PML Differentially Regulates Growth and Invasion in Brain Cancer
Abstract
1. Introduction
2. Results
2.1. Proliferation/Doubling Time Estimation
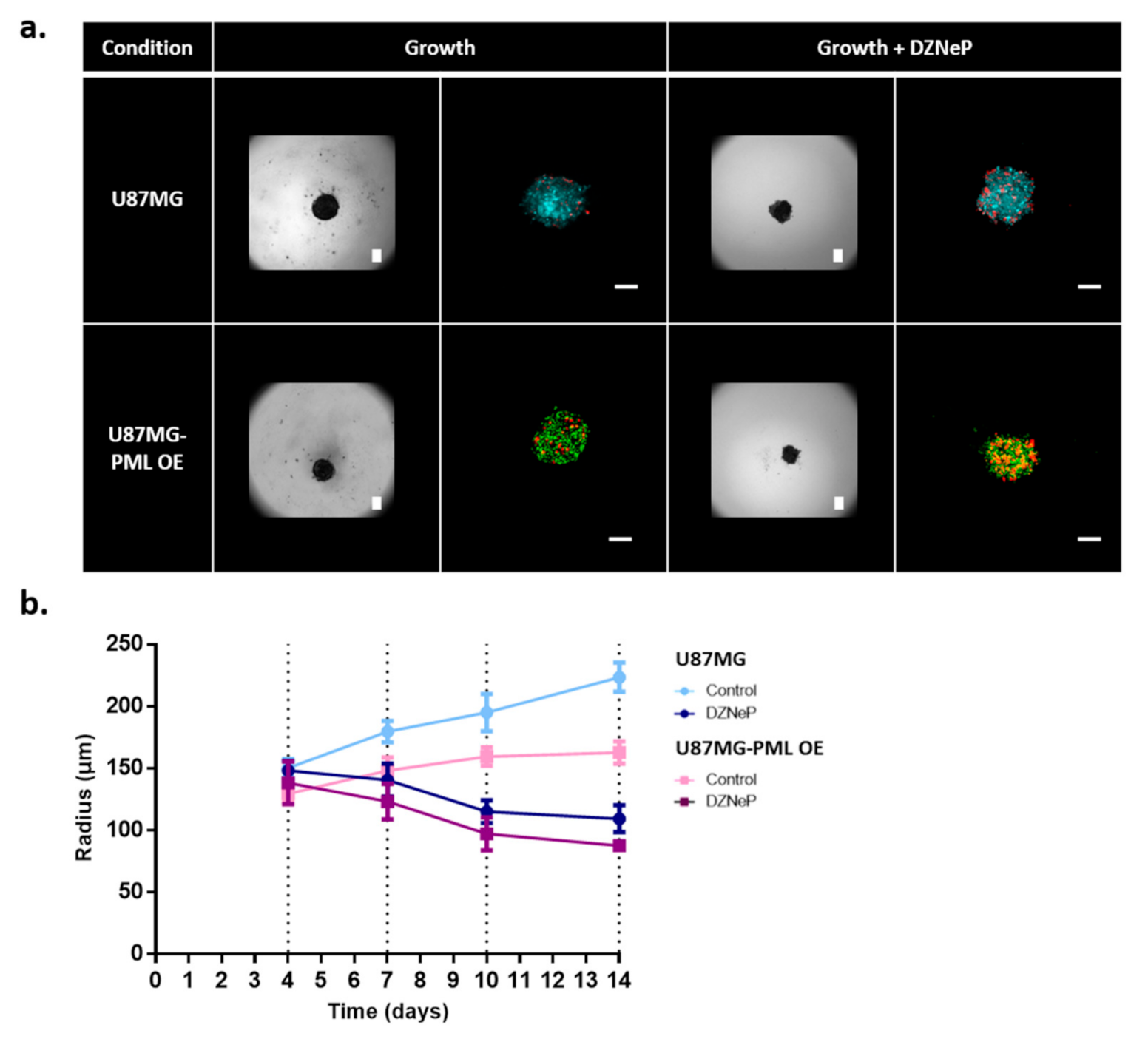
2.2. Tumor Growth Expansion
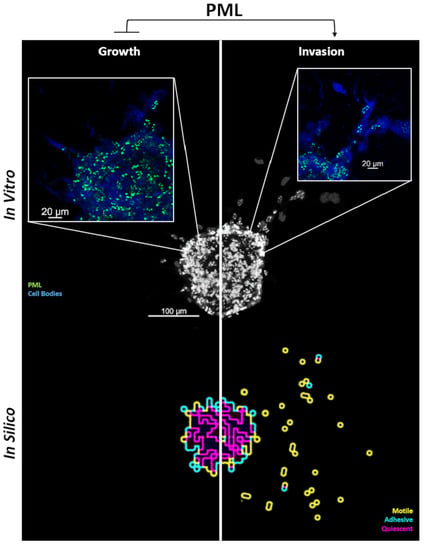
2.3. Tumor Invasion
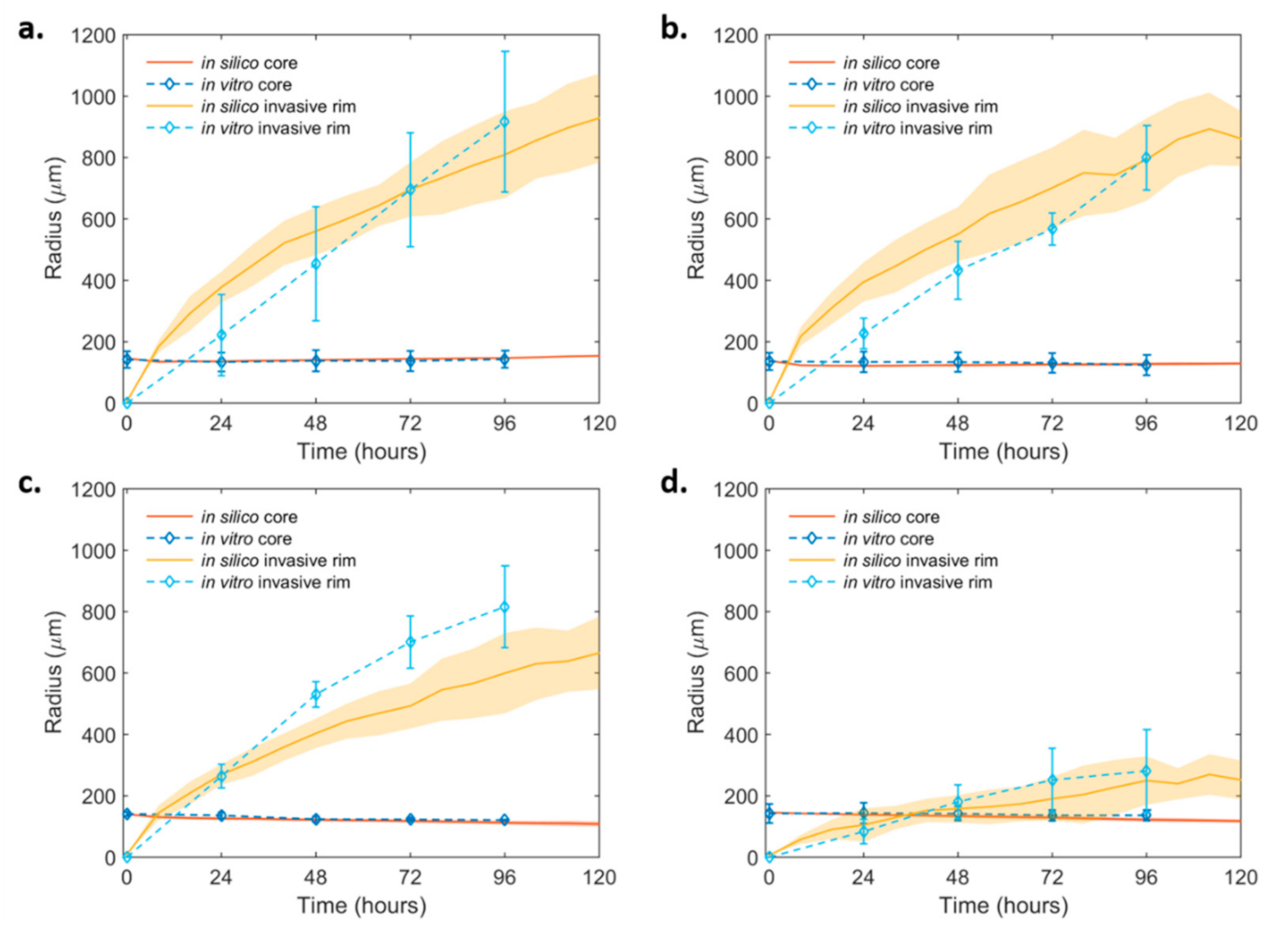
2.4. Mathematical Modeling and Analysis of Spheroid Growth and Invasion
2.4.1. Spheroid Growth Study
2.4.2. Spheroid Invasion Study
3. Discussion
4. Materials and Methods
4.1. Cell Lines
4.2. 2D Cell Cultures. Cell Proliferation, Cell Viability and Cytotoxicity Assays
4.2.1. Trypan Blue Viability Assay
4.2.2. Flow Cytometry
4.2.3. MTT Viability Assay
4.3. 3D Cell Cultures
4.3.1. Generation of 3D Multicellular Spheroids
4.3.2. Invasion Assay
4.3.3. Bright-Field Data Analysis
Spheroid Image Segmentation
Statistical Analysis
Tumor Growth Analysis
Tumor Invasion Analysis
4.3.4. Confocal Imaging and Analysis
4.4. In Silico Modeling
Parametrization of the Model
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Louis, D.N.; Perry, A.; Reifenberger, G.; von Deimling, A.; Figarella-Branger, D.; Cavenee, W.K.; Ohgaki, H.; Wiestler, O.D.; Kleihues, P.; Ellison, D.W. The 2016 World Health Organization Classification of Tumors of the Central Nervous System: A summary. Acta Neuropathol. 2016, 131, 803–820. [Google Scholar] [CrossRef]
- Hou, L.C.; Veeravagu, A.; Hsu, A.R.; Tse, V.C.K. Recurrent glioblastoma multiforme: A review of natural history and management options. Neurosurg. Focus 2006, 20, E3. [Google Scholar] [CrossRef]
- Mullins, C.S.; Schneider, B.; Stockhammer, F.; Krohn, M.; Classen, C.F.; Linnebacher, M. Establishment and Characterization of Primary Glioblastoma Cell Lines from Fresh and Frozen Material: A Detailed Comparison. PLoS ONE 2013, 8, e71070. [Google Scholar] [CrossRef]
- Hsu, K.S.; Kao, H.Y. PML: Regulation and multifaceted function beyond tumor suppression. Cell Biosci. 2018, 8, 1–21. [Google Scholar] [CrossRef] [PubMed]
- Jensen, K.; Shiels, C.; Freemont, P.S. PML Protein Isoforms and the RBCC/TRIM Motif. Oncogene 2001, 20, 7223–7233. [Google Scholar] [CrossRef]
