Genome-Wide Identification and Expression Analysis of Tomato ADK Gene Family during Development and Stress
Abstract
1. Introduction
2. Results
2.1. Identification of the ADK Family in Tomato
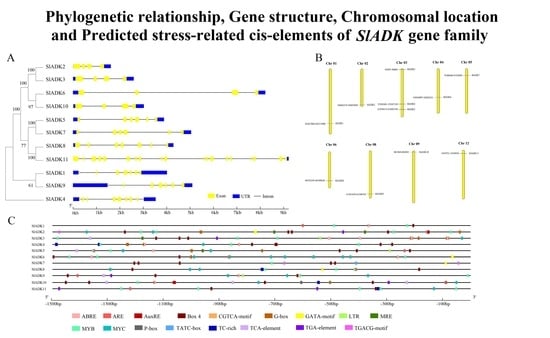
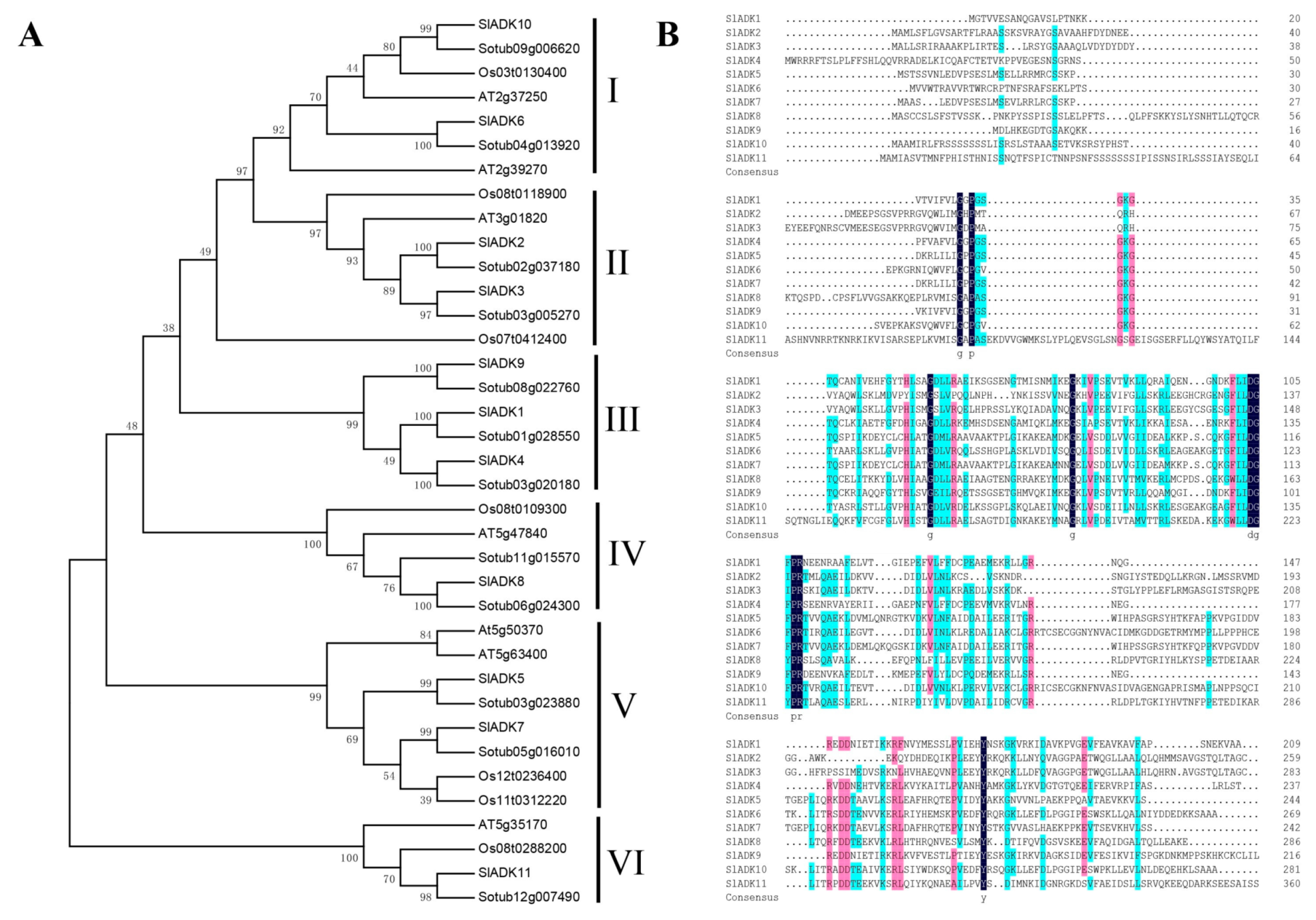
2.2. Phylogenetic Analysis and Multiple Sequence Alignment of SlADK Genes
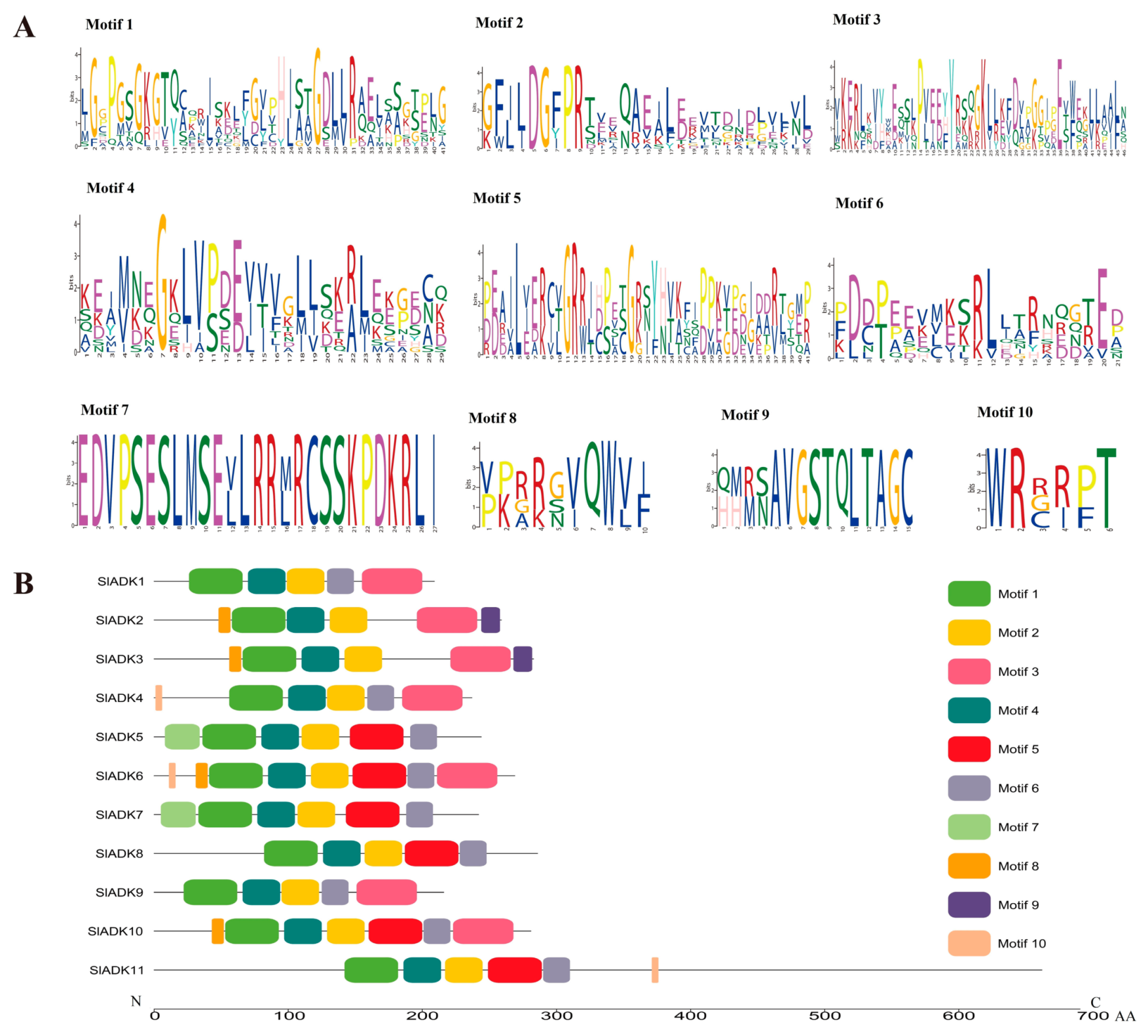
2.3. Motif Analysis
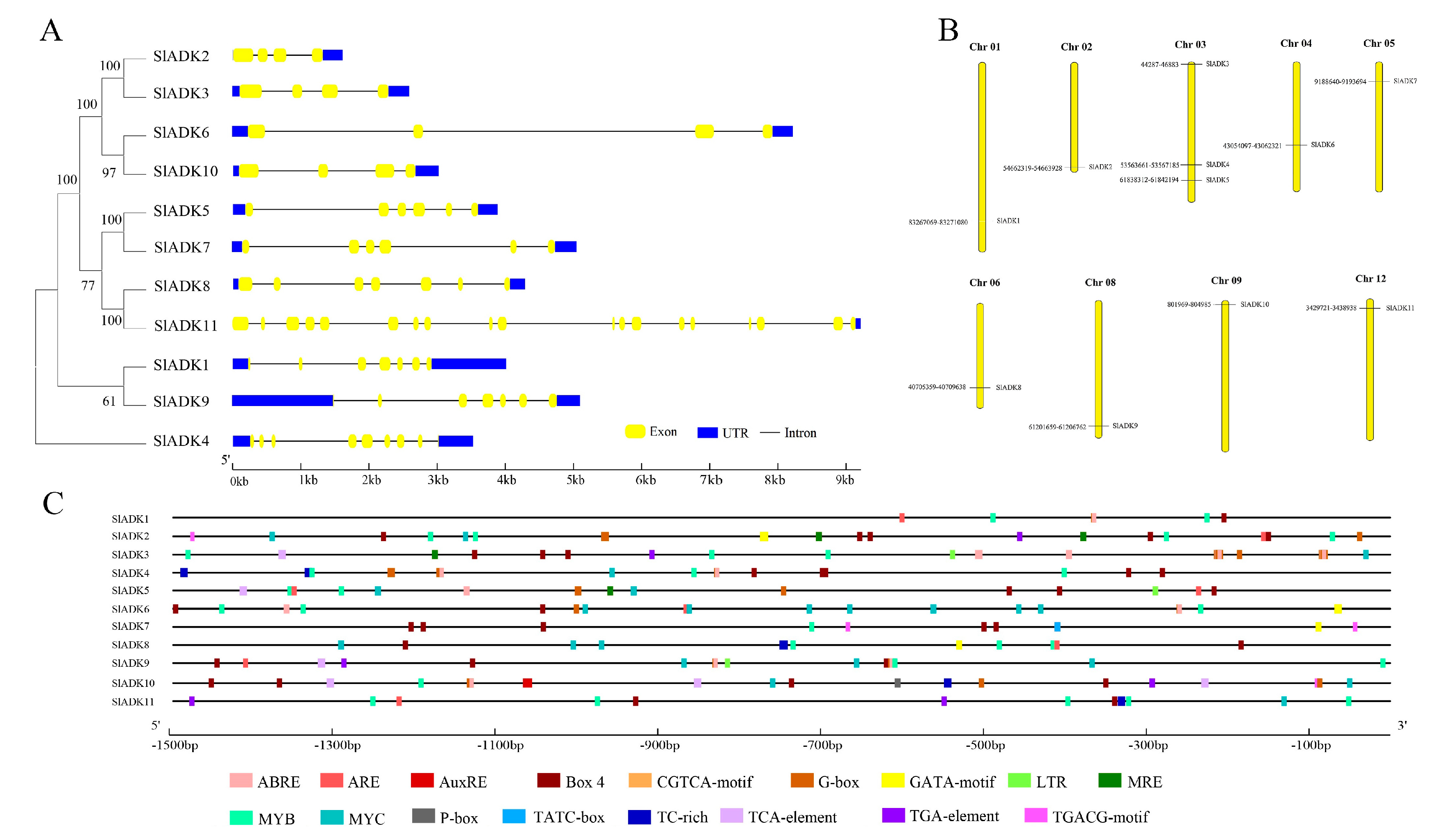
2.4. Gene Structure and Chromosomal Location Analysis of SlADKs
2.5. Cis-Regulatory Elements in SlADK Promoters
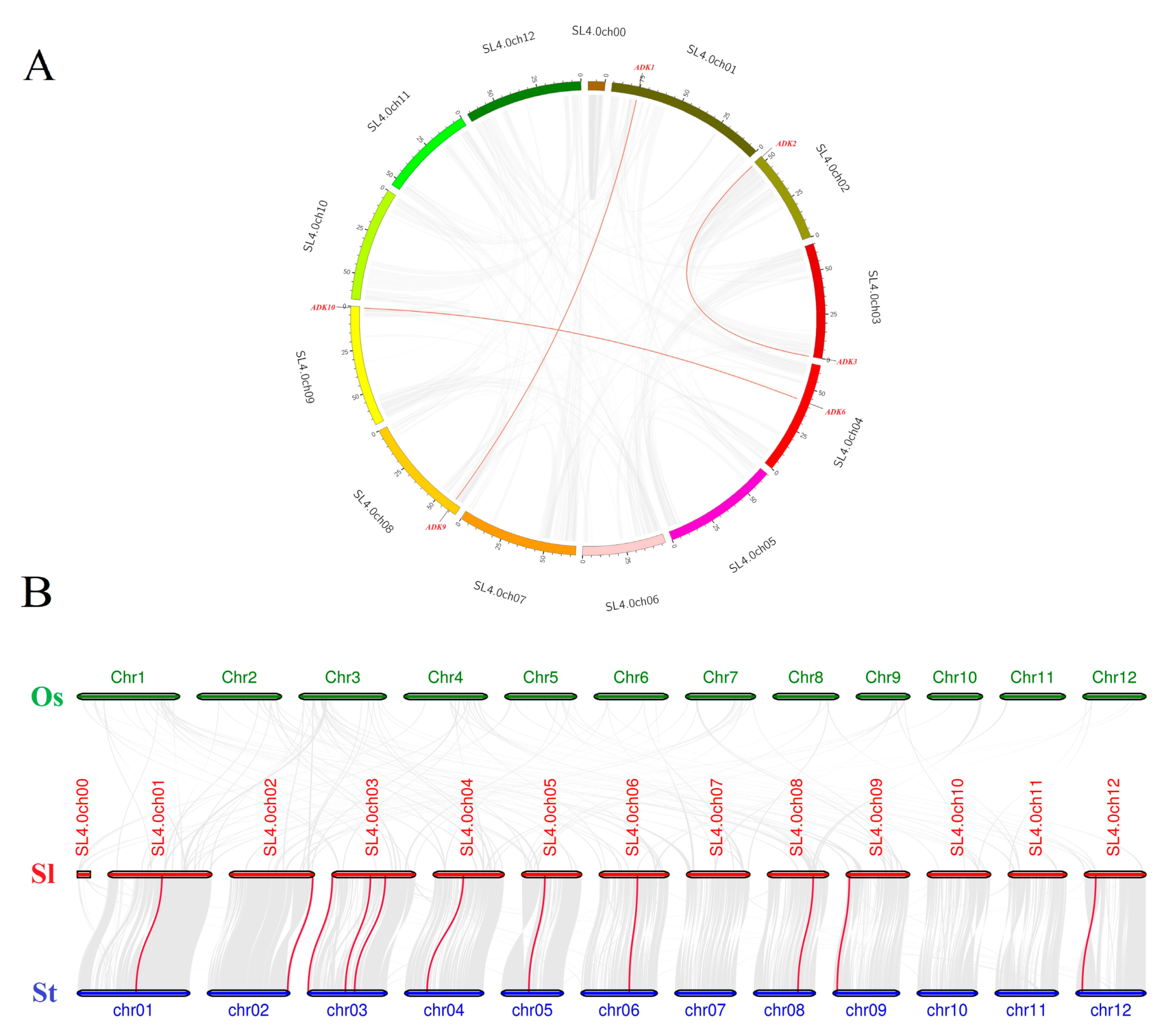
2.6. Synteny Analysis of SlADK Genes
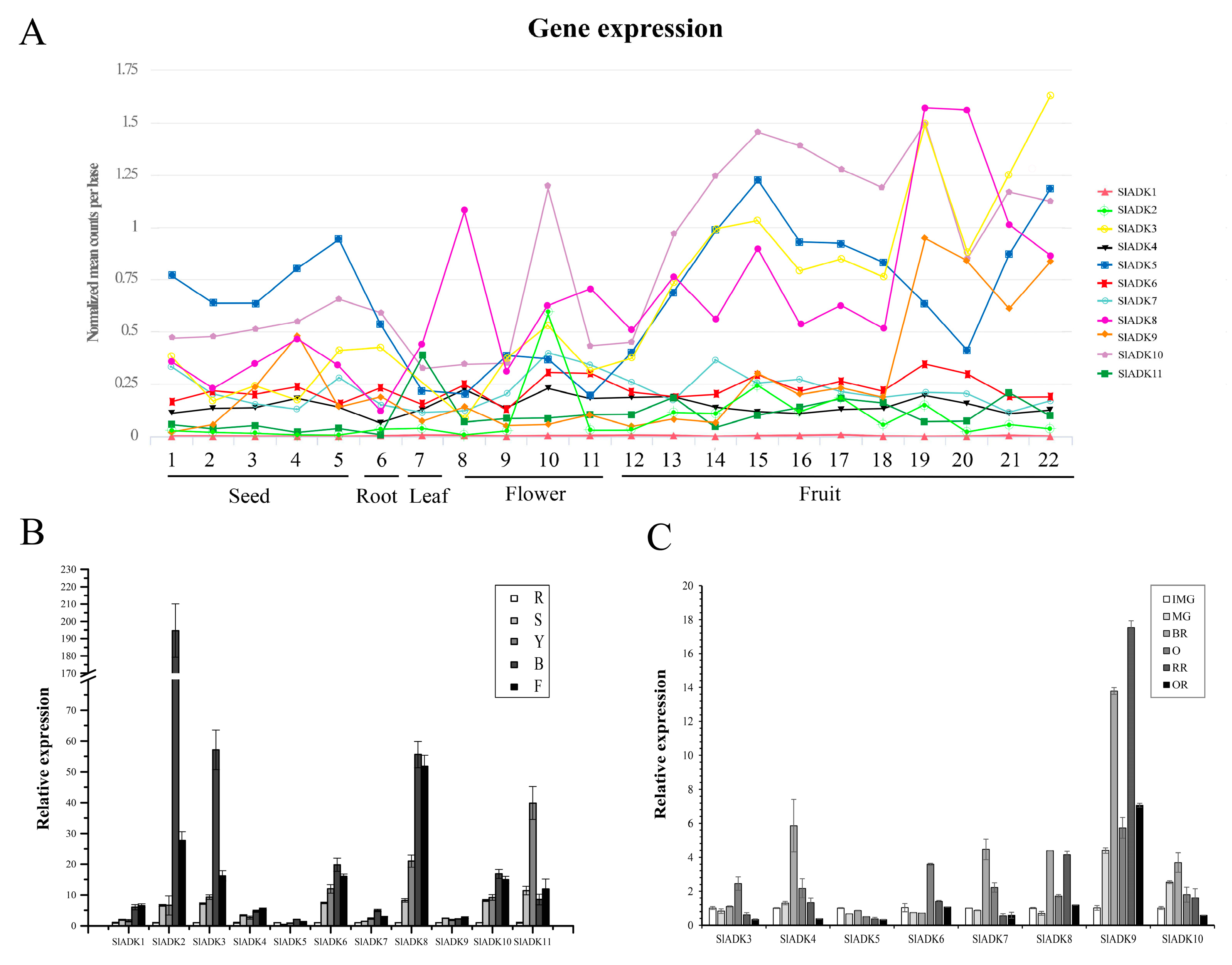
2.7. Expression Pattern Analysis of SlADKs
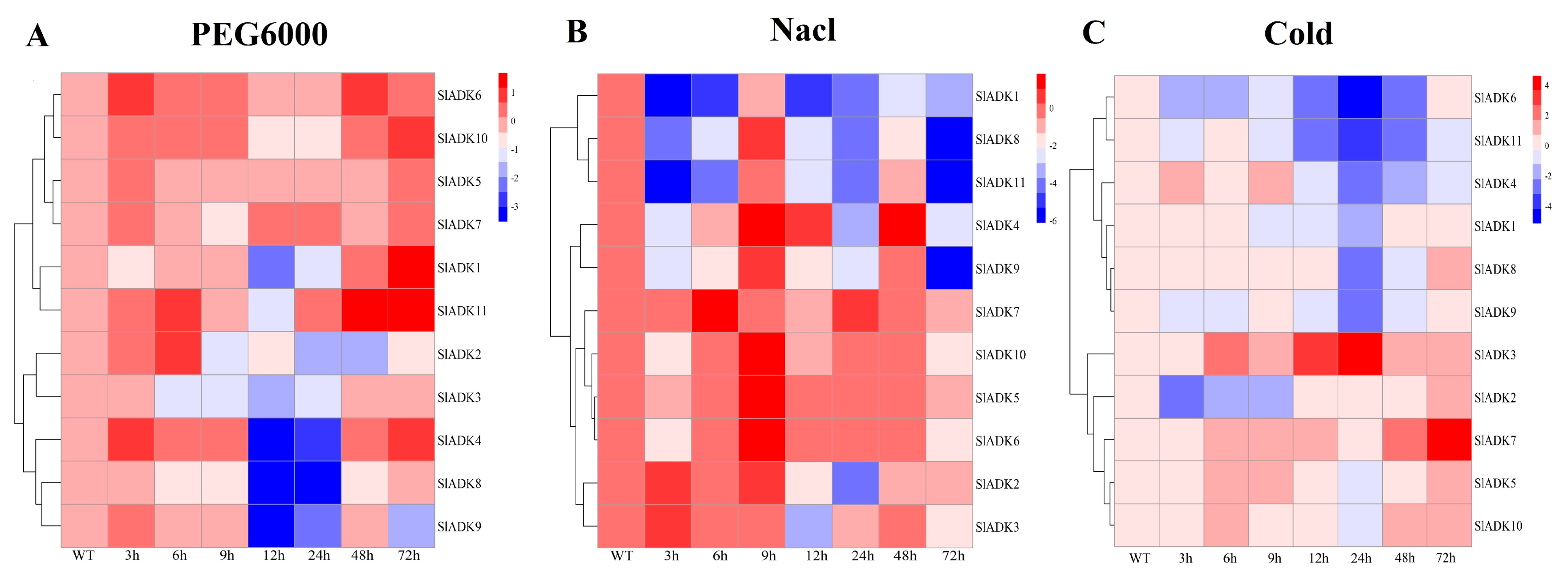
2.8. Expression Characteristics of SlADKs under Diverse Abiotic Stresses
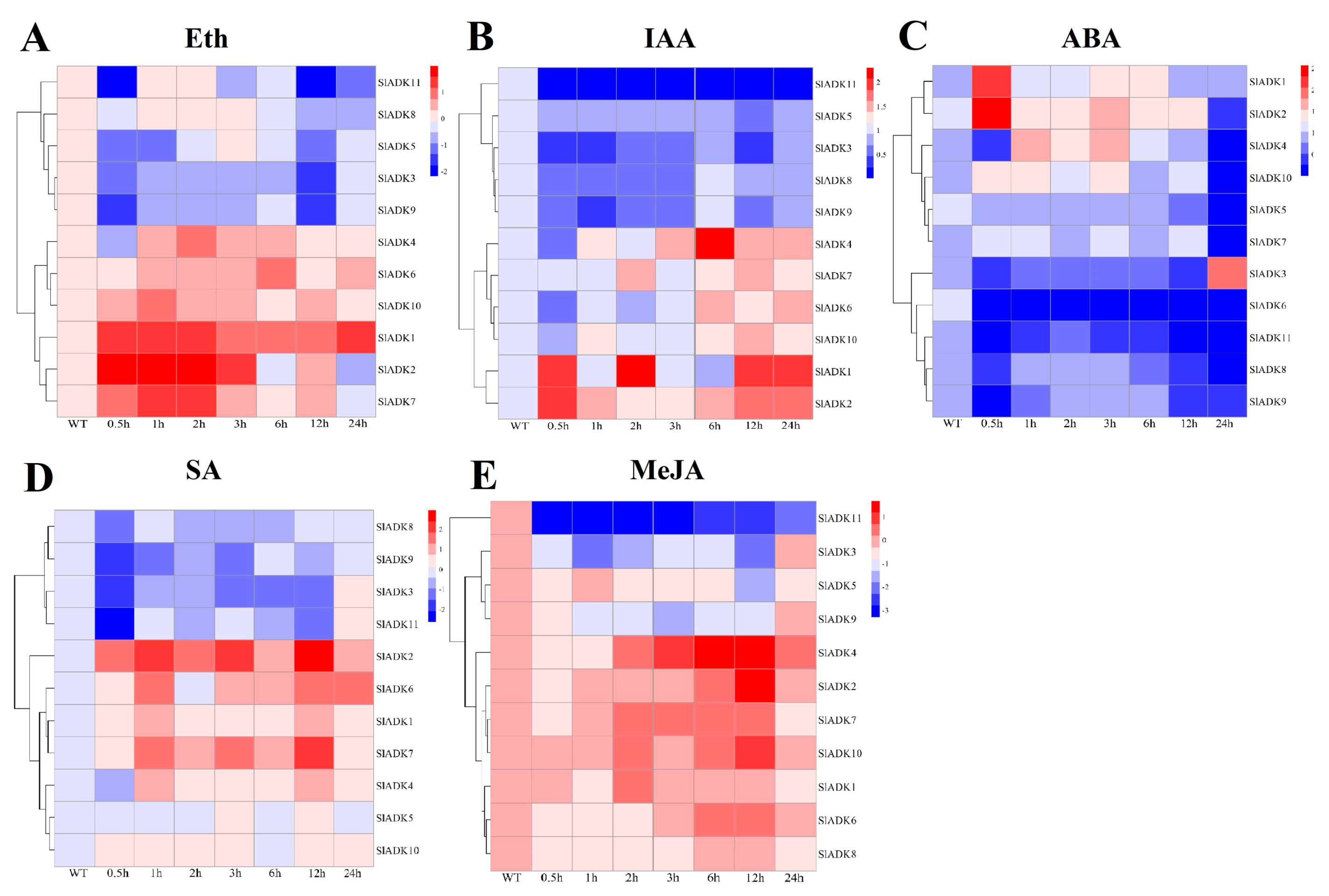
2.9. Expression Profiles of SlADKs in Response to Diverse Hormone Treatments
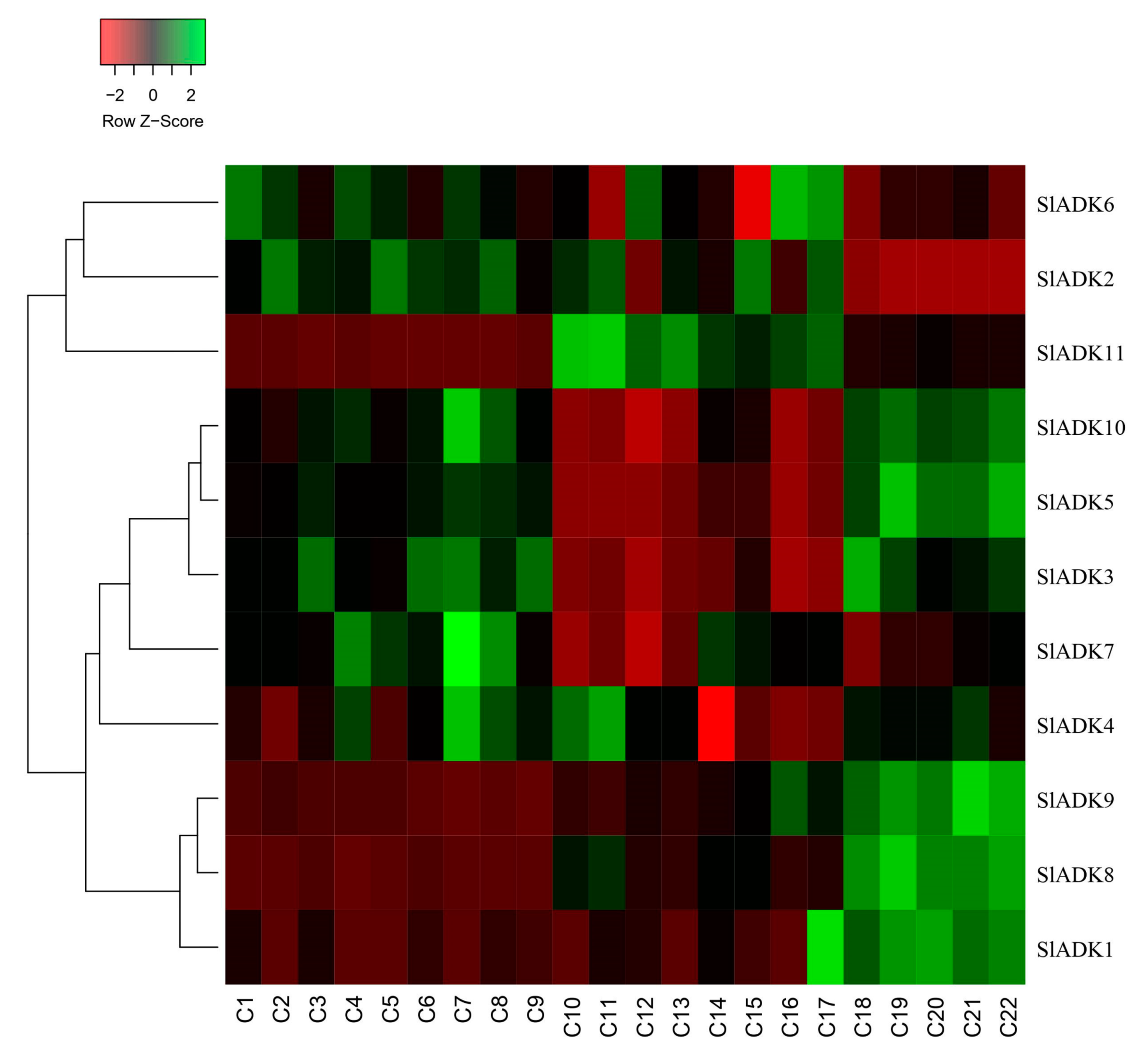
2.10. SlADK Expression Patterns under Stress Based on RNA-Seq Data
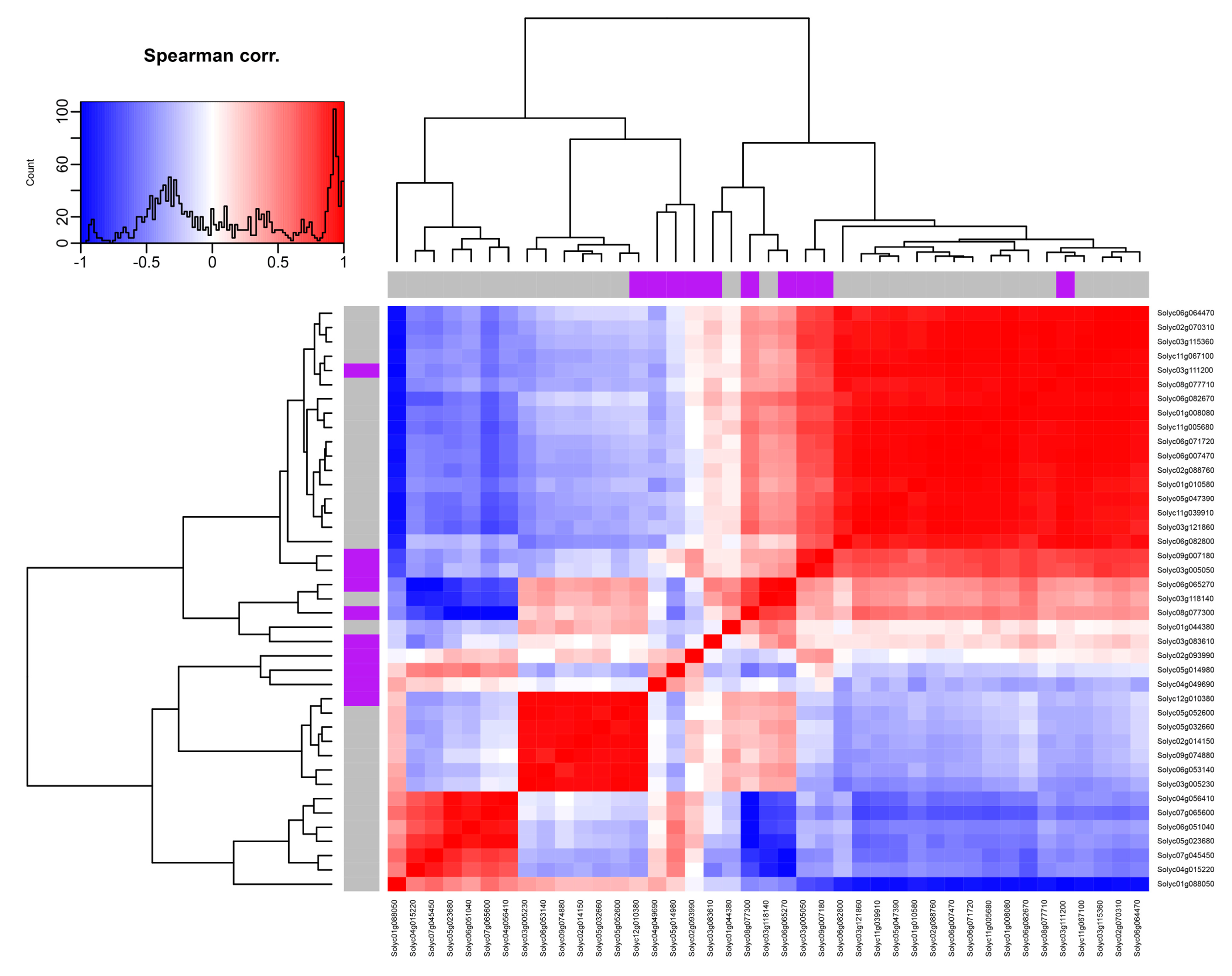
2.11. Coexpression and Correlation Network Analysis
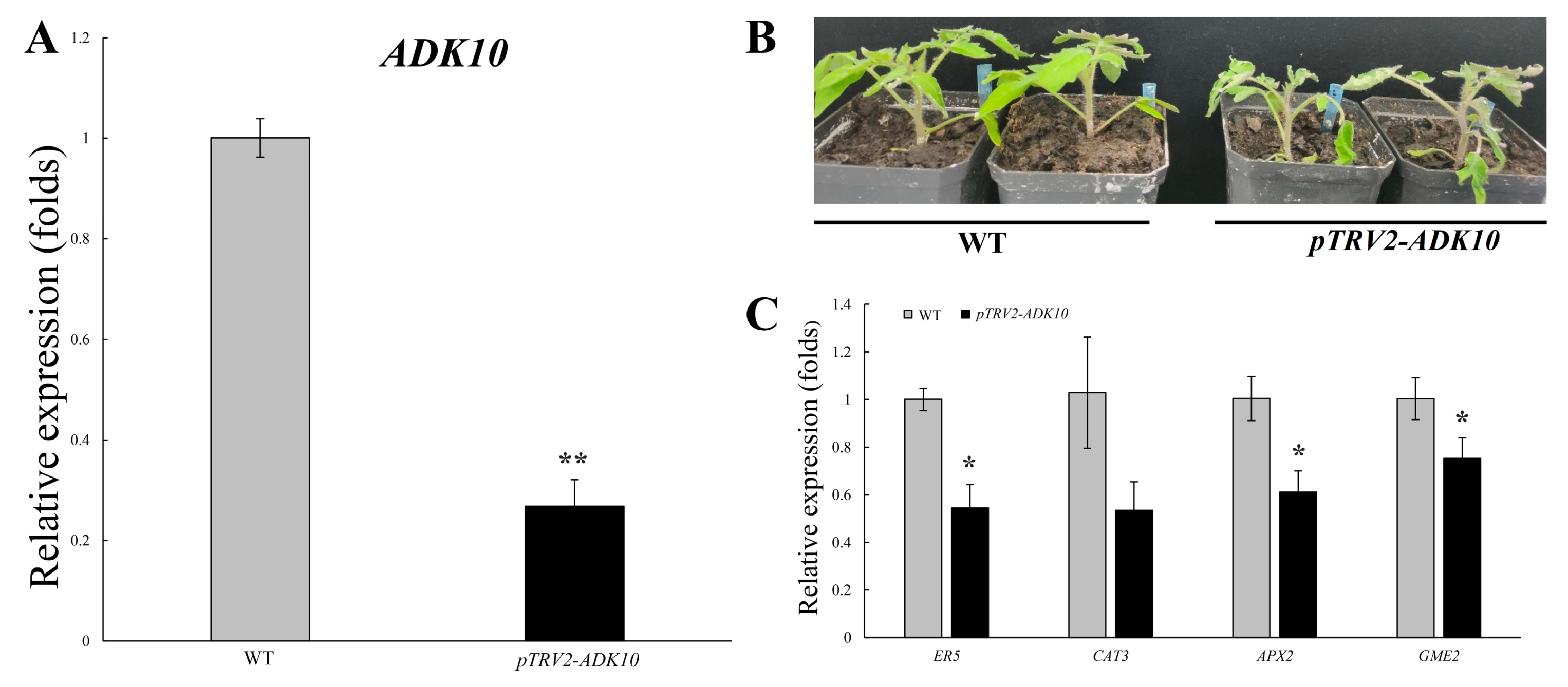
2.12. Functional Analysis of SlADK10 Silencing in Tomato under Drought Stress
3. Discussion
4. Methods
4.1. Plant Materials and Growth Conditions
4.2. Identification of Tomato ADK Genes
4.3. Characteristics, Phylogenetic Relationships, and Sequence Analysis of ADK Proteins
4.4. Gene Structure, Chromosome Location, and CREs of SlADK Genes
4.5. Synteny Analysis of SlADK Genes
4.6. Expression Data Mining of Tomato SlADK Genes
4.7. Hormone and Abiotic Stress Treatments
4.8. RNA Isolation and Quantitative Real-Time PCR Analysis
4.9. Virus-Induced Gene Silencing
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Dzheia, P.P.; Kal’venas, A.A.; Toleĭkis, A.I.; Prashkiavichius, A.K. The role of adenylate kinase in the regulation of the rate and effectiveness of energy transfer from mitochondria to hexokinase in vitro. Biokhimiia 1986, 51, 974–979. [Google Scholar] [PubMed]
