The Chloroplast of Chlamydomonas reinhardtii as a Testbed for Engineering Nitrogen Fixation into Plants
Abstract
:1. Introduction
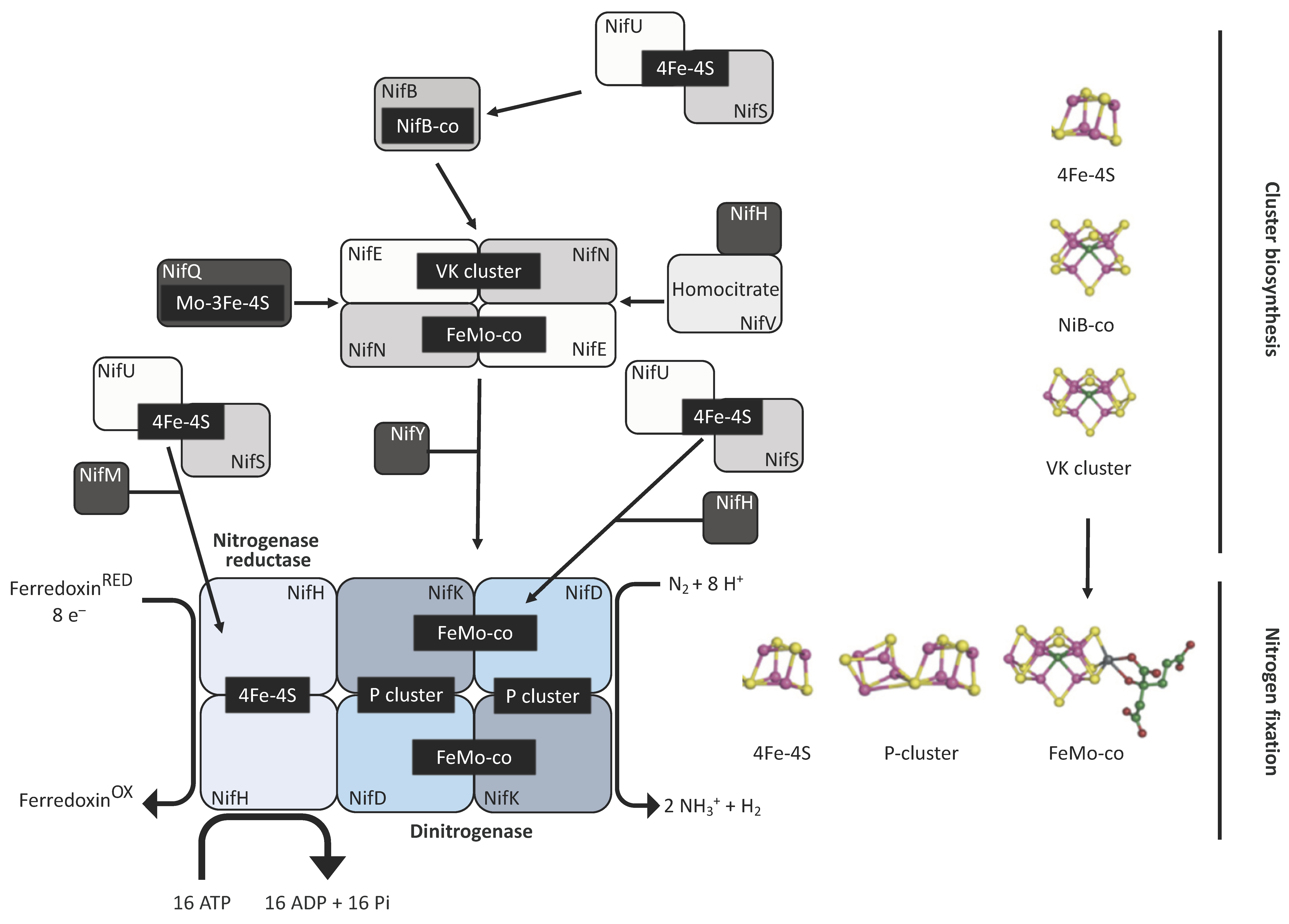
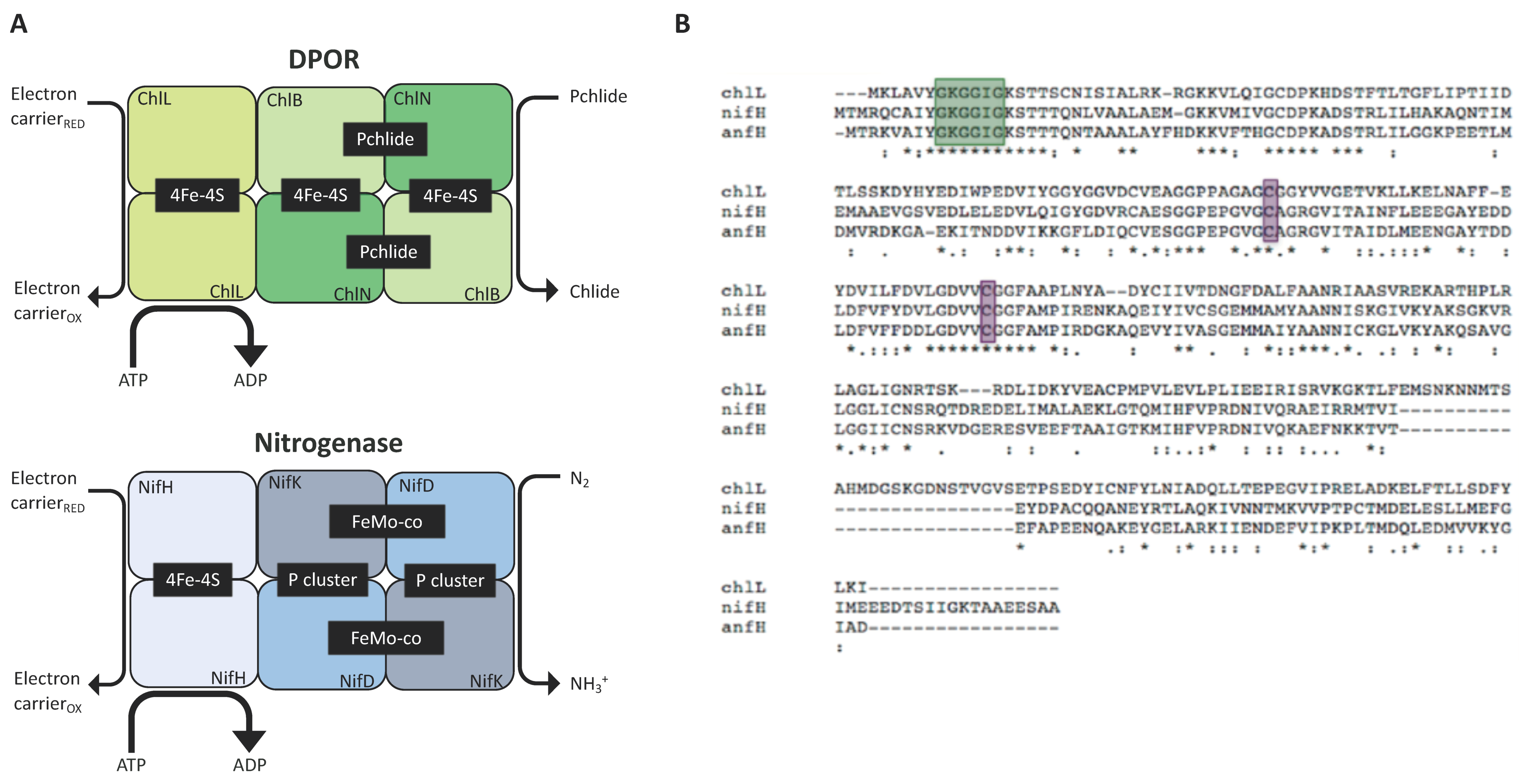
2. Nitrogenase in the Chloroplast
3. The Chlamydomonas Chloroplast as a Testbed
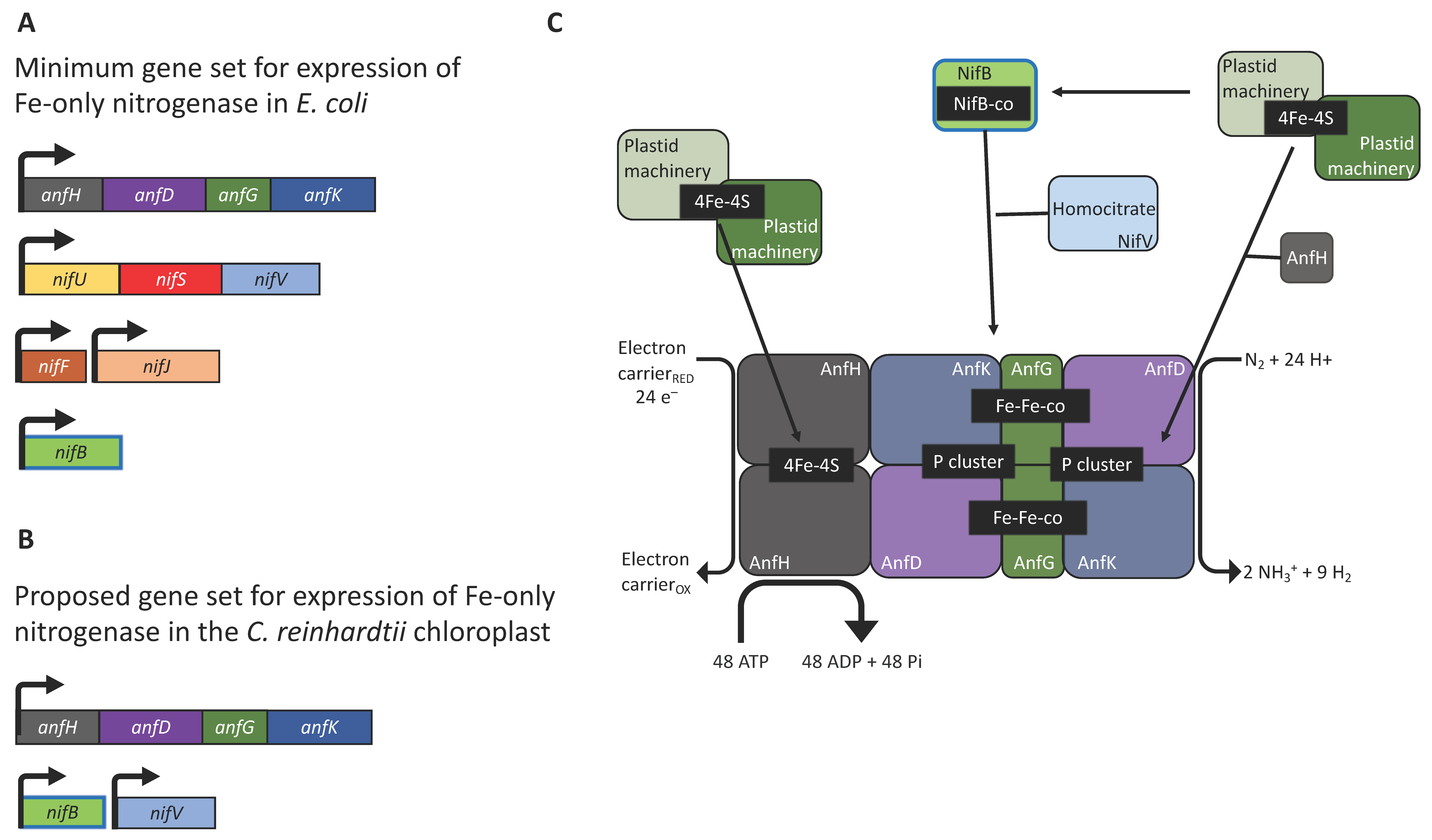
4. A Minimal Set of Nitrogenase Genes
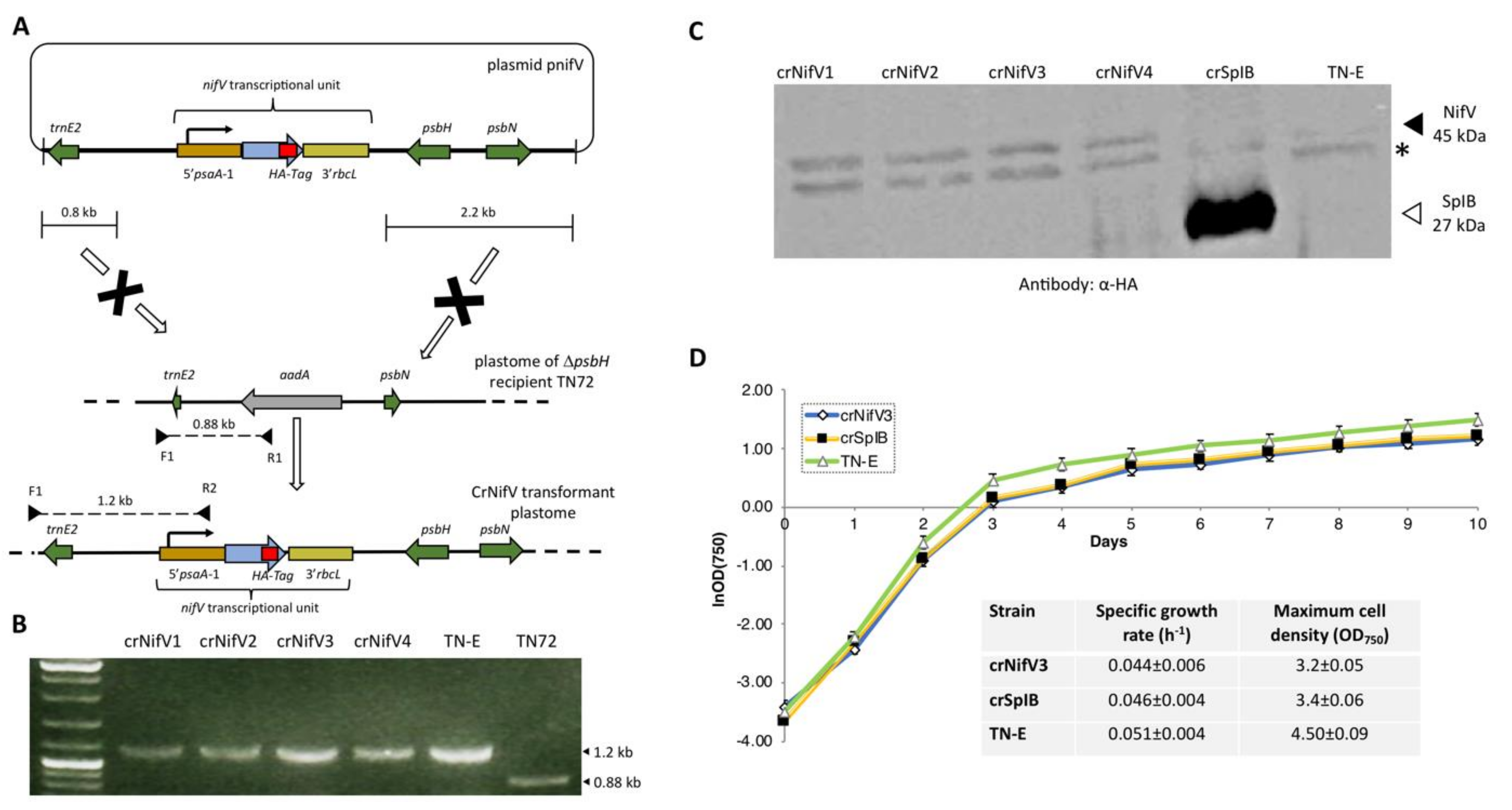
5. Expression of the anf/nif Genes in the Chloroplast of C. reinhardtii
6. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Conflicts of Interest
References
- Winter, H.C.; Burris, R.H. Nitrogenase. Annu. Rev. Biochem. 1976, 45, 409–426. [Google Scholar] [CrossRef] [PubMed]
- Hu, Y.; Ribbe, M.W. Nitrogenase assembly. Biochim. Biophys. Acta Bioenerg. 2012, 1827, 1112–1122. [Google Scholar] [CrossRef] [Green Version]
- Hu, Y.; Ribbe, M.W. Nitrogenase and homologs. J. Biol. Inorg. Chem. 2014, 20, 435–445. [Google Scholar] [CrossRef] [Green Version]
- Einsle, O.; Tezcan, F.A.; Andrade, S.L.A.; Schmid, B.; Yoshida, M.; Howard, J.B.; Rees, D.C. Nitrogenase MoFe-Protein at 1.16 A Resolution: A Central Ligand in the FeMo-Cofactor. Science 2002, 297, 1696–1700. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Duval, S.; Danyal, K.; Shaw, S.; Lytle, A.K.; Dean, D.R.; Hoffman, B.M.; Antony, E.; Seefeldt, L.C. Electron transfer precedes ATP hydrolysis during nitrogenase catalysis. Proc. Natl. Acad. Sci. USA 2013, 110, 16414–16419. [Google Scholar] [CrossRef] [Green Version]
- Smil, V. Nitrogen and Food Production: Proteins for Human Diets. Ambio 2002, 31, 126–131. [Google Scholar] [CrossRef]
- Good, A.; Beatty, P.H. Fertilizing Nature: A Tragedy of Excess in the Commons. PLoS Biol. 2011, 9, e1001124. [Google Scholar] [CrossRef]
- Sutton, M.A.; Oenema, O.; Erisman, J.W.; Leip, A.; Van Grinsven, H.; Winiwarter, W. Too much of a good thing. Nat. Cell Biol. 2011, 472, 159–161. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rogers, C.; Oldroyd, G.E.D. Synthetic biology approaches to engineering the nitrogen symbiosis in cereals. J. Exp. Bot. 2014, 65, 1939–1946. [Google Scholar] [CrossRef] [Green Version]
- Merrick, M.; Dixon, R. Why don’t plants fix nitrogen? Trends Biotechnol. 1984, 2, 162–166. [Google Scholar] [CrossRef]
