The Lung Microbiome during Health and Disease
Abstract
1. Introduction
2. The Respiratory Tract Microbiome during Health
2.1. The Lungs Are Not Sterile and Are Not the Gut
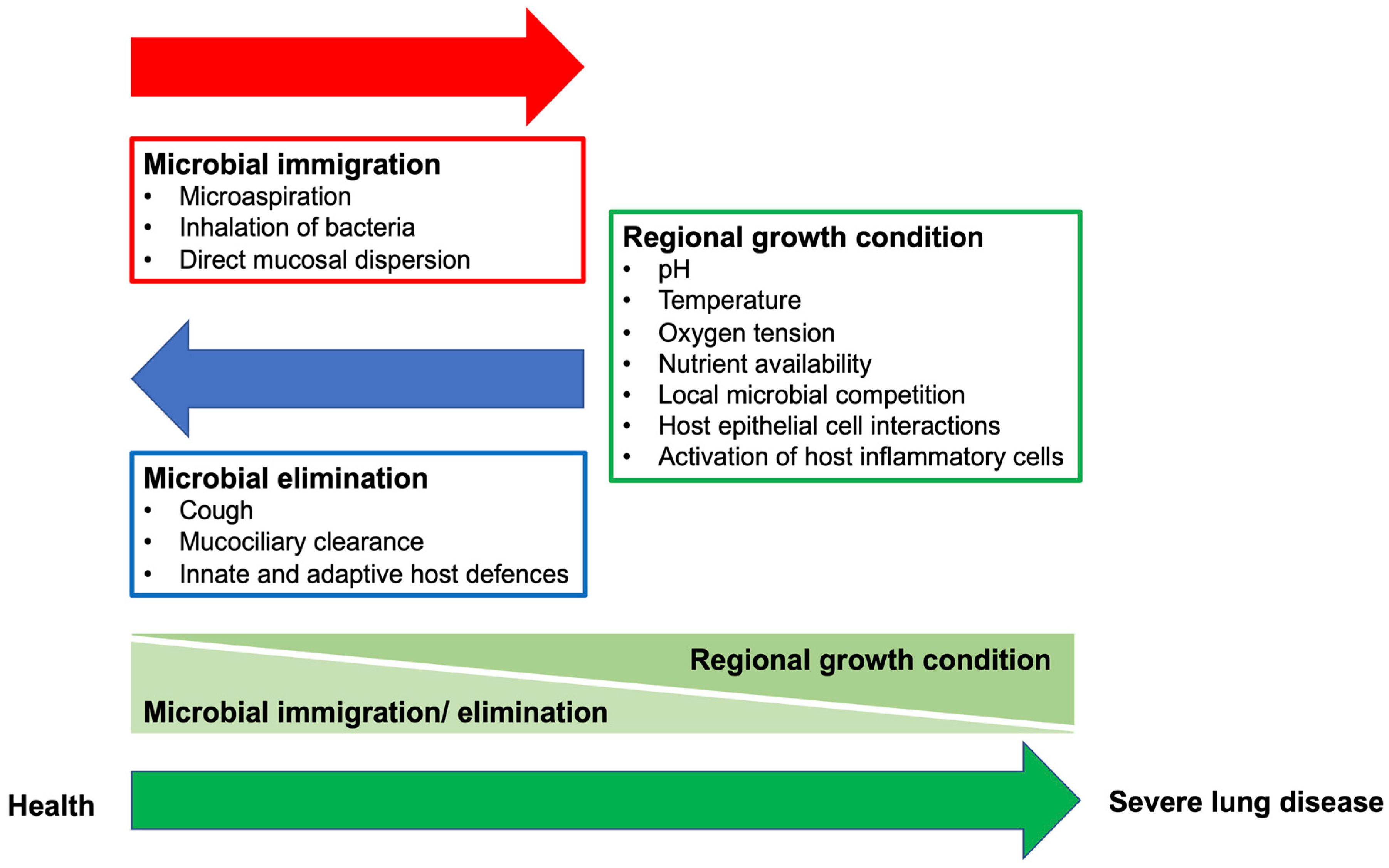
2.2. Key Ecological Factors Determining the Lung Microbiome
2.3. The Oral Microbiome and the Nasal Microbiome
3. The Lung Microbiome during Disease
3.1. Asthma
3.2. Chronic Obstrutive Pulmonary Disease (COPD)
3.3. Bronchiectasis
3.4. Lung Cancer
3.5. Respiratory Viral Infections
4. Perspectives of Lung Microbiome Research
4.1. Limitations of Lung Microbiome Research
4.2. Host-Microbe Interactions
4.3. Clinical Applicatiton of the Lung Microbiome
Author Contributions
Funding
Conflicts of Interest
References
- Huffnagle, G.B.; Dickson, R.P.; Lukacs, N.W. The respiratory tract microbiome and lung inflammation: A two-way street. Mucosal. Immunol. 2017, 10, 299–306. [Google Scholar] [CrossRef]
- Dickson, R.P.; Erb-Downward, J.R.; Martinez, F.J.; Huffnagle, G.B. The Microbiome and the Respiratory Tract. Ann. Rev. Physiol. 2016, 78, 481–504. [Google Scholar] [CrossRef]
- Gleeson, K.; Eggli, D.F.; Maxwell, S.L. Quantitative aspiration during sleep in normal subjects. Chest 1997, 111, 1266–1272. [Google Scholar] [CrossRef]
- Huxley, E.J.; Viroslav, J.; Gray, W.R.; Pierce, A.K. Pharyngeal aspiration in normal adults and patients with depressed consciousness. Am. J. Med. 1978, 64, 564–568. [Google Scholar] [CrossRef]
- Hilty, M.; Burke, C.; Pedro, H.; Cardenas, P.; Bush, A.; Bossley, C.; Davies, J.; Ervine, A.; Poulter, L.; Pachter, L.; et al. Disordered microbial communities in asthmatic airways. PLoS ONE 2010, 5, e8578. [Google Scholar] [CrossRef]
- Amberson, J.B. A clinical consideration of abscesses and cavities of the lung. Bull. Johns Hopkins Hosp. 1954, 94, 227–237. [Google Scholar]
- Lederberg, J.; McCray, A.T. ‘Ome Sweet’ Omics-A Genealogical Treasury of Words. Scientist 2001, 15, 8. [Google Scholar]
- Pattaroni, C.; Watzenboeck, M.L.; Schneidegger, S.; Kieser, S.; Wong, N.C.; Bernasconi, E.; Pernot, J.; Mercier, L.; Knapp, S.; Nicod, L.P.; et al. Early-Life Formation of the Microbial and Immunological Environment of the Human Airways. Cell Host Microbe 2018, 24, 857–865.e4. [Google Scholar] [CrossRef]
- Bassis, C.M.; Erb-Downward, J.R.; Dickson, R.P.; Freeman, C.M.; Schmidt, T.M.; Young, V.B.; Beck, J.M.; Curtis, J.L.; Huffnagle, G.B. Analysis of the upper respiratory tract microbiotas as the source of the lung and gastric microbiotas in healthy individuals. mBiology 2015, 6, e00037. [Google Scholar] [CrossRef] [PubMed]
- Yu, G.; Gail, M.H.; Consonni, D.; Carugno, M.; Humphrys, M.; Pesatori, A.C.; Caporaso, N.E.; Goedert, J.J.; Ravel, J.; Landi, M.T. Characterizing human lung tissue microbiota and its relationship to epidemiological and clinical features. Genome Biol. 2016, 17, 163. [Google Scholar] [CrossRef] [PubMed]
- Erb-Downward, J.R.; Thompson, D.L.; Han, M.K.; Freeman, C.M.; McCloskey, L.; Schmidt, L.A.; Young, V.B.; Toews, G.B.; Curtis, J.L.; Sundaram, B.; et al. Analysis of the lung microbiome in the “healthy” smoker and in COPD. PLoS ONE 2011, 6, e16384. [Google Scholar] [CrossRef]
- Morris, A.; Beck, J.M.; Schloss, P.D.; Campbell, T.B.; Crothers, K.; Curtis, J.L.; Flores, S.C.; Fontenot, A.P.; Ghedin, E.; Huang, L.; et al. Comparison of the respiratory microbiome in healthy nonsmokers and smokers. Am. J. Respir. Crit Care Med. 2013, 187, 1067–1075. [Google Scholar] [CrossRef] [PubMed]
- Segal, L.N.; Clemente, J.C.; Tsay, J.C.; Koralov, S.B.; Keller, B.C.; Wu, B.G.; Li, Y.; Shen, N.; Ghedin, E.; Morris, A.; et al. Enrichment of the lung microbiome with oral taxa is associated with lung inflammation of a Th17 phenotype. Nat. Microbiol. 2016, 1, 16031. [Google Scholar] [CrossRef] [PubMed]
