Tumor Suppressive Effects of miR-124 and Its Function in Neuronal Development
Abstract
1. Introduction
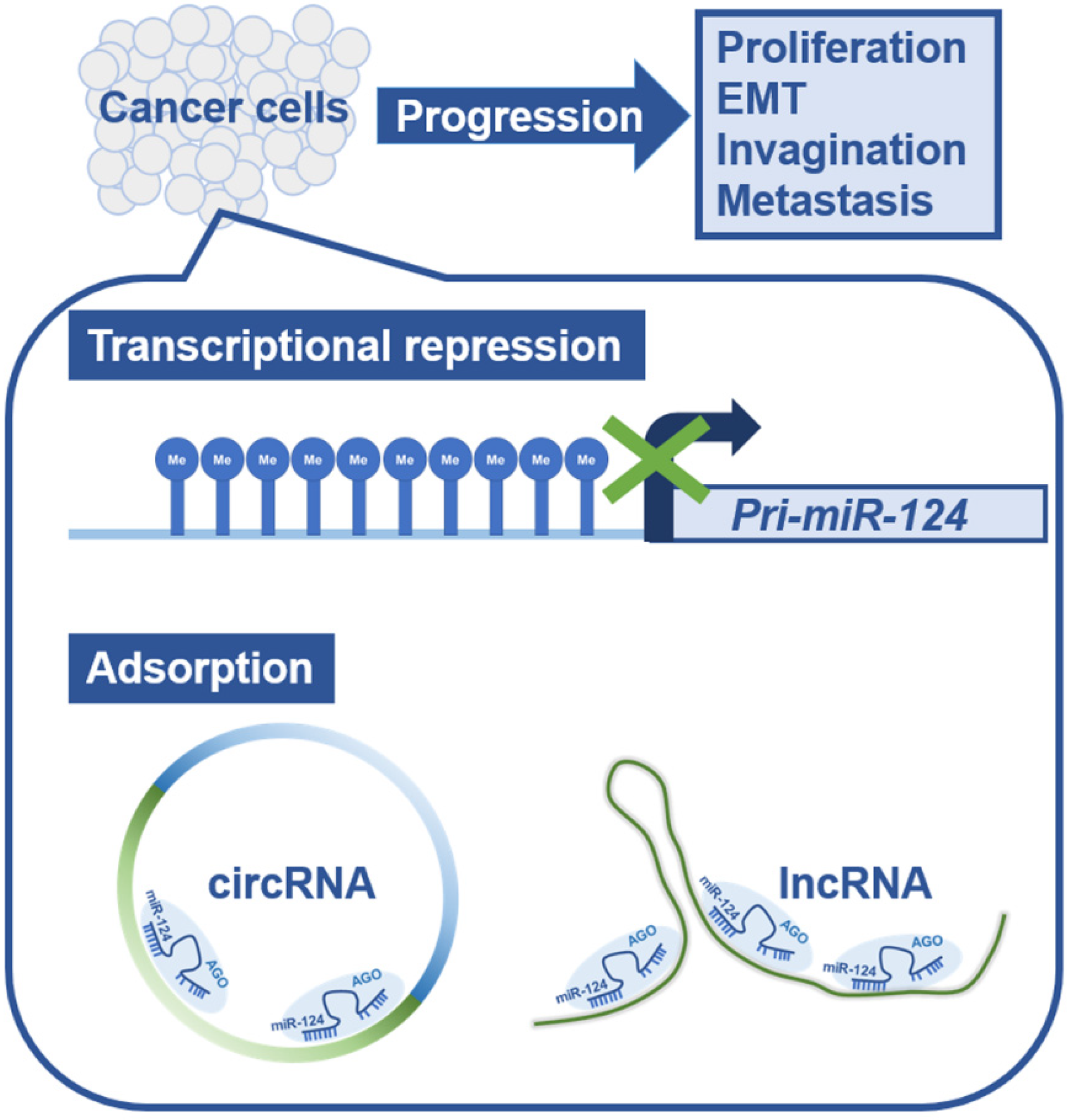
2. Suppression of miR-124 Functions in Cancer Cells
3. Tumor Suppressive Effects by miR-124 in Glial Lineage Cancers
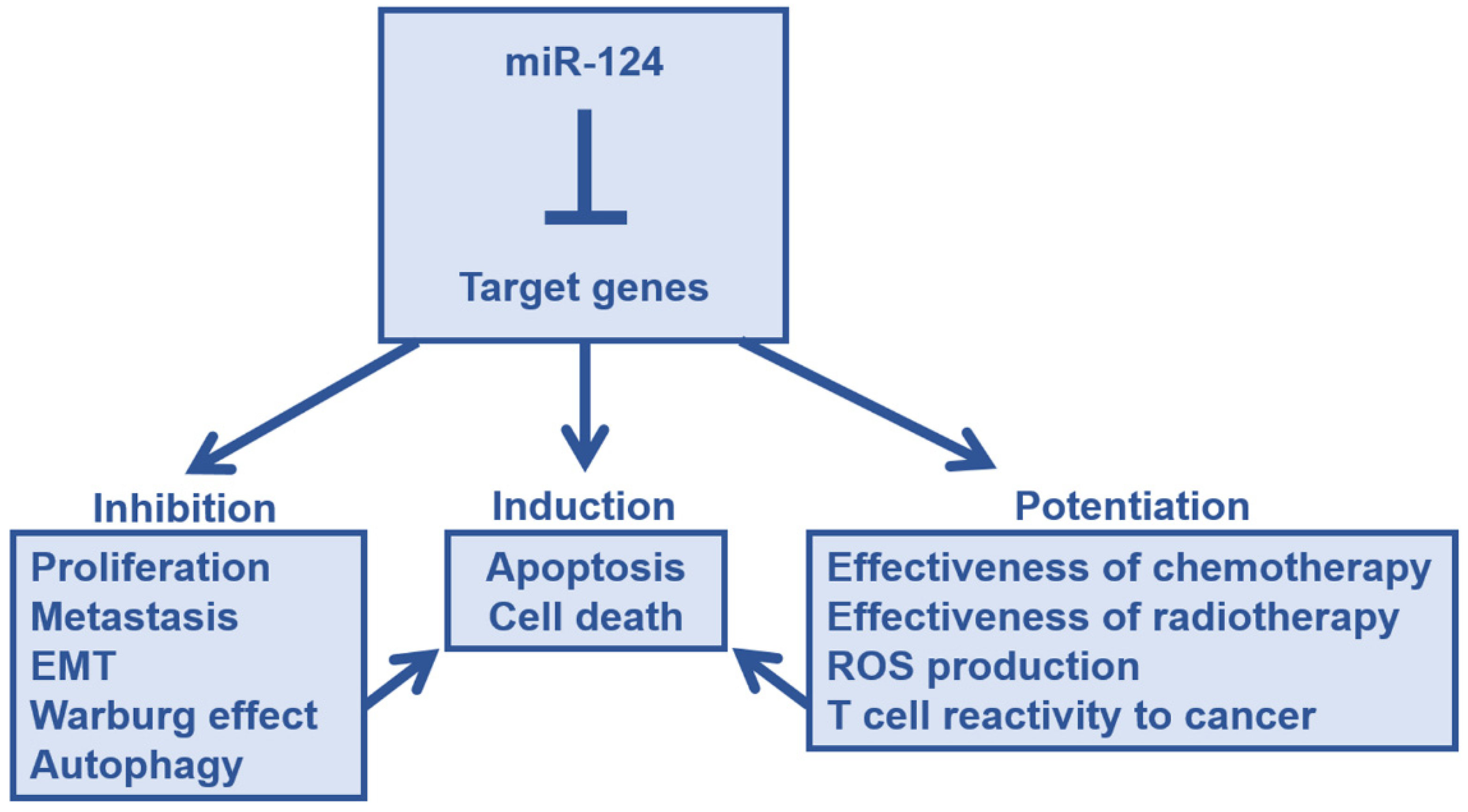
4. Tumor Suppressive Effects of miR-124 in Major Cancers
5. Target Genes Responsible for the Tumor Suppressor Effect of miR-124, STAT3, and EZH2
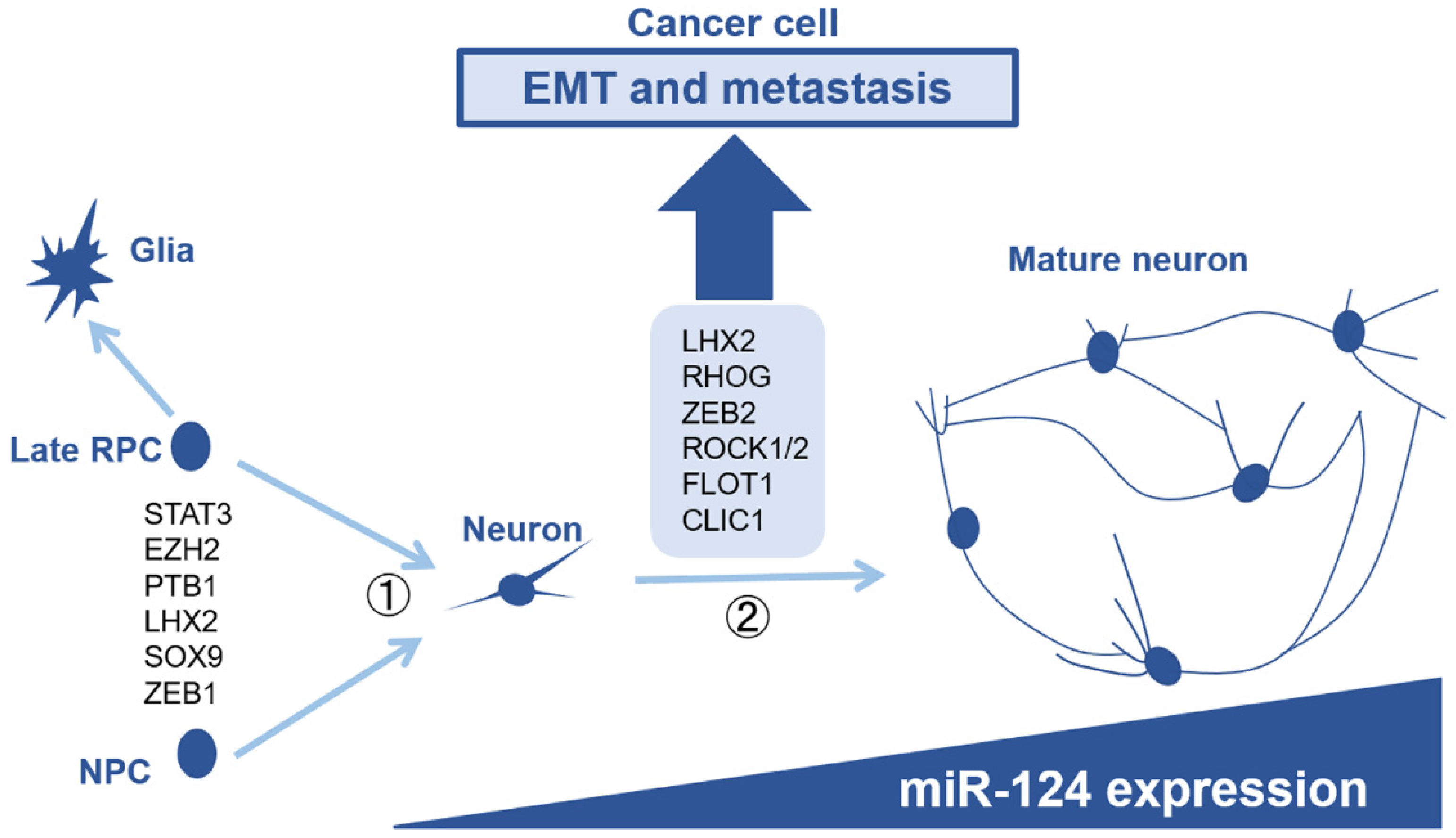
6. Comparison with Neuronal Development, the Original Function of miR-124
7. Discussion
8. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Lee, R.C.; Feinbaum, R.L.; Ambros, V. The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14. Cell 1993, 75, 843–854. [Google Scholar] [CrossRef]
- Wightman, B.; Ha, I.; Ruvkun, G. Posttranscriptional regulation of the heterochronic gene lin-14 by lin-4 mediates temporal pattern formation in C. elegans. Cell 1993, 75, 855–862. [Google Scholar] [CrossRef]
- Sammarco, G.; Gallo, G.; Vescio, G.; Picciariello, A.; De Paola, G.; Trompetto, M.; Curro, G.; Ammendola, M. Mast Cells, microRNAs and Others: The Role of Translational Research on Colorectal Cancer in the Forthcoming Era of Precision Medicine. J. Clin. Med. 2020, 9, 2852. [Google Scholar] [CrossRef]
- Shah, M.Y.; Ferrajoli, A.; Sood, A.K.; Lopez-Berestein, G.; Calin, G.A. microRNA Therapeutics in Cancer—An Emerging Concept. EBioMedicine 2016, 12, 34–42. [Google Scholar] [CrossRef] [PubMed]
- Danza, K.; Silvestris, N.; Simone, G.; Signorile, M.; Saragoni, L.; Brunetti, O.; Monti, M.; Mazzotta, A.; De Summa, S.; Mangia, A.; et al. Role of miR-27a, miR-181a and miR-20b in gastric cancer hypoxia-induced chemoresistance. Cancer Biol. Ther. 2016, 17, 400–406. [Google Scholar] [CrossRef] [PubMed]
- Gnoni, A.; Santini, D.; Scartozzi, M.; Russo, A.; Licchetta, A.; Palmieri, V.; Lupo, L.; Faloppi, L.; Palasciano, G.; Memeo, V.; et al. Hepatocellular carcinoma treatment over sorafenib: Epigenetics, microRNAs and microenvironment. Is there a light at the end of the tunnel? Expert Opin. Ther. Targets 2015, 19, 1623–1635. [Google Scholar] [CrossRef] [PubMed]
- Brunetti, O.; Russo, A.; Scarpa, A.; Santini, D.; Reni, M.; Bittoni, A.; Azzariti, A.; Aprile, G.; Delcuratolo, S.; Signorile, M.; et al. MicroRNA in pancreatic adenocarcinoma: Predictive/prognostic biomarkers or therapeutic targets? Oncotarget 2015, 6, 23323–23341. [Google Scholar] [CrossRef]
- Landgraf, P.; Rusu, M.; Sheridan, R.; Sewer, A.; Iovino, N.; Aravin, A.; Pfeffer, S.; Rice, A.; Kamphorst, A.O.; Landthaler, M.; et al. A mammalian microRNA expression atlas based on small RNA library sequencing. Cell 2007, 129, 1401–1414. [Google Scholar] [CrossRef] [PubMed]
- Sanuki, R.; Onishi, A.; Koike, C.; Muramatsu, R.; Watanabe, S.; Muranishi, Y.; Irie, S.; Uneo, S.; Koyasu, T.; Matsui, R.; et al. miR-124a is required for hippocampal axogenesis and retinal cone survival through Lhx2 suppression. Nat. Neurosci. 2011, 14, 1125–1134. [Google Scholar] [CrossRef] [PubMed]
- Kozuka, T.; Omori, Y.; Watanabe, S.; Tarusawa, E.; Yamamoto, H.; Chaya, T.; Furuhashi, M.; Morita, M.; Sato, T.; Hirose, S.; et al. miR-124 dosage regulates prefrontal cortex function by dopaminergic modulation. Sci. Rep. 2019, 9, 3445. [Google Scholar] [CrossRef]
- Lu, J.; Getz, G.; Miska, E.A.; Alvarez-Saavedra, E.; Lamb, J.; Peck, D.; Sweet-Cordero, A.; Ebert, B.L.; Mak, R.H.; Ferrando, A.A.; et al. MicroRNA expression profiles classify human cancers. Nature 2005, 435, 834–838. [Google Scholar] [CrossRef]
- Lujambio, A.; Ropero, S.; Ballestar, E.; Fraga, M.F.; Cerrato, C.; Setien, F.; Casado, S.; Suarez-Gauthier, A.; Sanchez-Cespedes, M.; Git, A.; et al. Genetic unmasking of an epigenetically silenced microRNA in human cancer cells. Cancer Res. 2007, 67, 1424–1429. [Google Scholar] [CrossRef] [PubMed]
- Grossel, M.J.; Hinds, P.W. Beyond the cell cycle: A new role for Cdk6 in differentiation. J. Cell Biochem. 2006, 97, 485–493. [Google Scholar] [CrossRef] [PubMed]
- Pierson, J.; Hostager, B.; Fan, R.; Vibhakar, R. Regulation of cyclin dependent kinase 6 by microRNA 124 in medulloblastoma. J. Neurooncol. 2008, 90, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Silber, J.; Lim, D.A.; Petritsch, C.; Persson, A.I.; Maunakea, A.K.; Yu, M.; Vandenberg, S.R.; Ginzinger, D.G.; James, C.D.; Costello, J.F.; et al. miR-124 and miR-137 inhibit proliferation of glioblastoma multiforme cells and induce differentiation of brain tumor stem cells. BMC Med. 2008, 6, 14. [Google Scholar] [CrossRef] [PubMed]
- Zheng, Q.; Bao, C.; Guo, W.; Li, S.; Chen, J.; Chen, B.; Luo, Y.; Lyu, D.; Li, Y.; Shi, G.; et al. Circular RNA profiling reveals an abundant circHIPK3 that regulates cell growth by sponging multiple miRNAs. Nat. Commun. 2016, 7, 11215. [Google Scholar] [CrossRef]
- Chen, G.; Shi, Y.; Zhang, Y.; Sun, J. CircRNA_100782 regulates pancreatic carcinoma proliferation through the IL6-STAT3 pathway. Onco Targets Ther. 2017, 10, 5783–5794. [Google Scholar] [CrossRef]