- Peoples, M.B.; Herridge, D.F.; Ladha, J.K. Biological nitrogen fixation: An efficient source of nitrogen for sustainable agricultural production? In Management of Biological Nitrogen Fixation for the Development of More Productive and Sustainable Agricultural Systems; Ladha, J.K., Peoples, M.B., Eds.; Springer: Dordrecht, The Netherlands, 1995; pp. 3–28. [Google Scholar] [CrossRef]
- Britto, D.T.; Kronzucker, H.J. Bioengineering nitrogen acquisition in rice: Can novel initiatives in rice genomics and physiology contribute to global food security? BioEssays 2004, 26, 683–692. [Google Scholar] [CrossRef]
- Vicente, E.J.; Dean, D.R. Keeping the nitrogen-fixation dream alive. Proc. Natl. Acad. Sci. USA 2017, 114, 3009–3011. [Google Scholar] [CrossRef] [Green Version]
- A Geddes, B.; Ryu, M.-H.; Mus, F.; Costas, A.G.; Peters, J.; A Voigt, C.; Poole, P. Use of plant colonizing bacteria as chassis for transfer of N2-fixation to cereals. Curr. Opin. Biotechnol. 2015, 32, 216–222. [Google Scholar] [CrossRef] [PubMed]
- Dixon, R.A.; Postgate, J.R. Genetic Transfer of Nitrogen Fixation from Klebsiella pneumoniae to Escherichia coli. Nat. Cell Biol. 1972, 237, 102–103. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.; Xie, X.; Wang, X.; Dixon, R.; Wang, Y. Reconstruction and minimal gene requirements for the alternative iron-only nitrogenase in Escherichia coli. Proc. Natl. Acad. Sci. USA 2014, 111, E3718–E3725. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, X.; Yang, J.-G.; Chen, L.; Wang, J.-L.; Cheng, Q.; Dixon, R.; Wang, Y.-P. Using Synthetic Biology to Distinguish and Overcome Regulatory and Functional Barriers Related to Nitrogen Fixation. PLoS ONE 2013, 8, e68677. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Temme, K.; Zhao, D.; Voigt, C.A. Refactoring the nitrogen fixation gene cluster from Klebsiella oxytoca. Proc. Natl. Acad. Sci. USA 2012, 109, 7085–7090. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Allen, R.; Tilbrook, K.; Warden, A.; Campbell, P.C.; Rolland, V.; Singh, S.P.; Wood, C.C. Expression of 16 Nitrogenase Proteins within the Plant Mitochondrial Matrix. Front. Plant Sci. 2017, 8, 287. [Google Scholar] [CrossRef] [PubMed]
- Burén, S.; Pratt, K.; Jiang, X.; Guo, Y.; Jimenez-Vicente, E.; Echavarri-Erasun, C.; Dean, D.R.; Saaem, I.; Gordon, D.B.; Voigt, C.A.; et al. Biosynthesis of the nitrogenase active-site cofactor precursor NifB-co inSaccharomyces cerevisiae. Proc. Natl. Acad. Sci. USA 2019, 116, 25078–25086. [Google Scholar] [CrossRef] [Green Version]