- Mathieu, E.; Escribano-Vazquez, U.; Descamps, D.; Cherbuy, C.; Langella, P.; Riffault, S.; Remot, A.; Thomas, M. Paradigms of Lung Microbiota Functions in Health and Disease, Particularly, in Asthma. Front. Physiol. 2018, 9, 1168. [Google Scholar] [CrossRef]
- Dickson, R.P.; Erb-Downward, J.R.; Freeman, C.M.; McCloskey, L.; Beck, J.M.; Huffnagle, G.B.; Curtis, J.L. Spatial Variation in the Healthy Human Lung Microbiome and the Adapted Island Model of Lung Biogeography. Ann. Am. Thorac Soc. 2015, 12, 821–830. [Google Scholar] [CrossRef]
- Faner, R.; Sibila, O.; Agusti, A.; Bernasconi, E.; Chalmers, J.D.; Huffnagle, G.B.; Manichanh, C.; Molyneaux, P.L.; Paredes, R.; Perez Brocal, V.; et al. The microbiome in respiratory medicine: Current challenges and future perspectives. Eur. Respir. J. 2017, 49, 1602086. [Google Scholar] [CrossRef]
- Magalhaes, A.P.; Azevedo, N.F.; Pereira, M.O.; Lopes, S.P. The cystic fibrosis microbiome in an ecological perspective and its impact in antibiotic therapy. Appl. Microbiol. Biotechnol. 2016, 100, 1163–1181. [Google Scholar] [CrossRef] [PubMed]
- Vatanen, T.; Kostic, A.D.; d’Hennezel, E.; Siljander, H.; Franzosa, E.A.; Yassour, M.; Kolde, R.; Vlamakis, H.; Arthur, T.D.; Hamalainen, A.M.; et al. Variation in Microbiome LPS Immunogenicity Contributes to Autoimmunity in Humans. Cell 2016, 165, 1551. [Google Scholar] [CrossRef] [PubMed]
- Blanton, L.V.; Charbonneau, M.R.; Salih, T.; Barratt, M.J.; Venkatesh, S.; Ilkaveya, O.; Subramanian, S.; Manary, M.J.; Trehan, I.; Jorgensen, J.M.; et al. Gut bacteria that prevent growth impairments transmitted by microbiota from malnourished children. Science 2016, 351, 854. [Google Scholar] [CrossRef] [PubMed]
- Guarner, F.; Malagelada, J.R. Gut flora in health and disease. Lancet 2003, 361, 512–519. [Google Scholar] [CrossRef]
- Dickson, R.P.; Huffnagle, G.B. The Lung Microbiome: New Principles for Respiratory Bacteriology in Health and Disease. PLoS Pathog. 2015, 11, e1004923. [Google Scholar] [CrossRef]
- Wypych, T.P.; Wickramasinghe, L.C.; Marsland, B.J. The influence of the microbiome on respiratory health. Nat. Immunol. 2019, 20, 1279–1290. [Google Scholar] [CrossRef]
- Ingenito, E.P.; Solway, J.; McFadden, E.R., Jr.; Pichurko, B.; Bowman, H.F.; Michaels, D.; Drazen, J.M. Indirect assessment of mucosal surface temperatures in the airways: Theory and tests. J. Appl. Physiol. 1987, 63, 2075–2083. [Google Scholar] [CrossRef]
- Wu, H.; Kuzmenko, A.; Wan, S.; Schaffer, L.; Weiss, A.; Fisher, J.H.; Kim, K.S.; McCormack, F.X. Surfactant proteins A and D inhibit the growth of Gram-negative bacteria by increasing membrane permeability. J. Clin. Investig. 2003, 111, 1589–1602. [Google Scholar] [CrossRef]
- Pilette, C.; Ouadrhiri, Y.; Godding, V.; Vaerman, J.P.; Sibille, Y. Lung mucosal immunity: Immunoglobulin-A revisited. Eur. Respir. J. 2001, 18, 571–588. [Google Scholar] [CrossRef]
- Allard, B.; Panariti, A.; Martin, J.G. Alveolar Macrophages in the Resolution of Inflammation, Tissue Repair, and Tolerance to Infection. Front. Immunol. 2018, 9, 1777. [Google Scholar] [CrossRef] [PubMed]
- Dickson, R.P.; Martinez, F.J.; Huffnagle, G.B. The role of the microbiome in exacerbations of chronic lung diseases. Lancet 2014, 384, 691–702. [Google Scholar] [CrossRef]
- Lighthart, B.; Prier, K.; Loper, G.M.; Bromenshenk, J. Bees Scavenge Airborne Bacteria. Microb. Ecol. 2000, 39, 314–321. [Google Scholar] [PubMed]
- Dickson, R.P.; Erb-Downward, J.R.; Huffnagle, G.B. Towards an ecology of the lung: New conceptual models of pulmonary microbiology and pneumonia pathogenesis. Lancet Respir. Med. 2014, 2, 238–246. [Google Scholar] [CrossRef]
