Lipid Droplets and Their Autophagic Turnover via the Raft-Like Vacuolar Microdomains
Abstract
:1. Introduction
2. Lipid Droplets
2.1. Structure of Lipid Droplets
2.2. Biogenesis of Lipid Droplets
2.3. Induction of Lipid Droplet Biogenesis
2.4. Fusion and Fission of Lipid Droplets
2.5. Contact Sites of Lipid Droplets
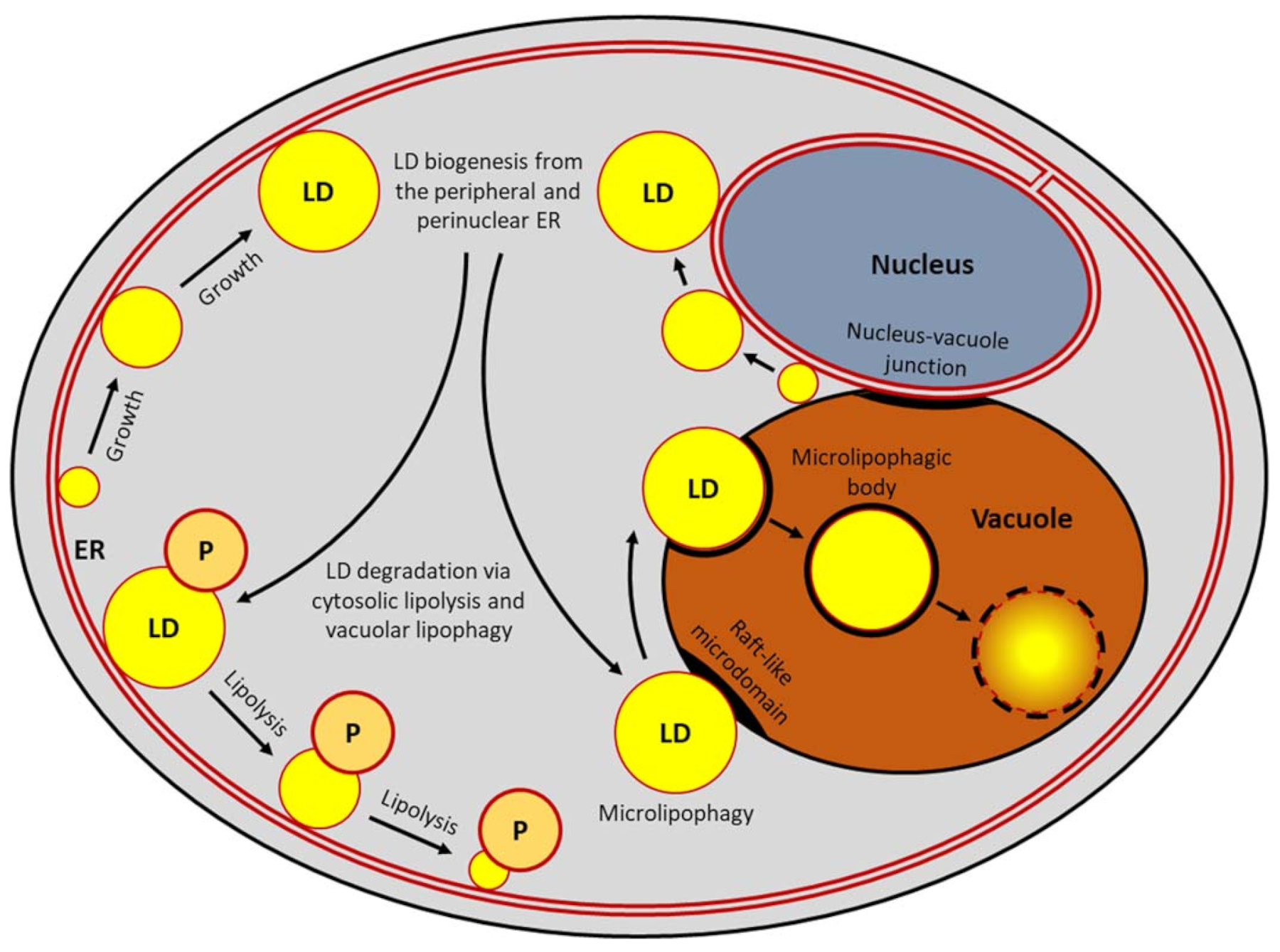
3. Lipid Droplet Turnover
3.1. Microlipophagy
3.2. Induction of Microlipophagy
3.3. Molecular Mechanism of Microlipophagy
3.3.1. Formation of Vacuolar Microdomains as a Prerequisite for Microlipophagy
3.3.2. The Requirement of Autophagic Proteins for Vacuolar Microdomain Formation and Microlipophagy
3.3.3. Role of ESCRT Machinery in Microlipophagy
3.3.4. Role of Nucleus-Vacuole Junction in Microlipophagy
4. Lipid Droplets and Diseases
5. Conclusions and Future Perspectives
- Is microlipophagy the only morphological mode of lipophagy in yeast?
- Why are microlipophagy mechanisms distinct under different conditions?
- What is the nature of the contact sites between the vacuolar microdomains and LDs?
- What are the exact functions of autophagic proteins in microlipophagy?
- What is the autophagic receptor protein during microlipophagy (if any)?
- What machinery is required to recycle lipids after degradation?
- How extensive is the crosstalk between lipophagy and lipolysis in yeast?
- How does the cell decide whether to proceed for lipolysis or lipophagy?
