Small Structural Proteins E and M Render the SARS-CoV-2 Pseudovirus More Infectious and Reveal the Phenotype of Natural Viral Variants
Abstract
:1. Introduction
2. Results
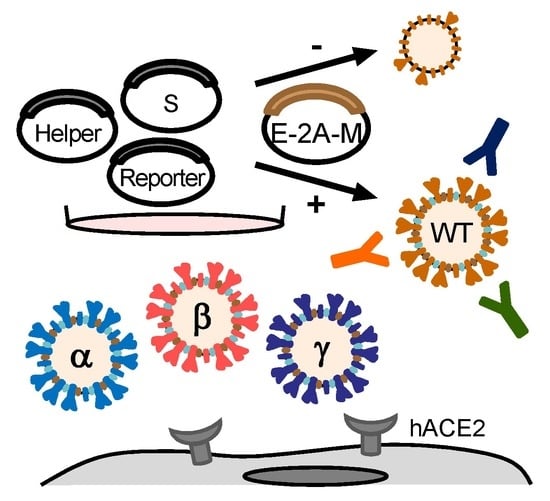
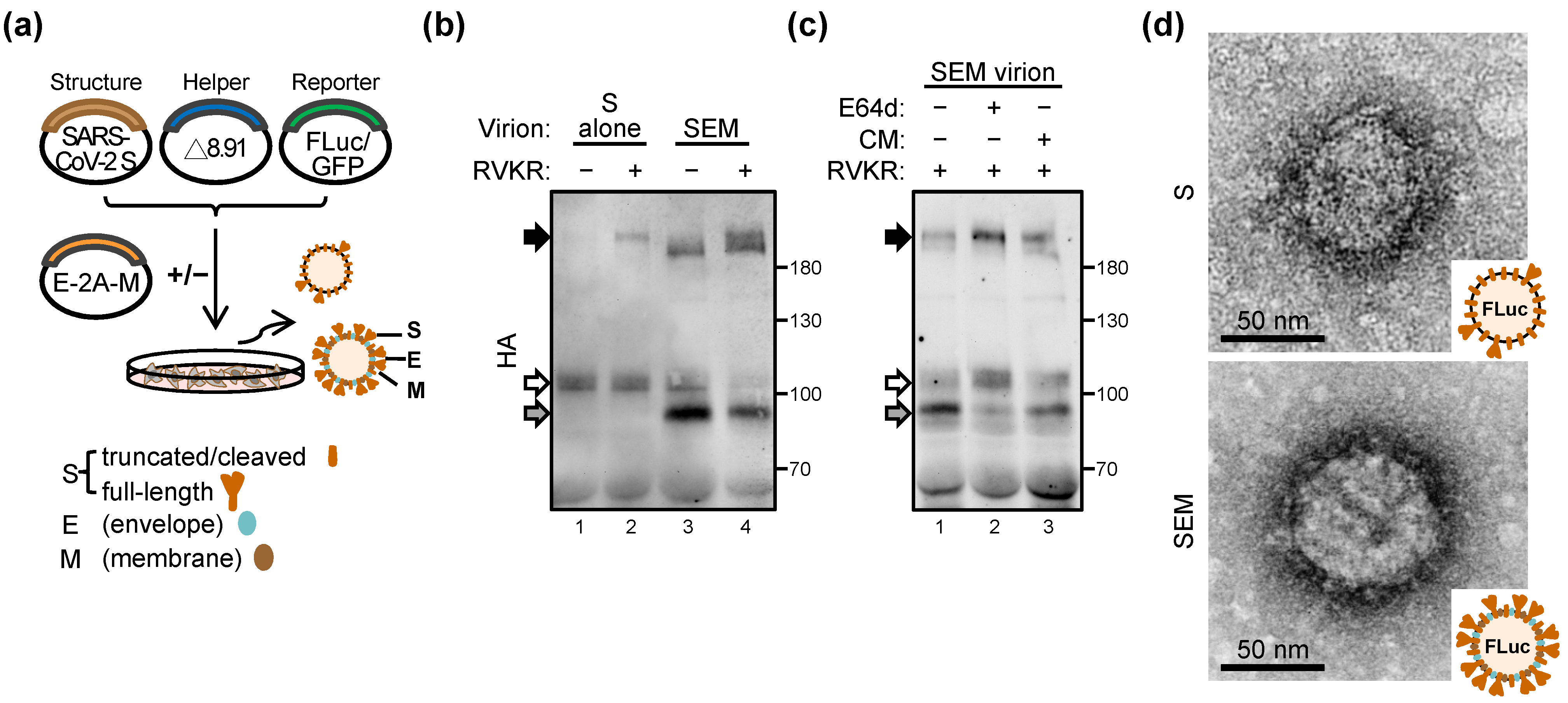
2.1. SARS-CoV-2 Structural Proteins E and M Enhance S Priming
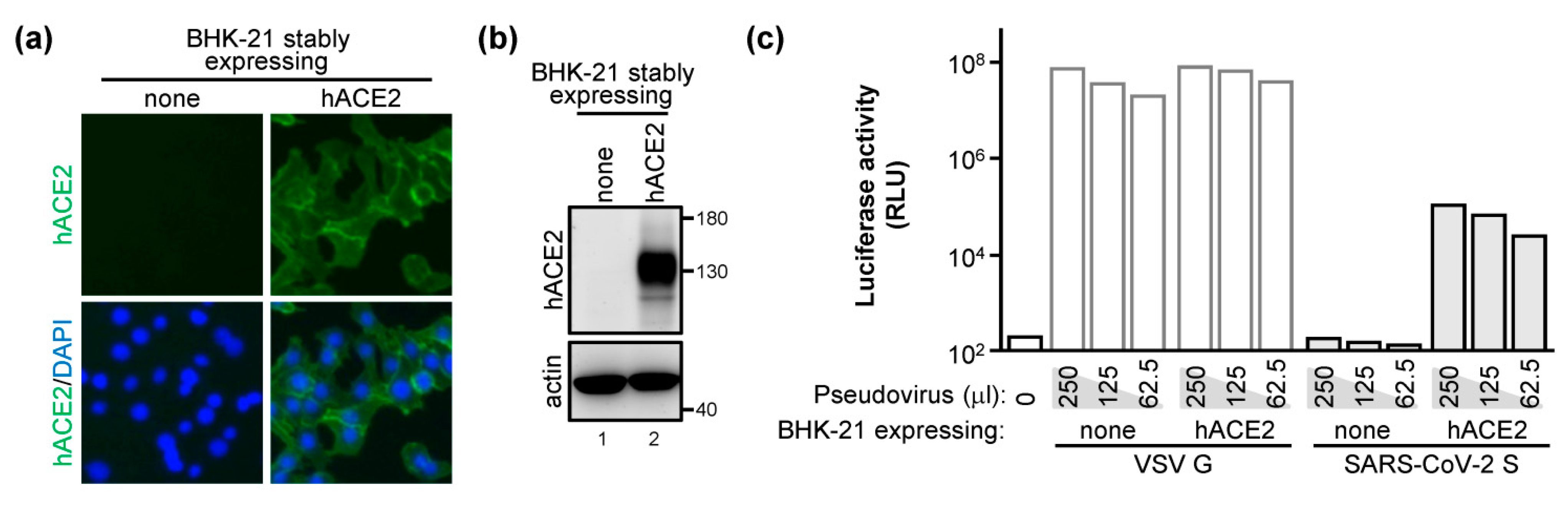
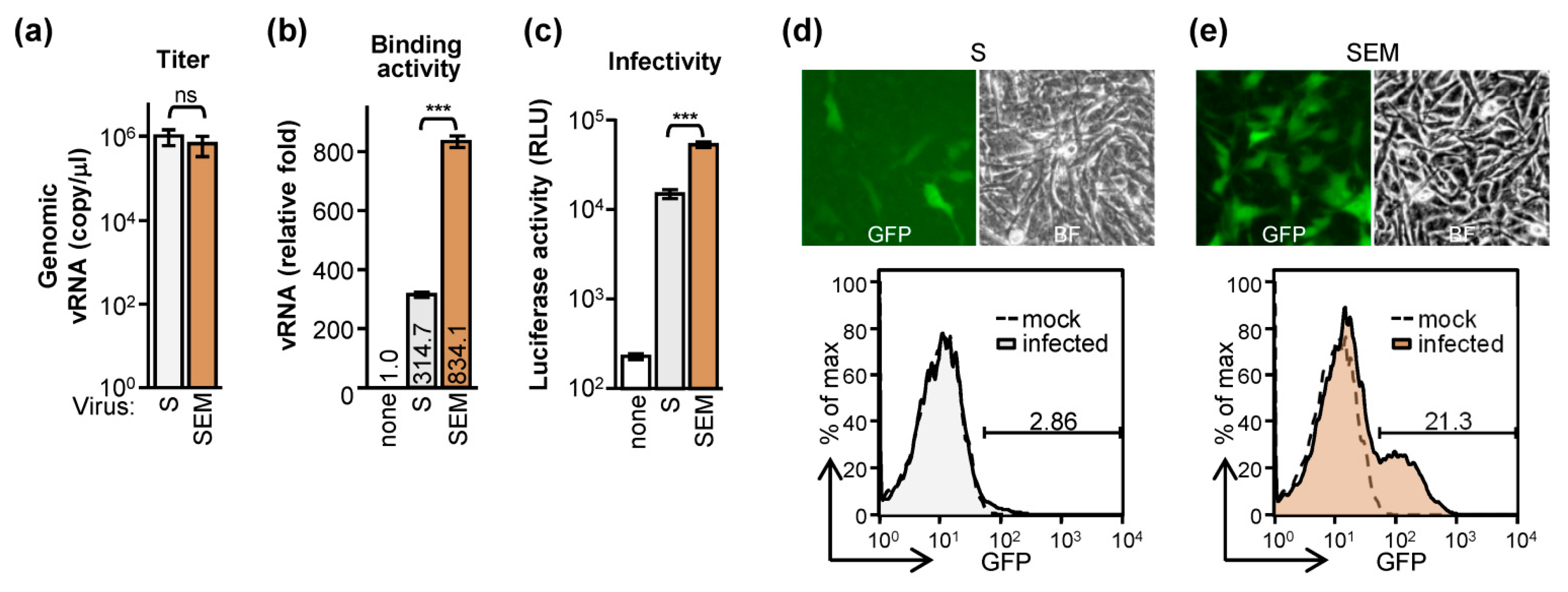
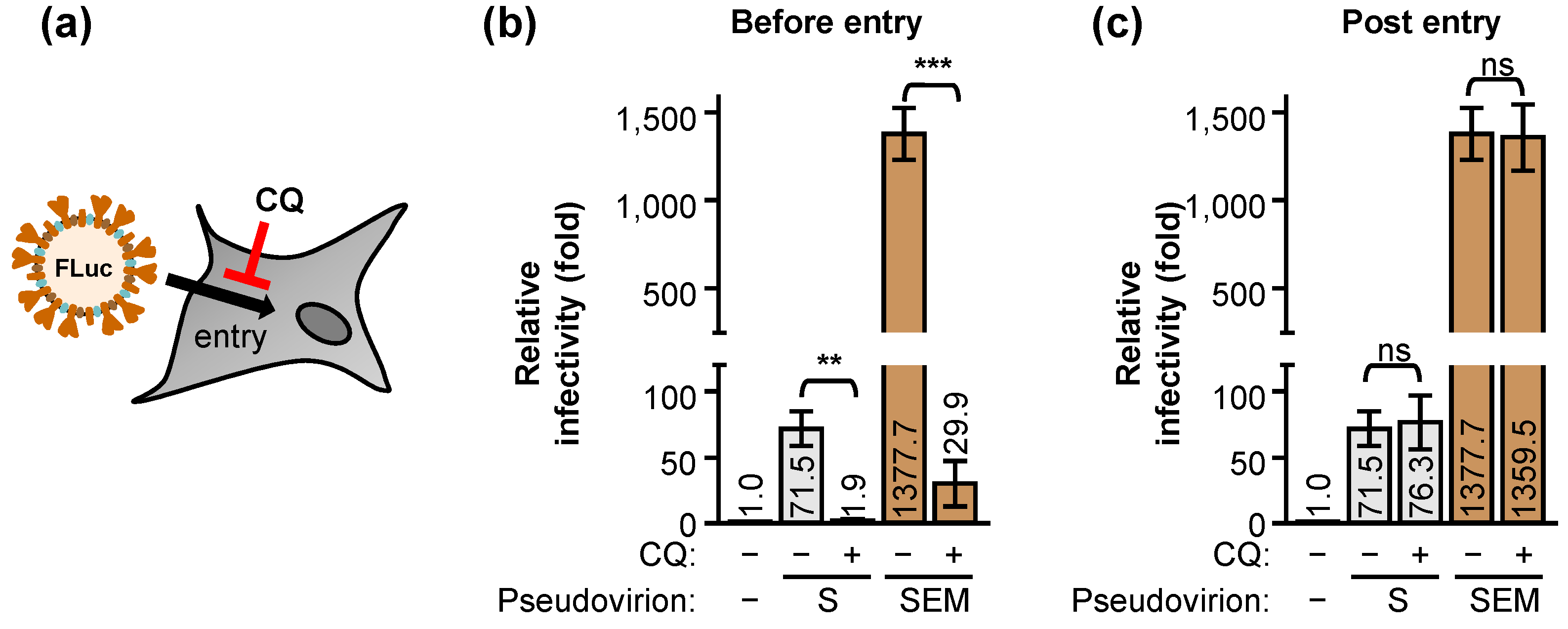
2.2. The E and M Proteins Make SARS-CoV-2 Pseudovirus Highly Infectious
2.3. The SEM Pseudovirus Is Highly Sensitive in the Bioassay Evaluating SARS-CoV-2 Neutralizing Antibodies
2.4. The SEM Pseudoviruses of Some Natural S Variants Showed the Capability to Resist Neutralizing Antibodies
3. Discussion
4. Materials and Methods
4.1. Inhibitors, Primary Antibodies, and Convalescent Plasma
4.2. Plasmid Constructs
4.3. Cell Lines
4.4. Immunofluorescence Assay
4.5. Western Blot Analysis
4.6. Pseudovirus Propagation
4.7. Transmission Electron Microscope (TEM)
4.8. Flow Cytometry Assay
4.9. Binding Assay
4.10. Neutralization Assay
4.11. Reporter Assay
4.12. Quantification and Statistical Analysis
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Zhu, N.; Zhang, D.; Wang, W.; Li, X.; Yang, B.; Song, J.; Zhao, X.; Huang, B.; Shi, W.; Lu, R.; et al. A Novel Coronavirus from Patients with Pneumonia in China, 2019. N. Engl. J. Med. 2020, 382, 727–733. [Google Scholar] [CrossRef]
