur. J. Immunol. 2014, 44, 1552–1562. [Google Scholar] [CrossRef]
- Anderson, S.; Bankier, A.T.; Barrell, B.G.; de Bruijn, M.H.; Coulson, A.R.; Drouin, J.; Eperon, I.C.; Nierlich, D.P.; Roe, B.A.; Sanger, F.; et al. Sequence and organization of the human mitochondrial genome. Nature 1981, 290, 457–465. [Google Scholar] [CrossRef] [PubMed]
- Trifunovic, A.; Larsson, N.G. Mitochondrial dysfunction as a cause of ageing. J. Intern. Med. 2008, 263, 167–178. [Google Scholar] [CrossRef] [PubMed]
- Larsson, N.G. Somatic mitochondrial DNA mutations in mammalian aging. Annu. Rev. Biochem. 2010, 79, 683–706. [Google Scholar] [CrossRef] [PubMed]
- Seo, A.Y.; Joseph, A.M.; Dutta, D.; Hwang, J.C.; Aris, J.P.; Leeuwenburgh, C. New insights into the role of mitochondria in aging: Mitochondrial dynamics and more. J. Cell Sci. 2010, 123, 2533–2542. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Beckman, K.B.; Ames, B.N. Endogenous oxidative damage of mtDNA. Mutat. Res. 1999, 424, 51–58. [Google Scholar] [CrossRef]
- Haag-Liautard, C.; Coffey, N.; Houle, D.; Lynch, M.; Charlesworth, B.; Keightley, P.D. Direct estimation of the mitochondrial DNA mutation rate in Drosophila melanogaster. PLoS Biol. 2008, 6, e204. [Google Scholar] [CrossRef] [Green Version]
- Radzvilavicius, A.L.; Hadjivasiliou, Z.; Pomiankowski, A.; Lane, N. Selection for Mitochondrial Quality Drives Evolution of the Germline. PLoS Biol. 2016, 14, e2000410. [Google Scholar] [CrossRef] [PubMed]
- Mengel-From, J.; Thinggaard, M.; Dalgard, C.; Kyvik, K.O.; Christensen, K.; Christiansen, L. Mitochondrial DNA copy number in peripheral blood cells declines with age and is associated with general health among elderly. Hum. Genet. 2014, 133, 1149–1159. [Google Scholar] [CrossRef] [Green Version]
- Knez, J.; Winckelmans, E.; Plusquin, M.; Thijs, L.; Cauwenberghs, N.; Gu, Y.; Staessen, J.A.; Nawrot, T.S.; Kuznetsova, T. Correlates of Peripheral Blood Mitochondrial DNA Content in a General Population. Am. J. Epidemiol. 2016, 183, 138–146. [Google Scholar] [CrossRef] [Green Version]
- Sondheimer, N.; Glatz, C.E.; Tirone, J.E.; Deardorff, M.A.; Krieger, A.M.; Hakonarson, H. Neutral mitochondrial heteroplasmy and the influence of aging. Hum. Mol. Genet. 2011, 20, 1653–1659. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, M.; Rothwell, R.; Vermaat, M.; Wachsmuth, M.; Schroder, R.; Laros, J.F.; van Oven, M.; de Bakker, P.I.; Bovenberg, J.A.; van Duijn, C.M.; et al. Transmission of human mtDNA heteroplasmy in the Genome of the Netherlands families: Support for a variable-size bottleneck. Genome Res. 2016, 26, 417–426. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tanaka, M.; Gong, J.S.; Zhang, J.; Yoneda, M.; Yagi, K. Mitochondrial genotype associated with longevity. Lancet 1998, 351, 185–186. [Google Scholar] [CrossRef]
- Takagi, K.; Yamada, Y.; Gong, J.S.; Sone, T.; Yokota, M.; Tanaka, M. Association of a 5178C-->A (Leu237Met) polymorphism in the mitochondrial DNA with a low prevalence of myocardial infarction in Japanese individuals. Atherosclerosis 2004, 175, 281–286. [Google Scholar] [CrossRef] [PubMed]