- Atkinson, D.E. The energy charge of the adenylate pool as a regulatory parameter. Interaction with feedback modifiers. Biochemistry 1968, 7, 4030–4034. [Google Scholar] [CrossRef]
- Ching, T.M.; Crane, J.M. Adenylate energy pool and energy charge in maturing rape seeds. Plant Physiol. 1974, 54, 748–751. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Raymond, P.; Pradet, A. Stabilization of adenine nucleotide ratios at various values by an oxygen limitation of respiration in germinating lettuce (Lactuca sativa) seeds. Biochem J. 1980, 190, 39–44. [Google Scholar] [CrossRef]
- Chang, H.Y.; Fu, C.Y. Adenylate Kinase. Encycl. Food Microbiol. 2014, 1, 16–24. [Google Scholar]
- Noda, L. Adenylate kinase. In The Enzymes; Boyer, P.D., Ed.; Academic Press: New York, NY, USA, 1973; Volume 8, pp. 279–305. [Google Scholar]
- Pradet, A.; Raymond, P. Adenine nucleotide ratios and adenylate energy charge in energy metabolism. Ann. Rev. Plant Physiol. 1983, 34, 199–224. [Google Scholar] [CrossRef]
- Maragakis, P.; Karplus, M. Large amplitude conformational change in proteins explored with a plastic network model: Adenylate kinase. J. Mol. Biol. 2005, 352, 807–822. [Google Scholar] [CrossRef]
- Arora, K.; Brooks, C.L. Large-scale allosteric conformational transitions of adenylate kinase appear to involve a population-shift mechanism. Proc. Natl. Acad. Sci. USA 2007, 104, 18496–18501. [Google Scholar] [CrossRef]
- Kawai, M.; Kidou, S.; Kato, A.; Uchimiya, H.; Biology, M. Molecular characterization of cDNA encoding for adenylate kinase of rice (Oryza sativa L.). Plant J. 1992, 2, 845–854. [Google Scholar]
- Chen, Z.; Fu, H.; Liu, D.; Chang, P.F.; Narasimhan, M.; Ferl, R.; Hasegawa, P.M.; Bressan, R.A. A NaCl-regulated plant gene encoding a brain protein homolog that activates ADP ribosyltransferase and inhibits protein kinase C. Plant J. 1994, 6, 729–740. [Google Scholar]