- Wang, R.; Zhang, S.; Chen, X.; Li, N.; Li, J.; Jia, R.; Pan, Y.; Liang, H. EIF4A3-induced circular RNA MMP9 (circMMP9) acts as a sponge of miR-124 and promotes glioblastoma multiforme cell tumorigenesis. Mol. Cancer 2018, 17, 166. [Google Scholar] [CrossRef]
- Liu, Y.Y.; Zhang, L.Y.; Du, W.Z. Circular RNA circ-PVT1 contributes to paclitaxel resistance of gastric cancer cells through the regulation of ZEB1 expression by sponging miR-124-3p. Biosci. Rep. 2019, 39, BSR20193045. [Google Scholar] [CrossRef]
- Zhang, S.; Liu, Y.; Liu, Z.; Zhang, Y.; Chen, G.; Li, K.; Tang, K. CircRNA_0000502 promotes hepatocellular carcinoma metastasis and inhibits apoptosis through targeting microRNA-124. J. BUON 2019, 24, 2402–2410. [Google Scholar]
- Cui, W.; Xue, J. Circular RNA DOCK1 downregulates microRNA-124 to induce the growth of human thyroid cancer cell lines. Biofactors 2020, 46, 591–599. [Google Scholar] [CrossRef]
- Sha, J.; Xia, L.; Han, Q.; Chi, C.; Zhu, Y.; Pan, J.; Huang, Y.; Xia, W.; Dong, B.; Xue, W.; et al. Downregulation of circ-TRPS1 suppressed prostatic cancer prognoses by regulating miR-124-3p/EZH2 axis-mediated stemness. Am. J. Cancer Res. 2020, 10, 4372–4385. [Google Scholar] [PubMed]
- Yang, X.; Wang, J.; Li, H.; Sun, Y.; Tong, X. Downregulation of hsa_circ_0026123 suppresses ovarian cancer cell metastasis and proliferation through the miR-124-3p/EZH2 signaling pathway. Int. J. Mol. Med. 2021, 47, 668–676. [Google Scholar] [CrossRef] [PubMed]
- Chen, G.; Shi, Y.; Liu, M.; Sun, J. circHIPK3 regulates cell proliferation and migration by sponging miR-124 and regulating AQP3 expression in hepatocellular carcinoma. Cell Death Dis. 2018, 9, 175. [Google Scholar] [CrossRef] [PubMed]
- Hu, D.; Zhang, Y. Circular RNA HIPK3 promotes glioma progression by binding to miR-124-3p. Gene 2019, 690, 81–89. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.; Guo, S.; Sun, H.; Bai, Y.; Song, Z.; Liu, X. Circular RNA CircHIPK3 Elevates CCND2 Expression and Promotes Cell Proliferation and Invasion Through miR-124 in Glioma. Front. Genet. 2020, 11, 1013. [Google Scholar] [CrossRef]
- Xia, L.; Yi, F.; Zhai, X.; Zhang, M. Circular RNA homeodomain-interacting protein kinase 3 (circHIPK3) promotes growth and metastasis of glioma cells by sponging miR-124-3p. Xi Bao Yu Fen Zi Mian Yi Xue Za Zhi 2020, 36, 609–615. [Google Scholar] [PubMed]
- Yu, H.; Chen, Y.; Jiang, P. Circular RNA HIPK3 exerts oncogenic properties through suppression of miR-124 in lung cancer. Biochem. Biophys. Res. Commun. 2018, 506, 455–462. [Google Scholar] [CrossRef]
- Cheng, J.; Zhuo, H.; Xu, M.; Wang, L.; Xu, H.; Peng, J.; Hou, J.; Lin, L.; Cai, J. Regulatory network of circRNA-miRNA-mRNA contributes to the histological classification and disease progression in gastric cancer. J. Transl. Med. 2018, 16, 216. [Google Scholar] [CrossRef]
- Kai, D.; Yannian, L.; Yitian, C.; Dinghao, G.; Xin, Z.; Wu, J. Circular RNA HIPK3 promotes gallbladder cancer cell growth by sponging microRNA-124. Biochem. Biophys. Res. Commun. 2018, 503, 863–869. [Google Scholar] [CrossRef]
- Wang, J.; Zhao, S.Y.; Ouyang, S.S.; Huang, Z.K.; Luo, Q.; Liao, L. Circular RNA circHIPK3 acts as the sponge of microRNA-124 to promote human oral squamous cell carcinoma cells proliferation. Zhonghua Kou Qiang Yi Xue Za Zhi 2018, 53, 546–551. [Google Scholar] [PubMed]
- Li, S.; Mei, Z.; Hu, H.B.; Zhang, X. The lncRNA MALAT1 contributes to non-small cell lung cancer development via modulating miR-124/STAT3 axis. J. Cell Physiol. 2018, 233, 6679–6688. [Google Scholar] [CrossRef] [PubMed]
- Wu, J.; Weng, Y.; He, F.; Liang, D.; Cai, L. LncRNA MALAT-1 competitively regulates miR-124 to promote EMT and development of non-small-cell lung cancer. Anticancer Drugs 2018, 29, 628–636. [Google Scholar] [CrossRef] [PubMed]
- Liu, S.; Song, L.; Zeng, S.; Zhang, L. MALAT1-miR-124-RBG2 axis is involved in growth and invasion of HR-HPV-positive cervical cancer cells. Tumour Biol. 2016, 37, 633–640. [Google Scholar] [CrossRef]
- Shi, B.; Wang, Y.; Yin, F. MALAT1/miR-124/Capn4 axis regulates proliferation, invasion and EMT in nasopharyngeal carcinoma cells. Cancer Biol. Ther. 2017, 18, 792–800. [Google Scholar] [CrossRef] [PubMed]
- Li, D.; Chen, P.; Li, X.Y.; Zhang, L.Y.; Xiong, W.; Zhou, M.; Xiao, L.; Zeng, F.; Li, X.L.; Wu, M.H.; et al. Grade-specific expression profiles of miRNAs/mRNAs and docking study in human grade I-III astrocytomas. OMICS 2011, 15, 673–682. [Google Scholar] [CrossRef]
- Lv, Z.; Yang, L. MiR-124 inhibits the growth of glioblastoma through the downregulation of SOS1. Mol. Med. Rep. 2013, 8, 345–349. [Google Scholar] [CrossRef]
- Silber, J.; Hashizume, R.; Felix, T.; Hariono, S.; Yu, M.; Berger, M.S.; Huse, J.T.; VandenBerg, S.R.; James, C.D.; Hodgson, J.G.; et al. Expression of miR-124 inhibits growth of medulloblastoma cells. Neuro-oncology 2013, 15, 83–90. [Google Scholar] [CrossRef]
- Wei, J.; Wang, F.; Kong, L.Y.; Xu, S.; Doucette, T.; Ferguson, S.D.; Yang, Y.; McEnery, K.; Jethwa, K.; Gjyshi, O.; et al. miR-124 inhibits STAT3 signaling to enhance T cell-mediated immune clearance of glioma. Cancer Res. 2013, 73, 3913–3926. [Google Scholar] [CrossRef]
