Autophagy Modulators in Cancer Therapy
Abstract
1. Introduction
2. Types of Autophagy
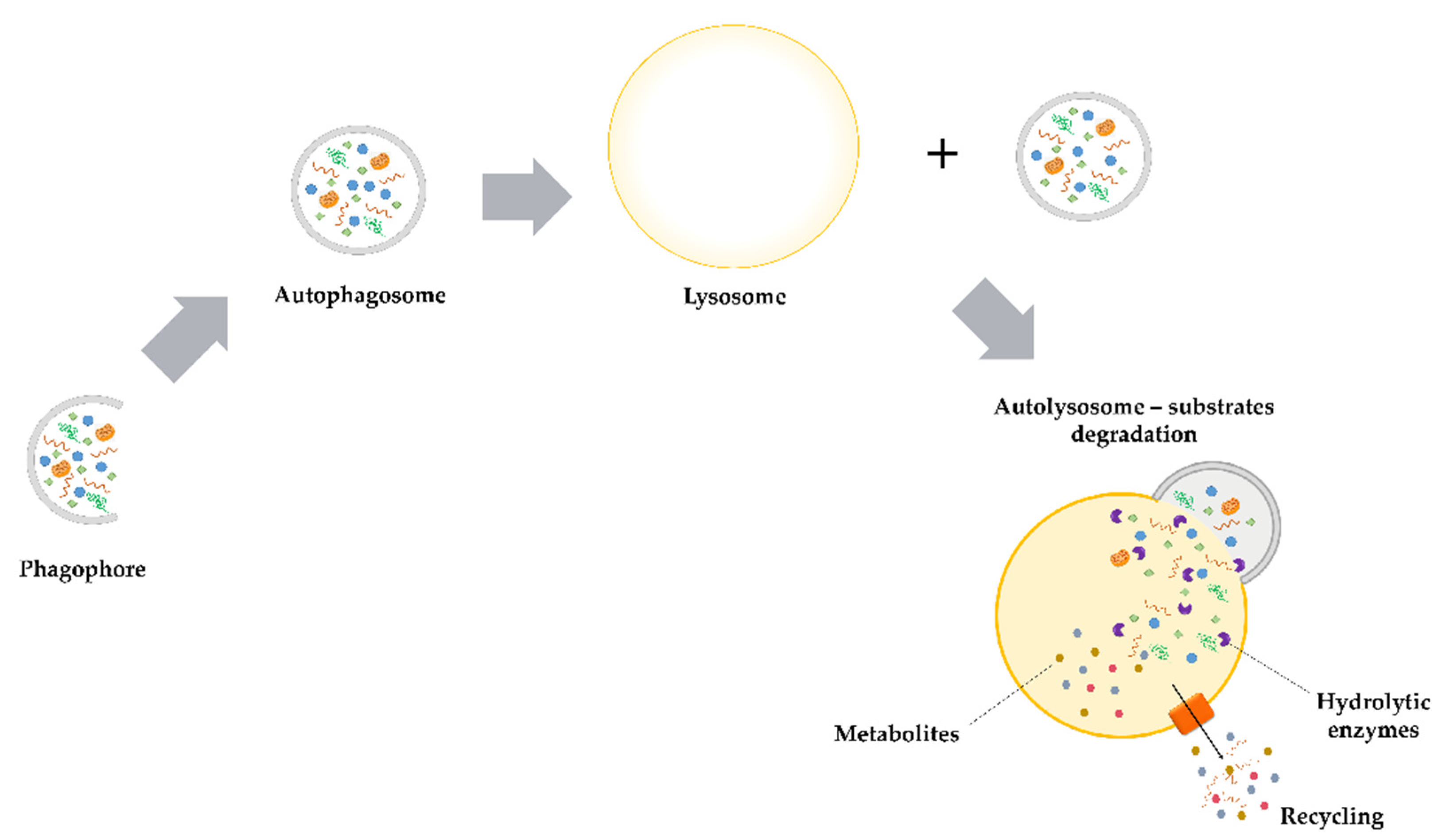
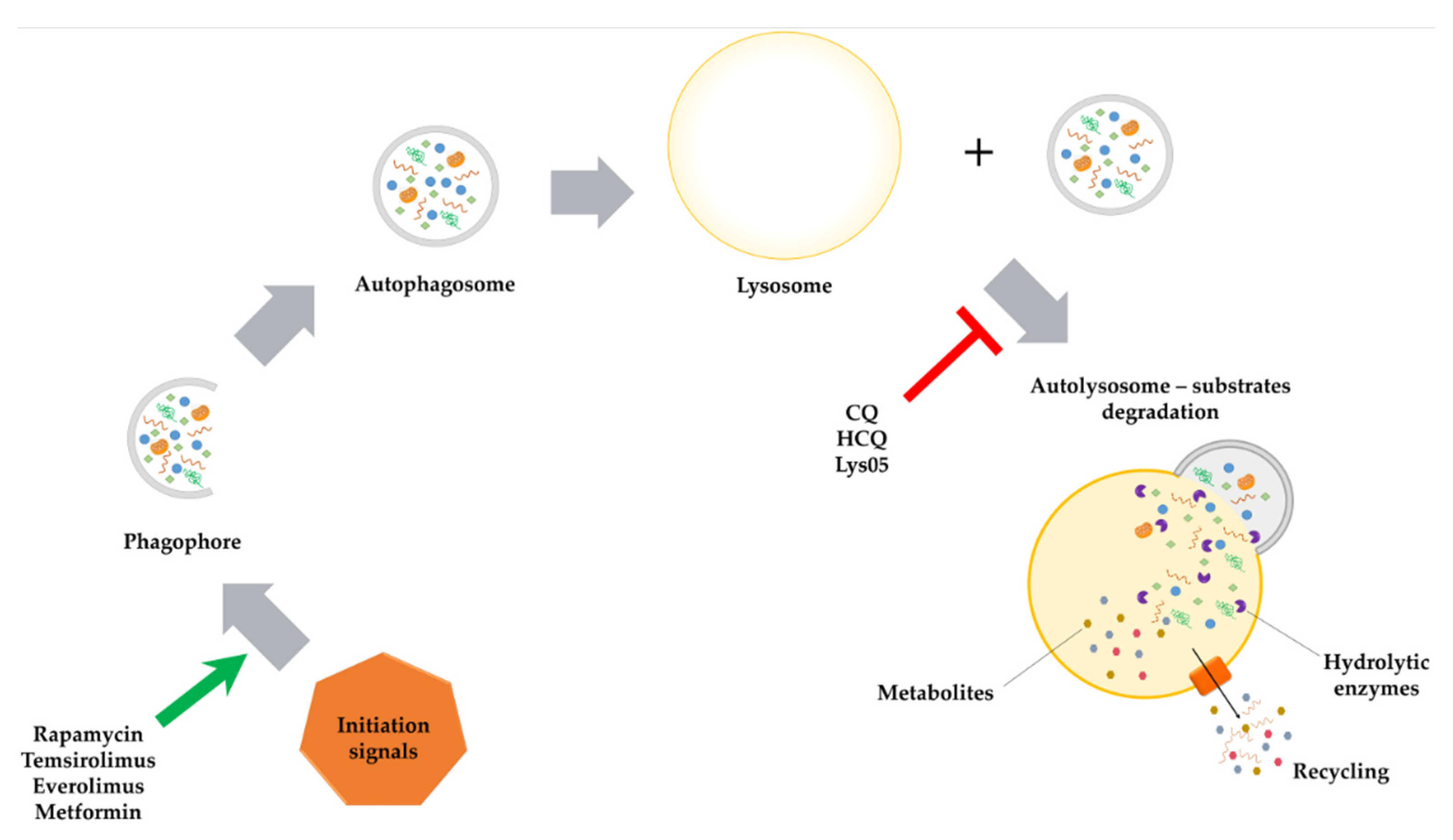
2.1. Macroautophagy
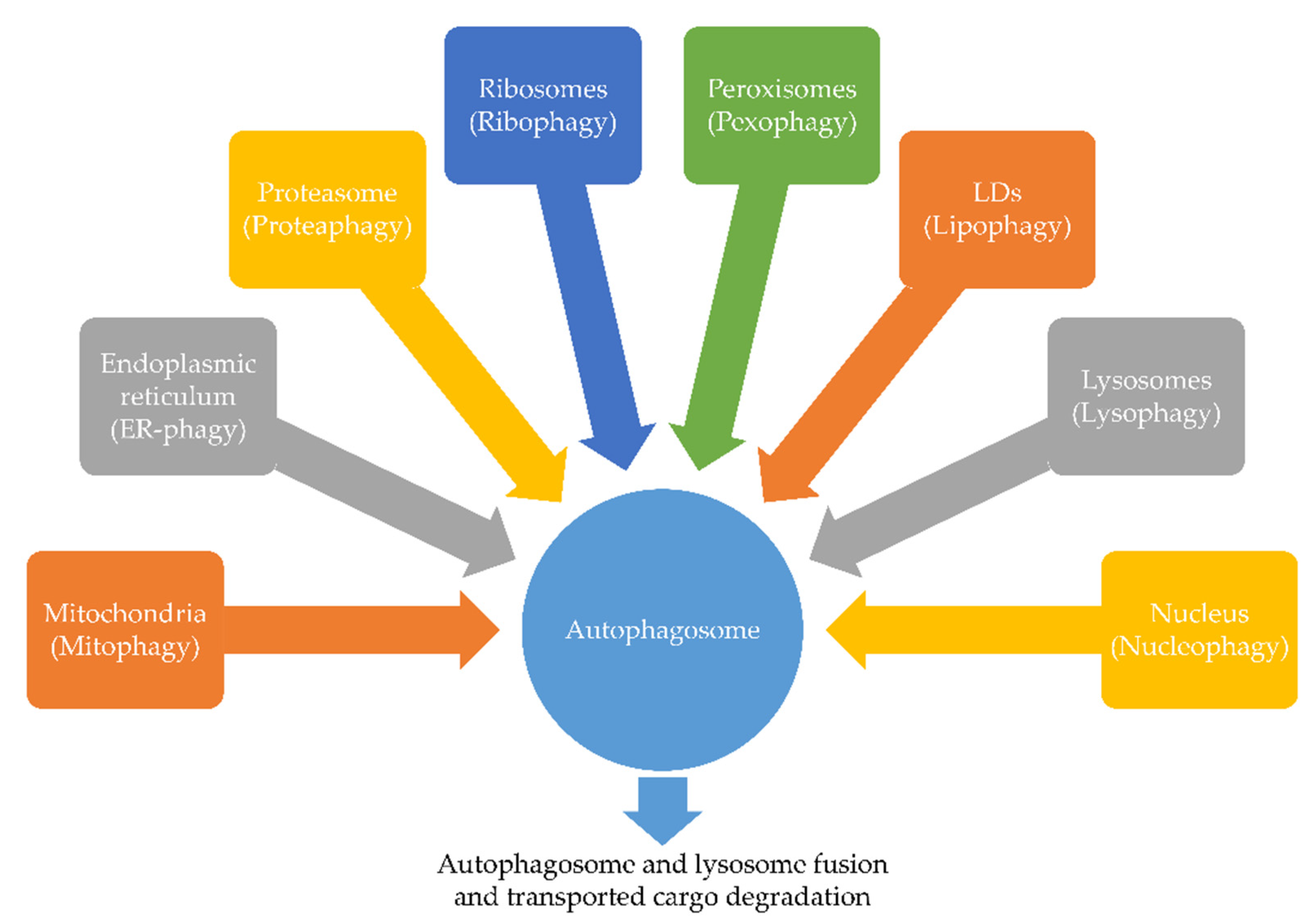
2.2. Selective Autophagy
2.2.1. ER-Phagy
2.2.2. Mitophagy
2.2.3. Proteaphagy
2.2.4. Ribophagy
2.2.5. Pexophagy
2.2.6. Lipophagy
2.2.7. Lysophagy
2.2.8. Nucleophagy
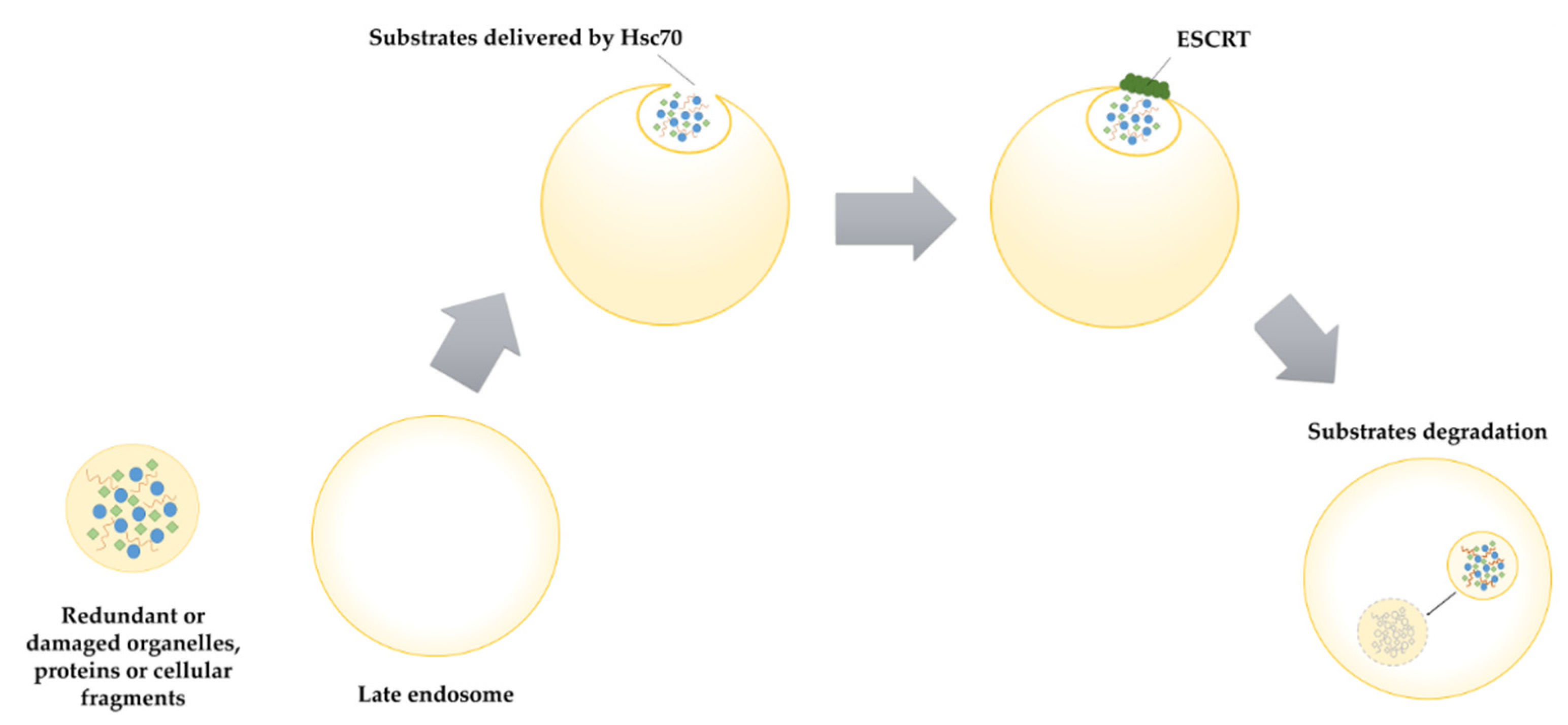
2.3. Microautophagy
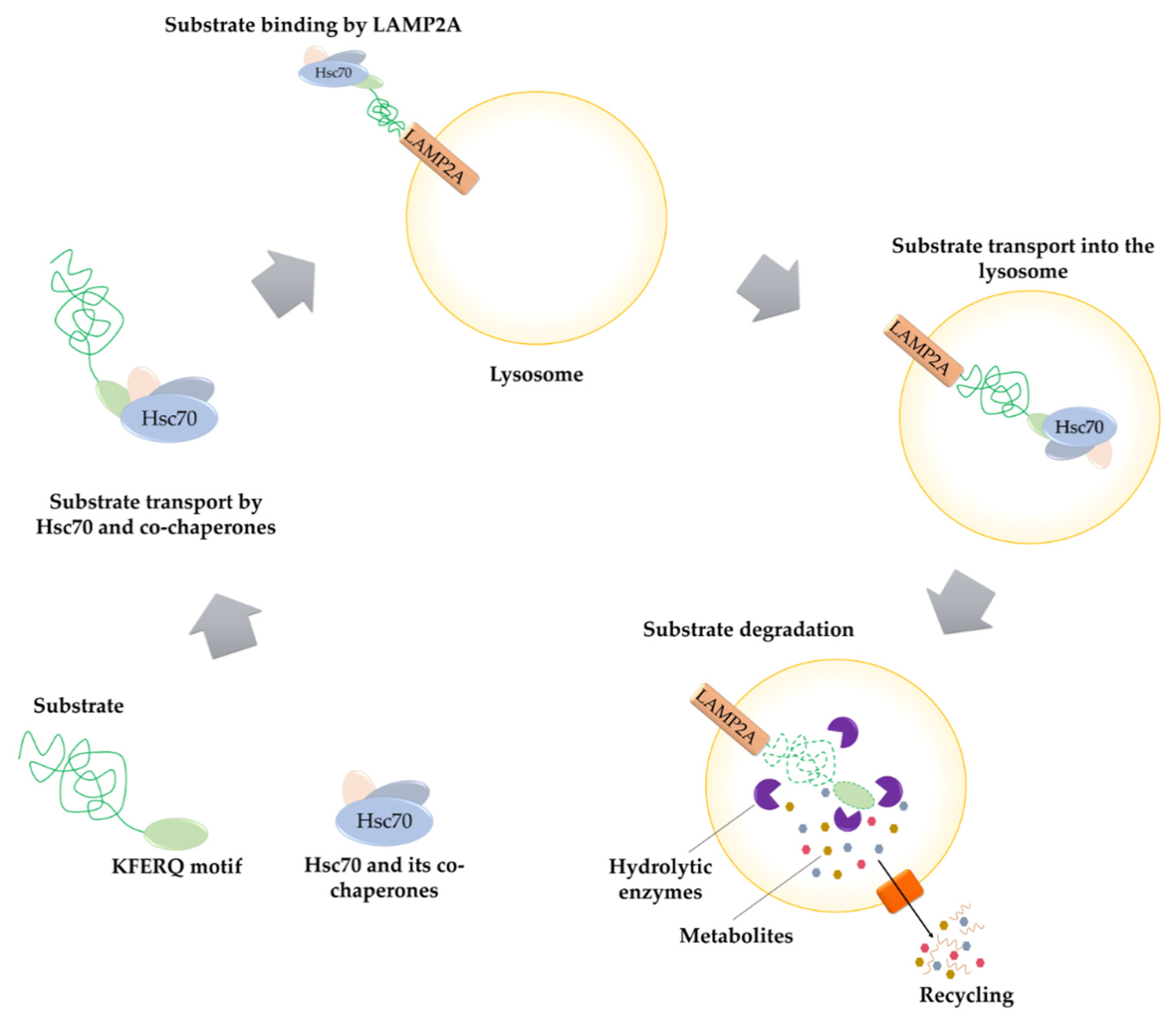
2.4. Chaperone-Mediated Autophagy
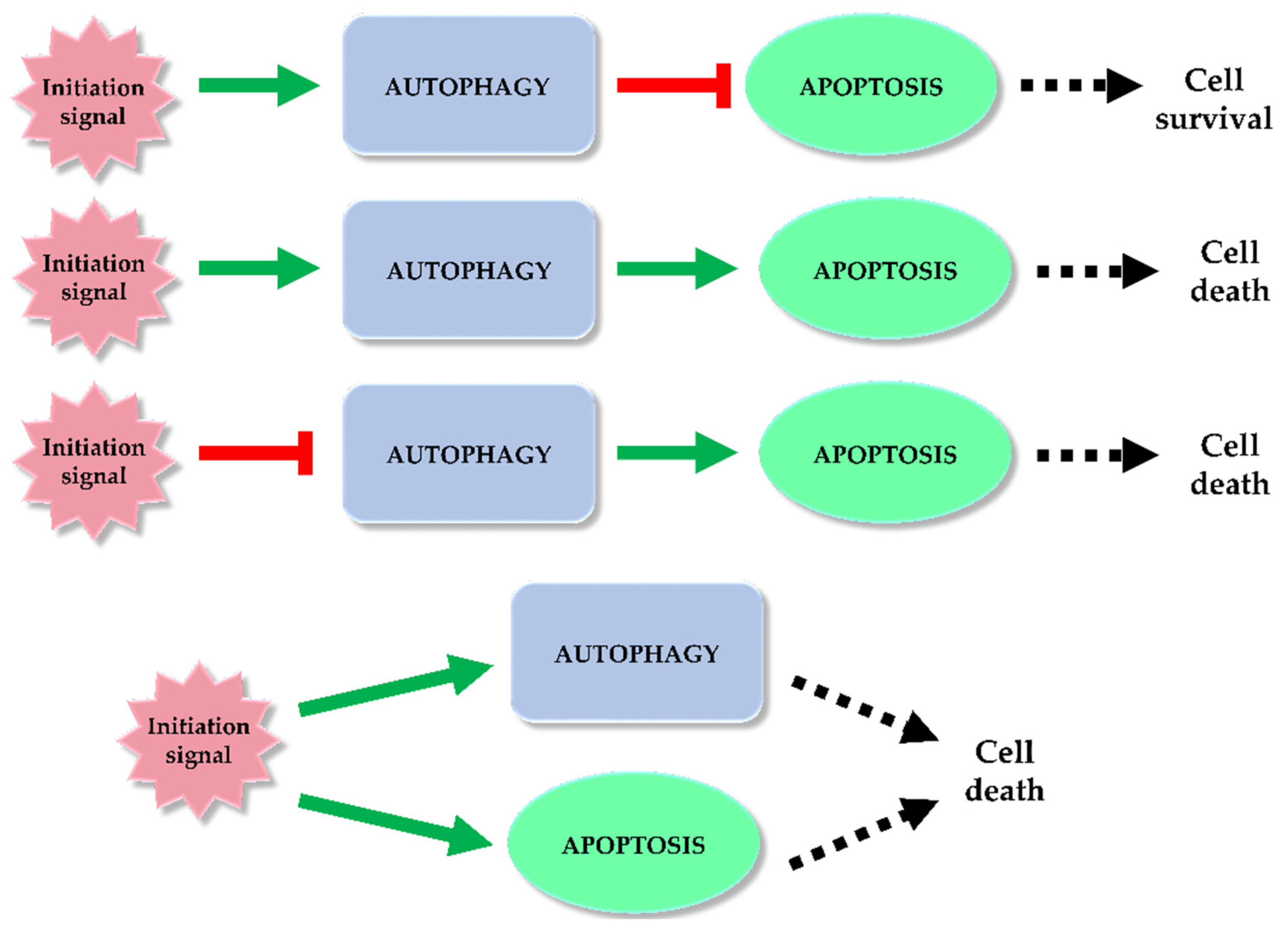
3. Autophagy and Programmed Cell Death—Double-Edged Sword Relationship
3.1. p53 in Apoptosis and Autophagy
3.2. Bcl-2/Beclin 1 in Apoptosis and Autophagy
3.3. Atg Proteins in Apoptosis and Autophagy
3.4. p62 in Apoptosis and Autophagy
3.5. Caspases in Apoptosis and Autophagy
4. Autophagy Inhibitors and Activators
4.1. Autophagy Inhibitors Undergoing Clinical Trials
4.1.1. Chloroquine
4.1.2. Hydroxychloroquine
4.1.3. Verteporfin
4.1.4. Clarithromycin
4.2. Autophagy Inhibiotors Undergoing Preclinical Trials
4.2.1. 3-Methyladenine
4.2.2. SAR405
4.2.3. Lys05
4.2.4. ROC-325
4.2.5. Spautin-1
4.2.6. MM124 and MM137
4.3. Autophagy Activators Undergoing Clinical Trials
4.3.1. Rapamycin
4.3.2. Temsirolimus
4.3.3. Everolimus
4.3.4. Metformin
4.4. Autophagy Activators Undergoing Preclinical Trials
4.4.1. Miconazole
4.4.2. CRO15
4.4.3. α-Hederin
4.4.4. MJ-33
5. Conclusions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| 3-MA | 3-Methyladenine |
| 5FUR | 5-fluorouracil-resistant |
| ADCD | autophagy-dependent cell death |
| AML | acute myeloid leukemia |
| AMPK | 5′ adenosine monophosphate-activated protein kinase |
| ATG | autophagy-related |
| AZA | azacitidine |
| C3G | cyaniding-3-O-glucoside |
| CAM | clarithromycin |
| CMA | chaperone-mediated autophagy |
| CML | chronic myeloid leukemia |
| CQ | chloroquine |
| DRAM | damage-regulated autophagy modulator |
| eMI | endosomal microphagy |
| ER | endoplasmic reticulum |
| ESCRT | endosomal sorting complex required for transport |
| FDA | Food and Drug Administration |
| HCQ | hydroxychloroquine |
| Hsc70 | heat shock cognate protein 70 |
| JNK-1 | C-Jun N-terminal protein kinase 1 |
| LAL | lysosomal acid lipase |
| LAMP2A | lysosome associated membrane protein type 2A |
| LC3 | light chain protein 3 |
| LDs | lipid droplets |
| Mcl-1 | myeloid cell leukemia 1 |
| MCZ | miconazole |
| mTOR | mammalian target of rapamycin |
| NSCLC | non-small cell lung cancer |
| NUFIP 1 | nuclear FMR1 interacting protein 1 |
| PIK3K | phosphoinositide 3-kinase |
| PMPs | peroxisomal membrane proteins |
| RAD001 | everolimus |
| RAPA | rapamycin |
| SQSTM1 | sequestosome 1 |
| TEM | temsirolimus |
| VP | verteporfin |
| α-HN | α-Hederin |
References
- Levine, B.; Klionsky, D.J. Development by self-digestion: Molecular mechanisms and biological functions of autophagy. Dev. Cell 2004, 6, 463–477. [Google Scholar] [CrossRef]
- Kroemer, G.; Galluzzi, L.; Vandenabeele, P.; Abrams, J.; Alnemri, E.S.; Baehrecke, E.H.; Blagosklonny, M.V.; El-Deiry, W.S.; Golstein, P.; Green, D.R.; et al. Classification of cell death: Recommendations of the Nomenclature Committee on Cell Death 2009. Cell Death Differ. 2009, 16, 3–11. [Google Scholar] [CrossRef] [PubMed]
- Kim, R. Recent advances in understanding the cell death pathways activated by anticancer therapy. Cancer 2005, 103, 1551–1560. [Google Scholar] [CrossRef] [PubMed]
- Polewska, J. Autophagy—molecular mechanism, apoptosis and cancer. Postepy Hig. Med. Dosw. 2012, 66, 921–936. [Google Scholar] [CrossRef] [PubMed]
- Dereń-Wagemann, I.; Kiełbiński, M.; Kuliczkowski, K. Autofagia—Proces o dwóch obliczach. Acta Haematol. Pol. 2013, 44, 383–391. [Google Scholar] [CrossRef]
- Galluzzi, L.; Vitale, I.; Aaronson, S.A.; Abrams, J.M.; Adam, D.; Agostinis, P.; Alnemri, E.S.; Altucci, L.; Amelio, I.; Andrews, D.W.; et al. Molecular mechanisms of cell death: Recommendations of the Nomenclature Committee on Cell Death 2018. Cell Death Differ. 2018, 25, 486–541. [Google Scholar] [CrossRef]
- Klionsky, D.J. The molecular machinery of autophagy: Unanswered questions. J. Cell Sci. 2005, 118, 7–18. [Google Scholar] [CrossRef]
- Yang, Y.-P.; Liang, Z.-Q.; Gu, Z.-L.; Qin, Z.-H. Molecular mechanism and regulation of autophagy. Acta Pharmacol. Sin. 2005, 26, 1421–1434. [Google Scholar] [CrossRef]
- Rabinowitz, J.D.; White, E. Autophagy and metabolism. Science 2010, 330, 1344–1348. [Google Scholar] [CrossRef]
- Eskelinen, E.-L.; Saftig, P. Autophagy: A lysosomal degradation pathway with a central role in health and disease. Biochim. Et Biophys. Acta (BBA) Mol. Cell Res. 2009, 1793, 664–673. [Google Scholar] [CrossRef]
- Andrade-Tomaz, M.; de Souza, I.; Rocha, C.R.R.; Gomes, L.R. The role of chaperone-mediated autophagy in cell cycle control and its implications in cancer. Cells 2020, 9, 2140. [Google Scholar] [CrossRef]
- Kon, M.; Kiffin, R.; Koga, H.; Chapochnick, J.; Macian, F.; Varticovski, L.; Cuervo, A.M. Chaperone-mediated autophagy is required for tumor growth. Sci. Transl. Med. 2011, 3, 109ra117. [Google Scholar] [CrossRef]
- Glick, D.; Barth, S.; Macleod, K.F. Autophagy: Cellular and molecular mechanisms. J. Pathol. 2010, 221, 3–12. [Google Scholar] [CrossRef] [PubMed]
- Mizushima, N. Autophagy: Process and function. Genes Dev. 2007, 21, 2861–2873. [Google Scholar] [CrossRef] [PubMed]
- Mizushima, N.; Yoshimori, T.; Ohsumi, Y. The role of Atg proteins in autophagosome formation. Annu. Rev. Cell Dev. Biol. 2011, 27, 107–132. [Google Scholar] [CrossRef]
- Kondo, Y.; Kanzawa, T.; Sawaya, R.; Kondo, S. The role of autophagy in cancer development and response to therapy. Nat. Rev. Cancer 2005, 5, 726–734. [Google Scholar] [CrossRef]
- Mehrpour, M.; Esclatine, A.; Beau, I.; Codogno, P. Overview of macroautophagy regulation in mammalian cells. Cell Res. 2010, 20, 748–762. [Google Scholar] [CrossRef]
- Yang, Y.; Klionsky, D.J. Autophagy and disease: Unanswered questions. Cell Death Differ. 2020, 27, 858–871. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; He, S.; Ma, B. Autophagy and autophagy-related proteins in cancer. Mol. Cancer 2020, 19, 12. [Google Scholar] [CrossRef] [PubMed]
- Liang, X.H.; Jackson, S.; Seaman, M.; Brown, K.; Kempkes, B.; Hibshoosh, H.; Levine, B. Induction of autophagy and inhibition of tumorigenesis by beclin 1. Nature 1999, 402, 672–676. [Google Scholar] [CrossRef]
- Jin, S.; White, E. Role of autophagy in cancer: Management of metabolic stress. Autophagy 2007, 3, 28–31. [Google Scholar] [CrossRef]
