Processed Animal Proteins from Insect and Poultry By-Products in a Fish Meal-Free Diet for Rainbow Trout: Impact on Intestinal Microbiota and Inflammatory Markers
Abstract
1. Introduction
2. Results
2.1. Growth Parameters
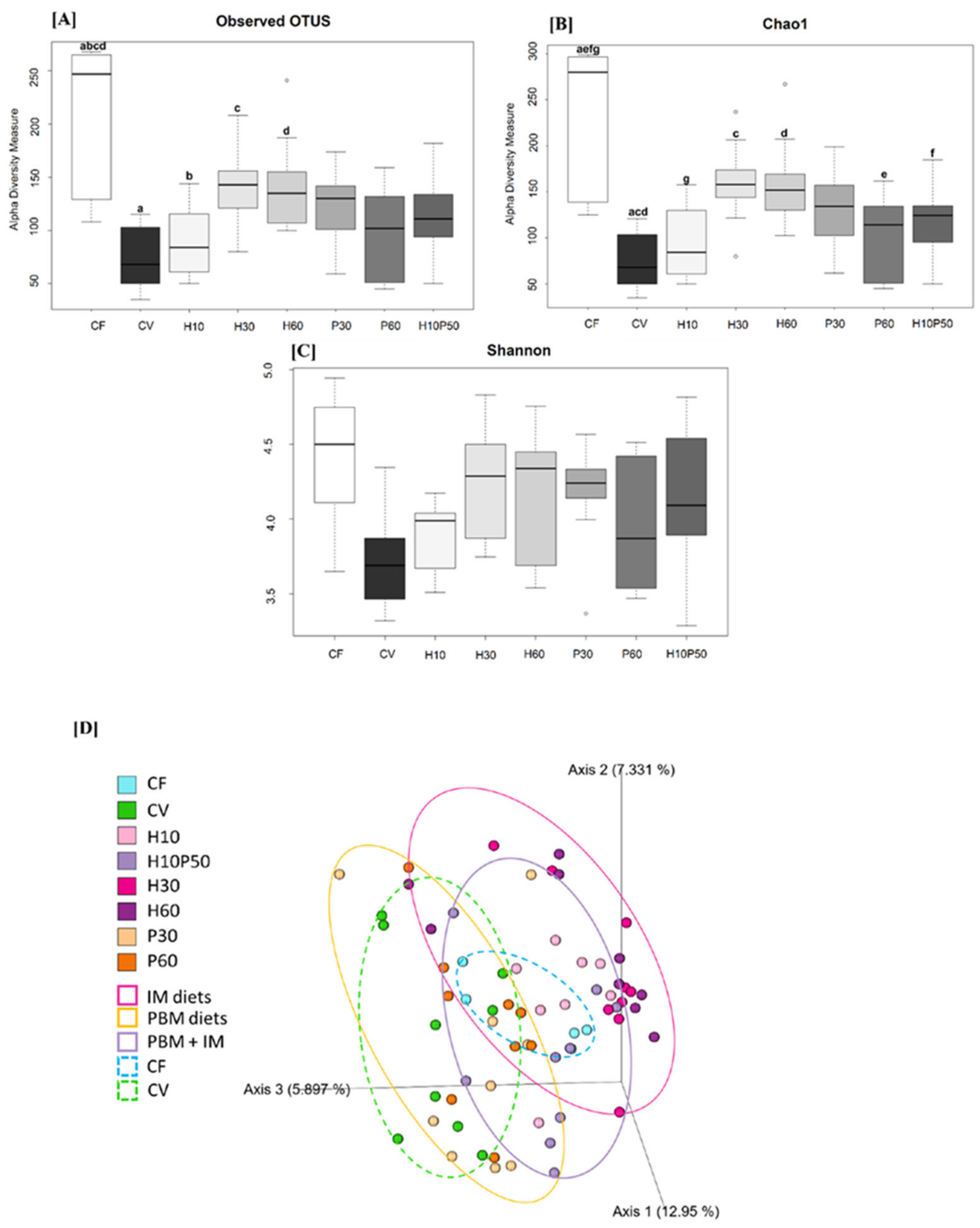
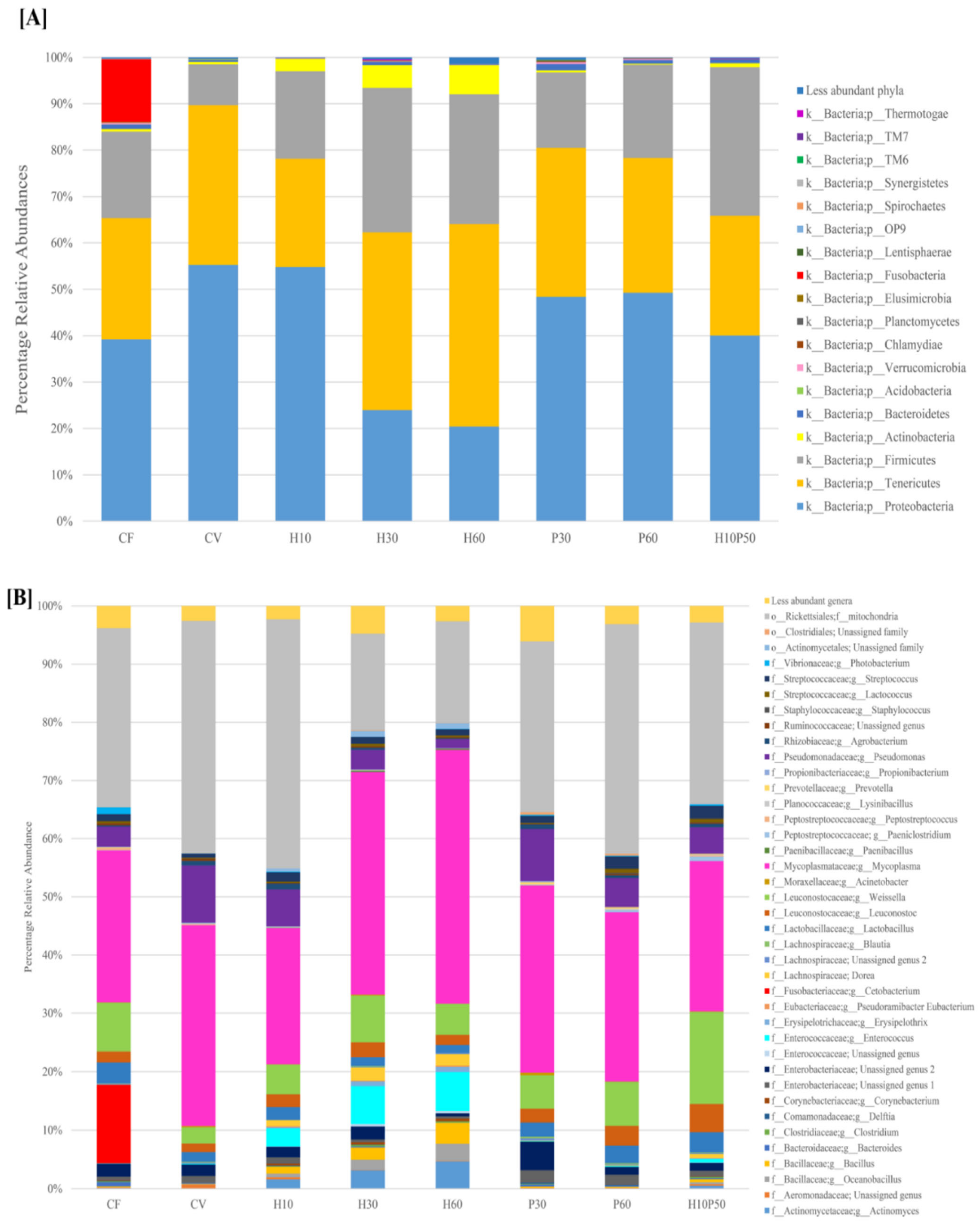
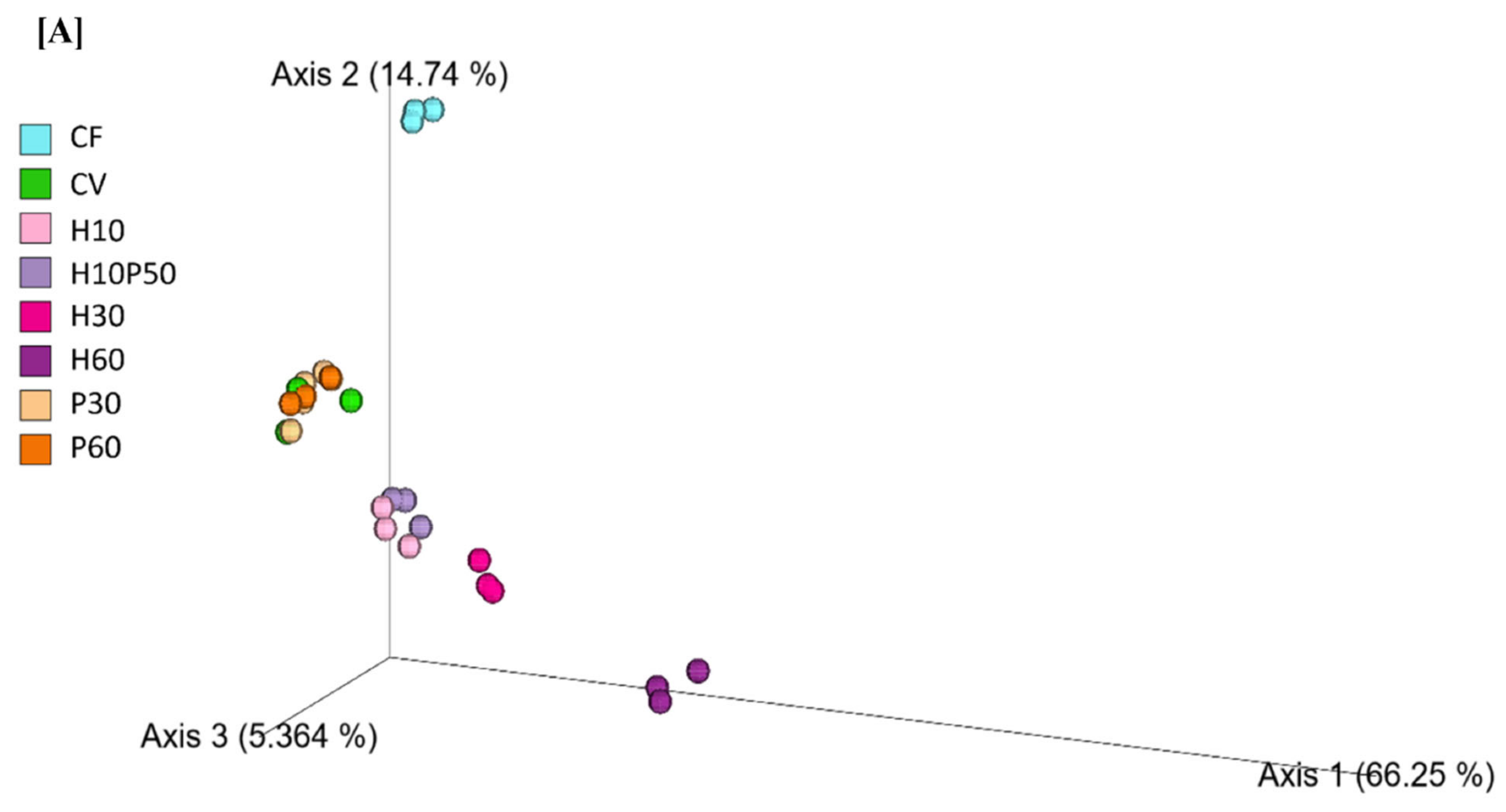
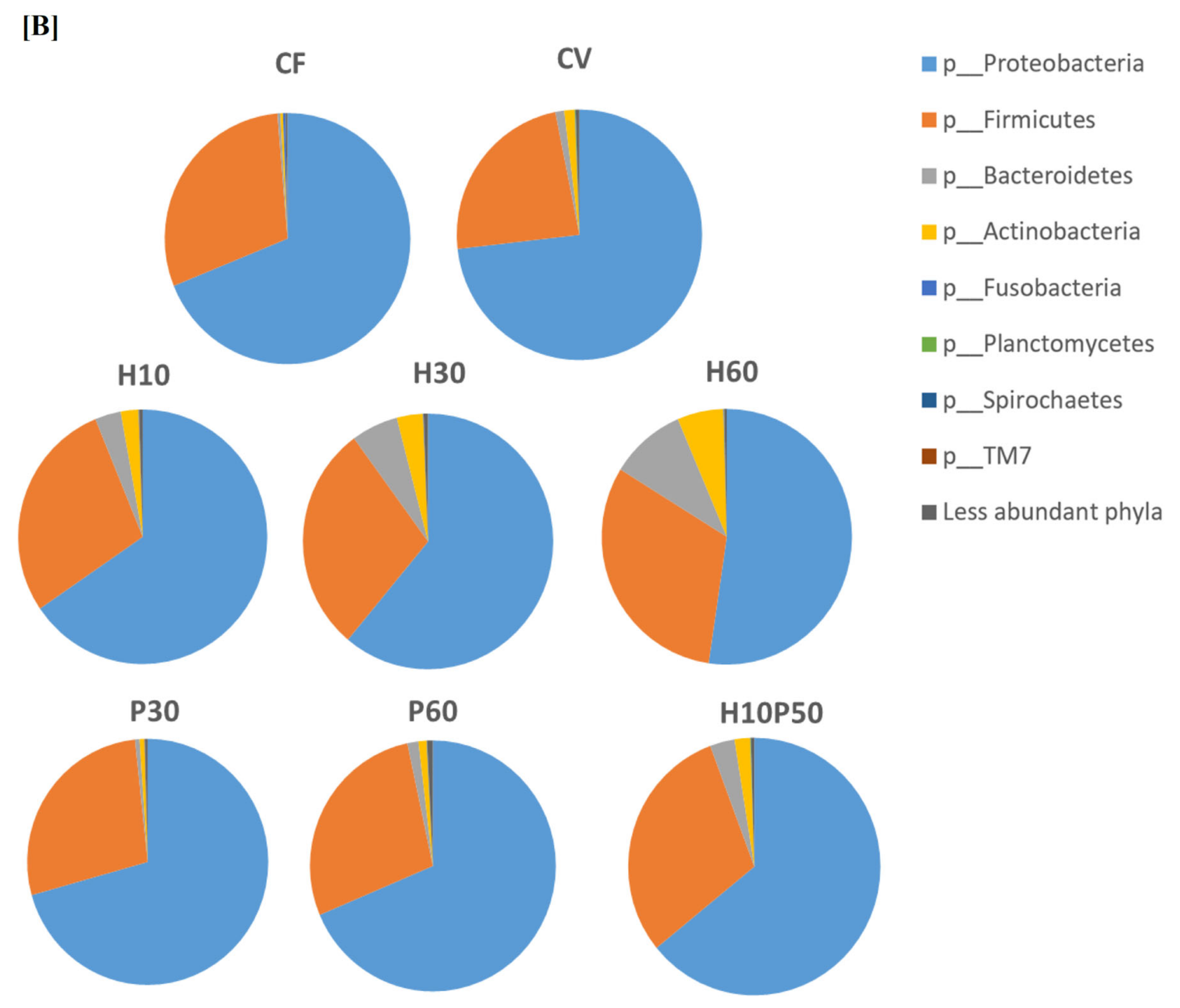
2.2. Gut Microbial Ecology
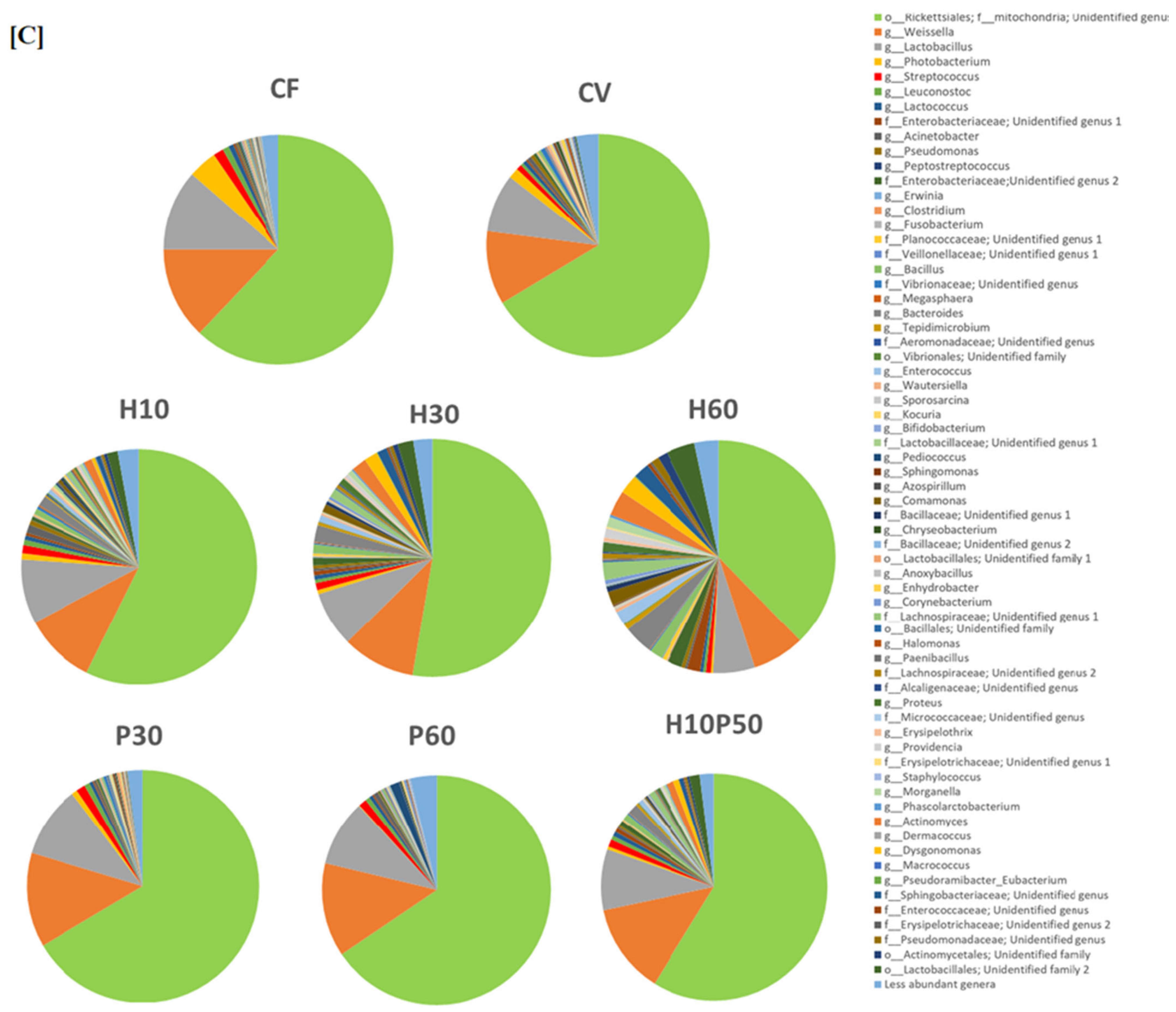
2.3. Microbiota Composition of Fish Feed
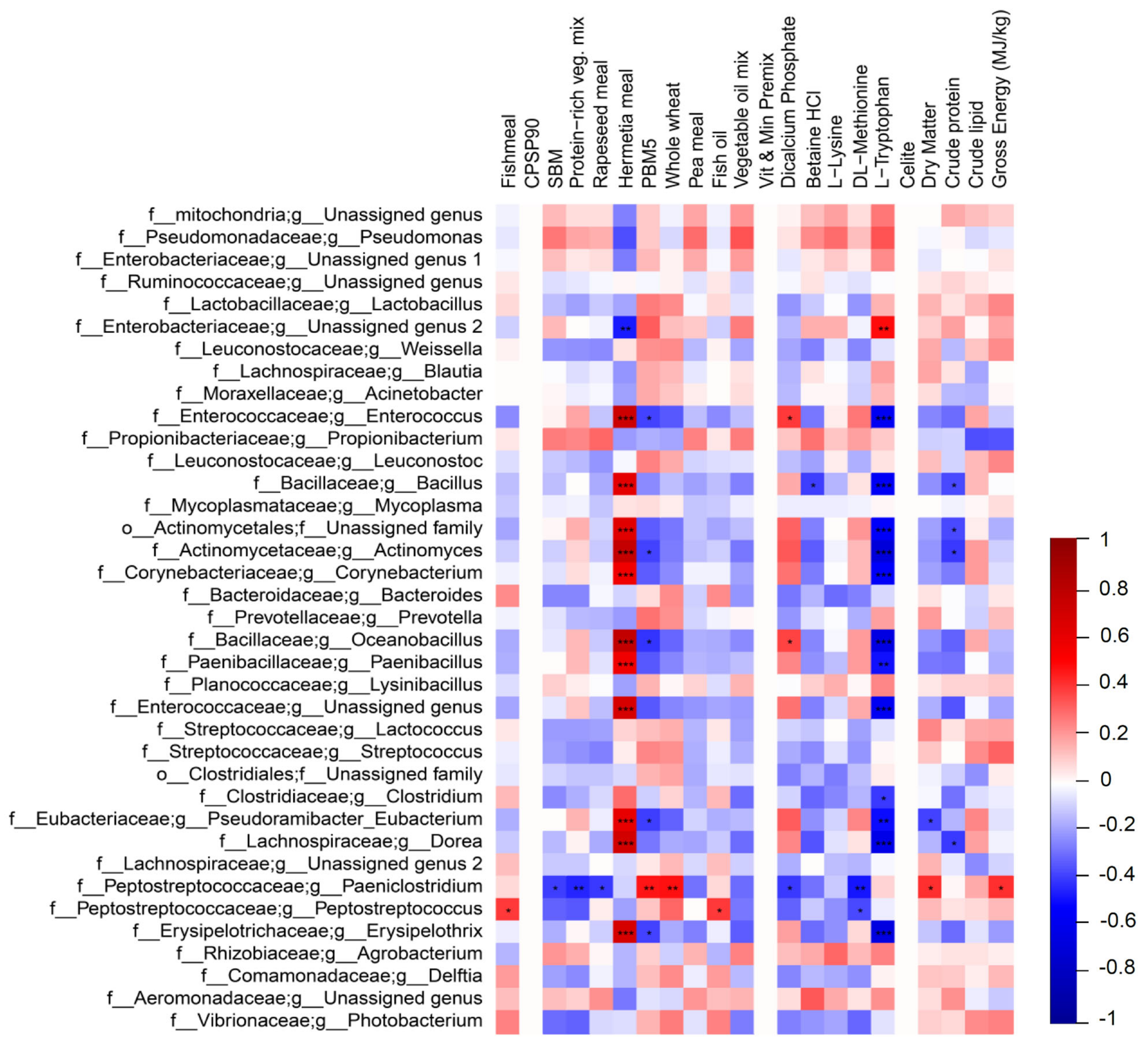
2.4. Correlation Analysis between GM and Ingredients
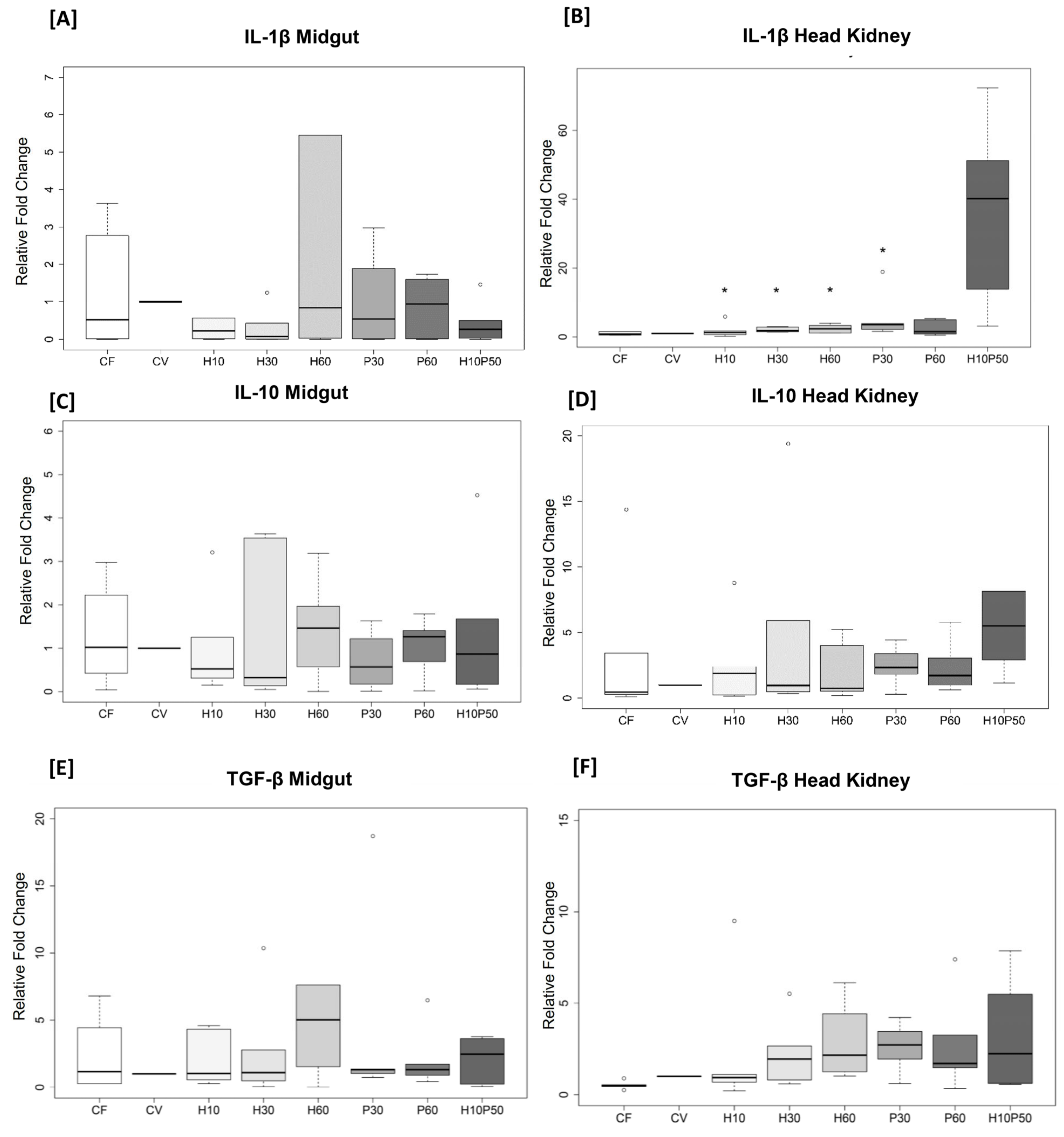
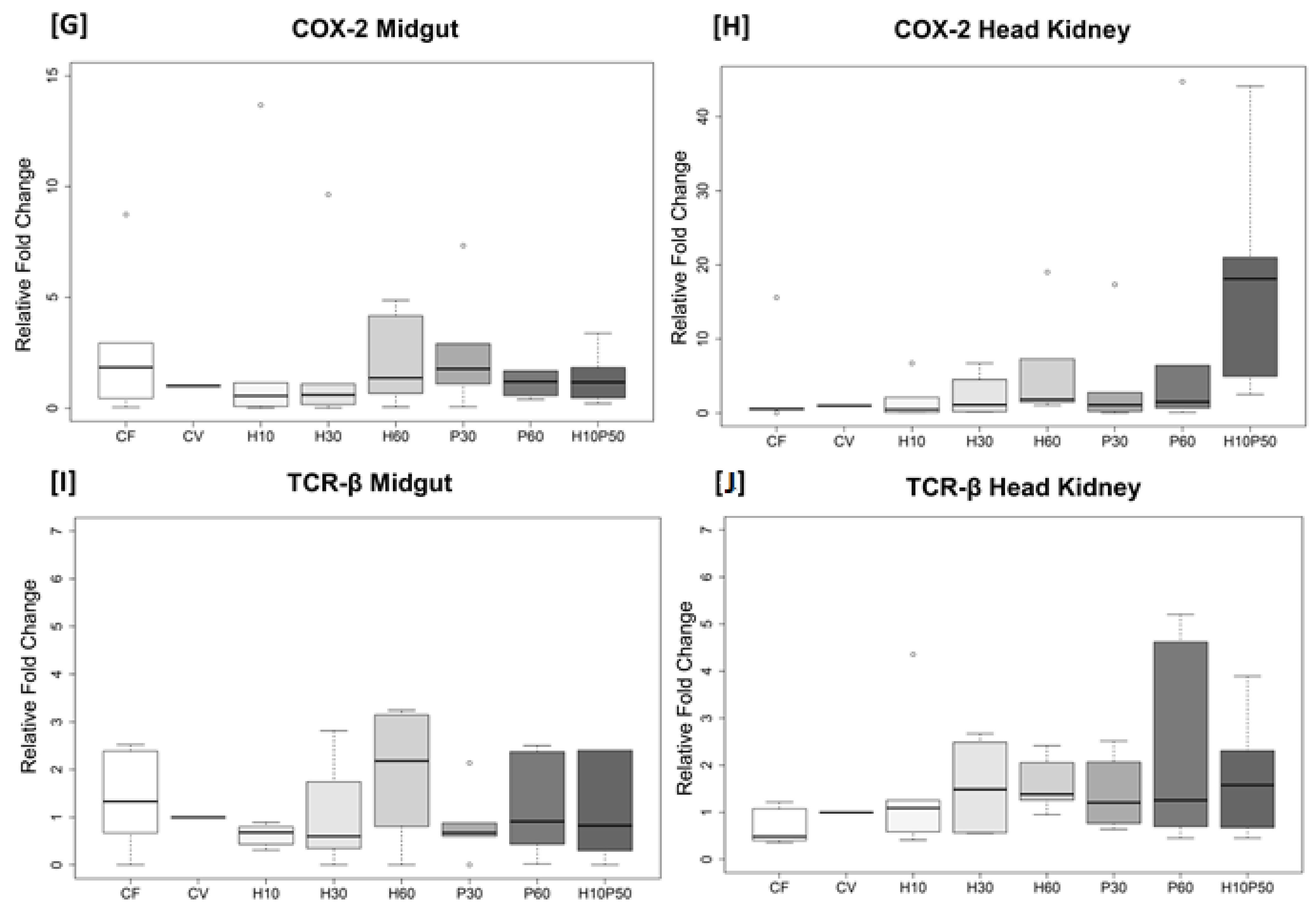
2.5. Gene Expression in the Midgut and Head Kidney
2.6. Plasma LPS as an Intestinal Permeability Marker
3. Discussion
4. Materials and Methods
4.1. Animal Ethics Statement
4.2. Fish, Experimental Diets, and Culture Conditions
4.3. Tissue Sampling and Calculation
4.4. Gut Microbiota Analysis
4.5. Gene Expression in the Midgut and Head Kidney
4.6. Plasma LPS Concentration
4.7. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Willett, W.; Rockström, J.; Loken, B.; Springmann, M.; Lang, T.; Vermeulen, S.; Garnett, T.; Tilman, D.; DeClerck, F.; Wood, A.; et al. Food in the Anthropocene: The EAT—Lancet Commission on healthy diets from sustainable food systems. Lancet 2019, 393, 447–492. [Google Scholar] [CrossRef]
- FAO. The State of World Fisheries and Aquaculture 2020. Sustainability in Action; FAO: Rome, Italy, 2020. [Google Scholar] [CrossRef]
- Den Hartog, L.A.; Sijtsma, S.R. Sustainable feed ingredients. In Proceedings of the 12th International Symposium of Australian Renderers Association “Rendering for Sustainability”, Victoria, Australia, 23–26 July 2013. [Google Scholar]
- Nasopoulou, C.; Zabetakis, I. Benefits of fish oil replacement by plant originated oils in compounded fish feeds. A review. LWT Food Sci. Technol. 2012, 47, 217–224. [Google Scholar] [CrossRef]
- Montero, D.; Izquierdo, M. Welfare and health of fish fed vegetable oils as alternative lipid sources to fish oil. In Fish Oil Replacement and Alternative Lipid Sources in Aquaculture Feeds; CRC Press: Boca Raton, FL, USA, 2010; pp. 439–485. [Google Scholar] [CrossRef]
- Jędrejek, D.; Levic, J.; Wallace, J.; Oleszek, W. Animal by-products for feed: Characteristics, European regulatory framework, and potential impacts on human and animal health and the environment. J. Anim. Feed Sci. 2016, 25, 189–202. [Google Scholar] [CrossRef]
- Hernández, C.; Osuna-Osuna, L.; Benitez-Hernandez, A.; Sanchez-Gutierrez, Y.; González-Rodríguez, B.; Dominguez-Jimenez, P. Replacement of fish meal by poultry by-product meal, food grade, in diets for juvenile spotted rose snapper (Lutjanus guttatus). Lat. Am. J. Aquat. Res. 2014, 42, 111–120. [Google Scholar] [CrossRef]
- Maiolo, S.; Parisi, G.; Biondi, N. Fishmeal partial substitution within aquafeed formulations: Life cycle assessment of four alternative protein sources. Int. J. Life Cycle Assess 2020, 25, 1455–1471. [Google Scholar] [CrossRef]
- Badillo, D.; Herzka, S.Z.; Viana, M.T. Protein Retention Assessment of Four Levels of Poultry By-Product Substitution of Fishmeal in Rainbow Trout (Oncorhynchus mykiss) Diets Using Stable Isotopes of Nitrogen (δ15N) as Natural Tracers. PLoS ONE 2014, 9, e107523. [Google Scholar] [CrossRef]
- Hatlen, B.; Jakobsen, J.V.; Crampton, V.; Alm, M.; Langmyhr, E.; Espe, M.; Hevrøy, E.M.; Torstensen, B.E.; Liland, N.; Waagbø, R. Growth, feed utilization and endocrine responses in Atlantic salmon (Salmo salar) fed diets added poultry by-product meal and blood meal in combination with poultry oil. Aquac. Nutr. 2015, 21, 714–725. [Google Scholar] [CrossRef]