- Bernardi, R.; Pandolfi, P.P. Structure, dynamics and functions of promyelocytic leukaemia nuclear bodies. Nat. Rev. Mol. Cell Biol. 2007, 8, 1006–1016. [Google Scholar] [CrossRef] [PubMed]
- Lee, H.E.; Do Jee, C.; Kim, M.A.; Lee, H.S.; Lee, Y.M.; Lee, B.L.; Kim, W.H. Loss of promyelocytic leukemia protein in human gastric cancers. Cancer Lett. 2007, 247, 103–109. [Google Scholar] [CrossRef] [PubMed]
- Martín-Martín, N.; Piva, M.; Urosevic, J.; Aldaz, P.; Sutherland, J.D.; Fernández-Ruiz, S.; Arreal, L.; Torrano, V.; Cortazar, A.R.; Planet, E.; et al. Stratification and therapeutic potential of PML in metastatic breast cancer. Nat. Commun. 2016, 7. [Google Scholar] [CrossRef] [PubMed]
- Regad, T.; Bellodi, C.; Nicotera, P.; Salomoni, P. The tumor suppressor Pml regulates cell fate in the developing neocortex. Nat. Neurosci. 2009, 12, 132–140. [Google Scholar] [CrossRef]
- Kaneko, N.; Marín, O.; Koike, M.; Hirota, Y.; Uchiyama, Y.; Wu, J.Y.; Lu, Q.; Tessier-Lavigne, M.; Alvarez-Buylla, A.; Okano, H.; et al. New neurons clear the path of astrocytic processes for their rapid migration in the adult brain. Neuron 2010. [Google Scholar] [CrossRef]
- Amodeo, V.; Deli, A.; Betts, J.; Bartesaghi, S.; Zhang, Y.; Richard-Londt, A.; Ellis, M.; Roshani, R.; Vouri, M.; Galavotti, S.; et al. A PML/Slit Axis Controls Physiological Cell Migration and Cancer Invasion in the CNS. Cell Rep. 2017, 20, 411–426. [Google Scholar] [CrossRef]
- Blockus, H.; Chédotal, A. The multifaceted roles of Slits and Robos in cortical circuits: From proliferation to axon guidance and neurological diseases. Curr. Opin. Neurobiol. 2014, 27, 82–88. [Google Scholar] [CrossRef]
- Margueron, R.; Reinberg, D. The Polycomb complex PRC2 and its mark in life. Nature 2011, 469, 343. [Google Scholar] [CrossRef]
- Cuddapah, V.A.; Robel, S.; Watkins, S.; Sontheimer, H. A neurocentric perspective on glioma invasion. Nat. Rev. Neurosci. 2014, 15, 455–465. [Google Scholar] [CrossRef]
- Faiz, M.; Sachewsky, N.; Gascón, S.; Bang, K.W.A.; Morshead, C.M.; Nagy, A. Adult Neural Stem Cells from the Subventricular Zone Give Rise to Reactive Astrocytes in the Cortex after Stroke. Cell Stem Cell 2015, 17, 624–634. [Google Scholar] [CrossRef]
