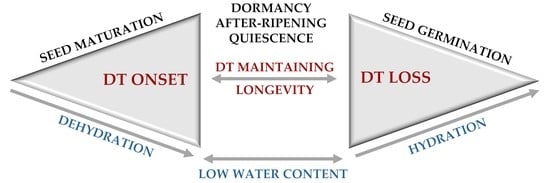
Desiccation Tolerance as the Basis of Long-Term Seed Viability
Abstract
1. Introduction
2. Mechanisms behind the Onset of Desiccation Tolerance during Seed Maturation
2.1. Late Embryogenesis Abundant (LEA) Proteins
2.2. Heat Shock Proteins (HSPs)
2.3. Non-Reducing Carbohydrates
3. Epigenetic Regulation of Desiccation Tolerance
4. Seed Longevity in the Context of Dormancy and after-Ripening
4.1. Seed Longevity
4.2. Seed Dormancy
4.3. Seed After-Ripening
5. Role of ABA and DOG1 in the Regulation of Seed Maturation
6. Mechanisms of Seed Antioxidant and Redox Protection
6.1. Seed Damages
6.2. Seed Redox Homeostasis and Antioxidant Protection
7. Repair of Nucleic Acids and Proteins in Germinating Seeds
8. Loss of Desiccation Tolerance during Seed Germination
9. Conclusions
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| ABA | Abscisic acid |
| ABI | ABA-INSENSITIVE |
| AGEs | Advanced glycoxidation end products |
| AGL | AGAMOUS-like |
| AHG | ABA Hypersensitive Germination |
| AKR | Aldo-ketoreductase |
| BER | Base excision repair |
| CMT | Chromomethylase |
| DME | Demeter |
| DOG | DELAY OF GERMINATION |
| DOGL | DELAY OF GERMINATION-like |
| DREB2D | Dehydration-responsive element-binding protein 2D |
| DT | Desiccation tolerance |
| DRM | Domains rearranged methyltransferase |
| FPG | Formamidopyrimidine-DNA glycosylase |
| FUS | FUSCA |
| GA | Gibberellic acid |
| GA3ox | Gibberellin 3-oxidase |
| GAAS | Germination Ability After STORAGE |
| HsfA | Heat shock transcription factor |
| HSI2 | High-level expression of Sugar-Inducible gene |
| HSL1 | HSI2-like1 |
| HSP | Heat shock proteins |
| LAFL | Network of transcription factors (LEC1, ABI3, FUS3 and LEC2) |
| LEA | Late embryogenesis abundant |
| LEC | LEAFY COTYLEDONS |
| LPP | Lipid phosphate phosphatase |
| MET | Methyltransferase |
| MSR | Methionine sulfoxide reductase |
| NCED | 9-cis-epoxycarotenoid dioxygenase |
| OGG 1 | 8-oxoguanine DNA glycosylase/lyase |
| PARP | Poly (ADP-ribose) polymerase |
| PIMT | L-isoaspartylmethyl transferase |
| PKL | PICKLE |
| PKR | PICKLE-RELATED |
| PLATZ | Plant AT-rich sequence- and Zinc-binding protein |
| PRC | Polycomb Repressive Complex |
| PYR/PYL/RCAR | Pyrabactin resistance/PYR-like/Regulatory component of abscisic acid receptor |
| RdDM | RNA directed DNA Methylation |
| RFO | Raffinose family oligosaccharides |
| ROS | Reactive oxygen species |
| ROS1 | Repressor of silencing1 |
| DML1 | Demeter-like1 |
| SMP | Seed maturation protein |
| SnRK | Sucrose-nonfermenting related kinase |
| TXN | Thioredoxin |
| VAL | VP1/ABI3-like |
References
- Marques, A.; Buijs, G.; Ligterink, W.; Hilhorst, H. Evolutionary ecophysiology of seed desiccation sensitivity. Funct. Plant Biol. 2018, 45, 1083. [Google Scholar] [CrossRef] [PubMed]
- Farrant, J.M.; Moore, J.P. Programming desiccation-tolerance: From plants to seeds to resurrection plants. Curr. Opin. Plant Biol. 2011, 14, 340–345. [Google Scholar] [CrossRef] [PubMed]
