Dual Activity BLEG-1 from Bacillus lehensis G1 Revealed Structural Resemblance to B3 Metallo-β-Lactamase and Glyoxalase II: An Insight into Its Enzyme Promiscuity and Evolutionary Divergence
Abstract
1. Introduction
2. Results
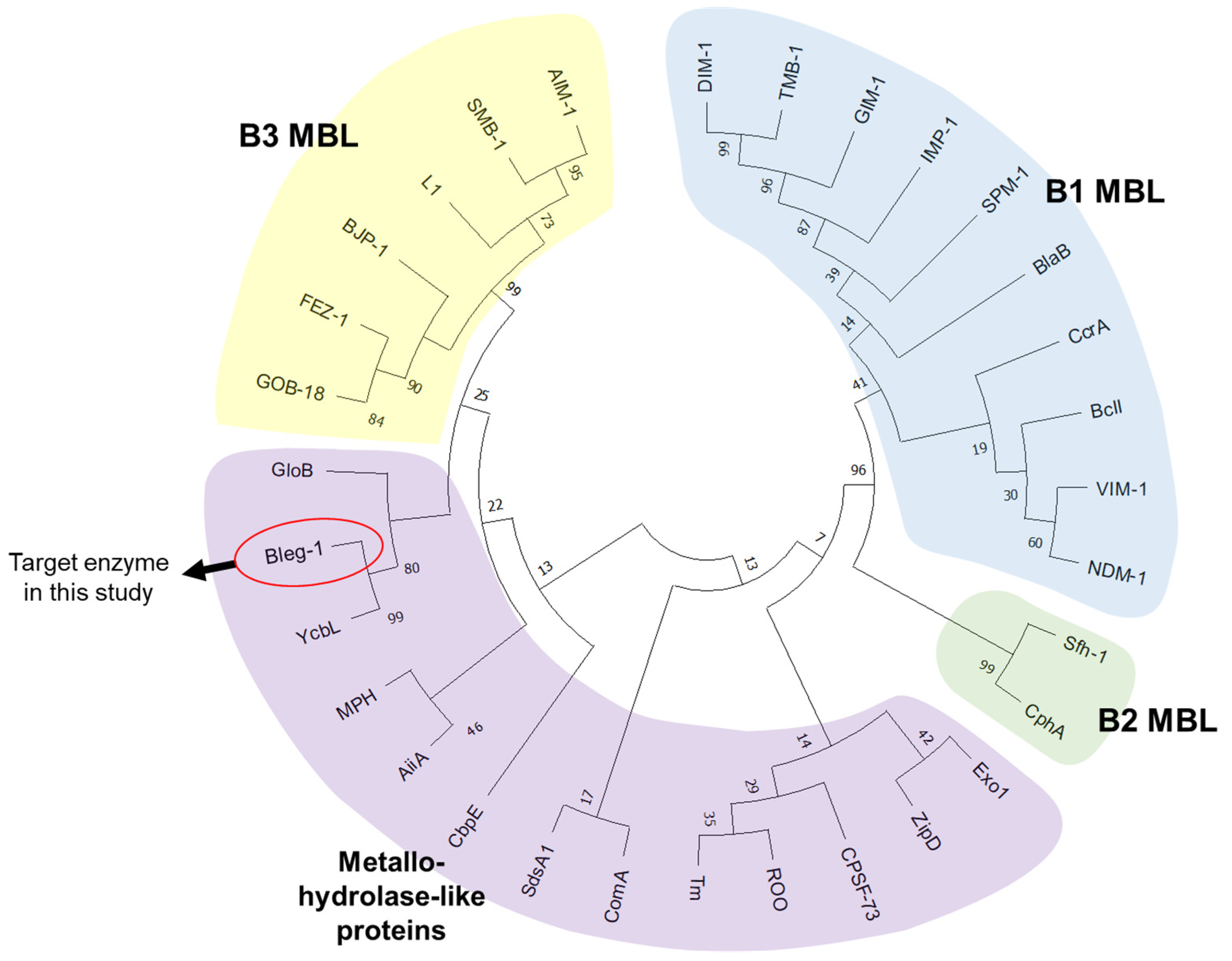
2.1. Phylogenetic Analysis and Sequence Alignment of BLEG-1
2.2. Steady-State Kinetics Analysis of BLEG-1
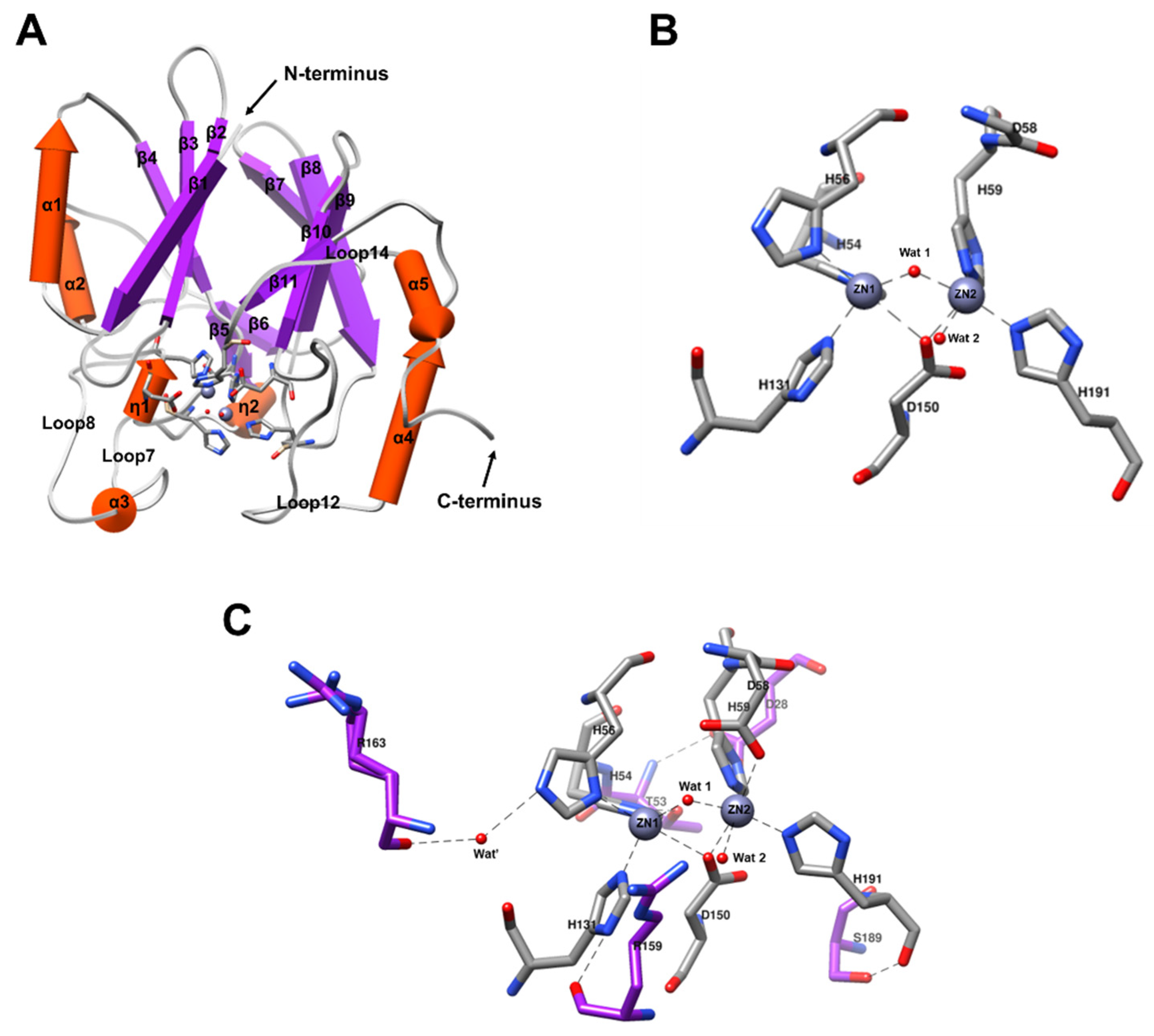
2.3. Overall X-ray Crystal Structure of BLEG-1
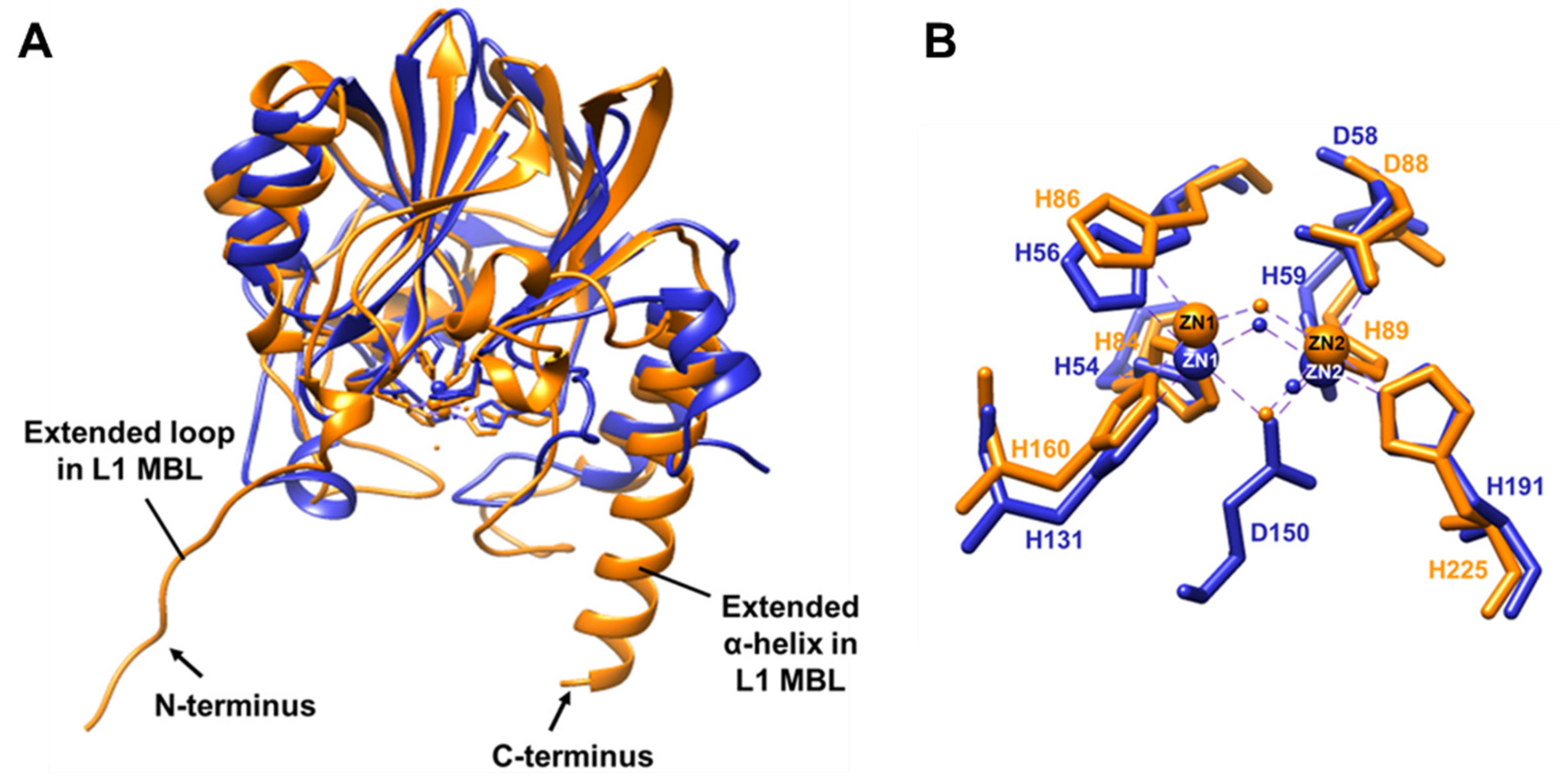
2.4. Structural Comparison of BLEG-1 with L1 B3 MBL Global Structure and Metal-Center
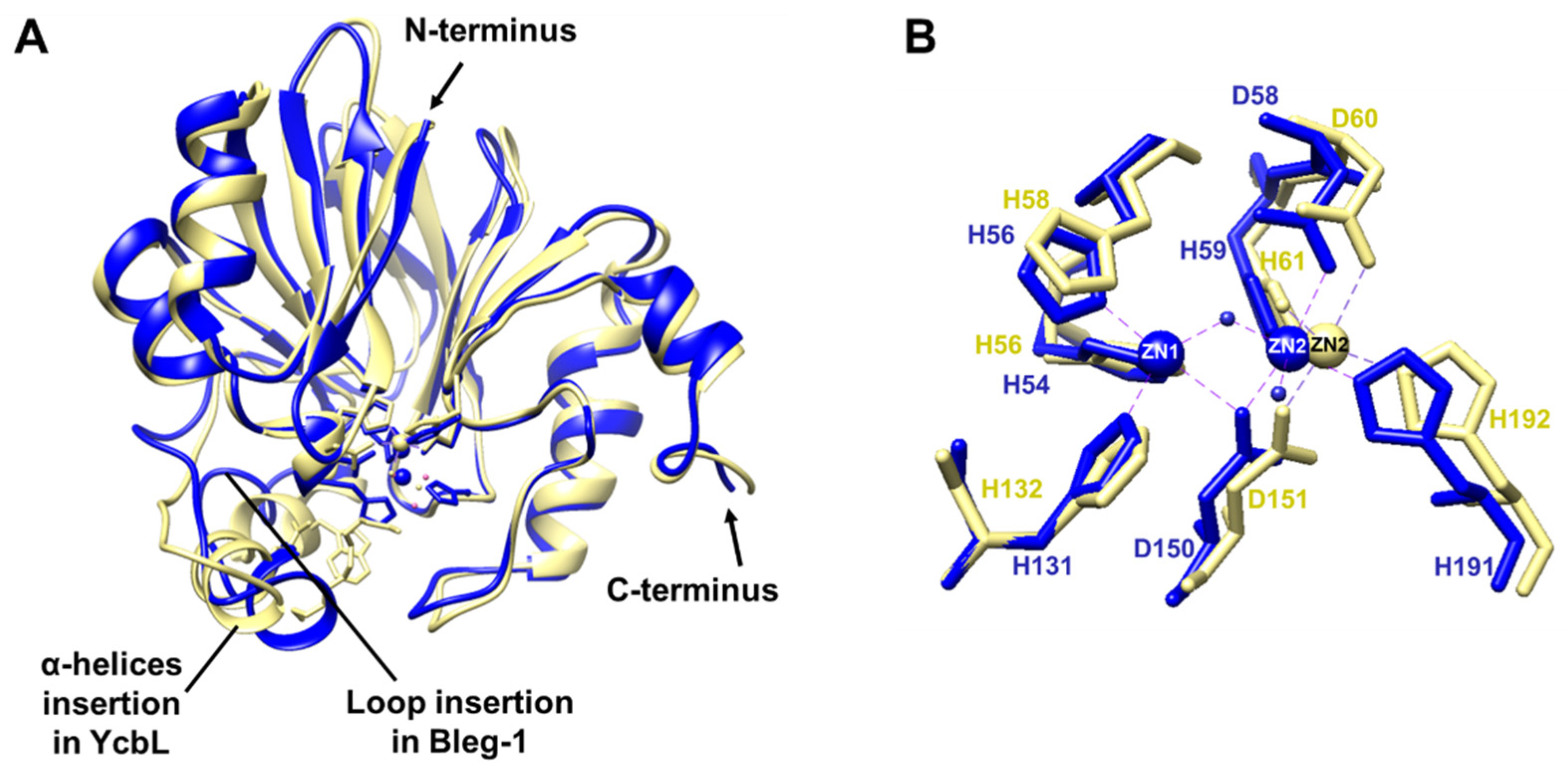
2.5. Structural Comparison of BLEG-1 with YcbL and GloB GLXIIs Global Structure and Metal-Center
2.6. Active Site Configuration of BLEG-1, L1 B3 MBL, YcbL, and GloB GLXIIs
2.7. In Silico Substrate Binding Analyses of BLEG-1, L1 MBL, YcbL GLXII, and GloB GLXII
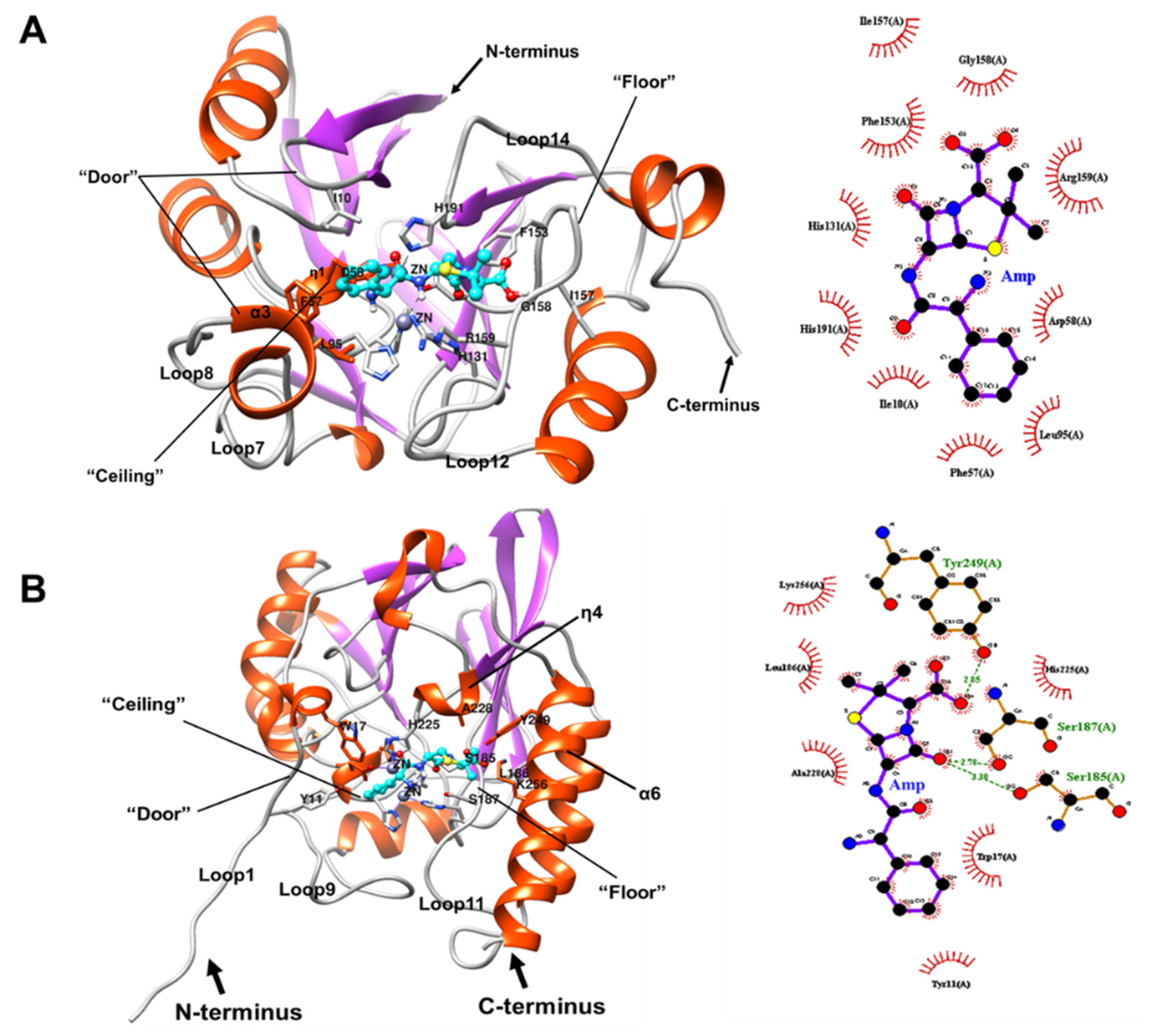
2.7.1. Docking of BLEG-1 and L1 MBL with Ampicillin
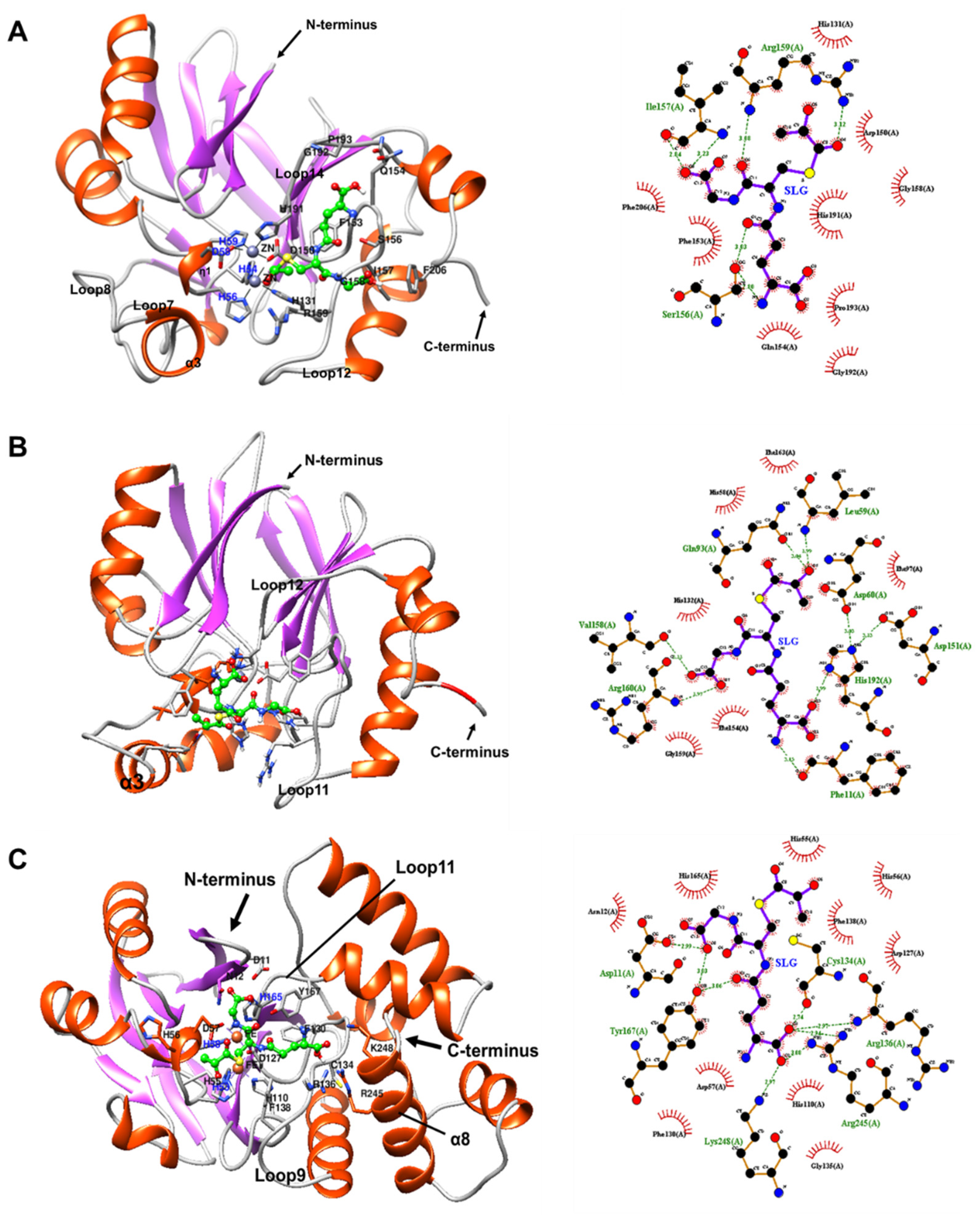
2.7.2. Docking of BLEG-1, YcbL, and GloB GLXII with S-D-Lactoylglutathione (SLG)
3. Discussion
4. Materials and Methods
4.1. Bacterial Strains and Plasmids
4.2. Phylogenetic Analysis of BLEG-1
4.3. Sequence Alignment of BLEG-1
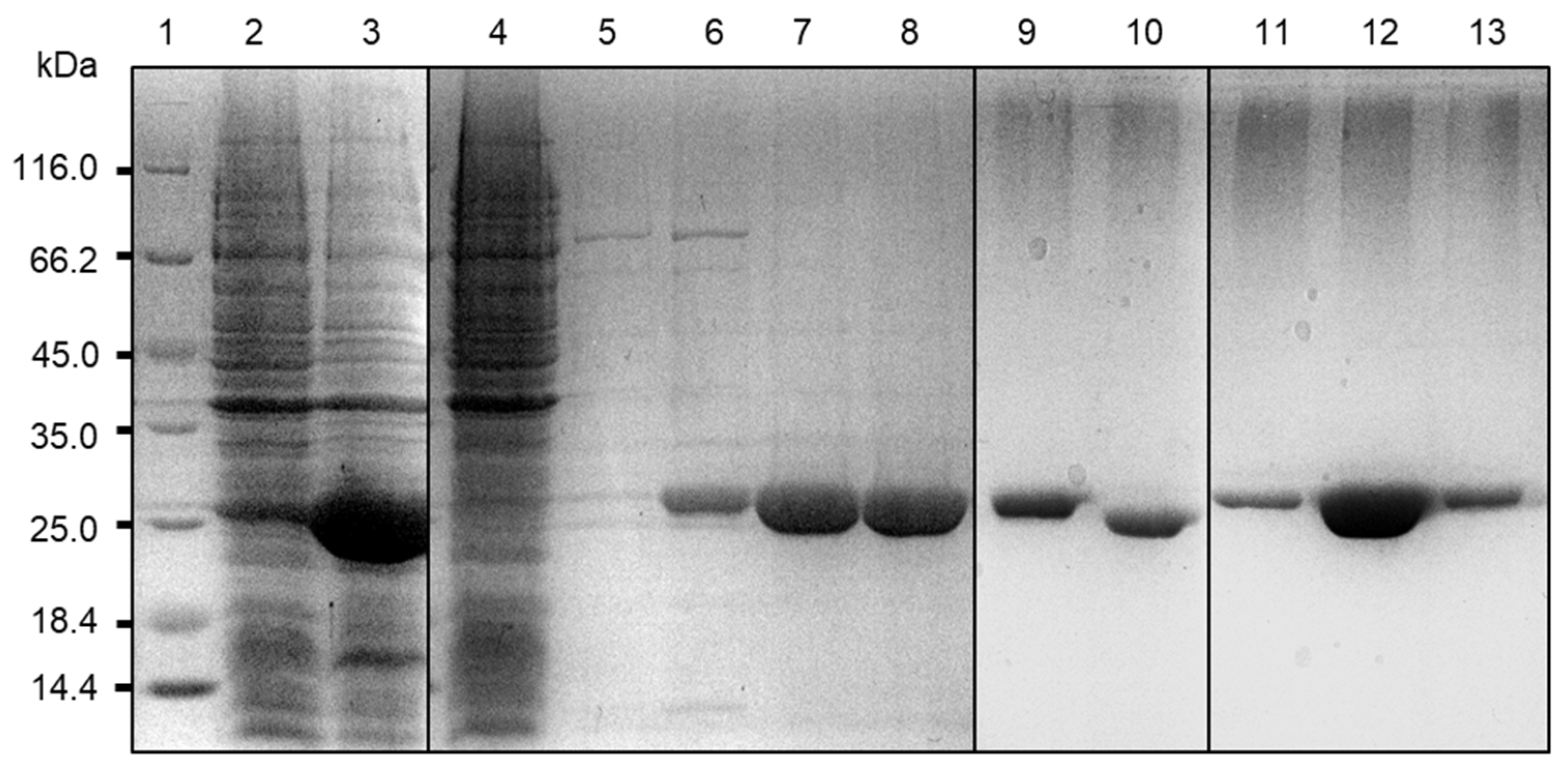
4.4. Overexpression and Purification of BLEG-1
4.5. Steady-State Kinetic Analysis of BLEG-1
4.6. Protein Crystallization, Diffraction, and Data Collection
4.7. Structure Determination, Refinement, and Validation
4.8. Docking Analyses of BLEG-1 and Selected MBL and GLXII
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Daiyasu, H.; Osaka, K.; Ishino, Y.; Toh, H. Expansion of the zinc metallo-hydrolase family of the β-lactamase fold. FEBS Lett. 2001, 503, 1–6. [Google Scholar] [CrossRef]
- Lu, S.; Wang, J.; Chitsaz, F.; Derbyshire, M.K.; Geer, R.C.; Gonzales, N.R.; Gwadz, M.; Hurwitz, D.I.; Marchler, G.H.; Song, J.S.; et al. CDD/SPARCLE: The conserved domain database in 2020. Nucleic Acids Res. 2020, 48, D265–D268. [Google Scholar] [CrossRef] [PubMed]
- González, J.M. Visualizing the superfamily of metallo-β-lactamases through sequence similarity network neighborhood connectivity analysis. Heliyon 2021, 7, e05867. [Google Scholar] [CrossRef] [PubMed]
