Multiplexed MRM-Based Proteomics Identified Multiple Biomarkers of Disease Severity in Human Heart Failure
Abstract
1. Introduction
2. Results
2.1. Characteristics of Study Participants
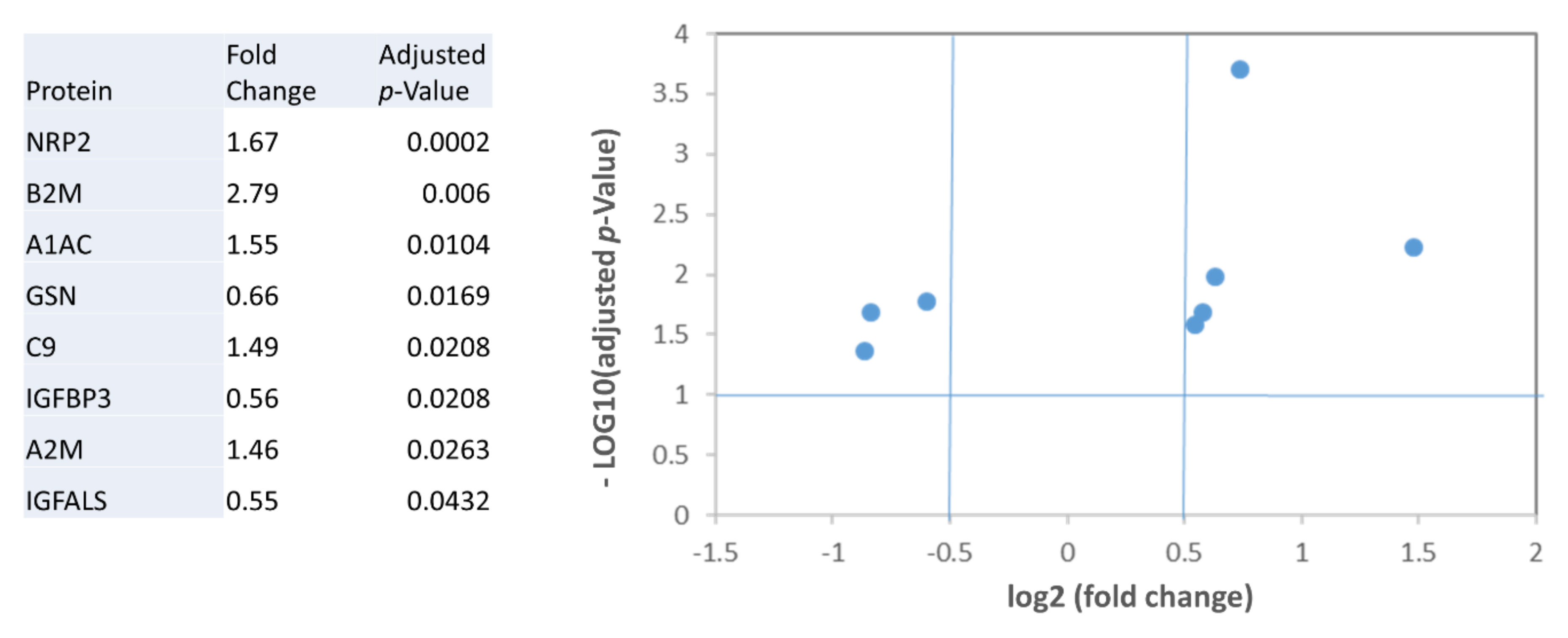
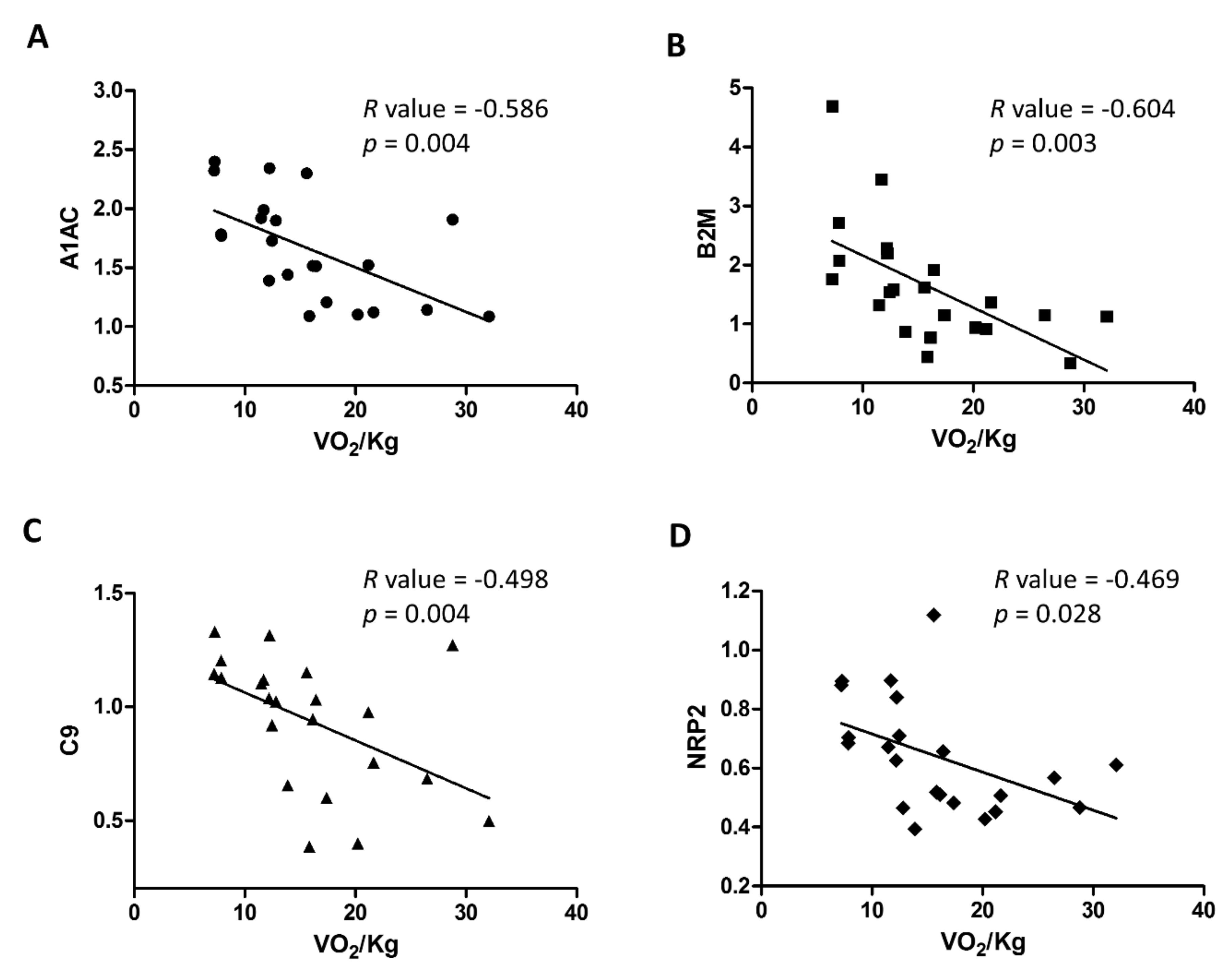
2.2. Analysis of Plasma Biomarkers in Relation to HF Severity
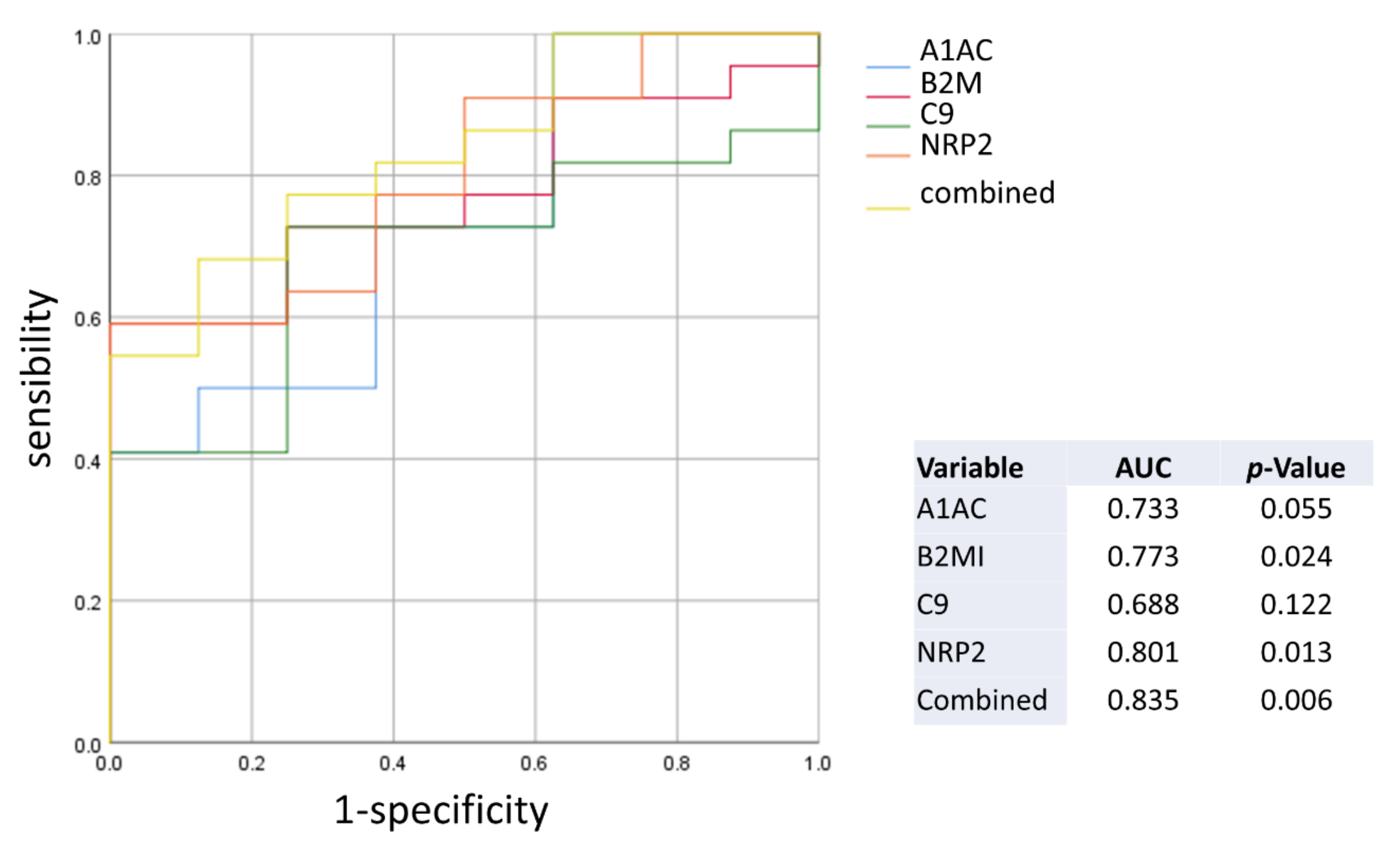
2.3. Diagnostic Performance of the Plasma Biomarkers
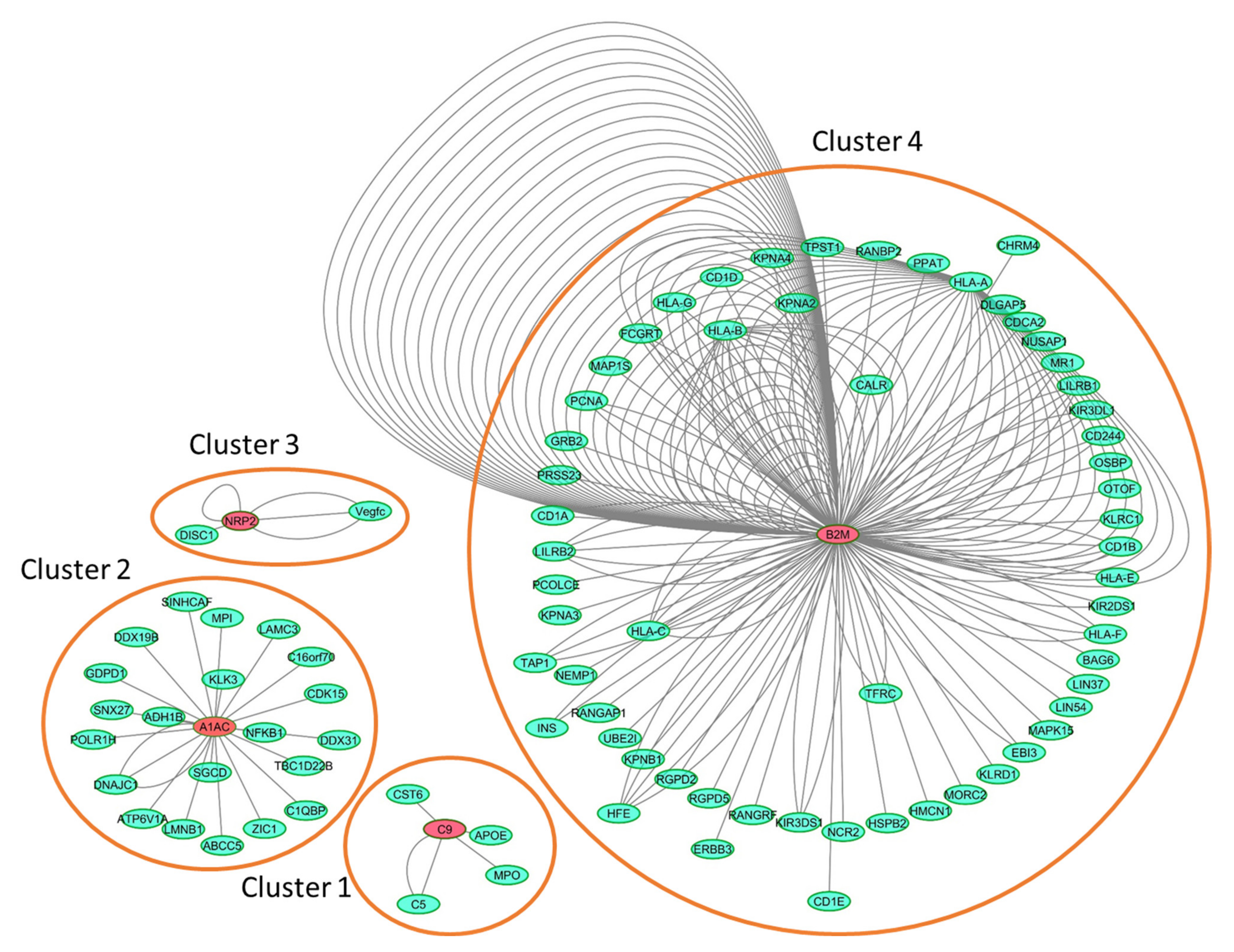
2.4. Interaction Network Analysis
3. Discussion
4. Materials and Methods
4.1. Study Population
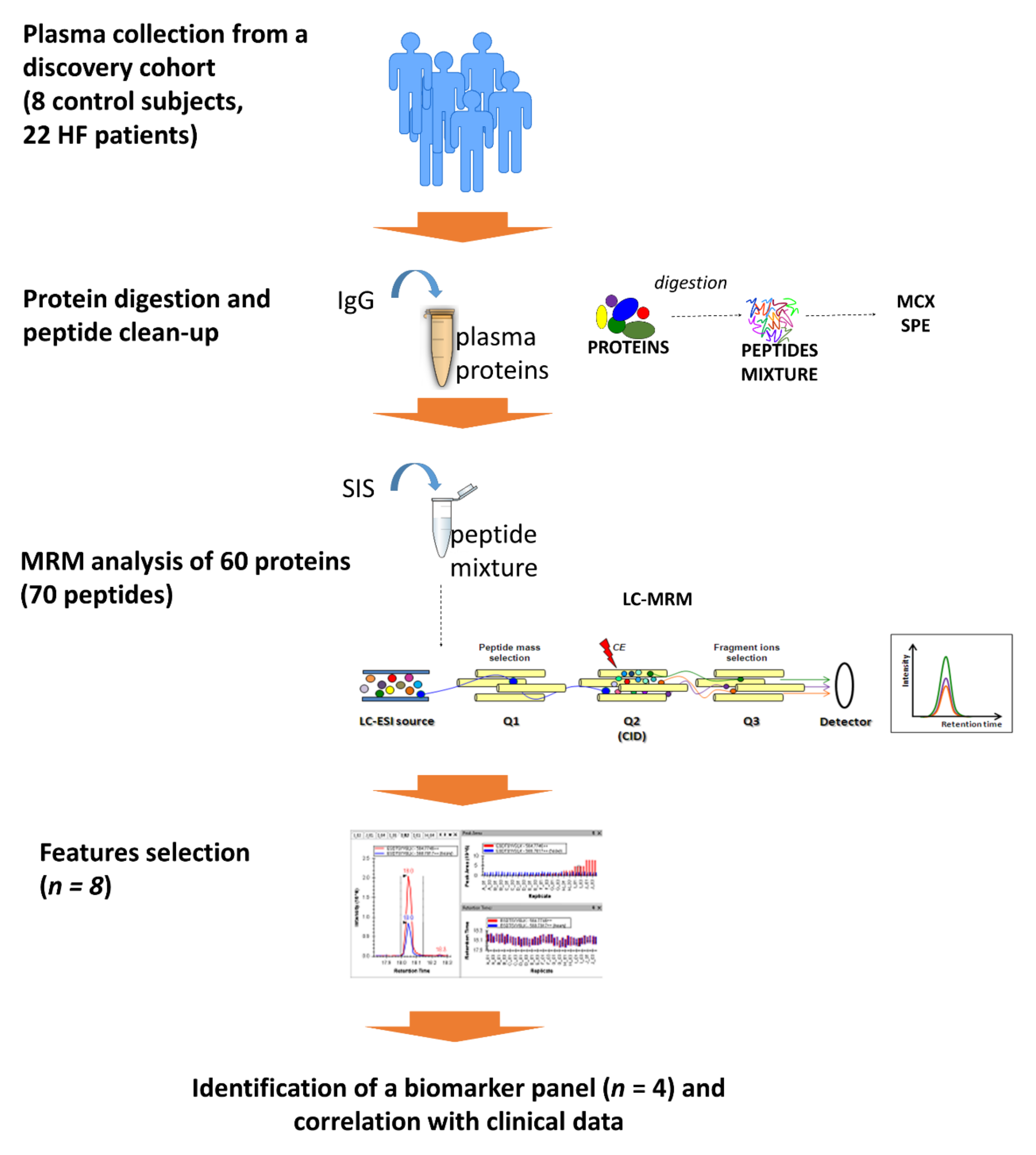
4.2. Sample Preparation
4.3. Liquid Chromatography Mass Spectrometry (LC-MS) Analysis
4.4. Computational Analysis
4.5. Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Braunwald, E. Heart failure. JACC Heart Fail 2013, 1, 1–20. [Google Scholar] [CrossRef]
- Ibrahim, N.E.; Januzzi, J.L., Jr. Established and Emerging Roles of Biomarkers in Heart Failure. Circ. Res. 2018, 123, 614–629. [Google Scholar] [CrossRef] [PubMed]
- Ky, B.; French, B.; Levy, W.C.; Sweitzer, N.K.; Fang, J.C.; Wu, A.H.; Goldberg, L.R.; Jessup, M.; Cappola, T.P. Multiple biomarkers for risk prediction in chronic heart failure. Circ. Heart Fail. 2012, 5, 183–190. [Google Scholar] [CrossRef] [PubMed]
- Sarhene, M.; Wang, Y.; Wei, J.; Huang, Y.; Li, M.; Li, L.; Acheampong, E.; Zhengcan, Z.; Xiaoyan, Q.; Yunsheng, X.; et al. Biomarkers in heart failure: The past, current and future. Heart Fail. Rev. 2019, 24, 867–903. [Google Scholar] [CrossRef] [PubMed]
- Rastogi, A.; Novak, E.; Platts, A.E.; Mann, D.L. Epidemiology, pathophysiology and clinical outcomes for heart failure patients with a mid-range ejection fraction. Eur. J. Heart Fail. 2017, 19, 1597–1605. [Google Scholar] [CrossRef]