- Gaff, D.F.; Oliver, M. The evolution of desiccation tolerance in angiosperm plants: A rare yet common phenomenon. Funct. Plant Biol. 2013, 40, 315. [Google Scholar] [CrossRef] [PubMed]
- Costa, M.-C.D.; Cooper, K.; Hilhorst, H.W.M.; Farrant, J.M. Orthodox seeds and resurrection plants: Two of a kind? Plant Physiol. 2017, 175, 589–599. [Google Scholar] [CrossRef] [PubMed]
- Walters, C. Orthodoxy, recalcitrance and in-between: Describing variation in seed storage characteristics using threshold responses to water loss. Planta 2015, 242, 397–406. [Google Scholar] [CrossRef] [PubMed]
- Azarkovich, M.I. Dehydrins in orthodox and recalcitrant seeds. Russ. J. Plant Physiol. 2020, 67, 221–230. [Google Scholar] [CrossRef]
- Börner, A.; Khlestkina, E.K. Ex-situ genebanks—Seed treasure chambers for the future. Russ. J. Genet. 2019, 55, 1299–1305. [Google Scholar] [CrossRef]
- Leprince, O.; Pellizzaro, A.; Berriri, S.; Buitink, J. Late seed maturation: Drying without dying. J. Exp. Bot. 2017, 68, 827–841. [Google Scholar] [CrossRef]
- Smolikova, G.N. Loss of tolerance to desiccation during germination of Brasicca oleraceae L. seeds with different content of residual chlorophylls. Trans. Karelian Res. Cent. Russ. Acad. Sci. 2011, 3, 105–111. [Google Scholar]
- Zhang, Q.; Bartels, D. Molecular responses to dehydration and desiccation in desiccation-tolerant angiosperm plants. J. Exp. Bot. 2018, 69, 3211–3222. [Google Scholar] [CrossRef]
- Capron, A.; Chatfield, S.; Provart, N.; Berleth, T. Embryogenesis: Pattern formation from a single cell. Arab. B. 2009, 7, e0126. [Google Scholar] [CrossRef] [PubMed]
- Lau, S.; Slane, D.; Herud, O.; Kong, J.; Jürgens, G. Early embryogenesis in flowering plants: Setting up the basic body pattern. Annu. Rev. Plant Biol. 2012, 63, 483–506. [Google Scholar] [CrossRef] [PubMed]
- Bewley, J.D.; Bradford, K.J.; Hilhorst, H.W.M.; Nonogaki, H. Seeds: Physiology of Development, Germination and Dormancy, 3rd ed.; Springer: New York, NY, USA, 2013; ISBN 978-1-4614-4692-7. [Google Scholar]
- Lima, J.J.P.; Buitink, J.; Lalanne, D.; Rossi, R.F.; Pelletier, S.; Da Silva, E.A.A.; Leprince, O. Molecular characterization of the acquisition of longevity during seed maturation in soybean. PLoS ONE 2017, 12, e0180282. [Google Scholar]
- Cairns, N.G.; Pasternak, M.; Wachter, A.; Cobbett, C.S.; Meyer, A.J. Maturation of Arabidopsis seeds is dependent on glutathione biosynthesis within the embryo. Plant Physiol. 2006, 141, 446–455. [Google Scholar] [CrossRef] [PubMed]
- Chen, D.; Li, Y.; Fang, T.; Shi, X.; Chen, X. Specific roles of tocopherols and tocotrienols in seed longevity and germination tolerance to abiotic stress in transgenic rice. Plant Sci. 2016, 244, 31–39. [Google Scholar] [CrossRef]
- Smolikova, G.N.; Medvedev, S.S. Seed carotenoids:synthesis, diversity, and functions. Russ. J. Plant Physiol. 2015, 62, 1–13. [Google Scholar] [CrossRef]
- Hundertmark, M.; Hincha, D.K. LEA (Late Embryogenesis Abundant) proteins and their encoding genes in Arabidopsis thaliana. BMC Genom. 2008, 9, 118. [Google Scholar] [CrossRef]