- Bahr, G.; González, L.J.; Vila, A.J. Metallo-β-lactamases in the Age of Multidrug Resistance: From Structure and Mechanism to Evolution, Dissemination, and Inhibitor Design. Chem. Rev. 2021, 121, 7957–8094. [Google Scholar] [CrossRef]
- Bebrone, C. Metallo-β-lactamases (classification, activity, genetic organization, structure, zinc coordination) and their superfamily. Biochem. Pharmacol. 2007, 74, 1686–1701. [Google Scholar] [CrossRef]
- Meini, M.-R.; Llarrull, L.I.; Vila, A.J. Evolution of Metallo-β-lactamases: Trends Revealed by Natural Diversity and in vitro Evolution. Antibiotics 2014, 3, 285–316. [Google Scholar] [CrossRef]
- Diene, S.M.; Pinault, L.; Keshri, V.; Armstrong, N.; Khelaifia, S.; Chabrière, E.; Caetano-Anolles, G.; Colson, P.; La Scola, B.; Rolain, J.-M.; et al. Human metallo-β-lactamase enzymes degrade penicillin. Sci. Rep. 2019, 9, 1–7. [Google Scholar] [CrossRef]
- Lee, J.-H.; Takahashi, M.; Jeon, J.H.; Kang, L.-W.; Seki, M.; Park, K.S.; Hong, M.-K.; Park, Y.S.; Kim, T.Y.; Karim, A.M.; et al. Dual activity of PNGM-1 pinpoints the evolutionary origin of subclass B3 metallo-β-lactamases: A molecular and evolutionary study. Emerg. Microbes Infect. 2019, 8, 1688–1700. [Google Scholar] [CrossRef]
- Park, H.-S.; Nam, S.-H.; Lee, J.K.; Yoon, C.N.; Mannervik, B.; Benkovic, S.J.; Kim, H.-S. Design and Evolution of New Catalytic Activity with an Existing Protein Scaffold. Science 2006, 311, 535–538. [Google Scholar] [CrossRef]
- Tan, S.H.; Normi, Y.M.; Leow, A.T.C.; Salleh, A.B.; Murad, A.M.A.; Mahadi, N.M.; Rahman, M.B.A. Danger lurking in the “unknowns”: Structure-to-function studies of hypothetical protein Bleg1_2437 fromBacillus lehensisG1 alkaliphile revealed an evolutionary divergent B3 metallo-beta-lactamase. J. Biochem. 2016, 161, 58–186. [Google Scholar] [CrossRef]
- Palzkill, T. Metallo-β-lactamase structure and function. Ann. N. Y. Acad. Sci. 2013, 1277, 91–104. [Google Scholar] [CrossRef] [PubMed]
- Thornalley, P. The glyoxalase system: New developments towards functional characterization of a metabolic pathway fundamental to biological life. Biochem. J. 1990, 269, 1–11. [Google Scholar] [CrossRef]
- Jagt, D.L.V. Glyoxalase II: Molecular characteristics, kinetics and mechanism. Biochem. Soc. Trans. 1993, 21, 522–527. [Google Scholar] [CrossRef]
- Suttisansanee, U.; Honek, J. Bacterial glyoxalase enzymes. Semin. Cell Dev. Biol. 2011, 22, 285–292. [Google Scholar] [CrossRef]