- Liu, Z.; Zheng, H.; Lin, H.; Li, M.; Yuan, R.; Peng, J.; Xiong, Q.; Sun, J.; Li, B.; Wu, J.; et al. Identification of Common Deletions in the Spike Protein of Severe Acute Respiratory Syndrome Coronavirus 2. J. Virol. 2020, 94, e00790-20. [Google Scholar] [CrossRef]
- Cui, J.; Li, F.; Shi, Z.L. Origin and evolution of pathogenic coronaviruses. Nat. Rev. Microbiol. 2019, 17, 181–192. [Google Scholar] [CrossRef] [Green Version]
- De Haan, C.A.; Rottier, P.J. Molecular interactions in the assembly of coronaviruses. Adv. Virus Res. 2005, 64, 165–230. [Google Scholar]
- De Haan, C.A.; Vennema, H.; Rottier, P.J. Assembly of the coronavirus envelope: Homotypic interactions between the M proteins. J. Virol. 2000, 74, 4967–4978. [Google Scholar] [CrossRef]
- Walls, A.C.; Tortorici, M.A.; Bosch, B.J.; Frenz, B.; Rottier, P.J.M.; DiMaio, F.; Rey, F.A.; Veesler, D. Cryo-electron microscopy structure of a coronavirus spike glycoprotein trimer. Nature 2016, 531, 114–117. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, F. Evidence for a common evolutionary origin of coronavirus spike protein receptor-binding subunits. J. Virol. 2012, 86, 2856–2858. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hoffmann, M.; Kleine-Weber, H.; Schroeder, S.; Kruger, N.; Herrler, T.; Erichsen, S.; Schiergens, T.S.; Herrler, G.; Wu, N.H.; Nitsche, A.; et al. SARS-CoV-2 Cell Entry Depends on ACE2 and TMPRSS2 and Is Blocked by a Clinically Proven Protease Inhibitor. Cell 2020, 181, 271–280.e8. [Google Scholar] [CrossRef] [PubMed]