- Amara, I.; Zaidi, I.; Masmoudi, K.; Ludevid, M.D.; Pagès, M.; Goday, A.; Brini, F.; Dolors Ludevid, M.; Pagès, M.; Goday, A.; et al. Insights into late embryogenesis abundant (LEA) proteins in plants: From structure to the functions. Am. J. Plant Sci. 2014, 5, 3440–3455. [Google Scholar] [CrossRef]
- Artur, M.A.S.; Zhao, T.; Ligterink, W.; Schranz, E.; Hilhorst, H.W.M. Dissecting the genomic diversification of late embryogenesis abundant (LEA) protein gene families in plants. Genome Biol. Evol. 2019, 11, 459–471. [Google Scholar] [CrossRef]
- Nagaraju, M.; Kumar, S.A.; Reddy, P.S.; Kumar, A.; Rao, D.M.; Kavi Kishor, P.B. Genome-scale identification, classification, and tissue specific expression analysis of late embryogenesis abundant (LEA) genes under abiotic stress conditions in Sorghum bicolor L. PLoS ONE 2019, 14, e0209980. [Google Scholar] [CrossRef]
- Chen, Y.; Li, C.; Zhang, B.; Yi, J.; Yang, Y.; Kong, C.; Lei, C.; Gong, M. The role of the late embryogenesis-abundant (LEA) protein family in development and the abiotic stress response: A comprehensive expression analysis of potato (Solanum Tuberosum). Genes 2019, 10, 148. [Google Scholar] [CrossRef] [PubMed]
- Battaglia, M.; Olvera-Carrillo, Y.; Garciarrubio, A.; Campos, F.; Covarrubias, A.A. The enigmatic LEA proteins and other hydrophilins. Plant Physiol. 2008, 148, 6–24. [Google Scholar] [CrossRef] [PubMed]
- Candat, A.; Paszkiewicz, G.; Neveu, M.; Gautier, R.; Logan, D.C.; Avelange-Macherel, M.-H.; Macherel, D. The ubiquitous distribution of late embryogenesis abundant proteins across cell compartments in Arabidopsis offers tailored protection against abiotic stress. Plant Cell 2014, 26, 3148–3166. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.; Zabad, S.; Liu, H.; Wang, W.; Jeffery, C. MoonProt 2.0: An expansion and update of the moonlighting proteins database. Nucleic Acids Res. 2018, 46, D640–D644. [Google Scholar] [CrossRef] [PubMed]
- Jin, X.; Cao, D.; Wang, Z.; Ma, L.; Tian, K.; Liu, Y.; Gong, Z.; Zhu, X.; Jiang, C.; Li, Y. Genome-wide identification and expression analyses of the LEA protein gene family in tea plant reveal their involvement in seed development and abiotic stress responses. Sci. Rep. 2019, 9, 14123. [Google Scholar] [CrossRef] [PubMed]
- Yu, Z.; Wang, X.; Zhang, L. Structural and functional dynamics of dehydrins: A plant protector protein under abiotic stress. Int. J. Mol. Sci. 2018, 19, 3420. [Google Scholar] [CrossRef] [PubMed]
- Radwan, A.; Hara, M.; Kleinwächter, M.; Selmar, D. Dehydrin expression in seeds and maturation drying: A paradigm change. Plant Biol. 2014, 16, 853–855. [Google Scholar] [CrossRef] [PubMed]
- Kalemba, E.M.; Pukacka, S. Possible role of LEA proteins and sHSPs in seed protection: A short review. Biol. Lett. 2007, 44, 3–16. [Google Scholar]
- Sano, N.; Rajjou, L.; North, H.M.; Debeaujon, I.; Marion-Poll, A.; Seo, M. Staying alive: Molecular aspects of seed longevity. Plant Cell Physiol. 2016, 57, 660–674. [Google Scholar] [CrossRef]