- Tawfik, O.K.A.D.S. Enzyme Promiscuity: A Mechanistic and Evolutionary Perspective. Annu. Rev. Biochem. 2010, 79, 471–505. [Google Scholar] [CrossRef] [PubMed]
- Stamp, A.L.; Owen, P.; El Omari, K.; Nichols, C.E.; Lockyer, M.; Lamb, H.K.; Charles, I.G.; Hawkins, A.R.; Stammers, D.K. Structural and functional characterization of Salmonella enterica serovar Typhimurium YcbL: An unusual Type II glyoxalase. Protein Sci. 2010, 19, 1897–1905. [Google Scholar] [CrossRef]
- Ullah, J.; Walsh, T.; Taylor, I.; Emery, D.; Verma, C.; Gamblin, S.; Spencer, J. The crystal structure of the L1 metallo-β-lactamase from Stenotrophomonas maltophilia at 1.7 å resolution. J. Mol. Biol. 1998, 284, 125–136. [Google Scholar] [CrossRef] [PubMed]
- Simm, A.M.; Higgins, C.S.; Carenbauer, A.L.; Crowder, M.W.; Bateson, J.H.; Bennett, P.M.; Clarke, A.R.; Halford, S.E.; Walsh, T. Characterization of Monomeric L1 Metallo-β-lactamase and the Role of the N-terminal Extension in Negative Cooperativity and Antibiotic Hydrolysis. J. Biol. Chem. 2002, 277, 24744–24752. [Google Scholar] [CrossRef]
- Crowder, M.W.; Spencer, J.; Vila, A.J. Metallo-β-lactamases: Novel Weaponry for Antibiotic Resistance in Bacteria. Accounts Chem. Res. 2006, 39, 721–728. [Google Scholar] [CrossRef]
- Leiros, H.-K.S.; Borra, P.S.; Brandsdal, B.O.; Edvardsen, K.S.W.; Spencer, J.; Walsh, T.; Samuelsen, Ø. Crystal Structure of the Mobile Metallo-β-Lactamase AIM-1 from Pseudomonas aeruginosa: Insights into Antibiotic Binding and the Role of Gln157. Antimicrob. Agents Chemother. 2012, 56, 4341–4353. [Google Scholar] [CrossRef]
- Wachino, J.-I.; Yamaguchi, Y.; Mori, S.; Kurosaki, H.; Arakawa, Y.; Shibayama, K. Structural Insights into the Subclass B3 Metallo-β-Lactamase SMB-1 and the Mode of Inhibition by the Common Metallo-β-Lactamase Inhibitor Mercaptoacetate. Antimicrob. Agents Chemother. 2012, 57, 101–109. [Google Scholar] [CrossRef] [PubMed]
- Morán-Barrio, J.; Lisa, M.-N.; Larrieux, N.; Drusin, S.; Viale, A.M.; Moreno, D.; Buschiazzo, A.; Vila, A.J. Crystal Structure of the Metallo-β-Lactamase GOB in the Periplasmic Dizinc Form Reveals an Unusual Metal Site. Antimicrob. Agents Chemother. 2016, 60, 6013–6022. [Google Scholar] [CrossRef] [PubMed]