- Doughty, K.H.; Garner, S.R.; Bernards, M.A.; Heath, J.W.; Neff, B.D. Effects of dietary fishmeal substitution with corn gluten meal and poultry meal on growth rate and flesh characteristics of Chinook salmon (Oncorhynchus tshawytscha). Int. Aquat. Res. 2019, 11, 325–334. [Google Scholar] [CrossRef]
- Barreto-Curiel, F.; Parés-Sierra, G.; Correa-Reyes, G.; Durazo-Beltrán, E.; Viana, M.T. Total and partial fishmeal substitution by poultry by-product meal (pet food grade) and enrichment with acid fish silage in aquafeeds for juveniles of rainbow trout Oncorhynchus mykiss. Lat. Am. J. Aquat. Res. 2016, 44, 327–335. [Google Scholar] [CrossRef]
- Nogales-Mérida, S.; Gobbi, P.; Jozefiak, D.; Mazurkiewicz, J.; Dudek, K.; Rawski, M.; Kieronczyk, B.; Jozefiak, A. Insect meal in fish nutrition. Rev. Aquacult. 2018, 11, 1080–1103. [Google Scholar] [CrossRef]
- Bruni, L.; Pastorelli, R.; Viti, C.; Gasco, L.; Parisi, G. Characterisation of the intestinal microbial communities of rainbow trout (Oncorhynchus mykiss) fed with Hermetia illucens (black soldier fly) partially defatted larva meal as partial dietary protein source. Aquaculture 2018, 487, 56–63. [Google Scholar] [CrossRef]
- Rimoldi, S.; Gini, E.; Iannini, F.; Gasco, L.; Terova, G. The effects of dietary insect meal from Hermetia illucens prepupae on autochthonous gut microbiota of rainbow trout (Oncorhynchus mykiss). Animals 2019, 9, 143. [Google Scholar] [CrossRef]
- Terova, G.; Rimoldi, S.; Ascione, C.; Gini, E.; Ceccotti, C.; Gasco, L. Rainbow trout (Oncorhynchus mykiss) gut microbiota is modulated by insect meal from Hermetia illucens prepupae in the diet. Rev. Fish Biol. Fish. 2019, 29, 465–486. [Google Scholar] [CrossRef]
- Mousavi, S.; Zahedinezhad, S.; Loh, J. A review on insect meals in aquaculture: The immunomodulatory and physiological effects. Int. Aquat. Res. 2020, 12, 100–115. [Google Scholar] [CrossRef]
- St-Hilaire, S.; Cranfill, K.; Mcguire, M.A.; Mosley, E.E.; Tomberlin, J.K.; Newton, L.; Sealey, W.; Sheppard, C.; Irving, S. Fish offal recycling by the black soldier fly produces a foodstuff high in omega-3 fatty acids. J. World Aquac. Soc. 2007, 38, 309–313. [Google Scholar] [CrossRef]
- Cebra, J.J. Influences of microbiota on intestinal immune system development. Am. J. Clin. Nutr. 1999, 69, 1046S–1051S. [Google Scholar] [CrossRef]
- Pérez, T.; Balcazar, J.L.; Ruiz-Zarzuela, I.; Halaihel, N.; Vendrell, D.; de Blas, I.; Muzquiz, J.L. Host-microbiota interactions within the fish intestinal ecosystem. Mucosal. Immunol. 2010, 3, 355–360. [Google Scholar] [CrossRef]
- Wang, A.R.; Ran, C.; Ringø, E.; Zhou, Z.G. Progress in fish gastrointestinal microbiota research. Rev. Aquacult. 2018, 10, 626–640. [Google Scholar] [CrossRef]
- Kim, S.; Covington, A.; Pamer, E.G. The intestinal microbiota: Antibiotics, colonization resistance, and enteric pathogens. Immunol. Rev. 2017, 279, 90–105. [Google Scholar] [CrossRef]
- Sakata, T. Microflora in the digestive tract of fish and shellfish, Microbiology in Poecilotherms. Elsevier 1990, 171–176. [Google Scholar]
- Estensoro, I.; Ballester-Lozano, G.; Benedito-Palos, L.; Grammes, F.; Martos-Sitcha, J.A.; Mydland, L.T.; Calduch-Giner, J.A.; Fuentes, J.; Karalazos, V.; Ortiz, A.; et al. Dietary Butyrate Helps to Restore the Intestinal Status of a Marine Teleost (Sparus aurata) Fed Extreme Diets Low in Fish Meal and Fish Oil. PLoS ONE 2016, 11, e0166564. [Google Scholar] [CrossRef]
- Wu, S.; Wang, G.; Angert, E.R.; Wang, W.; Li, W.; Zou, H. Composition, diversity and origin of the bacterial community in grass carp intestine. PLoS ONE 2012, 7, e30440. [Google Scholar] [CrossRef]
