Could Histamine H1 Receptor Antagonists Be Used for Treating COVID-19?
Abstract
1. Introduction
2. Drug Repurposing for COVID-19
3. Histamine and COVID-19
4. Inhibition of SARS-CoV-2 by H1 Receptor Antagonists
4.1. In Vitro and In Silico Study
4.2. Patient-Level Study
5. The Role of Other Histamine Receptor Antagonists
6. Potential Anti-SARS-CoV-2 Mechanisms of H1 Receptor Antagonists
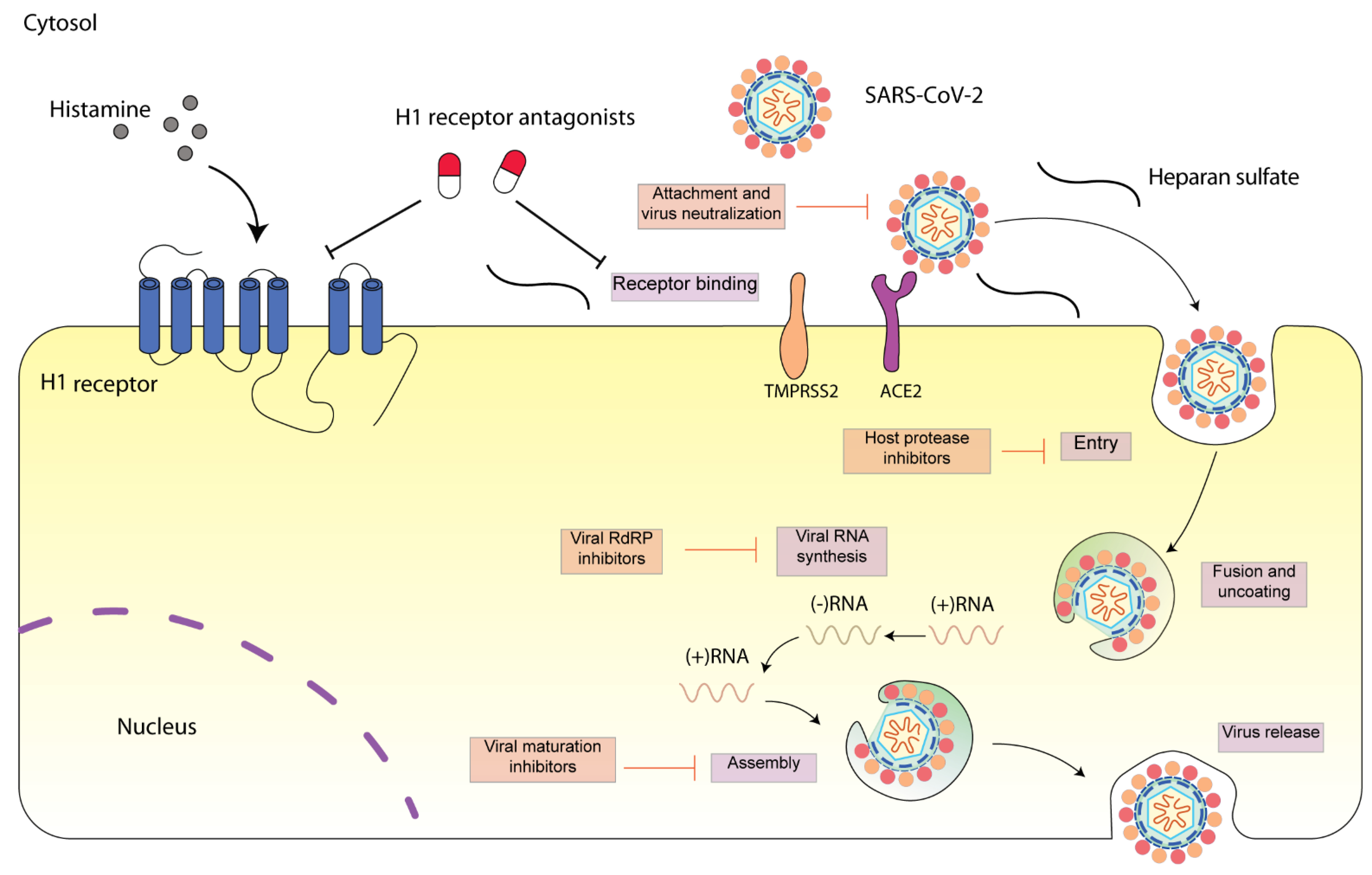
6.1. Intervention of Early Step of SARS-CoV-2 Infection
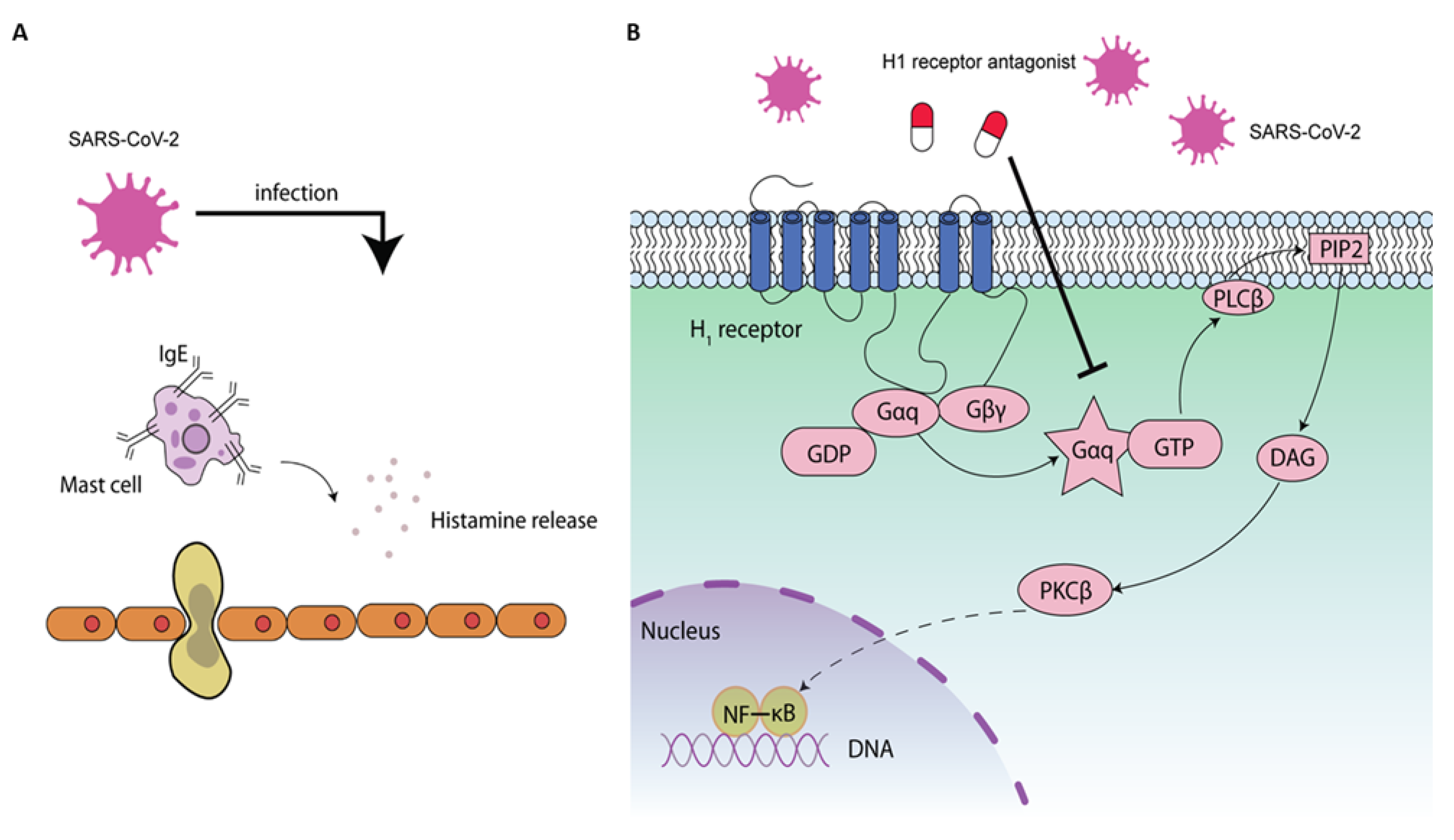
6.2. NF-κB-Mediated Antiviral Activity by H1 Receptor Antagonist
6.3. Other Potential Antiviral Mechanisms
7. Opportunities and Challenges
Author Contributions
Funding
Conflicts of Interest
References
- Huang, C.; Wang, Y.; Li, X.; Ren, L.; Zhao, J.; Hu, Y.; Zhang, L.; Fan, G.; Xu, J.; Gu, X. Clinical features of patients infected with 2019 novel coronavirus in Wuhan, China. Lancet 2020, 395, 497–506. [Google Scholar] [CrossRef]
- Xu, Z.; Shi, L.; Wang, Y.; Zhang, J.; Huang, L.; Zhang, C.; Liu, S.; Zhao, P.; Liu, H.; Zhu, L.; et al. Pathological findings of COVID-19 associated with acute respiratory distress syndrome. Lancet Respir. Med. 2020, 8, 420–422. [Google Scholar] [CrossRef]
- Chan, J.F.; To, K.K.; Tse, H.; Jin, D.Y.; Yuen, K.Y. Interspecies transmission and emergence of novel viruses: Lessons from bats and birds. Trends Microbiol. 2013, 21, 544–555. [Google Scholar] [CrossRef] [PubMed]
- Naqvi, A.A.T.; Fatima, K.; Mohammad, T.; Fatima, U.; Singh, I.K.; Singh, A.; Atif, S.M.; Hariprasad, G.; Hasan, G.M.; Hassan, I. Insights into SARS-CoV-2 genome, structure, evolution, pathogenesis and therapies: Structural genomics approach. Biochim. Biophys. Acta Mol. Basis Dis. 2020, 1866, 165878. [Google Scholar] [CrossRef]
- Zhu, Y.; Feng, F.; Hu, G.; Wang, Y.; Yu, Y.; Zhu, Y.; Xu, W.; Cai, X.; Sun, Z.; Han, W.; et al. A genome-wide CRISPR screen identifies host factors that regulate SARS-CoV-2 entry. Nat. Commun. 2021, 12, 961. [Google Scholar] [CrossRef]
- Hoffmann, M.; Kleine-Weber, H.; Schroeder, S.; Krüger, N.; Herrler, T.; Erichsen, S.; Schiergens, T.S.; Herrler, G.; Wu, N.H.; Nitsche, A.; et al. SARS-CoV-2 cell entry depends on ACE2 and TMPRSS2 and is blocked by a clinically proven protease inhibitor. Cell 2020, 181, 271–280.e8. [Google Scholar] [CrossRef] [PubMed]
- Aggarwal, N.R.; King, L.S.; D’Alessio, F.R. Diverse macrophage populations mediate acute lung inflammation and resolution. Am. J. Physiol. Cell. Mol. Physiol. 2014, 306, L709–L725. [Google Scholar] [CrossRef]
- Junior, J.D.S.M.; Miggiolaro, A.F.R.D.S.; Nagashima, S.; De Paula, C.B.V.; Baena, C.P.; Scharfstein, J.; De Noronha, L. Mast Cells in Alveolar Septa of COVID-19 Patients: A Pathogenic Pathway That May Link Interstitial Edema to Immunothrombosis. Front. Immunol. 2020, 11, 574862. [Google Scholar] [CrossRef]
- Kritas, S.K.; Ronconi, G.; Caraffa, A.; E Gallenga, C.; Ross, R.; Conti, P. Mast cells contribute to coronavirus-induced inflammation: New anti-inflammatory strategy. J. Biol. Regul. Homeost. Agents 2020, 34, 9–14. [Google Scholar] [PubMed]
- Kilinc, E.; Kilinc, Y.B. Mast cell stabilizers as a supportive therapy can contribute to alleviate fatal inflammatory responses and severity of pulmonary complications in COVID-19 infection. Anatol. Clin. J. Med. Sci. 2020, 25, 111–118. [Google Scholar] [CrossRef][Green Version]
- Zhou, Z.; Ren, L.; Zhang, L.; Zhong, J.; Xiao, Y.; Jia, Z.; Guo, L.; Yang, J.; Wang, C.; Jiang, S.; et al. Heightened Innate Immune Responses in the Respiratory Tract of COVID-19 Patients. Cell Host Microbe 2020, 27, 883–890.e2. [Google Scholar] [CrossRef] [PubMed]
- Theoharides, T.C. COVID-19, pulmonary mast cells, cytokine storms, and beneficial actions of luteolin. BioFactors 2020, 46, 306–308. [Google Scholar] [CrossRef]
