Successful Treatment of Persistent SARS-CoV-2 Infection in a B-Cell Depleted Patient with Activated Cytotoxic T and NK Cells: A Case Report
Abstract
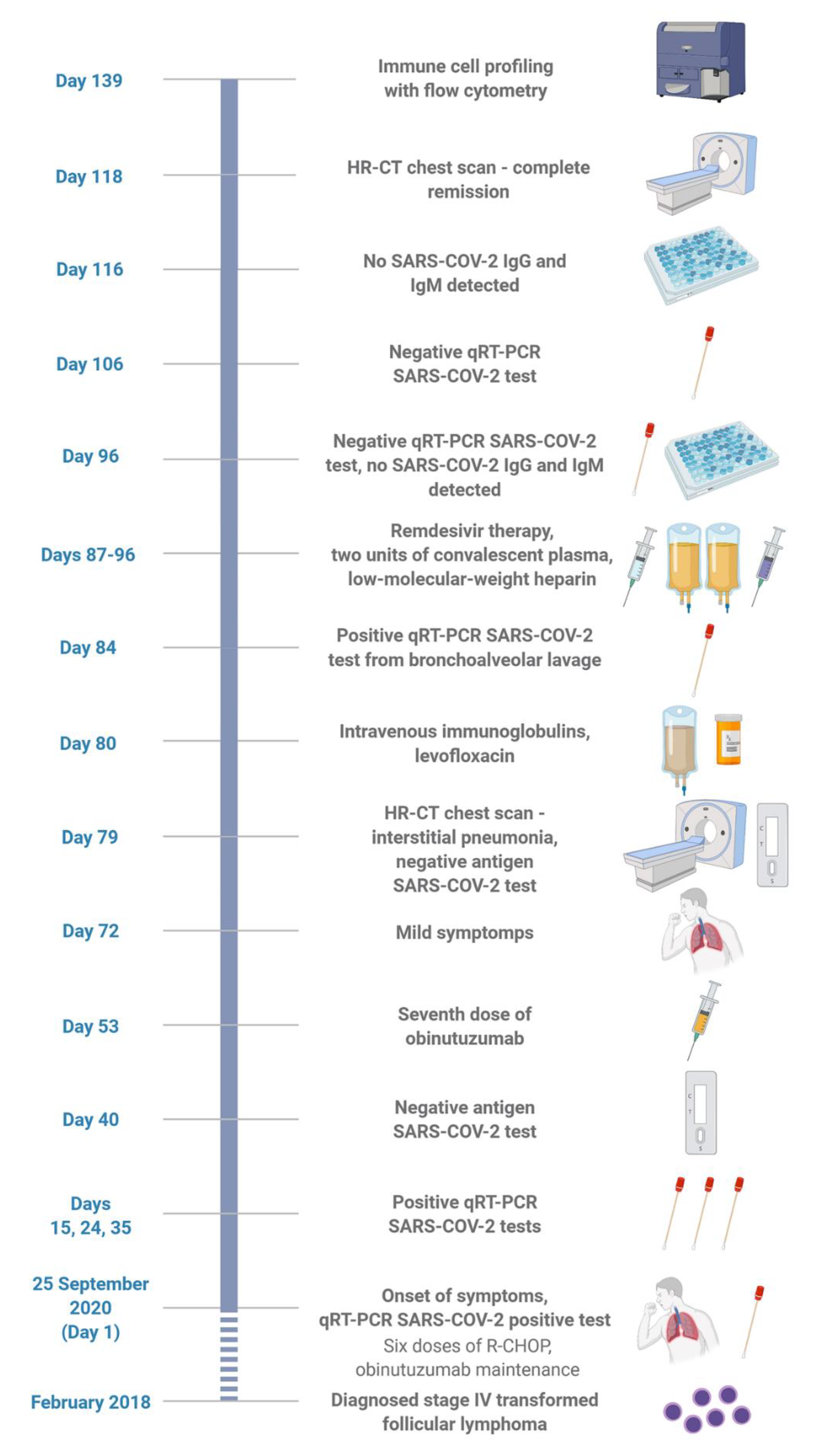
:1. Introduction
2. Results
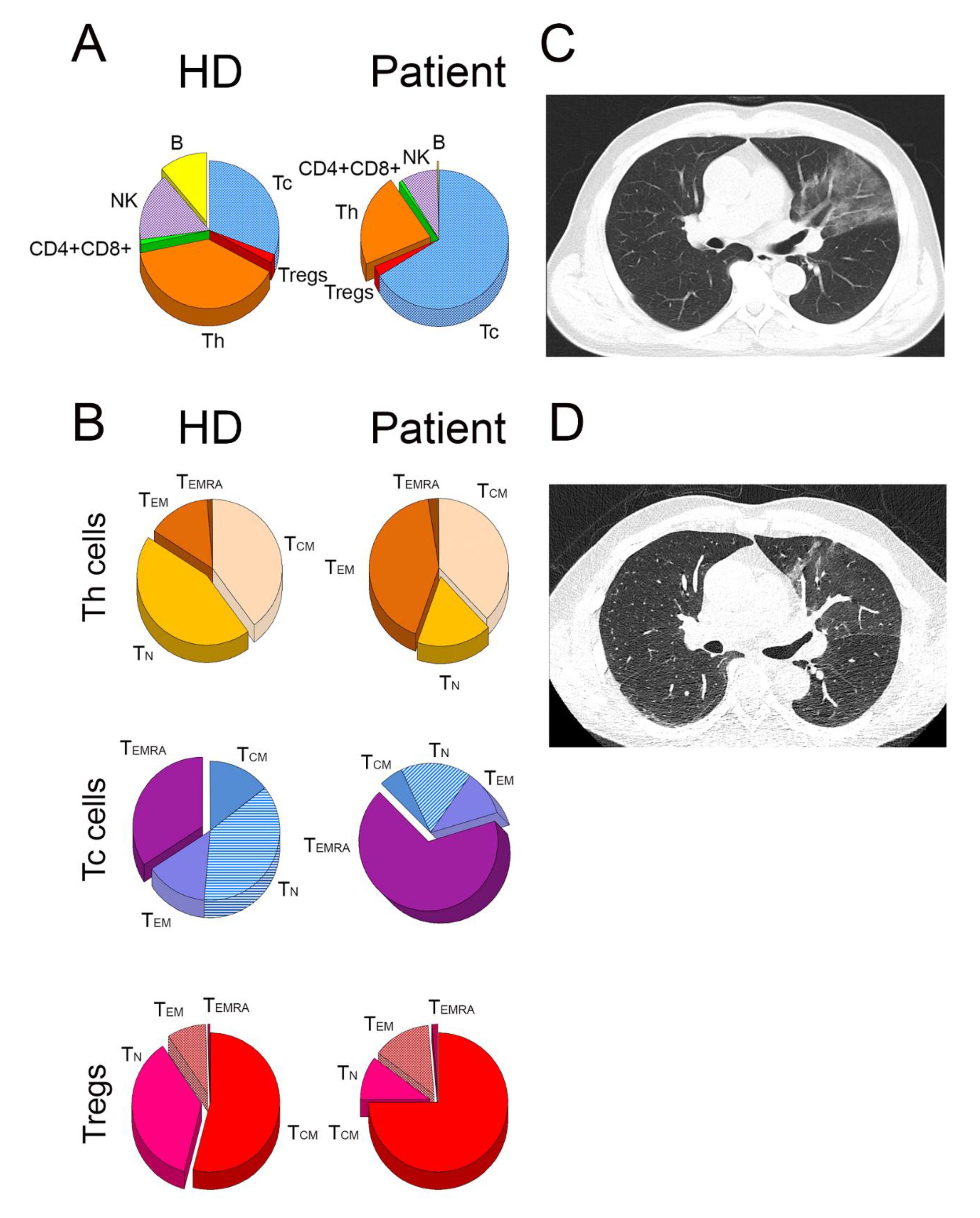
2.1. Distribution of Lymphocyte Subsets
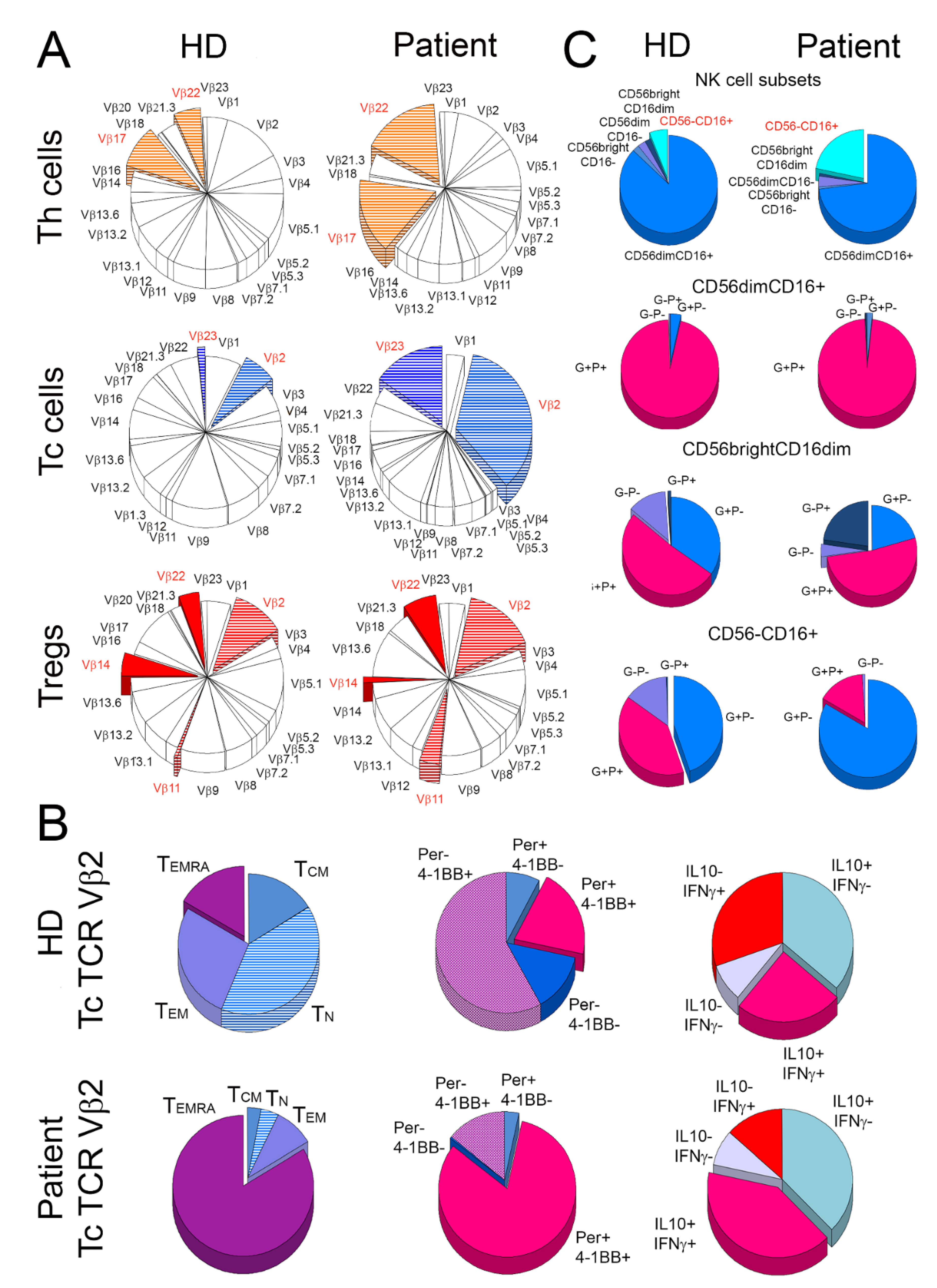
2.2. Expansion and Activation of TCR Vβ2 Clones of Tc Cells
2.3. Alterations in the Distribution of NK Cells and Their Activation Status
3. Discussion
4. Materials and Methods
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Jee, J.; Foote, M.B.; Lumish, M.; Stonestrom, A.J.; Wills, B.; Narendra, V.; Avutu, V.; Murciano-Goroff, Y.R.; Chan, J.E.; Derkach, A.; et al. Chemotherapy and COVID-19 Outcomes in Patients with Cancer. J. Clin. Oncol. 2020, 38, 3538–3546. [Google Scholar] [CrossRef]
- Avanzato, V.A.; Matson, M.J.; Seifert, S.N.; Pryce, R.; Williamson, B.N.; Anzick, S.L.; Barbian, K.; Judson, S.D.; Fischer, E.R.; Martens, C.; et al. Case Study: Prolonged Infectious SARS-CoV-2 Shedding from an Asymptomatic Immunocompromised Individual with Cancer. Cell 2020, 183, 1901–1912. [Google Scholar] [CrossRef] [PubMed]
- Vinay, D.S.; Kwon, B.S. 4-1BB (CD137), an inducible costimulatory receptor, as a specific target for cancer therapy. BMB Rep. 2014, 47, 122–129. [Google Scholar] [CrossRef] [Green Version]
- Poli, A.; Michel, T.; Thérésine, M.; Andrès, E.; Hentges, F.; Zimmer, J. CD56 bright natural killer (NK) cells: An important NK cell subset. Immunology 2009, 126, 458–465. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Zhang, D.; Du, G.; Du, R.; Zhao, J.; Jin, Y.; Fu, S.; Gao, L.; Cheng, Z.; Lu, Q.; et al. Remdesivir in adults with severe COVID-19: A randomised, double-blind, placebo-controlled, multicentre trial. Lancet 2020, 395, 1569–1578. [Google Scholar] [CrossRef]