- Bayes-Genis, A.; Nunez, J.; Nunez, E.; Martinez, J.B.; Ferrer, M.P.; de Antonio, M.; Zamora, E.; Sanchis, J.; Roses, J.L. Multi-Biomarker Profiling and Recurrent Hospitalizations in Heart Failure. Front. Cardiovasc. Med. 2016, 3, 37. [Google Scholar] [CrossRef]
- Agostoni, P.; Paolillo, S.; Mapelli, M.; Gentile, P.; Salvioni, E.; Veglia, F.; Bonomi, A.; Corra, U.; Lagioia, R.; Limongelli, G.; et al. Multiparametric prognostic scores in chronic heart failure with reduced ejection fraction: A long-term comparison. Eur. J. Heart Fail. 2018, 20, 700–710. [Google Scholar] [CrossRef]
- Spoletini, I.; Coats, A.J.S.; Senni, M.; Rosano, G.M.C. Monitoring of biomarkers in heart failure. Eur. Heart J. Suppl. 2019, 21, M5–M8. [Google Scholar] [CrossRef]
- Gianazza, E.; Tremoli, E.; Banfi, C. The selected reaction monitoring/multiple reaction monitoring-based mass spectrometry approach for the accurate quantitation of proteins: Clinical applications in the cardiovascular diseases. Expert Rev. Proteomics 2014, 11, 771–788. [Google Scholar] [CrossRef]
- Ignjatovic, V.; Geyer, P.E.; Palaniappan, K.K.; Chaaban, J.E.; Omenn, G.S.; Baker, M.S.; Deutsch, E.W.; Schwenk, J.M. Mass Spectrometry-Based Plasma Proteomics: Considerations from Sample Collection to Achieving Translational Data. J. Proteome Res. 2019, 18, 4085–4097. [Google Scholar] [CrossRef]
- Dudkina, N.V.; Spicer, B.A.; Reboul, C.F.; Conroy, P.J.; Lukoyanova, N.; Elmlund, H.; Law, R.H.; Ekkel, S.M.; Kondos, S.C.; Goode, R.J.; et al. Structure of the poly-C9 component of the complement membrane attack complex. Nat. Commun. 2016, 7, 10588. [Google Scholar] [CrossRef] [PubMed]
- Egerstedt, A.; Berntsson, J.; Smith, M.L.; Gidlof, O.; Nilsson, R.; Benson, M.; Wells, Q.S.; Celik, S.; Lejonberg, C.; Farrell, L.; et al. Profiling of the plasma proteome across different stages of human heart failure. Nat. Commun. 2019, 10, 5830. [Google Scholar] [CrossRef] [PubMed]