- Conti, P.; Caraffa, A.; Tetè, G.; Gallenga, C.E.; Ross, R.; Kritas, S.K.; Frydas, I.; Younes, A.; Di Emidio, P.; Ronconi, G. Mast cells activated by SARS-CoV-2 release histamine which increases IL-1 levels causing cytokine storm and inflammatory reaction in COVID-19. J. Biol. Regul. Homeost Agents 2020, 34, 1629–1632. [Google Scholar] [PubMed]
- Malone, R.W.; Tisdall, P.; Fremont-Smith, P.; Liu, Y.; Huang, X.-P.; White, K.M.; Miorin, L.; Moreno, E.; Alon, A.; Delaforge, E.; et al. COVID-19: Famotidine, Histamine, Mast Cells, and Mechanisms. Front. Pharmacol. 2021, 12, 633680. [Google Scholar] [CrossRef] [PubMed]
- Murgolo, N.; Therien, A.G.; Howell, B.; Klein, D.; Koeplinger, K.; Lieberman, L.A.; Adam, G.C.; Flynn, J.; McKenna, P.; Swaminathan, G.; et al. SARS-CoV-2 tropism, entry, replication, and propagation: Considerations for drug discovery and development. PLoS Pathog. 2021, 17, e1009225. [Google Scholar] [CrossRef] [PubMed]
- Clausen, T.M.; Sandoval, D.R.; Spliid, C.B.; Pihl, J.; Perrett, H.R.; Painter, C.D.; Narayanan, A.; Majowicz, S.A.; Kwong, E.M.; McVicar, R.N.; et al. SARS-CoV-2 infection depends on cellular heparan sulfate and ACE2. Cell 2020, 183, 1043–1057.e15. [Google Scholar] [CrossRef]
- Galimberti, F.; McBride, J.; Cronin, M.; Li, Y.; Fox, J.; Abrouk, M.; Herbst, A.; Kirsner, R.S. Evidence-based best practice advice for patients treated with systemic immunosuppressants in relation to COVID-19. Clin. Dermatol. 2020, 38, 775–780. [Google Scholar] [CrossRef]
- Ye, Q.; Wang, B.; Mao, J. The pathogenesis and treatment of the ‘Cytokine Storm’ in COVID-19. J. Infect. 2020, 80, 607–613. [Google Scholar] [CrossRef] [PubMed]
- Jamilloux, Y.; Henry, T.; Belot, A.; Viel, S.; Fauter, M.; El Jammal, T.; Walzer, T.; François, B.; Sève, P. Should we stimulate or suppress immune responses in COVID-19? Cytokine and anti-cytokine interventions. Autoimmun. Rev. 2020, 19, 102567. [Google Scholar] [CrossRef] [PubMed]
- Russell, B.; Moss, C.; Rigg, A.; Van Hemelrijck, M. COVID-19 and treatment with NSAIDs and corticosteroids: Should we be limiting their use in the clinical setting? Ecancer Med. Sci. 2020, 14, 1023. [Google Scholar] [CrossRef] [PubMed]
- Sridharan, G.K.; Kotagiri, R.; Chandiramani, V.H.; Mohan, B.P.; Vegunta, R.; Vegunta, R.; Rokkam, V.R.P. COVID-19 and Avoiding Ibuprofen. How Good Is the Evidence? Am. J. Ther. 2020, 27, e400–e402. [Google Scholar] [CrossRef]
- Horby, P.; Lim, W.S.; Emberson, J.R.; Mafham, M.; Bell, J.L.; Linsell, L.; Staplin, N.; Brightling, C.; Ustianowski, A.; Elmahi, E.; et al. Dexamethasone in hospitalized patients with COVID-19. N. Engl. J. Med. 2021, 384, 693–704. [Google Scholar]
- McGee, S.; Hirschmann, J. Use of corticosteroids in treating infectious diseases. Arch. Intern. Med. 2008, 168, 1034–1046. [Google Scholar] [CrossRef]
- Yang, J.-W.; Fan, L.-C.; Miao, X.-Y.; Mao, B.; Li, M.-H.; Lu, H.-W.; Liang, S.; Xu, J.-F. Corticosteroids for the treatment of human infection with influenza virus: A systematic review and meta-analysis. Clin. Microbiol. Infect. 2015, 21, 956–963. [Google Scholar] [CrossRef] [PubMed]
- Saghazadeh, A.; Rezaei, N. Towards treatment planning of COVID-19: Rationale and hypothesis for the use of multiple immunosuppressive agents: Anti-antibodies, immunoglobulins, and corticosteroids. Int. Immunopharmacol. 2020, 84, 106560. [Google Scholar] [CrossRef] [PubMed]
- Thurmond, R.L.; Gelfand, E.W.; Dunford, P.J. The role of histamine H1 and H4 receptors in allergic inflammation: The search for new antihistamines. Nat. Rev. Drug Discov. 2008, 7, 41–53. [Google Scholar] [CrossRef]
- Togias, A. H1-receptors: Localization and role in airway physiology and in immune functions. J. Allergy Clin. Immunol. 2003, 112 (Suppl. 4), S60–S68. [Google Scholar] [CrossRef]
- Qu, C.; Li, Y.; Li, Y.; Yu, P.; Li, P.; Donkers, J.M.; van de Graaf, S.F.J.; de Man, R.A.; Peppelenbosch, M.P.; Pan, Q. FDA-drug screening identifies deptropine inhibiting hepatitis E virus involving the NF-κB-RIPK1-caspase axis. Antivir. Res. 2019, 170, 104588. [Google Scholar] [CrossRef] [PubMed]