- FEAP Report. FEAP European Aquaculture Report 2014–2019; FEAP Secretariat: Brussels, Belgium, 2020.
- Desai, A.R.; Links, M.; Collins, S.A.; Mansfield, G.S.; Drew, M.; Kessel, A.; Hill, J. Effects of plant-based diets on the distal gut microbiome of rainbow trout (Oncorhynchus mykiss). Aquaculture 2012, 350, 134–142. [Google Scholar] [CrossRef]
- Rimoldi, S.; Terova, G.; Ascione, C.; Giannico, R.; Brambilla, F. Next generation sequencing for gut microbiome characterization in rainbow trout (Oncorhynchus mykiss) fed animal by-product meals as an alternative to fishmeal protein sources. PLoS ONE 2018, 13, e0193652. [Google Scholar] [CrossRef]
- Chaklader, M.R.; Siddik, M.A.B.; Fotedar, R.; Howieson, J. Insect larvae, Hermetia illucens in poultry by-product meal for barramundi, Lates calcarifer modulates histomorphology, immunity and resistance to Vibrio harveyi. Sci. Rep. 2019, 9, 16703. [Google Scholar] [CrossRef]
- Randazzo, B.; Zarantoniello, M.; Gioacchini, G.; Cardinaletti, G.; Belloni, A.; Giorgini, E.; Faccenda, F.; Cerri, R.; Tibaldi, E.; Olivotto, I. Physiological response of rainbow trout (Oncorhynchus mykiss) to graded levels of Hermetia illucens or poultry by-product meals as single or combined substitute ingredients to dietary plant proteins. Aquaculture 2021, 538, 736550. [Google Scholar] [CrossRef]
- Gómez, G.D.; Balcázar, J.L. A review on the interactions between gut microbiota and innate immunity of fish. FEMS Immunol Med. Microbiol. 2008, 52, 145–154. [Google Scholar] [CrossRef]
- Ringø, E.; Zhou, Z.; Vecino, J.; Wadsworth, S.; Romero, J.; Krogdahl, Å.; Olsen, R.; Dimitroglou, A.; Foey, A.; Davies, S.; et al. Effect of dietary components on the gut microbiota of aquatic animals. A never-ending story? Aquacult. Nutr. 2016, 22, 219–282. [Google Scholar] [CrossRef]
- Alami-Durante, H.; Médale, F.; Cluzeaud, M.; Kaushik, S.J. Skeletal muscle growth dynamics and expression of related genes in white and red muscles of rainbow trout fed diets with graded levels of a mixture of plant protein sources as substitutes for fishmeal. Aquaculture 2010, 303, 50–58. [Google Scholar] [CrossRef]
- Burr, G.S.; Wolters, W.R.; Barrows, F.T.; Hardy, R.W. Replacing fishmeal with blends of alternative proteins on growth performance of rainbow trout (Oncorhynchus mykiss), and early or late stage juvenile Atlantic salmon (Salmo salar). Aquaculture 2012, 334–337, 110–116. [Google Scholar] [CrossRef]
- Renna, M.; Schiavone, A.; Gai, F.; Dabbou, S.; Lussiana, C.; Malfatto, V.; Prearo, M.; Capucchio, M.T.; Biasato, I.; De Marco, M.; et al. Evaluation of the suitability of a partially defatted black soldier fly (Hermetia illucens L.) larvae meal as ingredient for rainbow trout (Oncorhynchus mykiss Walbaum) diets. J. Anim. Sci. Biotechnol. 2017, 8, 57. [Google Scholar] [CrossRef]
- Muddasir, J.; Imtiaz, A. Length weight relationship and condition factor of snow trout, Schizothorax plagiostomus (Heckel, 1838) from Lidder River, Kashmir. Int. J. Fish Aquat. Stud. 2016, 4, 131–136. [Google Scholar]
- Le Boucher, R.; Quillet, E.; Vandeputte, M.; Lecalvez, J.M.; Goardon, L.; Chatain, B.; Médale, F.; Dupont-Nivet, M. Plant-based diet in rainbow trout (Oncorhynchus mykiss Walbaum): Are there genotype-diet interactions for main production traits when fish are fed marine vs. plant-based diets from the first meal? Aquaculture 2011, 321, 41–48, ISSN 0044-8486. [Google Scholar] [CrossRef]
- Krogdahl, Å.; Penn, M.; Thorsen, J.; Refstie, S.; Bakke, A.M. Important antinutrients in plant feedstuffs for aquaculture: An update on recent findings regarding responses in salmonids. Aquac. Res. 2010, 41, 333–344. [Google Scholar] [CrossRef]