- Zong, H.; Parada, L.F.; Baker, S.J. Cell of origin for malignant gliomas and its implication in therapeutic development. Cold Spring Harb. Perspect. Biol. 2015, 7, a020610. [Google Scholar] [CrossRef] [PubMed]
- Gurrieri, C.; Capodieci, P.; Bernardi, R.; Scaglioni, P.P.; Nafa, K.; Rush, L.J.; Verbel, D.A.; Cordon-Cardo, C.; Pandolfi, P.P. Loss of the tumor suppressor PML in human cancers of multiple histologic origins. J. Natl. Cancer Inst. 2004, 96, 269–279. [Google Scholar] [CrossRef] [PubMed]
- Iwanami, A.; Gini, B.; Zanca, C.; Matsutani, T.; Assuncao, A.; Nael, A.; Dang, J.; Yang, H.; Zhu, S.; Kohyama, J.; et al. PML mediates glioblastoma resistance to mammalian target of rapamycin (mTOR)-targeted therapies. Proc. Natl. Acad. Sci. USA 2013, 110, 4339–4344. [Google Scholar] [CrossRef] [PubMed]
- Mazza, M.; Pelicci, P.G. Is PML a Tumor Suppressor? Front. Oncol. 2013, 3, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Roniotis, A.; Marias, K.; Sakkalis, V.; Tsibidis, G.D.; Zervakis, M. A complete mathematical study of a 3D model of heterogeneous and anisotropic glioma evolution. In Proceedings of the 2009 Annual International Conference of the IEEE Engineering in Medicine and Biology Society, Minneapolis, MN, USA, 3–6 September 2009; pp. 2807–2810. [Google Scholar]
- Marias, K.; Sakkalis, V.; Roniotis, A.; Farmaki, C.; Stamatakos, G.; Dionysiou, D.; Giatili, S.; Uzunoglou, N.; Graf, N.; Bohle, R.; et al. Clinically Oriented Translational Cancer Multilevel Modeling: The ContraCancrum Project. In Proceedings of the World Congress on Medical Physics and Biomedical Engineering, Munich, Germany, 7–12 September 2009; Dössel, O., Schlegel, W.C., Eds.; Springer: Berlin/Heidelberg, Germany, 2010; pp. 2124–2127. [Google Scholar]
- Jagiella, N.; Müller, B.; Müller, M.; Vignon-Clementel, I.E.; Drasdo, D. Inferring Growth Control Mechanisms in Growing Multi-cellular Spheroids of NSCLC Cells from Spatial-Temporal Image Data. PLoS Comput. Biol. 2016, 12. [Google Scholar] [CrossRef]
- Oraiopoulou, M.-E.; Tzamali, E.; Tzedakis, G.; Liapis, E.; Zacharakis, G.; Vakis, A.; Papamatheakis, J.; Sakkalis, V. Integrating in vitro experiments with in silico approaches for Glioblastoma invasion: The role of cell-to-cell adhesion heterogeneity. Sci. Rep. 2018, 8, 16200. [Google Scholar] [CrossRef]