- Ding, Z.-B.; Shi, Y.-H.; Zhou, J.; Qiu, S.-J.; Xu, Y.; Dai, Z.; Shi, G.-M.; Wang, X.-Y.; Ke, A.-W.; Wu, B.; et al. Association of autophagy defect with a malignant phenotype and poor prognosis of hepatocellular carcinoma. Cancer Res. 2008, 68, 9167–9175. [Google Scholar] [CrossRef]
- Birgisdottir, Å.B.; Lamark, T.; Johansen, T. The LIR motif—Crucial for selective autophagy. J. Cell Sci. 2013, 126, 3237–3247. [Google Scholar] [CrossRef]
- Zaffagnini, G.; Martens, S. Mechanisms of selective autophagy. J. Mol. Biol. 2016, 428, 1714–1724. [Google Scholar] [CrossRef]
- Rogov, V.V.; Stolz, A.; Ravichandran, A.C.; Rios-Szwed, D.O.; Suzuki, H.; Kniss, A.; Löhr, F.; Wakatsuki, S.; Dötsch, V.; Dikic, I.; et al. Structural and functional analysis of the GABARAP interaction motif (GIM). EMBO Rep. 2018, 19, e47268. [Google Scholar] [CrossRef]
- Noda, N.N.; Kumeta, H.; Nakatogawa, H.; Satoo, K.; Adachi, W.; Ishii, J.; Fujioka, Y.; Ohsumi, Y.; Inagaki, F. Structural basis of target recognition by Atg8/LC3 during selective autophagy. Genes Cells 2008, 13, 1211–1218. [Google Scholar] [CrossRef] [PubMed]
- Johansen, T.; Lamark, T. Selective autophagy mediated by autophagic adapter proteins. Autophagy 2011, 7, 279–296. [Google Scholar] [CrossRef] [PubMed]
- Li, W.; He, P.; Huang, Y.; Li, Y.-F.; Lu, J.; Li, M.; Kurihara, H.; Luo, Z.; Meng, T.; Onishi, M.; et al. Selective autophagy of intracellular organelles: Recent research advances. Theranostics 2021, 11, 222–256. [Google Scholar] [CrossRef] [PubMed]
- Lei, Y.; Zhang, D.; Yu, J.; Dong, H.; Zhang, J.; Yang, S. Targeting autophagy in cancer stem cells as an anticancer therapy. Cancer Lett. 2017, 393, 33–39. [Google Scholar] [CrossRef]
- Guo, J.Y.; White, E. Autophagy, metabolism, and cancer. Cold Spring Harb. Symp. Quant. Biol. 2016, 81, 73–78. [Google Scholar] [CrossRef]
- Wang, F.; Jia, J.; Rodrigues, B. Autophagy, metabolic disease, and pathogenesis of heart dysfunction. Can. J. Cardiol. 2017, 33, 850–859. [Google Scholar] [CrossRef]
- Miettinen, T.P.; Björklund, M. The mevalonate pathway as a metabolic requirement for autophagy–implications for growth control, proteostasis, and disease. Mol. Cell. Oncol. 2016, 3, e1143546. [Google Scholar] [CrossRef] [PubMed]
- Zhong, Z.; Sanchez-Lopez, E.; Karin, M. Autophagy, NLRP3 inflammasome and auto-inflammatory/immune diseases. Clin. Exp. Rheumatol 2016, 34, 12–16. [Google Scholar]
- Menzies, F.M.; Fleming, A.; Caricasole, A.; Bento, C.F.; Andrews, S.P.; Ashkenazi, A.; Füllgrabe, J.; Jackson, A.; Jimenez Sanchez, M.; Karabiyik, C.; et al. Autophagy and neurodegeneration: Pathogenic mechanisms and therapeutic opportunities. Neuron 2017, 93, 1015–1034. [Google Scholar] [CrossRef] [PubMed]
- Plaza-Zabala, A.; Sierra-Torre, V.; Sierra, A. Autophagy and microglia: Novel partners in neurodegeneration and aging. Int. J. Mol. Sci. 2017, 18, 598. [Google Scholar] [CrossRef]
- Chino, H.; Mizushima, N. ER-Phagy: Quality control and turnover of endoplasmic reticulum. Trends Cell Biol. 2020, 30, 384–398. [Google Scholar] [CrossRef]
- Ruggiano, A.; Foresti, O.; Carvalho, P. ER-associated degradation: Protein quality control and beyond. J. Cell Biol. 2014, 204, 869–879. [Google Scholar] [CrossRef] [PubMed]
- Oikonomou, C.; Hendershot, L.M. Disposing of misfolded ER proteins: A troubled substrate’s way out of the ER. Mol. Cell. Endocrinol. 2020, 500, 110630. [Google Scholar] [CrossRef] [PubMed]
- Wilkinson, S. ER-phagy: Shaping up and destressing the endoplasmic reticulum. FEBS J. 2019, 286, 2645–2663. [Google Scholar] [CrossRef]
- Schuck, S.; Gallagher, C.M.; Walter, P. ER-phagy mediates selective degradation of endoplasmic reticulum independently of the core autophagy machinery. J. Cell Sci. 2014, 127, 4078–4088. [Google Scholar] [CrossRef]
- De Leonibus, C.; Cinque, L.; Settembre, C. Emerging lysosomal pathways for quality control at the endoplasmic reticulum. Febs Lett. 2019, 593, 2319–2329. [Google Scholar] [CrossRef]
- Islam, F.; Gopalan, V.; Law, S.; Tang, J.C.-o.; Lam, A.K.-y. FAM134B promotes esophageal squamous cell carcinoma in vitro and its correlations with clinicopathologic features. Hum. Pathol. 2019, 87, 1–10. [Google Scholar] [CrossRef]
- Islam, F.; Gopalan, V.; Wahab, R.; Smith, R.A.; Qiao, B.; Lam, A.K.-Y. Stage dependent expression and tumor suppressive function of FAM134B (JK1) in colon cancer. Mol. Carcinog. 2017, 56, 238–249. [Google Scholar] [CrossRef] [PubMed]
- Vara-Perez, M.; Felipe-Abrio, B.; Agostinis, P. Mitophagy in cancer: A tale of adaptation. Cells 2019, 8, 493. [Google Scholar] [CrossRef]
- Sorrentino, V.; Menzies, K.J.; Auwerx, J. Repairing mitochondrial dysfunction in disease. Annu. Rev. Pharmacol. Toxicol. 2018, 58, 353–389. [Google Scholar] [CrossRef] [PubMed]
- Kulikov, A.V.; Luchkina, E.A.; Gogvadze, V.; Zhivotovsky, B. Mitophagy: Link to cancer development and therapy. Biochem. Biophys. Res. Commun. 2017, 482, 432–439. [Google Scholar] [CrossRef] [PubMed]
- Bard, J.A.M.; Goodall, E.A.; Greene, E.R.; Jonsson, E.; Dong, K.C.; Martin, A. Structure and function of the 26S proteasome. Annu. Rev. Biochem. 2018, 87, 697–724. [Google Scholar] [CrossRef] [PubMed]
- Livneh, I.; Cohen-Kaplan, V.; Cohen-Rosenzweig, C.; Avni, N.; Ciechanover, A. The life cycle of the 26S proteasome: From birth, through regulation and function, and onto its death. Cell Res. 2016, 26, 869–885. [Google Scholar] [CrossRef]
- Grice, G.L.; Nathan, J.A. The recognition of ubiquitinated proteins by the proteasome. Cell. Mol. Life Sci. 2016, 73, 3497–3506. [Google Scholar] [CrossRef] [PubMed]
- Cuervo, A.M.; Palmer, A.; Rivett, A.J.; Knecht, E. Degradation of proteasomes by lysosomes in rat liver. Eur. J. Biochem. 1995, 227, 792–800. [Google Scholar] [CrossRef] [PubMed]
- Marshall, R.S.; Li, F.; Gemperline, D.C.; Book, A.J.; Vierstra, R.D. Autophagic degradation of the 26S proteasome is mediated by the dual ATG8/ubiquitin receptor RPN10 in Arabidopsis. Mol. Cell 2015, 58, 1053–1066. [Google Scholar] [CrossRef]
- Cohen-Kaplan, V.; Livneh, I.; Avni, N.; Fabre, B.; Ziv, T.; Kwon, Y.T.; Ciechanover, A. p62- and ubiquitin-dependent stress-induced autophagy of the mammalian 26S proteasome. Proc. Natl. Acad. Sci. USA 2016, 113, E7490–E7499. [Google Scholar] [CrossRef] [PubMed]
- Beese, C.J.; Brynjólfsdóttir, S.H.; Frankel, L.B. Selective autophagy of the protein homeostasis machinery: Ribophagy, proteaphagy and ER-phagy. Front. Cell Dev. Biol. 2020, 7, 373. [Google Scholar] [CrossRef] [PubMed]
- An, H.; Harper, J.W. Systematic analysis of ribophagy in human cells reveals bystander flux during selective autophagy. Nat. Cell Biol. 2018, 20, 135–143. [Google Scholar] [CrossRef] [PubMed]