- Hu, H.; Kortner, T.M.; Gajardo, K.; Chikwati, E.; Tinsley, J. Intestinal Fluid Permeability in Atlantic Salmon (Salmo salar L.) Is Affected by Dietary Protein Source. PLoS ONE 2016, 11, e0167515. [Google Scholar] [CrossRef]
- Rimoldi, S.; Antonini, M.; Gasco, L.; Moroni, F.; Terova, G. Intestinal microbial communities of rainbow trout (Oncorhynchus mykiss) may be improved by feeding a Hermetia illucens meal/low-fishmeal diet. Fish Physiol. Biochem. 2021. [Google Scholar] [CrossRef]
- Parshukov, A.N.; Kashinskaya, E.N.; Simonov, E.P.; Hlunov, O.V.; Izvekova, G.I.; Andree, K.B.; Solovyev, M.M. Variations of the intestinal gut microbiota of farmed rainbow trout, Oncorhynchus mykiss (Walbaum), depending on the infection status of the fish. J. Appl. Microbiol. 2019, 127, 379–395. [Google Scholar] [CrossRef]
- Li, Y.; Bruni, L.; Jaramillo-Torres, A.; Gajardo, K.; Kortner, T.M.; Krogdahl, Å. Differential Response of Digesta- and Mucosa-Associated Intestinal Microbiota to Dietary Black Soldier Fly (Hermetia illucens) Larvae Meal in Seawater Phase Atlantic Salmon (Salmo salar). Anim. Microbiome 2021, 3, 8. [Google Scholar] [CrossRef]
- Beier, S.; Bertilsson, S. Bacterial chitin degradation-mechanisms and ecophysiological strategies. Front. Microbiol. 2013, 4, 149. [Google Scholar] [CrossRef] [PubMed]
- Ng, W.K.; Koh, C.B. The utilization and mode of action of organic acids in the feeds of cultured aquatic animals. Rev Aquacult. 2017, 9, 342–368. [Google Scholar] [CrossRef]
- Oushani, A.K.; Soltani, M.; Sheikhzadeh, N.; Mehrgan, M.S.; Islami, H.R. Effects of dietary chitosan and nano-chitosan loaded clinoptilolite on growth and immune responses of rainbow trout (Oncorhynchus mykiss). Fish Shellfish. Immunol. 2020, 98, 210–217, ISSN 1050-4648. [Google Scholar] [CrossRef]
- Nawaz, A.; Javaid, A.B.; Irshad, S.; Hoseinifar, S.H.; Xionga, H. The functionality of prebiotics as immunostimulant: Evidences from trials on terrestrial and aquatic animals. Fish Shellfish. Immunol. 2018, 76, 272–278. [Google Scholar] [CrossRef]
- Zhou, Z.; Karlsen, Ø.; He, S.; Olsen, R.E.; Yao, B.; Ringø, E. The effect of dietary chitin on the autochthonous gut bacteria of Atlantic cod (Gadus morhua L.). Aquac. Res. 2013, 44, 1889–1900. [Google Scholar] [CrossRef]
- Esteban, M.A.; Cuesta, A.; Ortuño, J.; Meseguer, J. Immunomodulatory effects of dietary intake of chitin on gilthead seabream (Sparus aurata L.) innate immune system. Fish Shellfish. Immunol. 2001, 11, 303–315. [Google Scholar] [CrossRef] [PubMed]
- Casadidio, C.; Peregrina, D.V.; Gigliobianco, M.R.; Deng, S.; Censi, R.; Di Martino, P. Chitin and Chitosans: Characteristics, Eco-Friendly Processes, and Applications in Cosmetic Science. Mar. Drugs 2019, 17, 369. [Google Scholar] [CrossRef]
- Gonçalves, A.T.; Gallardo-Escarate, C. Microbiome dynamic modulation through functional diets based on pre- and probiotics (mannan-oligosaccharides and Saccharomyces cerevisiae) in juvenile rainbow trout (Oncorhynchus mykiss). J. Appl. Microbiol. 2017, 122, 1333–1347. [Google Scholar] [CrossRef]
- Lyons, P.P.; Turnbull, J.F.; Dawson, K.A.; Crumlish, M. Effects of low-level dietary microalgae supplementation on the distal intestinal microbiome of farmed rainbow trout Oncorhynchus mykiss (Walbaum). Aquac. Res. 2017, 48, 2438–2452. [Google Scholar] [CrossRef]
- Pollack, J.D.; Williams, M.V.; McElhaney, R.N. The comparative metabolism of the mollicutes (Mycoplasmas): The utility for taxonomic classification and the relationship of putative gene annotation and phylogeny to enzymatic function in the smallest free-living cells. Crit. Rev. Microbiol. 1997, 23, 269–354. [Google Scholar] [CrossRef]