- Anderson, A.R.A. A hybrid mathematical model of solid tumour invasion: The importance of cell adhesion. Math. Med. Biol. J. IMA 2005, 22, 163–186. [Google Scholar] [CrossRef]
- Alfonso, J.C.L.; Talkenberger, K.; Seifert, M.; Klink, B.; Hawkins-Daarud, A.; Swanson, K.R.; Hatzikirou, H.; Deutsch, A. The biology and mathematical modelling of glioma invasion: A review. J. R. Soc. Interface 2017, 14. [Google Scholar] [CrossRef] [PubMed]
- Cortesi, M.; Liverani, C.; Mercatali, L.; Ibrahim, T.; Giordano, E. An in-silico study of cancer cell survival and spatial distribution within a 3D microenvironment. Sci. Rep. 2020, 10, 12976. [Google Scholar] [CrossRef]
- Frieboes, H.B.; Zheng, X.; Sun, C.-H.; Tromberg, B.; Gatenby, R.; Cristini, V. An Integrated Computational/Experimental Model of Tumor Invasion. Cancer Res. 2006, 66, 1597–1604. [Google Scholar] [CrossRef] [PubMed]
- Sachini, N.; Arampatzi, P.; Klonizakis, A.; Nikolaou, C.; Makatounakis, T.; Lam, E.W.F.; Kretsovali, A.; Papamatheakis, J. Promyelocytic leukemia protein (PML) controls breast cancer cell proliferation by modulating Forkhead transcription factors. Mol. Oncol. 2019, 13, 1369–1387. [Google Scholar] [CrossRef]
- Ivanschitz, L.; Takahashi, Y.; Jollivet, F.; Ayrault, O.; Le Bras, M.; de Thé, H. PML IV/ARF interaction enhances p53 SUMO-1 conjugation, activation, and senescence. Proc. Natl. Acad. Sci. USA 2015, 112, 14278–14283. [Google Scholar] [CrossRef]
- Nisole, S.; Maroui, M.A.; Mascle, X.H.; Aubry, M.; Chelbi-Alix, M.K. Differential Roles of PML Isoforms. Front. Oncol. 2013, 3, 1–17. [Google Scholar] [CrossRef]
- Oraiopoulou, M.-E.; Tzamali, E.; Tzedakis, G.; Vakis, A.; Papamatheakis, J.; Sakkalis, V. In Vitro/In Silico Study on the Role of Doubling Time Heterogeneity among Primary Glioblastoma Cell Lines. BioMed Res. Int. 2017, 2017, 8569328. [Google Scholar] [CrossRef] [PubMed]
- Larson, B. BioTek White Paper—3D Cell Culture: A Review of Current Techniques. 2015. Available online: https://www.biotek.com/resources/white-papers/3d-cell-culture-a-review-of-current-techniques/ (accessed on 9 June 2021).