- Kaur, H.; Petla, B.P.; Kamble, N.U.; Singh, A.; Rao, V.; Salvi, P.; Ghosh, S.; Majee, M. Differentially expressed seed aging responsive heat shock protein OsHSP18.2 implicates in seed vigor, longevity and improves germination and seedling establishment under abiotic stress. Front. Plant Sci. 2015, 6, 713. [Google Scholar] [CrossRef]
- Nover, L.; Bharti, K.; Döring, P.; Mishra, S.K.; Ganguli, A.; Scharf, K.-D. Arabidopsis and the heat stress transcription factor world: How many heat stress transcription factors do we need? Cell Stress Chaperones 2001, 6, 177. [Google Scholar] [CrossRef]
- Kotak, S.; Vierling, E.; Bäumlein, H.; von Koskull-Döring, P.; Von Koskull-Dörlng, P. A novel transcriptional cascade regulating expression of heat stress proteins during seed development of Arabidopsis. Plant Cell 2007, 19, 182–195. [Google Scholar] [CrossRef] [PubMed]
- Dekkers, B.J.W.; He, H.; Hanson, J.; Willems, L.A.J.; Jamar, D.C.L.; Cueff, G.; Rajjou, L.; Hilhorst, H.W.M.; Bentsink, L. The Arabidopsis DELAY OF GERMINATION 1 gene affects ABSCISIC ACID INSENSITIVE 5 (ABI5) expression and genetically interacts with ABI3 during Arabidopsis seed development. Plant J. 2016, 85, 451–465. [Google Scholar] [CrossRef] [PubMed]
- Neto, V.G.; Barbosa, R.R.; Carosio, M.G.A.; Ferreira, A.G.; Fernandez, L.G.; de Castro, R.D.; Ligterink, W.; Hilhorst, H.; Ribeiro, P.R. Sequence analysis of Ricinus communis small heat-shock protein (sHSP) subfamily and its role in abiotic stress responses. Ind. Crops Prod. 2020, 152, 112541. [Google Scholar] [CrossRef]
- Leonova, T.; Popova, V.; Tsarev, A.; Henning, C.; Antonova, K.; Rogovskaya, N.; Vikhnina, M.; Baldensperger, T.; Soboleva, A.; Dinastia, E.; et al. Does protein glycation impact on the drought-related changes in metabolism and nutritional properties of mature pea (Pisum sativum L.) seeds? Int. J. Mol. Sci. 2020, 21, 567. [Google Scholar] [CrossRef]
- Bilova, T.; Lukasheva, E.; Brauch, D.; Greifenhagen, U.; Paudel, G.; Tarakhovskaya, E.; Frolova, N.; Mittasch, J.; Balcke, G.U.G.U.; Tissier, A.; et al. A snapshot of the plant glycated proteome: Structural, functional and mechanistic aspect. J. Biol. Chem. 2016, 291, 7621–7636. [Google Scholar] [CrossRef]
- Soboleva, A.; Schmidt, R.; Vikhnina, M.; Grishina, T.; Frolov, A. Maillard proteomics: Opening new pages. Int. J. Mol. Sci. 2017, 18, 2677. [Google Scholar] [CrossRef]
- Bilova, T.; Greifenhagen, U.; Paudel, G.; Lukasheva, E.; Brauch, D.; Osmolovskaya, N.; Tarakhovskaya, E.; Balcke, G.U.; Tissier, A.; Vogt, T.; et al. Glycation of plant proteins under environmental stress—Methodological approaches, potential mechanisms and biological role. In Abiotic and Biotic Stress in Plants Recent Advances and Future Perspectives; IntechOpen: London, UK, 2016; pp. 295–316. [Google Scholar]