- Lui, C.T. Prescription practice of antihistamines for acute upper respiratory tract infections in pediatric patients in a local emergency department in Hong Kong. World J. Emerg. Med. 2017, 8, 47–54. [Google Scholar] [CrossRef]
- He, S.; Lin, B.; Chu, V.; Hu, Z.; Hu, X.; Xiao, J.; Wang, A.Q.; Schweitzer, C.J.; Li, Q.; Imamura, M.; et al. Repurposing of the antihistamine chlorcyclizine and related compounds for treatment of hepatitis C virus infection. Sci. Transl. Med. 2015, 7, 282ra49. [Google Scholar] [CrossRef] [PubMed]
- Olival, K.J.; Hosseini, P.R.; Zambrana-Torrelio, C.; Ross, N.; Bogich, T.L.; Daszak, P. Host and viral traits predict zoonotic spillover from mammals. Nature 2017, 546, 646–650. [Google Scholar] [CrossRef]
- Holshue, M.L.; DeBolt, C.; Lindquist, S.; Lofy, K.H.; Wiesman, J.; Bruce, H.; Spitters, C.; Ericson, K.; Wilkerson, S.; Tural, A.; et al. First Case of 2019 Novel Coronavirus in the United States. N. Engl. J. Med. 2020, 382, 929–936. [Google Scholar] [CrossRef]
- Cai, Q.; Yang, M.; Liu, D.; Chen, J.; Shu, D.; Xia, J.; Liao, X.; Gu, Y.; Cai, Q.; Yang, Y.; et al. Experimental Treatment with Favipiravir for COVID-19: An Open-Label Control Study. Engineering 2020. [Google Scholar] [CrossRef]
- Pan, Q.; Tilanus, H.W.; Janssen, H.L.; Van Der Laan, L.J.W. Ribavirin enhances interferon-stimulated gene transcription by activation of the interferon-stimulated response element. Hepatology 2011, 53, 1400–1401. [Google Scholar] [CrossRef] [PubMed]
- Gong, W.-J.; Zhou, T.; Wu, S.-L.; Ye, J.-L.; Xu, J.-Q.; Zeng, F.; Su, Y.-Y.; Han, Y.; Lv, Y.-N.; Zhang, Y.; et al. A retrospective analysis of clinical efficacy of ribavirin in adults hospitalized with severe COVID-19. J. Infect. Chemother. 2021, 27, 876–881. [Google Scholar] [CrossRef] [PubMed]
- Elfiky, A.A. Anti-HCV, nucleotide inhibitors, repurposing against COVID-19. Life Sci. 2020, 248, 117477. [Google Scholar] [CrossRef] [PubMed]
- Freedman, H.; Winter, P.; Tuszynski, J.; Tyrrell, D.L.; Houghton, M. A computational approach for predicting off-target toxicity of antiviral ribonucleoside analogues to mitochondrial RNA polymerase. J. Biol. Chem. 2018, 293, 9696–9705. [Google Scholar] [CrossRef]
- Álvarez, M.; Nevot, M.; Mendieta, J.; Martínez, M.A.; Menéndez-Arias, L. Amino acid residues in HIV-2 reverse transcriptase that restrict the development of nucleoside analogue resistance through the excision pathway. J. Biol. Chem. 2018, 293, 2247–2259. [Google Scholar] [CrossRef]
- Bourlière, M.; Gordon, S.C.; Flamm, S.L.; Cooper, C.L.; Ramji, A.; Tong, M.; Ravendhran, N.; Vierling, J.M.; Tran, T.T.; Pianko, S.; et al. Sofosbuvir, Velpatasvir, and Voxilaprevir for Previously Treated HCV Infection. N. Engl. J. Med. 2017, 376, 2134–2146. [Google Scholar] [CrossRef]
- Hayden, F.G.; Sugaya, N.; Hirotsu, N.; Lee, N.; De Jong, M.D.; Hurt, A.C.; Ishida, T.; Sekino, H.; Yamada, K.; Portsmouth, S.; et al. Baloxavir Marboxil for Uncomplicated Influenza in Adults and Adolescents. N. Engl. J. Med. 2018, 379, 913–923. [Google Scholar] [CrossRef]
- Williamson, B.N.; Feldmann, F.; Schwarz, B.; Meade-White, K.; Porter, D.P.; Schulz, J.; van Doremalen, N.; Leighton, I.; Yinda, C.K.; Pérez-Pérez, L.; et al. Clinical benefit of remdesivir in rhesus macaques infected with SARS-CoV-2. Nature 2020, 585, 273–276. [Google Scholar] [CrossRef]
- Tang, W.; Cao, Z.; Han, M.; Wang, Z.; Chen, J.; Sun, W.; Wu, Y.; Xiao, W.; Liu, S.; Chen, E.; et al. Hydroxychloroquine in patients with mainly mild to moderate coronavirus disease 2019: Open label, randomised controlled trial. BMJ 2020, 369, m1849. [Google Scholar] [CrossRef] [PubMed]
- Cao, B.; Wang, Y.; Wen, D.; Liu, W.; Wang, J.; Fan, G.; Ruan, L.; Song, B.; Cai, Y.; Wei, M.; et al. A Trial of Lopinavir–Ritonavir in Adults Hospitalized with Severe Covid-19. N. Engl. J. Med. 2020, 382, 1787–1799. [Google Scholar] [CrossRef]