- Joyner, M.J.; Senefeld, J.W.; Klassen, S.A.; Mills, J.R.; Johnson, P.W.; Theel, E.S.; Wiggins, C.C.; Bruno, K.A.; Klompas, A.M.; Lesser, E.R.; et al. Effect of convalescent plasma on mortality among hospitalized patients with COVID-19: Initial three-month experience. medRxiv 2020, 1–31. [Google Scholar] [CrossRef]
- Beigel, J.H.; Tomashek, K.M.; Dodd, L.E.; Mehta, A.K.; Zingman, B.S.; Kalil, A.C.; Hohmann, E.; Chu, H.Y.; Luetkemeyer, A.; Kline, S.; et al. Remdesivir for the Treatment of Covid-19—Final Report. N. Engl. J. Med. 2020, 383, 1813–1826. [Google Scholar] [CrossRef]
- Abdul-Jawad, S.; Baù, L.; Alaguthurai, T.; del Molino del Barrio, I.; Laing, A.G.; Hayday, T.S.; Monin, L.; Muñoz-Ruiz, M.; McDonald, L.; Francos Quijorna, I.; et al. Acute Immune Signatures and Their Legacies in Severe Acute Respiratory Syndrome Coronavirus-2 Infected Cancer Patients. Cancer Cell 2021, 39, 257–275. [Google Scholar] [CrossRef]
- Helleberg, M.; Niemann, C.U.; Moestrup, K.S.; Kirk, O.; Lebech, A.M.; Lane, C.; Lundgren, J. Persistent COVID-19 in an immunocompromised patient temporarily responsive to two courses of remdesivir therapy. J. Infect. Dis. 2020, 222, 1103–1107. [Google Scholar] [CrossRef]
- Martinot, M.; Jary, A.; Fafi-Kremer, S.; Leducq, V.; Delagreverie, H.; Garnier, M.; Pacanowski, J.; Mékinian, A.; Pirenne, F.; Tiberghien, P.; et al. Emerging RNA-Dependent RNA Polymerase Mutation in a Remdesivir-Treated B-cell Immunodeficient Patient With Protracted Coronavirus Disease 2019. Clin. Infect. Dis. 2021, 73, e1762–e1765. [Google Scholar] [CrossRef]
- Camprubí, D.; Gaya, A.; Marcos, M.A.; Martí-Soler, H.; Soriano, A.; del Mosquera, M.M.; Oliver, A.; Santos, M.; Muñoz, J.; García-Vidal, C. Persistent replication of SARS-CoV-2 in a severely immunocompromised patient treated with several courses of remdesivir. Int. J. Infect. Dis. 2021, 104, 379–381. [Google Scholar] [CrossRef]