- Foty, R. A Simple Hanging Drop Cell Culture Protocol for Generation of 3D Spheroids. J. Vis. Exp. 2011, 20, 4–7. [Google Scholar] [CrossRef]
- Leung, B.M.; Lesher-Perez, S.C.; Matsuoka, T.; Moraes, C.; Takayama, S. Media additives to promote spheroid circularity and compactness in hanging drop platform. Biomater. Sci. 2015, 3, 336–344. [Google Scholar] [CrossRef]
- Cao, Q.; Yu, J.; Dhanasekaran, S.M.; Kim, J.H.; Mani, R.-S.; Tomlins, S.A.; Mehra, R.; Laxman, B.; Cao, X.; Yu, J.; et al. Repression of E-cadherin by the polycomb group protein EZH2 in cancer. Oncogene 2008, 27, 7274–7284. [Google Scholar] [CrossRef] [PubMed]
- De Rosa, J.; Iraburu, M.; Gallo-oller, G.; Shahi, M.H.; Meléndez, B.; Rey, J.A.; Idoate, M.A.; Castresana, J.S. EZH2 as a Therapeutic Target in Glioblastoma: A Cellular and Molecular Study Carcinogenesis Car cinogenesis & Mutagenesis. Clin. Res. 2016, 7, 6–13. [Google Scholar] [CrossRef]
- Anderson, A.R.A.; Rejniak, K.A.; Gerlee, P.; Quaranta, V. Modelling of cancer growth, evolution and invasion: Bridging scales and models. Math. Model. Nat. Phenom. 2007, 2, 1–29. [Google Scholar] [CrossRef]
- Deisboeck, T.S.; Zhang, L.; Yoon, J.; Costa, J. In silico cancer modeling: Is it ready for prime time? Nat. Clin. Pract. Oncol. 2009, 6, 34–42. [Google Scholar] [CrossRef] [PubMed]
- Gallaher, J.A.; Massey, S.C.; Hawkins-Daarud, A.; Noticewala, S.S.; Rockne, R.C.; Johnston, S.K.; Gonzalez-Cuyar, L.; Juliano, J.; Gil, O.; Swanson, K.R.; et al. From cells to tissue: How cell scale heterogeneity impacts glioblastoma growth and treatment response. PLoS Comput. Biol. 2020, 16, e1007672. [Google Scholar] [CrossRef]
- Alexeyev, M.F.; Fayzulin, R.; Shokolenko, I.N.; Pastukh, V. A retro-lentiviral system for doxycycline-inducible gene expression and gene knockdown in cells with limited proliferative capacity. Mol. Biol. Rep. 2010, 37, 1987–1991. [Google Scholar] [CrossRef]
- Oraiopoulou, M.-E.; Tampakaki, M.; Tzamali, E.; Tamiolakis, T.; Makatounakis, V.; Vakis, A.F.; Zacharakis, G.; Sakkalis, V.; Papamatheakis, J. A 3D tumor spheroid model for the T98G Glioblastoma cell line phenotypic characterization. Tissue Cell 2019, 59, 39–43. [Google Scholar] [CrossRef] [PubMed]