- Wyant, G.A.; Abu-Remaileh, M.; Frenkel, E.M.; Laqtom, N.N.; Dharamdasani, V.; Lewis, C.A.; Chan, S.H.; Heinze, I.; Ori, A.; Sabatini, D.M. NUFIP1 is a ribosome receptor for starvation-induced ribophagy. Science 2018, 360, 751–758. [Google Scholar] [CrossRef] [PubMed]
- Devenport, S.N.; Shah, Y.M. Functions and implications of autophagy in colon cancer. Cells 2019, 8, 1349. [Google Scholar] [CrossRef]
- Huybrechts, S.J.; Van Veldhoven, P.P.; Brees, C.; Mannaerts, G.P.; Los, G.V.; Fransen, M. Peroxisome dynamics in cultured mammalian cells. Traffic 2009, 10, 1722–1733. [Google Scholar] [CrossRef]
- Mancias, J.D.; Kimmelman, A.C. Mechanisms of selective autophagy in normal physiology and cancer. J. Mol. Biol. 2016, 428, 1659–1680. [Google Scholar] [CrossRef]
- Deosaran, E.; Larsen, K.B.; Hua, R.; Sargent, G.; Wang, Y.; Kim, S.; Lamark, T.; Jauregui, M.; Law, K.; Lippincott-Schwartz, J.; et al. NBR1 acts as an autophagy receptor for peroxisomes. J. Cell Sci. 2013, 126, 939–952. [Google Scholar] [CrossRef]
- Kim, P.K.; Hailey, D.W.; Mullen, R.T.; Lippincott-Schwartz, J. Ubiquitin signals autophagic degradation of cytosolic proteins and peroxisomes. Proc. Natl. Acad. Sci. USA 2008, 105, 20567–20574. [Google Scholar] [CrossRef]
- Eberhart, T.; Kovacs, W.J. Pexophagy in yeast and mammals: An update on mysteries. Histochem. Cell Biol. 2018, 150, 473–488. [Google Scholar] [CrossRef] [PubMed]
- Kounakis, K.; Chaniotakis, M.; Markaki, M.; Tavernarakis, N. Emerging roles of lipophagy in health and disease. Front. Cell Dev. Biol. 2019, 7. [Google Scholar] [CrossRef]
- Rogov, V.; Dötsch, V.; Johansen, T.; Kirkin, V. Interactions between autophagy receptors and ubiquitin-like proteins form the molecular basis for selective autophagy. Mol. Cell 2014, 53, 167–178. [Google Scholar] [CrossRef]
- Singh, R.; Kaushik, S.; Wang, Y.; Xiang, Y.; Novak, I.; Komatsu, M.; Tanaka, K.; Cuervo, A.M.; Czaja, M.J. Autophagy regulates lipid metabolism. Nature 2009, 458, 1131–1135. [Google Scholar] [CrossRef] [PubMed]
- Zhao, T.; Du, H.; Ding, X.; Walls, K.; Yan, C. Activation of mTOR pathway in myeloid-derived suppressor cells stimulates cancer cell proliferation and metastasis in lal−/− mice. Oncogene 2015, 34, 1938–1948. [Google Scholar] [CrossRef] [PubMed]
- Gómez de Cedrón, M.; Ramírez de Molina, A. Microtargeting cancer metabolism: Opening new therapeutic windows based on lipid metabolism. J. Lipid Res. 2016, 57, 193–206. [Google Scholar] [CrossRef] [PubMed]
- Saftig, P.; Klumperman, J. Lysosome biogenesis and lysosomal membrane proteins: Trafficking meets function. Nat. Rev. Mol. Cell Biol. 2009, 10, 623–635. [Google Scholar] [CrossRef] [PubMed]
- Aits, S.; Jäättelä, M. Lysosomal cell death at a glance. J. Cell Sci. 2013, 126, 1905–1912. [Google Scholar] [CrossRef]
- Boya, P.; Kroemer, G. Lysosomal membrane permeabilization in cell death. Oncogene 2008, 27, 6434–6451. [Google Scholar] [CrossRef]
- Maejima, I.; Takahashi, A.; Omori, H.; Kimura, T.; Takabatake, Y.; Saitoh, T.; Yamamoto, A.; Hamasaki, M.; Noda, T.; Isaka, Y.; et al. Autophagy sequesters damaged lysosomes to control lysosomal biogenesis and kidney injury. Embo J. 2013, 32, 2336–2347. [Google Scholar] [CrossRef]
- Papadopoulos, C.; Kravic, B.; Meyer, H. Repair or lysophagy: Dealing with damaged lysosomes. J. Mol. Biol. 2020, 432, 231–239. [Google Scholar] [CrossRef] [PubMed]
- Bo Otto, F.; Thumm, M. Nucleophagy-implications for microautophagy and health. Int. J. Mol. Sci. 2020, 21, 4506. [Google Scholar] [CrossRef]
- Park, Y.-E.; Hayashi, Y.K.; Bonne, G.; Arimura, T.; Noguchi, S.; Nonaka, I.; Nishino, I. Autophagic degradation of nuclear components in mammalian cells. Autophagy 2009, 5, 795–804. [Google Scholar] [CrossRef] [PubMed]
- Zhao, L.; Li, W.; Luo, X.; Sheng, S. The multifaceted roles of nucleophagy in cancer development and therapy. Cell Biol. Int. 2021, 45, 246–257. [Google Scholar] [CrossRef] [PubMed]
- Duve, C.d.; Wattiaux, R. Functions of lysosomes. Annu. Rev. Physiol. 1966, 28, 435–492. [Google Scholar] [CrossRef] [PubMed]
- Yim, W.W.-Y.; Mizushima, N. Lysosome biology in autophagy. Cell Discov. 2020, 6, 6. [Google Scholar] [CrossRef]
- Klionsky, D.J.; Eskelinen, E.L. The vacuole versus the lysosome: When size matters. Autophagy 2014, 10, 185–187. [Google Scholar] [CrossRef]
- Schuck, S. Microautophagy—distinct molecular mechanisms handle cargoes of many sizes. J. Cell Sci. 2020, 133, jcs246322. [Google Scholar] [CrossRef]
- Sahu, R.; Kaushik, S.; Clement, C.C.; Cannizzo, E.S.; Scharf, B.; Follenzi, A.; Potolicchio, I.; Nieves, E.; Cuervo, A.M.; Santambrogio, L. Microautophagy of cytosolic proteins by late endosomes. Dev. Cell 2011, 20, 131–139. [Google Scholar] [CrossRef]
- Mukherjee, A.; Patel, B.; Koga, H.; Cuervo, A.M.; Jenny, A. Selective endosomal microautophagy is starvation-inducible in Drosophila. Autophagy 2016, 12, 1984–1999. [Google Scholar] [CrossRef]
- Uytterhoeven, V.; Lauwers, E.; Maes, I.; Miskiewicz, K.; Melo, M.N.; Swerts, J.; Kuenen, S.; Wittocx, R.; Corthout, N.; Marrink, S.-J.; et al. Hsc70-4 deforms membranes to promote synaptic protein turnover by endosomal microautophagy. Neuron 2015, 88, 735–748. [Google Scholar] [CrossRef]
- Morozova, K.; Clement, C.C.; Kaushik, S.; Stiller, B.; Arias, E.; Ahmad, A.; Rauch, J.N.; Chatterjee, V.; Melis, C.; Scharf, B.; et al. Structural and biological interaction of hsc-70 protein with phosphatidylserine in endosomal microautophagy. J. Biol. Chem. 2016, 291, 18096–18106. [Google Scholar] [CrossRef] [PubMed]
- Chauhan, A.S.; Kumar, M.; Chaudhary, S.; Dhiman, A.; Patidar, A.; Jakhar, P.; Jaswal, P.; Sharma, K.; Sheokand, N.; Malhotra, H.; et al. Trafficking of a multifunctional protein by endosomal microautophagy: Linking two independent unconventional secretory pathways. FASEB J. 2019, 33, 5626–5640. [Google Scholar] [CrossRef] [PubMed]
- Mejlvang, J.; Olsvik, H.; Svenning, S.; Bruun, J.-A.; Abudu, Y.P.; Larsen, K.B.; Brech, A.; Hansen, T.E.; Brenne, H.; Hansen, T.; et al. Starvation induces rapid degradation of selective autophagy receptors by endosomal microautophagy. J. Cell Biol. 2018, 217, 3640–3655. [Google Scholar] [CrossRef]
- Rios, J.; Sequeida, A.; Albornoz, A.; Budini, M. Chaperone mediated autophagy substrates and components in cancer. Front. Oncol. 2021, 10. [Google Scholar] [CrossRef]
- Liao, Z.; Wang, B.; Liu, W.; Xu, Q.; Hou, L.; Song, J.; Guo, Q.; Li, N. Dysfunction of chaperone-mediated autophagy in human diseases. Mol. Cell. Biochem. 2021. [Google Scholar] [CrossRef]
- Cuervo, A.M.; Wong, E. Chaperone-mediated autophagy: Roles in disease and aging. Cell Res. 2014, 24, 92–104. [Google Scholar] [CrossRef]
