Establishment of Bovine-Induced Pluripotent Stem Cells
Abstract
:1. Introduction
2. Results
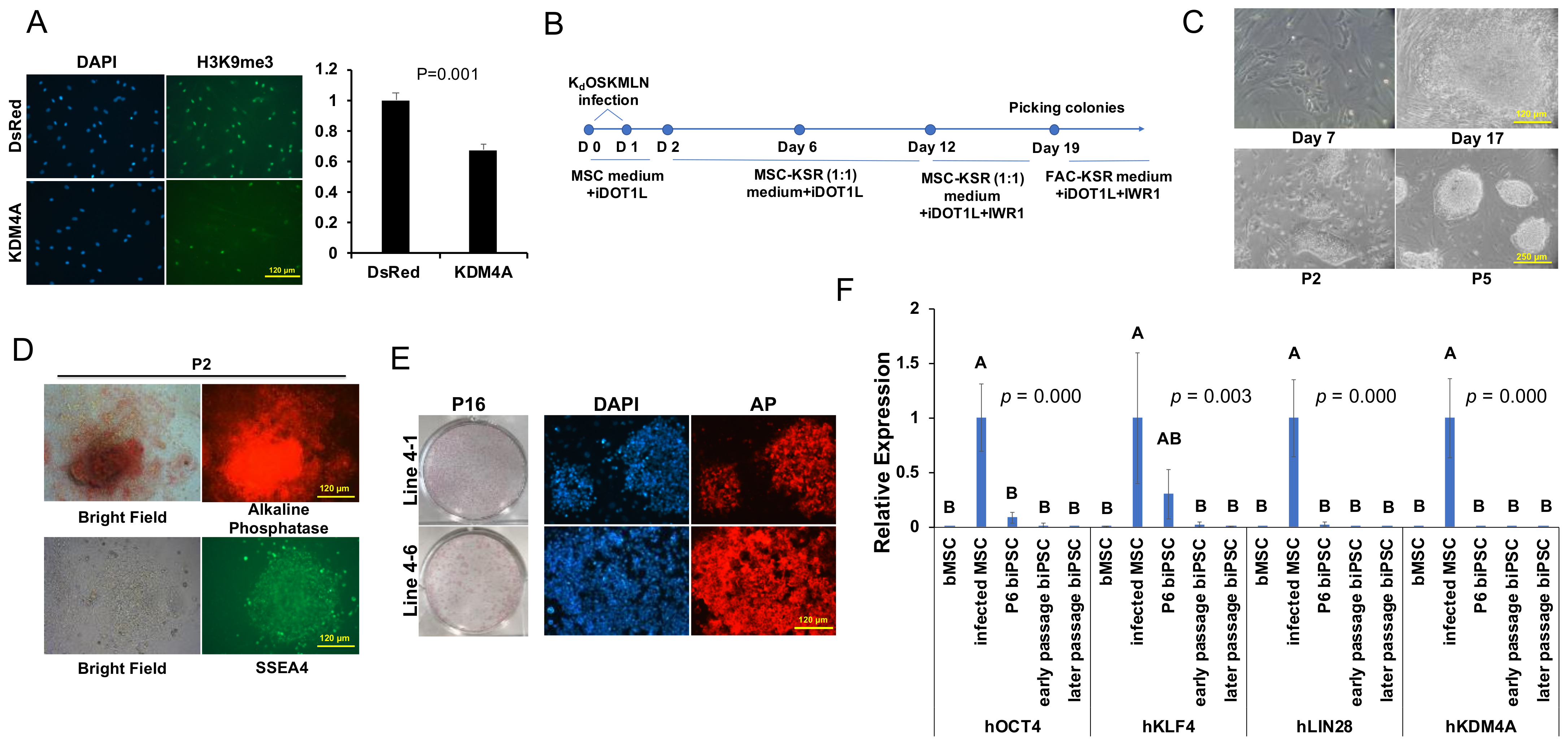
2.1. Establishment of Bovine iPSCs
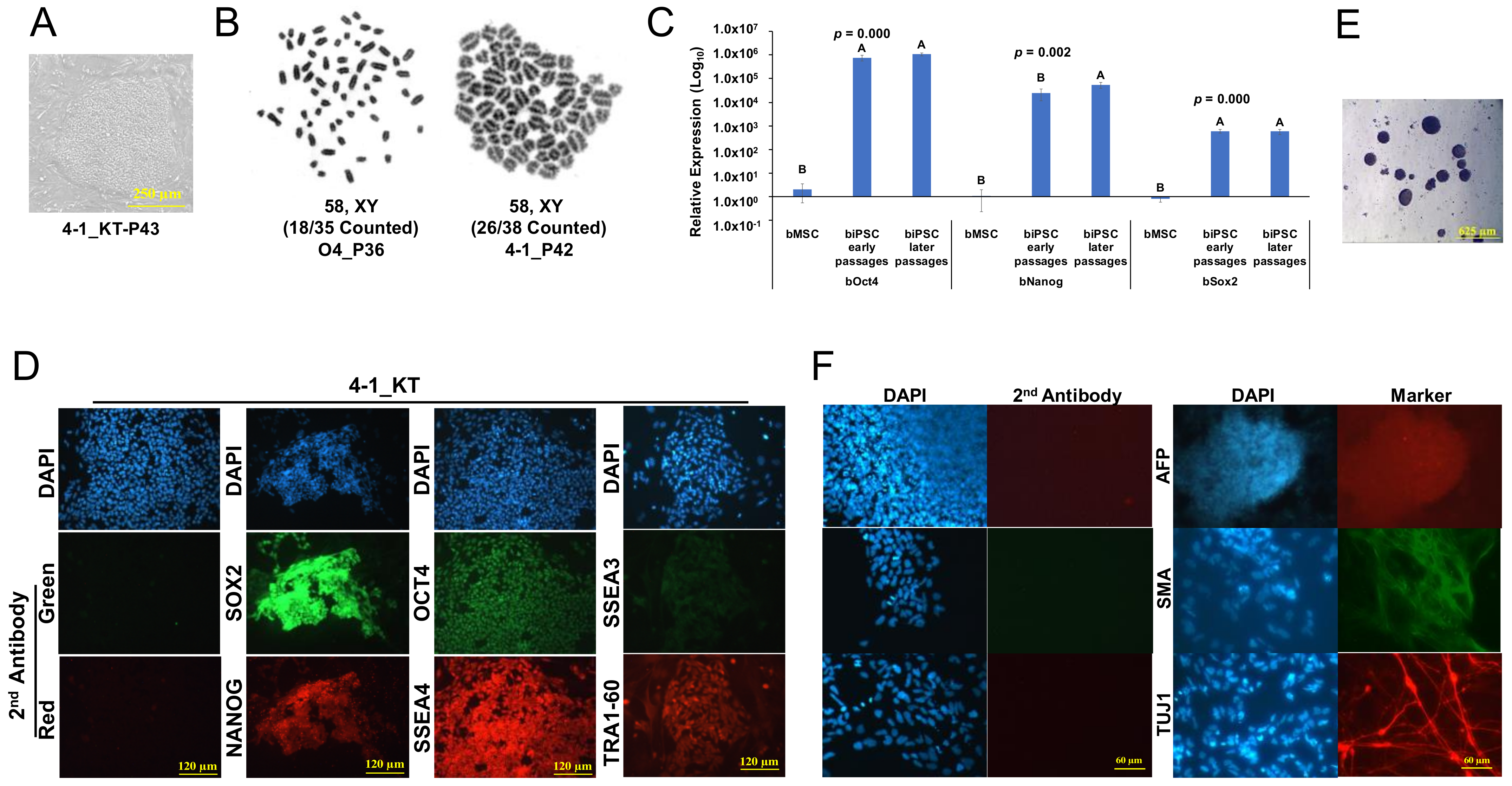
2.2. Characterization of Primed-like Bovine iPSCs
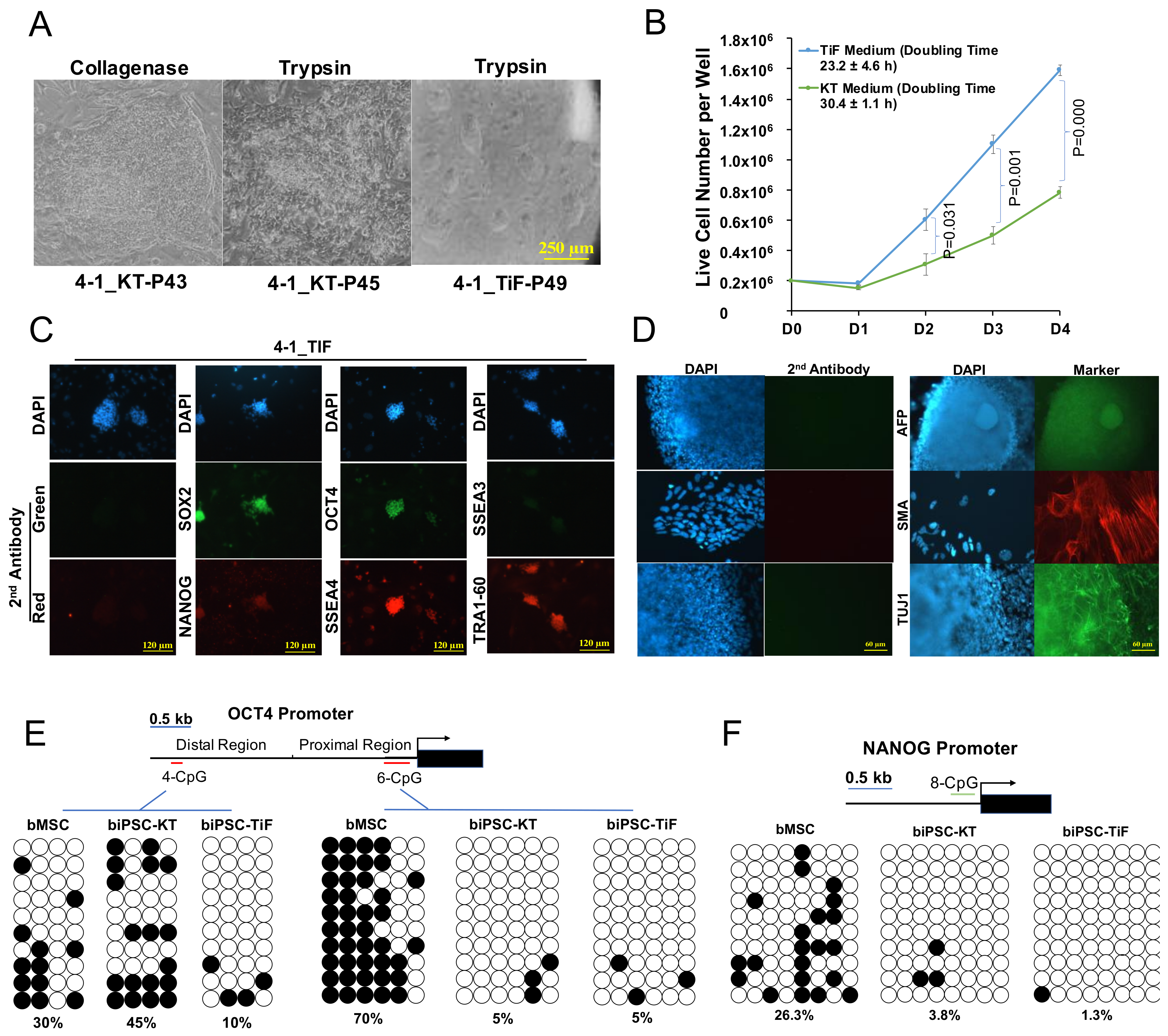
2.3. Development and Characterization of Naïve-Like Bovine iPSCs
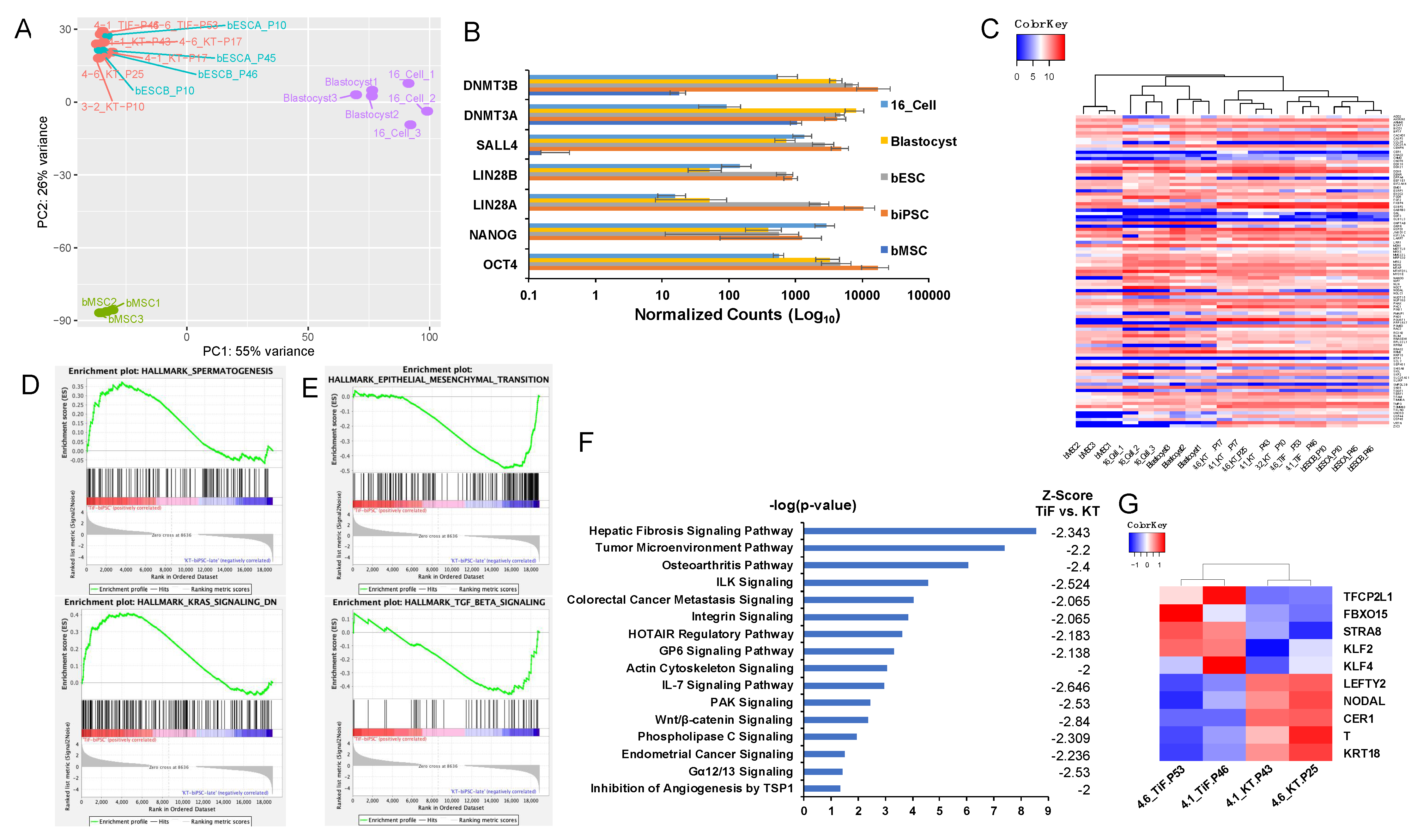
2.4. Global Transcriptome Analysis of Bovine iPSCs
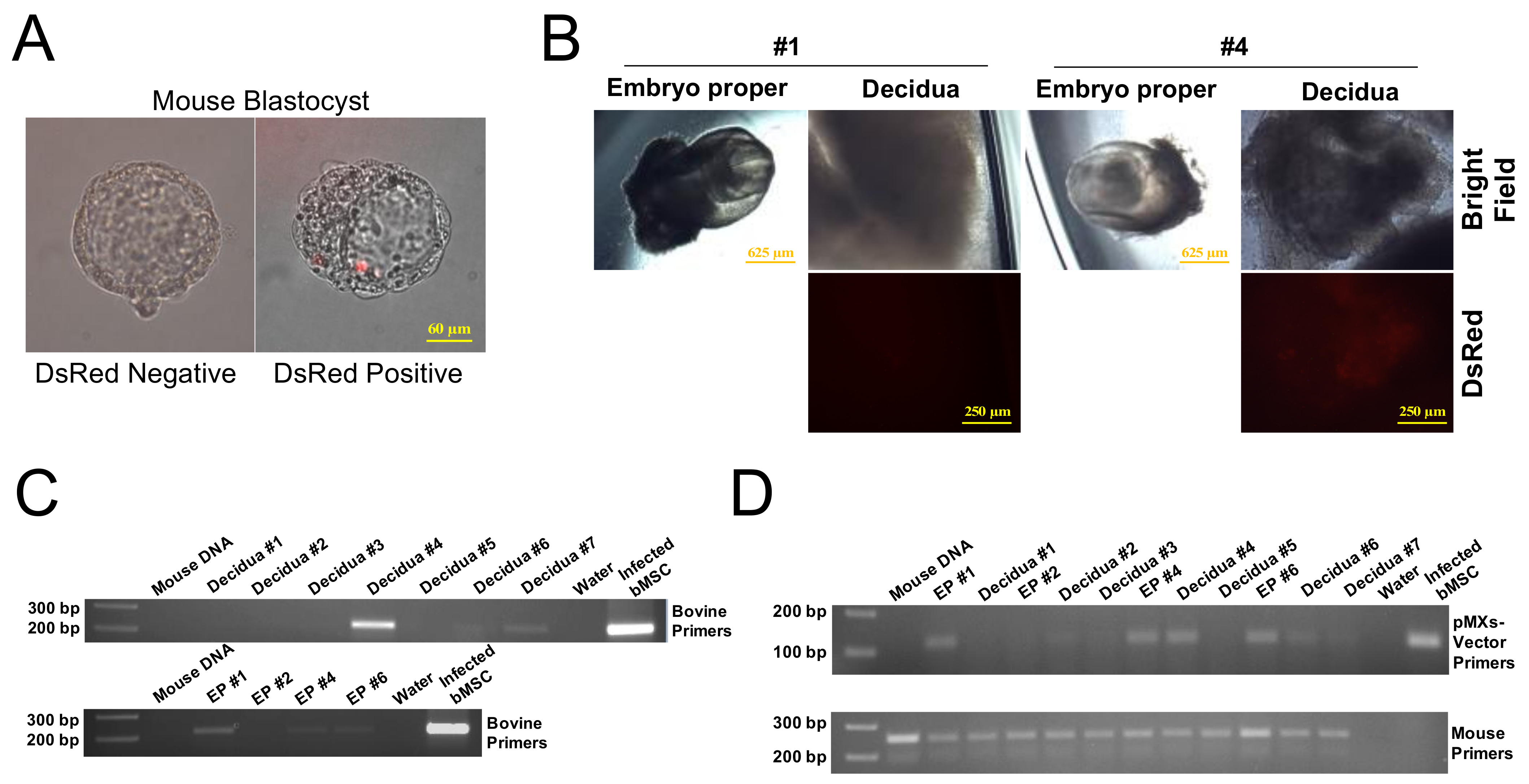
2.5. In Vivo Chimerism Capacity of Bovine iPSCs in Mouse Embryos
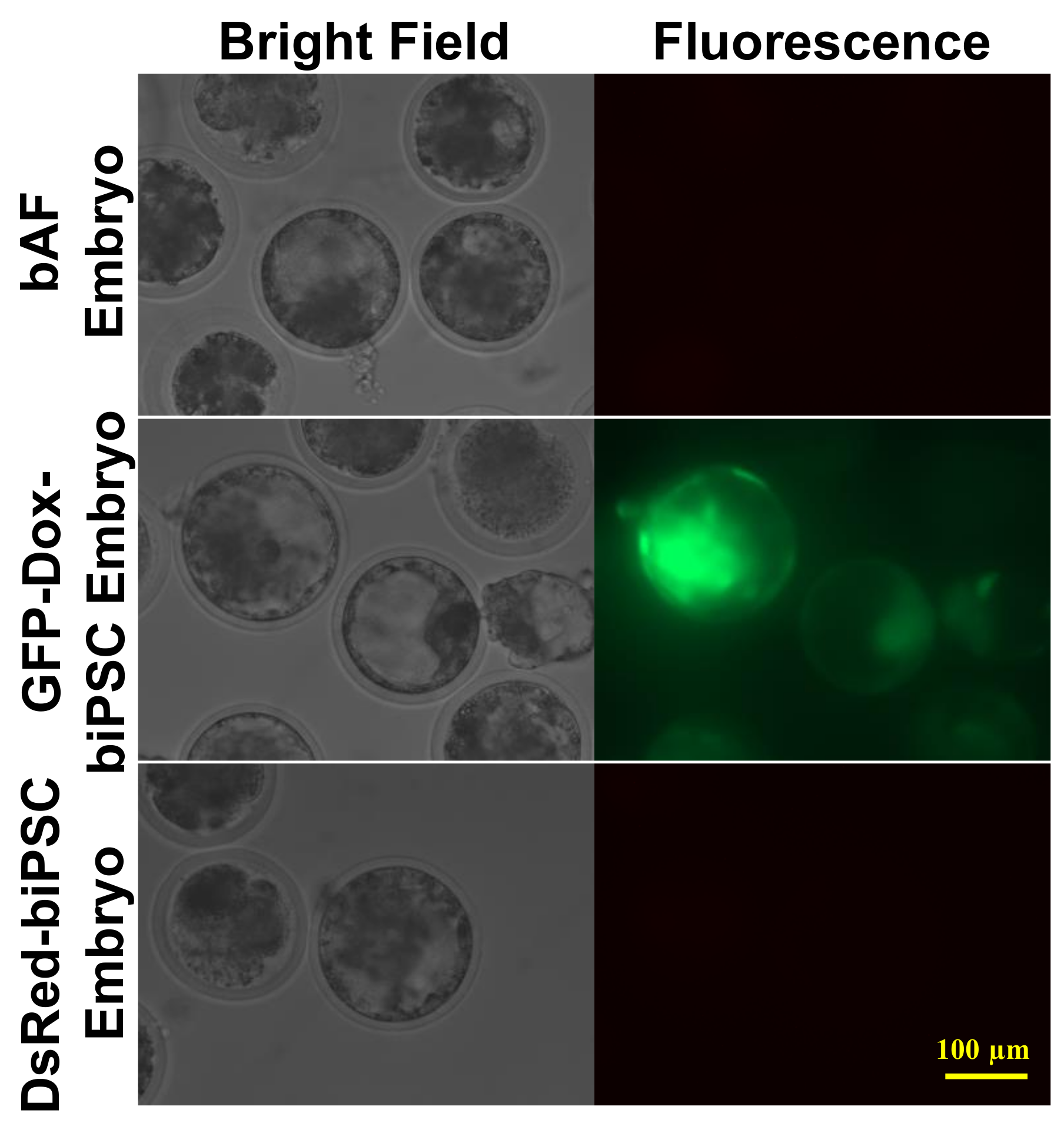
2.6. Efficiency of biPSCs as Donors for Somatic Cell Nuclear Transfer (SCNT)
3. Discussion
4. Materials and Methods
4.1. Chemicals, DNA Constructs, and Primary Bovine Cells
4.2. Reprogramming of bMSCs
4.3. Quantitative Reverse Transcription-PCR (qRT-PCR) Analysis
4.4. Embryoid Body (EB) Differentiation
4.5. Immunostaining
4.6. RNA-seq Data Analysis
4.7. Karyotyping
4.8. Bisulfite Sequencing
4.9. Aggregation to Generate Chimeric Embryos
4.10. Nuclear Transfer Using biPSCs as Nuclear Donors
4.11. Statistial Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Data Availability Statement
Conflicts of Interest
References
- Okita, K.; Ichisaka, T.; Yamanaka, S. Generation of germline-competent induced pluripotent stem cells. Nat. Cell Biol. 2007, 448, 313–317. [Google Scholar] [CrossRef]
- Takahashi, K.; Yamanaka, S. Induction of Pluripotent Stem Cells from Mouse Embryonic and Adult Fibroblast Cultures by Defined Factors. Cell 2006, 126, 663–676. [Google Scholar] [CrossRef] [Green Version]
- Yu, J.; Vodyanik, M.A.; Smuga-Otto, K.; Antosiewicz-Bourget, J.; Frane, J.L.; Tian, S.; Nie, J.; Jonsdottir, G.A.; Ruotti, V.; Stewart, R.; et al. Induced pluripotent stem cell lines derived from human somatic cells. Science 2007, 318, 1917–1920. [Google Scholar] [CrossRef]
- Stice, S.L.; Strelchenko, N.S.; Keefer, C.; Matthews, L. Pluripotent bovine embryonic cell lines direct embryonic development following nuclear transfer. Biol. Reprod. 1996, 54, 100–110. [Google Scholar] [CrossRef] [Green Version]
- Gong, G.; Roach, M.L.; Jiang, L.; Yang, X.; Tian, X.C. Culture Conditions and Enzymatic Passaging of Bovine ESC-Like Cells. Cell. Reprogram. 2010, 12, 151–160. [Google Scholar] [CrossRef]
- Verma, V.; Huang, B.; Kallingappa, P.K.; Oback, B. Dual Kinase Inhibition Promotes Pluripotency in Finite Bovine Embryonic Cell Lines. Stem Cells Dev. 2013, 22, 1728–1742. [Google Scholar] [CrossRef]
- Maruotti, J.; Munoz, M.; Degrelle, S.A.; Gomez, E.; Louet, C.; Diez, C.; de Longchamp, P.H.; Brochard, V.; Hue, I.; Caamano, J.N.; et al. Efficient derivation of bovine embryonic stem cells needs more than active core pluripotency factors. Mol. Reprod. Dev. 2012, 79, 461–477. [Google Scholar] [CrossRef]