- Lian, N.; Xie, H.; Lin, S.; Huang, J.; Zhao, J.; Lin, Q. Umifenovir treatment is not associated with improved outcomes in patients with coronavirus disease 2019: A retrospective study. Clin. Microbiol. Infect. 2020, 26, 917–921. [Google Scholar] [CrossRef] [PubMed]
- Naydenova, K.; Muir, K.W.; Wu, L.F.; Zhang, Z.; Coscia, F.; Peet, M.J.; Castro-Hartmann, P.; Qian, P.; Sader, K.; Dent, K.; et al. Structure of the SARS-CoV-2 RNA-dependent RNA polymerase in the presence of favipiravir-RTP. Proc. Natl. Acad. Sci. USA 2021, 118, e2021946118. [Google Scholar] [CrossRef] [PubMed]
- Dittmar, M.; Lee, J.S.; Whig, K.; Segrist, E.; Li, M.; Kamalia, B.; Castellana, L.; Ayyanathan, K.; Truitt, R.; Yang, W.; et al. Drug repurposing screens reveal cell-type specific entry pathways and FDA approved drugs active against SARS-Cov-2. Cell Rep. 2021, 35, 108959. [Google Scholar] [CrossRef] [PubMed]
- Nitulescu, G.M.; Paunescu, H.; Moschos, S.A.; Petrakis, D.; Nitulescu, G.; Ion, G.N.D.; Spandidos, D.A.; Nikolouzakis, T.K.; Drakoulis, N.; Tsatsakis, A. Comprehensive analysis of drugs to treat SARS-CoV-2 infection: Mechanistic insights into current COVID-19 therapies (Review). Int. J. Mol. Med. 2020, 46, 467–488. [Google Scholar] [CrossRef]
- Badary, O.A. Pharmacogenomics and COVID-19: Clinical implications of human genome interactions with repurposed drugs. Pharm. J. 2021, 1–10. [Google Scholar] [CrossRef]
- Nourian, A.; Khalili, H. Sofosbuvir as a potential option for the treatment of COVID-19. Acta Biomed. 2020, 91, 239–241. [Google Scholar]
- Stebbing, J.; Sánchez Nievas, G.; Falcone, M.; Youhanna, S.; Richardson, P.; Ottaviani, S.; Shen, J.X.; Sommerauer, C.; Tiseo, G.; Ghiadoni, L.; et al. JAK inhibition reduces SARS-CoV-2 liver infectivity and modulates inflammatory responses to reduce morbidity and mortality. Sci. Adv. 2021, 7, eabe4724. [Google Scholar] [CrossRef]
- Gordon, D.E.; Jang, G.M.; Bouhaddou, M.; Xu, J.; Obernier, K.; White, K.M.; O’Meara, M.J.; Rezelj, V.V.; Guo, J.Z.; Swaney, D.L.; et al. A SARS-CoV-2 protein interaction map reveals targets for drug repurposing. Nature 2020, 583, 459–468. [Google Scholar] [CrossRef] [PubMed]
- Hippensteel, J.A.; Lariviere, W.B.; Colbert, J.F.; Langouët-Astrié, C.J.; Schmidt, E.P. Heparin as a therapy for COVID-19: Current evidence and future possibilities. Am. J. Physiol. Cell. Mol. Physiol. 2020, 319, L211–L217. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.S.; Shin, E.-C. The type I interferon response in COVID-19: Implications for treatment. Nat. Rev. Immunol. 2020, 20, 585–586. [Google Scholar] [CrossRef] [PubMed]
- Ziegler, C.G.K.; Allon, S.J.; Nyquist, S.K.; Mbano, I.M.; Miao, V.N.; Tzouanas, C.N.; Cao, Y.; Yousif, A.S.; Bals, J.; Hauser, B.M.; et al. SARS-CoV-2 receptor ACE2 is an interferon-stimulated gene in human airway epithelial cells and is detected in specific cell subsets across tissues. Cell 2020, 181, 1016–1035.e19. [Google Scholar] [CrossRef]
- Blume, C.; Jackson, C.L.; Spalluto, C.M.; Legebeke, J.; Nazlamova, L.; Conforti, F.; Perotin, J.-M.; Frank, M.; Butler, J.; Crispin, M.; et al. A novel ACE2 isoform is expressed in human respiratory epithelia and is upregulated in response to interferons and RNA respiratory virus infection. Nat. Genet. 2021, 53, 205–214. [Google Scholar] [CrossRef]
- Maisonnasse, P.; Guedj, J.; Contreras, V.; Behillil, S.; Solas, C.; Marlin, R.; Naninck, T.; Pizzorno, A.; Lemaitre, J.; Gonçalves, A.; et al. Hydroxychloroquine use against SARS-CoV-2 infection in non-human primates. Nature 2020, 585, 584–587. [Google Scholar] [CrossRef] [PubMed]
- Ou, T.; Mou, H.; Zhang, L.; Ojha, A.; Choe, H.; Farzan, M. Hydroxychloroquine-mediated inhibition of SARS-CoV-2 entry is attenuated by TMPRSS2. PLoS Pathog. 2021, 17, e1009212. [Google Scholar] [CrossRef] [PubMed]
- Satarker, S.; Ahuja, T.; Banerjee, M.; Dogra, S.; Agarwal, T.; Nampoothiri, M. Hydroxychloroquine in COVID-19: Potential mechanism of action against SARS-CoV-2. Curr. Pharmacol. Rep. 2020, 6, 203–211. [Google Scholar] [CrossRef] [PubMed]