- Yu, X.; Gimsa, U.; Wester-Rosenlof, L.; Kanitz, E.; Otten, W.; Kunz, M.; Ibrahim, S.M. Dissecting the effects of mtDNA variations on complex traits using mouse conplastic strains. Genome Res. 2009, 19, 159–165. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kretzschmar, C.; Roolf, C.; Timmer, K.; Sekora, A.; Knubel, G.; Murua Escobar, H.; Fuellen, G.; Ibrahim, S.M.; Tiedge, M.; Baltrusch, S.; et al. Polymorphisms of the murine mitochondrial ND4, CYTB and COX3 genes impact hematopoiesis during aging. Oncotarget 2016, 7, 74460–74472. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Taylor, R.W.; Barron, M.J.; Borthwick, G.M.; Gospel, A.; Chinnery, P.F.; Samuels, D.C.; Taylor, G.A.; Plusa, S.M.; Needham, S.J.; Greaves, L.C.; et al. Mitochondrial DNA mutations in human colonic crypt stem cells. J. Clin. Investig. 2003, 112, 1351–1360. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Norddahl, G.L.; Pronk, C.J.; Wahlestedt, M.; Sten, G.; Nygren, J.M.; Ugale, A.; Sigvardsson, M.; Bryder, D. Accumulating mitochondrial DNA mutations drive premature hematopoietic aging phenotypes distinct from physiological stem cell aging. Cell Stem Cell 2011, 8, 499–510. [Google Scholar] [CrossRef] [Green Version]
- Ahlqvist, K.J.; Hamalainen, R.H.; Yatsuga, S.; Uutela, M.; Terzioglu, M.; Gotz, A.; Forsstrom, S.; Salven, P.; Angers-Loustau, A.; Kopra, O.H.; et al. Somatic progenitor cell vulnerability to mitochondrial DNA mutagenesis underlies progeroid phenotypes in Polg mutator mice. Cell Metab. 2012, 15, 100–109. [Google Scholar] [CrossRef] [Green Version]
- Ahlqvist, K.J.; Leoncini, S.; Pecorelli, A.; Wortmann, S.B.; Ahola, S.; Forsstrom, S.; Guerranti, R.; De Felice, C.; Smeitink, J.; Ciccoli, L.; et al. MtDNA mutagenesis impairs elimination of mitochondria during erythroid maturation leading to enhanced erythrocyte destruction. Nat. Commun. 2015, 6, 6494. [Google Scholar] [CrossRef]
- Kujoth, G.C.; Hiona, A.; Pugh, T.D.; Someya, S.; Panzer, K.; Wohlgemuth, S.E.; Hofer, T.; Seo, A.Y.; Sullivan, R.; Jobling, W.A.; et al. Mitochondrial DNA mutations, oxidative stress, and apoptosis in mammalian aging. Science 2005, 309, 481–484. [Google Scholar] [CrossRef] [PubMed]
- Trifunovic, A.; Wredenberg, A.; Falkenberg, M.; Spelbrink, J.N.; Rovio, A.T.; Bruder, C.E.; Bohlooly, Y.M.; Gidlof, S.; Oldfors, A.; Wibom, R.; et al. Premature ageing in mice expressing defective mitochondrial DNA polymerase. Nature 2004, 429, 417–423. [Google Scholar] [CrossRef]
- Edgar, D.; Shabalina, I.; Camara, Y.; Wredenberg, A.; Calvaruso, M.A.; Nijtmans, L.; Nedergaard, J.; Cannon, B.; Larsson, N.G.; Trifunovic, A. Random point mutations with major effects on protein-coding genes are the driving force behind premature aging in mtDNA mutator mice. Cell Metab. 2009, 10, 131–138. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Trifunovic, A.; Hansson, A.; Wredenberg, A.; Rovio, A.T.; Dufour, E.; Khvorostov, I.; Spelbrink, J.N.; Wibom, R.; Jacobs, H.T.; Larsson, N.G. Somatic mtDNA mutations cause aging phenotypes without affecting reactive oxygen species production. Proc. Natl. Acad. Sci. USA 2005, 102, 17993–17998. [Google Scholar] [CrossRef] [Green Version]