- Zhao, W.H.; Wu, S.Q.; Zhang, Y.D. Downregulation of miR-124 promotes the growth and invasiveness of glioblastoma cells involving upregulation of PPP1R13L. Int. J. Mol. Med. 2013, 32, 101–107. [Google Scholar] [CrossRef]
- Mucaj, V.; Lee, S.S.; Skuli, N.; Giannoukos, D.N.; Qiu, B.; Eisinger-Mathason, T.S.; Nakazawa, M.S.; Shay, J.E.; Gopal, P.P.; Venneti, S.; et al. MicroRNA-124 expression counteracts pro-survival stress responses in glioblastoma. Oncogene 2015, 34, 2204–2214. [Google Scholar] [CrossRef]
- Cai, J.J.; Qi, Z.X.; Chen, L.C.; Yao, Y.; Gong, Y.; Mao, Y. miR-124 suppresses the migration and invasion of glioma cells in vitro via Capn4. Oncol. Rep. 2016, 35, 284–290. [Google Scholar] [CrossRef]
- Deng, D.; Wang, L.; Chen, Y.; Li, B.; Xue, L.; Shao, N.; Wang, Q.; Xia, X.; Yang, Y.; Zhi, F. MicroRNA-124-3p regulates cell proliferation, invasion, apoptosis, and bioenergetics by targeting PIM1 in astrocytoma. Cancer Sci. 2016, 107, 899–907. [Google Scholar] [CrossRef]
- Marisetty, A.L.; Singh, S.K.; Nguyen, T.N.; Coarfa, C.; Liu, B.; Majumder, S. REST represses miR-124 and miR-203 to regulate distinct oncogenic properties of glioblastoma stem cells. Neuro-oncology 2017, 19, 514–523. [Google Scholar] [CrossRef] [PubMed]
- Qiao, W.; Guo, B.; Zhou, H.; Xu, W.; Chen, Y.; Liang, Y.; Dong, B. miR-124 suppresses glioblastoma growth and potentiates chemosensitivity by inhibiting AURKA. Biochem. Biophys. Res. Commun. 2017, 486, 43–48. [Google Scholar] [CrossRef] [PubMed]
- Lin, J.; Wen, X.; Zhang, X.; Sun, X.; Yunzhi, L.; Peng, R.; Zhu, M.; Wang, M.; Zhang, Y.; Luo, W.; et al. miR-135a-5p and miR-124-3p Inhibit Malignancy of Glioblastoma by Downregulation of Syndecan Binding Protein. J. Biomed. Nanotechnol. 2018, 14, 1317–1329. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Kang, J.; Sun, S.; Luo, Y.; Ji, X.; Zeng, X.; Zhao, S. iASPP, a microRNA124 target, is aberrantly expressed in astrocytoma and regulates malignant glioma cell migration and viability. Mol. Med. Rep. 2018, 17, 1970–1978. [Google Scholar] [PubMed]
- Sharif, S.; Ghahremani, M.H.; Soleimani, M. Delivery of Exogenous miR-124 to Glioblastoma Multiform Cells by Wharton’s Jelly Mesenchymal Stem Cells Decreases Cell Proliferation and Migration, and Confers Chemosensitivity. Stem Cell Rev. Rep. 2018, 14, 236–246. [Google Scholar] [CrossRef]
- Zhang, G.; Chen, L.; Khan, A.A.; Li, B.; Gu, B.; Lin, F.; Su, X.; Yan, J. miRNA-124-3p/neuropilin-1(NRP-1) axis plays an important role in mediating glioblastoma growth and angiogenesis. Int. J. Cancer 2018, 143, 635–644. [Google Scholar] [CrossRef]
- Bhaskaran, V.; Nowicki, M.O.; Idriss, M.; Jimenez, M.A.; Lugli, G.; Hayes, J.L.; Mahmoud, A.B.; Zane, R.E.; Passaro, C.; Ligon, K.L.; et al. The functional synergism of microRNA clustering provides therapeutically relevant epigenetic interference in glioblastoma. Nat. Commun. 2019, 10, 442. [Google Scholar] [CrossRef]
- Deng, X.; Ma, L.; Wu, M.; Zhang, G.; Jin, C.; Guo, Y.; Liu, R. miR-124 radiosensitizes human glioma cells by targeting CDK4. J. Neurooncol. 2013, 114, 263–274. [Google Scholar] [CrossRef] [PubMed]
- Hirose, Y.; Ohb, S. Biological Responses of Glioma Cell to Chemotherapeutic Agents. In Advances in the Biology, Imaging and Therapies for Glioblastoma; IntechOpen: London, UK, 2011; pp. 61–76. ISBN 978-953-307-284-5. [Google Scholar]
- Kumar, M.S.; Lu, J.; Mercer, K.L.; Golub, T.R.; Jacks, T. Impaired microRNA processing enhances cellular transformation and tumorigenesis. Nat. Genet. 2007, 39, 673–677. [Google Scholar] [CrossRef]
- Wang, D.; Zhang, H.; Li, M.; Frid, M.G.; Flockton, A.R.; McKeon, B.A.; Yeager, M.E.; Fini, M.A.; Morrell, N.W.; Pullamsetti, S.S.; et al. MicroRNA-124 controls the proliferative, migratory, and inflammatory phenotype of pulmonary vascular fibroblasts. Circ. Res. 2014, 114, 67–78. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Yu, Z.; Li, Y.; Liu, S.; Gao, C.; Hou, X.; Yao, R.; Cui, L. The tumor suppressor miR-124 inhibits cell proliferation by targeting STAT3 and functions as a prognostic marker for postoperative NSCLC patients. Int. J. Oncol. 2015, 46, 798–808. [Google Scholar] [CrossRef] [PubMed]
- Cui, Z.; Hu, Y. MicroRNA-124 suppresses Slug-mediated lung cancer metastasis. Eur. Rev. Med. Pharmacol. Sci. 2016, 20, 3802–3811. [Google Scholar]
- Li, Z.; Wang, X.; Li, W.; Wu, L.; Chang, L.; Chen, H. miRNA-124 modulates lung carcinoma cell migration and invasion. Int. J. Clin. Pharmacol. Ther. 2016, 54, 603–612. [Google Scholar] [CrossRef] [PubMed]
- Lin, J.; Xu, K.; Wei, J.; Heimberger, A.B.; Roth, J.A.; Ji, L. MicroRNA-124 suppresses tumor cell proliferation and invasion by targeting CD164 signaling pathway in non-small cell lung cancer. J. Gene. Ther. 2016, 2, 6. [Google Scholar]
- Mehta, A.K.; Hua, K.; Whipple, W.; Nguyen, M.T.; Liu, C.T.; Haybaeck, J.; Weidhaas, J.; Settleman, J.; Singh, A. Regulation of autophagy, NF-kappaB signaling, and cell viability by miR-124 in KRAS mutant mesenchymal-like NSCLC cells. Sci. Signal. 2017, 10, eaam6291. [Google Scholar] [CrossRef]
