Functional Validation of cas9/GuideRNA Constructs for Site-Directed Mutagenesis of Triticale ABA8′OH1 loci
Abstract
:1. Introduction
2. Results
2.1. Selection of Target Motifs
2.2. Utility of Cas9/gRNA Target Motifs in ABA8′OH-1 of Triticale
2.3. Selection of Target Motifs under Consideration of Predicted On- and Off-Targets
2.4. Modeling the Secondary Structure of Guide RNAs
2.5. Assessment of Gene Editing Frequency via Frameshift Restitution in a Transiently Expressed Reporter Gene
2.6. Assessment of Gene Editing Frequency in Triticale Protoplasts Using T7E1 Assay
2.7. Assessment of Targeted Mutagenesis Frequency in Triticale Protoplasts Using Deep-Sequencing of Short Reads
3. Discussion
3.1. Off-Target and On-Target SNP Evaluation
3.2. Protoplast Transfection as a Convenient Test for Cas9/gRNA Functionality
3.3. Gene Editing Frequency in Plants Can Be Estimated via Transient Expression
3.4. The Effect of TREX2 Co-Expression on Cas9-Mediated Gene Editing
3.5. Prospects of Triticale Cas9-Mediated Gene Editing
4. Materials and Methods
4.1. Plant Material and ABA8′OH-1 Identification in Triticale
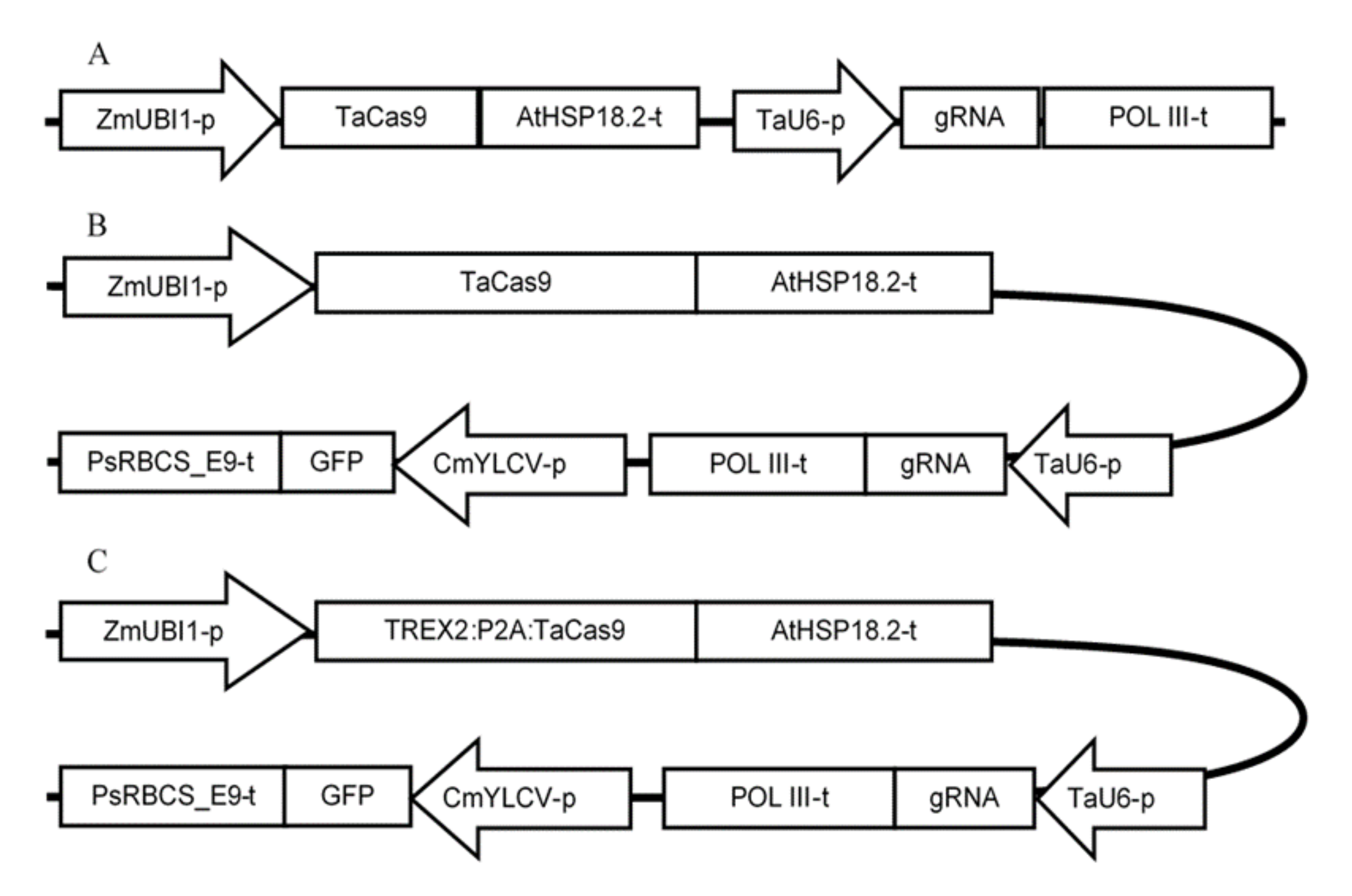
4.2. gRNA Design, 2D Modeling, and Vectors Construction
4.3. Transient Expression Method
4.4. Protoplast Transfection Assay
4.5. Deep-Sequencing of Amplicons
4.6. Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Li, C.; Brant, E.; Budak, H.; Zhang, B. CRISPR/Cas: A Nobel Prize award-winning precise genome editing technology for gene therapy and crop improvement. J. Zhejiang Univ. Sci. B 2021, 22, 253–284. [Google Scholar] [CrossRef] [PubMed]
- Ma, X.; Liu, Y.G. CRISPR/Cas9-Based Multiplex Genome Editing in Monocot and Dicot Plants. Curr. Protoc. Mol. Biol. 2016, 115, 31.6.1–31.6.21. [Google Scholar] [CrossRef] [PubMed]
- Hisano, H.; Abe, F.; Hoffie, R.E.; Kumlehn, J. Targeted genome modification in cereals. Breed. Sci. 2021, BS21019, in press. [Google Scholar]
- Sánchez-León, S.; Gil-Humanes, J.; Ozuna, C.V.; Giménez, M.J.; Sousa, C.; Voytas, D.F.; Barro, F. Low-gluten, nontransgenic wheat engineered with CRISPR/Cas9. Plant. Biotechnol. J. 2018, 16, 902–910. [Google Scholar] [CrossRef]
- Abe, F.; Haque, E.; Hisano, H.; Tanaka, T.; Kamiya, Y.; Mikami, M.; Kawaura, K.; Endo, M.; Onishi, K.; Hayashi, T.; et al. Genome-Edited Triple-Recessive Mutation Alters Seed Dormancy in Wheat. Cell Rep. 2019, 28, 1362–1369.e1364. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, J.; Jiao, G.; Sun, Y.; Chen, J.; Zhong, Y.; Yan, L.; Jiang, D.; Ma, Y.; Xia, L. Modification of starch composition, structure and properties through editing of TaSBEIIa in both winter and spring wheat varieties by CRISPR/Cas9. Plant. Biotechnol. J. 2020, 19, 937–951. [Google Scholar] [CrossRef]