- Bogliotti, Y.S.; Wu, J.; Vilarino, M.; Okamura, D.; Soto, D.A.; Zhong, C.; Sakurai, M.; Sampaio, R.; Suzuki, K.; Belmonte, J.C.I.; et al. Efficient derivation of stable primed pluripotent embryonic stem cells from bovine blastocysts. Proc. Natl. Acad. Sci. USA 2018, 115, 2090–2095. [Google Scholar] [CrossRef] [Green Version]
- Deng, Y.; Liu, Q.; Luo, C.; Chen, S.; Li, X.; Wang, C.; Liu, Z.; Lei, X.; Zhang, H.; Sun, H.; et al. Generation of Induced Pluripotent Stem Cells from Buffalo (Bubalus bubalis) Fetal Fibroblasts with Buffalo Defined Factors. Stem Cells Dev. 2012, 21, 2485–2494. [Google Scholar] [CrossRef] [Green Version]
- Cao, H.; Yang, P.; Pu, Y.; Sun, X.; Yin, H.; Zhang, Y.; Zhang, Y.; Li, Y.; Liu, Y.; Fang, F.; et al. Characterization of Bovine Induced Pluripotent Stem Cells by Lentiviral Transduction of Reprogramming Factor Fusion Proteins. Int. J. Biol. Sci. 2012, 8, 498–511. [Google Scholar] [CrossRef]
- Sumer, H.; Liu, J.; Malaver-Ortega, L.F.; Lim, M.L.; Khodadadi, K.; Verma, P.J. NANOG is a key factor for induction of pluripotency in bovine adult fibroblasts1. J. Anim. Sci. 2011, 89, 2708–2716. [Google Scholar] [CrossRef] [Green Version]
- Han, X.; Han, J.; Ding, F.; Cao, S.; Lim, S.S.; Dai, Y.; Zhang, R.; Zhang, Y.; Lim, B.; Li, N. Generation of induced pluripotent stem cells from bovine embryonic fibroblast cells. Cell Res. 2011, 21, 1509–1512. [Google Scholar] [CrossRef] [Green Version]
- Talluri, T.R.; Kumar, D.; Glage, S.; Garrels, W.; Ivics, Z.; Debowski, K.; Behr, R.; Niemann, H.; Kues, W.A. Derivation and Characterization of Bovine Induced Pluripotent Stem Cells by Transposon-Mediated Reprogramming. Cell. Reprogram. 2015, 17, 131–140. [Google Scholar] [CrossRef] [Green Version]
- Malaver-Ortega, L.F.; Sumer, H.; Liu, J.; Verma, P.J. Inhibition of JAK-STAT ERK/MAPK and Glycogen Synthase Kinase-3 Induces a Change in Gene Expression Profile of Bovine Induced Pluripotent Stem Cells. Stem Cells Int. 2016, 2016, 5127984. [Google Scholar] [CrossRef] [Green Version]
- Kawaguchi, T.; Tsukiyama, T.; Kimura, K.; Matsuyama, S.; Minami, N.; Yamada, M.; Imai, H. Generation of Naïve Bovine Induced Pluripotent Stem Cells Using PiggyBac Transposition of Doxycycline-Inducible Transcription Factors. PLoS ONE 2015, 10, e0135403. [Google Scholar] [CrossRef] [Green Version]
- Heo, Y.T.; Quan, X.; Xu, Y.N.; Baek, S.; Choi, H.; Kim, N.-H.; Kim, J. CRISPR/Cas9 Nuclease-Mediated Gene Knock-In in Bovine-Induced Pluripotent Cells. Stem Cells Dev. 2015, 24, 393–402. [Google Scholar] [CrossRef]