- Zhao, X.; Lu, C.; Chu, W.; Zhang, B.; Zhen, Q.; Wang, R.; Zhang, Y.; Li, Z.; Lv, B.; Li, H.; et al. MicroRNA-124 suppresses proliferation and glycolysis in non-small cell lung cancer cells by targeting AKT-GLUT1/HKII. Tumour Biol. 2017, 39, 1010428317706215. [Google Scholar] [CrossRef]
- Sun, Y.; Ai, X.; Shen, S.; Lu, S. NF-kappaB-mediated miR-124 suppresses metastasis of non-small-cell lung cancer by targeting MYO10. Oncotarget 2015, 6, 8244–8254. [Google Scholar] [CrossRef]
- Hao, C.; Xu, X.; Ma, J.; Xia, J.; Dai, B.; Liu, L.; Ma, Y. MicroRNA-124 regulates the radiosensitivity of non-small cell lung cancer cells by targeting TXNRD1. Oncol. Lett. 2017, 13, 2071–2078. [Google Scholar] [CrossRef]
- Wang, M.; Meng, B.; Liu, Y.; Yu, J.; Chen, Q.; Liu, Y. MiR-124 Inhibits Growth and Enhances Radiation-Induced Apoptosis in Non-Small Cell Lung Cancer by Inhibiting STAT3. Cell Physiol. Biochem. 2017, 44, 2017–2028. [Google Scholar] [CrossRef]
- Yang, Q.; Wan, L.; Xiao, C.; Hu, H.; Wang, L.; Zhao, J.; Lei, Z.; Zhang, H.T. Inhibition of LHX2 by miR-124 suppresses cellular migration and invasion in non-small cell lung cancer. Oncol. Lett. 2017, 14, 3429–3436. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Qi, M.M.; Ge, F.; Chen, X.J.; Tang, C.; Ma, J. MiR-124 changes the sensitivity of lung cancer cells to cisplatin through targeting STAT3. Eur. Rev. Med. Pharmacol. Sci. 2019, 23, 5242–5250. [Google Scholar]
- Liu, T.; Zhu, J.; Du, W.; Ning, W.; Zhang, Y.; Zeng, Y.; Liu, Z.; Huang, J.A. AKT2 drives cancer progression and is negatively modulated by miR-124 in human lung adenocarcinoma. Respir. Res. 2020, 21, 227. [Google Scholar] [CrossRef] [PubMed]
- Wu, J.; Li, L.; Zhang, Y.; Zhu, J. Decreased miR-124 contributes to the epithelial-mesenchymal transition phenotype formation of lung adenocarcinoma cells via targeting enhancer of zeste homolog 2. Pathol. Res. Pract. 2020, 216, 152976. [Google Scholar] [CrossRef] [PubMed]
- Ma, T.; Zhao, Y.; Wei, K.; Yao, G.; Pan, C.; Liu, B.; Xia, Y.; He, Z.; Qi, X.; Li, Z.; et al. MicroRNA-124 Functions as a Tumor Suppressor by Regulating CDH2 and Epithelial-Mesenchymal Transition in Non-Small Cell Lung Cancer. Cell Physiol. Biochem. 2016, 38, 1563–1574. [Google Scholar] [CrossRef]
- Park, S.Y.; Kim, H.; Yoon, S.; Bae, J.A.; Choi, S.Y.; Jung, Y.D.; Kim, K.K. KITENIN-targeting microRNA-124 suppresses colorectal cancer cell motility and tumorigenesis. Mol. Ther. 2014, 22, 1653–1664. [Google Scholar] [CrossRef]
- Zhou, L.; Xu, Z.; Ren, X.; Chen, K.; Xin, S. MicroRNA-124 (MiR-124) Inhibits Cell Proliferation, Metastasis and Invasion in Colorectal Cancer by Downregulating Rho-Associated Protein Kinase 1(ROCK1). Cell Physiol. Biochem. 2016, 38, 1785–1795. [Google Scholar] [CrossRef] [PubMed]
- Liu, K.; Yao, H.; Lei, S.; Xiong, L.; Qi, H.; Qian, K.; Liu, J.; Wang, P.; Zhao, H. The miR-124-p63 feedback loop modulates colorectal cancer growth. Oncotarget 2017, 8, 29101–29115. [Google Scholar] [CrossRef] [PubMed]
- Taniguchi, K.; Sugito, N.; Kumazaki, M.; Shinohara, H.; Yamada, N.; Nakagawa, Y.; Ito, Y.; Otsuki, Y.; Uno, B.; Uchiyama, K.; et al. MicroRNA-124 inhibits cancer cell growth through PTB1/PKM1/PKM2 feedback cascade in colorectal cancer. Cancer Lett. 2015, 363, 17–27. [Google Scholar] [CrossRef] [PubMed]
- Gatenby, R.A.; Gillies, R.J. Why do cancers have high aerobic glycolysis? Nat. Rev. Cancer 2004, 4, 891–899. [Google Scholar] [CrossRef] [PubMed]
- Taniguchi, K.; Sugito, N.; Kumazaki, M.; Shinohara, H.; Yamada, N.; Matsuhashi, N.; Futamura, M.; Ito, Y.; Otsuki, Y.; Yoshida, K.; et al. Positive feedback of DDX6/c-Myc/PTB1 regulated by miR-124 contributes to maintenance of the Warburg effect in colon cancer cells. Biochim. Biophys. Acta 2015, 1852, 1971–1980. [Google Scholar] [CrossRef]
- Li, M.; Wan, G.; Yu, H.; Xiong, W. High-intensity focused ultrasound inhibits invasion and metastasis of colon cancer cells by enhancing microRNA-124-mediated suppression of STAT3. FEBS Open Bio 2019, 9, 1128–1136. [Google Scholar] [CrossRef]
- Zhang, J.; Lu, Y.; Yue, X.; Li, H.; Luo, X.; Wang, Y.; Wang, K.; Wan, J. MiR-124 suppresses growth of human colorectal cancer by inhibiting STAT3. PLoS ONE 2013, 8, e70300. [Google Scholar] [CrossRef] [PubMed]
- Furuta, M.; Kozaki, K.I.; Tanaka, S.; Arii, S.; Imoto, I.; Inazawa, J. miR-124 and miR-203 are epigenetically silenced tumor-suppressive microRNAs in hepatocellular carcinoma. Carcinogenesis 2010, 31, 766–776. [Google Scholar] [CrossRef]
- Lang, Q.; Ling, C. MiR-124 suppresses cell proliferation in hepatocellular carcinoma by targeting PIK3CA. Biochem. Biophys. Res. Commun. 2012, 426, 247–252. [Google Scholar] [CrossRef] [PubMed]
- Zheng, F.; Liao, Y.J.; Cai, M.Y.; Liu, Y.H.; Liu, T.H.; Chen, S.P.; Bian, X.W.; Guan, X.Y.; Lin, M.C.; Zeng, Y.X.; et al. The putative tumour suppressor microRNA-124 modulates hepatocellular carcinoma cell aggressiveness by repressing ROCK2 and EZH2. Gut 2012, 61, 278–289. [Google Scholar] [CrossRef]