- Regierer, B.; Fernie, A.R.; Springer, F.; Perez-Melis, A.; Leisse, A.; Koehl, K.; Willmitzer, L.; Geigenberger, P.; Kossmann, J. Starch content and yield increase as a result of altering adenylate pools in transgenic plants. Nat. Biotechnol. 2002, 20, 1256–1260. [Google Scholar] [CrossRef]
- Carrari, F.; Coll-Garcia, D.; Schauer, N.; Lytovchenko, A.; Palacios-Rojas, N.; Balbo, I.; Rosso, M.; Fernie, A.R. Deficiency of a plastidial adenylate kinase in Arabidopsis results in elevated photosynthetic amino acid biosynthesis and enhanced growth. Plant Physiol. 2005, 137, 70–82. [Google Scholar] [CrossRef]
- Lange, P.R.; Geserick, C.; Tischendorf, G.; Zrenner, R. Functions of chloroplastic adenylate kinases in Arabidopsis. Plant Physiol. 2008, 146, 492–504. [Google Scholar] [CrossRef]
- Zhang, Y.Z.; Launay, H.; Liu, F.; Lebrun, R.; Gontero, B. Interaction between adenylate kinase 3 and glyceraldehyde-3-phosphate dehydrogenase from Chlamydomonas reinhardtii. FEBS J. 2018, 285, 2495–2503. [Google Scholar] [CrossRef] [PubMed]
- Peterson, T.A.; NIieman, R.H.; Clark, R.A. Nucleotide metabolism in salt-stressed Zea mays L. Root tips: I. Adenine and uridine nucleotides. Plant Physiol. 1987, 85, 984–989. [Google Scholar] [CrossRef]
- Zhou, S.; Wei, S.; Boone, B.; Levy, S. Microarray analysis of genes affected by salt stress in tomato. Afr. J. Environ. Sci. Technol. 2007, 1, 014–026. [Google Scholar]
- Gong, P.; Zhang, J.; Li, H.; Yang, C.; Zhang, C.; Zhang, X.; Khurram, Z.; Zhang, Y.; Wang, T.; Fei, Z.; et al. Transcriptional profiles of drought-responsive genes in modulating transcription signal transduction, and biochemical pathways in tomato. J. Exp. Bot. 2010, 61, 3563–3575. [Google Scholar] [CrossRef]
- Raveneau, M.P.; Benamar, A.; Macherel, D. Water content, adenylate kinase, and mitochondria drive adenylate balance in dehydrating and imbibing seeds. J. Exp. Bot. 2017, 68, 3501–3512. [Google Scholar] [CrossRef]
- Giovannoni, J. Molecular biology of fruit maturation and ripening. Annu. Rev. Plant Physiol. Plant Mol. Biol. 2001, 52, 725–749. [Google Scholar] [CrossRef]