- Cohen, M.S. Hydroxychloroquine for the Prevention of Covid-19—Searching for Evidence. N. Engl. J. Med. 2020, 383, 585–586. [Google Scholar] [CrossRef] [PubMed]
- Lu, H. Drug treatment options for the 2019-new coronavirus (2019-nCoV). Biosci. Trends 2020, 14, 69–71. [Google Scholar] [CrossRef]
- Horby, P.W.; Mafham, M.; Bell, J.L.; Linsell, L.; Staplin, N.; Emberson, J.; Palfreeman, A.; Raw, J.; Elmahi, E.; Prudon, B.; et al. Lopinavir-ritonavir in patients admitted to hospital with COVID-19 (RECOVERY): A randomised, controlled, open-label, platform trial. Lancet 2020, 396, 1345–1352. [Google Scholar] [CrossRef]
- Huang, D.; Yu, H.; Wang, T.; Yang, H.; Yao, R.; Liang, Z. Efficacy and safety of umifenovir for coronavirus disease 2019 (COVID-19): A systematic review and meta-analysis. J. Med. Virol. 2021, 93, 481–490. [Google Scholar] [CrossRef] [PubMed]
- Tan, Q.; Duan, L.; Ma, Y.; Wu, F.; Huang, Q.; Mao, K.; Xiao, W.; Xia, H.; Zhang, S.; Zhou, E.; et al. Is oseltamivir suitable for fighting against COVID-19: In silico assessment, in vitro and retrospective study. Bioorganic Chem. 2020, 104, 104257. [Google Scholar] [CrossRef] [PubMed]
- Horby, P.W.; Pessoa-Amorim, G.; Peto, L.; Brightling, C.E.; Sarkar, R.; Thomas, K.; Jeebun, V.; Ashish, A.; Tully, R.; Chadwick, D. Tocilizumab in patients admitted to hospital with COVID-19 (RECOVERY): Preliminary results of a randomised, controlled, open-label, platform trial. Medrxiv 2021, 397, 1637–1645. [Google Scholar]
- Lundh, A. Tocilizumab in patients hospitalized with COVID-19 pneumonia. N. Engl. J. Med. 2021, 384, 1473. [Google Scholar]
- Shamburek, R.D.; Schubert, M.L. Control of gastric acid secretion. Histamine H2-receptor antagonists and H+K(+)-ATPase inhibitors. Gastroenterol. Clin. N. Am. 1992, 21, 527–550. [Google Scholar] [CrossRef]
- A Esbenshade, T.; E Browman, K.; Bitner, R.S.; Strakhova, M.; Cowart, M.D.; Brioni, J.D. The histamine H3 receptor: An attractive target for the treatment of cognitive disorders. Br. J. Pharmacol. 2008, 154, 1166–1181. [Google Scholar] [CrossRef] [PubMed]
- Bryce, P.J.; Mathias, C.B.; Harrison, K.L.; Watanabe, T.; Geha, R.S.; Oettgen, H.C. The H1 histamine receptor regulates allergic lung responses. J. Clin. Investig. 2006, 116, 1624–1632. [Google Scholar] [CrossRef]
- Jutel, M.; Watanabe, T.; Klunker, S.; Akdis, M.; Thomet, O.A.R.; Malolepszy, J.; Zak-Nejmark, T.; Koga, R.; Kobayashi, T.; Blaser, K.; et al. Histamine regulates T-cell and antibody responses by differential expression of H1 and H2 receptors. Nat. Cell Biol. 2001, 413, 420–425. [Google Scholar] [CrossRef]
- Tachoua, W.; Kabrine, M.; Mushtaq, M.; Ul-Haq, Z. An in-silico evaluation of COVID-19 main protease with clinically approved drugs. J. Mol. Graph. Model. 2020, 101, 107758. [Google Scholar] [CrossRef]
- Ge, S.; Wang, X.; Hou, Y.; Lv, Y.; Wang, C.; He, H. Repositioning of histamine H(1) receptor antagonist: Doxepin inhibits viropexis of SARS-CoV-2 Spike pseudovirus by blocking ACE2. Eur. J. Pharmacol. 2021, 896, 173897. [Google Scholar] [CrossRef]
- Shah, Y.D.; Stringel, V.; Pavkovic, I.; Kothare, S.V. Doxepin in children and adolescents with symptoms of insomnia: A single-center experience. J. Clin. Sleep Med. 2020, 16, 743–747. [Google Scholar] [CrossRef] [PubMed]
- Reznikov, L.R.; Norris, M.H.; Vashisht, R.; Bluhm, A.P.; Li, D.; Liao, Y.-S.J.; Brown, A.; Butte, A.J.; Ostrov, D.A. Identification of antiviral antihistamines for COVID-19 repurposing. Biochem. Biophys. Res. Commun. 2021, 538, 173–179. [Google Scholar] [CrossRef] [PubMed]
- Jeon, S.; Ko, M.; Lee, J.; Choi, I.; Byun, S.Y.; Park, S.; Shum, D.; Kim, S. Identification of antiviral drug candidates against SARS-CoV-2 from FDA-approved drugs. Antimicrob. Agents Chemother 2020, 64, e00819–e00820. [Google Scholar] [CrossRef] [PubMed]