- Kaushik, S.; Bandyopadhyay, U.; Sridhar, S.; Kiffin, R.; Martinez-Vicente, M.; Kon, M.; Orenstein, S.J.; Wong, E.; Cuervo, A.M. Chaperone-mediated autophagy at a glance. J. Cell Sci. 2011, 124, 495–499. [Google Scholar] [CrossRef]
- Gomes, L.R.; Menck, C.F.M.; Cuervo, A.M. Chaperone-mediated autophagy prevents cellular transformation by regulating MYC proteasomal degradation. Autophagy 2017, 13, 928–940. [Google Scholar] [CrossRef]
- Arias, E.; Cuervo, A.M. Pros and cons of chaperone-mediated autophagy in cancer biology. Trends Endocrinol. Metab. 2020, 31, 53–66. [Google Scholar] [CrossRef]
- Xia, H.-g.; Najafov, A.; Geng, J.; Galan-Acosta, L.; Han, X.; Guo, Y.; Shan, B.; Zhang, Y.; Norberg, E.; Zhang, T.; et al. Degradation of HK2 by chaperone-mediated autophagy promotes metabolic catastrophe and cell death. J. Cell Biol. 2015, 210, 705–716. [Google Scholar] [CrossRef] [PubMed]
- Yang, T.; Ren, C.; Qiao, P.; Han, X.; Wang, L.; Lv, S.; Sun, Y.; Liu, Z.; Du, Y.; Yu, Z. PIM2-mediated phosphorylation of hexokinase 2 is critical for tumor growth and paclitaxel resistance in breast cancer. Oncogene 2018, 37, 5997–6009. [Google Scholar] [CrossRef] [PubMed]
- Condello, M.; Pellegrini, E.; Caraglia, M.; Meschini, S. Targeting autophagy to overcome human diseases. Int. J. Mol. Sci. 2019, 20, 725. [Google Scholar] [CrossRef] [PubMed]
- Chen, Q.; Kang, J.; Fu, C. The independence of and associations among apoptosis, autophagy, and necrosis. Signal. Transduct. Target. Ther. 2018, 3, 18. [Google Scholar] [CrossRef] [PubMed]
- Mariño, G.; Niso-Santano, M.; Baehrecke, E.H.; Kroemer, G. Self-consumption: The interplay of autophagy and apoptosis. Nat. Rev. Mol. Cell Biol. 2014, 15, 81–94. [Google Scholar] [CrossRef] [PubMed]
- Hu, C.-A.A.; White, K.; Torres, S.; Ishak, M.-A.; Sillerud, L.; Miao, Y.; Liu, Z.; Wu, Z.; Sklar, L.; Berwick, M. Chapter 10—Apoptosis and autophagy: The yin–yang of homeostasis in cell death in cancer. In Autophagy: Cancer, Other Pathologies, Inflammation, Immunity, Infection, and Aging; Hayat, M.A., Ed.; Academic Press: Amsterdam, The Netherlands, 2015; pp. 161–181. [Google Scholar]
- Gaglia, G.; Lahav, G. Constant rate of p53 tetramerization in response to DNA damage controls the p53 response. Mol. Syst. Biol. 2014, 10, 753. [Google Scholar] [CrossRef] [PubMed]
- Leszczynska, K.B.; Foskolou, I.P.; Abraham, A.G.; Anbalagan, S.; Tellier, C.; Haider, S.; Span, P.N.; O’Neill, E.E.; Buffa, F.M.; Hammond, E.M. Hypoxia-induced p53 modulates both apoptosis and radiosensitivity via AKT. J. Clin. Investig. 2015, 125, 2385–2398. [Google Scholar] [CrossRef]
- Reid, M.A.; Wang, W.I.; Rosales, K.R.; Welliver, M.X.; Pan, M.; Kong, M. The B55α subunit of PP2A drives a p53-dependent metabolic adaptation to glutamine deprivation. Mol. Cell 2013, 50, 200–211. [Google Scholar] [CrossRef]
- Li, M.; Gao, P.; Zhang, J. Crosstalk between autophagy and apoptosis: Potential and emerging therapeutic targets for cardiac diseases. Int. J. Mol. Sci. 2016, 17, 332. [Google Scholar] [CrossRef]
- Fridman, J.S.; Lowe, S.W. Control of apoptosis by p53. Oncogene 2003, 22, 9030–9040. [Google Scholar] [CrossRef]
- Crighton, D.; Wilkinson, S.; O’Prey, J.; Syed, N.; Smith, P.; Harrison, P.R.; Gasco, M.; Garrone, O.; Crook, T.; Ryan, K.M. DRAM, a p53-induced modulator of autophagy, is critical for apoptosis. Cell 2006, 126, 121–134. [Google Scholar] [CrossRef] [PubMed]
- Tasdemir, E.; Maiuri, M.C.; Galluzzi, L.; Vitale, I.; Djavaheri-Mergny, M.; D’Amelio, M.; Criollo, A.; Morselli, E.; Zhu, C.; Harper, F.; et al. Regulation of autophagy by cytoplasmic p53. Nat. Cell Biol. 2008, 10, 676–687. [Google Scholar] [CrossRef]
- Scherz-Shouval, R.; Weidberg, H.; Gonen, C.; Wilder, S.; Elazar, Z.; Oren, M. p53-dependent regulation of autophagy protein LC3 supports cancer cell survival under prolonged starvation. Proc. Natl. Acad. Sci. USA 2010, 107, 18511–18516. [Google Scholar] [CrossRef] [PubMed]
- Kilbride, S.M.; Prehn, J.H.M. Central roles of apoptotic proteins in mitochondrial function. Oncogene 2013, 32, 2703–2711. [Google Scholar] [CrossRef]
- McKnight, N.C.; Yue, Z. Beclin 1, an essential component and master regulator of PI3K-III in health and disease. Curr. Pathobiol. Rep. 2013, 1, 231–238. [Google Scholar] [CrossRef] [PubMed]
- Decuypere, J.-P.; Parys, J.B.; Bultynck, G. Regulation of the autophagic Bcl-2/Beclin 1 interaction. Cells 2012, 1, 284–312. [Google Scholar] [CrossRef] [PubMed]
- Kang, R.; Zeh, H.J.; Lotze, M.T.; Tang, D. The Beclin 1 network regulates autophagy and apoptosis. Cell Death Differ. 2011, 18, 571–580. [Google Scholar] [CrossRef]
- Maiuri, M.C.; Criollo, A.; Tasdemir, E.; Vicencio, J.M.; Tajeddine, N.; Hickman, J.A.; Geneste, O.; Kroemer, G. BH3-only proteins and BH3 mimetics induce autophagy by competitively disrupting the interaction between Beclin 1 and Bcl-2/Bcl-XL. Autophagy 2007, 3, 374–376. [Google Scholar] [CrossRef]
- Wei, Y.; Pattingre, S.; Sinha, S.; Bassik, M.; Levine, B. JNK1-mediated phosphorylation of Bcl-2 regulates starvation-induced autophagy. Mol. Cell 2008, 30, 678–688. [Google Scholar] [CrossRef]
- Ruvolo, P.P.; Deng, X.; May, W.S. Phosphorylation of Bcl2 and regulation of apoptosis. Leukemia 2001, 15, 515–522. [Google Scholar] [CrossRef]
- Wei, Y.; Sinha, S.C.; Levine, B. Dual role of JNK1-mediated phosphorylation of Bcl-2 in autophagy and apoptosis regulation. Autophagy 2008, 4, 949–951. [Google Scholar] [CrossRef]
- van Delft, M.F.; Huang, D.C.S. How the Bcl-2 family of proteins interact to regulate apoptosis. Cell Res. 2006, 16, 203–213. [Google Scholar] [CrossRef] [PubMed]
- Cooper, K.F. Till death do us part: The marriage of autophagy and apoptosis. Oxidative Med. Cell. Longev. 2018, 2018, 4701275. [Google Scholar] [CrossRef] [PubMed]
- Rubinstein, A.D.; Eisenstein, M.; Ber, Y.; Bialik, S.; Kimchi, A. The autophagy protein Atg12 associates with antiapoptotic Bcl-2 family members to promote mitochondrial apoptosis. Mol. Cell 2011, 44, 698–709. [Google Scholar] [CrossRef]
- Yousefi, S.; Perozzo, R.; Schmid, I.; Ziemiecki, A.; Schaffner, T.; Scapozza, L.; Brunner, T.; Simon, H.-U. Calpain-mediated cleavage of Atg5 switches autophagy to apoptosis. Nat. Cell Biol. 2006, 8, 1124–1132. [Google Scholar] [CrossRef] [PubMed]
- Huang, S.; Okamoto, K.; Yu, C.; Sinicrope, F.A. p62/sequestosome-1 up-regulation promotes ABT-263-induced caspase-8 aggregation/activation on the autophagosome *. J. Biol. Chem. 2013, 288, 33654–33666. [Google Scholar] [CrossRef] [PubMed]