- García-Sáez, I.; Mercuri, P.; Papamicael, C.; Kahn, R.; Frère, J.; Galleni, M.; Rossolini, G.M.; Dideberg, O. Three-dimensional Structure of FEZ-1, a Monomeric Subclass B3 Metallo-β-lactamase from Fluoribacter gormanii, in Native Form and in Complex with d-Captopril. J. Mol. Biol. 2003, 325, 651–660. [Google Scholar] [CrossRef]
- Cameron, A.D.; Ridderström, M.; Olin, B.; Mannervik, B. Crystal structure of human glyoxalase II and its complex with a glutathione thiolester substrate analogue. Structure 1999, 7, 1067–1078. [Google Scholar] [CrossRef]
- Marasinghe, G.P.K.; Sander, I.M.; Bennett, B.; Periyannan, G.; Yang, K.-W.; Makaroff, C.; Crowder, M.W. Structural Studies on a Mitochondrial Glyoxalase II. J. Biol. Chem. 2005, 280, 40668–40675. [Google Scholar] [CrossRef] [PubMed]
- Campos-Bermudez, V.A.; Leite, N.R.; Krog, R.; Costa-Filho, A.; Soncini, F.C.; Oliva, G.; Vila, A. Biochemical and Structural Characterization of Salmonella typhimurium Glyoxalase II: New Insights into Metal Ion Selectivity. Biochemistry 2007, 46, 11069–11079. [Google Scholar] [CrossRef] [PubMed]
- Levin, E.J.; Kondrashov, D.A.; Wesenberg, G.E.; Phillips, G.N. Ensemble Refinement of Protein Crystal Structures: Validation and Application. Structure 2007, 15, 1040–1052. [Google Scholar] [CrossRef] [PubMed]
- Selvaraju, G.; Leow, T.C.; Salleh, A.B.; Normi, Y.M. Design and Characterisation of Inhibitory Peptides against Bleg1_2478, an Evolutionary Divergent B3 Metallo-β-lactamase. Molecules 2020, 25, 5797. [Google Scholar] [CrossRef]
- Kim, Y.; Maltseva, N.; Wilamowski, M.; Tesar, C.; Endres, M.; Joachimiak, A. Structural and biochemical analysis of the metallo-β-lactamase L1 from emerging pathogen Stenotrophomonas maltophilia revealed the subtle but distinct di-metal scaffold for catalytic activity. Protein Sci. 2020, 29, 723–743. [Google Scholar] [CrossRef]
- Garrity, J.D.; Pauff, J.M.; Crowder, M.W.; Alonso, A.; Narisawa, S.; Bogetz, J.; Tautz, L.; Hadzic, R.; Huynh, H.; Williams, S.; et al. Probing the Dynamics of a Mobile Loop above the Active Site of L1, a Metallo-β-lactamase from Stenotrophomonas maltophilia, via Site-directed Mutagenesis and Stopped-flow Fluorescence Spectroscopy. J. Biol. Chem. 2004, 279, 39663–39670. [Google Scholar] [CrossRef]
- Guo, Y.; Wang, J.; Niu, G.; Shui, W.; Sun, Y.; Zhou, H.; Zhang, Y.; Yang, C.; Lou, Z.; Rao, Z. A structural view of the antibiotic degradation enzyme NDM-1 from a superbug. Protein Cell 2011, 2, 384–394. [Google Scholar] [CrossRef]