- Budhagatapalli, N.; Halbach, T.; Hiekel, S.; Büchner, H.; Müller, A.E.; Kumlehn, J. Site-directed mutagenesis in bread and durum wheat via pollination by cas9/guide RNA-transgenic maize used as haploidy inducer. Plant. Biotechnol. J. 2020, 18, 2376–2378. [Google Scholar] [CrossRef]
- Lin, Q.; Zong, Y.; Xue, C.; Wang, S.; Jin, S.; Zhu, Z.; Wang, Y.; Anzalone, A.V.; Raguram, A.; Doman, J.L.; et al. Prime genome editing in rice and wheat. Nat. Biotechnol. 2020, 38, 582–585. [Google Scholar] [CrossRef] [PubMed]
- Zhang, R.; Liu, J.; Chai, Z.; Chen, S.; Bai, Y.; Zong, Y.; Chen, K.; Li, J.; Jiang, L.; Gao, C. Generation of herbicide tolerance traits and a new selectable marker in wheat using base editing. Nat. Plants 2019, 5, 480–485. [Google Scholar] [CrossRef] [PubMed]
- Kumar, R.; Kaur, A.; Pandey, A.; Mamrutha, H.M.; Singh, G.P. CRISPR-based genome editing in wheat: A comprehensive review and future prospects. Mol. Biol. Rep. 2019, 46, 3557–3569. [Google Scholar] [CrossRef]
- Arndell, T.; Sharma, N.; Langridge, P.; Baumann, U.; Watson-Haigh, N.S.; Whitford, R. gRNA validation for wheat genome editing with the CRISPR-Cas9 system. BMC Biotechnol. 2019, 19, 71. [Google Scholar] [CrossRef] [PubMed]
- Paul, J.W., 3rd; Qi, Y. CRISPR/Cas9 for plant genome editing: Accomplishments, problems and prospects. Plant. Cell Rep. 2016, 35, 1417–1427. [Google Scholar] [CrossRef]
- Kumlehn, J.; Pietralla, J.; Hensel, G.; Pacher, M.; Puchta, H. The CRISPR/Cas revolution continues: From efficient gene editing for crop breeding to plant synthetic biology. J. Integr. Plant Biol. 2018, 60, 1127–1153. [Google Scholar] [CrossRef]
- Shockey, J. Gene editing in plants: Assessing the variables through a simplified case study. Plant Mol. Biol. 2020, 103, 75–89. [Google Scholar] [CrossRef] [Green Version]
- Koeppel, I.; Hertig, C.; Hoffie, R.; Kumlehn, J. Cas endonuclease technology—A quantum leap in the advancement of barley and wheat genetic engineering. Int. J. Mol. Sci. 2019, 20, 2647. [Google Scholar] [CrossRef] [Green Version]