- López-Torrejón, G.; Jiménez-Vicente, E.; Buesa, J.M.; Hernandez, J.A.; Verma, H.K.; Rubio, L.M. Expression of a functional oxygen-labile nitrogenase component in the mitochondrial matrix of aerobically grown yeast. Nat. Commun. 2016, 7, 11426. [Google Scholar] [CrossRef] [Green Version]
- Okada, S.; Gregg, C.M.; Allen, R.S.; Menon, A.; Hussain, D.; Gillespie, V.; Johnston, E.; Byrne, K.; Colgrave, M.L.; Wood, C.C. A Synthetic Biology Workflow Reveals Variation in Processing and Solubility of Nitrogenase Proteins Targeted to Plant Mitochondria, and Differing Tolerance of Targeting Sequences in a Bacterial Nitrogenase Assay. Front. Plant Sci. 2020, 11. [Google Scholar] [CrossRef]
- Xiang, N.; Guo, C.; Liu, J.; Xu, H.; Dixon, R.; Yang, J.; Wang, Y.-P. Using synthetic biology to overcome barriers to stable expression of nitrogenase in eukaryotic organelles. Proc. Natl. Acad. Sci. USA 2020, 117, 16537–16545. [Google Scholar] [CrossRef]
- Burén, S.; Young, E.M.; Sweeny, E.; Lopez-Torrejón, G.; Veldhuizen, M.; Voigt, C.A.; Rubio, L.M. Formation of Nitrogenase NifDK Tetramers in the Mitochondria of Saccharomyces cerevisiae. ACS Synth. Biol. 2017, 6, 1043–1055. [Google Scholar] [CrossRef]
- Dixon, R.; Cheng, Q.; Shen, G.-F.; Day, A.; Dowson-Day, M. Nif gene transfer and expression in chloroplasts: Prospects and problems. Plant Soil 1997, 194, 193–203. [Google Scholar] [CrossRef]
- Yang, J.; Xie, X.; Yang, M.; Dixon, R.; Wang, Y. Modular electron-transport chains from eukaryotic organelles function to support nitrogenase activity. Proc. Natl. Acad. Sci. USA 2017, 114, E2460–E2465. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cheng, Q.; Day, A.; Dowson-Day, M.; Shen, G.-F.; Dixon, R. The Klebsiella pneumoniae nitrogenase Fe protein gene (nifH) functionally substitutes for the chlL gene in Chlamydomonas reinhardtii. Biochem. Biophys. Res. Commun. 2005, 329, 966–975. [Google Scholar] [CrossRef] [PubMed]
- Ivleva, N.B.; Groat, J.; Staub, J.M.; Stephens, M. Expression of Active Subunit of Nitrogenase via Integration into Plant Organelle Genome. PLoS ONE 2016, 11, e0160951. [Google Scholar] [CrossRef] [Green Version]
- Adem, M.; Beyene, D.; Feyissa, T. Recent achievements obtained by chloroplast transformation. Plant Methods 2017, 13, 1–11. [Google Scholar] [CrossRef] [Green Version]