- Xu, G.; Guo, C.; Shan, H.; Kong, H. Divergence of duplicate genes in exon-intron structure. Proc. Natl. Acad. Sci. USA 2012, 109, 1187–1192. [Google Scholar] [CrossRef]
- Lorenzo, O.; Solano, R. Molecular players regulating the jasmonate signalling network. Curr. Opin. Plant Biol. 2005, 8, 532–540. [Google Scholar] [CrossRef] [PubMed]
- Mauch-Mani, B.; Mauch, F. The role of abscisic acid in plant-pathogen interactions. Curr. Opin. Plant Biol. 2005, 8, 409–414. [Google Scholar] [CrossRef] [PubMed]
- Hirayama, T.; Shinozaki, K. Research on plant abiotic stress responses in the post-genome era: Past, present and future. Plant J. 2010, 61, 1041–1052. [Google Scholar] [CrossRef]
- Zhao, P.; Wang, D.; Wang, R.; Kong, N.; Zhang, C.; Yang, C.; Wu, W.; Ma, H.; Chen, Q. Genome-wide analysis of the potato Hsp20 gene family: Identification, genomic organization and expression profiles in response to heat stress. BMC Genomics 2018, 19, 61. [Google Scholar] [CrossRef] [PubMed]
- Birkenhead, K.; Walker, D.; Foyer, C. The intracellular distribution of adenylate kinase in the leaves of spinach, wheat and barley. Planta 1982, 156, 171–175. [Google Scholar] [CrossRef] [PubMed]
- Hampp, R.; Goller, M.; Ziegler, H. Adenylate levels, energy charge, and phosphorylation potential during dark-light and light-dark transition in chloroplasts, mitochondria, and cytosol of mesophyll protoplasts from Avena sativa L. Plant Physiol. 1982, 69, 448–455. [Google Scholar] [CrossRef]
- Stitt, M.; Lilley, R.M.; Heldt, H.W. Adenine nucleotide levels in the cytosol, chloroplasts, and mitochondria of wheat leaf protoplasts. Plant Physiol. 1982, 70, 971–977. [Google Scholar] [CrossRef]
- Wang, X.; Xue, S.; Chen, S.; Ma, C.; Xu, S. Evolutionary origin, gradual accumulation and functional divergence of heat shock factor gene family with plant evolution. Front. Plant Sci. 2018, 9, 71. [Google Scholar] [CrossRef] [PubMed]