- Uusi-Mäkelä, M.I.E.; Barker, H.R.; Bäuerlein, C.A.; Häkkinen, T.; Nykter, M.; Rämet, M. Chromatin accessibility is associated with CRISPR-Cas9 efficiency in the zebrafish (Danio rerio). PLoS ONE 2018, 13, e0196238. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Budhagatapalli, N.; Schedel, S.; Gurushidze, M.; Pencs, S.; Hiekel, S.; Rutten, T.; Kusch, S.; Morbitzer, R.; Lahaye, T.; Panstruga, R.; et al. A simple test for the cleavage activity of customized endonucleases in plants. Plant Methods 2016, 12, 18. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chen, M.J.; Ma, S.M.; Dumitrache, L.C.; Hasty, P. Biochemical and cellular characteristics of the 3′-> 5′ exonuclease TREX2. Nucleic Acids Res. 2007, 35, 2682–2694. [Google Scholar] [CrossRef] [Green Version]
- Mazur, D.J.; Perrino, F.W. Identification and expression of the TREX1 and TREX2 cDNA sequences encoding mammalian 3-prime-to-5-prime exonucleases. J. Biol. Chem. 1999, 274, 19655–19660. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hasty, P. Trex2 responds to damaged replication forks in diverse ways. Mol. Cell Oncol. 2021, 8, 1881394. [Google Scholar] [CrossRef]
- Ko, J.H.; Son, M.Y.; Zhou, Q.; Molnarova, L.; Song, L.; Mlcouskova, J.; Jekabsons, A.; Montagna, C.; Krejci, L.; Hasty, P. TREX2 exonuclease causes spontaneous mutations and stress-induced replication fork defects in cells expressing RAD51(K133A). Cell Rep. 2020, 33, 108543. [Google Scholar] [CrossRef]
- Okamoto, M.; Kuwahara, A.; Seo, A.; Kushiro, T.; Asami, A.; Hira Ni Yuji Kamiy, Y.; Koshiba, T.; Nambara, E. CYP707A1 and CYP707A2, Which Encode Abscisic Acid 8′-Hydroxylases, Are Indispensable for Proper Control of Seed Dormancy and Germination in Arabidopsis. Plant Physiol. 2006, 141, 97–107. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fidler, J.; Grabowska, A.; Prabucka, B.; Więsyk, A.; Góra-Sochacka, A.; Bielawski, W.; Pojmaj, M.; Zdunek-Zastocka, E. The varied ability of grains to synthesize and catabolize ABA is one of the factors affecting dormancy and its release by afterripening in imbibed triticale grains of cultivars with different pre-harvest sprouting susceptibilities. J. Plant Physiol. 2018, 226, 48–55. [Google Scholar] [CrossRef] [PubMed]
- Park, J.; Bae, S.; Kim, J.-S. Cas-Designer: A web-based tool for choice of CRISPR-Cas9 target sites. Bioinformatics 2015, 31, 4014–4016. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wong, N.; Liu, W.; Wang, X. WU-CRISPR: Characteristics of functional guide RNAs for the CRISPR/Cas9 system. Genome Biol. 2015, 16, 218. [Google Scholar] [CrossRef] [Green Version]
- Gruber, A.R.; Lorenz, R.; Bernhart, S.H.; Neuböck, R.; Hofacker, I.L. The Vienna RNA Websuite. Nucleic Acids Res. 2008, 36, W70–W74. [Google Scholar] [CrossRef] [Green Version]
- Shan, Q.W.; Wang, Y.P.; Li, J.; Gao, C.X. Genome editing in rice and wheat using the CRISPR/Cas system. Nat. Protoc. 2014, 9, 2395–2410. [Google Scholar] [CrossRef]