Funding
Conflicts of Interest
References
- Klionsky, D.J. Autophagy revisited: A conversation with Christian de Duve. Autophagy 2008, 4, 740–743. [Google Scholar] [CrossRef] [Green Version]
- Abeliovich, H.; Klionsky, D.J. Autophagy in yeast: Mechanistic insights and physiological function. Microbiol. Mol. Biol. Rev. 2001, 65, 463–479. [Google Scholar] [CrossRef] [Green Version]
- Ohsumi, Y. Historical landmarks of autophagy research. Cell Res. 2014, 24, 9–23. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, F.; Vierstra, R.D. Autophagy: A multifaceted intracellular system for bulk and selective recycling. Trends Plant Sci. 2012, 17, 526–537. [Google Scholar] [CrossRef]
- Shimizu, S.; Kanaseki, T.; Mizushima, N.; Mizuta, T.; Arakawa-Kobayashi, S.; Thompson, C.B.; Tsujimoto, Y. Role of Bcl-2 family proteins in a non-apoptotic programmed cell death dependent on autophagy genes. Nat. Cell. Biol. 2004, 6, 1221–1228. [Google Scholar] [CrossRef]
- Tsukamoto, S.; Kuma, A.; Murakami, M.; Kishi, C.; Yamamoto, A.; Mizushima, N. Autophagy is essential for preimplantation development of mouse embryos. Science 2008, 321, 117–120. [Google Scholar] [CrossRef]
- Meléndez, A.; Levine, B. Autophagy in C. elegans. WormBook Online Rev. C. elegans Biol. 2009. [Google Scholar] [CrossRef] [Green Version]
- Mizushima, N.; Komatsu, M. Autophagy: Renovation of cells and tissues. Cell 2011, 147, 728–741. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- White, E. The role for autophagy in cancer. J. Clin. Invest. 2015, 125, 42–46. [Google Scholar] [CrossRef] [Green Version]
- Quan, W.; Lim, Y.M.; Lee, M.S. Role of autophagy in diabetes and endoplasmic reticulum stress of pancreatic β-cells. Exp. Mol. Med. 2012, 44, 81–88. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Deretic, V.; Saitoh, T.; Akira, S. Autophagy in infection, inflammation and immunity. Nat. Rev. Immunol. 2013, 13, 722–737. [Google Scholar] [CrossRef]
- Nixon, R.A. The role of autophagy in neurodegenerative disease. Nat. Med. 2013, 19, 983–997. [Google Scholar] [CrossRef]
- Farré, J.C.; Subramani, S. Mechanistic insights into selective autophagy pathways: Lessons from yeast. Nat. Rev. Mol. Cell Biol. 2016, 17, 537–552. [Google Scholar] [CrossRef] [PubMed]
- Li, L.; Tong, M.; Fu, Y.; Chen, F.; Zhang, S.; Chen, H.; Ma, X.; Li, D.; Liu, X.; Zhong, Q. Lipids and membrane-associated proteins in autophagy. Protein Cell 2021, 12, 520–544. [Google Scholar] [CrossRef]
- Fujimoto, T.; Parton, R.G. Not just fat: The structure and function of the lipid droplet. Cold Spring Harb. Perspect. Biol. 2011, 3, a004838. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Walther, T.C.; Farese, R.V., Jr. Lipid droplets and cellular lipid metabolism. Annu. Rev. Biochem. 2012, 81, 687–714. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Brasaemle, D.L.; Wolins, N.E. Packaging of fat: An evolving model of lipid droplet assembly and expansion. J. Biol. Chem. 2012, 287, 2273–2279. [Google Scholar] [CrossRef] [Green Version]
- Bartz, R.; Li, W.H.; Venables, B.; Zehmer, J.K.; Roth, M.R.; Welti, R.; Anderson, R.G.; Liu, P.; Chapman, K.D. Lipidomics reveals that adiposomes store ether lipids and mediate phospholipid traffic. J. Lipid Res. 2007, 48, 837–847. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Martin, S.; Parton, R.G. Lipid droplets: A unified view of a dynamic organelle. Nat. Rev. Mol. Cell Biol. 2006, 7, 373–378. [Google Scholar] [CrossRef] [PubMed]
- Zechner, R.; Zimmermann, R.; Eichmann, T.O.; Kohlwein, S.D.; Haemmerle, G.; Lass, A.; Madeo, F. FAT SIGNALS--lipases and lipolysis in lipid metabolism and signaling. Cell Metab. 2012, 15, 279–291. [Google Scholar] [CrossRef] [Green Version]
- Cermelli, S.; Guo, Y.; Gross, S.P.; Welte, M.A. The lipid-droplet proteome reveals that droplets are a protein-storage depot. Curr. Biol. 2006, 16, 1783–1795. [Google Scholar] [CrossRef] [Green Version]
- Fujimoto, T.; Ohsaki, Y.; Cheng, J.; Suzuki, M.; Shinohara, Y. Lipid droplets: A classic organelle with new outfits. Histochem Cell Biol. 2008, 130, 263–279. [Google Scholar] [CrossRef] [Green Version]
- McLauchlan, J. Lipid droplets and hepatitis C virus infection. Biochim. Biophys. Acta. 2009, 1791, 552–559. [Google Scholar] [CrossRef] [PubMed]
- Liu, P.; Bartz, R.; Zehmer, J.K.; Ying, Y.S.; Zhu, M.; Serrero, G.; Anderson, R.G. Rab-regulated interaction of early endosomes with lipid droplets. Biochim. Biophys. Acta. 2007, 1773, 784–793. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Xu, S.; Zhang, X.; Liu, P. Lipid droplet proteins and metabolic diseases. Biochim. Biophys. Acta. Mol. Basis. Dis. 2018, 1864, 1968–1983. [Google Scholar] [CrossRef] [PubMed]
- Bozza, P.T.; Viola, J.P. Lipid droplets in inflammation and cancer. Prostaglandins Leukot Essent Fat. Acids 2010, 82, 243–250. [Google Scholar] [CrossRef]
- Connerth, M.; Grillitsch, K.; Köfeler, H.; Daum, G. Analysis of lipid particles from yeast. Methods Mol. Biol. 2009, 579, 359–374. [Google Scholar] [CrossRef]