- Lin, Y.-C.; Kuo, K.-K.; Wuputra, K.; Lin, S.-H.; Ku, C.-C.; Yang, Y.-H.; Wang, S.-W.; Wang, S.-W.; Wu, D.-C.; Wu, C.-C.; et al. Bovine Induced Pluripotent Stem Cells Are More Resistant to Apoptosis than Testicular Cells in Response to Mono-(2-ethylhexyl) Phthalate. Int. J. Mol. Sci. 2014, 15, 5011–5031. [Google Scholar] [CrossRef] [Green Version]
- Wang, S.-W.; Wang, S.S.-W.; Wu, D.-C.; Lin, Y.-C.; Ku, C.-C.; Wu, C.-C.; Chai, C.-Y.; Lee, J.-N.; Tsai, E.-M.; Lin, C.-L.; et al. Androgen receptor-mediated apoptosis in bovine testicular induced pluripotent stem cells in response to phthalate esters. Cell Death Dis. 2013, 4, e907. [Google Scholar] [CrossRef] [Green Version]
- Bressan, F.F.; Bassanezze, V.; Pessôa, L.V.D.F.; Sacramento, C.B.; Malta, T.M.; Kashima, S.; Neto, P.F.; Strefezzi, R.D.F.; Pieri, N.C.G.; Krieger, J.E.; et al. Generation of induced pluripotent stem cells from large domestic animals. Stem Cell Res. Ther. 2020, 11, 247. [Google Scholar] [CrossRef]
- Pillai, V.V.; Kei, T.G.; Reddy, S.E.; Das, M.; Abratte, C.; Cheong, S.H.; Selvaraj, V. Induced pluripotent stem cell generation from bovine somatic cells indicates unmet needs for pluripotency sustenance. Anim. Sci. J. 2019, 90, 1149–1160. [Google Scholar] [CrossRef]
- Kawaguchi, T.; Cho, D.; Hayashi, M.; Tsukiyama, T.; Kimura, K.; Matsuyama, S.; Minami, N.; Yamada, M.; Imai, H. Derivation of Induced Trophoblast Cell Lines in Cattle by Doxycycline-Inducible piggyBac Vectors. PLoS ONE 2016, 11, e0167550. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Canizo, J.R.; Vazquez Echegaray, C.; Klisch, D.; Aller, J.F.; Paz, D.A.; Alberio, R.H.; Alberio, R.; Guberman, A.S. Exogenous human OKSM factors maintain pluripotency gene expression of bovine and porcine iPS-like cells obtained with STEMCCA delivery system. BMC Res. Notes 2018, 11, 509. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Talbot, N.C.; Sparks, W.O.; Phillips, C.E.; Ealy, A.D.; Powell, A.M.; Caperna, T.J.; Garrett, W.M.; Donovan, D.M.; Blomberg, L.A. Bovine trophectoderm cells induced from bovine fibroblasts with induced pluripotent stem cell reprogramming factors. Mol. Reprod. Dev. 2017, 84, 468–485. [Google Scholar] [CrossRef] [PubMed]
- Su, Y.; Zhu, J.; Salman, S.; Tang, Y. Induced pluripotent stem cells from farm animals. J. Anim. Sci. 2020, 98, skaa343. [Google Scholar] [CrossRef]
- Zhao, L.; Gao, X.; Zheng, Y.; Wang, Z.; Zhao, G.; Ren, J.; Zhang, J.; Wu, J.; Wu, B.; Chen, Y.; et al. Establishment of bovine expanded potential stem cells. Proc. Natl. Acad. Sci. USA 2021, 118, 15. [Google Scholar] [CrossRef]
- Van Eenennaam, A.L. Application of genome editing in farm animals: Cattle. Transgenic Res. 2019, 28, 93–100. [Google Scholar] [CrossRef]