- Čermák, T.; Curtin, S.J.; Gil-Humanes, J.; Čegan, R.; Kono, T.J.Y.; Konecna, E.; Belanto, J.J.; Starker, C.G.; Mathre, J.W.; Greenstein, R.L.; et al. A multi-purpose toolkit to enable advanced genome engineering in plants. Plant Cell 2017, 18. [Google Scholar] [CrossRef] [Green Version]
- Vouillot, L.; Thelie, A.; Pollet, N. Comparison of T7E1 and surveyor mismatch cleavage assays to detect mutations triggered by engineered nucleases. G3 2015, 5, 407–415. [Google Scholar] [CrossRef] [Green Version]
- Gagnon, J.A.; Valen, E.T.; Thyme, S.B.; Huang, P.; Ahkmetova, L.; Pauli, A.; Montague, T.G.; Zimmerman, S.; Richter, C.; Schier, A.F. Efficient mutagenesis by Cas9 protein-mediated oligonucleotide insertion and large-scale assessment of single-guide RNAs. PLoS ONE 2014, 9, e98186. [Google Scholar] [CrossRef]
- Zischewski, J.; Fischer, R.; Bortesi, L. Detection of on-target and off-target mutations generated by CRISPR/Cas9 and other sequence-specific nucleases. Biotechnol. Adv. 2017, 35, 95–104. [Google Scholar] [CrossRef] [PubMed]
- Arora, L.; Narula, A. Gene Editing and Crop Improvement Using CRISPR-Cas9 System. Front. Plant Sci. 2017, 8, 1932. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Liang, G.; Zhang, H.; Lou, D.; Yu, D. Selection of highly efficient sgRNAs for CRISPR/Cas9-based plant genome editing. Sci. Rep. 2016, 6, 21451. [Google Scholar] [CrossRef]
- Xing, H.L.; Dong, L.; Wang, Z.P.; Zhang, H.Y.; Han, C.Y.; Liu, B.; Wang, X.C.; Chen, Q.J. A CRISPR/Cas9 toolkit for multiplex genome editing in plants. BMC Plant Biol. 2014, 14, 327. [Google Scholar] [CrossRef] [Green Version]
- Fauser, F.; Schiml, S.; Puchta, H. Both CRISPR/Cas-based nucleases and nickases can be used efficiently for genome engineering in Arabidopsis thaliana. Plant J. 2014, 79, 348–359. [Google Scholar] [CrossRef]
- Lowder, L.G.; Paul, J.W.; Baltes, N.J.; Voytas, D.F.; Zhang, Y.; Zhang, D.; Tang, X.; Zheng, X.; Hsieh, T.F.; Qi, Y. A CRISPR/Cas9 toolbox for multiplexed plant genome editing and transcriptional regulation. Plant Physiol. 2015, 169, 971–985. [Google Scholar] [CrossRef] [Green Version]
- Zong, Y.; Wang, Y.; Li, C.; Zhang, R.; Chen, K.; Ran, Y.; Qiu, J.L.; Wang, D.; Gao, C. Precise base editing in rice, wheat and maize with a Cas9-cytidine deaminase fusion. Nat. Biotechnol. 2017, 35, 438–440. [Google Scholar] [CrossRef]
- Weiss, T.; Wang, C.H.; Kang, X.; Zhao, H.; Gamo, M.E.; Starker, C.G.; Crisp, P.A.; Zhou, P.; Springer, N.M.; Voytas, D.F.; et al. Optimization of multiplexed CRISPR/Cas9 system for highly efficient genome editing in Setaria viridis. Biorxiv 2020. [Google Scholar] [CrossRef]