- Leber, R.; Zinser, E.; Zellnig, G.; Paltauf, F.; Daum, G. Characterization of lipid particles of the yeast, Saccharomyces cerevisiae. Yeast 1994, 10, 1421–1428. [Google Scholar] [CrossRef]
- Athenstaedt, K.; Zweytick, D.; Jandrositz, A.; Kohlwein, S.D.; Daum, G. Identification and characterization of major lipid particle proteins of the yeast Saccharomyces cerevisiae. J. Bacteriol. 1999, 181, 6441–6448. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Czabany, T.; Wagner, A.; Zweytick, D.; Lohner, K.; Leitner, E.; Ingolic, E.; Daum, G. Structural and biochemical properties of lipid particles from the yeast Saccharomyces cerevisiae. J. Biol. Chem. 2008, 283, 17065–17074. [Google Scholar] [CrossRef] [Green Version]
- Kimmel, A.R.; Sztalryd, C. The Perilipins: Major Cytosolic Lipid Droplet-Associated Proteins and Their Roles in Cellular Lipid Storage, Mobilization, and Systemic Homeostasis. Annu. Rev. Nutr. 2016, 36, 471–509. [Google Scholar] [CrossRef]
- Yang, H.; Galea, A.; Sytnyk, V.; Crossley, M. Controlling the size of lipid droplets: Lipid and protein factors. Curr. Opin. Cell Biol. 2012, 24, 509–516. [Google Scholar] [CrossRef] [PubMed]
- Wang, C.W.; Miao, Y.H.; Chang, Y.S. A sterol-enriched vacuolar microdomain mediates stationary phase lipophagy in budding yeast. J. Cell Biol. 2014, 206, 357–366. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- van Zutphen, T.; Todde, V.; de Boer, R.; Kreim, M.; Hofbauer, H.F.; Wolinski, H.; Veenhuis, M.; van der Klei, I.J.; Kohlwein, S.D. Lipid droplet autophagy in the yeast Saccharomyces cerevisiae. Mol. Biol. Cell 2014, 25, 290–301. [Google Scholar] [CrossRef] [Green Version]
- Casanovas, A.; Sprenger, R.R.; Tarasov, K.; Ruckerbauer, D.E.; Hannibal-Bach, H.K.; Zanghellini, J.; Jensen, O.N.; Ejsing, C.S. Quantitative analysis of proteome and lipidome dynamics reveals functional regulation of global lipid metabolism. Chem. Biol. 2015, 22, 412–425. [Google Scholar] [CrossRef] [Green Version]
- Currie, E.; Guo, X.; Christiano, R.; Chitraju, C.; Kory, N.; Harrison, K.; Haas, J.; Walther, T.C.; Farese, R.V., Jr. High confidence proteomic analysis of yeast LDs identifies additional droplet proteins and reveals connections to dolichol synthesis and sterol acetylation. J. Lipid Res. 2014, 55, 1465–1477. [Google Scholar] [CrossRef] [Green Version]
- Hodges, B.D.; Wu, C.C. Proteomic insights into an expanded cellular role for cytoplasmic lipid droplets. J. Lipid Res. 2010, 51, 262–273. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Farese, R.V., Jr.; Walther, T.C. Lipid droplets finally get a little R-E-S-P-E-C-T. Cell 2009, 139, 855–860. [Google Scholar] [CrossRef] [Green Version]
- Radulovic, M.; Knittelfelder, O.; Cristobal-Sarramian, A.; Kolb, D.; Wolinski, H.; Kohlwein, S.D. The emergence of lipid droplets in yeast: Current status and experimental approaches. Curr. Genet. 2013, 59, 231–242. [Google Scholar] [CrossRef] [Green Version]
- Thiam, A.R.; Farese, R.V., Jr.; Walther, T.C. The biophysics and cell biology of lipid droplets. Nat. Rev. Mol. Cell Biol. 2013, 14, 775–786. [Google Scholar] [CrossRef] [Green Version]
- Jacquier, N.; Choudhary, V.; Mari, M.; Toulmay, A.; Reggiori, F.; Schneiter, R. Lipid droplets are functionally connected to the endoplasmic reticulum in Saccharomyces cerevisiae. J. Cell Sci. 2011, 124, 2424–2437. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mishra, S.; Khaddaj, R.; Cottier, S.; Stradalova, V.; Jacob, C.; Schneiter, R. Mature lipid droplets are accessible to ER luminal proteins. J. Cell Sci. 2016, 129, 3803–3815. [Google Scholar] [CrossRef] [Green Version]
- Hariri, H.; Rogers, S.; Ugrankar, R.; Liu, Y.L.; Feathers, J.R.; Henne, W.M. Lipid droplet biogenesis is spatially coordinated at ER-vacuole contacts under nutritional stress. EMBO Rep. 2018, 19, 57–72. [Google Scholar] [CrossRef]
- Henne, W.M.; Hariri, H. Endoplasmic Reticulum-Vacuole Contact Sites “Bloom” With Stress-Induced Lipid Droplets. Contact 2018, 1, 2515256418756112. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Adeyo, O.; Horn, P.J.; Lee, S.; Binns, D.D.; Chandrahas, A.; Chapman, K.D.; Goodman, J.M. The yeast lipin orthologue Pah1p is important for biogenesis of lipid droplets. J. Cell Biol. 2011, 192, 1043–1055. [Google Scholar] [CrossRef] [Green Version]
- Barbosa, A.D.; Sembongi, H.; Su, W.M.; Abreu, S.; Reggiori, F.; Carman, G.M.; Siniossoglou, S. Lipid partitioning at the nuclear envelope controls membrane biogenesis. Mol. Biol. Cell 2015, 26, 3641–3657. [Google Scholar] [CrossRef]
- Santos-Rosa, H.; Leung, J.; Grimsey, N.; Peak-Chew, S.; Siniossoglou, S. The yeast lipin Smp2 couples phospholipid biosynthesis to nuclear membrane growth. EMBO J. 2005, 24, 1931–1941. [Google Scholar] [CrossRef] [Green Version]
- Stone, S.J.; Levin, M.C.; Farese, R.V., Jr. Membrane topology and identification of key functional amino acid residues of murine acyl-CoA:diacylglycerol acyltransferase-2. J. Biol. Chem. 2006, 281, 40273–40282. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Oelkers, P.; Cromley, D.; Padamsee, M.; Billheimer, J.T.; Sturley, S.L. The DGA1 gene determines a second triglyceride synthetic pathway in yeast. J. Biol. Chem. 2002, 277, 8877–8881. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sorger, D.; Daum, G. Synthesis of triacylglycerols by the acyl-coenzyme A:diacyl-glycerol acyltransferase Dga1p in lipid particles of the yeast Saccharomyces cerevisiae. J. Bacteriol. 2002, 184, 519–524. [Google Scholar] [CrossRef] [Green Version]