- Shinozaki, K.; Yamaguchi-Shinozaki, K. Molecular responses to drought and cold stress. Curr. Opin. Biotechnol. 1996, 7, 161–167. [Google Scholar] [CrossRef]
- Bray, E.A. Plant responses to water deficit. Trends. Plant Sci. 1997, 2, 48–54. [Google Scholar] [CrossRef]
- Zhu, J.K. Salt and drought stress signal transduction in plants. Annu. Rev. Plant Biol. 2002, 53, 247–273. [Google Scholar] [CrossRef]
- Shinozaki, K.; Yamaguchi-Shinozaki, K. Gene networks involved in drought stress response and tolerance. J. Exp. Bot. 2007, 58, 221–227. [Google Scholar] [CrossRef]
- Shen, Q.; Ho, T.H. Functional dissection of an abscisic acid (ABA)-inducible gene reveals two independent ABA-responsive complexes each containing a G-box and a novel cis-acting element. Plant Cell. 1995, 7, 295–307. [Google Scholar]
- Yamaguchi-Shinozaki, K.; Urao, T.; Shinozaki, K. Regulation of genes that are induced by drought stress in Arabidopsis thaliana. J. Plant Res. 1995, 108, 127–136. [Google Scholar] [CrossRef]
- Agarwal, M.; Hao, Y.; Kapoor, A.; Dong, C.; Fujii, H.; Zheng, X.; Zhu, J. A R2R3 type MYB transcription factor is involved in the cold regulation of CBF genes and in acquired freezing tolerance. J. Biol. Chem. 2006, 281, 37636–37645. [Google Scholar] [CrossRef]
- Onishi, M.; Tachi, H.; Kojima, T.; Shiraiwa, M.; Takahara, H. Molecular cloning and characterization of a novel salt-inducible gene encoding an acidic isoform of PR-5 protein in soybean (Glycine max [L.] Merr.). Plant Physiol. Biochem. 2006, 44, 574–580. [Google Scholar] [CrossRef]
- Moore, R.C.; Purugganan, M.D. The early stages of duplicate gene evolution. Proc. Natl. Acad. Sci. USA 2003, 100, 15682–15687. [Google Scholar] [CrossRef]
- Kong, H.; Landherr, L.L.; Frohlich, M.W.; Leebens-Mack, J.; Ma, H.; DePamphilis, C.W. Patterns of gene duplication in the plant SKP1 gene family in angiosperms: Evidence for multiple mechanisms of rapid gene birth. Plant J. 2007, 50, 873–885. [Google Scholar] [CrossRef]