- Ellinger, B.; Bojkova, D.; Zaliani, A.; Cinatl, J.; Claussen, C.; Westhaus, S.; Keminer, O.; Reinshagen, J.; Kuzikov, M.; Wolf, M.; et al. A SARS-CoV-2 cytopathicity dataset generated by high-content screening of a large drug repurposing collection. Sci. Data 2021, 8, 70. [Google Scholar] [CrossRef]
- Hou, Y.; Ge, S.; Li, X.; Wang, C.; He, H.; He, L. Testing of the inhibitory effects of loratadine and desloratadine on SARS-CoV-2 spike pseudotyped virus viropexis. Chem. Biol. Interact 2021, 338, 109420. [Google Scholar] [CrossRef]
- Touret, F.; Gilles, M.; Barral, K.; Nougairède, A.; van Helden, J.; Decroly, E.; de Lamballerie, X.; Coutard, B. In vitro screening of a FDA approved chemical library reveals potential inhibitors of SARS-CoV-2 replication. Sci. Rep. 2020, 10, 13093. [Google Scholar] [CrossRef]
- Glebov, O.O. Understanding SARS-CoV-2 endocytosis for COVID-19 drug repurposing. FEBS J. 2020, 287, 3664–3671. [Google Scholar] [CrossRef]
- Ennis, M.; Tiligada, K. Histamine receptors and COVID-19. Inflamm. Res. 2021, 70, 67–75. [Google Scholar] [CrossRef] [PubMed]
- Theoharides, T.C.; Antonopoulou, S.; Demopoulos, C.A. Coronavirus 2019, Microthromboses, and Platelet Activating Factor. Clin. Ther. 2020, 42, 1850–1852. [Google Scholar] [CrossRef]
- Kircheis, R.; Haasbach, E.; Lueftenegger, D.; Heyken, W.T.; Ocker, M.; Planz, O. NF-κB pathway as a potential target for treatment of critical stage COVID-19 patients. Front Immunol. 2020, 11, 598444. [Google Scholar] [CrossRef] [PubMed]
- Smit, M.J.; Hoffmann, M.; Timmerman, H.; Leurs, R. Molecular properties and signalling pathways of the histamine H1 receptor. Clin. Exp. Allergy 1999, 29, 19–28. [Google Scholar] [CrossRef] [PubMed]
- Wu, R.L.; Anthes, J.C.; Kreutner, W.; Harris, A.G.; West, R.E., Jr. Desloratadine inhibits constitutive and histamine-stimulated nuclear factor-kappaB activity consistent with inverse agonism at the histamine H1 Receptor. Int. Arch. Allergy Immunol. 2004, 135, 313–318. [Google Scholar] [CrossRef] [PubMed]
- Roumestan, C.; Henriquet, C.; Gougat, C.; Michel, A.; Bichon, F.; Portet, K.; Jaffuel, D.; Mathieu, M. Histamine H1-receptor antagonists inhibit nuclear factor-kappaB and activator protein-1 activities via H1-receptor-dependent and -independent mechanisms. Clin. Exp. Allergy 2008, 38, 947–956. [Google Scholar] [CrossRef]
- Vila-Córcoles, A.; Ochoa-Gondar, O.; Satué-Gracia, E.M.; Torrente-Fraga, C.; Gomez-Bertomeu, F.; Vila-Rovira, A.; Hospital-Guardiola, I.; De Diego-Cabanes, C.; Bejarano-Romero, F.; Basora-Gallisà, J. Influence of prior comorbidities and chronic medications use on the risk of COVID-19 in adults: A population-based cohort study in Tarragona, Spain. BMJ Open 2020, 10, e041577. [Google Scholar] [CrossRef] [PubMed]
- Blanco, J.I.M.; Bonilla, J.A.A.; Homma, S.; Suzuki, K.; Fremont-Smith, P.; Heras, K.V.G.D.L. Antihistamines and azithromycin as a treatment for COVID-19 on primary health care—A retrospective observational study in elderly patients. Pulm. Pharmacol. Ther. 2021, 67, 101989. [Google Scholar] [CrossRef]
- Ii, R.B.H.; Cannon, T.; Rappai, M.; Studdard, J.; Paul, D.; Dooley, T.P. Dual-histamine receptor blockade with cetirizine—famotidine reduces pulmonary symptoms in COVID-19 patients. Pulm. Pharmacol. Ther. 2020, 63, 101942. [Google Scholar] [CrossRef]
- Pagali, S.; Parikh, R.S. Severe urticarial rash as the initial symptom of COVID-19 infection. BMJ Case Rep. 2021, 14, e241793. [Google Scholar] [CrossRef]
- Ishola, A.A.; Joshi, T.; Abdulai, S.I.; Tijjani, H.; Pundir, H.; Chandra, S. Molecular basis for the repurposing of histamine H2-receptor antagonist to treat COVID-19. J. Biomol. Struct. Dyn. 2021, 1–18. [Google Scholar] [CrossRef]
- Freedberg, D.E.; Conigliaro, J.; Wang, T.C.; Tracey, K.J.; Callahan, M.V.; Abrams, J.A.; Sobieszczyk, M.E.; Markowitz, D.D.; Gupta, A.; O’Donnell, M.R.; et al. Famotidine Use Is Associated With Improved Clinical Outcomes in Hospitalized COVID-19 Patients: A Propensity Score Matched Retrospective Cohort Study. Gastroenterology 2020, 159, 1129–1131.e3. [Google Scholar] [CrossRef]