- Zweytick, D.; Leitner, E.; Kohlwein, S.D.; Yu, C.; Rothblatt, J.; Daum, G. Contribution of Are1p and Are2p to steryl ester synthesis in the yeast Saccharomyces cerevisiae. Eur. J. Biochem. 2000, 267, 1075–1082. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Moir, R.D.; Gross, D.A.; Silver, D.L.; Willis, I.M. SCS3 and YFT2 link transcription of phospholipid biosynthetic genes to ER stress and the UPR. PLoS. Genet. 2012, 8, e1002890. [Google Scholar] [CrossRef] [Green Version]
- Binns, D.; Lee, S.; Hilton, C.L.; Jiang, Q.X.; Goodman, J.M. Seipin is a discrete homooligomer. Biochemistry 2010, 49, 10747–10755. [Google Scholar] [CrossRef] [Green Version]
- Kadereit, B.; Kumar, P.; Wang, W.J.; Miranda, D.; Snapp, E.L.; Severina, N.; Torregroza, I.; Evans, T.; Silver, D.L. Evolutionarily conserved gene family important for fat storage. Proc. Natl. Acad. Sci. USA 2008, 105, 94–99. [Google Scholar] [CrossRef] [Green Version]
- Wang, C.W.; Miao, Y.H.; Chang, Y.S. Control of lipid droplet size in budding yeast requires the collaboration between Fld1 and Ldb16. J. Cell Sci. 2014, 127, 1214–1228. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bailey, A.P.; Koster, G.; Guillermier, C.; Hirst, E.M.; MacRae, J.I.; Lechene, C.P.; Postle, A.D.; Gould, A.P. Antioxidant Role for Lipid Droplets in a Stem Cell Niche of Drosophila. Cell 2015, 163, 340–353. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bensaad, K.; Favaro, E.; Lewis, C.A.; Peck, B.; Lord, S.; Collins, J.M.; Pinnick, K.E.; Wigfield, S.; Buffa, F.M.; Li, J.L.; et al. Fatty acid uptake and lipid storage induced by HIF-1α contribute to cell growth and survival after hypoxia-reoxygenation. Cell Rep. 2014, 9, 349–365. [Google Scholar] [CrossRef] [Green Version]
- Cotte, A.K.; Aires, V.; Fredon, M.; Limagne, E.; Derangère, V.; Thibaudin, M.; Humblin, E.; Scagliarini, A.; de Barros, J.P.; Hillon, P.; et al. Lysophosphatidylcholine acyltransferase 2-mediated lipid droplet production supports colorectal cancer chemoresistance. Nat. Commun. 2018, 9, 322. [Google Scholar] [CrossRef]
- Jarc, E.; Kump, A.; Malavašič, P.; Eichmann, T.O.; Zimmermann, R.; Petan, T. Lipid droplets induced by secreted phospholipase A(2) and unsaturated fatty acids protect breast cancer cells from nutrient and lipotoxic stress. Biochim. Biophys. Acta. Mol. Cell Biol. Lipids 2018, 1863, 247–265. [Google Scholar] [CrossRef]
- Pucer, A.; Brglez, V.; Payré, C.; Pungerčar, J.; Lambeau, G.; Petan, T. Group X secreted phospholipase A(2) induces lipid droplet formation and prolongs breast cancer cell survival. Mol. Cancer. 2013, 12, 111. [Google Scholar] [CrossRef] [Green Version]
- Qiu, B.; Ackerman, D.; Sanchez, D.J.; Li, B.; Ochocki, J.D.; Grazioli, A.; Bobrovnikova-Marjon, E.; Diehl, J.A.; Keith, B.; Simon, M.C. HIF2α-Dependent Lipid Storage Promotes Endoplasmic Reticulum Homeostasis in Clear-Cell Renal Cell Carcinoma. Cancer Discov. 2015, 5, 652–667. [Google Scholar] [CrossRef] [Green Version]
- Rambold, A.S.; Cohen, S.; Lippincott-Schwartz, J. Fatty acid trafficking in starved cells: Regulation by lipid droplet lipolysis, autophagy, and mitochondrial fusion dynamics. Dev. Cell 2015, 32, 678–692. [Google Scholar] [CrossRef] [Green Version]
- Listenberger, L.L.; Han, X.; Lewis, S.E.; Cases, S.; Farese, R.V., Jr.; Ory, D.S.; Schaffer, J.E. Triglyceride accumulation protects against fatty acid-induced lipotoxicity. Proc. Natl. Acad. Sci. USA 2003, 100, 3077–3082. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Nguyen, T.B.; Louie, S.M.; Daniele, J.R.; Tran, Q.; Dillin, A.; Zoncu, R.; Nomura, D.K.; Olzmann, J.A. DGAT1-Dependent Lipid Droplet Biogenesis Protects Mitochondrial Function during Starvation-Induced Autophagy. Dev. Cell 2017, 42, 9–21.e5. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Roa-Mansergas, X.; Fadó, R.; Atari, M.; Mir, J.F.; Muley, H.; Serra, D.; Casals, N. CPT1C promotes human mesenchymal stem cells survival under glucose deprivation through the modulation of autophagy. Sci. Rep. 2018, 8, 6997. [Google Scholar] [CrossRef] [PubMed]
- Singh, R.; Kaushik, S.; Wang, Y.; Xiang, Y.; Novak, I.; Komatsu, M.; Tanaka, K.; Cuervo, A.M.; Czaja, M.J. Autophagy regulates lipid metabolism. Nature 2009, 458, 1131–1135. [Google Scholar] [CrossRef] [Green Version]
- Boren, J.; Brindle, K.M. Apoptosis-induced mitochondrial dysfunction causes cytoplasmic lipid droplet formation. Cell Death Differ. 2012, 19, 1561–1570. [Google Scholar] [CrossRef] [Green Version]
- Lee, S.J.; Zhang, J.; Choi, A.M.; Kim, H.P. Mitochondrial dysfunction induces formation of lipid droplets as a generalized response to stress. Oxid. Med. Cell Longev. 2013, 2013, 327167. [Google Scholar] [CrossRef]