- Alexanian, R.; Barlogie, B.; Fritsche, H. Beta 2 microglobulin in multiple myeloma. Am. J. Hematol. 1985, 20, 345–351. [Google Scholar] [CrossRef] [PubMed]
- Wilson, A.M.; Kimura, E.; Harada, R.K.; Nair, N.; Narasimhan, B.; Meng, X.Y.; Zhang, F.; Beck, K.R.; Olin, J.W.; Fung, E.T.; et al. Beta2-microglobulin as a biomarker in peripheral arterial disease: Proteomic profiling and clinical studies. Circulation 2007, 116, 1396–1403. [Google Scholar] [CrossRef] [PubMed]
- Klappacher, G.; Mundigler, G.; Papousek, A.; Pacher, R.; Woloszczuk, W.; Ullrich, R.; Buxbaum, P.; Glogar, D. Elevated circulating levels of beta 2-microglobulin in patients with idiopathic dilated cardiomyopathy. Am. J. Cardiol. 1993, 71, 119–122. [Google Scholar] [CrossRef]
- Amighi, J.; Hoke, M.; Mlekusch, W.; Schlager, O.; Exner, M.; Haumer, M.; Pernicka, E.; Koppensteiner, R.; Minar, E.; Rumpold, H.; et al. Beta 2 microglobulin and the risk for cardiovascular events in patients with asymptomatic carotid atherosclerosis. Stroke 2011, 42, 1826–1833. [Google Scholar] [CrossRef]
- Shinkai, S.; Chaves, P.H.; Fujiwara, Y.; Watanabe, S.; Shibata, H.; Yoshida, H.; Suzuki, T. Beta2-microglobulin for risk stratification of total mortality in the elderly population: Comparison with cystatin C and C-reactive protein. Arch. Intern. Med. 2008, 168, 200–206. [Google Scholar] [CrossRef][Green Version]
- Xie, J.; Yi, Q. Beta2-microglobulin as a potential initiator of inflammatory responses. Trends Immunol. 2003, 24, 228–229. [Google Scholar] [CrossRef]
- Prentice, R.L.; Paczesny, S.; Aragaki, A.; Amon, L.M.; Chen, L.; Pitteri, S.J.; McIntosh, M.; Wang, P.; Buson Busald, T.; Hsia, J.; et al. Novel proteins associated with risk for coronary heart disease or stroke among postmenopausal women identified by in-depth plasma proteome profiling. Genome Med. 2010, 2, 48. [Google Scholar] [CrossRef]
