Current Advances on the Important Roles of Enhancer RNAs in Molecular Pathways of Cancer
Abstract
1. Introduction
2. eRNA in Oncogene Expression Regulation and Tumorigenesis
3. eRNA-Associated Signaling Pathways
4. eRNA and Cancer Therapy
5. Computational and Experimental Methods
6. Current eRNA Data Resouces
7. Conclusions and Perspectives
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Conflicts of Interest
References
- Kim, T.-K.; Hemberg, M.; Gray, J.M.; Costa, A.M.; Bear, D.M.; Wu, J.; Harmin, D.A.; Laptewicz, M.; Barbara-Haley, K.; Kuersten, S.; et al. Widespread transcription at neuronal activity-regulated enhancers. Nature 2010, 465, 182–187. [Google Scholar]
- Andersson, R.; The FANTOM Consortium; Gebhard, C.; Miguel-Escalada, I.; Hoof, I.; Bornholdt, J.; Boyd, M.; Chen, Y.; Zhao, X.; Schmidl, C.; et al. An atlas of active enhancers across human cell types and tissues. Nature 2014, 507, 455–461. [Google Scholar] [CrossRef]
- Kaikkonen, M.U.; Spann, N.J.; Heinz, S.; Romanoski, C.E.; Allison, K.A.; Stender, J.D.; Chun, H.B.; Tough, D.F.; Prinjha, R.K.; Benner, C.; et al. Remodeling of the Enhancer Landscape during Macrophage Activation Is Coupled to Enhancer Transcription. Mol. Cell 2013, 51, 310–325. [Google Scholar] [CrossRef]
- Djebali, S.; Davis, C.A.; Merkel, A.; Dobin, A.; Lassmann, T.; Mortazavi, A.; Tanzer, A.; Lagarde, J.; Lin, W.; Schlesinger, F.; et al. Landscape of transcription in human cells. Nature 2012, 489, 101–108. [Google Scholar] [CrossRef]
- Wu, H.; Nord, A.S.; Akiyama, J.A.; Shoukry, M.; Afzal, V.; Rubin, E.M.; Pennacchio, L.A.; Visel, A. Tissue-Specific RNA Expression Marks Distant-Acting Developmental Enhancers. PLoS Genet. 2014, 10, e1004610. [Google Scholar] [CrossRef]
- Franco, H.L.; Nagari, A.; Malladi, V.S.; Li, W.; Xi, Y.; Richardson, D.; Allton, K.L.; Tanaka, K.; Li, J.; Murakami, S.; et al. Enhancer transcription reveals subtype-specific gene expression programs controlling breast cancer pathogenesis. Genome Res. 2018, 28, 159–170. [Google Scholar] [CrossRef]
- Hah, N.; Murakami, S.; Nagari, A.; Danko, C.; Kraus, W.L. Enhancer transcripts mark active estrogen receptor binding sites. Genome Res. 2013, 23, 1210–1223. [Google Scholar] [CrossRef]
- Hah, N.; Benner, C.; Chong, L.-W.; Yu, R.T.; Downes, M.; Evans, R.M. Inflammation-sensitive super enhancers form domains of coordinately regulated enhancer RNAs. Proc. Natl. Acad. Sci. USA 2015, 112, E297–E302. [Google Scholar] [CrossRef]