- Li, N.; Lizardo, D.Y.; Atilla-Gokcumen, G.E. Specific Triacylglycerols Accumulate via Increased Lipogenesis During 5-FU-Induced Apoptosis. ACS Chem. Biol. 2016, 11, 2583–2587. [Google Scholar] [CrossRef]
- Chen, E.; Tsai, T.H.; Li, L.; Saha, P.; Chan, L.; Chang, B.H. PLIN2 is a Key Regulator of the Unfolded Protein Response and Endoplasmic Reticulum Stress Resolution in Pancreatic β Cells. Sci. Rep. 2017, 7, 40855. [Google Scholar] [CrossRef]
- Mylonis, I.; Sembongi, H.; Befani, C.; Liakos, P.; Siniossoglou, S.; Simos, G. Hypoxia causes triglyceride accumulation by HIF-1-mediated stimulation of lipin 1 expression. J. Cell Sci. 2012, 125, 3485–3493. [Google Scholar] [CrossRef] [Green Version]
- Schlaepfer, I.R.; Nambiar, D.K.; Ramteke, A.; Kumar, R.; Dhar, D.; Agarwal, C.; Bergman, B.; Graner, M.; Maroni, P.; Singh, R.P.; et al. Hypoxia induces triglycerides accumulation in prostate cancer cells and extracellular vesicles supporting growth and invasiveness following reoxygenation. Oncotarget 2015, 6, 22836–22856. [Google Scholar] [CrossRef]
- Dichlberger, A.; Schlager, S.; Lappalainen, J.; Käkelä, R.; Hattula, K.; Butcher, S.J.; Schneider, W.J.; Kovanen, P.T. Lipid body formation during maturation of human mast cells. J. Lipid Res. 2011, 52, 2198–2208. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gao, G.; Chen, F.J.; Zhou, L.; Su, L.; Xu, D.; Xu, L.; Li, P. Control of lipid droplet fusion and growth by CIDE family proteins. Biochim. Biophys. Acta Mol. Cell Biol. Lipids 2017, 1862, 1197–1204. [Google Scholar] [CrossRef]
- Gong, J.; Sun, Z.; Wu, L.; Xu, W.; Schieber, N.; Xu, D.; Shui, G.; Yang, H.; Parton, R.G.; Li, P. Fsp27 promotes lipid droplet growth by lipid exchange and transfer at lipid droplet contact sites. J. Cell Biol. 2011, 195, 953–963. [Google Scholar] [CrossRef]
- Jambunathan, S.; Yin, J.; Khan, W.; Tamori, Y.; Puri, V. FSP27 promotes lipid droplet clustering and then fusion to regulate triglyceride accumulation. PLoS ONE 2011, 6, e28614. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sun, Z.; Gong, J.; Wu, H.; Xu, W.; Wu, L.; Xu, D.; Gao, J.; Wu, J.W.; Yang, H.; Yang, M.; et al. Perilipin1 promotes unilocular lipid droplet formation through the activation of Fsp27 in adipocytes. Nat. Commun. 2013, 4, 1594. [Google Scholar] [CrossRef] [Green Version]
- Guo, Y.; Walther, T.C.; Rao, M.; Stuurman, N.; Goshima, G.; Terayama, K.; Wong, J.S.; Vale, R.D.; Walter, P.; Farese, R.V. Functional genomic screen reveals genes involved in lipid-droplet formation and utilization. Nature 2008, 453, 657–661. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Krahmer, N.; Guo, Y.; Wilfling, F.; Hilger, M.; Lingrell, S.; Heger, K.; Newman, H.W.; Schmidt-Supprian, M.; Vance, D.E.; Mann, M.; et al. Phosphatidylcholine synthesis for lipid droplet expansion is mediated by localized activation of CTP:phosphocholine cytidylyltransferase. Cell Metab. 2011, 14, 504–515. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Murphy, S.; Martin, S.; Parton, R.G. Quantitative analysis of lipid droplet fusion: Inefficient steady state fusion but rapid stimulation by chemical fusogens. PLoS ONE 2010, 5, e15030. [Google Scholar] [CrossRef] [Green Version]
- Marcinkiewicz, A.; Gauthier, D.; Garcia, A.; Brasaemle, D.L. The phosphorylation of serine 492 of perilipin a directs lipid droplet fragmentation and dispersion. J. Biol. Chem. 2006, 281, 11901–11909. [Google Scholar] [CrossRef] [Green Version]
- Long, A.P.; Manneschmidt, A.K.; VerBrugge, B.; Dortch, M.R.; Minkin, S.C.; Prater, K.E.; Biggerstaff, J.P.; Dunlap, J.R.; Dalhaimer, P. Lipid droplet de novo formation and fission are linked to the cell cycle in fission yeast. Traffic 2012, 13, 705–714. [Google Scholar] [CrossRef]
- Hashimoto, T.; Segawa, H.; Okuno, M.; Kano, H.; Hamaguchi, H.O.; Haraguchi, T.; Hiraoka, Y.; Hasui, S.; Yamaguchi, T.; Hirose, F.; et al. Active involvement of micro-lipid droplets and lipid-droplet-associated proteins in hormone-stimulated lipolysis in adipocytes. J. Cell Sci. 2012, 125, 6127–6136. [Google Scholar] [CrossRef] [Green Version]
- Paar, M.; Jüngst, C.; Steiner, N.A.; Magnes, C.; Sinner, F.; Kolb, D.; Lass, A.; Zimmermann, R.; Zumbusch, A.; Kohlwein, S.D.; et al. Remodeling of lipid droplets during lipolysis and growth in adipocytes. J. Biol. Chem. 2012, 287, 11164–11173. [Google Scholar] [CrossRef] [Green Version]
- Chitraju, C.; Mejhert, N.; Haas, J.T.; Diaz-Ramirez, L.G.; Grueter, C.A.; Imbriglio, J.E.; Pinto, S.; Koliwad, S.K.; Walther, T.C.; Farese, R.V., Jr. Triglyceride Synthesis by DGAT1 Protects Adipocytes from Lipid-Induced ER Stress during Lipolysis. Cell Metab. 2017, 26, 407–418.e3. [Google Scholar] [CrossRef] [PubMed]
- Szymanski, K.M.; Binns, D.; Bartz, R.; Grishin, N.V.; Li, W.P.; Agarwal, A.K.; Garg, A.; Anderson, R.G.; Goodman, J.M. The lipodystrophy protein seipin is found at endoplasmic reticulum lipid droplet junctions and is important for droplet morphology. Proc. Natl. Acad Sci. USA 2007, 104, 20890–20895. [Google Scholar] [CrossRef] [Green Version]