- Yuan, Y. Capturing bovine pluripotency. Proc. Natl. Acad. Sci. USA 2018, 115, 1962–1963. [Google Scholar] [CrossRef] [Green Version]
- Kehrli, M.E.; Ackermann, M.R.; Shuster, D.E.; Van Der Maaten, M.J.; Schmalstieg, F.C.; Anderson, D.C.; Hughes, B.J. Bovine leukocyte adhesion deficiency. Beta 2 integrin deficiency in young Holstein cattle. Am. J. Pathol. 1992, 140, 1489–1492. [Google Scholar]
- Casal, M.; Haskins, M. Large animal models and gene therapy. Eur. J. Hum. Genet. 2006, 14, 266–272. [Google Scholar] [CrossRef]
- Wang, L.; Su, Y.; Huang, C.; Yin, Y.; Chu, A.; Knupp, A.; Tang, Y. NANOG and LIN28 dramatically improve human cell reprogramming by modulating LIN41 and canonical WNT activities. Biol. Open 2019, 8, 12. [Google Scholar] [CrossRef] [Green Version]
- Ludwig, T.E.; Levenstein, M.E.; Jones, J.M.; Berggren, W.T.; Mitchen, E.R.; Frane, J.L.; Crandall, L.J.; Daigh, C.A.; Conard, K.R.; Piekarczyk, M.S.; et al. Derivation of human embryonic stem cells in defined conditions. Nat. Biotechnol. 2006, 24, 185–187. [Google Scholar] [CrossRef] [PubMed]
- Tsukiyama, T.; Ohinata, Y. A Modified EpiSC Culture Condition Containing a GSK3 Inhibitor Can Support Germline-Competent Pluripotency in Mice. PLoS ONE 2014, 9, e95329. [Google Scholar] [CrossRef] [PubMed]
- Wu, J.; Luengo, A.P.; Sakurai, M.; Sugawara, A.; Gil, M.A.; Yamauchi, T.; Suzuki, K.; Bogliotti, Y.S.; Cuello, C.; Valencia, M.M.; et al. Interspecies Chimerism with Mammalian Pluripotent Stem Cells. Cell 2017, 168, 473–486.e15. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Matoba, S.; Liu, Y.; Lu, F.; Iwabuchi, K.A.; Shen, L.; Inoue, A.; Zhang, Y. Embryonic Development following Somatic Cell Nuclear Transfer Impeded by Persisting Histone Methylation. Cell 2014, 159, 884–895. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chen, J.; Liu, H.; Liu, J.; Qi, J.; Wei, B.; Yang, J.; Liang, H.; Chen, Y.; Chen, J.; Wu, Y.; et al. H3K9 methylation is a barrier during somatic cell reprogramming into iPSCs. Nat. Genet. 2013, 45, 34–42. [Google Scholar] [CrossRef] [PubMed]
- Onder, T.T.; Kara, N.; Cherry, A.; Sinha, A.U.; Zhu, N.; Bernt, K.M.; Cahan, P.; Mancarci, B.O.; Unternaehrer, J.; Gupta, P.B.; et al. Chromatin-modifying enzymes as modulators of reprogramming. Nature 2012, 483, 598–602. [Google Scholar] [CrossRef]
- Soufi, A.; Donahue, G.; Zaret, K.S. Facilitators and Impediments of the Pluripotency Reprogramming Factors’ Initial Engagement with the Genome. Cell 2012, 151, 994–1004. [Google Scholar] [CrossRef] [Green Version]
- Maherali, N.; Sridharan, R.; Xie, W.; Utikal, J.; Eminli, S.; Arnold, K.; Stadtfeld, M.; Yachechko, R.; Tchieu, J.; Jaenisch, R.; et al. Directly Reprogrammed Fibroblasts Show Global Epigenetic Remodeling and Widespread Tissue Contribution. Cell Stem Cell 2007, 1, 55–70. [Google Scholar] [CrossRef] [Green Version]