- Alvarez-Dominguez, J.R.; Knoll, M.; Gromatzky, A.A.; Lodish, H.F. The Super-Enhancer-Derived alncRNA-EC7/Bloodlinc Potentiates Red Blood Cell Development in trans. Cell Rep. 2017, 19, 2503–2514. [Google Scholar]
- Pulakanti, K.; Pinello, L.; Stelloh, C.; Blinka, S.; Allred, J.; Milanovich, S.; Kiblawi, S.; Peterson, J.; Wang, A.; Yuan, G.-C.; et al. Enhancer transcribed RNAs arise from hypomethylated, Tet-occupied genomic regions. Epigenetics 2013, 8, 1303–1320. [Google Scholar] [CrossRef]
- Lee, J.-E.; Park, Y.-K.; Park, S.; Jang, Y.; Waring, N.; Dey, A.; Ozato, K.; Lai, B.; Peng, W.; Ge, K. Brd4 binds to active enhancers to control cell identity gene induction in adipogenesis and myogenesis. Nat. Commun. 2017, 8, 1–12. [Google Scholar] [CrossRef]
- Hnisz, D.; Abraham, B.; Lee, T.I.; Lau, A.; Saint-André, V.; Sigova, A.A.; Hoke, H.A.; Young, R.A. Super-Enhancers in the Control of Cell Identity and Disease. Cell 2013, 155, 934–947. [Google Scholar] [CrossRef]
- Hsieh, C.-L.; Fei, T.; Chen, Y.; Li, T.; Gao, Y.; Wang, X.; Sun, T.; Sweeney, C.J.; Lee, G.-S.M.; Chen, S.; et al. Enhancer RNAs participate in androgen receptor-driven looping that selectively enhances gene activation. Proc. Natl. Acad. Sci. USA 2014, 111, 7319–7324. [Google Scholar] [CrossRef]
- Schaukowitch, K.; Joo, J.-Y.; Liu, X.; Watts, J.K.; Martinez, C.; Kim, T.-K. Enhancer RNA Facilitates NELF Release from Immediate Early Genes. Mol. Cell 2014, 56, 29–42. [Google Scholar] [CrossRef]
- Lai, F.; Orom, U.A.; Cesaroni, M.; Beringer, M.; Taatjes, D.J.; Blobel, G.A.; Shiekhattar, R. Activating RNAs associate with Mediator to enhance chromatin architecture and transcription. Nature 2013, 494, 497–501. [Google Scholar] [CrossRef]
- Lai, F.; Shiekhattar, R. Enhancer RNAs: The new molecules of transcription. Curr. Opin. Genet. Dev. 2014, 25, 38–42. [Google Scholar] [CrossRef]
- Rennie, S.; Dalby, M.; Lloret-Llinares, M.; Bakoulis, S.; Vaagensø, C.D.; Jensen, T.H.; Andersson, R. Transcription start site analysis reveals widespread divergent transcription in D. melanogaster and core promoter-encoded enhancer activities. Nucleic Acids Res. 2018, 46, 5455–5469. [Google Scholar] [CrossRef]
- Melo, C.A.; Drost, J.; Wijchers, P.J.; van de Werken, H.; de Wit, E.; Oude Vrielink, J.A.; Elkon, R.; Melo, S.A.; Léveillé, N.; Kalluri, R.; et al. eRNAs Are Required for p53-Dependent Enhancer Activity and Gene Transcription. Mol. Cell 2013, 49, 524–535. [Google Scholar]