- Venkataraman, A.; Bassis, C.M.; Beck, J.M.; Young, V.B.; Curtis, J.L.; Huffnagle, G.B.; Schmidt, T.M. Application of a neutral community model to assess structuring of the human lung microbiome. mBiology 2015, 6, e02284-14. [Google Scholar] [CrossRef]
- Dickson, R.P.; Erb-Downward, J.R.; Freeman, C.M.; Walker, N.; Scales, B.S.; Beck, J.M.; Martinez, F.J.; Curtis, J.L.; Lama, V.N.; Huffnagle, G.B. Changes in the lung microbiome following lung transplantation include the emergence of two distinct Pseudomonas species with distinct clinical associations. PLoS ONE 2014, 9, e97214. [Google Scholar] [CrossRef]
- Dickson, R.P.; Erb-Downward, J.R.; Huffnagle, G.B. The role of the bacterial microbiome in lung disease. Expert Rev. Respir. Med. 2013, 7, 245–257. [Google Scholar] [CrossRef]
- Molyneaux, P.L.; Cox, M.J.; Willis-Owen, S.A.; Mallia, P.; Russell, K.E.; Russell, A.M.; Murphy, E.; Johnston, S.L.; Schwartz, D.A.; Wells, A.U.; et al. The role of bacteria in the pathogenesis and progression of idiopathic pulmonary fibrosis. Am. J. Respir. Crit. Care Med. 2014, 190, 906–913. [Google Scholar] [CrossRef]
- Muhlebach, M.S.; Zorn, B.T.; Esther, C.R.; Hatch, J.E.; Murray, C.P.; Turkovic, L.; Ranganathan, S.C.; Boucher, R.C.; Stick, S.M.; Wolfgang, M.C. Initial acquisition and succession of the cystic fibrosis lung microbiome is associated with disease progression in infants and preschool children. PLoS Pathog. 2018, 14, e1006798. [Google Scholar] [CrossRef] [PubMed]
- Philley, J.V.; Kannan, A.; Olusola, P.; McGaha, P.; Singh, K.P.; Samten, B.; Griffith, D.E.; Dasgupta, S. Microbiome Diversity in Sputum of Nontuberculous Mycobacteria Infected Women with a History of Breast Cancer. Cell Physiol. Biochem. 2019, 52, 263–279. [Google Scholar] [PubMed]
- Tunney, M.M.; Einarsson, G.G.; Wei, L.; Drain, M.; Klem, E.R.; Cardwell, C.; Ennis, M.; Boucher, R.C.; Wolfgang, M.C.; Elborn, J.S. Lung microbiota and bacterial abundance in patients with bronchiectasis when clinically stable and during exacerbation. Am. J. Respir. Crit Care Med. 2013, 187, 1118–1126. [Google Scholar] [CrossRef] [PubMed]
- Cevhertas, L.; Ogulur, I.; Maurer, D.J.; Burla, D.; Ding, M.; Jansen, K.; Koch, J.; Liu, C.; Ma, S.; Mitamura, Y.; et al. Advances and recent developments in asthma in 2020. Allergy 2020, 75, 3124–3146. [Google Scholar] [CrossRef] [PubMed]
- Pavord, I.D.; Beasley, R.; Agusti, A.; Anderson, G.P.; Bel, E.; Brusselle, G.; Cullinan, P.; Custovic, A.; Ducharme, F.M.; Fahy, J.V.; et al. After asthma: Redefining airways diseases. Lancet 2018, 391, 350–400. [Google Scholar] [CrossRef]
- Lukacs, N.W.; Huang, Y.J. Microbiota-immune interactions in asthma pathogenesis and phenotype. Curr. Opin. Immunol. 2020, 66, 22–26. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.J.; Nariya, S.; Harris, J.M.; Lynch, S.V.; Choy, D.F.; Arron, J.R.; Boushey, H. The airway microbiome in patients with severe asthma: Associations with disease features and severity. J. Allergy Clin. Immunol. 2015, 136, 874–884. [Google Scholar] [CrossRef]
- Taylor, S.L.; Leong, L.E.X.; Mobegi, F.M.; Choo, J.M.; Wesselingh, S.; Yang, I.A.; Upham, J.W.; Reynolds, P.N.; Hodge, S.; James, A.L.; et al. Long-Term Azithromycin Reduces Haemophilus influenzae and Increases Antibiotic Resistance in Severe Asthma. Am. J. Respir. Crit Care Med. 2019, 200, 309–317. [Google Scholar] [CrossRef] [PubMed]
- Simpson, J.L.; Daly, J.; Baines, K.J.; Yang, I.A.; Upham, J.W.; Reynolds, P.N.; Hodge, S.; James, A.L.; Hugenholtz, P.; Willner, D.; et al. Airway dysbiosis: Haemophilus influenzae and Tropheryma in poorly controlled asthma. Eur. Respir. J. 2016, 47, 792–800. [Google Scholar] [CrossRef] [PubMed]