- Wernig, M.; Meissner, A.; Foreman, R.; Brambrink, T.; Ku, M.; Hochedlinger, K.; Bernstein, B.E.; Jaenisch, R. In vitro reprogramming of fibroblasts into a pluripotent ES-cell-like state. Nat. Cell Biol. 2007, 448, 318–324. [Google Scholar] [CrossRef]
- Thomson, J.A.; Itskovitz-Eldor, J.; Shapiro, S.S.; Waknitz, M.A.; Swiergiel, J.J.; Marshall, V.S.; Jones, J.M. Embryonic Stem Cell Lines Derived from Human Blastocysts. Science 1998, 282, 1145–1147. [Google Scholar] [CrossRef] [Green Version]
- Camus, A.; Perea-Gomez, A.; Moreau, A.; Collignon, J. Absence of Nodal signaling promotes precocious neural differentiation in the mouse embryo. Dev. Biol. 2006, 295, 743–755. [Google Scholar] [CrossRef] [Green Version]
- James, D.; Levine, A.J.; Besser, D.; Hemmati-Brivanlou, A. TGFbeta/activin/nodal signaling is necessary for the maintenance of pluripotency in human embryonic stem cells. Development 2005, 132, 1273–1282. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Vallier, L.; Reynolds, D.; Pedersen, R.A. Nodal inhibits differentiation of human embryonic stem cells along the neuroectodermal default pathway. Dev. Biol. 2004, 275, 403–421. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ying, Q.-L.; Wray, J.; Nichols, J.; Batlle-Morera, L.; Doble, B.; Woodgett, J.; Cohen, P.; Smith, A. The ground state of embryonic stem cell self-renewal. Nat. Cell Biol. 2008, 453, 519–523. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Silva, J.C.R.; Barrandon, O.; Nichols, J.; Kawaguchi, J.; Theunissen, T.; Smith, A. Promotion of Reprogramming to Ground State Pluripotency by Signal Inhibition. PLoS Biol. 2008, 6, e253. [Google Scholar] [CrossRef] [PubMed]
- Smith, A.; Heath, J.K.; Donaldson, D.D.; Wong, G.G.; Moreau, J.; Stahl, M.; Rogers, D. Inhibition of pluripotential embryonic stem cell differentiation by purified polypeptides. Nat. Cell Biol. 1988, 336, 688–690. [Google Scholar] [CrossRef] [PubMed]
- Niwa, H.; Burdon, T.; Chambers, I.; Smith, A. Self-renewal of pluripotent embryonic stem cells is mediated via activation of STAT3. Genes Dev. 1998, 12, 2048–2060. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lipsitz, Y.Y.; Woodford, C.; Yin, T.; Hanna, J.H.; Zandstra, P.W. Modulating cell state to enhance suspension expansion of human pluripotent stem cells. Proc. Natl. Acad. Sci. USA 2018, 115, 6369–6374. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Smith, A.; Guo, G.; Rostovskaya, M.; Clarke, J.; Dietmann, S.; Myers, S.; Bertone, P.; Reik, W. Epigenetic resetting of human pluripotency. Development 2017, 144, 2748–2763. [Google Scholar] [CrossRef]