- Jiang, Y.; Jiang, Y.-Y.; Xie, J.-J.; Mayakonda, A.; Hazawa, M.; Chen, L.; Xiao, J.-F.; Li, C.-Q.; Huang, M.-L.; Ding, L.-W.; et al. Co-activation of super-enhancer-driven CCAT1 by TP63 and SOX2 promotes squamous cancer progression. Nat. Commun. 2018, 9, 1–13. [Google Scholar] [CrossRef]
- Li, W.; Notani, D.; Ma, Q.; Tanasa, B.; Nunez, E.; Chen, A.Y.; Merkurjev, D.; Zhang, J.; Ohgi, K.; Song, X.; et al. Functional roles of enhancer RNAs for oestrogen-dependent transcriptional activation. Nature 2013, 498, 516–520. [Google Scholar] [CrossRef]
- Xiang, J.-F.; Yin, Q.-F.; Chen, T.; Zhang, Y.; Zhang, X.-O.; Wu, Z.; Zhang, S.; Wang, H.-B.; Ge, J.; Lu, X.; et al. Human colorectal cancer-specific CCAT1-L lncRNA regulates long-range chromatin interactions at the MYC locus. Cell Res. 2014, 24, 513–531. [Google Scholar] [CrossRef]
- Ling, J.; Ainol, L.; Zhang, L.; Yu, X.; Pi, W.; Tuan, D. HS2 Enhancer Function Is Blocked by a Transcriptional Terminator Inserted between the Enhancer and the Promoter. J. Biol. Chem. 2004, 279, 51704–51713. [Google Scholar] [CrossRef]
- Racko, D.; Benedetti, F.; Dorier, J.; Stasiak, A. Transcription-induced supercoiling as the driving force of chromatin loop ex-trusion during formation of TADs in interphase chromosomes. Nucleic Acids Res. 2018, 46, 1648–1660. [Google Scholar]
- Liang, J.; Zhou, H.; Gerdt, C.; Tan, M.; Colson, T.; Kaye, K.M.; Kieff, E.; Zhao, B. Epstein–Barr virus super-enhancer eRNAs are essential for MYC oncogene expression and lymphoblast proliferation. Proc. Natl. Acad. Sci. USA 2016, 113, 14121–14126. [Google Scholar] [CrossRef]
- Lewis, M.W.; Li, S.; Franco, H.L. Transcriptional control by enhancers and enhancer RNAs. Transcription 2019, 10, 171–186. [Google Scholar] [CrossRef]
- Mousavi, K.; Zare, H.; Dell’Orso, S.; Grontved, L.; Gutierrez-Cruz, G.; Derfoul, A.; Hager, G.L.; Sartorelli, V. eRNAs Promote Transcription by Establishing Chromatin Accessibility at Defined Genomic Loci. Mol. Cell 2013, 51, 606–617. [Google Scholar] [CrossRef]
- Zhang, Z.; Lee, J.-H.; Ruan, H.; Ye, Y.; Krakowiak, J.; Hu, Q.; Xiang, Y.; Gong, J.; Zhou, B.; Wang, L.; et al. Transcriptional landscape and clinical utility of enhancer RNAs for eRNA-targeted therapy in cancer. Nat. Commun. 2019, 10, 4562. [Google Scholar]
- Qin, N.; Ma, Z.; Wang, C.; Zhang, E.; Li, Y.; Huang, M.; Chen, C.; Zhang, C.; Fan, J.; Gu, Y.; et al. Comprehensive characterization of functional eRNAs in lung ade-nocarcinoma reveals novel regulators and a prognosis-related molecular subtype. Theranostics 2020, 10, 11264–11277. [Google Scholar]