- Fei, W.; Shui, G.; Gaeta, B.; Du, X.; Kuerschner, L.; Li, P.; Brown, A.J.; Wenk, M.R.; Parton, R.G.; Yang, H. Fld1p, a functional homologue of human seipin, regulates the size of lipid droplets in yeast. J. Cell Biol. 2008, 180, 473–482. [Google Scholar] [CrossRef] [PubMed]
- Bouchez, I.; Pouteaux, M.; Canonge, M.; Genet, M.; Chardot, T.; Guillot, A.; Froissard, M. Regulation of lipid droplet dynamics in Saccharomyces cerevisiae depends on the Rab7-like Ypt7p, HOPS complex and V1-ATPase. Biol. Open. 2015, 4, 764–775. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kumar, R.; Rahman, M.A.; Nazarko, T.Y. Nitrogen Starvation and Stationary Phase Lipophagy Have Distinct Molecular Mechanisms. Int. J. Mol. Sci. 2020, 21, 9094. [Google Scholar] [CrossRef]
- Barbosa, A.D.; Siniossoglou, S. Spatial distribution of lipid droplets during starvation: Implications for lipophagy. Commun. Integr. Biol. 2016, 9, e1183854. [Google Scholar] [CrossRef]
- Schrader, M. Tubulo-reticular clusters of peroxisomes in living COS-7 cells: Dynamic behavior and association with lipid droplets. J. Histochem. Cytochem. 2001, 49, 1421–1429. [Google Scholar] [CrossRef] [Green Version]
- Binns, D.; Januszewski, T.; Chen, Y.; Hill, J.; Markin, V.S.; Zhao, Y.; Gilpin, C.; Chapman, K.D.; Anderson, R.G.; Goodman, J.M. An intimate collaboration between peroxisomes and lipid bodies. J. Cell Biol. 2006, 173, 719–731. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Valm, A.M.; Cohen, S.; Legant, W.R.; Melunis, J.; Hershberg, U.; Wait, E.; Cohen, A.R.; Davidson, M.W.; Betzig, E.; Lippincott-Schwartz, J. Applying systems-level spectral imaging and analysis to reveal the organelle interactome. Nature 2017, 546, 162–167. [Google Scholar] [CrossRef] [PubMed]
- Kohlwein, S.D.; Veenhuis, M.; van der Klei, I.J. Lipid droplets and peroxisomes: Key players in cellular lipid homeostasis or a matter of fat--store ‘em up or burn ‘em down. Genetics 2013, 193, 1–50. [Google Scholar] [CrossRef] [Green Version]
- Chang, C.L.; Weigel, A.V.; Ioannou, M.S.; Pasolli, H.A.; Xu, C.S.; Peale, D.R.; Shtengel, G.; Freeman, M.; Hess, H.F.; Blackstone, C.; et al. Spastin tethers lipid droplets to peroxisomes and directs fatty acid trafficking through ESCRT-III. J. Cell Biol. 2019, 218, 2583–2599. [Google Scholar] [CrossRef] [PubMed]
- Kong, J.; Ji, Y.; Jeon, Y.G.; Han, J.S.; Han, K.H.; Lee, J.H.; Lee, G.; Jang, H.; Choe, S.S.; Baes, M.; et al. Spatiotemporal contact between peroxisomes and lipid droplets regulates fasting-induced lipolysis via PEX5. Nat. Commun. 2020, 11, 578. [Google Scholar] [CrossRef] [PubMed]
- Joshi, A.S.; Cohen, S. Lipid Droplet and Peroxisome Biogenesis: Do They Go Hand-in-Hand? Front. Cell Dev. Biol. 2019, 7, 92. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Joshi, A.S.; Huang, X.; Choudhary, V.; Levine, T.P.; Hu, J.; Prinz, W.A. A family of membrane-shaping proteins at ER subdomains regulates pre-peroxisomal vesicle biogenesis. J. Cell Biol. 2016, 215, 515–529. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Joshi, A.S.; Nebenfuehr, B.; Choudhary, V.; Satpute-Krishnan, P.; Levine, T.P.; Golden, A.; Prinz, W.A. Lipid droplet and peroxisome biogenesis occur at the same ER subdomains. Nat. Commun. 2018, 9, 2940. [Google Scholar] [CrossRef] [Green Version]
- Pu, J.; Ha, C.W.; Zhang, S.; Jung, J.P.; Huh, W.K.; Liu, P. Interactomic study on interaction between lipid droplets and mitochondria. Protein Cell 2011, 2, 487–496. [Google Scholar] [CrossRef] [Green Version]
- Zechner, R.; Madeo, F.; Kratky, D. Cytosolic lipolysis and lipophagy: Two sides of the same coin. Nat. Rev. Mol. Cell Biol. 2017, 18, 671–684. [Google Scholar] [CrossRef] [PubMed]
- Kaushik, S.; Cuervo, A.M. Degradation of lipid droplet-associated proteins by chaperone-mediated autophagy facilitates lipolysis. Nat. Cell Biol. 2015, 17, 759–770. [Google Scholar] [CrossRef] [Green Version]
- Kaushik, S.; Cuervo, A.M. AMPK-dependent phosphorylation of lipid droplet protein PLIN2 triggers its degradation by CMA. Autophagy 2016, 12, 432–438. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shpilka, T.; Welter, E.; Borovsky, N.; Amar, N.; Mari, M.; Reggiori, F.; Elazar, Z. Lipid droplets and their component triglycerides and steryl esters regulate autophagosome biogenesis. EMBO J. 2015, 34, 2117–2131. [Google Scholar] [CrossRef] [Green Version]
- Velázquez, A.P.; Tatsuta, T.; Ghillebert, R.; Drescher, I.; Graef, M. Lipid droplet-mediated ER homeostasis regulates autophagy and cell survival during starvation. J. Cell Biol. 2016, 212, 621–631. [Google Scholar] [CrossRef]
- Maeda, Y.; Oku, M.; Sakai, Y. A defect of the vacuolar putative lipase Atg15 accelerates degradation of lipid droplets through lipolysis. Autophagy 2015, 11, 1247–1258. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Nakatogawa, H.; Suzuki, K.; Kamada, Y.; Ohsumi, Y. Dynamics and diversity in autophagy mechanisms: Lessons from yeast. Nat. Rev. Mol. Cell Biol. 2009, 10, 458–467. [Google Scholar] [CrossRef] [Green Version]