- Prud’homme, G.J.; Glinka, Y. Neuropilins are multifunctional coreceptors involved in tumor initiation, growth, metastasis and immunity. Oncotarget 2012, 3, 921–939. [Google Scholar] [CrossRef]
- Tromp, J.; Khan, M.A.; Klip, I.T.; Meyer, S.; de Boer, R.A.; Jaarsma, T.; Hillege, H.; van Veldhuisen, D.J.; van der Meer, P.; Voors, A.A. Biomarker Profiles in Heart Failure Patients with Preserved and Reduced Ejection Fraction. J. Am. Heart Assoc. 2017, 6, e003989. [Google Scholar] [CrossRef] [PubMed]
- Lok, S.I.; van Mil, A.; Bovenschen, N.; van der Weide, P.; van Kuik, J.; van Wichen, D.; Peeters, T.; Siera, E.; Winkens, B.; Sluijter, J.P.; et al. Post-transcriptional regulation of alpha-1-antichymotrypsin by microRNA-137 in chronic heart failure and mechanical support. Circ. Heart Fail. 2013, 6, 853–861. [Google Scholar] [CrossRef] [PubMed]
- Braghin, E.; Galimberti, D.; Scarpini, E.; Bresolin, N.; Baron, P. Alpha1-antichymotrypsin induces TNF-alpha production and NF-kappaB activation in the murine N9 microglial cell line. Neurosci. Lett. 2009, 467, 40–42. [Google Scholar] [CrossRef] [PubMed]
- Yokoyama, T.; Nakano, M.; Bednarczyk, J.L.; McIntyre, B.W.; Entman, M.; Mann, D.L. Tumor necrosis factor-alpha provokes a hypertrophic growth response in adult cardiac myocytes. Circulation 1997, 95, 1247–1252. [Google Scholar] [CrossRef] [PubMed]
- Miller, S.A.; Kolpakov, M.A.; Guo, X.; Du, B.; Nguyen, Y.; Wang, T.; Powel, P.; Dell’Italia, L.J.; Sabri, A. Intracardiac administration of neutrophil protease cathepsin G activates noncanonical inflammasome pathway and promotes inflammation and pathological remodeling in non-injured heart. J. Mol. Cell. Cardiol. 2019, 134, 29–39. [Google Scholar] [CrossRef] [PubMed]