- Young, M.M.; Takahashi, Y.; Khan, O.; Park, S.; Hori, T.; Yun, J.; Sharma, A.K.; Amin, S.; Hu, C.-D.; Zhang, J.; et al. Autophagosomal membrane serves as platform for intracellular Death-inducing Signaling Complex (iDISC)-mediated caspase-8 activation and apoptosis *. J. Biol. Chem. 2012, 287, 12455–12468. [Google Scholar] [CrossRef] [PubMed]
- Gornowicz, A.; Bielawska, A.; Szymanowski, W.; Gabryel-Porowska, H.; Czarnomysy, R.; Bielawski, K. Mechanism of anticancer action of novel berenil complex of platinum(II) combined with anti-MUC1 in MCF-7 breast cancer cells. Oncol. Lett. 2018, 15, 2340–2348. [Google Scholar] [CrossRef]
- Pawłowska, N.; Gornowicz, A.; Bielawska, A.; Surażyński, A.; Szymanowska, A.; Czarnomysy, R.; Bielawski, K. The molecular mechanism of anticancer action of novel octahydropyrazino[2,1-a:5,4-a′]diisoquinoline derivatives in human gastric cancer cells. Investig. New Drugs 2018, 36, 970–984. [Google Scholar] [CrossRef]
- Gornowicz, A.; Pawłowska, N.; Czajkowska, A.; Czarnomysy, R.; Bielawska, A.; Bielawski, K.; Michalak, O.; Staszewska-Krajewska, O.; Kałuża, Z. Biological evaluation of octahydropyrazin[2,1-a:5,4-a′]diisoquinoline derivatives as potent anticancer agents. Tumor Biol. 2017, 39, 1010428317701641. [Google Scholar] [CrossRef] [PubMed]
- Wu, H.; Che, X.; Zheng, Q.; Wu, A.; Pan, K.; Shao, A.; Wu, Q.; Zhang, J.; Hong, Y. Caspases: A molecular switch node in the crosstalk between autophagy and apoptosis. Int. J. Biol. Sci. 2014, 10, 1072–1083. [Google Scholar] [CrossRef]
- Parrish, A.B.; Freel, C.D.; Kornbluth, S. Cellular mechanisms controlling caspase activation and function. Cold Spring Harb. Perspect Biol. 2013, 5. [Google Scholar] [CrossRef] [PubMed]
- Oral, O.; Oz-Arslan, D.; Itah, Z.; Naghavi, A.; Deveci, R.; Karacali, S.; Gozuacik, D. Cleavage of Atg3 protein by caspase-8 regulates autophagy during receptor-activated cell death. Apoptosis 2012, 17, 810–820. [Google Scholar] [CrossRef]
- Wirawan, E.; Vande Walle, L.; Kersse, K.; Cornelis, S.; Claerhout, S.; Vanoverberghe, I.; Roelandt, R.; De Rycke, R.; Verspurten, J.; Declercq, W.; et al. Caspase-mediated cleavage of Beclin-1 inactivates Beclin-1-induced autophagy and enhances apoptosis by promoting the release of proapoptotic factors from mitochondria. Cell Death Dis. 2010, 1, e18. [Google Scholar] [CrossRef] [PubMed]
- Han, J.; Hou, W.; Goldstein, L.A.; Stolz, D.B.; Watkins, S.C.; Rabinowich, H. A complex between Atg7 and caspase-9: A novel mechanism of cross-regulation between autophagy and apoptosis*. J. Biol. Chem. 2014, 289, 6485–6497. [Google Scholar] [CrossRef]
- Mizushima, N.; Levine, B. Autophagy in mammalian development and differentiation. Nat. Cell Biol. 2010, 12, 823–830. [Google Scholar] [CrossRef]
- Wu, W.K.K.; Coffelt, S.B.; Cho, C.H.; Wang, X.J.; Lee, C.W.; Chan, F.K.L.; Yu, J.; Sung, J.J.Y. The autophagic paradox in cancer therapy. Oncogene 2012, 31, 939–953. [Google Scholar] [CrossRef] [PubMed]
- Roy, S.; Debnath, J. Autophagy and tumorigenesis. Semin. Immunopathol. 2010, 32, 383–396. [Google Scholar] [CrossRef] [PubMed]
- Drugs@FDA: FDA-Approved Drugs (Chloroquine). Available online: https://www.accessdata.fda.gov/scripts/cder/daf/index.cfm?event=overview.process&ApplNo=006002 (accessed on 1 March 2021).
- Solomon, V.R.; Lee, H. Chloroquine and its analogs: A new promise of an old drug for effective and safe cancer therapies. Eur. J. Pharm. 2009, 625, 220–233. [Google Scholar] [CrossRef] [PubMed]
- Pellegrini, P.; Strambi, A.; Zipoli, C.; Hägg-Olofsson, M.; Buoncervello, M.; Linder, S.; De Milito, A. Acidic extracellular pH neutralizes the autophagy-inhibiting activity of chloroquine. Autophagy 2014, 10, 562–571. [Google Scholar] [CrossRef]
- Erkisa, M.; Aydinlik, S.; Cevatemre, B.; Aztopal, N.; Akar, R.O.; Celikler, S.; Yilmaz, V.T.; Ari, F.; Ulukaya, E. A promising therapeutic combination for metastatic prostate cancer: Chloroquine as autophagy inhibitor and palladium(II) barbiturate complex. Biochimie 2020, 175, 159–172. [Google Scholar] [CrossRef] [PubMed]
- Lopiccolo, J.; Kawabata, S.; Gills, J.J.; Dennis, P.A. Combining nelfinavir with chloroquine inhibits in vivo growth of human lung cancer xenograft tumors. Vivo 2021, 35, 141–145. [Google Scholar] [CrossRef]
- Wei, T.; Ji, X.; Xue, J.; Gao, Y.; Zhu, X.; Xiao, G. Cyanidin-3-O-glucoside represses tumor growth and invasion in vivo by suppressing autophagy via inhibition of the JNK signaling pathways. Food Funct. 2021, 12, 387–396. [Google Scholar] [CrossRef] [PubMed]
- Wolf, R.; Wolf, D.; Ruocco, V. Antimalarials: Unapproved uses or indications. Clin. Dermatol. 2000, 18, 17–35. [Google Scholar] [CrossRef]
- Drugs@FDA: FDA-Approved Drugs (Hydroxychloroquine). Available online: https://www.accessdata.fda.gov/scripts/cder/daf/index.cfm?event=overview.process&ApplNo=009768 (accessed on 1 March 2021).
- Shi, T.-T.; Yu, X.-X.; Yan, L.-J.; Xiao, H.-T. Research progress of hydroxychloroquine and autophagy inhibitors on cancer. Cancer Chemother. Pharmacol. 2017, 79, 287–294. [Google Scholar] [CrossRef]
- White, E. Deconvoluting the context-dependent role for autophagy in cancer. Nat. Rev. Cancer 2012, 12, 401–410. [Google Scholar] [CrossRef]
- Drug Approval Package (Verteporfin). Available online: https://www.accessdata.fda.gov/drugsatfda_docs/nda/2002/21-119s004_Visudyne.cfm (accessed on 5 March 2021).
- Donohue, E.; Tovey, A.; Vogl, A.W.; Arns, S.; Sternberg, E.; Young, R.N.; Roberge, M. Inhibition of autophagosome formation by the benzoporphyrin derivative verteporfin. J. Biol. Chem. 2011, 286, 7290–7300. [Google Scholar] [CrossRef]
- Donohue, E.; Thomas, A.; Maurer, N.; Manisali, I.; Zeisser-Labouebe, M.; Zisman, N.; Anderson, H.J.; Ng, S.S.W.; Webb, M.; Bally, M.; et al. The autophagy inhibitor verteporfin moderately enhances the antitumor activity of gemcitabine in a pancreatic ductal adenocarcinoma model. J. Cancer 2013, 4, 585–596. [Google Scholar] [CrossRef]
- Gavini, J.; Dommann, N.; Jakob, M.O.; Keogh, A.; Bouchez, L.C.; Karkampouna, S.; Julio, M.K.-d.; Medova, M.; Zimmer, Y.; Schläfli, A.M.; et al. Verteporfin-induced lysosomal compartment dysregulation potentiates the effect of sorafenib in hepatocellular carcinoma. Cell Death Dis. 2019, 10, 749. [Google Scholar] [CrossRef]
- Saini, H.; Sharma, H.; Mukherjee, S.; Chowdhury, S.; Chowdhury, R. Verteporfin disrupts multiple steps of autophagy and regulates p53 to sensitize osteosarcoma cells. Cancer Cell Int. 2021, 21, 52. [Google Scholar] [CrossRef]
- Drug Approval Package (Clarithromycin). Available online: https://www.accessdata.fda.gov/drugsatfda_docs/nda/2000/50775S1_Biaxin.cfm (accessed on 1 March 2021).