- Colebatch, A.J.; Dobrovic, A.; A Cooper, W. TERT gene: Its function and dysregulation in cancer. J. Clin. Pathol. 2019, 72, 281–284. [Google Scholar] [CrossRef]
- Zhao, Y.; Wang, L.; Ren, S.; Wang, L.; Blackburn, P.R.; McNulty, M.S.; Gao, X.; Qiao, M.; Vessella, R.L.; Kohli, M.; et al. Acti-vation of P-TEFb by Androgen Receptor-Regulated Enhancer RNAs in Castration-Resistant Prostate Cancer. Cell Rep. 2016, 15, 599–610. [Google Scholar]
- Zhang, X.; Pang, P.; Jiang, M.; Cao, Q.; Li, H.; Xu, Y.; Li, Y.; Chen, X.; Han, J. eRNAs and Superenhancer lncRNAs Are Functional in Human Prostate Cancer. Dis. Markers 2020, 2020, 1–17. [Google Scholar] [CrossRef]
- Guo, Y.; Pan, W.; Liu, S.; Shen, Z.; Xu, Y.; Hu, L. ERK/MAPK signalling pathway and tumorigenesis (Review). Exp. Ther. Med. 2020, 19, 1997–2007. [Google Scholar] [CrossRef]
- Caunt, C.J.; Keyse, S.M. Dual-specificity MAP kinase phosphatases (MKPs): Shaping the outcome of MAP kinase signalling. FEBS J. 2013, 280, 489–504. [Google Scholar]
- Hoxhaj, G.; Manning, B.D. The PI3K–AKT network at the interface of oncogenic signalling and cancer metabolism. Nat. Rev. Cancer 2019, 20, 74–88. [Google Scholar] [CrossRef]
- Yang, J.; Nie, J.; Ma, X.; Wei, Y.; Peng, Y.; Wei, X. Targeting PI3K in cancer: Mechanisms and advances in clinical trials. Mol. Cancer 2019, 18, 1–28. [Google Scholar] [CrossRef]
- Jiang, N.; Dai, Q.; Su, X.; Fu, J.; Feng, X.; Peng, J. Role of PI3K/AKT pathway in cancer: The framework of malignant behavior. Mol. Biol. Rep. 2020, 47, 4587–4629. [Google Scholar] [CrossRef]
- Ledoux, A.C.; Perkins, N.D. NF-κB and the cell cycle. Biochem. Soc. Trans. 2014, 42, 76–81. [Google Scholar]
- Oeckinghaus, A.; Ghosh, S. The NF-kappaB family of transcription factors and its regulation. Cold Spring Harb. Perspect. Biol. 2009, 1, a000034. [Google Scholar]
- Chen, L.-F.; Greene, W.C. Shaping the nuclear action of NF-κB. Nat. Rev. Mol. Cell Biol. 2004, 5, 392–401. [Google Scholar]
- Vallabhapurapu, S.; Karin, M. Regulation and function of NF-kappaB transcription factors in the immune system. Annu. Rev. Immunol. 2009, 27, 693–733. [Google Scholar]
- Sun, S.C. Non-canonical NF-κB signaling pathway. Cell Res. 2011, 21, 71–85. [Google Scholar]
- Heward, J.A.; Roux, B.T.; Lindsay, M.A. Divergent signalling pathways regulate lipopolysaccharide-induced eRNA ex-pression in human monocytic THP1 cells. FEBS Lett. 2015, 589, 396–406. [Google Scholar]