- Krüger, J.; Groß, R.; Conzelmann, C.; Müller, J.A.; Koepke, L.; Sparrer, K.M.; Weil, T.; Schütz, D.; Seufferlein, T.; Barth, T.F. Drug inhibition of SARS-CoV-2 replication in human pluripotent stem cell–derived intestinal organoids. Cell. Mol. Gastroenterol. Hepatol. 2021, 11, 935–948. [Google Scholar] [CrossRef] [PubMed]
- Janowitz, T.; Gablenz, E.; Pattinson, D.; Wang, T.C.; Conigliaro, J.; Tracey, K.; Tuveson, D. Famotidine use and quantitative symptom tracking for COVID-19 in non-hospitalised patients: A case series. Gut 2020, 69, 1592–1597. [Google Scholar] [CrossRef] [PubMed]
- Chazot, P.L.; Johnston, L.; McAuley, E.; Bonner, S. Histamine and Delirium: Current Opinion. Front. Pharmacol. 2019, 10, 299. [Google Scholar] [CrossRef] [PubMed]
- Mirmazloomi, S.R.; Gholinia, N.; Peymani, A. Histamine H4 receptor is a potential target for COVID-19 treatment. Electron. Physician 2020, 12, 7776–7783. [Google Scholar] [CrossRef]
- Cheng, H.; Lear-Rooney, C.M.; Johansen, L.; Varhegyi, E.; Chen, Z.W.; Olinger, G.G.; Rong, L. Inhibition of Ebola and Marburg Virus Entry by G Protein-Coupled Receptor Antagonists. J. Virol. 2015, 89, 9932–9938. [Google Scholar] [CrossRef]
- Rolt, A.; Le, D.; Hu, Z.; Wang, A.Q.; Shah, P.; Singleton, M.; Hughes, E.; E Dulcey, A.; He, S.; Imamura, M.; et al. Preclinical Pharmacological Development of Chlorcyclizine Derivatives for the Treatment of Hepatitis C Virus Infection. J. Infect. Dis. 2018, 217, 1761–1769. [Google Scholar] [CrossRef]
- Pietschmann, T. Clinically Approved Ion Channel Inhibitors Close Gates for Hepatitis C Virus and Open Doors for Drug Repurposing in Infectious Viral Diseases. J. Virol. 2016, 91, 01914–01916. [Google Scholar] [CrossRef]
- Yu, M.; Zhang, T.; Zhang, W.; Sun, Q.; Li, H.; Li, J.P. Elucidating the interactions between heparin/heparan sulfate and SARS-CoV-2-related proteins-an important strategy for developing novel therapeutics for the COVID-19 pandemic. Front. Mol. Biosci. 2020, 7, 628551. [Google Scholar] [CrossRef] [PubMed]
- Jooyandah, F.; Moore, J.; Davies, J.; Jooyandeh, F. The Mechanism of Histamine Binding to Heparin. Int. J. Radiat. Biol. Relat. Stud. Phys. Chem. Med. 1979, 35, 177–182. [Google Scholar] [CrossRef] [PubMed]
- Rabenstein, D.L.; Bratt, P.; Peng, J. Quantitative Characterization of the Binding of Histamine by Heparin†. Biochemistry 1998, 37, 14121–14127. [Google Scholar] [CrossRef] [PubMed]
- Loffredo, M.; Lucero, H.; Chen, D.Y.; O’Connell, A.; Bergqvist, S.; Munawar, A.; Bandara, A.; De Graef, S.; Weeks, S.D.; Douam, F.; et al. The in-vitro effect of famotidine on SARS-CoV-2 proteases and virus replication. Sci. Rep. 2021, 11, 5433. [Google Scholar] [CrossRef] [PubMed]
- Matsubara, M.; Tamura, T.; Ohmori, K.; Hasegawa, K. Histamine H1 receptor antagonist blocks histamine-induced proinflammatory cytokine production through inhibition of Ca2+-dependent protein kinase C, Raf/MEK/ERK and IKK/IκB/NF-κB signal cascades. Biochem. Pharmacol. 2005, 69, 433–449. [Google Scholar] [CrossRef]
- Misto, A.; Provensi, G.; Vozella, V.; Passani, M.B.; Piomelli, D. Mast Cell-Derived Histamine Regulates Liver Ketogenesis via Oleoylethanolamide Signaling. Cell Metab. 2019, 29, 91–102.e5. [Google Scholar] [CrossRef] [PubMed]
- Tabarean, I.V. Histamine receptor signaling in energy homeostasis. Neuropharmacology 2016, 106, 13–19. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Wang, K.-Y.; Tanimoto, A.; Yamada, S.; Guo, X.; Ding, Y.; Watanabe, T.; Watanabe, T.; Kohno, K.; Hirano, K.-I.; Tsukada, H.; et al. Histamine Regulation in Glucose and Lipid Metabolism via Histamine Receptors: Model for Nonalcoholic Steatohepatitis in Mice. Am. J. Pathol. 2010, 177, 713–723. [Google Scholar] [CrossRef]