- Lu, Y.; Yue, X.; Cui, Y.; Zhang, J.; Wang, K. MicroRNA-124 suppresses growth of human hepatocellular carcinoma by targeting STAT3. Biochem. Biophys. Res. Commun. 2013, 441, 873–879. [Google Scholar] [CrossRef]
- Xu, L.; Dai, W.; Li, J.; He, L.; Wang, F.; Xia, Y.; Chen, K.; Li, S.; Liu, T.; Lu, J.; et al. Methylation-regulated miR-124-1 suppresses tumorigenesis in hepatocellular carcinoma by targeting CASC3. Oncotarget 2016, 7, 26027–26041. [Google Scholar] [CrossRef]
- Cai, Q.Q.; Dong, Y.W.; Wang, R.; Qi, B.; Guo, J.X.; Pan, J.; Liu, Y.Y.; Zhang, C.Y.; Wu, X.Z. MiR-124 inhibits the migration and invasion of human hepatocellular carcinoma cells by suppressing integrin alphaV expression. Sci. Rep. 2017, 7, 40733. [Google Scholar] [CrossRef] [PubMed]
- Yue, X.; Cui, Y.; You, Q.; Lu, Y.; Zhang, J. MicroRNA124 negatively regulates chloride intracellular channel 1 to suppress the migration and invasion of liver cancer cells. Oncol. Rep. 2019, 42, 1380–1390. [Google Scholar] [PubMed]
- Xia, J.; Wu, Z.; Yu, C.; He, W.; Zheng, H.; He, Y.; Jian, W.; Chen, L.; Zhang, L.; Li, W. miR-124 inhibits cell proliferation in gastric cancer through down-regulation of SPHK1. J. Pathol. 2012, 227, 470–480. [Google Scholar] [CrossRef] [PubMed]
- Xie, L.M.; He, R.F.; Zhang, Z.W.; Tang, Y.Y.; Luo, Z.Y. miR-124 suppresses cell proliferation and invasion in gastric carcinoma and its mechanism. Zhonghua Zhong Liu Za Zhi 2013, 35, 497–500. [Google Scholar] [PubMed]
- Hu, C.B.; Li, Q.L.; Hu, J.F.; Zhang, Q.; Xie, J.P.; Deng, L. miR-124 inhibits growth and invasion of gastric cancer by targeting ROCK1. Asian Pac. J. Cancer Prev. 2014, 15, 6543–6546. [Google Scholar] [CrossRef]
- Xie, L.; Zhang, Z.; Tan, Z.; He, R.; Zeng, X.; Xie, Y.; Li, S.; Tang, G.; Tang, H.; He, X. MicroRNA-124 inhibits proliferation and induces apoptosis by directly repressing EZH2 in gastric cancer. Mol. Cell Biochem. 2014, 392, 153–159. [Google Scholar] [CrossRef]
- Pan, Y.; Wu, A.; Xu, F.; Chen, C.; Jiang, L.; Jin, R. Lentivirus-mediated overexpression of miR-124 suppresses growth and invasion by targeting JAG1 and EZH2 in gastric cancer. Oncol. Lett. 2018, 15, 7450–7458. [Google Scholar] [CrossRef]
- Lv, X.B.; Jiao, Y.; Qing, Y.; Hu, H.; Cui, X.; Lin, T.; Song, E.; Yu, F. miR-124 suppresses multiple steps of breast cancer metastasis by targeting a cohort of pro-metastatic genes in vitro. Chin. J. Cancer 2011, 30, 821–830. [Google Scholar] [CrossRef]
- Han, Z.B.; Yang, Z.; Chi, Y.; Zhang, L.; Wang, Y.; Ji, Y.; Wang, J.; Zhao, H.; Han, Z.C. MicroRNA-124 suppresses breast cancer cell growth and motility by targeting CD151. Cell Physiol. Biochem. 2013, 31, 823–832. [Google Scholar] [CrossRef]
- Li, L.; Luo, J.; Wang, B.; Wang, D.; Xie, X.; Yuan, L.; Guo, J.; Xi, S.; Gao, J.; Lin, X.; et al. Microrna-124 targets flotillin-1 to regulate proliferation and migration in breast cancer. Mol. Cancer 2013, 12, 163. [Google Scholar] [CrossRef] [PubMed]
- Liang, Y.J.; Wang, Q.Y.; Zhou, C.X.; Yin, Q.Q.; He, M.; Yu, X.T.; Cao, D.X.; Chen, G.Q.; He, J.R.; Zhao, Q. MiR-124 targets Slug to regulate epithelial-mesenchymal transition and metastasis of breast cancer. Carcinogenesis 2013, 34, 713–722. [Google Scholar] [CrossRef]
- Feng, T.; Xu, D.; Tu, C.; Li, W.; Ning, Y.; Ding, J.; Wang, S.; Yuan, L.; Xu, N.; Qian, K.; et al. MiR-124 inhibits cell proliferation in breast cancer through downregulation of CDK4. Tumour Biol. 2015, 36, 5987–5997. [Google Scholar] [CrossRef] [PubMed]
- Jiang, C.F.; Li, D.M.; Shi, Z.M.; Wang, L.; Liu, M.M.; Ge, X.; Liu, X.; Qian, Y.C.; Wen, Y.Y.; Zhen, L.L.; et al. Estrogen regulates miRNA expression: Implication of estrogen receptor and miR-124/AKT2 in tumor growth and angiogenesis. Oncotarget 2016, 7, 36940–36955. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Chen, L.; Wu, Z.; Wang, M.; Jin, F.; Wang, N.; Hu, X.; Liu, Z.; Zhang, C.Y.; Zen, K.; et al. miR-124-3p functions as a tumor suppressor in breast cancer by targeting CBL. BMC Cancer 2016, 16, 826. [Google Scholar] [CrossRef] [PubMed]
- Cai, W.L.; Huang, W.D.; Li, B.; Chen, T.R.; Li, Z.X.; Zhao, C.L.; Li, H.Y.; Wu, Y.M.; Yan, W.J.; Xiao, J.R. microRNA-124 inhibits bone metastasis of breast cancer by repressing Interleukin-11. Mol. Cancer 2018, 17, 9. [Google Scholar] [CrossRef] [PubMed]
- Shi, P.; Chen, C.; Li, X.; Wei, Z.; Liu, Z.; Liu, Y. MicroRNA124 suppresses cell proliferation and invasion of triple negative breast cancer cells by targeting STAT3. Mol. Med. Rep. 2019, 19, 3667–3675. [Google Scholar]
- Ji, H.; Sang, M.; Liu, F.; Ai, N.; Geng, C. miR-124 regulates EMT based on ZEB2 target to inhibit invasion and metastasis in triple-negative breast cancer. Pathol. Res. Pract. 2019, 215, 697–704. [Google Scholar] [CrossRef]
- Goltzman, D. Osteolysis and cancer. J. Clin. Investig. 2001, 107, 1219–1220. [Google Scholar] [CrossRef]