- Newton, K.; Dixit, V.M. Signaling in Innate Immunity and Inflammation. Cold Spring Harb. Perspect. Biol. 2012, 4, a006049. [Google Scholar] [CrossRef]
- McCleland, M.L.; Mesh, K.; Lorenzana, E.; Chopra, V.S.; Segal, E.; Watanabe, C.; Haley, B.; Mayba, O.; Yaylaoglu, M.; Gnad, F.; et al. CCAT1 is an enhancer-templated RNA that predicts BET sensitivity in colorectal cancer. J. Clin. Investig. 2016, 126, 639–652. [Google Scholar] [CrossRef]
- Younger, S.T.; Rinn, J.L. ’Lnc’-ing enhancers to MYC regulation. Cell Res. 2014, 24, 643–644. [Google Scholar] [CrossRef]
- Ferraldeschi, R.; Welti, J.; Luo, J.; Attard, G.; De Bono, J.S. Targeting the androgen receptor pathway in castration-resistant prostate cancer: Progresses and prospects. Oncogene 2015, 34, 1745–1757. [Google Scholar] [CrossRef]
- Heinlein, C.A.; Chang, C. Androgen receptor in prostate cancer. Endocr. Rev. 2004, 25, 276–308. [Google Scholar]
- Nelson, P.S. Molecular States Underlying Androgen Receptor Activation: A Framework for Therapeutics Targeting Androgen Signaling in Prostate Cancer. J. Clin. Oncol. Off. J. Am. Soc. Clin. Oncol. 2012, 30, 644–646. [Google Scholar]
- Debes, J.D.; Tindall, D.J. Mechanisms of Androgen-Refractory Prostate Cancer. N. Engl. J. Med. 2004, 351, 1488–1490. [Google Scholar] [CrossRef]
- Ding, M.; Zhan, H.; Liao, X.; Li, A.; Zhong, Y.; Gao, Q.; Liu, Y.; Huang, W.; Cai, Z. Enhancer RNA—P2RY2e induced by estrogen promotes malignant behaviors of bladder cancer. Int. J. Biol. Sci. 2018, 14, 1268–1276. [Google Scholar] [CrossRef]
- Che, W.; Ye, S.; Cai, A.; Cui, X.; Sun, Y. CRISPR-Cas13a Targeting the Enhancer RNA-SMAD7e Inhibits Bladder Cancer Development Both in vitro and in vivo. Front. Mol. Biosci. 2020, 7, 607740. [Google Scholar] [CrossRef]
- Maeda, E.; Akahane, M.; Kiryu, S.; Kato, N.; Yoshikawa, T.; Hayashi, N.; Aoki, S.; Minami, M.; Uozaki, H.; Fukayama, M.; et al. Spectrum of Epstein-Barr virus-related diseases: A pictorial review. Jpn. J. Radiol. 2009, 27, 4–19. [Google Scholar] [CrossRef]
- Rezk, S.A.; Zhao, X.; Weiss, L.M. Epstein-Barr virus (EBV)–associated lymphoid proliferations, a 2018 update. Hum. Pathol. 2018, 79, 18–41. [Google Scholar] [CrossRef]
- Hussain, T.; Mulherkar, R. Lymphoblastoid Cell lines: A Continuous in Vitro Source of Cells to Study Carcinogen Sensitivity and DNA Repair. Int. J. Mol. Cell. Med. 2012, 1, 75–87. [Google Scholar]
- Guo, X.-Y.; Zhong, S.; Wang, Z.-N.; Xie, T.; Duan, H.; Zhang, J.-Y.; Zhang, G.-H.; Liang, L.; Cui, R.; Hu, H.-R.; et al. Immunogenomic Profiling Demonstrate AC003092.1 as an Immune-Related eRNA in Glio-blastoma Multiforme. Front. Genet. 2021, 12, 633812. [Google Scholar]