- Wachino, J.-I.; Yamaguchi, Y.; Mori, S.; Jin, W.; Kimura, K.; Kurosaki, H.; Arakawa, Y. Structural Insights into Recognition of Hydrolyzed Carbapenems and Inhibitors by Subclass B3 Metallo-β-Lactamase SMB-1. Antimicrob. Agents Chemother. 2016, 60, 4274–4282. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Giles, N.M.; Watts, A.B.; Giles, G.; Fry, F.H.; Littlechild, J.A.; Jacob, C. Metal and Redox Modulation of Cysteine Protein Function. Chem. Biol. 2003, 10, 677–693. [Google Scholar] [CrossRef]
- Ninković, D.B.; Malenov, D.P.; Petrović, P.V.; Brothers, E.N.; Niu, S.; Hall, M.B.; Belić, M.R.; Zarić, S.D. Unexpected Importance of Aromatic-Aliphatic and Aliphatic Side Chain-Backbone Interactions in the Stability of Amyloids. Chem. A Eur. J. 2017, 23, 11046–11053. [Google Scholar] [CrossRef] [PubMed]
- Reiger, M.; Lassak, J.; Jung, K. Deciphering the role of the type II glyoxalase isoenzyme YcbL (GlxII-2) in Escherichia coli. FEMS Microbiol. Lett. 2014, 362, 1–7. [Google Scholar] [CrossRef]
- Meini, M.-R.; Llarrull, L.I.; Vila, A.J. Overcoming differences: The catalytic mechanism of metallo-β-lactamases. FEBS Lett. 2015, 589, 3419–3432. [Google Scholar] [CrossRef]
- Zang, T.M.; Hollman, D.A.; Crawford, P.A.; Crowder, M.W.; Makaroff, C. Arabidopsis Glyoxalase II Contains a Zinc/Iron Binuclear Metal Center That Is Essential for Substrate Binding and Catalysis. J. Biol. Chem. 2001, 276, 4788–4795. [Google Scholar] [CrossRef]
- Tomatis, P.E.; Fabiane, S.M.; Simona, F.; Carloni, P.; Sutton, B.J.; Vila, A. Adaptive protein evolution grants organismal fitness by improving catalysis and flexibility. Proc. Natl. Acad. Sci. USA 2008, 105, 20605–20610. [Google Scholar] [CrossRef]
- Garau, G.; Di Guilmi, A.M.; Hall, B.G. Structure-Based Phylogeny of the Metallo-β-Lactamases. Antimicrob. Agents Chemother. 2005, 49, 2778–2784. [Google Scholar] [CrossRef]
- Lee, S.Y.; Brem, J.; Pettinati, I.; Claridge, T.D.W.; Gileadi, O.; Schofield, C.J.; McHugh, P.J. Cephalosporins inhibit human metallo β-lactamase fold DNA repair nucleases SNM1A and SNM1B/apollo. Chem. Commun. 2016, 52, 6727–6730. [Google Scholar] [CrossRef]
- Tan, S.H.; Normi, Y.M. 14th October 2016 Progress Report; Universiti Putra Malaysia: Serdang, Malaysia, 2016; unpublished work. [Google Scholar]