- Cannon, S.B.; Mitra, A.; Baumgarten, A.; Young, N.D.; May, G. The roles of segmental and tandem gene duplication in the evolution of large gene families in Arabidopsis thaliana. BMC Plant Biol. 2004, 4, 10. [Google Scholar] [CrossRef]
- Bray, E.A.; Bailey-Serres, J.; Weretilnyk, E. Responses to abiotic stress. In Biochemistry and Molecular Biology of Plants; Buchanan, B., Gruissem, W., Jones, R., Eds.; The American Society of Plant Physiologists: Rockville, MD, USA, 2000; pp. 1158–1203. [Google Scholar]
- Najami, N.; Janda, T.; Barriah, W.; Kayam, G.; Tal, M.; Guy, M.; Volokita, M. Ascorbate peroxidase gene family in tomato: Its identifcation and characterization. Mol. Genet. Genomics 2008, 279, 171–182. [Google Scholar] [CrossRef]
- Zhang, C.; Liu, J.; Zhang, Y.; Cai, X.; Gong, P.; Zhang, J.; Wang, T.; Li, H.; Ye, Z. Overexpression of SlGMEs leads to ascorbate accumulation with enhanced oxidative stress, cold, and salt tolerance in tomato. Plant Cell Reports 2011, 30, 389–398. [Google Scholar] [CrossRef] [PubMed]
- Zegzouti, H.; Jones, B.; Marty, C.; Lelievre, J.M.; Latche, A.; Pech, J.C.; Bouzayen, M. ER5, a tomato cDNA encoding an ethylene-responsive LEA-like protein: Characterization and expression in response to drought, ABA and wounding. Plant Mol. Biol. 1997, 35, 847–854. [Google Scholar] [CrossRef] [PubMed]
- Li, F.F.; Chen, X.Y.; Zhou, S.G.; Xie, Q.L.; Wang, Y.S.; Xiang, X.X.; Hu, Z.L.; Chen, G.P. Overexpression of SlMBP22 in tomato affects plant growth and enhances tolerance to drought stress. Plant Sci. 2020, 301, 110672. [Google Scholar] [CrossRef]
- Atkin, O.K.; Macherel, D. The crucial role of plant mitochondria in orchestrating drought tolerance. Ann. Bot. 2009, 103, 581–597. [Google Scholar] [CrossRef] [PubMed]
- Woodward, A.W.; Bartel, B. Auxin: Regulation, action, and interaction. Ann. Bot. 2005, 95, 707–735. [Google Scholar] [CrossRef]
- Kumar, R.; Khurana, A.; Sharma, A.K. Role of plant hormones and their interplay in development and ripening of fleshy fruits. J. Exp. Bot. 2014, 65, 4561–4575. [Google Scholar] [CrossRef]