- Hoffmann, M.; Hofmann-Winkler, H.; Pöhlmann, S. Priming Time: How Cellular Proteases Arm Coronavirus Spike Proteins. In Activation of Viruses by Host Proteases; Böttcher-Friebertshäuser, E., Garten, W., Klenk, H.D., Eds.; Springer International Publishing: Cham, Switzerland, 2018; pp. 71–98. [Google Scholar]
- Glowacka, I.; Bertram, S.; Muller, M.A.; Allen, P.; Soilleux, E.; Pfefferle, S.; Steffen, I.; Tsegaye, T.S.; He, Y.; Gnirss, K.; et al. Evidence that TMPRSS2 activates the severe acute respiratory syndrome coronavirus spike protein for membrane fusion and reduces viral control by the humoral immune response. J. Virol. 2011, 85, 4122–4134. [Google Scholar] [CrossRef] [Green Version]
- Simmons, G.; Gosalia, D.N.; Rennekamp, A.J.; Reeves, J.D.; Diamond, S.L.; Bates, P. Inhibitors of cathepsin L prevent severe acute respiratory syndrome coronavirus entry. Proc. Natl. Acad. Sci. USA 2005, 102, 11876–11881. [Google Scholar] [CrossRef] [Green Version]
- Federico, M. Lentivirus Gene Engineering Protocols; Humana Press: Rome, Italy, 2003; Volume 229, pp. 1–15. [Google Scholar]
- Hu, J.; Gao, Q.; He, C.; Huang, A.; Tang, N.; Wang, K. Development of cell-based pseudovirus entry assay to identify potential viral entry inhibitors and neutralizing antibodies against SARS-CoV-2. Genes Dis. 2020, 7, 551–557. [Google Scholar] [CrossRef]
- Johnson, M.C.; Lyddon, T.D.; Suarez, R.; Salcedo, B.; LePique, M.; Graham, M.; Ricana, C.; Robinson, C.; Ritter, D.G. Optimized Pseudotyping Conditions for the SARS-COV-2 Spike Glycoprotein. J. Virol. 2020, 94, e01062-20. [Google Scholar] [CrossRef]