- Wang, T.; Wei, J.J.; Sabatini, D.M.; Lander, E.S. Genetic screens in human cells using the CRISPR-Cas9 system. Science 2014, 343, 80–84. [Google Scholar] [CrossRef] [Green Version]
- Moreno-Mateos, M.A.; Vejnar, C.E.; Beaudoin, J.D.; Fernandez, J.P.; Mis, E.K.; Khokha, M.K.; Giraldez, A.J. CRISPRscan: Designing highly efficient sgRNAs for CRISPR-Cas9 targeting in vivo. Nat. Methods 2015, 12, 982–988. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Graf, R.; Li, X.; Chu, V.T.; Rajewsky, K. sgRNA Sequence Motifs Blocking Efficient CRISPR/Cas9-Mediated Gene Editing. Cell Rep. 2019, 26, 1098–1103.e3. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kleinstiver, B.P.; Pattanayak, V.; Prew, M.S.; Tsai, S.Q.; Nguyen, N.T.; Zheng, Z.; Joung, J.K. High-fidelity CRISPR-Cas9 nucleases with no detectable genome-wide off-target effects. Nature 2016, 529, 490–495. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kocak, D.D.; Josephs, E.A.; Bhandarkar, V.; Adkar, S.S.; Kwon, J.B.; Gersbach, C.A. Increasing the specificity of CRISPR systems with engineered RNA secondary structures. Nat. Biotechnol. 2019, 37, 657–666. [Google Scholar] [CrossRef] [PubMed]
- Zheng, T.; Hou, Y.; Zhang, P.; Zhang, Z.; Xu, Y.; Zhang, L.; Niu, L.; Yang, Y.; Liang, D.; Yi, F.; et al. Profiling single-guide RNA specificity reveals a mismatch sensitive core sequence. Sci. Rep. 2017, 7, 40638. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cermak, T.; Baltes, N.J.; Cegan, R.; Zhang, Y.; Voytas, D.F. High frequency, precise modification of the tomato genome. Genome Biol. 2015, 16, 232. [Google Scholar] [CrossRef] [Green Version]
- Liang, Z.; Zhang, K.; Chen, K.L.; Gao, C.X. Targeted mutagenesis in Zea mays using TALENs and the CRISPR/Cas system. J. Genet. Genom. 2014, 41, 63–68. [Google Scholar] [CrossRef] [PubMed]
- Woo, J.W.; Kim, J.; Kwon, S.I.; Corvalan, C.; Cho, S.W.; Kim, H.; Kim, S.-G.; Kim, S.-T.; Choe, S.; Kim, J.-S. DNA-free genome editing in plants with preassembled CRISPRCas9 ribonucleoproteins. Nat. Biotechnol. 2015, 33, 1162–1164. [Google Scholar] [CrossRef] [PubMed]
- Murovec, J.; Guček, K.; Bohanec, B.; Avbelj, M.; Jerala, R. DNA-Free Genome Editing of Brassica oleracea and B. rapa Protoplasts Using CRISPR-Cas9 Ribonucleoprotein Complexes. Front. Plant. Sci. 2018, 9, 159. [Google Scholar] [CrossRef]