- Matamoros, M.A.; Kim, A.; Peñuelas, M.; Ihling, C.; Griesser, E.; Hoffmann, R.; Fedorova, M.; Frolov, A.; Becana, M. Protein carbonylation and clycation in legume nodules. Plant Physiol. 2018, 177, 1510–1528. [Google Scholar] [CrossRef]
- Baud, S.; Dubreucq, B.; Miquel, M.; Rochat, C.; Lepiniec, L. Storage reserve accumulation in Arabidopsis: Metabolic and developmental control of seed filling. Arab. B. 2008, 6, e0113. [Google Scholar] [CrossRef]
- Hoekstra, F.; Golovina, E.; Tetteroo, F.; Wolkers, W.F. Induction of desiccation tolerance in plant somatic embryos: How exclusive is the protective role of sugars? Cryobiology 2001, 43, 140–150. [Google Scholar] [CrossRef]
- Buitink, J.; Leprince, O. Intracellular glasses and seed survival in the dry state. C. R. Biol. 2008, 331, 788–795. [Google Scholar] [CrossRef] [PubMed]
- Berr, A.; Shen, W.H. Molecular mechanisms in epigenetic regulation of plant growth and development. In Plant Developmental Biology Biotechnological Perspectives; Pua, E.C., Davey, M.R., Eds.; Springer: Berlin/Heidelberg, Germany, 2010; Volume 2, pp. 325–344. ISBN 9783642046698. [Google Scholar]
- Cabej, N.R. Plant epigenetics. In Epigenetic Principles of Evolution; Elsevier Inc.: Amsterdam, The Netherlands, 2019; pp. 733–781. ISBN 9780128140680. [Google Scholar]
- Bartels, A.; Han, Q.; Nair, P.; Stacey, L.; Gaynier, H.; Mosley, M.; Huang, Q.; Pearson, J.; Hsieh, T.-F.; An, Y.-Q.; et al. Dynamic DNA methylation in plant growth and development. Int. J. Mol. Sci. 2018, 19, 2144. [Google Scholar] [CrossRef] [PubMed]
- Cui, X.; Cao, X. Epigenetic regulation and functional exaptation of transposable elements in higher plants. Curr. Opin. Plant Biol. 2014, 21, 83–88. [Google Scholar] [CrossRef] [PubMed]
- Lebedeva, M.A.; Tvorogova, V.E.; Tikhodeyev, O.N. Epigenetic mechanisms and their role in plant development. Russ. J. Genet. 2017, 53, 1057–1071. [Google Scholar] [CrossRef]
- Lin, J.-Y.; Le, B.H.; Chen, M.; Henry, K.F.; Hur, J.; Hsieh, T.-F.; Chen, P.-Y.; Pelletier, J.M.; Pellegrini, M.; Fischer, R.L.; et al. Similarity between soybean and Arabidopsis seed methylomes and loss of non-CG methylation does not affect seed development. Proc. Natl. Acad. Sci. USA 2017, 114, E9730–E9739. [Google Scholar] [CrossRef] [PubMed]
- Kawakatsu, T.; Nery, J.R.; Castanon, R.; Ecker, J.R. Dynamic DNA methylation reconfiguration during seed development and germination. Genome Biol. 2017, 18, 1–12. [Google Scholar] [CrossRef]
- Plitta-Michalak, B.P.; Naskręt-Barciszewska, M.Z.; Kotlarski, S.; Tomaszewski, D.; Tylkowski, T.; Barciszewski, J.; Chmielarz, P.; Michalak, M. Changes in genomic 5-methylcytosine level mirror the response of orthodox (Acer platanoides L.) and recalcitrant (Acer pseudoplatanus L.) seeds to severe desiccation. Tree Physiol. 2018, 38, 617–629. [Google Scholar] [CrossRef]
- Kurek, K.; Plitta-Michalak, B.; Ratajczak, E. Reactive oxygen species as potential drivers of the seed aging process. Plants 2019, 8, 174. [Google Scholar] [CrossRef]
- Bouyer, D.; Kramdi, A.; Kassam, M.; Heese, M.; Schnittger, A.; Roudier, F.; Colot, V. DNA methylation dynamics during early plant life. Genome Biol. 2017, 18, 1–12. [Google Scholar] [CrossRef]