- Malsy, J.; Veletzky, L.; Heide, J.; Hennigs, A.; Gil-Ibanez, I.; Stein, A.; Lütgehetmann, M.; Rosien, U.; Jasper, D.; Peine, S.; et al. Sustained Response after Remdesivir and Convalescent Plasma Therapy in a B-Cell–Depleted Patient With Protracted Coronavirus Disease 2019 (COVID-19). Clin. Infect. Dis. 2020, 1–7. [Google Scholar] [CrossRef]
- Buckland, M.S.; Galloway, J.B.; Fhogartaigh, C.N.; Meredith, L.; Provine, N.M.; Bloor, S.; Ogbe, A.; Zelek, W.M.; Smielewska, A.; Yakovleva, A.; et al. Treatment of COVID-19 with remdesivir in the absence of humoral immunity: A case report. Nat. Commun. 2020, 11, 6385. [Google Scholar] [CrossRef] [PubMed]
- Baang, J.H.; Smith, C.; Mirabelli, C.; Valesano, A.L.; Manthei, D.M.; Bachman, M.A.; Wobus, C.E.; Adams, M.; Washer, L.; Martin, E.T.; et al. Prolonged Severe Acute Respiratory Syndrome Coronavirus 2 Replication in an Immunocompromised Patient. J. Infect. Dis. 2021, 223, 23–27. [Google Scholar] [CrossRef] [PubMed]
- Kowalik, M.M.; Trzonkowski, P.; Łasińska-Kowara, M.; Mital, A.; Smiatacz, T.; Jaguszewski, M. COVID-19—Toward a comprehensive understanding of the disease. Cardiol. J. 2020, 27, 99–114. [Google Scholar] [CrossRef] [PubMed]
- Tian, Y.; Babor, M.; Lane, J.; Schulten, V.; Patil, V.S.; Seumois, G.; Rosales, S.L.; Fu, Z.; Picarda, G.; Burel, J.; et al. Unique phenotypes and clonal expansions of human CD4 effector memory T cells re-expressing CD45RA. Nat. Commun. 2017, 8, 1473. [Google Scholar] [CrossRef]
- Meraviglia, S.; Di Carlo, P.; Pampinella, D.; Guadagnino, G.; Presti, E.L.; Orlando, V.; Marchetti, G.; Dieli, F.; Sergi, C. T-cell subsets (TCM, TEM, TEMRA) and poly-functional immune response in patients with human immunodeficiency virus (HIV) infection and different T-CD4 cell response. Ann. Clin. Lab. Sci. 2019, 49, 519–528. [Google Scholar] [PubMed]