- Liu, C.; Xing, H.; Guo, C.; Yang, Z.; Wang, Y.; Wang, Y. MiR-124 reversed the doxorubicin resistance of breast cancer stem cells through STAT3/HIF-1 signaling pathways. Cell Cycle 2019, 18, 2215–2227. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Zhang, Z.; Liu, X.; Huang, T.; He, W.; Shen, Y.; Liu, X.; Hong, K.; Cao, Q. miR-124 functions as a tumor suppressor in the endometrial carcinoma cell line HEC-1B partly by suppressing STAT3. Mol. Cell Biochem. 2014, 388, 219–231. [Google Scholar] [CrossRef]
- Cheng, Y.; Li, Y.; Nian, Y.; Liu, D.; Dai, F.; Zhang, J. STAT3 is involved in miR-124-mediated suppressive effects on esophageal cancer cells. BMC Cancer 2015, 15, 306. [Google Scholar] [CrossRef]
- Li, Z.; Qin, X.; Bian, W.; Li, Y.; Shan, B.; Yao, Z.; Li, S. Exosomal lncRNA ZFAS1 regulates esophageal squamous cell carcinoma cell proliferation, invasion, migration and apoptosis via microRNA-124/STAT3 axis. J. Exp. Clin. Cancer Res. 2019, 38, 477. [Google Scholar] [CrossRef]
- Xu, S.; Zhao, N.; Hui, L.; Song, M.; Miao, Z.W.; Jiang, X.J. MicroRNA-124-3p inhibits the growth and metastasis of nasopharyngeal carcinoma cells by targeting STAT3. Oncol. Rep. 2016, 35, 1385–1394. [Google Scholar] [CrossRef]
- Liu, S.; Hu, C.; Wang, Y.; Shi, G.; Li, Y.; Wu, H. miR-124 inhibits proliferation and invasion of human retinoblastoma cells by targeting STAT3. Oncol. Rep. 2016, 36, 2398–2404. [Google Scholar] [CrossRef]
- Wu, Z.; Huang, W.; Chen, B.; Bai, P.D.; Wang, X.G.; Xing, J.C. Up-regulation of miR-124 inhibits invasion and proliferation of prostate cancer cells through mediating JAK-STAT3 signaling pathway. Eur. Rev. Med. Pharmacol. Sci. 2020, 24, 7546. [Google Scholar]
- Ma, J.; Weng, L.; Wang, Z.; Jia, Y.; Liu, B.; Wu, S.; Cao, Y.; Sun, X.; Yin, X.; Shang, M.; et al. MiR-124 induces autophagy-related cell death in cholangiocarcinoma cells through direct targeting of the EZH2-STAT3 signaling axis. Exp. Cell Res. 2018, 366, 103–113. [Google Scholar] [CrossRef]
- Zheng, Y.B.; Xiao, G.C.; Tong, S.L.; Ding, Y.; Wang, Q.S.; Li, S.B.; Hao, Z.N. Paeoniflorin inhibits human gastric carcinoma cell proliferation through up-regulation of microRNA-124 and suppression of PI3K/Akt and STAT3 signaling. World J. Gastroenterol. 2015, 21, 7197–7207. [Google Scholar] [CrossRef]
- Lee, H.; Herrmann, A.; Deng, J.H.; Kujawski, M.; Niu, G.; Li, Z.; Forman, S.; Jove, R.; Pardoll, D.M.; Yu, H. Persistently activated Stat3 maintains constitutive NF-kappaB activity in tumors. Cancer Cell 2009, 15, 283–293. [Google Scholar] [CrossRef] [PubMed]
- Frank, D.A. STAT3 as a central mediator of neoplastic cellular transformation. Cancer Lett. 2007, 251, 199–210. [Google Scholar] [CrossRef] [PubMed]
- Kamran, M.Z.; Patil, P.; Gude, R.P. Role of STAT3 in cancer metastasis and translational advances. Biomed. Res. Int. 2013, 2013, 421821. [Google Scholar] [CrossRef] [PubMed]
- Ahmed-Choudhury, J.; Williams, K.T.; Young, L.S.; Adams, D.H.; Afford, S.C. CD40 mediated human cholangiocyte apoptosis requires JAK2 dependent activation of STAT3 in addition to activation of JNK1/2 and ERK1/2. Cell Signal. 2006, 18, 456–468. [Google Scholar] [CrossRef]
- de la Iglesia, N.; Konopka, G.; Puram, S.V.; Chan, J.A.; Bachoo, R.M.; You, M.J.; Levy, D.E.; Depinho, R.A.; Bonni, A. Identification of a PTEN-regulated STAT3 brain tumor suppressor pathway. Genes Dev. 2008, 22, 449–462. [Google Scholar] [CrossRef]
- Xiao, D.; Cui, X.; Wang, X. Long noncoding RNA XIST increases the aggressiveness of laryngeal squamous cell carcinoma by regulating miR-124-3p/EZH2. Exp. Cell Res. 2019, 381, 172–178. [Google Scholar] [CrossRef]
- Sabour Takanlu, J.; Aghaie Fard, A.; Mohammdi, S.; Hosseini Rad, S.M.A.; Abroun, S.; Nikbakht, M. Indirect Tumor Inhibitory Effects of MicroRNA-124 through Targeting EZH2 in The Multiple Myeloma Cell Line. Cell J. 2020, 22, 23–29. [Google Scholar]
- Bracken, A.P.; Pasini, D.; Capra, M.; Prosperini, E.; Colli, E.; Helin, K. EZH2 is downstream of the pRB-E2F pathway, essential for proliferation and amplified in cancer. EMBO J. 2003, 22, 5323–5335. [Google Scholar] [CrossRef] [PubMed]
- Tiwari, N.; Tiwari, V.K.; Waldmeier, L.; Balwierz, P.J.; Arnold, P.; Pachkov, M.; Meyer-Schaller, N.; Schubeler, D.; van Nimwegen, E.; Christofori, G. Sox4 is a master regulator of epithelial-mesenchymal transition by controlling Ezh2 expression and epigenetic reprogramming. Cancer Cell 2013, 23, 768–783. [Google Scholar] [CrossRef] [PubMed]
- Papagiannakopoulos, T.; Kosik, K.S. MicroRNA-124: Micromanager of neurogenesis. Cell Stem Cell 2009, 4, 375–376. [Google Scholar] [CrossRef] [PubMed]
- Tuttle, M.; Dalman, M.R.; Liu, Q.; Londraville, R.L. Leptin-a mediates transcription of genes that participate in central endocrine and phosphatidylinositol signaling pathways in 72-hour embryonic zebrafish (Danio rerio). PeerJ 2019, 7, e6848. [Google Scholar] [CrossRef]
- Neo, W.H.; Yap, K.; Lee, S.H.; Looi, L.S.; Khandelia, P.; Neo, S.X.; Makeyev, E.V.; Su, I.H. MicroRNA miR-124 controls the choice between neuronal and astrocyte differentiation by fine-tuning Ezh2 expression. J. Biol. Chem. 2014, 289, 20788–20801. [Google Scholar] [CrossRef] [PubMed]