- Hanna, J.; Cheng, A.; Saha, K.; Kim, J.; Lengner, C.; Soldner, F.; Cassady, J.P.; Muffat, J.; Carey, B.W.; Jaenisch, R. Human embryonic stem cells with biological and epigenetic characteristics similar to those of mouse ESCs. Proc. Natl. Acad. Sci. USA 2010, 107, 9222–9227. [Google Scholar] [CrossRef] [Green Version]
- Gafni, O.; Weinberger, L.; Mansour, A.A.; Manor, Y.S.; Chomsky, E.; Ben-Yosef, D.; Kalma, Y.; Viukov, S.; Maza, I.; Zviran, A.; et al. Derivation of novel human ground state naive pluripotent stem cells. Nature 2013, 504, 246–282. [Google Scholar] [CrossRef]
- Theunissen, T.W.; Powell, B.E.; Wang, H.; Mitalipova, M.; Faddah, D.A.; Reddy, J.; Fan, Z.P.; Maetzel, D.; Ganz, K.; Shi, L.; et al. Systematic identification of culture conditions for induction and maintenance of naive human pluripotency. Cell Stem Cell 2014, 15, 471–487. [Google Scholar] [CrossRef] [Green Version]
- Yeom, Y.I.; Fuhrmann, G.; Ovitt, C.E.; Brehm, A.; Ohbo, K.; Gross, M.; Hubner, K.; Scholer, H.R. Germline regulatory element of Oct-4 specific for the totipotent cycle of embryonal cells. Development 1996, 122, 881–894. [Google Scholar] [CrossRef]
- Tesar, P.J.; Chenoweth, J.G.; Brook, F.A.; Davies, T.J.; Evans, E.P.; Mack, D.L.; Gardner, R.L.; McKay, R.D.G. New cell lines from mouse epiblast share defining features with human embryonic stem cells. Nat. Cell Biol. 2007, 448, 196–199. [Google Scholar] [CrossRef]
- Hanna, J.; Markoulaki, S.; Mitalipova, M.; Cheng, A.W.; Cassady, J.P.; Staerk, J.; Carey, B.W.; Lengner, C.J.; Foreman, R.; Love, J.; et al. Metastable pluripotent states in NOD-mouse-derived ESCs. Cell Stem Cell 2009, 4, 513–524. [Google Scholar] [CrossRef] [Green Version]
- Graf, A.; Krebs, S.; Zakhartchenko, V.; Schwalb, B.; Blum, H.; Wolf, E. Fine mapping of genome activation in bovine embryos by RNA sequencing. Proc. Natl. Acad. Sci. USA 2014, 111, 4139–4144. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mallon, B.S.; Chenoweth, J.G.; Johnson, K.R.; Hamilton, R.S.; Tesar, P.J.; Yavatkar, A.S.; Tyson, L.J.; Park, K.; Chen, K.; Fann, Y.C.; et al. StemCellDB: The Human Pluripotent Stem Cell Database at the National Institutes of Health. Stem Cell Res. 2013, 10, 57–66. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Subramanian, A.; Tamayo, P.; Mootha, V.K.; Mukherjee, S.; Ebert, B.L.; Gillette, M.A.; Paulovich, A.; Pomeroy, S.L.; Golub, T.R.; Lander, E.S.; et al. Gene set enrichment analysis: A knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. USA 2005, 102, 15545–15550. [Google Scholar] [CrossRef] [Green Version]