- Durack, J.; Lynch, S.V.; Nariya, S.; Bhakta, N.R.; Beigelman, A.; Castro, M.; Dyer, A.M.; Israel, E.; Kraft, M.; Martin, R.J.; et al. Features of the bronchial bacterial microbiome associated with atopy, asthma, and responsiveness to inhaled corticosteroid treatment. J. Allergy Clin. Immunol. 2017, 140, 63–75. [Google Scholar] [CrossRef]
- Arrieta, M.C.; Arevalo, A.; Stiemsma, L.; Dimitriu, P.; Chico, M.E.; Loor, S.; Vaca, M.; Boutin, R.C.T.; Morien, E.; Jin, M.; et al. Associations between infant fungal and bacterial dysbiosis and childhood atopic wheeze in a nonindustrialized setting. J. Allergy Clin. Immunol. 2018, 142, 424–434.e10. [Google Scholar] [CrossRef]
- Begley, L.; Madapoosi, S.; Opron, K.; Ndum, O.; Baptist, A.; Rysso, K.; Erb-Downward, J.R.; Huang, Y.J. Gut microbiota relationships to lung function and adult asthma phenotype: A pilot study. BMJ Open Respir. Res. 2018, 5, e000324. [Google Scholar] [CrossRef] [PubMed]
- Pragman, A.A.; Kim, H.B.; Reilly, C.S.; Wendt, C.; Isaacson, R.E. The lung microbiome in moderate and severe chronic obstructive pulmonary disease. PLoS ONE 2012, 7, e47305. [Google Scholar] [CrossRef]
- Wilkinson, T.M.A.; Aris, E.; Bourne, S.; Clarke, S.C.; Peeters, M.; Pascal, T.G.; Schoonbroodt, S.; Tuck, A.C.; Kim, V.; Ostridge, K.; et al. A prospective, observational cohort study of the seasonal dynamics of airway pathogens in the aetiology of exacerbations in COPD. Thorax 2017, 72, 919–927. [Google Scholar] [CrossRef]
- Wang, Z.; Maschera, B.; Lea, S.; Kolsum, U.; Michalovich, D.; Van Horn, S.; Traini, C.; Brown, J.R.; Hessel, E.M.; Singh, D. Airway host-microbiome interactions in chronic obstructive pulmonary disease. Respir. Res. 2019, 20, 113. [Google Scholar] [CrossRef]
- Ghebre, M.A.; Pang, P.H.; Diver, S.; Desai, D.; Bafadhel, M.; Haldar, K.; Kebadze, T.; Cohen, S.; Newbold, P.; Rapley, L.; et al. Biological exacerbation clusters demonstrate asthma and chronic obstructive pulmonary disease overlap with distinct mediator and microbiome profiles. J. Allergy Clin. Immunol. 2018, 141, 2027–2036.e12. [Google Scholar] [CrossRef]
- Haldar, K.; George, L.; Wang, Z.; Mistry, V.; Ramsheh, M.Y.; Free, R.C.; John, C.; Reeve, N.F.; Miller, B.E.; Tal-Singer, R.; et al. The sputum microbiome is distinct between COPD and health, independent of smoking history. Respir. Res. 2020, 21, 183. [Google Scholar] [CrossRef]
- Sze, M.A.; Dimitriu, P.A.; Suzuki, M.; McDonough, J.E.; Campbell, J.D.; Brothers, J.F.; Erb-Downward, J.R.; Huffnagle, G.B.; Hayashi, S.; Elliott, W.M.; et al. Host Response to the Lung Microbiome in Chronic Obstructive Pulmonary Disease. Am. J. Respir. Crit. Care Med. 2015, 192, 438–445. [Google Scholar] [CrossRef] [PubMed]
- Wu, Y.; Luo, Z.; Liu, C. Variations in fecal microbial profiles of acute exacerbations and stable chronic obstructive pulmonary disease. Life Sci. 2021, 265, 118738. [Google Scholar] [CrossRef]
- Jang, Y.O.; Lee, S.H.; Choi, J.J.; Kim, D.H.; Choi, J.M.; Kang, M.J.; Oh, Y.M.; Park, Y.J.; Shin, Y.; Lee, S.W. Fecal microbial transplantation and a high fiber diet attenuates emphysema development by suppressing inflammation and apoptosis. Exp. Mol. Med. 2020, 52, 1128–1139. [Google Scholar] [CrossRef] [PubMed]
- Chalmers, J.D.; Aliberti, S.; Blasi, F. Management of bronchiectasis in adults. Eur. Respir. J. 2015, 45, 1446–1462. [Google Scholar] [CrossRef] [PubMed]
- Chalmers, J.D.; Chang, A.B.; Chotirmall, S.H.; Dhar, R.; McShane, P.J. Bronchiectasis. Nat. Rev. Dis. Primers. 2018, 4, 45. [Google Scholar] [CrossRef]