- Suzuki, F.; Okuno, M.; Tanaka, T.; Sanuki, R. Overexpression of neural miRNAs miR-9/9* and miR-124 suppresses differentiation to Muller glia and promotes differentiation to neurons in mouse retina in vivo. Genes Cells 2020, 25, 741–752. [Google Scholar] [CrossRef] [PubMed]
- Yoo, A.S.; Sun, A.X.; Li, L.; Shcheglovitov, A.; Portmann, T.; Li, Y.; Lee-Messer, C.; Dolmetsch, R.E.; Tsien, R.W.; Crabtree, G.R. MicroRNA-mediated conversion of human fibroblasts to neurons. Nature 2011, 476, 228–231. [Google Scholar] [CrossRef] [PubMed]
- Lee, S.W.; Oh, Y.M.; Lu, Y.L.; Kim, W.K.; Yoo, A.S. MicroRNAs Overcome Cell Fate Barrier by Reducing EZH2-Controlled REST Stability during Neuronal Conversion of Human Adult Fibroblasts. Dev. Cell 2018, 46, 73–84. [Google Scholar] [CrossRef] [PubMed]
- Xue, Y.; Ouyang, K.; Huang, J.; Zhou, Y.; Ouyang, H.; Li, H.; Wang, G.; Wu, Q.; Wei, C.; Bi, Y.; et al. Direct conversion of fibroblasts to neurons by reprogramming PTB-regulated microRNA circuits. Cell 2013, 152, 82–96. [Google Scholar] [CrossRef] [PubMed]
- Makeyev, E.V.; Zhang, J.; Carrasco, M.A.; Maniatis, T. The MicroRNA miR-124 promotes neuronal differentiation by triggering brain-specific alternative pre-mRNA splicing. Mol. Cell 2007, 27, 435–448. [Google Scholar] [CrossRef]
- Mangale, V.S.; Hirokawa, K.E.; Satyaki, P.R.; Gokulchandran, N.; Chikbire, S.; Subramanian, L.; Shetty, A.S.; Martynoga, B.; Paul, J.; Mai, M.V.; et al. Lhx2 selector activity specifies cortical identity and suppresses hippocampal organizer fate. Science 2008, 319, 304–309. [Google Scholar] [CrossRef]
- Wilson, S.I.; Shafer, B.; Lee, K.J.; Dodd, J. A molecular program for contralateral trajectory: Rig-1 control by LIM homeodomain transcription factors. Neuron 2008, 59, 413–424. [Google Scholar] [CrossRef]
- Subramanian, L.; Sarkar, A.; Shetty, A.S.; Muralidharan, B.; Padmanabhan, H.; Piper, M.; Monuki, E.S.; Bach, I.; Gronostajski, R.M.; Richards, L.J.; et al. Transcription factor Lhx2 is necessary and sufficient to suppress astrogliogenesis and promote neurogenesis in the developing hippocampus. Proc. Natl. Acad. Sci. USA 2011, 108, E265–E274. [Google Scholar] [CrossRef]
- Franke, K.; Otto, W.; Johannes, S.; Baumgart, J.; Nitsch, R.; Schumacher, S. miR-124-regulated RhoG reduces neuronal process complexity via ELMO/Dock180/Rac1 and Cdc42 signalling. EMBO J. 2012, 31, 2908–2921. [Google Scholar] [CrossRef]
- Swanwick, C.C.; Shapiro, M.E.; Vicini, S.; Wenthold, R.J. Flotillin-1 promotes formation of glutamatergic synapses in hippocampal neurons. Dev. Neurobiol. 2010, 70, 875–883. [Google Scholar] [CrossRef] [PubMed]
- Averaimo, S.; Gritti, M.; Barini, E.; Gasparini, L.; Mazzanti, M. CLIC1 functional expression is required for cAMP-induced neurite elongation in post-natal mouse retinal ganglion cells. J. Neurochem. 2014, 131, 444–456. [Google Scholar] [CrossRef] [PubMed]
- Newell-Litwa, K.A.; Badoual, M.; Asmussen, H.; Patel, H.; Whitmore, L.; Horwitz, A.R. ROCK1 and 2 differentially regulate actomyosin organization to drive cell and synaptic polarity. J. Cell Biol. 2015, 210, 225–242. [Google Scholar] [CrossRef] [PubMed]
- Epifanova, E.; Babaev, A.; Newman, A.G.; Tarabykin, V. Role of Zeb2/Sip1 in neuronal development. Brain Res. 2019, 1705, 24–31. [Google Scholar] [CrossRef] [PubMed]
- O’Neill, C.P.; Dwyer, R.M. Nanoparticle-Based Delivery of Tumor Suppressor microRNA for Cancer Therapy. Cells 2020, 9, 521. [Google Scholar] [CrossRef] [PubMed]
| TARGET GENES | EFFECTS | REF. |
|---|---|---|
| AURKA | Inhibition of cell proliferation and potentiation of the temozolomide TMZ-based chemosensitivity. | [45] |
| CAPNS1 | Inhibition of cell migration and invasion. | [42] |
| CDK4 | Conferring radio-sensitivity. | [51] |
| CDK4, CDK6, and PPP1R13L | Inhibition of cell migration, decrease in cell viability, and cell cycle arrest at the G0/G1 phase. | [47] |
| CDK6 | Induce G1 cell cycle arrest. | [15] |
| CDK6 | Decrease in cell proliferation and migration and conferring chemosensitivity to TMZ. | [48] |
| CDK6 | Inhibition of cell proliferation. | [38] |
| TEAD1, MAPK14, and SERP1 | Increase in cell death. | [41] |
| EZH2 | Expression of miR-124/128/137 of artificial miRNA clusters, reduction of cell proliferation both in vitro and in vivo, and prolongation of survival in a mouse model. | [50] |
| KITLG, NRP2, SEMA6D, and THBS1 | Regulation of self-renewal, apoptosis, and invasion. | [44] |
| NRP1 | Arresting of cell cycle, inhibition of cell proliferation and migration, inhibition of tumor angiogenesis, and induction of apoptosis. | [49] |
| PIM1 | Inhibition of cell proliferation, invasion, and aerobic glycolysis and promotion of apoptosis. | [43] |
| PPP1R13L | Inhibition of cell proliferation, G1/S transition, and invasiveness. | [40] |
| SDCBP | Decrease in malignancy of glioblastoma cells, inhibition of cell proliferation, migration, and invasion. | [46] |
| SOS1 | Inhibition of cell proliferation. | [37] |
| STAT3 | Reversing immunosuppression in the tumor microenvironment, enhancing T cell-mediated immune clearance, and inhibition of glioma growth. | [39] |