- Chen, M.; Lin, J.Y.; Hur, J.; Pelletier, J.M.; Baden, R.; Pellegrini, M.; Harada, J.J.; Goldberg, R.B. Seed genome hypomethylated regions are enriched in transcription factor genes. Proc. Natl. Acad. Sci. USA 2018, 115, E8315–E8322. [Google Scholar] [CrossRef]
- An, Y.Q.C.; Goettel, W.; Han, Q.; Bartels, A.; Liu, Z.; Xiao, W. Dynamic changes of genome-wide DNA methylation during soybean seed development. Sci. Rep. 2017, 7, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Michalak, M.; Barciszewska, M.Z.; Barciszewski, J.; Plitta, B.P.; Chmielarz, P. Global changes in DNA methylation in seeds and seedlings of Pyrus communis after seed desiccation and storage. PLoS ONE 2013, 8, e70693. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Kumar, S.; Qian, W. Active DNA demethylation: Mechanism and role in plant development. Plant Cell Rep. 2018, 37, 77–85. [Google Scholar] [CrossRef] [PubMed]
- Zhu, J.-K. Active DNA demethylation mediated by DNA glycosylases. Annu. Rev. Genet. 2009, 43, 143–166. [Google Scholar] [CrossRef]
- Liu, R.; Lang, Z. The mechanism and function of active DNA demethylation in plants. J. Integr. Plant Biol. 2020, 62, 148–159. [Google Scholar] [CrossRef]
- Nonogaki, H. Seed germination and dormancy: The classic story, new puzzles, and evolution. J. Integr. Plant Biol. 2019, 61, 541–563. [Google Scholar] [CrossRef] [PubMed]
- Lepiniec, L.; Devic, M.; Roscoe, T.J.; Bouyer, D.; Zhou, D.-X.; Boulard, C.; Baud, S.; Dubreucq, B. Molecular and epigenetic regulations and functions of the LAFL transcriptional regulators that control seed development. Plant Reprod. 2018, 31, 291–307. [Google Scholar] [CrossRef]
- Huo, H.; Wei, S.; Bradford, K.J. DELAY OF GERMINATION1 (DOG1) regulates both seed dormancy and flowering time through microRNA pathways. Proc. Natl. Acad. Sci. USA 2016, 113, E2199–E2206. [Google Scholar] [CrossRef]
- Colville, L.; Pritchard, H.W. Seed life span and food security. New Phytol. 2019, 224, 557–562. [Google Scholar] [CrossRef]
- Zinsmeister, J.; Leprince, O.; Buitink, J. Molecular and environmental factors regulating seed longevity. Biochem. J. 2020, 477, 305–323. [Google Scholar] [CrossRef]
- Shvachko, N.A.; Khlestkina, E.K. Molecular genetic bases of seed resistance to oxidative stress during storage. Vavilov J. Genet. Breed. 2020, 24, 451–458. [Google Scholar] [CrossRef]
- Sallon, S.; Solowey, E.; Cohen, Y.; Korchinsky, R.; Egli, M.; Woodhatch, I.; Simchoni, O.; Kislev, M. Germination, genetics, and growth of an ancient date seed. Science 2008, 320, 1464. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, T.-P.; Keizer, P.; van Eeuwijk, F.; Smeekens, S.; Bentsink, L. Natural variation for seed longevity and seed dormancy are negatively correlated in Arabidopsis. Plant Physiol. 2012, 160, 2083–2092. [Google Scholar] [CrossRef]
- Finkelstein, R.; Reeves, W.; Ariizumi, T.; Steber, C. Molecular aspects of seed dormancy. Annu. Rev. Plant Biol. 2008, 59, 387–415. [Google Scholar] [CrossRef] [PubMed]