- Reggiori, F.; Klionsky, D.J. Autophagic processes in yeast: Mechanism, machinery and regulation. Genetics 2013, 194, 341–361. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hamasaki, M.; Furuta, N.; Matsuda, A.; Nezu, A.; Yamamoto, A.; Fujita, N.; Oomori, H.; Noda, T.; Haraguchi, T.; Hiraoka, Y.; et al. Autophagosomes form at ER-mitochondria contact sites. Nature 2013, 495, 389–393. [Google Scholar] [CrossRef]
- Ravikumar, B.; Moreau, K.; Jahreiss, L.; Puri, C.; Rubinsztein, D.C. Plasma membrane contributes to the formation of pre-autophagosomal structures. Nat. Cell Biol. 2010, 12, 747–757. [Google Scholar] [CrossRef]
- Kunz, J.B.; Schwarz, H.; Mayer, A. Determination of four sequential stages during microautophagy in vitro. J. Biol. Chem. 2004, 279, 9987–9996. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Müller, O.; Sattler, T.; Flötenmeyer, M.; Schwarz, H.; Plattner, H.; Mayer, A. Autophagic tubes: Vacuolar invaginations involved in lateral membrane sorting and inverse vesicle budding. J. Cell Biol. 2000, 151, 519–528. [Google Scholar] [CrossRef] [PubMed]
- Sattler, T.; Mayer, A. Cell-free reconstitution of microautophagic vacuole invagination and vesicle formation. J. Cell Biol. 2000, 151, 529–538. [Google Scholar] [CrossRef]
- Kissová, I.; Salin, B.; Schaeffer, J.; Bhatia, S.; Manon, S.; Camougrand, N. Selective and non-selective autophagic degradation of mitochondria in yeast. Autophagy 2007, 3, 329–336. [Google Scholar] [CrossRef] [Green Version]
- Roberts, P.; Moshitch-Moshkovitz, S.; Kvam, E.; O’Toole, E.; Winey, M.; Goldfarb, D.S. Piecemeal microautophagy of nucleus in Saccharomyces cerevisiae. Mol. Biol. Cell 2003, 14, 129–141. [Google Scholar] [CrossRef] [Green Version]
- Sakai, Y.; Koller, A.; Rangell, L.K.; Keller, G.A.; Subramani, S. Peroxisome degradation by microautophagy in Pichia pastoris: Identification of specific steps and morphological intermediates. J. Cell Biol. 1998, 141, 625–636. [Google Scholar] [CrossRef] [Green Version]
- Schuck, S.; Gallagher, C.M.; Walter, P. ER-phagy mediates selective degradation of endoplasmic reticulum independently of the core autophagy machinery. J. Cell Sci. 2014, 127, 4078–4088. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Seo, A.Y.; Lau, P.W.; Feliciano, D.; Sengupta, P.; Gros, M.A.L.; Cinquin, B.; Larabell, C.A.; Lippincott-Schwartz, J. AMPK and vacuole-associated Atg14p orchestrate μ-lipophagy for energy production and long-term survival under glucose starvation. Elife 2017, 6, e21690. [Google Scholar] [CrossRef] [PubMed]
- Zhang, A.; Meng, Y.; Li, Q.; Liang, Y. The endosomal sorting complex required for transport complex negatively regulates Erg6 degradation under specific glucose restriction conditions. Traffic 2020, 21, 488–502. [Google Scholar] [CrossRef]
- Oku, M.; Maeda, Y.; Kagohashi, Y.; Kondo, T.; Yamada, M.; Fujimoto, T.; Sakai, Y. Evidence for ESCRT- and clathrin-dependent microautophagy. J. Cell Biol. 2017, 216, 3263–3274. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kurokawa, Y.; Konishi, R.; Yoshida, A.; Tomioku, K.; Tanabe, K.; Fujita, A. Microautophagy in the yeast vacuole depends on the activities of phosphatidylinositol 4-kinases, Stt4p and Pik1p. Biochim. Biophys. Acta. Biomembr. 2020, 1862, 183416. [Google Scholar] [CrossRef]
- Vevea, J.D.; Garcia, E.J.; Chan, R.B.; Zhou, B.; Schultz, M.; Di Paolo, G.; McCaffery, J.M.; Pon, L.A. Role for Lipid Droplet Biogenesis and Microlipophagy in Adaptation to Lipid Imbalance in Yeast. Dev. Cell 2015, 35, 584–599. [Google Scholar] [CrossRef] [Green Version]
- Garcia, E.J.; Liao, P.C.; Tan, G.; Vevea, J.D.; Sing, C.N.; Tsang, C.A.; McCaffery, J.M.; Boldogh, I.R.; Pon, L.A. Membrane dynamics and protein targets of lipid droplet microautophagy during ER stress-induced proteostasis in the budding yeast, Saccharomyces cerevisiae. Autophagy 2020, 1–21. [Google Scholar] [CrossRef] [PubMed]
- Simons, K.; Toomre, D. Lipid rafts and signal transduction. Nat. Rev. Mol. Cell Biol. 2000, 1, 31–39. [Google Scholar] [CrossRef] [PubMed]
- Edidin, M. The state of lipid rafts: From model membranes to cells. Annu. Rev. Biophys. Biomol. Struct. 2003, 32, 257–283. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Levental, I.; Veatch, S. The Continuing Mystery of Lipid Rafts. J. Mol. Biol. 2016, 428, 4749–4764. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Goni, F.M. “Rafts”: A nickname for putative transient nanodomains. Chem. Phys. Lipids 2019, 218, 34–39. [Google Scholar] [CrossRef]
- Toulmay, A.; Prinz, W.A. Direct imaging reveals stable, micrometer-scale lipid domains that segregate proteins in live cells. J. Cell Biol. 2013, 202, 35–44. [Google Scholar] [CrossRef] [PubMed]
- Rayermann, S.P.; Rayermann, G.E.; Cornell, C.E.; Merz, A.J.; Keller, S.L. Hallmarks of Reversible Separation of Living, Unperturbed Cell Membranes into Two Liquid Phases. Biophys. J. 2017, 113, 2425–2432. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Murley, A.; Sarsam, R.D.; Toulmay, A.; Yamada, J.; Prinz, W.A.; Nunnari, J. Ltc1 is an ER-localized sterol transporter and a component of ER-mitochondria and ER-vacuole contacts. J. Cell Biol. 2015, 209, 539–548. [Google Scholar] [CrossRef] [Green Version]