- Rochaix, J. Chlamydomonas reinhardtii as the Photosynthetic Yeast. Annu. Rev. Genet. 1995, 29, 209–230. [Google Scholar] [CrossRef] [PubMed]
- Harris, E.H. The Chlamydomonas Sourcebook: Introduction to Chlamydomonas and Its Laboratory Use; Academic Press: Cambridge, MA, USA, 2013; Volume 1. [Google Scholar] [CrossRef]
- Salomé, P.A.; Merchant, S.S. A Series of Fortunate Events: Introducing Chlamydomonas as a Reference Organism. Plant Cell 2019, 31, 1682–1707. [Google Scholar] [CrossRef] [Green Version]
- Scaife, M.A.; Nguyen, G.T.; Rico, J.; Lambert, D.; Helliwell, K.; Smith, A. Establishing Chlamydomonas reinhardtii as an industrial biotechnology host. Plant J. 2015, 82, 532–546. [Google Scholar] [CrossRef] [PubMed]
- Dyo, Y.M.; Purton, S. The algal chloroplast as a synthetic biology platform for production of therapeutic proteins. Microbiology 2018, 164, 113–121. [Google Scholar] [CrossRef]
- Rolland, N.; Curien, G.; Finazzi, G.; Kuntz, M.; Maréchal, E.; Matringe, M.; Ravanel, S.; Seigneurin-Berny, D. The Biosynthetic Capacities of the Plastids and Integration Between Cytoplasmic and Chloroplast Processes. Annu. Rev. Genet. 2012, 46, 233–264. [Google Scholar] [CrossRef]
- Siddiqui, A.; Wei, Z.; Boehm, M.; Ahmad, N. Engineering microalgae through chloroplast transformation to produce high-value industrial products. Biotechnol. Appl. Biochem. 2019, 67, 30–40. [Google Scholar] [CrossRef]
- Yang, W.; Catalanotti, C.; Wittkopp, T.M.; Posewitz, M.C.; Grossman, A.R. Algae after dark: Mechanisms to cope with anoxic/hypoxic conditions. Plant J. 2015, 82, 481–503. [Google Scholar] [CrossRef]
- Grossman, A.R.; Catalanotti, C.; Yang, W.; Dubini, A.; Magneschi, L.; Subramanian, V.; Posewitz, M.C.; Seibert, M. Multiple facets of anoxic metabolism and hydrogen production in the unicellular green alga Chlamydomonas reinhardtii. New Phytol. 2010, 190, 279–288. [Google Scholar] [CrossRef]
- López-Torrejón, G.; Burén, S.; Veldhuizen, M.; Rubio, L.M. Biosynthesis of cofactor-activatable iron-only nitrogenase in Saccharomyces cerevisiae. Microb. Biotechnol. 2021, 14, 1073–1083. [Google Scholar] [CrossRef]
- Bock, R. Genetic engineering of the chloroplast: Novel tools and new applications. Curr. Opin. Biotechnol. 2014, 26, 7–13. [Google Scholar] [CrossRef] [PubMed]
- Rojas, M.; Yu, Q.; Williams-Carrier, R.; Maliga, P.; Barkan, A. Engineered PPR proteins as inducible switches to activate the expression of chloroplast transgenes. Nat. Plants 2019, 5, 505–511. [Google Scholar] [CrossRef] [PubMed]