- Anvari, E.; Wang, X.; Sandler, S.; Welsh, N. The H1-receptor antagonist cetirizine ameliorates high-fat diet-induced glucose intolerance in male C57BL/6 mice, but not diabetes outcome in female non-obese diabetic (NOD) mice. Upsala J. Med. Sci. 2015, 120, 40–46. [Google Scholar] [CrossRef] [PubMed]
- Estelle, F.; Simons, R. H1-receptor antagonists: Safety issues. Ann Allergy Asthma Immunol. 1999, 83, 481–488. [Google Scholar] [PubMed]
- Criado, P.R.; Criado, R.F.; Maruta, C.W.; Machado Filho, C. Histamine, histamine receptors and antihistamines: New concepts. An. Bras. Dermatol. 2010, 85, 195–210. [Google Scholar] [CrossRef]
- Takano, T.; Sakaue, Y.; Sokoda, T.; Sawai, C.; Akabori, S.; Maruo, Y.; Taga, T.; Ohno, M.; Takeuchi, Y. Seizure Susceptibility Due to Antihistamines in Febrile Seizures. Pediatr. Neurol. 2010, 42, 277–279. [Google Scholar] [CrossRef] [PubMed]
- Metz, M.; Doyle, E.; Bindslev-Jensen, C.; Watanabe, T.; Zuberbier, T.; Maurer, M. Effects of Antihistamines on Innate Immune Responses to Severe Bacterial Infection in Mice. Int. Arch. Allergy Immunol. 2011, 155, 355–360. [Google Scholar] [CrossRef] [PubMed]
| Drug | Mode of Action | References |
|---|---|---|
| Remdesivir | Binds to the viral-RNA dependent RNA polymerase (RdRp), terminating transcription of viral RNA | [41] |
| Hydroxychloroquine | Increases the endosomal pH, suppressing the fusion of SARS-CoV-2 with the host cell membrane | [42] |
| Lopinavir/Ritonavir | Inhibits the protein 3CLpro, required for cleaving poly protein into RNA dependent RNA polymerase and helicase | [43] |
| Umifenovir | Blocks the fusion of virus to the cell/endosome by interfering with the hydrogen bond network in the phospholipid | [44] |
| Favipiravir | Destroys the conservative catalytic domain of RdRp, interrupting the nucleotide incorporation | [45] |
| Cyclosporine | Targets cyclophilin rather than calcineurin | [46] |
| Recombinant human ACE2 | Blocks virus cell entry | [47] |
| Oseltamivir | Interferes with viral exocytosis | [48] |
| Sofosbuvir | Binds to the viral-RNA dependent RNA polymerase (RdRp) | [49] |
| Valinomycin | Inhibits S-phase kinase-associated protein | [47] |
| Baricitinib | Targets both viral entry and the cytokine storm | [50] |
| Zotatifin | Inhibits cap-dependent mRNA translation through the host translationmachinery | [51] |
| Heparin | Inhibits viral spike protein-cell receptor interaction and block viral entry | [16,52] |
| Drug | Doses | Clinical Studies |
|---|---|---|
| Cyproheptadine | 4 mg three times a day for 10 days | NCT04876573 |
| Famotidine & N-Acetyl Cysteine | N-Acetyl Cysteine 600~1800 mg three times daily; Famotidine 20~80 mg three times daily | NCT04545008 |
| Celecoxib & Famotidine plus Remdesivir | 80 mg four times daily for 7 days and then 40 mg twice daily for a course of 14 days | NCT04488081 |
| Cetirizine & Famotidine | 10 mg of cetirizine once a day and 20 mg of famotidine twice a day for 10 days | NCT04836806 |
| Famotidine | 80 mg three times a day for a maximum of 14 days | NCT04724720 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Qu, C.; Fuhler, G.M.; Pan, Y. Could Histamine H1 Receptor Antagonists Be Used for Treating COVID-19? Int. J. Mol. Sci. 2021, 22, 5672. https://doi.org/10.3390/ijms22115672
Qu C, Fuhler GM, Pan Y. Could Histamine H1 Receptor Antagonists Be Used for Treating COVID-19? International Journal of Molecular Sciences. 2021; 22(11):5672. https://doi.org/10.3390/ijms22115672
Chicago/Turabian StyleQu, Changbo, Gwenny M. Fuhler, and Yihang Pan. 2021. "Could Histamine H1 Receptor Antagonists Be Used for Treating COVID-19?" International Journal of Molecular Sciences 22, no. 11: 5672. https://doi.org/10.3390/ijms22115672
APA StyleQu, C., Fuhler, G. M., & Pan, Y. (2021). Could Histamine H1 Receptor Antagonists Be Used for Treating COVID-19? International Journal of Molecular Sciences, 22(11), 5672. https://doi.org/10.3390/ijms22115672