- Norfield, J.W.; Loo, C.P.; Barbour, J.D.; Spotts, G.; Hecht, F.M.; Klenerman, P.; Nixon, D.F.; Michaëlsson, J. Human immunodeficiency virus type 1 (HIV-1)-specific CD8+ T(EMRA) cells in early infection are linked to control of HIV-1 viremia and predict the subsequent viral load set point. J. Virol. 2007, 81, 5759–5765. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gaitzsch, E.; Passerini, V.; Khatamzas, E.; Strobl, C.D.; Muenchhoff, M.; Scherer, C.; Osterman, A.; Heide1, M.; Reischer, A.; Subklewe, M.; et al. COVID-19 in patients receiving CD20-depleting immunochemotherapy for B-cell lymphoma. Hemasphere 2021, 5, e603. [Google Scholar] [CrossRef]
- Cope, A.; Le Friec, G.; Cardone, J.; Kemper, C. The Th1 life cycle: Molecular control of IFN-γ to IL-10 switching. Trends Immunol. 2011, 32, 278–286. [Google Scholar] [CrossRef]
- Lugli, E.; Marcenaro, E.; Mavilio, D. NK Cell Subset Redistribution during the Course of Viral Infections. Front. Immunol. 2014, 5, 390. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- de Jong, L.C.; Crnko, S.; ten Broeke, T.; Bovenschen, N. Noncytotoxic functions of killer cell granzymes in viral infections. PLoS Pathog. 2021, 17, e1009818. [Google Scholar] [CrossRef] [PubMed]
- Shimizu, K.; Yamasaki, S.; Sakurai, M.; Yumoto, N.; Ikeda, M.; Mishima-Tsumagari, C.; Kukimoto-Niino, M.; Watanabe, T.; Kawamura, M.; Shirouzu, M.; et al. Granzyme A stimulates pDCs to promote adaptive immunity via induction of type I IFN. Front Immunol. 2019, 10, 1450. [Google Scholar] [CrossRef] [Green Version]
- Hueso, T.; Pouderoux, C.; Péré, H.; Beaumont, A.-L.; Raillon, L.-A.; Ader, F.; Chatenoud, L.; Eshagh, D.; Szwebel, T.-A.; Martinot, M.; et al. Convalescent plasma therapy for B-cell–depleted patients with protracted COVID-19. Blood 2020, 136, 2290–2295. [Google Scholar] [CrossRef]
- Gliwiński, M.; Iwaszkiewicz-Grześ, D.; Wołoszyn-Durkiewicz, A.; Tarnowska, M.; Żalińska, M.; Hennig, M.; Zielińska, H.; Dukat-Mazurek, A.; Zielkowska-Dębska, J.; Zieliński, M.; et al. Proinsulin-specific T regulatory cells may control immune responses in type 1 diabetes: Implications for adoptive therapy. BMJ Open Diabetes Res. Care 2020, 8, e000873. [Google Scholar] [CrossRef] [Green Version]