- Brown, R.M.; Wiens, G.D.; Salinas, I. Analysis of the gut and gill microbiome of resistant and susceptible lines of rainbow trout (Oncorhynchus mykiss). Fish Shellfish. Immunol. 2019, 86, 497–506, ISSN 1050-4648. [Google Scholar] [CrossRef]
- Lacombe-Harvey, M.È.; Brzezinski, R.; Beaulieu, C. Chitinolytic functions in actinobacteria: Ecology, enzymes, and evolution. Appl. Microbiol. Biotechnol. 2018, 102, 7219–7230. [Google Scholar] [CrossRef]
- Kim, D.H.; Brunt, J.; Austin, B. Microbial diversity of intestinal contents and mucus in rainbow trout (Oncorhynchus mykiss). J. Appl. Microbiol. 2007, 102, 1654–1664. [Google Scholar] [CrossRef]
- Tarnecki, A.M.; Wafapoor, M.; Phillips, R.N. Benefits of a Bacillus probiotic to larval fish survival and transport stress resistance. Sci. Rep. 2019, 9, 4892. [Google Scholar] [CrossRef]
- Veliz, E.A.; Martínez-Hidalgo, P.; Hirsch, A.M. Chitinase-producing bacteria and their role in biocontrol. AIMS microbiology 2017, 3, 689–705. [Google Scholar] [CrossRef]
- Parris, D.J.; Morgan, M.M.; Stewart, F.J. Feeding rapidly alters microbiome composition and gene transcription in the clownfish gut. Appl. Environ. Microbiol. 2019, 23, 85. [Google Scholar] [CrossRef]
- Jin, X.; Chen, Z.; Shi, Y.; Gui, J.F.; Zhao, Z. Response of gut microbiota to feed-borne bacteria depends on fish growth rate: A snapshot survey of farmed juvenile Takifugu Obscurus. Microb. Biotechnol. 2021. [CrossRef] [PubMed]
- Pomaranski, E.K.; Griffin, M.J.; Camus, A.C.; Armwood, A.R.; Shelley, J.; Waldbieser, G.C.; LaFrentz, B.R.; García, J.C.; Yanong, R.; Soto, E. Description of Erysipelothrix piscisicarius sp. nov., an emergent fish pathogen, and assessment of virulence using a tiger barb (Puntigrus tetrazona) infection model. Int. J. Syst. Evol. Microbiol. 2020, 70, 857–867. [Google Scholar] [CrossRef]
- Hartviksen, M.; Vecino, J.L.G.; Ringo, E.; Bakke, A.M.; Wadsworth, S.; Krogdahl, A.; Ruohonen, K.; Kettunen, A. Alternative dietary protein sources for Atlantic salmon (Salmo salar L.) effect on intestinal microbiota, intestinal and liver histology and growth. Aquac. Nutr. 2014, 20, 381–398. [Google Scholar] [CrossRef]
- Amaretti, A.; Gozzoli, C.; Simone, M.; Raimondi, S.; Righini, L.; Pérez-Brocal, V.; García-López, R.; Moya, A.; Rossi, M. Profiling of Protein Degraders in Cultures of Human Gut Microbiota. Front Microbiol. 2019, 10, 2614. [Google Scholar] [CrossRef]
- Chapagain, P.; Arivett, B.; Cleveland, B.M.; Walker, D.M.; Salem, M. Analysis of the fecal microbiota of fast- and slow-growing rainbow trout (Oncorhynchus mykiss). BMC Genom. 2019, 20, 788. [Google Scholar] [CrossRef]
- Sigh, J.; Lindenstrøm, T.; Buchmann, K. Expression of pro-inflammatory cytokines in rainbow trout (Oncorhynchus mykiss) during an infection with Ichthyophthirius multifiliis. Fish Shellfish. Immunol. 2004, 17, 75–86. [Google Scholar] [CrossRef] [PubMed]
- Secombes, C.J.; Wang, T.; Hong, S.; Peddie, S.; Crampe, M.; Laing, K. Cytokines and innate immunity of fish. Dev. Comp. Immunol. 2001, 25, 713–723. [Google Scholar] [CrossRef]
- Low, C.; Wadsworth, S.; Burrells, C.; Secombes, C.J. Expression of immune genes in turbot (Scophthalmus maximus) fed a nucleotide-supplemented diet. Aquaculture 2003, 221, 23–40. [Google Scholar] [CrossRef]
- Wang, X.; Lupardus, P.; Laporte, S.L.; Garcia, K.C. Structural biology of shared cytokine receptors. Annu. Rev. Immunol. 2009, 27, 29–60. [Google Scholar] [CrossRef]