- Haworth, C.S.; Banks, J.; Capstick, T.; Fisher, A.J.; Gorsuch, T.; Laurenson, I.F.; Leitch, A.; Loebinger, M.R.; Milburn, H.J.; Nightingale, M.; et al. British Thoracic Society guidelines for the management of non-tuberculous mycobacterial pulmonary disease (NTM-PD). Thorax 2017, 72, ii1–ii64. [Google Scholar] [CrossRef]
- Flume, P.A.; Chalmers, J.D.; Olivier, K.N. Advances in bronchiectasis: Endotyping, genetics, microbiome, and disease heterogeneity. Lancet 2018, 392, 880–890. [Google Scholar] [CrossRef]
- Quint, J.K.; Millett, E.R.; Joshi, M.; Navaratnam, V.; Thomas, S.L.; Hurst, J.R.; Smeeth, L.; Brown, J.S. Changes in the incidence, prevalence and mortality of bronchiectasis in the UK from 2004 to 2013: A population-based cohort study. Eur. Respir. J. 2016, 47, 186–193. [Google Scholar] [CrossRef]
- Aksamit, T.R.; O’Donnell, A.E.; Barker, A.; Olivier, K.N.; Winthrop, K.L.; Daniels, M.L.A.; Johnson, M.; Eden, E.; Griffith, D.; Knowles, M.; et al. Bronchiectasis Research Registry, C. Adult Patients With Bronchiectasis: A First Look at the US Bronchiectasis Research Registry. Chest 2017, 151, 982–992. [Google Scholar] [CrossRef]
- Araujo, D.; Shteinberg, M.; Aliberti, S.; Goeminne, P.C.; Hill, A.T.; Fardon, T.C.; Obradovic, D.; Stone, G.; Trautmann, M.; Davis, A.; et al. The independent contribution of Pseudomonas aeruginosa infection to long-term clinical outcomes in bronchiectasis. Eur. Respir. J. 2018, 51, 1701953. [Google Scholar] [CrossRef]
- Finch, S.; McDonnell, M.J.; Abo-Leyah, H.; Aliberti, S.; Chalmers, J.D. A Comprehensive Analysis of the Impact of Pseudomonas aeruginosa Colonization on Prognosis in Adult Bronchiectasis. Ann. Am. Thorac Soc. 2015, 12, 1602–1611. [Google Scholar]
- Chalmers, J.D.; Goeminne, P.; Aliberti, S.; McDonnell, M.J.; Lonni, S.; Davidson, J.; Poppelwell, L.; Salih, W.; Pesci, A.; Dupont, L.J.; et al. The bronchiectasis severity index. An international derivation and validation study. Am. J. Respir. Crit Care Med. 2014, 189, 576–585. [Google Scholar] [CrossRef]
- Rogers, G.B.; van der Gast, C.J.; Cuthbertson, L.; Thomson, S.K.; Bruce, K.D.; Martin, M.L.; Serisier, D.J. Clinical measures of disease in adult non-CF bronchiectasis correlate with airway microbiota composition. Thorax 2013, 68, 731–737. [Google Scholar] [CrossRef]
- Rogers, G.B.; Zain, N.M.; Bruce, K.D.; Burr, L.D.; Chen, A.C.; Rivett, D.W.; McGuckin, M.A.; Serisier, D.J. A novel microbiota stratification system predicts future exacerbations in bronchiectasis. Ann. Am. Thorac Soc. 2014, 11, 496–503. [Google Scholar] [CrossRef] [PubMed]
- Cox, M.J.; Turek, E.M.; Hennessy, C.; Mirza, G.K.; James, P.L.; Coleman, M.; Jones, A.; Wilson, R.; Bilton, D.; Cookson, W.O.; et al. Longitudinal assessment of sputum microbiome by sequencing of the 16S rRNA gene in non-cystic fibrosis bronchiectasis patients. PLoS ONE 2017, 12, e0170622. [Google Scholar] [CrossRef]
- Chalmers, J.D.; Moffitt, K.L.; Suarez-Cuartin, G.; Sibila, O.; Finch, S.; Furrie, E.; Dicker, A.; Wrobel, K.; Elborn, J.S.; Walker, B.; et al. Neutrophil Elastase Activity Is Associated with Exacerbations and Lung Function Decline in Bronchiectasis. Am. J. Respir. Crit Care Med. 2017, 195, 1384–1393. [Google Scholar] [CrossRef] [PubMed]
- Taylor, S.L.; Rogers, G.B.; Chen, A.C.; Burr, L.D.; McGuckin, M.A.; Serisier, D.J. Matrix metalloproteinases vary with airway microbiota composition and lung function in non-cystic fibrosis bronchiectasis. Ann. Am. Thorac Soc. 2015, 12, 701–707. [Google Scholar] [CrossRef] [PubMed]
- Rogers, G.B.; Bruce, K.D.; Martin, M.L.; Burr, L.D.; Serisier, D.J. The effect of long-term macrolide treatment on respiratory microbiota composition in non-cystic fibrosis bronchiectasis: An analysis from the randomised, double-blind, placebo-controlled BLESS trial. Lancet Respir. Med. 2014, 2, 988–996. [Google Scholar] [CrossRef]