- Marek-Trzonkowska, N.; Piekarska, K.; Filipowicz, N.; Piotrowski, A.; Gucwa, M.; Vogt, K.; Sawitzki, B.; Siebert, J.; Trzonkowski, P. Mild hypothermia provides Treg stability. Sci. Rep. 2017, 7, 11915. [Google Scholar] [CrossRef] [Green Version]
- Šustić, M.; Cokarić Brdovčak, M.; Lisnić, B.; Materljan, J.; Juranić Lisnić, V.; Rožmanić, C.; Indenbirken, D.; Hiršl, L.; Busch, D.H.; Brizić, I.; et al. Memory CD8 T Cells Generated by Cytomegalovirus Vaccine Vector Expressing NKG2D Ligand Have Effector-Like Phenotype and Distinct Functional Features. Front. Immunol. 2021, 12, 2035. [Google Scholar] [CrossRef] [PubMed]
| Author | Age; Sex | Diagnosis | Reason for B-Cell Immune Deficit | Infection Duration (Days) | ARDS/Multiorgan Involvement | Efficacy of Remdesivir (Treatment Duration) | Effect of Convalescent Plasma | Seroconversion |
|---|---|---|---|---|---|---|---|---|
| Helleberg et al. [9] | 50+; M | CLL | 6 cycles of CFR | 60 | no | Reduced viral load after 2nd attempt (10 days each) | Viral clearance (2 infusions) | no |
| Avanzato et al. [2] | 71; F | CLL | Long-term CLL therapy; hypogammaglobulinemia | 105 | no | NA | Viral clearance (2 infusions) | no |
| Martinot et al. [10] | 76; F | CLL | 4 cycles of RB | 41 | no | Short-term clinical improvement | Viral clearance (4 infusions) | no |
| Camprubí-Ferrer et al. [11] | 37; F | Relapsed FL | 3 cycles of R-ESHAP | 63 | no | Viral clearance after 2nd attempt (10 days each) | NE 1 | no |
| Malsy et al. [12] | 53; F | FP | O-CHOP; obinutuzumab maintenance | 94 | no | Clinical improvement (10 days, 5 days) | Viral clearance 2 (2 courses of 6 units) | no |
| Buckland et al. [13] | 31; M | XLA | Primary immunodeficiency | 64 | no | Short-term clinical improvement after 1st course, viral clearance after 2nd attempt (10 days each) | NE 1 | no |