- Jahanyar, J.; Youker, K.A.; Loebe, M.; Assad-Kottner, C.; Koerner, M.M.; Torre-Amione, G.; Noon, G.P. Mast cell-derived cathepsin g: A possible role in the adverse remodeling of the failing human heart. J. Surg. Res. 2007, 140, 199–203. [Google Scholar] [CrossRef]
- Aimo, A.; Vergaro, G.; Barison, A.; Maffei, S.; Borrelli, C.; Morrone, D.; Cameli, M.; Palazzuoli, A.; Ambrosio, G.; Coiro, S.; et al. Sex-related differences in chronic heart failure. Int. J. Cardiol. 2018, 255, 145–151. [Google Scholar] [CrossRef]
- Zannad, F.; Ferreira, J.P.; Pocock, S.J.; Anker, S.D.; Butler, J.; Filippatos, G.; Brueckmann, M.; Ofstad, A.P.; Pfarr, E.; Jamal, W.; et al. SGLT2 inhibitors in patients with heart failure with reduced ejection fraction: A meta-analysis of the EMPEROR-Reduced and DAPA-HF trials. Lancet 2020, 396, 819–829. [Google Scholar] [CrossRef]
- Bonnet, F.; Scheen, A.J. Effects of SGLT2 inhibitors on systemic and tissue low-grade inflammation: The potential contribution to diabetes complications and cardiovascular disease. Diabetes Metab. 2018, 44, 457–464. [Google Scholar] [CrossRef]
- Gerszten, R.E.; Wang, T.J. The search for new cardiovascular biomarkers. Nature 2008, 451, 949–952. [Google Scholar] [CrossRef]
- Wang, T.J.; Gona, P.; Larson, M.G.; Tofler, G.H.; Levy, D.; Newton-Cheh, C.; Jacques, P.F.; Rifai, N.; Selhub, J.; Robins, S.J.; et al. Multiple biomarkers for the prediction of first major cardiovascular events and death. N. Engl. J. Med. 2006, 355, 2631–2639. [Google Scholar] [CrossRef] [PubMed]
- Chirinos, J.A.; Orlenko, A.; Zhao, L.; Basso, M.D.; Cvijic, M.E.; Li, Z.; Spires, T.E.; Yarde, M.; Wang, Z.; Seiffert, D.A.; et al. Multiple Plasma Biomarkers for Risk Stratification in Patients with Heart Failure and Preserved Ejection Fraction. J. Am. Coll. Cardiol. 2020, 75, 1281–1295. [Google Scholar] [CrossRef] [PubMed]