- Gerasimova, S.V.; Hertig, C.; Korotkova, A.M.; Kolosovskaya, E.V.; Otto, I.; Hiekel, S.; Kochetov, A.V.; Khlestkina, E.K.; Kumlehn, J. Conversion of hulled into naked barley by Cas endonuclease-mediated knockout of the NUD gene. BMC Plant Biol. 2020, 20, 255. [Google Scholar] [CrossRef] [PubMed]
- Zuo, Z.; Liu, J. Cas9-catalyzed DNA Cleavage Generates Staggered Ends: Evidence from Molecular Dynamics Simulations. Sci. Rep. 2016, 6, 37584. [Google Scholar] [CrossRef]
- Chari, R.; Mali, P.; Moosburner, M.; Church, G.M. Unraveling CRISPR-Cas9 genome engineering parameters via a library-on-library approach. Nat. Methods 2015, 12, 823–826. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sanjana, N.E.; Shalem, O.; Zhang, F. Improved vectors and genome-wide libraries for CRISPR screening. Nat. Methods 2014, 11, 783–784. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sievers, F.; Wilm, A.; Dineen, D.; Gibson, T.J.; Karplus, K.; Li, W.; Lopez, R.; McWilliam, H.; Remmert, M.; Söding, J.; et al. Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol. Syst. Biol. 2011, 7, 539. [Google Scholar] [CrossRef] [PubMed]
- Brown, N.P.; Leroy, C.; Sander, C. MView: A web-compatible database search or multiple alignment viewer. Bioinformatics 1998, 14, 380–381. [Google Scholar] [CrossRef]
- Geneious Prime (v2020.2.4) Software. Available online: https://www.geneious.com/ (accessed on 3 April 2021).
- Statistica (Data Analysis Software System), Version 13. TIBCO Software Inc., 2017. Available online: http://statistica.io (accessed on 3 April 2021).



| Chromosome | Species | Sequence 5′-3′ | Target |
|---|---|---|---|
| 6A/6B | triticale | GCCTACCTTCCCGGCCAGCAAGG | On-target motif ABA/1/364 |
| 6R | triticale | GCCCACCTTCCCGGCCAGCAAGG | On-target motif ABA/1/364 |
| 6R | rye | GCCGACGTTCCCGGCCAGCAAGG | Potential off-target motif ABA/1/364 |
| 7B | wheat | GCATACCTTCCCG-CCAGCATGG | Potential off-target motif ABA/1/364 |
| 6A/6B/6R | triticale | GACCATCCACGTGAGCACGCTGG | On-target motif ABA/2/323 |
| 3A | wheat | GACCATCCACGTGAG-ACTCAGG | Potential off-target motif ABA/2/323 |
| 3B | wheat | GACCAGCC-CGTGAGCACGCCGG | Potential off-target motif ABA/2/323 |
| 4A | wheat | G-CCAGCCACGTGAGCACGCCGG | Potential off-target motif ABA/2/323 |
| 5A/5B/5R | wheat, rye | GACGATCCAGGTGAGCACGCTGG | Potential off-target motif ABA/2/323 |
| Number of Red Fluorescing Cells | Number of Yellow Fluorescing Cells | Proportion of Yellow out of Total Fluorescing Cells | |
|---|---|---|---|
| gRNA-ABA/1/364 | 33 | 2 | 6.1% |
| 110 | 6 | 5.5% | |
| 38 | 2 | 5.3% | |
| 5.6 ± 0.42% | |||
| gRNA-ABA/2/323 | 62 | 14 | 22.6% |
| 101 | 27 | 26.7% | |