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X: Molecular Evolutionary Genetics Analysis across Computing Platforms. Mol. Biol. Evol. 2018, 35, 1547–1549. [Google Scholar] [CrossRef]
- Larkin, M.A.; Blackshields, G.; Brown, N.P.; Chenna, R.; McGettigan, P.A.; McWilliam, H.; Valentin, F.; Wallace, I.; Wilm, A.; Lopez, R.; et al. Clustal W and Clustal X version 2.0. Bioinformation 2007, 23, 2947–2948. [Google Scholar] [CrossRef]
- Robert, X.; Gouet, P. Deciphering key features in protein structures with the new ENDscript server. Nucleic Acids Res. 2014, 42, W320–W324. [Google Scholar] [CrossRef] [PubMed]
- Bradford, M.M. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-Dye binding. Anal. Biochem. 1976, 72, 248–254. [Google Scholar] [CrossRef]
- Racker, E. The mechanism of action of glyoxalase. J. Biol. Chem. 1951, 190, 685–696. [Google Scholar] [CrossRef]
- Lineweaver, H.; Burk, D. The Determination of Enzyme Dissociation Constants. J. Am. Chem. Soc. 1934, 56, 658–666. [Google Scholar] [CrossRef]
- Minor, W.; Cymborowski, M.; Otwinowski, Z.; Chruszcz, M. HKL-3000: The integration of data reduction and structure solution–from diffraction images to an initial model in minutes. Acta Crystallogr. Sect. D Biol. Crystallogr. 2006, 62, 859–866. [Google Scholar] [CrossRef]
- McCoy, A.J.; Grosse-Kunstleve, R.W.; Adams, P.; Winn, M.D.; Storoni, L.C.; Read, R. Phasercrystallographic software. J. Appl. Crystallogr. 2007, 40, 658–674. [Google Scholar] [CrossRef]
- Echols, N.; Grosse-Kunstleve, R.W.; Afonine, P.V.; Bunkóczi, G.; Chen, V.; Headd, J.J.; McCoy, A.J.; Moriarty, N.W.; Read, R.; Richardson, D.C.; et al. Graphical tools for macromolecular crystallography in PHENIX. J. Appl. Crystallogr. 2012, 45, 581–586. [Google Scholar] [CrossRef]
- Emsley, P.; Lohkamp, B.; Scott, W.; Cowtan, K.D. Features and development of Coot. Acta Crystallogr. Sect. D Biol. Crystallogr. 2010, 66, 486–501. [Google Scholar] [CrossRef]
- Afonine, P.V.; Grosse-Kunstleve, R.W.; Echols, N.; Headd, J.J.; Moriarty, N.W.; Mustyakimov, M.; Terwilliger, T.C.; Urzhumtsev, A.; Zwart, P.H.; Adams, P.D. Towards automated crystallographic structure refinement with phenix.refine. Acta Crystallogr. Sect. D Biol. Crystallogr. 2012, 68, 352–367. [Google Scholar] [CrossRef] [PubMed]