- Gaggin, H.K.; Truong, Q.A.; Gandhi, P.U.; Motiwala, S.R.; Belcher, A.M.; Weiner, R.B.; Baggish, A.L.; Januzzi, J.L., Jr. Systematic Evaluation of Endothelin 1 Measurement Relative to Traditional and Modern Biomarkers for Clinical Assessment and Prognosis in Patients with Chronic Systolic Heart Failure: Serial Measurement and Multimarker Testing. Am. J. Clin. Pathol. 2017, 147, 461–472. [Google Scholar] [CrossRef] [PubMed]
- Gargiulo, P.; Banfi, C.; Ghilardi, S.; Magri, D.; Giovannardi, M.; Bonomi, A.; Salvioni, E.; Battaia, E.; Filardi, P.P.; Tremoli, E.; et al. Surfactant-derived proteins as markers of alveolar membrane damage in heart failure. PLoS ONE 2014, 9, e115030. [Google Scholar] [CrossRef] [PubMed]
- Magri, D.; Banfi, C.; Maruotti, A.; Farina, S.; Vignati, C.; Salvioni, E.; Morosin, M.; Brioschi, M.; Ghilardi, S.; Tremoli, E.; et al. Plasma immature form of surfactant protein type B correlates with prognosis in patients with chronic heart failure. A pilot single-center prospective study. Int. J. Cardiol. 2015, 201, 394–399. [Google Scholar] [CrossRef]
- Magri, D.; Brioschi, M.; Banfi, C.; Schmid, J.P.; Palermo, P.; Contini, M.; Apostolo, A.; Bussotti, M.; Tremoli, E.; Sciomer, S.; et al. Circulating plasma surfactant protein type B as biological marker of alveolar-capillary barrier damage in chronic heart failure. Circ. Heart Fail. 2009, 2, 175–180. [Google Scholar] [CrossRef]
- Banfi, C.; Agostoni, P. Surfactant protein B: From biochemistry to its potential role as diagnostic and prognostic marker in heart failure. Int. J. Cardiol. 2016, 221, 456–462. [Google Scholar] [CrossRef]