- Liberzon, A.; Subramanian, A.; Pinchback, R.; Thorvaldsdóttir, H.; Tamayo, P.; Mesirov, J.P. Molecular signatures database (MSigDB) 3.0. Bioinformatics 2011, 27, 1739–1740. [Google Scholar] [CrossRef] [PubMed]
- Liberzon, A.; Birger, C.; Thorvaldsdottir, H.; Ghandi, M.; Mesirov, J.P.; Tamayo, P. The Molecular Signatures Database (MSigDB) hallmark gene set collection. Cell Syst. 2015, 1, 417–425. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Krämer, A.; Green, J.; Pollard, J., Jr.; Tugendreich, S. Causal analysis approaches in Ingenuity Pathway Analysis. Bioinformatics 2014, 30, 523–530. [Google Scholar] [CrossRef]
- Takashima, Y.; Guo, G.; Loos, R.; Nichols, J.; Ficz, G.; Krueger, F.; Oxley, D.; Santos, F.; Clarke, J.; Mansfield, W.; et al. Resetting transcription factor control circuitry toward ground-state pluripotency in human. Cell 2018, 158, 1254–1269. [Google Scholar] [CrossRef] [Green Version]
- Brons, I.G.M.; Smithers, L.E.; Trotter, M.; Rugg-Gunn, P.; Sun, B.; Lopes, S.M.C.D.S.; Howlett, S.K.; Clarkson, A.; Ahrlund-Richter, L.; Pedersen, R.A.; et al. Derivation of pluripotent epiblast stem cells from mammalian embryos. Nat. Cell Biol. 2007, 448, 191–195. [Google Scholar] [CrossRef]
- Ghimire, S.; Van Der Jeught, M.; Neupane, J.; Roost, M.S.; Anckaert, J.; Popovic, M.; Van Nieuwerburgh, F.; Mestdagh, P.; Vandesompele, J.; Deforce, D.; et al. Comparative analysis of naive, primed and ground state pluripotency in mouse embryonic stem cells originating from the same genetic background. Sci. Rep. 2018, 8, 5884. [Google Scholar] [CrossRef] [Green Version]
- Rugg-Gunn, P.; Cox, B.J.; Lanner, F.; Sharma, P.; Ignatchenko, V.; McDonald, A.C.; Garner, J.; Gramolini, A.O.; Rossant, J.; Kislinger, T. Cell-Surface Proteomics Identifies Lineage-Specific Markers of Embryo-Derived Stem Cells. Dev. Cell 2012, 22, 887–901. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Xu, J.; Guo, Z.; Su, L.; Nedambale, T.; Zhang, J.; Schenk, J.; Moreno, J.; Dinnyes, A.; Ji, W.; Tian, X.; et al. Developmental Potential of Vitrified Holstein Cattle Embryos Fertilized In Vitro with Sex-Sorted Sperm. J. Dairy Sci. 2006, 89, 2510–2518. [Google Scholar] [CrossRef]
- Xiang, J.; Wang, H.; Zhang, Y.; Wang, J.; Liu, F.; Han, X.; Lu, Z.; Li, C.; Li, Z.; Gao, Y.; et al. LCDM medium supports the derivation of bovine extended pluripotent stem cells with embryonic and extraembryonic potency in bovine–mouse chimeras from iPSCs and bovine fetal fibroblasts. FEBS J. 2021, 288, 4394–4411. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Huang, D.; Huang, C.; Yin, Y.; Vali, K.; Zhang, M.; Tang, Y. Enhanced human somatic cell reprogramming efficiency by fusion of the MYC transactivation domain and OCT4. Stem Cell Res. 2017, 25, 88–97. [Google Scholar] [CrossRef]