- Lee, S.C.; Luan, S. ABA signal transduction at the crossroad of biotic and abiotic stress responses. Plant Cell Environ. 2012, 35, 53–60. [Google Scholar] [CrossRef]
- Yu, X.; Zhang, W.; Zhang, Y.; Zhang, X.; Lang, D.; Zhang, X. The roles of methyl jasmonate to stress in plants. Funct. Plant Biol. 2019, 46, 197–212. [Google Scholar] [CrossRef]
- Tanaka, Y.; Sano, T.; Tamaoki, M.; Nakajima, N.; Kondo, N.; Hasezawa, S. Ethylene inhibits abscisic acid-induced stomatal closure in Arabidopsis. Plant Physiol. 2005, 138, 2337–2343. [Google Scholar] [CrossRef]
- Fujita, M.; Fujita, Y.; Noutoshi, Y.; Takahashi, F.; Narusaka, Y.; Yamaguchi-Shinozaki, K.; Shinozaki, K. Crosstalk between abiotic and biotic stress responses: A current view from the points of convergence in the stress signaling networks. Curr. Opin. Plant Biol. 2006, 9, 436–442. [Google Scholar] [CrossRef]
- Fuller, T.; Langfelder, P.; Presson, A.; Horvath, S. Review of weighted gene coexpression network analysis. In Handbook of Statistical Bioinformatics; Springer: Berlin/Heidelberg, Germany, 2011; pp. 369–388. [Google Scholar]
- Ruan, J.; Dean, A.K.; Zhang, W. A general co-expression network-based approach to gene expression analysis: Comparison and applications. BMC Syst. Biol. 2010, 4, 8. [Google Scholar] [CrossRef]
- Tomato Genome Consortium. The tomato genome sequence provides insights into fleshy fruit evolution. Nature 2012, 485, 635–641. [Google Scholar] [CrossRef] [PubMed]
- Finn, R.D.; Coggill, P.; Eberhardt, R.Y.; Eddy, S.R.; Mistry, J.; Mitchell, A.L.; Potter, S.C.; Punta, M.; Qureshi, M.; Sangrador-Vegas, A.; et al. The Pfam protein families database: Towards a more sustainable future. Nucleic. Acid. Res. 2016, 44, D279–D285. [Google Scholar] [CrossRef]
- Johnson, L.S.; Eddy, S.R.; Portugaly, E. Hidden Markov model speed heuristic and iterative HMM search procedure. BMC Bioinform. 2010, 11, 431. [Google Scholar] [CrossRef]
- Letunic, I. SMART. Available online: http://smart.embl-heidelberg.de/ (accessed on 14 August 2019).