- Van Nuffel, A.M.; Sukhatme, V.; Pantziarka, P.; Meheus, L.; Sukhatme, V.P.; Bouche, G. Repurposing Drugs in Oncology (ReDO)-clarithromycin as an anti-cancer agent. Ecancermedicalscience 2015, 9, 513. [Google Scholar] [CrossRef] [PubMed]
- Nakamura, M.; Kikukawa, Y.; Takeya, M.; Mitsuya, H.; Hata, H. Clarithromycin attenuates autophagy in myeloma cells. Int. J. Oncol. 2010, 37, 815–820. [Google Scholar] [CrossRef] [PubMed]
- Kimura, T.; Takabatake, Y.; Takahashi, A.; Isaka, Y. Chloroquine in cancer therapy: A double-edged sword of autophagy. Cancer Res. 2013, 73, 3–7. [Google Scholar] [CrossRef] [PubMed]
- Huggett, M.T.; Jermyn, M.; Gillams, A.; Illing, R.; Mosse, S.; Novelli, M.; Kent, E.; Bown, S.G.; Hasan, T.; Pogue, B.W.; et al. Phase I/II study of verteporfin photodynamic therapy in locally advanced pancreatic cancer. Br. J. Cancer 2014, 110, 1698–1704. [Google Scholar] [CrossRef]
- Seglen, P.O.; Gordon, P.B. 3-Methyladenine: Specific inhibitor of autophagic/lysosomal protein degradation in isolated rat hepatocytes. Proc. Natl. Acad. Sci. USA 1982, 79, 1889–1892. [Google Scholar] [CrossRef]
- Vinod, V.; Padmakrishnan, C.J.; Vijayan, B.; Gopala, S. ‘How can I halt thee?’ The puzzles involved in autophagic inhibition. Pharmacol. Res. 2014, 82, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Petiot, A.; Ogier-Denis, E.; Blommaart, E.F.C.; Meijer, A.J.; Codogno, P. Distinct classes of phosphatidylinositol 3′-kinases are involved in signaling pathways that control macroautophagy in HT-29 cells. J. Biol. Chem. 2000, 275, 992–998. [Google Scholar] [CrossRef] [PubMed]
- Pasquier, B. Autophagy inhibitors. Cell. Mol. Life Sci. 2016, 73, 985–1001. [Google Scholar] [CrossRef]
- Wu, Y.-T.; Tan, H.-L.; Shui, G.; Bauvy, C.; Huang, Q.; Wenk, M.R.; Ong, C.-N.; Codogno, P.; Shen, H.-M. Dual role of 3-methyladenine in modulation of autophagy via different temporal patterns of inhibition on class I and III phosphoinositide 3-kinase. J. Biol. Chem. 2010, 285, 10850–10861. [Google Scholar] [CrossRef]
- Wang, H.; Peng, Y.; Wang, J.; Gu, A.; Li, Q.; Mao, D.; Guo, L. Effect of autophagy on the resveratrol-induced apoptosis of ovarian cancer SKOV3 cells. J. Cell. Biochem. 2019, 120, 7788–7793. [Google Scholar] [CrossRef]
- Zhao, F.; Feng, G.; Zhu, J.; Su, Z.; Guo, R.; Liu, J.; Zhang, H.; Zhai, Y. 3-Methyladenine-enhanced susceptibility to sorafenib in hepatocellular carcinoma cells by inhibiting autophagy. Anti Cancer Drugs 2021. Publish Ahead of Print. [Google Scholar] [CrossRef]
- Ronan, B.; Flamand, O.; Vescovi, L.; Dureuil, C.; Durand, L.; Fassy, F.; Bachelot, M.-F.; Lamberton, A.; Mathieu, M.; Bertrand, T.; et al. A highly potent and selective Vps34 inhibitor alters vesicle trafficking and autophagy. Nat. Chem. Biol. 2014, 10, 1013–1019. [Google Scholar] [CrossRef]
- Pasquier, B. SAR405, a PIK3C3/Vps34 inhibitor that prevents autophagy and synergizes with mTOR inhibition in tumor cells. Autophagy 2015, 11, 725–726. [Google Scholar] [CrossRef] [PubMed]
- Janji, B.; Hasmim, M.; Parpal, S.; De Milito, A.; Berchem, G.; Noman, M.Z. Lighting up the fire in cold tumors to improve cancer immunotherapy by blocking the activity of the autophagy-related protein PIK3C3/VPS34. Autophagy 2020, 16, 2110–2111. [Google Scholar] [CrossRef] [PubMed]
- McAfee, Q.; Zhang, Z.; Samanta, A.; Levi, S.M.; Ma, X.-H.; Piao, S.; Lynch, J.P.; Uehara, T.; Sepulveda, A.R.; Davis, L.E.; et al. Autophagy inhibitor Lys05 has single-agent antitumor activity and reproduces the phenotype of a genetic autophagy deficiency. Proc. Natl. Acad. Sci. USA 2012, 109, 8253–8258. [Google Scholar] [CrossRef] [PubMed]
- DeVorkin, L.; Hattersley, M.; Kim, P.; Ries, J.; Spowart, J.; Anglesio, M.S.; Levi, S.M.; Huntsman, D.G.; Amaravadi, R.K.; Winkler, J.D.; et al. Autophagy inhibition enhances sunitinib efficacy in clear cell ovarian carcinoma. Mol. Cancer Res. 2017, 15, 250–258. [Google Scholar] [CrossRef] [PubMed]
- Baquero, P.; Dawson, A.; Mukhopadhyay, A.; Kuntz, E.M.; Mitchell, R.; Olivares, O.; Ianniciello, A.; Scott, M.T.; Dunn, K.; Nicastri, M.C.; et al. Targeting quiescent leukemic stem cells using second generation autophagy inhibitors. Leukemia 2019, 33, 981–994. [Google Scholar] [CrossRef]
- Carew, J.S.; Espitia, C.M.; Zhao, W.; Han, Y.; Visconte, V.; Phillips, J.; Nawrocki, S.T. Disruption of autophagic degradation with ROC-325 antagonizes renal cell carcinoma pathogenesis. Clin. Cancer Res. 2017, 23, 2869–2879. [Google Scholar] [CrossRef]
- Nawrocki, S.T.; Han, Y.; Visconte, V.; Przychodzen, B.; Espitia, C.M.; Phillips, J.; Anwer, F.; Advani, A.; Carraway, H.E.; Kelly, K.R.; et al. The novel autophagy inhibitor ROC-325 augments the antileukemic activity of azacitidine. Leukemia 2019, 33, 2971–2974. [Google Scholar] [CrossRef]
- Takase, Y.; Saeki, T.; Watanabe, N.; Adachi, H.; Souda, S.; Saito, I. Cyclic GMP phosphodiesterase inhibitors. 2. Requirement of 6-substitution of quinazoline derivatives for potent and selective inhibitory activity. J. Med. Chem. 1994, 37, 2106–2111. [Google Scholar] [CrossRef] [PubMed]
- MacPherson, J.D.; Gillespie, T.D.; Dunkerley, H.A.; Maurice, D.H.; Bennett, B.M. Inhibition of phosphodiesterase 5 selectively reverses nitrate tolerance in the venous circulation. J. Pharmacol. Exp. Ther. 2006, 317, 188–195. [Google Scholar] [CrossRef]
- Liu, J.; Xia, H.; Kim, M.; Xu, L.; Li, Y.; Zhang, L.; Cai, Y.; Norberg, H.V.; Zhang, T.; Furuya, T.; et al. Beclin1 controls the levels of p53 by regulating the deubiquitination activity of USP10 and USP13. Cell 2011, 147, 223–234. [Google Scholar] [CrossRef]
- Correa, R.J.M.; Valdes, Y.R.; Peart, T.M.; Fazio, E.N.; Bertrand, M.; McGee, J.; Préfontaine, M.; Sugimoto, A.; DiMattia, G.E.; Shepherd, T.G. Combination of AKT inhibition with autophagy blockade effectively reduces ascites-derived ovarian cancer cell viability. Carcinogenesis 2014, 35, 1951–1961. [Google Scholar] [CrossRef]
- Shao, S.; Li, S.; Qin, Y.; Wang, X.; Yang, Y.; Bai, H.; Zhou, L.; Zhao, C.; Wang, C. Spautin-1, a novel autophagy inhibitor, enhances imatinib-induced apoptosis in chronic myeloid leukemia. Int. J. Oncol. 2014, 44, 1661–1668. [Google Scholar] [CrossRef]
- Liao, Y.; Guo, Z.; Xia, X.; Liu, Y.; Huang, C.; Jiang, L.; Wang, X.; Liu, J.; Huang, H. Inhibition of EGFR signaling with Spautin-1 represents a novel therapeutics for prostate cancer. J. Exp. Clin. Cancer Res. 2019, 38, 157. [Google Scholar] [CrossRef]
- Gornowicz, A.; Szymanowska, A.; Mojzych, M.; Bielawski, K.; Bielawska, A. The effect of novel 7-methyl-5-phenyl-pyrazolo[4,3-e]tetrazolo[4,5-b][1,2,4]triazine sulfonamide derivatives on apoptosis and autophagy in DLD-1 and HT-29 colon cancer cells. Int. J. Mol. Sci. 2020, 21, 5221. [Google Scholar] [CrossRef]
- Fu, Y.; Hong, L.; Xu, J.; Zhong, G.; Gu, Q.; Gu, Q.; Guan, Y.; Zheng, X.; Dai, Q.; Luo, X.; et al. Discovery of a small molecule targeting autophagy via ATG4B inhibition and cell death of colorectal cancer cells in vitro and in vivo. Autophagy 2019, 15, 295–311. [Google Scholar] [CrossRef]
- Das, C.K.; Banerjee, I.; Mandal, M. Pro-survival autophagy: An emerging candidate of tumor progression through maintaining hallmarks of cancer. Semin. Cancer Biol. 2020, 66, 59–74. [Google Scholar] [CrossRef]
- De Mei, C.; Ercolani, L.; Parodi, C.; Veronesi, M.; Vecchio, C.L.; Bottegoni, G.; Torrente, E.; Scarpelli, R.; Marotta, R.; Ruffili, R.; et al. Dual inhibition of REV-ERBβ and autophagy as a novel pharmacological approach to induce cytotoxicity in cancer cells. Oncogene 2015, 34, 2597–2608. [Google Scholar] [CrossRef]
- Kurdi, A.; Cleenewerck, M.; Vangestel, C.; Lyssens, S.; Declercq, W.; Timmermans, J.-P.; Stroobants, S.; Augustyns, K.; De Meyer, G.R.Y.; Van Der Veken, P.; et al. ATG4B inhibitors with a benzotropolone core structure block autophagy and augment efficiency of chemotherapy in mice. Biochem. Pharmacol. 2017, 138, 150–162. [Google Scholar] [CrossRef]
- Chen, X.-l.; Liu, P.; Zhu, W.-l.; Lou, L.-g. DCZ5248, a novel dual inhibitor of Hsp90 and autophagy, exerts antitumor activity against colon cancer. Acta Pharmacol. Sin. 2021, 42, 132–141. [Google Scholar] [CrossRef]
- Zhang, L.; Qiang, P.; Yu, J.; Miao, Y.; Chen, Z.; Qu, J.; Zhao, Q.; Chen, Z.; Liu, Y.; Yao, X.; et al. Identification of compound CA-5f as a novel late-stage autophagy inhibitor with potent anti-tumor effect against non-small cell lung cancer. Autophagy 2019, 15, 391–406. [Google Scholar] [CrossRef]
- Rebecca, V.W.; Nicastri, M.C.; McLaughlin, N.; Fennelly, C.; McAfee, Q.; Ronghe, A.; Nofal, M.; Lim, C.-Y.; Witze, E.; Chude, C.I.; et al. A unified approach to targeting the lysosome’s degradative and growth signaling roles. Cancer Discov. 2017, 7, 1266–1283. [Google Scholar] [CrossRef]
- Hu, P.; Wang, J.; Qing, Y.; Li, H.; Sun, W.; Yu, X.; Hui, H.; Guo, Q.; Xu, J. FV-429 induces autophagy blockage and lysosome-dependent cell death of T-cell malignancies via lysosomal dysregulation. Cell Death Dis. 2021, 12, 80. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Y.; Wei, L.; Zhang, H.; Dai, Q.; Li, Z.; Yu, B.; Guo, Q.; Lu, N. FV-429 induced apoptosis through ROS-mediated ERK2 nuclear translocation and p53 activation in gastric cancer cells. J. Cell. Biochem. 2015, 116, 1624–1637. [Google Scholar] [CrossRef]
- Miura, K.; Kawano, S.; Suto, T.; Sato, T.; Chida, N.; Simizu, S. Identification of madangamine A as a novel lysosomotropic agent to inhibit autophagy. Bioorganic Med. Chem. 2021, 34, 116041. [Google Scholar] [CrossRef]
- Park, S.R.; Yoo, Y.J.; Ban, Y.-H.; Yoon, Y.J. Biosynthesis of rapamycin and its regulation: Past achievements and recent progress. J. Antibiot. 2010, 63, 434–441. [Google Scholar] [CrossRef]
- Drug Approval Package (Rapamycin). Available online: https://www.accessdata.fda.gov/drugsatfda_docs/nda/99/21083A.cfm (accessed on 1 March 2021).
- Steiner, J.P.; Connolly, M.A.; Valentine, H.L.; Hamilton, G.S.; Dawson, T.M.; Hester, L.; Snyder, S.H. Neurotrophic actions of nonimmunosuppressive analogues of immunosuppressive drugs FK506, rapamycin and cyclosporin A. Nat. Med. 1997, 3, 421–428. [Google Scholar] [CrossRef] [PubMed]
- Douros, J.; Suffness, M. New antitumor substances of natural origin. Cancer Treat. Rev. 1981, 8, 63–87. [Google Scholar] [CrossRef]
- Harrison, D.E.; Strong, R.; Sharp, Z.D.; Nelson, J.F.; Astle, C.M.; Flurkey, K.; Nadon, N.L.; Wilkinson, J.E.; Frenkel, K.; Carter, C.S.; et al. Rapamycin fed late in life extends lifespan in genetically heterogeneous mice. Nature 2009, 460, 392–395. [Google Scholar] [CrossRef]
- Calne, R.Y.; Lim, S.; Samaan, A.; Collier, D.S.J.; Pollard, S.G.; White, D.J.G.; Thiru, S. Rapamycin for immunosuppression in organ allografting. Lancet 1989, 334, 227. [Google Scholar] [CrossRef]
- Demain, A.L. Importance of microbial natural products and the need to revitalize their discovery. J. Ind. Microbiol. Biotechnol. 2014, 41, 185–201. [Google Scholar] [CrossRef]
- Chiarini, F.; Evangelisti, C.; McCubrey, J.A.; Martelli, A.M. Current treatment strategies for inhibiting mTOR in cancer. Trends Pharmacol. Sci. 2015, 36, 124–135. [Google Scholar] [CrossRef]
- Zhou, C.; Zhong, W.; Zhou, J.; Sheng, F.; Fang, Z.; Wei, Y.; Chen, Y.; Deng, X.; Xia, B.; Lin, J. Monitoring autophagic flux by an improved tandem fluorescent-tagged LC3 (mTagRFP-mWasabi-LC3) reveals that high-dose rapamycin impairs autophagic flux in cancer cells. Autophagy 2012, 8, 1215–1226. [Google Scholar] [CrossRef]
- Drug Approval Package (Temsirolimus). Available online: https://www.accessdata.fda.gov/drugsatfda_docs/nda/2007/022088s000TOC.cfm (accessed on 9 March 2021).