- Baskin, J.M.; Baskin, C.C. A classification system for seed dormancy. Seed Sci. Res. 2007, 14, 1–16. [Google Scholar] [CrossRef]
- Nikolaeva, M.G. Factors controlling the seed dormancy pattern. In The Physiology and Biochemistry of Seed Dormancy; Khan, A.A., Ed.; North Holland: Amsterdam, The Netherlands, 1977; pp. 51–74. [Google Scholar]
- Graeber, K.; Nakabayashi, K.; Miatton, E.; Leubner-Metzger, G.; Soppe, W.J.J. Molecular mechanisms of seed dormancy. Plant. Cell Environ. 2012, 35, 1769–1786. [Google Scholar] [CrossRef]
- Chahtane, H.; Kim, W.; Lopez-Molina, L. Primary seed dormancy: A temporally multilayered riddle waiting to be unlocked. J. Exp. Bot. 2017, 68, 857–869. [Google Scholar] [CrossRef]
- Finch-Savage, W.E.; Leubner-Metzger, G. Seed dormancy and the control of germination. New Phytol. 2006, 171, 501–523. [Google Scholar] [CrossRef]
- Leubner-Metzger, G. β-1,3-Glucanase gene expression in low-hydrated seeds as a mechanism for dormancy release during tobacco after-ripening. Plant J. 2005, 41, 133–145. [Google Scholar] [CrossRef]
- Carrera, E.; Holman, T.; Medhurst, A.; Dietrich, D.; Footitt, S.; Theodoulou, F.L.; Holdsworth, M.J. Seed after-ripening is a discrete developmental pathway associated with specific gene networks in Arabidopsis. Plant J. 2008, 53, 214–224. [Google Scholar] [CrossRef]
- Ishikawa, S.; Barrero, J.M.; Takahashi, F.; Nakagami, H.; Peck, S.C.; Gubler, F.; Shinozaki, K.; Umezawa, T. Comparative phosphoproteomic analysis reveals a decay of ABA signaling in barley embryos during after-ripening. Plant Cell Physiol. 2019, 60, 2758–2768. [Google Scholar] [CrossRef]
- Holdsworth, M.J.; Bentsink, L.; Soppe, W.J.J. Molecular networks regulating Arabidopsis seed maturation, after-ripening, dormancy and germination. New Phytol. 2008, 179, 33–54. [Google Scholar] [CrossRef] [PubMed]
- Iglesias-Fernández, R.; del Carmen Rodríguez-Gacio, M.; Matilla, A.J. Progress in research on dry afterripening. Seed Sci. Res. 2011, 21, 69–80. [Google Scholar] [CrossRef]
- Oracz, K.; Bouteau, H.E.-M.; Farrant, J.M.; Cooper, K.; Belghazi, M.; Job, C.; Job, D.; Corbineau, F.; Bailly, C. ROS production and protein oxidation as a novel mechanism for seed dormancy alleviation. Plant J. 2007, 50, 452–465. [Google Scholar] [CrossRef] [PubMed]
- Finch-Savage, W.E.; Footitt, S. Seed dormancy cycling and the regulation of dormancy mechanisms to time germination in variable field environments. J. Exp. Bot. 2017, 68, 843–856. [Google Scholar] [CrossRef]
- Gutierrez, L.; Van Wuytswinkel, O.; Castelain, M.; Bellini, C. Combined networks regulating seed maturation. Trends Plant Sci. 2007, 12, 294–300. [Google Scholar] [CrossRef]
- Chen, K.; Li, G.; Bressan, R.A.; Song, C.; Zhu, J.; Zhao, Y. Abscisic acid dynamics, signaling, and functions in plants. J. Integr. Plant Biol. 2020, 62, 25–54. [Google Scholar] [CrossRef]
- Nambara, E.; Marion-Poll, A. Abscisic acid biosynthesis and catabolism. Annu. Rev. Plant Biol. 2005, 56, 165–185. [Google Scholar] [CrossRef]