| 72 | 17 | 23.6% | |
| 24.3 ± 0.42% | |||
| Editing Efficiency [%] | |||
|---|---|---|---|
| Genome A | Genome B | Genome R | |
| gRNA-ABA/1/364 | 6.3 | 17.1 | 27.3 |
| 6.8 | 11.1 | 13.9 | |
| 5.3 | 18.6 | 32.3 | |
| 6.2 ± 0.8 a | 15.6 ± 4.0 a,b | 24.4 ± 9.5 b | |
| gRNA-ABA/1/364 +TREX2 | 14.9 | 35.2 | 32.7 |
| 27.5 | 22.0 | 22.5 | |
| 17.8 | 38.9 | 14.1 | |
| 20.1 ± 6.6 a | 32.0 ± 8.9 a | 23.1 ± 9.3 a | |
| gRNA-ABA/2/323 | 0.3 | 1.9 | ND |
| ND | 1.4 | ND | |
| 0.4 | 1.1 | ND | |
| 0.2 ± 0.2 a | 1.4 ± 0.4 b | ND | |
| gRNA-ABA/2/323 +TREX2 | 26.2 | 35.4 | 22.2 |
| 15.6 | 19.2 | 34.7 | |
| 27.5 | 19.0 | 27.3 | |
| 23.1 ± 6.5 a | 24.5 ± 9.4 a | 28.1 ± 6.3 a | |
| Editing Efficiency [%] | |||
|---|---|---|---|
| Genome A | Genome B | Genome R | |
| gRNA-ABA/1/364 | 25.6 | 33.2 | 32.0 |
| 27.6 | 20.7 | 23.6 | |
| 26.8 | 37.3 | 31.7 | |
| 26.7 ± 1.0 | 30.4 ± 8.6 | 29.1 ± 4.8 | |
| gRNA-ABA/1/364 +TREX2 | 50.4 | 36.7 | 38.6 |
| 63.4 | 48.4 | 62.9 | |
| 46.7 | 43.5 | 44.8 | |
| 53.5 ± 8.7 | 42.9 ± 5.9 | 48.8 ± 12.6 | |
| gRNA-ABA/2/323 | 0.4 | 0.2 | 1.7 |
| 2.4 | 2.4 | 5.5 | |
| 3.3 | 4.4 | 2.6 | |
| 2.0 ± 1.5 | 2.3 ± 2.1 | 3.3 ± 2.0 | |
| gRNA-ABA/2/323 +TREX2 | 31.2 | 50.7 | 43.2 |
| 15.2 | 33.0 | 22.8 | |
| 45.9 | 48.9 | 49.8 | |
| 30.7 ± 15.3 | 44.2 ± 9.7 | 38.6 ± 14.1 | |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Michalski, K.; Hertig, C.; Mańkowski, D.R.; Kumlehn, J.; Zimny, J.; Linkiewicz, A.M. Functional Validation of cas9/GuideRNA Constructs for Site-Directed Mutagenesis of Triticale ABA8′OH1 loci. Int. J. Mol. Sci. 2021, 22, 7038. https://doi.org/10.3390/ijms22137038
Michalski K, Hertig C, Mańkowski DR, Kumlehn J, Zimny J, Linkiewicz AM. Functional Validation of cas9/GuideRNA Constructs for Site-Directed Mutagenesis of Triticale ABA8′OH1 loci. International Journal of Molecular Sciences. 2021; 22(13):7038. https://doi.org/10.3390/ijms22137038
Chicago/Turabian StyleMichalski, Krzysztof, Christian Hertig, Dariusz R. Mańkowski, Jochen Kumlehn, Janusz Zimny, and Anna M. Linkiewicz. 2021. "Functional Validation of cas9/GuideRNA Constructs for Site-Directed Mutagenesis of Triticale ABA8′OH1 loci" International Journal of Molecular Sciences 22, no. 13: 7038. https://doi.org/10.3390/ijms22137038
APA StyleMichalski, K., Hertig, C., Mańkowski, D. R., Kumlehn, J., Zimny, J., & Linkiewicz, A. M. (2021). Functional Validation of cas9/GuideRNA Constructs for Site-Directed Mutagenesis of Triticale ABA8′OH1 loci. International Journal of Molecular Sciences, 22(13), 7038. https://doi.org/10.3390/ijms22137038