- Gu, X.; Wang, L.; Boldrup, L.; Coates, P.J.; Fahraeus, R.; Sgaramella, N.; Wilms, T.; Nylander, K. AP001056.1, A Progno-sis-Related Enhancer RNA in Squamous Cell Carcinoma of the Head and Neck. Cancers 2019, 11, 347. [Google Scholar]
- Plumb, J.A.; Finn, P.W.; Williams, R.J.; Bandara, M.J.; Romero, M.R.; Watkins, C.J.; La Thangue, N.B.; Brown, R. Pharmacodynamic response and inhibition of growth of human tumor xenografts by the novel histone deacetylase inhibitor PXD. Mol. Cancer Ther. 2003, 2, 721–728. [Google Scholar]
- Gardini, A. Global Run-On Sequencing (GRO-Seq). In Enhancer RNAs; Clifton, N.J., Ed.; Humana Press: New York, NY, USA, 2017; pp. 111–120. [Google Scholar]
- Mahat, D.B.; Kwak, H.; Booth, G.T.; Jonkers, I.H.; Danko, C.G.; Patel, R.K.; Waters, C.T.; Munson, K.; Core, L.J.; Lis, J.T. Base-pair-resolution genome-wide mapping of active RNA polymerases using precision nuclear run-on (PRO-seq). Nat. Protoc. 2016, 11, 1455–1476. [Google Scholar] [CrossRef]
- Core, L.J.; Martins, A.L.; Danko, C.G.; Waters, C.T.; Siepel, A.; Lis, J.T. Analysis of nascent RNA identifies a unified ar-chitecture of initiation regions at mammalian promoters and enhancers. Nat. Genet. 2014, 46, 1311–1320. [Google Scholar]
- Core, L.J.; Waterfall, J.; Lis, J.T. Nascent RNA Sequencing Reveals Widespread Pausing and Divergent Initiation at Human Promoters. Science 2008, 322, 1845–1848. [Google Scholar] [CrossRef]
- Lam, M.T.; Cho, H.; Lesch, H.P.; Gosselin, D.; Heinz, S.; Tanaka-Oishi, Y.; Benner, C.; Kaikkonen, M.U.; Kim, A.S.; Kosaka, M.; et al. Rev-Erbs repress macrophage gene ex-pression by inhibiting enhancer-directed transcription. Nature 2013, 498, 511–515. [Google Scholar]
- Lai, F.; Gardini, A.; Zhang, A.; Shiekhattar, R. Integrator mediates the biogenesis of enhancer RNAs. Nature 2015, 525, 399–403. [Google Scholar] [CrossRef]
- Zhang, Z.; Hong, W.; Ruan, H.; Jing, Y.; Li, S.; Liu, Y.; Wang, J.; Li, W.; Diao, L.; Han, L. HeRA: An atlas of enhancer RNAs across human tissues. Nucleic Acids Res. 2021, 49, D932–D938. [Google Scholar] [CrossRef]
- Chen, H.; Liang, H. A High-Resolution Map of Human Enhancer RNA Loci Characterizes Super-enhancer Activities in Cancer. Cancer Cell 2020, 38, 701–715. [Google Scholar] [CrossRef]
| eRNA | Cancer Type | Correlated Signaling Pathway | Reference |
|---|---|---|---|
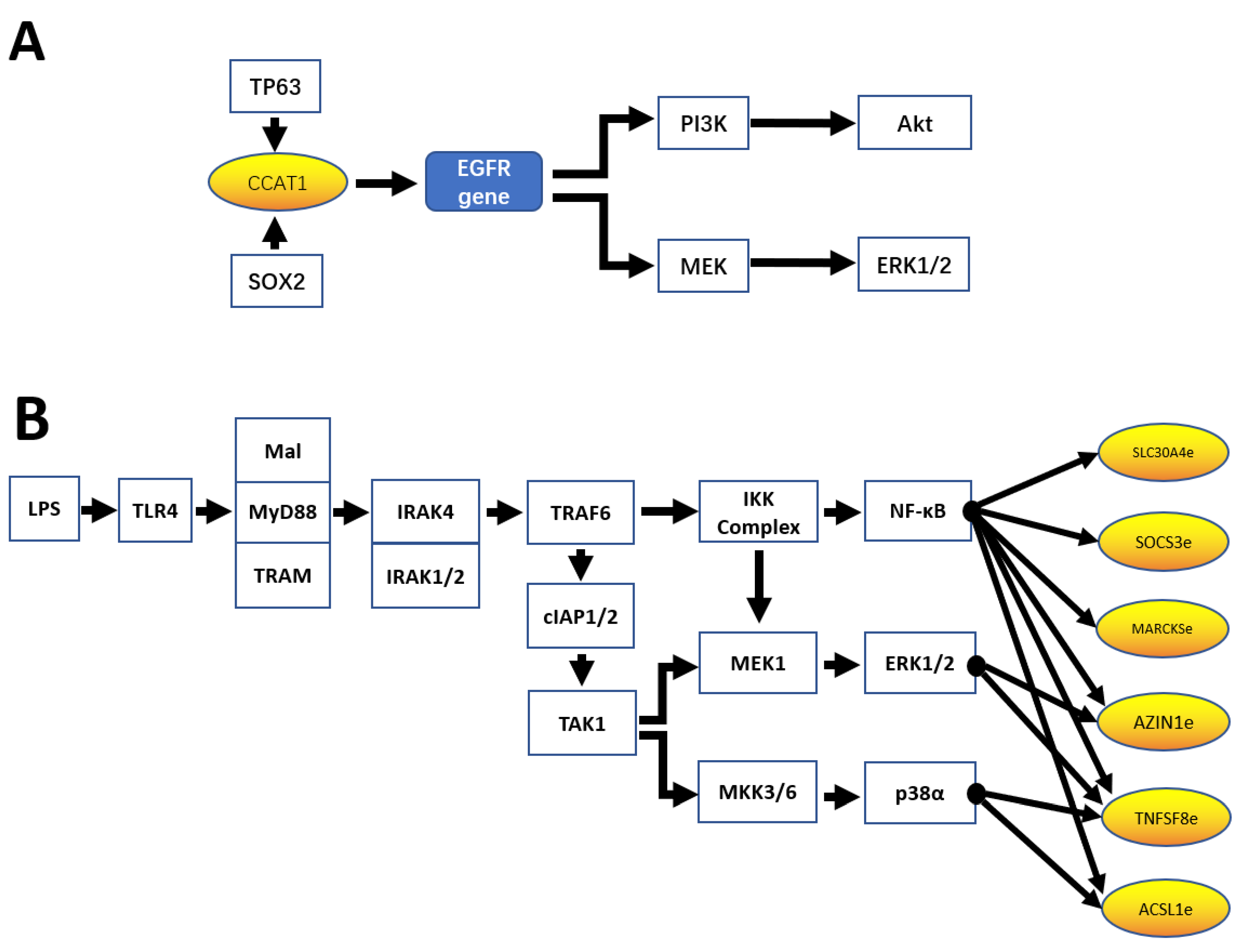