- Meng, L.-h.; Zheng, X.F.S. Toward rapamycin analog (rapalog)-based precision cancer therapy. Acta Pharmacol. Sin. 2015, 36, 1163–1169. [Google Scholar] [CrossRef] [PubMed]
- Raymond, E.; Alexandre, J.; Faivre, S.; Vera, K.; Materman, E.; Boni, J.; Leister, C.; Korth-Bradley, J.; Hanauske, A.; Armand, J.-P. Safety and pharmacokinetics of escalated doses of weekly intravenous infusion of CCI-779, a novel mTOR inhibitor, in patients with cancer. J. Clin. Oncol. 2004, 22, 2336–2347. [Google Scholar] [CrossRef]
- Shiratori, H.; Kawai, K.; Hata, K.; Tanaka, T.; Nishikawa, T.; Otani, K.; Sasaki, K.; Kaneko, M.; Murono, K.; Emoto, S.; et al. The combination of temsirolimus and chloroquine increases radiosensitivity in colorectal cancer cells. Oncol. Rep. 2019, 42, 377–385. [Google Scholar] [CrossRef]
- Inamura, S.; Ito, H.; Taga, M.; Tsuchiyama, K.; Hoshino, H.; Kobayashi, M.; Yokoyama, O. Low-dose docetaxel enhanced the anticancer effect of temsirolimus by overcoming autophagy in prostate cancer cells. Anticancer Res. 2019, 39, 5417–5425. [Google Scholar] [CrossRef] [PubMed]
- Kondo, S.; Hirakawa, H.; Ikegami, T.; Uehara, T.; Agena, S.; Uezato, J.; Kinjyo, H.; Kise, N.; Yamashita, Y.; Tanaka, K.; et al. Raptor and rictor expression in patients with human papillomavirus-related oropharyngeal squamous cell carcinoma. Bmc Cancer 2021, 21, 87. [Google Scholar] [CrossRef]
- Trivedi, N.D.; Armstrong, S.; Wang, H.; Hartley, M.; Deeken, J.; Ruth He, A.; Subramaniam, D.; Melville, H.; Albanese, C.; Marshall, J.L.; et al. A phase I trial of the mTOR inhibitor temsirolimus in combination with capecitabine in patients with advanced malignancies. Cancer Med. 2021, 10, 1944–1954. [Google Scholar] [CrossRef] [PubMed]
- Drugs@FDA: FDA-Approved Drugs (Everolimus). Available online: https://www.accessdata.fda.gov/scripts/cder/daf/index.cfm?event=overview.process&varApplNo=022334 (accessed on 10 March 2021).
- Feldman, D.R.; Ged, Y.; Lee, C.-H.; Knezevic, A.; Molina, A.M.; Chen, Y.-B.; Chaim, J.; Coskey, D.T.; Murray, S.; Tickoo, S.K.; et al. Everolimus plus bevacizumab is an effective first-line treatment for patients with advanced papillary variant renal cell carcinoma: Final results from a phase II trial. Cancer 2020, 126, 5247–5255. [Google Scholar] [CrossRef]
- El Guerrab, A.; Bamdad, M.; Bignon, Y.-J.; Penault-Llorca, F.; Aubel, C. Co-targeting EGFR and mTOR with gefitinib and everolimus in triple-negative breast cancer cells. Sci. Rep. 2020, 10, 6367. [Google Scholar] [CrossRef]
- Zhu, M.; Molina, J.R.; Dy, G.K.; Croghan, G.A.; Qi, Y.; Glockner, J.; Hanson, L.J.; Roos, M.M.; Tan, A.D.; Adjei, A.A. A phase I study of the VEGFR kinase inhibitor vatalanib in combination with the mTOR inhibitor, everolimus, in patients with advanced solid tumors. Investig. New Drugs 2020, 38, 1755–1762. [Google Scholar] [CrossRef] [PubMed]
- Werner, E.A.; Bell, J. CCXIV.—The preparation of methylguanidine, and of ββ-dimethylguanidine by the interaction of dicyanodiamide, and methylammonium and dimethylammonium chlorides respectively. J. Chem. Soc. Trans. 1922, 121, 1790–1794. [Google Scholar] [CrossRef]
- Bailey, C.J. Metformin: Historical overview. Diabetologia 2017, 60, 1566–1576. [Google Scholar] [CrossRef] [PubMed]
- Drug Approval Package (Metformin). Available online: https://www.accessdata.fda.gov/drugsatfda_docs/nda/98/020357s010.cfm (accessed on 1 March 2021).
- Pernicova, I.; Korbonits, M. Metformin—mode of action and clinical implications for diabetes and cancer. Nat. Rev. Endocrinol. 2014, 10, 143–156. [Google Scholar] [CrossRef] [PubMed]
- Lu, G.; Wu, Z.; Shang, J.; Xie, Z.; Chen, C.; Zhang, C. The effects of metformin on autophagy. Biomed. Pharmacother. 2021, 137, 111286. [Google Scholar] [CrossRef]
- Tomic, T.; Botton, T.; Cerezo, M.; Robert, G.; Luciano, F.; Puissant, A.; Gounon, P.; Allegra, M.; Bertolotto, C.; Bereder, J.M.; et al. Metformin inhibits melanoma development through autophagy and apoptosis mechanisms. Cell Death Dis. 2011, 2, e199. [Google Scholar] [CrossRef] [PubMed]
- Takahashi, A.; Kimura, F.; Yamanaka, A.; Takebayashi, A.; Kita, N.; Takahashi, K.; Murakami, T. Metformin impairs growth of endometrial cancer cells via cell cycle arrest and concomitant autophagy and apoptosis. Cancer Cell Int. 2014, 14, 53. [Google Scholar] [CrossRef]
- Diamanti-Kandarakis, E.; Economou, F.; Palimeri, S.; Christakou, C. Metformin in polycystic ovary syndrome. Ann. N.Y. Acad. Sci. 2010, 1205, 192–198. [Google Scholar] [CrossRef] [PubMed]
- Patel, R.; Shah, G. Effect of metformin on clinical, metabolic and endocrine outcomes in women with polycystic ovary syndrome: A meta-analysis of randomized controlled trials. Curr. Med. Res. Opin. 2017, 33, 1545–1557. [Google Scholar] [CrossRef]
- Gandini, S.; Puntoni, M.; Heckman-Stoddard, B.M.; Dunn, B.K.; Ford, L.; DeCensi, A.; Szabo, E. Metformin and cancer risk and mortality: A systematic review and meta-analysis taking into account biases and confounders. Cancer Prev. Res. 2014, 7, 867–885. [Google Scholar] [CrossRef]
- Hong, J.; Zhang, Y.; Lai, S.; Lv, A.; Su, Q.; Dong, Y.; Zhou, Z.; Tang, W.; Zhao, J.; Cui, L.; et al. Effects of metformin versus glipizide on cardiovascular outcomes in patients with type 2 diabetes and coronary artery disease. Diabetes Care 2013, 36, 1304–1311. [Google Scholar] [CrossRef]
- Piskovatska, V.; Stefanyshyn, N.; Storey, K.B.; Vaiserman, A.M.; Lushchak, O. Metformin as a geroprotector: Experimental and clinical evidence. Biogerontology 2019, 20, 33–48. [Google Scholar] [CrossRef]
- Bannister, C.A.; Holden, S.E.; Jenkins-Jones, S.; Morgan, C.L.; Halcox, J.P.; Schernthaner, G.; Mukherjee, J.; Currie, C.J. Can people with type 2 diabetes live longer than those without? A comparison of mortality in people initiated with metformin or sulphonylurea monotherapy and matched, non-diabetic controls. Diabetes Obes. Metab. 2014, 16, 1165–1173. [Google Scholar] [CrossRef]
- Drugs@FDA: FDA-Approved Drugs (Miconazole). Available online: https://www.accessdata.fda.gov/scripts/cder/daf/index.cfm?event=overview.process&ApplNo=017450 (accessed on 15 March 2021).
- Piérard, G.E.; Wallace, R.; De Doncker, P. Biometrological assessment of the preventive effect of a miconazole spray powder on athlete’s foot. Clin. Exp. Derm. 1996, 21, 344–346. [Google Scholar] [CrossRef]
- Tanenbaum, L.; Anderson, C.; Rosenberg, M.J.; Akers, W. 1% Sulconazole Cream v 2% Miconazole Cream in the Treatment of Tinea Versicolor: A Double-blind, Multicenter Study. Arch. Dermatol. 1984, 120, 216–219. [Google Scholar] [CrossRef] [PubMed]
- Park, J.-Y.; Jung, H.-J.; Seo, I.; Jha, B.K.; Suh, S.-I.; Suh, M.-H.; Baek, W.-K. Translational suppression of HIF-1α by miconazole through the mTOR signaling pathway. Cell. Oncol. 2014, 37, 269–279. [Google Scholar] [CrossRef] [PubMed]
- Shahbazfar, A.; Zare, P.; Ranjbaran, M.; Tayefi-Nasrabadi, H.; Fakhri, O.; Farshi, Y.; Shadi, S.; Khoshkerdar, A. A survey on anticancer effects of artemisinin, iron, miconazole, and butyric acid on 5637 (bladder cancer) and 4T1 (Breast cancer) cell lines. J. Cancer Res. Ther. 2014, 10, 1057–1062. [Google Scholar] [CrossRef]
- Chang, H.-T.; Chen, W.-C.; Chen, J.-S.; Lu, Y.-C.; Hsu, S.-S.; Wang, J.-L.; Cheng, H.-H.; Cheng, J.-S.; Jiann, B.-P.; Chiang, A.-J.; et al. Effect of miconazole on intracellular Ca2+ levels and proliferation in human osteosarcoma cells. Life Sci. 2005, 76, 2091–2101. [Google Scholar] [CrossRef]
- Jung, H.-J.; Seo, I.; Jha, B.K.; Suh, S.-I.; Baek, W.-K. Miconazole induces autophagic death in glioblastoma cells via reactive oxygen species-mediated endoplasmic reticulum stress. Oncol. Lett. 2021, 21, 335. [Google Scholar] [CrossRef]
- Ho, C.-Y.; Chang, A.-C.; Hsu, C.-H.; Tsai, T.-F.; Lin, Y.-C.; Chou, K.-Y.; Chen, H.-E.; Lin, J.-F.; Chen, P.-C.; Hwang, T.I.-S. Miconazole induces protective autophagy in bladder cancer cells. Environ. Toxicol. 2021, 36, 185–193. [Google Scholar] [CrossRef]
- Jaune, E.; Cavazza, E.; Ronco, C.; Grytsai, O.; Abbe, P.; Tekaya, N.; Zerhouni, M.; Beranger, G.; Kaminski, L.; Bost, F.; et al. Discovery of a new molecule inducing melanoma cell death: Dual AMPK/MELK targeting for novel melanoma therapies. Cell Death Dis. 2021, 12, 64. [Google Scholar] [CrossRef]
- Adamska, A.; Stefanowicz-Hajduk, J.; Ochocka, J.R. Alpha-hederin, the active saponin of Nigella sativa, as an anticancer agent inducing apoptosis in the SKOV-3 cell line. Molecules 2019, 24, 2958. [Google Scholar] [CrossRef]
- Li, J.; Wu, D.-D.; Zhang, J.-X.; Wang, J.; Ma, J.-J.; Hu, X.; Dong, W.-G. Mitochondrial pathway mediated by reactive oxygen species involvement in α-hederin-induced apoptosis in hepatocellular carcinoma cells. World J. Gastroenterol. 2018, 24, 1901–1910. [Google Scholar] [CrossRef] [PubMed]
- Sun, J.; Feng, Y.; Wang, Y.; Ji, Q.; Cai, G.; Shi, L.; Wang, Y.; Huang, Y.; Zhang, J.; Li, Q. α-hederin induces autophagic cell death in colorectal cancer cells through reactive oxygen species dependent AMPK/mTOR signaling pathway activation. Int. J. Oncol. 2019, 54, 1601–1612. [Google Scholar] [CrossRef]
- Ha, H.A.; Chiang, J.H.; Tsai, F.J.; Bau, D.T.; Juan, Y.N.; Lo, Y.H.; Hour, M.J.; Yang, J.S. Novel quinazolinone MJ-33 induces AKT/mTOR-mediated autophagy-associated apoptosis in 5FU-resistant colorectal cancer cells. Oncol. Rep. 2021, 45, 680–692. [Google Scholar] [CrossRef] [PubMed]
- Liu, T.; Zhang, J.; Li, K.; Deng, L.; Wang, H. Combination of an autophagy inducer and an autophagy inhibitor: A smarter strategy emerging in cancer therapy. Front. Pharm. 2020, 11, 408. [Google Scholar] [CrossRef]
- Hour, M.-J.; Tsai, S.-C.; Wu, H.-C.; Lin, M.-W.; Chung, J.G.; Wu, J.-B.; Chiang, J.-H.; Tsuzuki, M.; Yang, J.-S. Antitumor effects of the novel quinazolinone MJ-33: Inhibition of metastasis through the MAPK, AKT, NF-κB and AP-1 signaling pathways in DU145 human prostate cancer cells. Int. J. Oncol. 2012, 41, 1513–1519. [Google Scholar] [CrossRef]
- Omoruyi, S.I.; Ekpo, O.E.; Semenya, D.M.; Jardine, A.; Prince, S. Exploitation of a novel phenothiazine derivative for its anti-cancer activities in malignant glioblastoma. Apoptosis 2020, 25, 261–274. [Google Scholar] [CrossRef]
- De, U.; Son, J.Y.; Sachan, R.; Park, Y.J.; Kang, D.; Yoon, K.; Lee, B.M.; Kim, I.S.; Moon, H.R.; Kim, H.S. A new synthetic histone deacetylase inhibitor, MHY2256, induces apoptosis and autophagy cell death in endometrial cancer cells via p53 acetylation. Int. J. Mol. Sci. 2018, 19, 2743. [Google Scholar] [CrossRef]
- Kim, M.J.; Kang, Y.J.; Sung, B.; Jang, J.Y.; Ahn, Y.R.; Oh, H.J.; Choi, H.; Choi, I.; Im, E.; Moon, H.R.; et al. Novel SIRT inhibitor, MHY2256, induces cell cycle arrest, apoptosis, and autophagic cell death in HCT116 human colorectal cancer cells. Biomol. Ther. 2020, 28, 561–568. [Google Scholar] [CrossRef] [PubMed]