- Harun, N.O.; Costa, M.M.; Secombes, C.J.; Wang, T. Sequencing of a second interleukin-10 gene in rainbow trout Oncorhynchus mykiss and comparative investigation of the expression and modulation of the paralogues in vitro and in vivo. Fish Shellfish. Immunol. 2011, 31, 107–117, ISSN 1050-4648. [Google Scholar] [CrossRef]
- Heinecke, R.D.; Buchmann, K. Inflammatory response of rainbow trout Oncorhynchus mykiss (Walbaum, 1792) larvae against Ichthyophthirius multifiliis. Fish Shellfish. Immunol. 2013, 34, 521–528. [Google Scholar] [CrossRef]
- Ghosh, S.S.; Wang, J.; Yannie, P.J.; Ghosh, S. Intestinal Barrier Dysfunction, LPS Translocation, and Disease Development. J. Endocr. Soc. 2020, 4, bvz039. [Google Scholar] [CrossRef]
- Klindworth, A.; Pruesse, E.; Schweer, T.; Peplies, J.; Quast, C.; Horn, M.; Glockner, F.O. Evaluation of general 16S ribosomal RNA gene PCR primers for classical and next-generation sequencing-based diversity studies. Nucleic Acids Res. 2013, 41, e1. [Google Scholar] [CrossRef]
- Bolyen, E.; Rideout, J.R.; Dillon, M.R.; Bokulich, N.A.; Abnet, C.C.; Al-Ghalith, G.A.; Alexander, H.; Alm, E.J.; Arumugam, M.; Asnicar, F.; et al. Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat. Biotechnol. 2019, 37, 852–857. [Google Scholar] [CrossRef]
- De Santis, T.Z.; Hugenholtz, P.; Larsen, N.; Rojas, M.; Brodie, E.L.; Keller, K.; Huber, T.; Dalevi, D.; Hu, P.; Andersen, G.L. Greengenes, a Chimera-Checked 16S rRNA Gene Database and Workbench Compatible with ARB. Appl. Environ. Microbiol. 2006, 72, 5069–5072. [Google Scholar] [CrossRef]
- McMurdie, P.J.; Holmes, S. phyloseq: An R Package for Reproducible Interactive Analysis and Graphics of Microbiome Census Data. PLoS ONE 2013, 8, e61217. [Google Scholar] [CrossRef]
- Tacchi, L.; Lowrey, L.; Musharrafieh, R.; Crossey, K.; Larragoite, E.T.; Salinas, I. Effects of transportation stress and addition of salt to transport water on the skin mucosal homeostasis of rainbow trout (Oncorhynchus mykiss). Aquaculture 2015, 435, 120–127, ISSN 0044-8486. [Google Scholar] [CrossRef]
- Inoue, Y.; Kamota, S.; Ito, K.; Yoshiura, Y.; Ototake, M.; Moritomo, T.; Nakanishi, T. Molecular cloning and expression analysis of rainbow trout (Oncorhynchus mykiss) interleukin-10 cDNAs. Fish Shellfish. Immunol. 2005, 18, 335–344. [Google Scholar] [CrossRef]
- Zou, J.; Neuman, N.; Holland, J.; Belosevic, M.; Cunningham, C.; Secombes, C.J.; Rowley, A.F. Fish macrophages express a cyclooxygenase-2 homologue following activation. Biochem. J. 1999, 340, 153–159. [Google Scholar] [CrossRef] [PubMed]
- Kim, D.H.; Austin, B. Cytokine expression in leukocytes and gut cells of rainbow trout, Oncorhynchus mykiss Walbaum, induced by probiotics. Vet. Immunol. Immunopathol. 2006, 114, 297–304. [Google Scholar] [CrossRef]
- Henriksen, M.M.M.; Kania, P.W.; Buchmann, K.; Dalsgaard, I. Effect of hydrogen peroxide and/or Flavobacterium psychrophilum on the gills of rainbow trout, Oncorhynchus mykiss (Walbaum). J. Fish Dis. 2014, 38, 259–270. [Google Scholar] [CrossRef]
- Zhang, Z.; Swain, T.; Bøgwald, J.; Dalmo, R.A.; Kumari, J. Bath immunostimulation of rainbow trout (Oncorhynchus mykiss) fry induces enhancement of inflammatory cytokine transcripts, while repeated bath induce no changes. Fish Shellfish. Immunol. 2009, 26, 677–684. [Google Scholar] [CrossRef]
- Oksanen, J.; Blanchet, F.G.; Friendly, M.; Kindt, R.; Legendre, P.; McGlinn, D.; Minchin, P.R.; O’Hara, R.B.; Simpson, G.L.; Solymos, P.; et al. Vegan: Community Ecology Package. R package version 2.5–2.6. 2019. Available online: https://CRAN.R-project.org/package=vegan (accessed on 13 July 2020).
| Diet | Final Weight (g) | Total Length (cm) | K (g/cm3) |
|---|---|---|---|