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol. Biol Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef]
- Hall, B.G. Building phylogenetic trees from molecular data with MEGA. Mol. Biol. Evol. 2013, 30, 1229–1235. [Google Scholar] [CrossRef]
- Bailey, T.L.; Boden, M.; Buske, F.A.; Frith, M.; Grant, C.E.; Clementi, L.; Ren, J.; Li, W.W.; Noble, W.S. MEME SUITE: Tools for motif discovery and searching. Nucleic. Acids. Res. 2009, 37, W202–W208. [Google Scholar] [CrossRef]
- Artimo, P.; Jonnalagedda, M.; Arnold, K.; Baratin, D.; Csardi, G.; de Castro, E.; Duvaud, S.; Flegel, V.; Fortier, A.; Gasteiger, E.; et al. ExPASy: SIB bioinformatics resource portal. Nucleic Acids Res. 2012, 40, W597–W603. [Google Scholar] [CrossRef]
- Hu, B.; Jin, J.; Guo, A.Y.; Zhang, H.; Luo, J.; Gao, G. GSDS 2.0: An upgraded gene feature visualization server. Bioinformatics 2015, 31, 1296–1297. [Google Scholar] [CrossRef] [PubMed]
- Voorrips, R.E. MapChart: Software for the graphical presentation of linkage maps and QTLs. J. Hered. 2002, 93, 77–78. [Google Scholar] [CrossRef]
- Lescot, M.; Déhais, P.; Thijs, G.; Marchal, K.; Moreau, Y.; Van de Peer, Y.; Rouze, P.; Rombauts, S. Plantcare, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acids Res. 2002, 30, 325–327. [Google Scholar] [CrossRef]
- Wang, Y.; Tang, H.; DeBarry, J.D.; Tan, X.; Li, J.; Wang, X.; Lee, T.H.; Jin, H.; Marler, B.; Guo, H. MCScanX: A toolkit for detection and evolutionary analysis of gene synteny and collinearity. Nucleic Acids Res. 2012, 40, e49. [Google Scholar] [CrossRef] [PubMed]
- Krzywinski, M.; Schein, J.; Birol, I.; Connors, J.; Gascoyne, R.; Horsman, D.; Jones, S.J.; Marra, M.A. Circos: An information aesthetic for comparative genomics. Genome Res. 2009, 19, 1639–1645. [Google Scholar] [CrossRef] [PubMed]
- Liu, M.; Gomes, B.L.; Mila, I.; Purgatto, E.; Peres, L.E.; Frasse, P.; Maza, E.; Zouine, M.; Roustan, J.P.; Bouzayen, M.; et al. Comprehensive profiling of ethylene response factor expression identifies ripening-associated ERF genes and their link to key regulators of fruit ripening in Tomato. Plant Physiol. 2016, 170, 1732–1744. [Google Scholar] [CrossRef]
- Zouine, M.; Maza, E.; Djari, A.; Lauvernier, M.; Frasse, P.; Smouni, A.; Pirrello, J.; Bouzayen, M. Tomexpress, a unified tomato RNA-Seq platform for visualization of expression data, clustering and correlation networks. Plant J. 2017, 92, 727–735. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Schiff, M.; Dinesh-Kumar, S.P. Virus-induced gene silencing in tomato. Plant J. 2002, 31, 777–786. [Google Scholar] [CrossRef]
- Senthil-Kumar, M.; Mysore, K.S. Tobacco rattle virus–based virus induced gene silencing in Nicotiana benthamiana. Nat. Protoc. 2014, 9, 1549–1562. [Google Scholar] [CrossRef]
| Name | Gene ID | Chr | Genomic Location | Strand | ORF | Exon | AA | MW (kDa) | PI |
|---|---|---|---|---|---|---|---|---|---|
| SlADK1 | Solyc01g088480.2.1 | 1 | 83267069–83271080 | – | 630 | 10 | 209 | 22872.10 | 5.76 |
| SlADK2 | Solyc02g093990.2.1 | 2 | 54662319–54663928 | – | 780 | 4 | 259 | 29003.16 | 6.45 |
| SlADK3 | Solyc03g005050.2.1 | 3 | 44287–46883 | + | 852 | 4 | 283 | 31654.06 | 6.46 |
| SlADK4 | Solyc03g083610.2.1 | 3 | 53563661–53567185 | + | 714 | 10 | 237 | 26768.76 | 8.83 |
| SlADK5 | Solyc03g111200.2.1 | 3 | 61838312–61842194 | – | 735 | 6 | 244 | 26669.91 | 8.57 |
| SlADK6 | Solyc04g049690.2.1 | 4 | 43054097–43062321 | – | 810 | 4 | 269 | 30011.55 | 6.90 |
| SlADK7 | Solyc05g014980.2.1 | 5 | 9188640–9193694 | – | 729 | 6 | 242 | 26541.56 | 7.01 |
| SlADK8 | Solyc06g065270.2.1 | 6 | 40705359–40709638 | – | 861 | 7 | 286 | 31970.68 | 6.96 |
| SlADK9 | Solyc08g077300.2.1 | 8 | 61201659–61206762 | – | 651 | 8 | 216 | 24411.10 | 7.63 |
| SlADK10 | Solyc09g007180.2.1 | 9 | 801969–804985 | – | 846 | 4 | 281 | 30520.98 | 6.36 |
| SlADK11 | Solyc12g010380.2.1 | 12 | 3429721–3438938 | – | 1989 | 19 | 662 | 74110.62 | 6.67 |
| Predicted Subcellular Localization | |||||
|---|---|---|---|---|---|
| Locus Name | Name | Wolf PSort | Predotar | TargetP | CELLO |
| Solyc01g088480 | SlADK1 | cyto: 13 | none | other | cyto(2.70) |
| Solyc02g093990 | SlADK2 | mito: 6 | mitochondria | mitochondrial transfer peptide | mito(1.76)/cyto(1.28) |
| Solyc03g005050 | SlADK3 | mito: 8 | mitochondria | mitochondrial transfer peptide | cyto(1.49)/nucl(1.46)/mito(1.15) |
| Solyc03g083610 | SlADK4 | mito: 7.5 | mitochondria | mitochondrial transfer peptide | cyto(2.93) |
| Solyc03g111200 | SlADK5 | cyto: 8 | none | other | cyto(1.83)/mito(1.70) |
| Solyc04g049690 | SlADK6 | mito: 10 | mitochondria | mitochondrial transfer peptide | mito(1.81)/nucl(1.52) |
| Solyc05g014980 | SlADK7 | cyto: 6 | none | other | mito(1.83)/cyto(1.51) |
| Solyc06g065270 | SlADK8 | chlo: 14 | plastid | chloroplast transfer peptide | chlo(2.32) |
| Solyc08g077300 | SlADK9 | cyto: 9 | none | other | nucl(2.74) |
| Solyc09g007180 | SlADK10 | chlo: 11 | mitochondria | mitochondrial transfer peptide | chlo(3.70) |
| Solyc12g010380 | SlADK11 | chlo: 11 | plastid | chloroplast transfer peptide | chlo(3.42) |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yang, L.; Cao, H.; Zhang, X.; Gui, L.; Chen, Q.; Qian, G.; Xiao, J.; Li, Z. Genome-Wide Identification and Expression Analysis of Tomato ADK Gene Family during Development and Stress. Int. J. Mol. Sci. 2021, 22, 7708. https://doi.org/10.3390/ijms22147708
Yang L, Cao H, Zhang X, Gui L, Chen Q, Qian G, Xiao J, Li Z. Genome-Wide Identification and Expression Analysis of Tomato ADK Gene Family during Development and Stress. International Journal of Molecular Sciences. 2021; 22(14):7708. https://doi.org/10.3390/ijms22147708
Chicago/Turabian StyleYang, Lu, Haohao Cao, Xiaoping Zhang, Liangxian Gui, Qiang Chen, Gui Qian, Jiaxin Xiao, and Zhengguo Li. 2021. "Genome-Wide Identification and Expression Analysis of Tomato ADK Gene Family during Development and Stress" International Journal of Molecular Sciences 22, no. 14: 7708. https://doi.org/10.3390/ijms22147708
APA StyleYang, L., Cao, H., Zhang, X., Gui, L., Chen, Q., Qian, G., Xiao, J., & Li, Z. (2021). Genome-Wide Identification and Expression Analysis of Tomato ADK Gene Family during Development and Stress. International Journal of Molecular Sciences, 22(14), 7708. https://doi.org/10.3390/ijms22147708