- Daniels, R.W.; Rossano, A.J.; Macleod, G.T.; Ganetzky, B. Expression of multiple transgenes from a single construct using viral 2A peptides in Drosophila. PLoS ONE 2014, 9, e100637. [Google Scholar] [CrossRef]
- Liu, Z.; Chen, O.; Wall, J.B.J.; Zheng, M.; Zhou, Y.; Wang, L.; Vaseghi, H.R.; Qian, L.; Liu, J. Systematic comparison of 2A peptides for cloning multi-genes in a polycistronic vector. Sci. Rep. 2017, 7, 2193. [Google Scholar] [CrossRef]
- Ou, X.; Liu, Y.; Lei, X.; Li, P.; Mi, D.; Ren, L.; Guo, L.; Guo, R.; Chen, T.; Hu, J.; et al. Characterization of spike glycoprotein of SARS-CoV-2 on virus entry and its immune cross-reactivity with SARS-CoV. Nat. Commun. 2020, 11, 1620. [Google Scholar] [CrossRef] [Green Version]
- Yang, H.C.; Chen, C.H.; Wang, J.H.; Liao, H.C.; Yang, C.T.; Chen, C.W.; Lin, Y.C.; Kao, C.H.; Lu, M.J.; Liao, J.C. Analysis of genomic distributions of SARS-CoV-2 reveals a dominant strain type with strong allelic associations. Proc. Natl. Acad. Sci. USA 2020, 117, 30679–30686. [Google Scholar] [CrossRef]
- Laamarti, M.; Alouane, T.; Kartti, S.; Chemao-Elfihri, M.W.; Hakmi, M.; Essabbar, A.; Hlali, H.; Bendani, H.; Boumajdi, N.; Benhrif, O.; et al. Large scale genomic analysis of 3067 SARS-CoV-2 genomes reveals a clonal geo-distribution and a rich genetic variations of hotspots mutations. PLoS ONE 2020, 15, e0240345. [Google Scholar] [CrossRef]