- Caroca, R.; Howell, K.A.; Hasse, C.; Ruf, S.; Bock, R. Design of chimeric expression elements that confer high-level gene activity in chromoplasts. Plant J. 2012, 73, 368–379. [Google Scholar] [CrossRef] [PubMed]
- Fuentes, P.; Armarego-Marriott, T.; Bock, R. Plastid transformation and its application in metabolic engineering. Curr. Opin. Biotechnol. 2018, 49, 10–15. [Google Scholar] [CrossRef]
- Schindel, H.S.; Piatek, A.A.; Stewart, C.N.; Lenaghan, S.C. The plastid genome as a chassis for synthetic biology-enabled metabolic engineering: Players in gene expression. Plant Cell Rep. 2018, 37, 1419–1429. [Google Scholar] [CrossRef]
- Green, B.R. Chloroplast genomes of photosynthetic eukaryotes. Plant J. 2011, 66, 34–44. [Google Scholar] [CrossRef]
- Gallaher, S.D.; Fitz-Gibbon, S.T.; Strenkert, D.; Purvine, S.O.; Pellegrini, M.; Merchant, S.S. High-throughput sequencing of the chloroplast and mitochondrion of Chlamydomonas reinhardtii to generate improved de novo assemblies, analyze expression patterns and transcript speciation, and evaluate diversity among laboratory strains and wild isolates. Plant J. 2018, 93, 545–565. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Purton, S.; Szaub, J.B.; Wannathong, T.; Young, R.; Economou, C.K. Genetic engineering of algal chloroplasts: Progress and prospects. Russ. J. Plant Physiol. 2013, 60, 491–499. [Google Scholar] [CrossRef]
- Zedler, J.A.Z.; Gangl, D.; Hamberger, B.; Purton, S.; Robinson, C.; Hamberger, B. Stable expression of a bifunctional diterpene synthase in the chloroplast of Chlamydomonas reinhardtii. Environ. Boil. Fishes 2014, 27, 2271–2277. [Google Scholar] [CrossRef]
- Gangl, D.; Zedler, J.A.Z.; Wlodarczyk, A.; Jensen, P.E.; Purton, S.; Robinson, C. Expression and membrane-targeting of an active plant cytochrome P450 in the chloroplast of the green alga Chlamydomonas reinhardtii. Phytochemistry 2014, 110, 22–28. [Google Scholar] [CrossRef]
- Papaefthimiou, D.; Diretto, G.; Demurtas, O.C.; Mini, P.; Ferrante, P.; Giuliano, G.; Kanellis, A.K. Heterologous production of labdane-type diterpenes in the green alga Chlamydomonas reinhardtii. Phytochemistry 2019, 167, 112082. [Google Scholar] [CrossRef] [PubMed]
- Paik, S.-M.; Kim, J.; Jin, E.; Jeon, N.L. Overproduction of recombinant E. coli malate synthase enhances Chlamydomonas reinhardtii biomass by upregulating heterotrophic metabolism. Bioresour. Technol. 2018, 272, 594–598. [Google Scholar] [CrossRef]