| Baang et al. [14] | 60; M | Refractory MCL | COP + 2 consecutive B-cell directed antibodies | 156 3 | no | Short-term clinical improvement after each attempt (10 days each) 3 | Short-term clinical improvement 3 (2 infusions) | yes |
| Current case | 69; M | DLBCL | 6 cycles of R-CHOP; obinutuzumab maintenance | 96 | no | Clinical improvement, viral clearance (10 days) | Clinical improvement, viral clearance (2 infusions) | no |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Jassem, J.; Marek-Trzonkowska, N.M.; Smiatacz, T.; Arcimowicz, Ł.; Papak, I.; Jassem, E.; Zaucha, J.M. Successful Treatment of Persistent SARS-CoV-2 Infection in a B-Cell Depleted Patient with Activated Cytotoxic T and NK Cells: A Case Report. Int. J. Mol. Sci. 2021, 22, 10934. https://doi.org/10.3390/ijms222010934
Jassem J, Marek-Trzonkowska NM, Smiatacz T, Arcimowicz Ł, Papak I, Jassem E, Zaucha JM. Successful Treatment of Persistent SARS-CoV-2 Infection in a B-Cell Depleted Patient with Activated Cytotoxic T and NK Cells: A Case Report. International Journal of Molecular Sciences. 2021; 22(20):10934. https://doi.org/10.3390/ijms222010934
Chicago/Turabian StyleJassem, Jacek, Natalia Maria Marek-Trzonkowska, Tomasz Smiatacz, Łukasz Arcimowicz, Ines Papak, Ewa Jassem, and Jan Maciej Zaucha. 2021. "Successful Treatment of Persistent SARS-CoV-2 Infection in a B-Cell Depleted Patient with Activated Cytotoxic T and NK Cells: A Case Report" International Journal of Molecular Sciences 22, no. 20: 10934. https://doi.org/10.3390/ijms222010934
APA StyleJassem, J., Marek-Trzonkowska, N. M., Smiatacz, T., Arcimowicz, Ł., Papak, I., Jassem, E., & Zaucha, J. M. (2021). Successful Treatment of Persistent SARS-CoV-2 Infection in a B-Cell Depleted Patient with Activated Cytotoxic T and NK Cells: A Case Report. International Journal of Molecular Sciences, 22(20), 10934. https://doi.org/10.3390/ijms222010934