- Duan, L.; Zheng, Q.; Zhang, H.; Niu, Y.; Lou, Y.; Wang, H. The SARS-CoV-2 Spike Glycoprotein Biosynthesis, Structure, Function, and Antigenicity: Implications for the Design of Spike-Based Vaccine Immunogens. Front. Immunol. 2020, 11, 576622. [Google Scholar] [CrossRef] [PubMed]
- Ke, Z.; Oton, J.; Qu, K.; Cortese, M.; Zila, V.; McKeane, L.; Nakane, T.; Zivanov, J.; Neufeldt, C.J.; Cerikan, B.; et al. Structures and distributions of SARS-CoV-2 spike proteins on intact virions. Nature 2020, 588, 498–502. [Google Scholar] [CrossRef] [PubMed]
- Turonova, B.; Sikora, M.; Schurmann, C.; Hagen, W.J.H.; Welsch, S.; Blanc, F.E.C.; von Bulow, S.; Gecht, M.; Bagola, K.; Horner, C.; et al. In situ structural analysis of SARS-CoV-2 spike reveals flexibility mediated by three hinges. Science 2020, 370, 203–208. [Google Scholar] [CrossRef] [PubMed]
- Harvey, W.T.; Carabelli, A.M.; Jackson, B.; Gupta, R.K.; Thomson, E.C.; Harrison, E.M.; Ludden, C.; Reeve, R.; Rambaut, A.; Peacock, S.J.; et al. SARS-CoV-2 variants, spike mutations and immune escape. Nat. Rev. Microbiol. 2021, 19, 409–424. [Google Scholar] [CrossRef] [PubMed]
- Yu, C.Y.; Chang, T.H.; Liang, J.J.; Chiang, R.L.; Lee, Y.L.; Liao, C.L.; Lin, Y.L. Dengue virus targets the adaptor protein MITA to subvert host innate immunity. PLoS Pathog. 2012, 8, e1002780. [Google Scholar] [CrossRef] [PubMed]
- Makarova, O.; Kamberov, E.; Margolis, B. Generation of deletion and point mutations with one primer in a single cloning step. Biotechniques 2000, 29, 970–972. [Google Scholar] [CrossRef] [PubMed] [Green Version]



Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wang, H.-I.; Chuang, Z.-S.; Kao, Y.-T.; Lin, Y.-L.; Liang, J.-J.; Liao, C.-C.; Liao, C.-L.; Lai, M.M.C.; Yu, C.-Y. Small Structural Proteins E and M Render the SARS-CoV-2 Pseudovirus More Infectious and Reveal the Phenotype of Natural Viral Variants. Int. J. Mol. Sci. 2021, 22, 9087. https://doi.org/10.3390/ijms22169087
Wang H-I, Chuang Z-S, Kao Y-T, Lin Y-L, Liang J-J, Liao C-C, Liao C-L, Lai MMC, Yu C-Y. Small Structural Proteins E and M Render the SARS-CoV-2 Pseudovirus More Infectious and Reveal the Phenotype of Natural Viral Variants. International Journal of Molecular Sciences. 2021; 22(16):9087. https://doi.org/10.3390/ijms22169087
Chicago/Turabian StyleWang, Hsin-I, Zih-Shiuan Chuang, Yu-Ting Kao, Yi-Ling Lin, Jian-Jong Liang, Chun-Che Liao, Ching-Len Liao, Michael M. C. Lai, and Chia-Yi Yu. 2021. "Small Structural Proteins E and M Render the SARS-CoV-2 Pseudovirus More Infectious and Reveal the Phenotype of Natural Viral Variants" International Journal of Molecular Sciences 22, no. 16: 9087. https://doi.org/10.3390/ijms22169087
APA StyleWang, H.-I., Chuang, Z.-S., Kao, Y.-T., Lin, Y.-L., Liang, J.-J., Liao, C.-C., Liao, C.-L., Lai, M. M. C., & Yu, C.-Y. (2021). Small Structural Proteins E and M Render the SARS-CoV-2 Pseudovirus More Infectious and Reveal the Phenotype of Natural Viral Variants. International Journal of Molecular Sciences, 22(16), 9087. https://doi.org/10.3390/ijms22169087