- Williams, C.J.; Headd, J.J.; Moriarty, N.W.; Prisant, M.G.; Videau, L.L.; Deis, L.N.; Verma, V.; Keedy, D.A.; Hintze, B.; Chen, V.B.; et al. MolProbity: More and better reference data for improved all-atom structure validation. Protein Sci. 2018, 27, 293–315. [Google Scholar] [CrossRef] [PubMed]
- Pettersen, E.F.; Goddard, T.D.; Huang, C.C.; Couch, G.S.; Greenblatt, D.M.; Meng, E.C.; Ferrin, T.E. UCSF Chimera--A visualization system for exploratory research and analysis. J. Comput. Chem. 2004, 25, 1605–1612. [Google Scholar] [CrossRef] [PubMed]
- Martínez, L.; Andrade, R.; Birgin, E.G.; Martínez, J.M. Software News and Update Packmol: A Package for Building Initial Configurations for Molecular Dynamics Simulations. J. Comput. Chem. 2009, 30, 2157–2164. [Google Scholar] [CrossRef] [PubMed]
- Laskowski, R.A.; Swindells, M.B. LigPlot+: Multiple ligand-protein interaction diagrams for drug discovery. J. Chem. Inf. Model. 2011, 51, 2778–2786. [Google Scholar] [CrossRef]
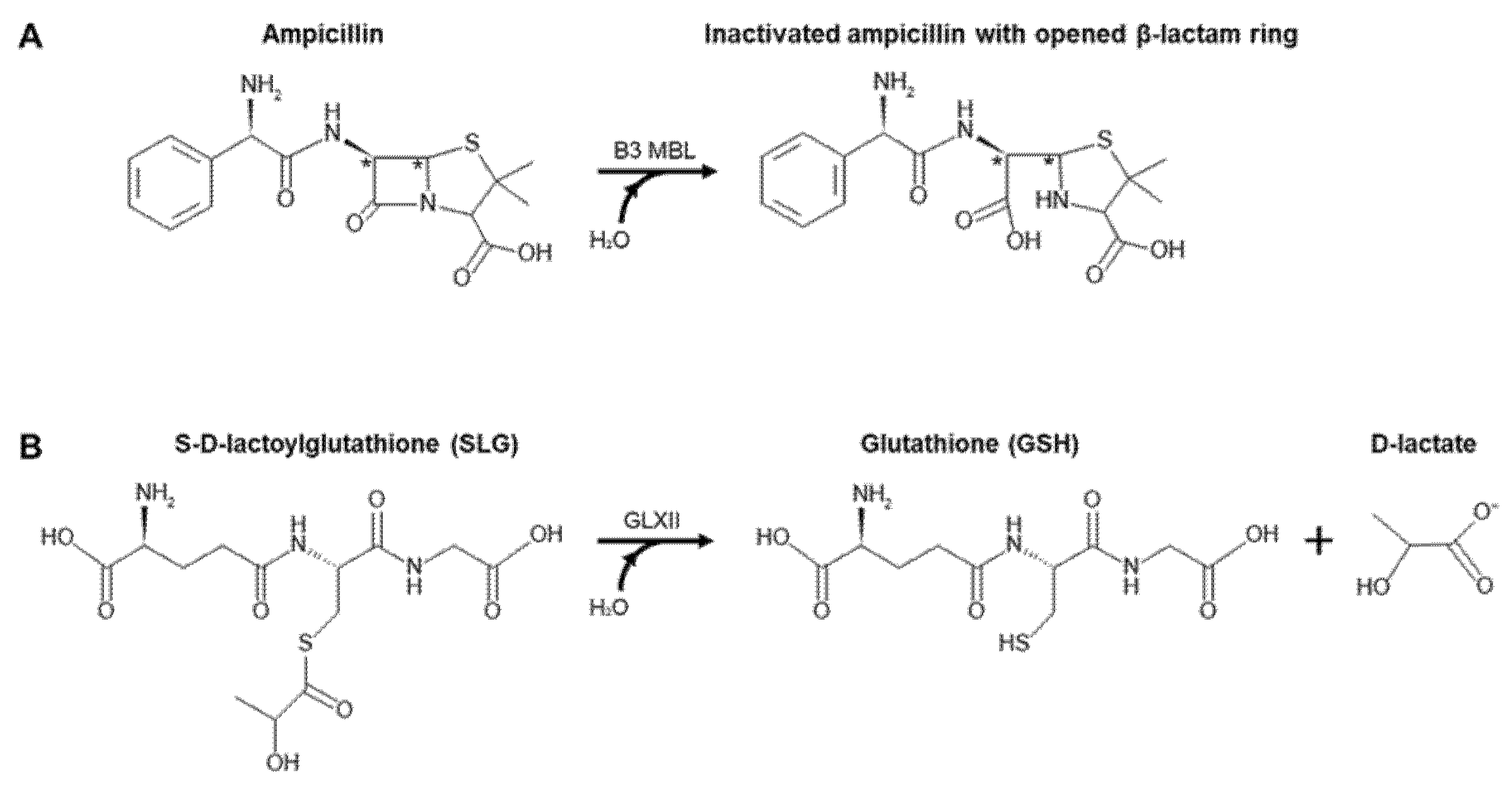



| With Ampicillin | With SLG | |
|---|---|---|
| Vmax (mM s−1) | (4.20 ± 0.16) × 10−5 | (3.55 ± 0.26) × 10−5 |
| KM (mM) | (7.27 ± 0.51) × 10−2 | (5.50 ± 0.12) × 10−2 |
| kcat (s−1) | (4.20 ± 0.16) × 10−2 | (3.55 ± 0.26) × 10−2 |
| kcat/KM (mM−1 s−1) | (5.78 ± 0.19) × 10−1 | (6.47 ± 0.77) × 10−1 |
| Data Collection Statistics | |
|---|---|
| Data collection site | Malaysia Genome Institute |
| Detector | CCD MSC PAD200sk |
| Wavelength (Å) | 1.5418 |
| Data resolution (Å) | 50.00–1.44 (1.46–1.44) |
| Space group | P41212 |
| Unit cell dimensions (a, b, c) (Å) | 55.36, 55.36, 143.09 |
| Total number of reflections | 205685 |
| Number of unique reflections | 40528 |
| Multiplicity | 5.1 (3.1) |
| Completeness (%) | 98.0 (97.9) |
| Rmerge | 0.045 (0.254) |
| <I/σ(I)> | 28.5 (4.0) |
| Refinement Statistics | |
| Resolution range (Å) | 30.26–1.44 (1.49–1.44) |
| No: residues/water/ion/ligand | 209/245/11/2 |
| R-factor (Rwork/Rfree) (%) | 18.44/20.85 |
| Rms bond length deviation (Å) | 0.005 |
| Rms bond angle deviation (°) | 0.804 |
| Ramachandran angles (favored/allowed/disallowed) (%) | 98.55/1.45/0 |
| Metal ion | Distance (Å) | First Shell Ligand | Second Shell Ligand | |
|---|---|---|---|---|
| Involved in Metal Coordination | Involved in Second Shell Effect | |||
| Zn1 | 2.07 | His-54 NE2 | His-54 ND1 | Thr-53 OG1 |
| Zn1 | 2.12 | His-56 ND1 | His-56 NE2 | Wat’ O, Arg-163 O |