- Li, L.-C.; Dahiya, R. MethPrimer: Designing primers for methylation PCRs. Bioinformatics 2002, 18, 1427–1431. [Google Scholar] [CrossRef] [Green Version]
- Bock, C.; Reither, S.; Mikeska, T.; Paulsen, M.; Walter, J.; Lengauer, T. BiQ Analyzer: Visualization and quality control for DNA methylation data from bisulfite sequencing. Bioinformatics 2005, 21, 4067–4068. [Google Scholar] [CrossRef]
- Behringer, R.; Gertsenstein, M.; Nagy, K.V.; Nagy, A. Manipulating the Mouse Embryo: A Laboratory Manual, 4th ed.; Cold Spring Harbor Laboratory Press: Cold Spring Harbor, NY, USA, 2014. [Google Scholar]
- Fan, Z.; Yang, M.; Regouski, M.; Polejaeva, I.A. Gene Knockouts in Goats Using CRISPR/Cas9 System and Somatic Cell Nuclear Transfer. Methods Mol. Biol. 2018, 1874, 373–390. [Google Scholar] [CrossRef]
- Kafi, M.; McGowan, M.; Kirkland, P. In vitro maturation and fertilization of bovine oocytes and in vitro culture of presumptive zygotes in the presence of bovine pestivirus. Anim. Reprod. Sci. 2002, 71, 169–179. [Google Scholar] [CrossRef]
| Donor Cells | No. of Oocytes | No. of Fused (mean ± sd%) | No. of Reconstructed Embryos | No. of Cleaved Embryos (mean ± sd%) | No. of Blastocysts (mean ± sd%) * |
|---|---|---|---|---|---|
| DsRed-biPSC | 194 | 170 (88.4 ± 5.4) a | 123 | 89 (70.5 ± 17.0) a | 22 (27.0 ± 9.0) a |
| GFP-Dox-biPSC | 206 | 183 (88.9 ± 1.9) a | 131 | 98 (74.9 ± 10.8) a | 24 (25.2 ± 15.2) a |
| bAF | 188 | 172 (91.5 ± 0.5) a | 125 | 111 (88.6 ± 2.0) a | 45 (39.3 ± 10.7) a |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Su, Y.; Wang, L.; Fan, Z.; Liu, Y.; Zhu, J.; Kaback, D.; Oudiz, J.; Patrick, T.; Yee, S.P.; Tian, X.; et al. Establishment of Bovine-Induced Pluripotent Stem Cells. Int. J. Mol. Sci. 2021, 22, 10489. https://doi.org/10.3390/ijms221910489
Su Y, Wang L, Fan Z, Liu Y, Zhu J, Kaback D, Oudiz J, Patrick T, Yee SP, Tian X, et al. Establishment of Bovine-Induced Pluripotent Stem Cells. International Journal of Molecular Sciences. 2021; 22(19):10489. https://doi.org/10.3390/ijms221910489
Chicago/Turabian StyleSu, Yue, Ling Wang, Zhiqiang Fan, Ying Liu, Jiaqi Zhu, Deborah Kaback, Julia Oudiz, Tayler Patrick, Siu Pok Yee, Xiuchun (Cindy) Tian, and et al. 2021. "Establishment of Bovine-Induced Pluripotent Stem Cells" International Journal of Molecular Sciences 22, no. 19: 10489. https://doi.org/10.3390/ijms221910489
APA StyleSu, Y., Wang, L., Fan, Z., Liu, Y., Zhu, J., Kaback, D., Oudiz, J., Patrick, T., Yee, S. P., Tian, X., Polejaeva, I., & Tang, Y. (2021). Establishment of Bovine-Induced Pluripotent Stem Cells. International Journal of Molecular Sciences, 22(19), 10489. https://doi.org/10.3390/ijms221910489