- Mac Aogain, M.; Chandrasekaran, R.; Lim, A.Y.H.; Low, T.B.; Tan, G.L.; Hassan, T.; Ong, T.H.; Hui Qi Ng, A.; Bertrand, D.; Koh, J.Y.; et al. Immunological corollary of the pulmonary mycobiome in bronchiectasis: The CAMEB study. Eur. Respir. J. 2018, 52, 1800766. [Google Scholar] [CrossRef]
- Namkoong, H.; Kurashima, A.; Morimoto, K.; Hoshino, Y.; Hasegawa, N.; Ato, M.; Mitarai, S. Epidemiology of Pulmonary Nontuberculous Mycobacterial Disease, Japan. Emerg. Infect. Dis. 2016, 22, 1116–1117. [Google Scholar] [CrossRef]
- Prevots, D.R.; Marras, T.K. Epidemiology of human pulmonary infection with nontuberculous mycobacteria: A review. Clin. Chest Med. 2015, 36, 13–34. [Google Scholar] [CrossRef]
- Yamasaki, K.; Mukae, H.; Kawanami, T.; Fukuda, K.; Noguchi, S.; Akata, K.; Naito, K.; Oda, K.; Ogoshi, T.; Nishida, C.; et al. Possible role of anaerobes in the pathogenesis of nontuberculous mycobacterial infection. Respirology 2015, 20, 758–765. [Google Scholar] [CrossRef]
- Sulaiman, I.; Wu, B.G.; Li, Y.; Scott, A.S.; Malecha, P.; Scaglione, B.; Wang, J.; Basavaraj, A.; Chung, S.; Bantis, K.; et al. Evaluation of the airway microbiome in nontuberculous mycobacteria disease. Eur. Respir. J. 2018, 52, 1800810. [Google Scholar] [CrossRef] [PubMed]
- Cowman, S.A.; James, P.; Wilson, R.; Cookson, W.O.C.; Moffatt, M.F.; Loebinger, M.R. Profiling mycobacterial communities in pulmonary nontuberculous mycobacterial disease. PLoS ONE 2018, 13, e0208018. [Google Scholar] [CrossRef] [PubMed]
- Aliberti, S.; Masefield, S.; Polverino, E.; De Soyza, A.; Loebinger, M.R.; Menendez, R.; Ringshausen, F.C.; Vendrell, M.; Powell, P.; Chalmers, J.D.; et al. Research priorities in bronchiectasis: A consensus statement from the EMBARC Clinical Research Collaboration. Eur. Respir. J. 2016, 48, 632–647. [Google Scholar] [CrossRef] [PubMed]
- Thornton, C.S.; Mellett, M.; Jarand, J.; Barss, L.; Field, S.K.; Fisher, D.A. The respiratory microbiome and nontuberculous mycobacteria: An emerging concern in human health. Eur. Respir. Rev. 2021, 30, 200299. [Google Scholar] [CrossRef] [PubMed]
- Perrone, F.; Belluomini, L.; Mazzotta, M.; Bianconi, M.; Di Noia, V.; Meacci, F.; Montrone, M.; Pignataro, D.; Prelaj, A.; Rinaldi, S.; et al. Exploring the role of respiratory microbiome in lung cancer: A systematic review. Crit Rev. Oncol. Hematol. 2021, 164, 103404. [Google Scholar] [CrossRef]
- Gomes, S.; Cavadas, B.; Ferreira, J.C.; Marques, P.I.; Monteiro, C.; Sucena, M.; Sousa, C.; Vaz Rodrigues, L.; Teixeira, G.; Pinto, P.; et al. Profiling of lung microbiota discloses differences in adenocarcinoma and squamous cell carcinoma. Sci. Rep. 2019, 9, 12838. [Google Scholar] [CrossRef]
- Jin, J.; Gan, Y.; Liu, H.; Wang, Z.; Yuan, J.; Deng, T.; Zhou, Y.; Zhu, Y.; Zhu, H.; Yang, S.; et al. Diminishing microbiome richness and distinction in the lower respiratory tract of lung cancer patients: A multiple comparative study design with independent validation. Lung Cancer 2019, 136, 129–135. [Google Scholar] [CrossRef]
- Huang, D.; Su, X.; Yuan, M.; Zhang, S.; He, J.; Deng, Q.; Qiu, W.; Dong, H.; Cai, S. The characterization of lung microbiome in lung cancer patients with different clinicopathology. Am. J. Cancer Res. 2019, 9, 2047–2063. [Google Scholar]
- Siegel, R.L.; Miller, K.D.; Jemal, A. Cancer statistics, 2020. CA Cancer J. Clin. 2020, 70, 7–30. [Google Scholar] [CrossRef]
- Bray, F.; Ferlay, J.; Soerjomataram, I.; Siegel, R.L.; Torre, L.A.; Jemal, A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 2018, 68, 394–424. [Google Scholar] [CrossRef]