| CF | 231.18 ± 32.11 b | 25.53 ± 1.35 d | 1.39 ± 0.11 cd |
| CV | 227.92 ± 35.22 b | 25.01 ± 1.30 e | 1.45 ± 0.11 a |
| H10 | 235.02 ± 30.41 ab | 25.46 ±1.24 d | 1.42 ± 0.11 abc |
| H30 | 239.1 ± 36.26 ab | 25.48 ± 1.42 d | 1.44 ± 0.11 ab |
| H60 | 241.07 ± 35.58 ab | 25.78 ± 1.31 c | 1.40 ± 0.10 bcd |
| P30 | 240.03 ± 37.82 ab | 25.81 ± 1.45 bc | 1.39 ± 0.10 cd |
| P60 | 244.06 ± 36.02 ab | 26.02 ± 1.25 ab | 1.38 ± 0.09 d |
| H10P50 | 254.81 ± 36.82 a | 26.13 ± 1.14 a | 1.42 ± 0.12 abc |
| [A] | Diet | |||||||
| CF | CV | H10 | H30 | H60 | P30 | P60 | H10P50 | |
| Phylum | ||||||||
| Actinobacteria | 0.51 ± 0.67 | 0.52 ± 0.46 | 2.62 ± 4.26 | 4.89 ± 4.23 | 6.29 ± 5.49 a | 0.48 ± 0.53 | 0.23 ± 0.32 a | 0.87 ± 0.65 |
| Bacteroidetes | 1.00 ± 1.22 | 0.41 ± 0.69 | 0.03 ± 0.06 | 0.72 ± 1.11 | 0.09 ± 0.09 a | 1.35 ± 1.26 a | 0.75 ± 0.82 | 0.77 ± 0.91 |
| Fusobacteria | 13.50 ± 29.88 a–g | <0.0001a | <0.0001 b | <0.0001c | <0.0001 d | <0.0001 e | <0.0001 f | 0.002 ± 0.01 g |
| Class | ||||||||
| Erysipelotrichi | 0.04 ± 0.06 | 0.002 ± 0.01 ab | 0.34 ± 0.37 | 0.91 ± 0.80 a | 1.11 ± 1.07 b | 0.58 ± 0.17 | 0.04 ± 0.10 | 0.18 ± 0.31 |
| Fusobacteriia | 13.50 ± 29.88 a | <0.0001 | <0.0001 a | <0.0001 | <0.0001 | <0.0001 | <0.0001 | 0.002 ± 0.01 |
| Gammaproteobacteria | 7.72 ± 7.05 | 14.02 ± 14.53a | 10.76 ± 10.82 | 6.47 ± 6.36 | 2.61 ± 2.74 a | 17.20 ± 15.41 | 8.52 ± 5.00 | 7.59 ± 6.12 |
| Order | ||||||||
| Actinomycetales | 0.49 ± 0.66 | 0.31 ± 0.29 | 2.50 ± 4.29 a | 4.83 ± 4.14 b | 6.25 ± 5.46 | 0.45 ± 0.52 | 0.20 ± 0.33 ab | 0.85 ± 0.65 |
| Bacteroidales | 1.00 ± 1.22 | 0.25 ± 0.55 | 0.03 ± 0.06 | 0.69 ± 1.13 | 0.04 ± 0.05 a | 1.14 ± 1.05 a | 0.69 ± 0.83 | 0.47 ± 0.52 |
| Erysipelotrichales | 0.04 ± 0.06 | 0.002 ± 0.001 abc | 0.34 ± 0.37 a | 0.91 ± 0.80 bde | 1.11 ± 1.07 cf | 0.06 ± 0.17 df | 0.04 ± 0.10 e | 0.18 ± 0.31 |
| Fusobacteriales | 13.50 ± 29.88 abc | <0.0001 | <0.0001 a | <0.0001 b | <0.0001 | <0.0001 c | <0.0001 d | 0.002 ± 0.01 |
| Pseudomonadales | 3.61 ± 2.94 | 10.08 ± 11.47a | 7.17 ± 7.51 | 3.56 ± 3.40 | 1.54 ± 1.88 a | 9.40 ± 8.19 | 5.06 ± 3.65 | 4.72 ± 3.73 |
| Family | ||||||||
| Actinomycetaceae | 0.10 ± 0.19 | 0.05 ± 0.16abcd | 1.57 ± 2.70 aef | 3.16 ± 2.86 bgh | 4.66 ± 3.93 cil | 0.02 ± 0.06 egim | <0.0001 fhln | 0.52 ± 0.40 dmn |
| Corynebacteriaceae | 0.10 ± 0.20 | 0.002 ± 0.007 | 0.32 ± 0.40 a | 0.41 ± 0.53 | 0.47 ± 0.63 | <0.0001 a | 0.004 ± 0.01 | 0.09 ± 0.12 |
| Bacillaceae | 0.31 ± 0.33 | 0.35 ± 0.70 | 1.79 ± 2.73 a | 3.83 ± 3.28 | 6.59 ± 5.60 | 0.30 ± 0.35 | 0.18 ± 0.26 a | 0.94 ± 0.74 |
| Paenibacillaceae | <0.0001 | <0.0001 a | 0.06 ± 0.11 | 0.21 ± 0.26 | 0.20 ± 0.28 a | 0.02 ± 0.07 | <0.0001 | 0.02 ± 0.06 |
| Clostridiaceae | 0.16 ± 0.18 | <0.0001 a | 0.09 ± 0.23 | 0.31 ± 0.29 | 0.24 ± 0.28 a | 0.22 ± 0.42 | 0.26 ± 0.49 | 0.34 ± 0.62 |
| Eubacteriaceae | <0.0001 | <0.0001 a | 0.16 ± 0.31 | 0.12 ± 0.18 bc | 0.19 ± 0.22 a | <0.0001 b | <0.0001 c | 0.02 ± 0.05 |
| Erysipelotrichaceae | 0.04 ± 0.06 | 0.002 ± 0.01 ab | 0.34 ± 0.37 a | 0.91 ± 0.81 cd | 1.11 ± 1.07 b | 0.06 ± 0.17 c | 0.04 ± 0.10 d | 0.18 ± 0.31 |
| Fusobacteriaceae | 13.49 ± 29.88 a | <0.0001 | <0.0001 a | <0.0001 | <0.0001 | <0.0001 | <0.0001 | <0.0001 |
| [B] | Diet | |||||||
| CF | CV | H10 | H30 | H60 | P30 | P60 | H10P50 | |
| Genus | ||||||||
| Actinomyces | 0.10 ± 0.18 | 0.05 ± 0.16 abcd | 1.58 ± 2.70 aef | 3.16 ± 2.86 bgh | 4.66 ± 3.98cil | 0.02 ± 0.06 egi | <0.0001 fhlm | 0.52 ± 0.39 dm |
| Oceanobacillus | 0.06 ± 0.14 | 0.04 ± 0.11 ab | 0.63 ± 1.03 cd | 1.75 ± 1.54 aef | 2.94 ± 2.58 bgh | <0.0001 ceg | <0.0001 dfh | 0.35 ± 0.37 |
| Bacillus | 0.16 ± 0.14 | 0.07 ± 0.14 a | 1.16 ± 1.75 | 2.08 ± 1.96 | 3.60 ± 3.09 a | 0.25 ± 0.27 | 0.18 ± 0.26 | 0.51 ± 0.44 |