- Campodonico, J.; Mapelli, M.; Spadafora, E.; Ghilardi, S.; Agostoni, P.; Banfi, C.; Sciomer, S. Surfactant proteins changes after acute hemodynamic improvement in patients with advanced chronic heart failure treated with Levosimendan. Respir. Physiol. Neurobiol. 2018, 252, 47–51. [Google Scholar] [CrossRef]
- Banfi, C.; Brioschi, M.; Karjalainen, M.K.; Huusko, J.M.; Gianazza, E.; Agostoni, P. Immature surfactant protein-B impairs the antioxidant capacity of HDL. Int. J. Cardiol. 2019, 285, 53–58. [Google Scholar] [CrossRef]
- Gaggin, H.K.; Januzzi, J.L., Jr. Biomarkers and diagnostics in heart failure. Biochim. Biophys. Acta 2013, 1832, 2442–2450. [Google Scholar] [CrossRef]
- Berezin, A.E. Circulating biomarkers in heart failure: Diagnostic and prognostic importance. J. Lab. Precis. Med. 2018, 3, 36. [Google Scholar] [CrossRef]
- Nadar, S.K.; Shaikh, M.M. Biomarkers in Routine Heart Failure Clinical Care. Card. Fail. Rev. 2019, 5, 50–56. [Google Scholar] [CrossRef] [PubMed]
- Orchard, S.; Ammari, M.; Aranda, B.; Breuza, L.; Briganti, L.; Broackes-Carter, F.; Campbell, N.H.; Chavali, G.; Chen, C.; del-Toro, N.; et al. The MIntAct project—IntAct as a common curation platform for 11 molecular interaction databases. Nucleic Acids Res. 2014, 42, D358–D363. [Google Scholar] [CrossRef] [PubMed]
- Franceschini, A.; Szklarczyk, D.; Frankild, S.; Kuhn, M.; Simonovic, M.; Roth, A.; Lin, J.; Minguez, P.; Bork, P.; von Mering, C.; et al. STRING v9.1: Protein-protein interaction networks, with increased coverage and integration. Nucleic Acids Res. 2013, 41, D808–D815. [Google Scholar] [CrossRef] [PubMed]
| Controls | Low Severity HF | Medium Severity HF | High Severity HF | |||
|---|---|---|---|---|---|---|
| Protein | VO2/Kg > 16 mL/min/Kg (n = 9) | 12 < VO2/Kg < 16 mL/min/Kg (n = 7) | VO2/Kg < 12 mL/min/Kg (n = 6) | p-Value | Adjusted p-Value | |
| A1AC | 1.332 ± 0.323 | 1.3466 ± 0.282 | 1.742 ± 0.471 | 2.030 ± 0.270 | 0.002 | 0.001 |