| CCAT1 | colon cancer; squamous cell carcinomas | PI3K/AKTK signaling; MEK/ERK signaling | Jiang et al. [19] McCleland et al. [44] |
| p53BER eRNA | p53-WT renal cancer | p53 signaling | Melo et al. [18] |
| PSA eRNA | prostate cancer | Androgen receptor signaling | Zhao et al. [30] |
| MYC ESE eRNAs | EBV-associated malignancies | MYC signaling | Liang et al. [24] |
| NET1e | breast cancer | / | Zhang et al. [27] |
| SMAD7e | bladder cancer | / | Che et al. [51] |
| P2RY2e | bladder cancer | / | Ding et al. [50] |
| AC003092.1 | glioblastoma multiforme | / | Guo et al. [55] |
| AP001056.1 | squamous cell carcinoma of the head and neck | / | Gu et al. [56] |
| Methods | Description |
|---|---|
| Global Run-On sequencing (GRO-seq) | GRO-seq is able to assess nascent low-abundant RNAs genome-wide independent of the effect of RNA stability. |
| Precision Global Run-On sequencing (PRO-seq) | PRO-seq is based on GRO-seq, and highly sensitive and can identify short unstable nascent RNAs. |
| PRO-cap; 5′GRO-seq | PRO-cap and 5′GRO-seq are developed basing on PRO-seq and GRO-seq, and are able to identify the capped nascent RNA from 5′ end. |
| small interfering RNAs | The simplest method for RNA interference, siRNAs can inactivate their target RNAs in a sequence-specific way. |
| short hairpin RNAs | Another method of RNA interference. shRNAs can be transfected by delivery of plasmid vectors or through virally produced vectors. |
| CRISPR-Cas13a | As a novel type of RNA targeting enzyme, CRISPR-Cas13a is proved to be efficacious for RNA knockdown in mammalian cells. |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wang, R.; Tang, Q. Current Advances on the Important Roles of Enhancer RNAs in Molecular Pathways of Cancer. Int. J. Mol. Sci. 2021, 22, 5640. https://doi.org/10.3390/ijms22115640
Wang R, Tang Q. Current Advances on the Important Roles of Enhancer RNAs in Molecular Pathways of Cancer. International Journal of Molecular Sciences. 2021; 22(11):5640. https://doi.org/10.3390/ijms22115640
Chicago/Turabian StyleWang, Rui, and Qianzi Tang. 2021. "Current Advances on the Important Roles of Enhancer RNAs in Molecular Pathways of Cancer" International Journal of Molecular Sciences 22, no. 11: 5640. https://doi.org/10.3390/ijms22115640
APA StyleWang, R., & Tang, Q. (2021). Current Advances on the Important Roles of Enhancer RNAs in Molecular Pathways of Cancer. International Journal of Molecular Sciences, 22(11), 5640. https://doi.org/10.3390/ijms22115640