- Tevatia, R.; Payne, S.; Allen, J.; White, D.; Clemente, T.E.; Cerutti, H.; Demirel, Y.; Blum, P. A synthetic cdo/csad taurine pathway in the green unicellular alga Chlamydomonas reinhardtii. Algal Res. 2019, 40, 101491. [Google Scholar] [CrossRef]
- Macedo-Osorio, K.S.; Pérez-España, V.H.; Garibay-Orijel, C.; Guzmán-Zapata, D.; Durán-Figueroa, N.V.; Badillo-Corona, J.A. Intercistronic expression elements (IEE) from the chloroplast of Chlamydomonas reinhardtii can be used for the expression of foreign genes in synthetic operons. Plant Mol. Biol. 2018, 98, 303–317. [Google Scholar] [CrossRef]
- Larrea-Alvarez, M.; Purton, S. Multigenic engineering of the chloroplast genome in the green alga Chlamydomonas reinhardtii. Microbiology 2020, 166, 510–515. [Google Scholar] [CrossRef]
- Jackson, H.O.; Taunt, H.N.; Pawel, M.M.; Smith, A.G.; Purton, S. The algal chloroplast as a testbed for synthetic biology designs aimed at radically rewiring plant metabolism. Front. Plant Sci. 2021, in press. [Google Scholar] [CrossRef]
- Young, R.; Purton, S. CITRIC: Cold-Inducible translational readthrough in the chloroplast of Chlamydomonas reinhardtii using a novel temperature-sensitive transfer RNA. Microb. Cell Factories 2018, 17, 1–12. [Google Scholar] [CrossRef] [Green Version]
- Carrera-Pacheco, S.E.; Hankamer, B.; Oey, M. Light and heat-shock mediated TDA1 overexpression as a tool for controlled high-yield recombinant protein production in Chlamydomonas reinhardtii chloroplasts. Algal Res. 2020, 48, 101921. [Google Scholar] [CrossRef]
- Crozet, P.; Navarro, F.J.; Willmund, F.; Mehrshahi, P.; Bakowski, K.; Lauersen, K.; Pérez-Pérez, M.-E.; Auroy, P.; Rovira, A.G.; Sauret-Gueto, S.; et al. Birth of a Photosynthetic Chassis: A MoClo Toolkit Enabling Synthetic Biology in the Microalga Chlamydomonas reinhardtii. ACS Synth. Biol. 2018, 7, 2074–2086. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Peden, E.A.; Boehm, M.; Mulder, D.W.; Davis, R.; Old, W.M.; King, P.W.; Ghirardi, M.L.; Dubini, A. Identification of Global Ferredoxin Interaction Networks in Chlamydomonas reinhardtii. J. Biol. Chem. 2013, 288, 35192–35209. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mosebach, L.; Heilmann, C.; Mutoh, R.; Gäbelein, P.; Steinbeck, J.; Happe, T.; Ikegami, T.; Hanke, G.; Kurisu, G.; Hippler, M. Association of Ferredoxin:NADP+ oxidoreductase with the photosynthetic apparatus modulates electron transfer in Chlamydomonas reinhardtii. Photosynth. Res. 2017, 134, 291–306. [Google Scholar] [CrossRef] [PubMed]