| A2M | 0.946 ± 0.163 | 1.055 ± 0.356 | 1.028 ± 0.169 | 1.398 ± 0.475 | ns | - |
| B2M | 0.873 ± 0.319 | 1.073 ± 0.429 | 1.5058 ± 0.663 | 2.665 ± 1.240 | <0.001 | 0.001 |
| C9 | 0.768 ± 0.192 | 0.7955 ± 0.282 | 0.9268 ± 0.314 | 1.171 ± 0.0854 | 0.020 | 0.014 |
| GSN | 1.034 ± 0.174 | 0.760 ± 0.185 | 0.822 ± 0.200 | 0.729 ± 0.216 | 0.020 | 0.043 |
| IGFBP3 | 0.929 ± 0.0930 | 0.774 ± 0.124 | 0.849 ± 0.340 | 0.589 ± 0.335 | ns | - |
| IGFALS | 1.101 ± 0.251 | 0.851 ± 0.250 | 0.963 ± 0.437 | 0.696 ± 0.390 | ns | - |
| NRP2 | 0.448 ± 0.0855 | 0.520 ± 0.076 | 0.667 ± 0.250 | 0.789 ± 0.112 | 0.001 | < 0.001 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Brioschi, M.; Gianazza, E.; Agostoni, P.; Zoanni, B.; Mallia, A.; Banfi, C. Multiplexed MRM-Based Proteomics Identified Multiple Biomarkers of Disease Severity in Human Heart Failure. Int. J. Mol. Sci. 2021, 22, 838. https://doi.org/10.3390/ijms22020838
Brioschi M, Gianazza E, Agostoni P, Zoanni B, Mallia A, Banfi C. Multiplexed MRM-Based Proteomics Identified Multiple Biomarkers of Disease Severity in Human Heart Failure. International Journal of Molecular Sciences. 2021; 22(2):838. https://doi.org/10.3390/ijms22020838
Chicago/Turabian StyleBrioschi, Maura, Erica Gianazza, Piergiuseppe Agostoni, Beatrice Zoanni, Alice Mallia, and Cristina Banfi. 2021. "Multiplexed MRM-Based Proteomics Identified Multiple Biomarkers of Disease Severity in Human Heart Failure" International Journal of Molecular Sciences 22, no. 2: 838. https://doi.org/10.3390/ijms22020838
APA StyleBrioschi, M., Gianazza, E., Agostoni, P., Zoanni, B., Mallia, A., & Banfi, C. (2021). Multiplexed MRM-Based Proteomics Identified Multiple Biomarkers of Disease Severity in Human Heart Failure. International Journal of Molecular Sciences, 22(2), 838. https://doi.org/10.3390/ijms22020838