- Tzamali, E.; Tzedakis, G.; Sakkalis, V. Modeling How Heterogeneity in Cell Cycle Length Affects Cancer Cell Growth Dynamics in Response to Treatment. Front. Oncol. 2020, 10, 1552. [Google Scholar] [CrossRef]


| Parameter | U87MG | U87MG-PML OE |
|---|---|---|
| Proliferation Time | 100 h | 160 h |
| Cell Size | 21.5 μm | 21.5 μm |
| Initial Cell Density | 90% | 90% |
| Random Cell Death Rate | 20% of the proliferation time | 20% of the proliferation time |
| DZNeP-Mediated Cell Death Rate | 0.0065 h−1 | 0.0065 h−1 |
| Parameter | U87MG | U87MG-PML OE |
|---|---|---|
| Diffusion Coefficient | 2.00 × 10−9 cm2/s | 2.00 × 10−9 cm2/s |
| Motile:Adhesive Phenotypic Ratio | 2:1 | 2:1 |
| Diffusion Coefficient upon DZNeP Treatment | 2.00 × 10−9 cm2/s | 1.00 × 10−9 cm2/s |
| Motile:Adhesive Phenotypic Ratio upon DZNeP Treatment | 1:1 | 1:4 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Tampakaki, M.; Oraiopoulou, M.-E.; Tzamali, E.; Tzedakis, G.; Makatounakis, T.; Zacharakis, G.; Papamatheakis, J.; Sakkalis, V. PML Differentially Regulates Growth and Invasion in Brain Cancer. Int. J. Mol. Sci. 2021, 22, 6289. https://doi.org/10.3390/ijms22126289
Tampakaki M, Oraiopoulou M-E, Tzamali E, Tzedakis G, Makatounakis T, Zacharakis G, Papamatheakis J, Sakkalis V. PML Differentially Regulates Growth and Invasion in Brain Cancer. International Journal of Molecular Sciences. 2021; 22(12):6289. https://doi.org/10.3390/ijms22126289
Chicago/Turabian StyleTampakaki, Maria, Mariam-Eleni Oraiopoulou, Eleftheria Tzamali, Giorgos Tzedakis, Takis Makatounakis, Giannis Zacharakis, Joseph Papamatheakis, and Vangelis Sakkalis. 2021. "PML Differentially Regulates Growth and Invasion in Brain Cancer" International Journal of Molecular Sciences 22, no. 12: 6289. https://doi.org/10.3390/ijms22126289
APA StyleTampakaki, M., Oraiopoulou, M.-E., Tzamali, E., Tzedakis, G., Makatounakis, T., Zacharakis, G., Papamatheakis, J., & Sakkalis, V. (2021). PML Differentially Regulates Growth and Invasion in Brain Cancer. International Journal of Molecular Sciences, 22(12), 6289. https://doi.org/10.3390/ijms22126289