- Hsieh, C.-L.; Huang, H.-S.; Chen, K.-C.; Saka, T.; Chiang, C.-Y.; Chung, L.W.K.; Sung, S.-Y. A novel salicylanilide derivative induces autophagy cell death in castration-resistant prostate cancer via ER stress-activated PERK signaling pathway. Mol. Cancer Ther. 2020, 19, 101–111. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Li, Z.; Song, Y.; Liu, W.; Liu, Z. The mTOR kinase inhibitor CZ415 inhibits human papillary thyroid carcinoma cell growth. Cell. Physiol. Biochem. 2018, 46, 579–590. [Google Scholar] [CrossRef]
- Zhou, X.; Yue, G.G.-L.; Chan, A.M.-L.; Tsui, S.K.-W.; Fung, K.-P.; Sun, H.; Pu, J.; Lau, C.B.-S. Eriocalyxin B, a novel autophagy inducer, exerts anti-tumor activity through the suppression of Akt/mTOR/p70S6K signaling pathway in breast cancer. Biochem. Pharmacol. 2017, 142, 58–70. [Google Scholar] [CrossRef]
| Autophagy Inhibitor | Chemical Structure | Study Type | References |
|---|---|---|---|
| Chloroquine | 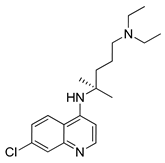 | Preclinical studies: metastatic prostate cancer, NSCLC treatment | [133,134,135] |
| Hydroxychloroquine | 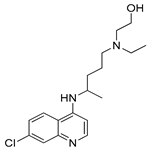 | Clinical trials: therapy of metastatic or advanced pancreatic cancer or HER2 negative breast cancer | NCT04524702, NCT03774472 |
| Verteporfin | 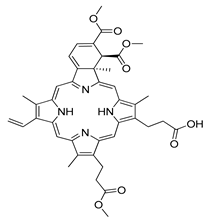 | Preclinical studies: treatment of pancreatic ductal adenocarcinoma, hepatocellular carcinoma or osteosarcoma Clinical trials: recurrent prostate cancer or pancreatic cancer treatment | [141,142,143,144] NCT03067051, NCT03033225 |
| Clarithromycin | 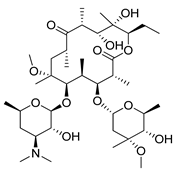 | Clinical trials: multiple myeloma, mucosa-associated lymphoid tissue lymphoma or previously untreated, advanced-stage indolent lymphoma therapy | NCT04302324, NCT04063189, NCT02542657, NCT03031483, NCT00461084 |
| Autophagy Inhibitor | Target Point | Adverse Effects | Therapeutic Combination or Single-Agent Treatment | Tumor Type | References |
|---|---|---|---|---|---|
| Chloroquine | lysosomes | headache, visual disturbances, pruritus or gastrointestinal upset, the risk of acute kidney injury due to kidney cells sensitization to chemotherapy [148] | CQ + temozolomide + radiation therapy | glioblastoma, gliosarcoma and astrocytoma (grade IV) | NCT04397679, NCT02432417 |
| CQ + taxane after anthracycline failure | advanced or metastatic breast cancer | NCT01446016 | |||
| Hydroxychloroquine | lysosomes | no adverse effects observed | HCQ + paclitaxel + carboplatin | advanced or recurrent NSCLC | NCT01649947 |
| decreased hemoglobin, diarrhea, nausea, vomiting, pain, fatigue, rash | HCQ + capecitabine, oxaliplatin and bevacizumab | metastatic colorectal cancer | NCT01006369 | ||
| Verteporfin | autophagy formation | no adverse effects observed | verteporfin photodynamic therapy | advanced pancreatic cancer | [149] |
| Clarithromycin | autophagy flux | anemia, gastrointestinal disorders, dyspnea | clarithromycin + abemaciclib | neoplasm | NCT02117648 |
| upper respiratory infections, sinus/acute otitis | clarithromycin, dexamethasone, lenalidomide therapy after stem cell transplant | multiple myeloma | NCT00445692 | ||
| Clarithromycin | autophagy flux | anemia, neutropenia, diarrhea, vomiting, fever, lung infection, renal insufficiency, dehydration, dyspnea | clarithromycin, dexamethasone, pomalidomide | relapsed or refractory myeloma | NCT01159574 |
| Autophagy Inhibitor | Chemical Structure | Preclinical Studies | References |
|---|---|---|---|
| 3-Methyladenine | 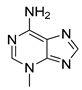 | therapy of human ovarian serous papillary cystadenocarcinoma or hepatocellular carcinoma | [155,156] |
| SAR405 | 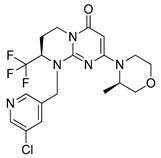 | colorectal and melanoma tumors treatment | [159] |
| Lys05 | 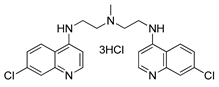 | ovarian carcinoma or CML therapy | [160,161,162] |
| ROC-325 | 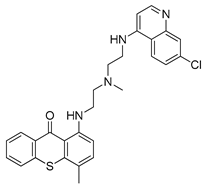 | treatment of AML | [164] |
| Spautin-1 | 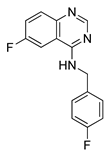 | ovarian cancers, CML or prostate cancer treatment | [168,169,170] |
| MM124 | 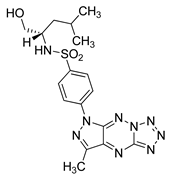 | in vitro studies on colon cancer cells | [171] |
| MM137 | 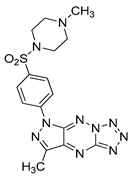 | in vitro studies on colon cancer cells | [171] |
| S130 | 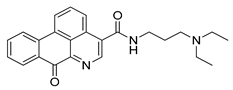 | colorectal cancer therapy | [172] |
| ARN5187 | 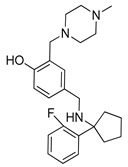 | breast cancer treatment | [173,174] |
| UAMC-2526 | 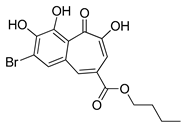 | human colon adenocarcinoma treatment | [175] |
| DCZ5248 | 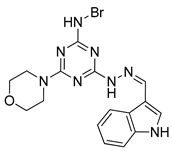 | colon cancer therapy | [176] |
| CA-5f | 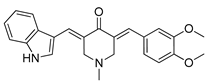 | NSCLC therapy | [177] |
| DQ661 | 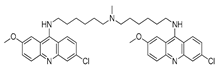 | melanoma, colon cancer or pancreatic cancer therapy | [178] |
| FV-429 | 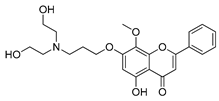 | gastric cancers or T-cell malignancies treatment | [179,180] |
| Madangamine A | 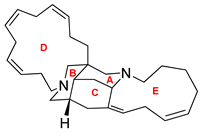 | human cervical carcinoma, human fibrosarcoma and human melanoma therapy | [181] |
| Autophagy Activator | Chemical Structure | Study Type | References |
|---|---|---|---|
| Rapamycin | 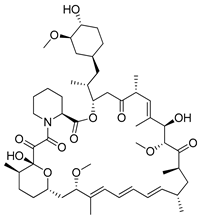 | Clinical trials: therapy of kaposiform hemangioendothelioma in children, bladder cancer, HER2+ metastatic breast cancer, refractory solid tumors, NSCLC and pediatric relapsed or refractory tumors | NCT04077515, NCT02753309, NCT04375813, NCT04736589, NCT02688881, NCT04348292, NCT02574728 |
| Temsirolimus | 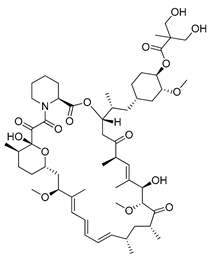 | Preclinical studies: colorectal cancer, prostate cancer, human papillomavirus-related oropharyngeal squamous cell carcinoma or advanced solid tumors treatment Clinical trials: advanced/metastatic malignancies, gynecological malignancies, rare tumors, diffuse intrinsic pontine glioma or solid tumors therapy | [194,195,196,197] NCT01552434, NCT01065662, NCT01396408, NCT02420613, NCT01375829 |
| Everolimus | 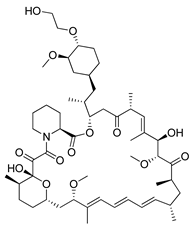 | Preclinical studies: treatment of advanced papillary variant renal cell carcinoma, triple-negative breast cancer or advanced solid tumors Clinical trials: recurrent or progressive ependymoma in children, Hodgkin lymphoma, metastatic transitional cell carcinoma of the urothelium, advanced gynecologic malignancies and breast cancers or recurrent low grade gliomas in young adults and pediatric patients | [199,200,201] NCT02155920, NCT03697408, NCT00805129, NCT03154281, NCT04485559 |
| Metformin | 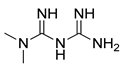 | Clinical trials: breast cancer, colon cancer, thoracic neoplasm or prostate cancer therapy | NCT04559308, NCT04387630, NCT01980823, NCT04741204, NCT03359681, NCT03477162, NCT02176161, NCT02339168 |
| Autophagy Activator | Target Point | Adverse Effects | Therapeutic Combination or Single-Agent Treatment | Tumor Type | References |
|---|---|---|---|---|---|
| Rapamycin | mTOR | blood and lymphatic system disorders e.g., anemia or leukopenia, nausea, fatigue, mucositis, rash | rapamycin + trastuzumab | HER2 receptor positive metastatic breast cancer | NCT00411788 |
| after treatment with high-dose RAPA (6 mg): neutropenia, diarrhea, fever, stomatitis | rapamycin + radical prostatectomy | advanced localized prostate cancer | NCT00311623 | ||
| Temsirolimus | mTOR | anemia, abdominal pain, diarrhea, nausea, fever, non-cardiac chest pain, dyspnea, headache, cough, metabolism and nutrition disorders | temsirolimus + sorafenib | thyroid cancer | NCT01025453 |
| blood and lymphatic system disorders, gastrointestinal disorders, back pain, dizziness, dry skin, pruritus, rash | temsirolimus + bevacizumab | prostate cancer | NCT01083368 | ||
| Temsirolimus | mTOR | mucositis oral, fatigue, dehydration, dyspnea | temsirolimus + cixutumumab | breast cancer | NCT00699491 |
| Everolimus | mTOR | no adverse effects observed | everolimus and pasireotide | thyroid cancer | NCT01270321 |
| anemia, vomiting, lower respiratory tract infection, hypercalcemia, confusional state | everolimus + exemestane | estrogen receptor positive advanced breast cancer | NCT01743560 | ||
| anemia, abdominal pain, diarrhea, mucositis oral, nausea, vomiting, fatigue, rash | everolimus + pazopanib | solid tumor, kidney cancer | NCT01184326 | ||
| Metformin | Beclin 1/mTOR | xerostomia, dysphagia, fatigue, dysgeusia | external beam radiation therapy + metformin | head and neck cancer | NCT03109873 |
| anemia, tinnitus, diarrhea, vomiting, nausea, white blood cell decreased | metformin + cisplatin and radiation therapy | locally advanced head and neck squamous cell carcinoma | NCT02325401 |
| Autophagy Activator | Chemical Structure | Preclinical Studies | References |
|---|---|---|---|
| Miconazole | 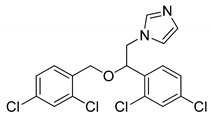 | glioblastoma and bladder cancer therapy | [221,222] |
| CRO15 | 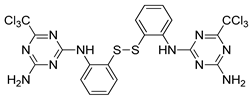 | treatment of melanoma | [223] |
| α-Hederin | 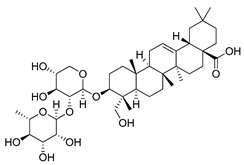 | in vitro and in vivo studies on colorectal cancer cells | [226] |
| MJ-33 |  | in vitro study on HT-29/5FUR cells | [227,229] |
| DS00329 | 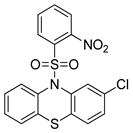 | in vitro study on malignant glioblastoma cells | [230] |
| MHY2256 | 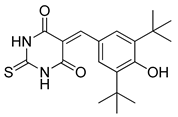 | endometrial and colorectal cancer treatment | [231,232] |
| LCC03 | 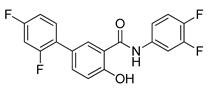 | in vitro and in vivo study on castration-resistant prostate cancer | [233] |
| CZ415 | 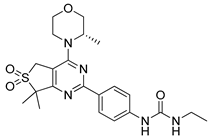 | in vitro and in vivo study on human papillary thyroid carcinoma cells | [234] |
| Eriocalyxin B | 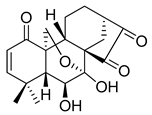 | in vitro and in vivo study on breast cancer | [235] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Buzun, K.; Gornowicz, A.; Lesyk, R.; Bielawski, K.; Bielawska, A. Autophagy Modulators in Cancer Therapy. Int. J. Mol. Sci. 2021, 22, 5804. https://doi.org/10.3390/ijms22115804
Buzun K, Gornowicz A, Lesyk R, Bielawski K, Bielawska A. Autophagy Modulators in Cancer Therapy. International Journal of Molecular Sciences. 2021; 22(11):5804. https://doi.org/10.3390/ijms22115804
Chicago/Turabian StyleBuzun, Kamila, Agnieszka Gornowicz, Roman Lesyk, Krzysztof Bielawski, and Anna Bielawska. 2021. "Autophagy Modulators in Cancer Therapy" International Journal of Molecular Sciences 22, no. 11: 5804. https://doi.org/10.3390/ijms22115804
APA StyleBuzun, K., Gornowicz, A., Lesyk, R., Bielawski, K., & Bielawska, A. (2021). Autophagy Modulators in Cancer Therapy. International Journal of Molecular Sciences, 22(11), 5804. https://doi.org/10.3390/ijms22115804