- Murley, A.; Yamada, J.; Niles, B.J.; Toulmay, A.; Prinz, W.A.; Powers, T.; Nunnari, J. Sterol transporters at membrane contact sites regulate TORC1 and TORC2 signaling. J. Cell. Biol. 2017, 216, 2679–2689. [Google Scholar] [CrossRef] [Green Version]
- Vaskovicova, K.; Vesela, P.; Zahumensky, J.; Folkova, D.; Balazova, M.; Malinsky, J. Plasma Membrane Protein Nce102 Modulates Morphology and Function of the Yeast Vacuole. Biomolecules 2020, 10, 1476. [Google Scholar] [CrossRef]
- Elbaz-Alon, Y.; Eisenberg-Bord, M.; Shinder, V.; Stiller, S.B.; Shimoni, E.; Wiedemann, N.; Geiger, T.; Schuldiner, M. Lam6 Regulates the Extent of Contacts between Organelles. Cell Rep. 2015, 12, 7–14. [Google Scholar] [CrossRef] [Green Version]
- Tomioku, K.N.; Shigekuni, M.; Hayashi, H.; Yoshida, A.; Futagami, T.; Tamaki, H.; Tanabe, K.; Fujita, A. Nanoscale domain formation of phosphatidylinositol 4-phosphate in the plasma and vacuolar membranes of living yeast cells. Eur. J. Cell Biol. 2018, 97, 269–278. [Google Scholar] [CrossRef] [PubMed]
- Tsuji, T.; Fujimoto, M.; Tatematsu, T.; Cheng, J.; Orii, M.; Takatori, S.; Fujimoto, T. Niemann-Pick type C proteins promote microautophagy by expanding raft-like membrane domains in the yeast vacuole. Elife 2017, 6, e25960. [Google Scholar] [CrossRef]
- Teixeira, V.; Johnsen, L.; Martínez-Montañés, F.; Grippa, A.; Buxó, L.; Idrissi, F.Z.; Ejsing, C.S.; Carvalho, P. Regulation of lipid droplets by metabolically controlled Ldo isoforms. J. Cell Biol. 2018, 217, 127–138. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Krick, R.; Muehe, Y.; Prick, T.; Bremer, S.; Schlotterhose, P.; Eskelinen, E.L.; Millen, J.; Goldfarb, D.S.; Thumm, M. Piecemeal microautophagy of the nucleus requires the core macroautophagy genes. Mol. Biol. Cell 2008, 19, 4492–4505. [Google Scholar] [CrossRef] [Green Version]
- Farre, J.C.; Subramani, S. Peroxisome turnover by micropexophagy: An autophagy-related process. Trends Cell Biol. 2004, 14, 515–523. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Barth, H.; Meiling-Wesse, K.; Epple, U.D.; Thumm, M. Mai1p is essential for maturation of proaminopeptidase I but not for autophagy. FEBS Lett 2002, 512, 173–179. [Google Scholar] [CrossRef] [Green Version]
- Tamura, N.; Oku, M.; Sakai, Y. Atg21 regulates pexophagy via its PI(3)P-binding activity in Pichia pastoris. FEMS Yeast Res. 2014, 14, 435–444. [Google Scholar] [CrossRef] [Green Version]
- Kanki, T.; Wang, K.; Baba, M.; Bartholomew, C.R.; Lynch-Day, M.A.; Du, Z.; Geng, J.; Mao, K.; Yang, Z.; Yen, W.L.; et al. A genomic screen for yeast mutants defective in selective mitochondria autophagy. Mol. Biol. Cell 2009, 20, 4730–4738. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pan, X.; Roberts, P.; Chen, Y.; Kvam, E.; Shulga, N.; Huang, K.; Lemmon, S.; Goldfarb, D.S. Nucleus-vacuole junctions in Saccharomyces cerevisiae are formed through the direct interaction of Vac8p with Nvj1p. Mol. Biol. Cell 2000, 11, 2445–2457. [Google Scholar] [CrossRef] [Green Version]
- Gross, D.A.; Silver, D.L. Cytosolic lipid droplets: From mechanisms of fat storage to disease. Crit. Rev. Biochem. Mol. Biol. 2014, 49, 304–326. [Google Scholar] [CrossRef]
- Reue, K. A thematic review series: Lipid droplet storage and metabolism: From yeast to man. J. Lipid Res. 2011, 52, 1865–1868. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Onal, G.; Kutlu, O.; Gozuacik, D.; Dokmeci Emre, S. Lipid Droplets in Health and Disease. Lipids Health Dis. 2017, 16, 128. [Google Scholar] [CrossRef] [PubMed] [Green Version]
| LD-Related Function | Proteins enriched in LDs and/or Co-Localized with LDs |
|---|---|
| LD biogenesis | Ldb16, Ldo16, Ldo45, Mdm1, Pln1, Sei1 |
| TAG synthesis | Ayr1, Dga1, Pah1, Pgc1, Slc1 |
| Ergosterol metabolism | Erg1, Erg6, Erg7, Erg27, Say1, Srt1 |
| Lipolysis-related | Ice2, Ldh1, Lpl1, Tgl1, Tgl3, Tgl4, Tgl5, Yeh1, Yju3, Ypr147c |
| FA/phospholipid-related | Eht1, Faa1, Faa4, Fat1, Hfd1, Loa1, Pdr16 |
| Other lipid-related functions | Atf1, Cab5, Nus1, Rer2, Tsc10, Ubx2 |
| LD-related role unknown | Anr2, Yim1, Lds1, Lds2, Rrt8 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Rahman, M.A.; Kumar, R.; Sanchez, E.; Nazarko, T.Y. Lipid Droplets and Their Autophagic Turnover via the Raft-Like Vacuolar Microdomains. Int. J. Mol. Sci. 2021, 22, 8144. https://doi.org/10.3390/ijms22158144
Rahman MA, Kumar R, Sanchez E, Nazarko TY. Lipid Droplets and Their Autophagic Turnover via the Raft-Like Vacuolar Microdomains. International Journal of Molecular Sciences. 2021; 22(15):8144. https://doi.org/10.3390/ijms22158144
Chicago/Turabian StyleRahman, Muhammad Arifur, Ravinder Kumar, Enrique Sanchez, and Taras Y. Nazarko. 2021. "Lipid Droplets and Their Autophagic Turnover via the Raft-Like Vacuolar Microdomains" International Journal of Molecular Sciences 22, no. 15: 8144. https://doi.org/10.3390/ijms22158144
APA StyleRahman, M. A., Kumar, R., Sanchez, E., & Nazarko, T. Y. (2021). Lipid Droplets and Their Autophagic Turnover via the Raft-Like Vacuolar Microdomains. International Journal of Molecular Sciences, 22(15), 8144. https://doi.org/10.3390/ijms22158144