| TARGET GENES | EFFECTS | REF. |
|---|---|---|
| AKT2 | Arresting of cell cycle at the G0/G1 phase and inhibition of cell growth, colony formation, and tumor growth without apoptosis.Inhibition of migratory and invasive abilities. | [66] |
| BECN1, RELA, and SQSTM1 | Disruption of autophagy and reduction of cell survival. | [59] |
| CD164 | Inhibition of tumor cell proliferation, colony formation, migration, and induction of apoptosis. | [58] |
| CDH2 | Inhibition of cell proliferation and invasion. | [68] |
| EZH2 | Inhibition of tumor cell proliferation and inhibition of the EMT process. | [67] |
| LHX2 | Attenuation of cellular migratory and invasive abilities. | [64] |
| MYO10 | Inhibition of migration and metastatic ability. | [61] |
| SNAI2 | Inhibition of invasion. | [56] |
| STAT3 | Inhibition of cell proliferation and induction of apoptosis. | [55] |
| STAT3 | Inhibition of cell growth and colony formation and induction of apoptosis. | [32] |
| STAT3 | Inhibition of cell growth and colony formation and induction of apoptosis. Increasing radio sensitivity. | [63] |
| STAT3 | Inhibition of invasion and metastasis capacities. Increasing sensitivity to cisplatin. | [65] |
| TXNRD1 | Improving sensitization of radiation-resistant cells to radiation. | [62] |
| ZEB1 | Inhibition of migration and invasion through suppressing EMT. | [57] |
| TARGET GENES | EFFECTS | REF. |
|---|---|---|
| DDX6 and PTB1 | Induction of apoptosis. Decreasing production of lactic acid, affecting the Warburg effect. | [74] |
| VANGL1, MYH9, and SOX9 | Inhibition of tumorigenicity. | [69] |
| PPP1R13L | Inhibition of cell proliferation and tumor formation. | [71] |
| PTB1 | Enhancement in oxidative stress and induction of apoptosis and autophagy. | [72] |
| ROCK1 | Inhibition of cell proliferation, migration, and invasion. | [70] |
| STAT3 | High-intensity focused ultrasound mediated inhibition of migration. | [75] |
| STAT3 | Induction of apoptosis and inhibition of tumor growth. | [76] |
| TARGET GENES | EFFECTS | REF. |
|---|---|---|
| CASC3 | Inhibition of tumor growth. | [81] |
| CDK6, IQGAP1, SMYD3, and VIM | Inhibition of cell growth. | [77] |
| CLIC1 | Inhibition of cell proliferation, migration and invasion. | [83] |
| EZH2 and ROCK2 | Inhibition of cell motility and invasion, and suppression of intrahepatic and pulmonary metastasis. Inhibition of EMT with impaired formation of stress fibers, filopodia, and lamellipodia. | [79] |
| ITGAV and SP1 | Inhibition of migration and tumor metastasis. | [82] |
| PIK3CA | Cell cycle arrest at the G0/G1 phase. | [78] |
| STAT3 | Inhibition of cell proliferation and induction of apoptosis. | [80] |
| TARGET GENES | EFFECTS | REF. |
|---|---|---|
| EZH2 | Inhibition of cell proliferation and colony formation and induction of apoptosis. Increased sensitization of 5-FU. | [87] |
| EZH2 and JAG1 | Inhibition of cell growth, migration, invasion, and tumor growth. | [88] |
| ROCK1 | Inhibition of cell proliferation, migration, and invasion. | [86] |
| SPHK1 | Inhibition of cell proliferation and tumorigenicity. | [84] |
| SPHK1 | Suppression of cell proliferation and invasion. | [85] |
| TARGET GENES | EFFECTS | REF. |
|---|---|---|
| AKT2 | Inhibition of cell proliferation, migration, and invasion. | [94] |
| CBL | Inhibition of cell proliferation and invasion. | [95] |
| CD151 | Inhibition of proliferation via cell cycle arrest but does not induce apoptosis. Reduction of invasive and metastatic potential. | [90] |
| CDK4 | Inhibition of cell proliferation. | [93] |
| FLOT1 | Inhibition of cell growth and migration. | [91] |
| IL11 | Inhibition of the survival and differentiation of osteoclast progenitor cells through cancer cell-derived microRNA-124. | [96] |
| CTGF, ITGB1, RHOG, and ROCK1 | Inhibition of metastasis. | [89] |
| SNAI2 | Inhibition of cell colony formation and pulmonary metastasis. | [92] |
| STAT3 | Inhibition of cell proliferation and invasion. | [97] |
| STAT3 | Improving sensitization of doxorubicin. | [100] |
| ZEB2 | Inhibition of cell growth and migration and EMT. | [98] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sanuki, R.; Yamamura, T. Tumor Suppressive Effects of miR-124 and Its Function in Neuronal Development. Int. J. Mol. Sci. 2021, 22, 5919. https://doi.org/10.3390/ijms22115919
Sanuki R, Yamamura T. Tumor Suppressive Effects of miR-124 and Its Function in Neuronal Development. International Journal of Molecular Sciences. 2021; 22(11):5919. https://doi.org/10.3390/ijms22115919
Chicago/Turabian StyleSanuki, Rikako, and Tomonori Yamamura. 2021. "Tumor Suppressive Effects of miR-124 and Its Function in Neuronal Development" International Journal of Molecular Sciences 22, no. 11: 5919. https://doi.org/10.3390/ijms22115919
APA StyleSanuki, R., & Yamamura, T. (2021). Tumor Suppressive Effects of miR-124 and Its Function in Neuronal Development. International Journal of Molecular Sciences, 22(11), 5919. https://doi.org/10.3390/ijms22115919