| Zn1 | 2.06 | His-131 NE2 | His-131 ND1 | Arg-159 O |
| Zn1 | 2.62 | Asp-150 OD2 | ||
| Zn1 | 1.94 | Wat1 O | ||
| Zn2 | 2.34 | Asp-58 OD2 | ||
| Zn2 | 2.06 | His-59 NE2 | His-59 ND1 | Asp-28 OD2 |
| Zn2 | 2.05 | Asp-150 OD2 | ||
| Zn2 | 2.03 | His-191 NE2 | His-191 O | Ser-189 OG |
| Zn2 | 2.00 | Wat1 O | ||
| Zn2 | 3.52 | Wat2 O | ||
| BLEG-1 | YcbL | GloB |
|---|---|---|
His-131 (Loop10)
| His-132 (Loop8)
| His-110 (Loop7)
|
Phe-153 (Loop12)
| Phe-154 (β10)
| Phe-130 (β11)
|
Ile-157 (Loop12)
| Val-158 (β11)
| Cys-134 (β12)
|
Gly-158 (Loop12)
| Gly-159 (Loop11)
| Gly-135 (Loop9)
|
Arg-159 (Loop12)
| Arg-160 (Loop11)
| Arg-136 (Loop9)
|
His-191 (Loop14)
| His-192 (Loop12)
| His-165 (Loop11)
|
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Au, S.X.; Dzulkifly, N.S.; Muhd Noor, N.D.; Matsumura, H.; Raja Abdul Rahman, R.N.Z.; Normi, Y.M. Dual Activity BLEG-1 from Bacillus lehensis G1 Revealed Structural Resemblance to B3 Metallo-β-Lactamase and Glyoxalase II: An Insight into Its Enzyme Promiscuity and Evolutionary Divergence. Int. J. Mol. Sci. 2021, 22, 9377. https://doi.org/10.3390/ijms22179377
Au SX, Dzulkifly NS, Muhd Noor ND, Matsumura H, Raja Abdul Rahman RNZ, Normi YM. Dual Activity BLEG-1 from Bacillus lehensis G1 Revealed Structural Resemblance to B3 Metallo-β-Lactamase and Glyoxalase II: An Insight into Its Enzyme Promiscuity and Evolutionary Divergence. International Journal of Molecular Sciences. 2021; 22(17):9377. https://doi.org/10.3390/ijms22179377
Chicago/Turabian StyleAu, Shaw Xian, Nur Syazana Dzulkifly, Noor Dina Muhd Noor, Hiroyoshi Matsumura, Raja Noor Zaliha Raja Abdul Rahman, and Yahaya M. Normi. 2021. "Dual Activity BLEG-1 from Bacillus lehensis G1 Revealed Structural Resemblance to B3 Metallo-β-Lactamase and Glyoxalase II: An Insight into Its Enzyme Promiscuity and Evolutionary Divergence" International Journal of Molecular Sciences 22, no. 17: 9377. https://doi.org/10.3390/ijms22179377
APA StyleAu, S. X., Dzulkifly, N. S., Muhd Noor, N. D., Matsumura, H., Raja Abdul Rahman, R. N. Z., & Normi, Y. M. (2021). Dual Activity BLEG-1 from Bacillus lehensis G1 Revealed Structural Resemblance to B3 Metallo-β-Lactamase and Glyoxalase II: An Insight into Its Enzyme Promiscuity and Evolutionary Divergence. International Journal of Molecular Sciences, 22(17), 9377. https://doi.org/10.3390/ijms22179377