- Sawyer, A.; Winkler, M. Evolution of Chlamydomonas reinhardtii ferredoxins and their interactions with [FeFe]-hydrogenases. Photosynth. Res. 2017, 134, 307–316. [Google Scholar] [CrossRef]
- Jasniewski, A.J.; Lee, C.C.; Ribbe, M.W.; Hu, Y. Reactivity, Mechanism, and Assembly of the Alternative Nitrogenases. Chem. Rev. 2020, 120, 5107–5157. [Google Scholar] [CrossRef]
- Eseverri, Á.; López-Torrejón, G.; Jiang, X.; Burén, S.; Rubio, L.M.; Caro, E. Use of synthetic biology tools to optimize the production of active nitrogenase Fe protein in chloroplasts of tobacco leaf cells. Plant Biotechnol. J. 2020, 18, 1882–1896. [Google Scholar] [CrossRef] [Green Version]
- Moser, J.; Bröcker, M.J. Enzymatic Systems with Homology to Nitrogenase. Methods Mol. Biol. 2011, 766, 67–77. [Google Scholar] [CrossRef]
- Taunt, H.N.; Stoffels, L.; Purton, S. Green biologics: The algal chloroplast as a platform for making biopharmaceuticals. Bioengineered 2017, 9, 48–54. [Google Scholar] [CrossRef] [Green Version]
- Armstrong, G.A. Greening in the dark: Light-Independent chlorophyll biosynthesis from anoxygenic photosynthetic bacteria to gymnosperms. J. Photochem. Photobiol. B Biol. 1998, 43, 87–100. [Google Scholar] [CrossRef]
- Jiang, X.; Payá-Tormo, L.; Coroian, D.; García-Rubio, I.; Castellanos-Rueda, R.; Eseverri, Á.; López-Torrejón, G.; Burén, S.; Rubio, L.M. Exploiting genetic diversity and gene synthesis to identify superior nitrogenase NifH protein variants to engineer N2-fixation in plants. Commun. Biol. 2021, 4, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Zheng, L.; White, R.H.; Dean, D.R. Purification of the Azotobacter vinelandii nifV-encoded homocitrate synthase. J. Bacteriol. 1997, 179, 5963–5966. [Google Scholar] [CrossRef] [Green Version]
- Liu, X.; Wang, M.; Song, Y.; Li, Y.; Liu, P.; Shi, H.; Li, Y.; Hao, T.; Zhang, H.; Jiang, W.; et al. Combined Assembly and Targeted Integration of Multigene for Nitrogenase Biosynthetic Pathway inSaccharomyces cerevisiae. ACS Synth. Biol. 2019, 8, 1766–1775. [Google Scholar] [CrossRef] [PubMed]