- Asai, N.; Ohkuni, Y.; Kaneko, N.; Yamaguchi, E.; Kubo, A. Relapsed small cell lung cancer: Treatment options and latest developments. Ther Adv. Med. Oncol. 2014, 6, 69–82. [Google Scholar] [CrossRef]
- Wang, K.; Huang, Y.; Zhang, Z.; Liao, J.; Ding, Y.; Fang, X.; Liu, L.; Luo, J.; Kong, J. A Preliminary Study of Microbiota Diversity in Saliva and Bronchoalveolar Lavage Fluid from Patients with Primary Bronchogenic Carcinoma. Med. Sci. Monit. 2019, 25, 2819–2834. [Google Scholar] [CrossRef] [PubMed]
- Weiss, G.; Cocking, J.; Bilagody, C.; Hornstra, H.; Kaufman, E.; Nader, D.; Turner, J.; Chandrika, S.; Caporaso, J.; Keim, P.; et al. P1.04-019 Final Analysis of Lung Microbiome from Patients Undergoing Bronchoscopy. J. Thoracic Oncol. 2017, 12, S606–S607. [Google Scholar] [CrossRef][Green Version]
- Zhang, C.; Wang, J.; Sun, Z.; Cao, Y.; Mu, Z.; Ji, X. Commensal microbiota contributes to predicting the response to immune checkpoint inhibitors in non-small-cell lung cancer patients. Cancer Sci. 2021, 112, 3005. [Google Scholar] [CrossRef]
- Huang, C.; Wang, Y.; Li, X.; Ren, L.; Zhao, J.; Hu, Y.; Zhang, L.; Fan, G.; Xu, J.; Gu, X.; et al. Clinical features of patients infected with 2019 novel coronavirus in Wuhan, China. Lancet 2020, 395, 497–506. [Google Scholar] [CrossRef]
- Lai, C.C.; Shih, T.P.; Ko, W.C.; Tang, H.J.; Hsueh, P.R. Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) and coronavirus disease-2019 (COVID-19): The epidemic and the challenges. Int J. Antimicrob. Agents 2020, 55, 105924. [Google Scholar] [CrossRef] [PubMed]
- Sulaiman, I.; Chung, M.; Angel, L.; Koralov, S.; Wu, B.; Yeung, S.; Krolikowski, K.; Li, Y.; Duerr, R.; Schluger, R.; et al. Microbial signatures in the lower airways of mechanically ventilated COVID19 patients associated with poor clinical outcome. Res. Sq. 2021. [Google Scholar] [CrossRef]
- Khatiwada, S.; Subedi, A. Lung microbiome and coronavirus disease 2019 (COVID-19): Possible link and implications. Hum. Microb. J. 2020, 17, 100073. [Google Scholar] [CrossRef]
- Gaibani, P.; Viciani, E.; Bartoletti, M.; Lewis, R.E.; Tonetti, T.; Lombardo, D.; Castagnetti, A.; Bovo, F.; Horna, C.S.; Ranieri, M.; et al. The lower respiratory tract microbiome of critically ill patients with COVID-19. Sci. Rep. 2021, 11, 10103. [Google Scholar] [CrossRef] [PubMed]
- Man, W.H.; de Steenhuijsen Piters, W.A.; Bogaert, D. The microbiota of the respiratory tract: Gatekeeper to respiratory health. Nat. Rev. Microbiol. 2017, 15, 259–270. [Google Scholar] [CrossRef] [PubMed]
- Yeoh, Y.K.; Zuo, T.; Lui, G.C.; Zhang, F.; Liu, Q.; Li, A.Y.; Chung, A.C.; Cheung, C.P.; Tso, E.Y.; Fung, K.S.; et al. Gut microbiota composition reflects disease severity and dysfunctional immune responses in patients with COVID-19. Gut 2021, 70, 698–706. [Google Scholar] [CrossRef] [PubMed]
- Gu, S.; Chen, Y.; Wu, Z.; Chen, Y.; Gao, H.; Lv, L.; Guo, F.; Zhang, X.; Luo, R.; Huang, C.; et al. Alterations of the Gut Microbiota in Patients With Coronavirus Disease 2019 or H1N1 Influenza. Clin. Infect. Dis. 2020, 71, 2669–2678. [Google Scholar] [CrossRef] [PubMed]
- Lee, S.; Ishitsuka, A.; Noguchi, M.; Hirohama, M.; Fujiyasu, Y.; Petric, P.P.; Schwemmle, M.; Staeheli, P.; Nagata, K.; Kawaguchi, A. Influenza restriction factor MxA functions as inflammasome sensor in the respiratory epithelium. Sci. Immunol. 2019, 4. [Google Scholar] [CrossRef]
- Tang, H.; Bohannon, L.; Lew, M.; Jensen, D.; Jung, S.H.; Zhao, A.; Sung, A.D.; Wischmeyer, P.E. Randomised, double-blind, placebo-controlled trial of Probiotics To Eliminate COVID-19 Transmission in Exposed Household Contacts (PROTECT-EHC): A clinical trial protocol. BMJ Open 2021, 11, e047069. [Google Scholar] [CrossRef]