| Pseudoramibacter Eubacterium | <0.0001 | <0.0001 a | 0.16 ± 0.31 | 0.12 ± 0.18 | 0.19 ± 0.22 bc | <0.0001b | <0.0001 ac | 0.02 ± 0.05 |
| Dorea | 0.03 ± 0.04 | <0.0001 abcd | 1.05 ± 1.72 ae | 2.26 ± 1.89 bf | 1.99 ± 1.71 cg | 0.08 ± 0.25 | 0.09 ± 0.26 efgh | 0.71 ± 0.79 dh |
| Paeniclostridium | 0.05 ± 0.05 | <0.0001 a | 0.05 ± 0.13 | 0.04 ± 0.08 | 0.02 ± 0.07 | 0.05 ± 0.10 | 0.36 ± 0.39 | 0.77 ± 1.35 a |
| Erysipelothrix | 0.04 ± 0.06 | <0.0001 ab | 0.21 ± 0.41 | 0.80 ± 0.63 acd | 0.89 ± 0.79 bef | <0.0001 ce | 0.01 ± 0.03 df | 0.13 ± 0.30 |
| Enterococcus | 0.06 ± 0.09 | 0.15 ± 0.27 ab | 3.09 ± 4.70 bfg | 6.48 ± 5.12 cdf | 6.68 ± 5.45 eg | 0.10 ± 0.21 c | 0.17 ± 0.20 ade | 0.62 ± 0.81 |
| Cetobacterium | 13.44 ± 29.91 a | <0.0001 | <0.0001 a | <0.0001 | <0.0001 | <0.0001 | <0.0001 | <0.0001 |
| Pseudomonas | 3.48 ± 2.85 | 9.80 ± 11.06 a | 6.36 ± 6.69 | 3.38 ± 3.10 | 1.53 ± 1.89 a | 8.94 ± 7.85 | 4.99 ± 3.67 | 4.56 ± 3.65 |
| Diet | LPS Concentration (ng/μL) |
|---|---|
| CF | 240.13 ± 150.13 |
| CV | 130.22 ± 111.60 |
| H10 | 160.23 ± 163.10 |
| H30 | 117.72 ± 175.90 |
| H60 | 297.22 ± 347.59 |
| P30 | 139.84 ± 130.44 |
| P60 | 176.33 ± 215.96 |
| H10P50 | 210.21 ± 178.07 |
| Gene | Accession Number | Forward Primer (5′-3′) | Reverse Primer (5′-3′) | References |
|---|---|---|---|---|
| IL-1β | AJ557021 | ACATTGCCAACCTCATCATCG | TTGAGCAGGTCCTTGTCCTTG | [75] |
| IL-10 | NM001246350 | CGACTTTAAATCTCCCATCGAC | GCATTGGACGATCTCTTTCTT | [76] |
| COX-2 | AJ238307 | ATCCTTACTCACTACAAAGG | GCTGGTCCTTTCATGAAGTCTG | [77] |
| TGF-β | X99303 | AGATAAATCGGAGAGTTGCTGTG | CCTGCTCCACCTTGTGTTGT | [78] |
| TCR-β | AF329700 | TCACCAGCAGACTGAGAGTCC | AAGCTGACAATGCAGGTGAATC | [79] |
| EF-1α | AF498320 | ACCCTCCTCTTGGTCGTTTC | TGATGACACCAACAGCAACA | [79] |
| 18S | AF308735 | GATCCATTGGAGGGCAAGTCT | CGAGCTTTTTAACTGCAGCAACTTT | [80] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gaudioso, G.; Marzorati, G.; Faccenda, F.; Weil, T.; Lunelli, F.; Cardinaletti, G.; Marino, G.; Olivotto, I.; Parisi, G.; Tibaldi, E.; et al. Processed Animal Proteins from Insect and Poultry By-Products in a Fish Meal-Free Diet for Rainbow Trout: Impact on Intestinal Microbiota and Inflammatory Markers. Int. J. Mol. Sci. 2021, 22, 5454. https://doi.org/10.3390/ijms22115454
Gaudioso G, Marzorati G, Faccenda F, Weil T, Lunelli F, Cardinaletti G, Marino G, Olivotto I, Parisi G, Tibaldi E, et al. Processed Animal Proteins from Insect and Poultry By-Products in a Fish Meal-Free Diet for Rainbow Trout: Impact on Intestinal Microbiota and Inflammatory Markers. International Journal of Molecular Sciences. 2021; 22(11):5454. https://doi.org/10.3390/ijms22115454
Chicago/Turabian StyleGaudioso, Giulia, Giulia Marzorati, Filippo Faccenda, Tobias Weil, Fernando Lunelli, Gloriana Cardinaletti, Giovanna Marino, Ike Olivotto, Giuliana Parisi, Emilio Tibaldi, and et al. 2021. "Processed Animal Proteins from Insect and Poultry By-Products in a Fish Meal-Free Diet for Rainbow Trout: Impact on Intestinal Microbiota and Inflammatory Markers" International Journal of Molecular Sciences 22, no. 11: 5454. https://doi.org/10.3390/ijms22115454
APA StyleGaudioso, G., Marzorati, G., Faccenda, F., Weil, T., Lunelli, F., Cardinaletti, G., Marino, G., Olivotto, I., Parisi, G., Tibaldi, E., Tuohy, K. M., & Fava, F. (2021). Processed Animal Proteins from Insect and Poultry By-Products in a Fish Meal-Free Diet for Rainbow Trout: Impact on Intestinal Microbiota and Inflammatory Markers. International Journal of Molecular Sciences, 22(11), 5454. https://doi.org/10.3390/ijms22115454