- Young, R.E.B.; Purton, S. Cytosine deaminase as a negative selectable marker for the microalgal chloroplast: A strategy for the isolation of nuclear mutations that affect chloroplast gene expression. Plant J. 2014, 80, 915–925. [Google Scholar] [CrossRef] [Green Version]
- Wannathong, T.; Waterhouse, J.C.; Young, R.E.B.; Economou, C.K.; Purton, S. New tools for chloroplast genetic engineering allow the synthesis of human growth hormone in the green alga Chlamydomonas reinhardtii. Appl. Microbiol. Biotechnol. 2016, 100, 5467–5477. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Larrea-Alvarez, M.; Young, R.; Purton, S. A Simple Technology for Generating Marker-Free Chloroplast Transformants of the Green Alga Chlamydomonas reinhardtii. Methods Mol. Biol. 2021, 2317, 293–304. [Google Scholar] [CrossRef]
- Esland, L.; Larrea-Alvarez, M.; Purton, S. Selectable Markers and Reporter Genes for Engineering the Chloroplast of Chlamydomonas reinhardtii. Biology 2018, 7, 46. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rochaix, J.-D.; Surzycki, R.; Ramundo, S. Tools for Regulated Gene Expression in the Chloroplast of Chlamydomonas. Methods Mol. Biol. 2014, 1132, 413–424. [Google Scholar] [CrossRef] [PubMed]
- Mus, F.; Dubini, A.; Seibert, M.; Posewitz, M.C.; Grossman, A.R. Anaerobic Acclimation in Chlamydomonas reinhardtii. J. Biol. Chem. 2007, 282, 25475–25486. [Google Scholar] [CrossRef] [PubMed] [Green Version]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Larrea-Álvarez, M.; Purton, S. The Chloroplast of Chlamydomonas reinhardtii as a Testbed for Engineering Nitrogen Fixation into Plants. Int. J. Mol. Sci. 2021, 22, 8806. https://doi.org/10.3390/ijms22168806
Larrea-Álvarez M, Purton S. The Chloroplast of Chlamydomonas reinhardtii as a Testbed for Engineering Nitrogen Fixation into Plants. International Journal of Molecular Sciences. 2021; 22(16):8806. https://doi.org/10.3390/ijms22168806
Chicago/Turabian StyleLarrea-Álvarez, Marco, and Saul Purton. 2021. "The Chloroplast of Chlamydomonas reinhardtii as a Testbed for Engineering Nitrogen Fixation into Plants" International Journal of Molecular Sciences 22, no. 16: 8806. https://doi.org/10.3390/ijms22168806
APA StyleLarrea-Álvarez, M., & Purton, S. (2021). The Chloroplast of Chlamydomonas reinhardtii as a Testbed for Engineering Nitrogen Fixation into Plants. International Journal of Molecular Sciences, 22(16), 8806. https://doi.org/10.3390/ijms22168806