- Peng, J.; Zhang, M.; Yao, G.; Kwok, L.Y.; Zhang, W. Probiotics as Adjunctive Treatment for Patients Contracted COVID-19: Current Understanding and Future Needs. Front. Nutr. 2021, 8, 669808. [Google Scholar] [CrossRef] [PubMed]
- Carney, S.M.; Clemente, J.C.; Cox, M.J.; Dickson, R.P.; Huang, Y.J.; Kitsios, G.D.; Kloepfer, K.M.; Leung, J.M.; LeVan, T.D.; Molyneaux, P.L.; et al. Methods in Lung Microbiome Research. Am. J. Respir. Cell Mol. Biol. 2020, 62, 283–299. [Google Scholar] [CrossRef] [PubMed]
- Cox, M.J.; Ege, M.J.; von Mutius, E.; Ege, M.J.; von Mutius, E.; von Mutius, E. Challenges, impact and the future. ERS Monogr. ERS Monograph. 2019, 2019, 240–244. [Google Scholar]
- Segal, L.N.; Alekseyenko, A.V.; Clemente, J.C.; Kulkarni, R.; Wu, B.; Gao, Z.; Chen, H.; Berger, K.I.; Goldring, R.M.; Rom, W.N.; et al. Enrichment of lung microbiome with supraglottic taxa is associated with increased pulmonary inflammation. Microbiome 2013, 1, 19. [Google Scholar] [CrossRef]
- Wu, B.G.; Sulaiman, I.; Tsay, J.J.; Perez, L.; Franca, B.; Li, Y.; Wang, J.; Gonzalez, A.N.; El-Ashmawy, M.; Carpenito, J.; et al. Episodic Aspiration with Oral Commensals Induces a MyD88-dependent, Pulmonary T-Helper Cell Type 17 Response that Mitigates Susceptibility to Streptococcus pneumoniae. Am. J. Respir. Crit Care Med. 2021, 203, 1099–1111. [Google Scholar] [CrossRef] [PubMed]
- Tsay, J.J.; Wu, B.G.; Badri, M.H.; Clemente, J.C.; Shen, N.; Meyn, P.; Li, Y.; Yie, T.A.; Lhakhang, T.; Olsen, E.; et al. Airway Microbiota Is Associated with Upregulation of the PI3K Pathway in Lung Cancer. Am. J. Respir. Crit. Care Med. 2018, 198, 1188–1198. [Google Scholar] [CrossRef] [PubMed]
- Garg, N.; Wang, M.; Hyde, E.; da Silva, R.R.; Melnik, A.V.; Protsyuk, I.; Bouslimani, A.; Lim, Y.W.; Wong, R.; Humphrey, G.; et al. Three-Dimensional Microbiome and Metabolome Cartography of a Diseased Human Lung. Cell Host Microbe 2017, 22, 705–716.e4. [Google Scholar] [CrossRef] [PubMed]
- Dolma, K.; Summerlin, T.L.; Wongprasert, H.; Lal, C.V.; Philips Iii, J.B.; Winter, L. Early-Onset Neonatal Sepsis with Extended Spectrum Beta-Lactamase Producing Escherichia Coli in Infants Born to South and South East Asian Immigrants: A Case Series. AJP Rep. 2018, 8, e277–e279. [Google Scholar] [CrossRef] [PubMed]
- Wei, X.; Jiang, P.; Liu, J.; Sun, R.; Zhu, L. Association between probiotic supplementation and asthma incidence in infants: A meta-analysis of randomized controlled trials. J. Asthma 2020, 57, 167–178. [Google Scholar] [CrossRef] [PubMed]
- Van de Pol, M.A.; Lutter, R.; Smids, B.S.; Weersink, E.J.; van der Zee, J.S. Synbiotics reduce allergen-induced T-helper 2 response and improve peak expiratory flow in allergic asthmatics. Allergy 2011, 66, 39–47. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yagi, K.; Huffnagle, G.B.; Lukacs, N.W.; Asai, N. The Lung Microbiome during Health and Disease. Int. J. Mol. Sci. 2021, 22, 10872. https://doi.org/10.3390/ijms221910872
Yagi K, Huffnagle GB, Lukacs NW, Asai N. The Lung Microbiome during Health and Disease. International Journal of Molecular Sciences. 2021; 22(19):10872. https://doi.org/10.3390/ijms221910872
Chicago/Turabian StyleYagi, Kazuma, Gary B. Huffnagle, Nicholas W. Lukacs, and Nobuhiro Asai. 2021. "The Lung Microbiome during Health and Disease" International Journal of Molecular Sciences 22, no. 19: 10872. https://doi.org/10.3390/ijms221910872
APA StyleYagi, K., Huffnagle, G. B., Lukacs, N. W., & Asai, N. (2021). The Lung Microbiome during Health and Disease. International Journal of Molecular Sciences, 22(19), 10872. https://doi.org/10.3390/ijms221910872