- Chen, M.L.; Logan, T.D.; Hochberg, M.L.; Shelat, S.G.; Yu, X.; Wilding, G.E.; Tan, W.; Kujoth, G.C.; Prolla, T.A.; Selak, M.A.; et al. Erythroid dysplasia, megaloblastic anemia, and impaired lymphopoiesis arising from mitochondrial dysfunction. Blood 2009, 114, 4045–4053. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rossmann, M.P.; Dubois, S.M.; Agarwal, S.; Zon, L.I. Mitochondrial function in development and disease. Dis. Models Mech. 2021, 14, dmm048912. [Google Scholar] [CrossRef]
- Kornblum, C.; Nicholls, T.J.; Haack, T.B.; Scholer, S.; Peeva, V.; Danhauser, K.; Hallmann, K.; Zsurka, G.; Rorbach, J.; Iuso, A.; et al. Loss-of-function mutations in MGME1 impair mtDNA replication and cause multisystemic mitochondrial disease. Nat. Genet. 2013, 45, 214–219. [Google Scholar] [CrossRef] [Green Version]
- Matic, S.; Jiang, M.; Nicholls, T.J.; Uhler, J.P.; Dirksen-Schwanenland, C.; Polosa, P.L.; Simard, M.L.; Li, X.; Atanassov, I.; Rackham, O.; et al. Mice lacking the mitochondrial exonuclease MGME1 accumulate mtDNA deletions without developing progeria. Nat. Commun. 2018, 9, 1202. [Google Scholar] [CrossRef] [Green Version]
- Rossi, D.J.; Bryder, D.; Weissman, I.L. Hematopoietic stem cell aging: Mechanism and consequence. Exp. Gerontol. 2007, 42, 385–390. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Liang, R.; Ghaffari, S. Stem cells, redox signaling, and stem cell aging. Antioxid. Redox. Signal. 2014, 20, 1902–1916. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Filippi, M.D.; Ghaffari, S. Mitochondria in the maintenance of hematopoietic stem cells: New perspectives and opportunities. Blood 2019, 133, 1943–1952. [Google Scholar] [CrossRef] [PubMed]
- Ludwig, L.S.; Lareau, C.A.; Ulirsch, J.C.; Christian, E.; Muus, C.; Li, L.H.; Pelka, K.; Ge, W.; Oren, Y.; Brack, A.; et al. Lineage Tracing in Humans Enabled by Mitochondrial Mutations and Single-Cell Genomics. Cell 2019, 176, 1325–1339. [Google Scholar] [CrossRef] [Green Version]
- Lareau, C.A.; Ludwig, L.S.; Muus, C.; Gohil, S.H.; Zhao, T.; Chiang, Z.; Pelka, K.; Verboon, J.M.; Luo, W.; Christian, E.; et al. Massively parallel single-cell mitochondrial DNA genotyping and chromatin profiling. Nat. Biotechnol. 2021, 39, 451–461. [Google Scholar] [CrossRef]
- Jaiswal, S.; Fontanillas, P.; Flannick, J.; Manning, A.; Grauman, P.V.; Mar, B.G.; Lindsley, R.C.; Mermel, C.H.; Burtt, N.; Chavez, A.; et al. Age-related clonal hematopoiesis associated with adverse outcomes. N. Engl. J. Med. 2014, 371, 2488–2498. [Google Scholar] [CrossRef] [Green Version]
- Chen, J.; Kao, Y.R.; Sun, D.; Todorova, T.I.; Reynolds, D.; Narayanagari, S.R.; Montagna, C.; Will, B.; Verma, A.; Steidl, U. Myelodysplastic syndrome progression to acute myeloid leukemia at the stem cell level. Nat. Med. 2019, 25, 103–110. [Google Scholar] [CrossRef]
- Abelson, S.; Collord, G.; Ng, S.W.K.; Weissbrod, O.; Mendelson Cohen, N.; Niemeyer, E.; Barda, N.; Zuzarte, P.C.; Heisler, L.; Sundaravadanam, Y.; et al. Prediction of acute myeloid leukaemia risk in healthy individuals. Nature 2018, 559, 400–404. [Google Scholar] [CrossRef]
- Wu, H.C.; Rerolle, D.; Berthier, C.; Hleihel, R.; Sakamoto, T.; Quentin, S.; Benhenda, S.; Morganti, C.; Wu, C.; Conte, L.; et al. Actinomycin D targets NPM1c-primed mitochondria to restore PML-driven senescence in AML therapy. Cancer Discov. 2021. online ahead of print. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).