- Finkelstein, R. Abscisic acid synthesis and response. Arab. B. 2013, 11, e0166. [Google Scholar] [CrossRef]
- Sall, K.; Dekkers, B.J.W.; Nonogaki, M.; Katsuragawa, Y.; Koyari, R.; Hendrix, D.; Willems, L.A.J.; Bentsink, L.; Nonogaki, H. DELAY OF GERMINATION 1-LIKE 4 acts as an inducer of seed reserve accumulation. Plant J. 2019, 100, 7–19. [Google Scholar] [CrossRef]
- Jo, L.; Pelletier, J.M.; Harada, J.J. Central role of the LEAFY COTYLEDON1 transcription factor in seed development. J. Integr. Plant Biol. 2019, 61, 564–580. [Google Scholar] [CrossRef] [PubMed]
- Soppe, W.J.J.; Bentsink, L. Seed dormancy back on track; its definition and regulation by DOG1. New Phytol. 2020, 228, 816–819. [Google Scholar] [CrossRef] [PubMed]
- Zinsmeister, J.; Lalanne, D.; Terrasson, E.; Chatelain, E.; Vandecasteele, C.; Vu, B.L.; Dubois-Laurent, C.; Geoffriau, E.; Le Signor, C.; Dalmais, M.; et al. ABI5 is a regulator of seed maturation and longevity in legumes. Plant Cell 2016, 28, 2735–2754. [Google Scholar] [CrossRef]
- Smolikova, G.; Dolgikh, E.; Vikhnina, M.; Frolov, A.; Medvedev, S. Genetic and hormonal regulation of chlorophyll degradation during maturation of seeds with green embryos. Int. J. Mol. Sci. 2017, 18, 1993. [Google Scholar] [CrossRef]
- Smolikova, G.; Shiroglazova, O.; Vinogradova, G.; Leppyanen, I.; Dinastiya, E.; Yakovleva, O.; Dolgikh, E.; Titova, G.; Frolov, A.; Medvedev, S. Comparative analysis of the plastid conversion, photochemical activity and chlorophyll degradation in developing embryos of green-seeded and yellow-seeded pea (Pisum sativum) cultivars. Funct. Plant Biol. 2020, 47, 409–424. [Google Scholar] [CrossRef] [PubMed]
- Smolikova, G.N.; Medvedev, S.S. Photosynthesis in the seeds of chloroembryophytes. Russ. J. Plant Physiol. 2016, 63, 1–12. [Google Scholar] [CrossRef]
- Smolikova, G.; Kreslavski, V.; Shiroglazova, O.; Bilova, T.; Sharova, E.; Frolov, A.; Medvedev, S. Photochemical activity changes accompanying the embryogenesis of pea (Pisum sativum) with yellow and green cotyledons. Funct. Plant Biol. 2018, 45, 228. [Google Scholar] [CrossRef] [PubMed]
- Mamontova, T.; Lukasheva, E.; Mavropolo-Stolyarenko, G.; Proksch, C.; Bilova, T.; Kim, A.; Babakov, V.; Grishina, T.; Hoehenwarter, W.; Medvedev, S.; et al. Proteome map of pea (Pisum sativum L.) embryos containing different amounts of residual chlorophylls. Int. J. Mol. Sci. 2018, 19, 4066. [Google Scholar] [CrossRef] [PubMed]
- González-Morales, S.I.; Chávez-Montes, R.A.; Hayano-Kanashir, C.; Alejo-Jacuind, G.; Rico-Cambron, T.Y.; De Folte, S.; Herrera-Estrell, L. Regulatory network analysis reveals novel regulators of seed desiccation tolerance in Arabidopsis thaliana. Proc. Natl. Acad. Sci. USA 2016, 113, E5232–E5241. [Google Scholar] [CrossRef]
- Braybrook, S.; Harada, J. LECs go crazy in embryo development. Trends Plant Sci. 2008, 13, 624–630. [Google Scholar] [CrossRef]
- Nambara, E.; Okamoto, M.; Tatematsu, K.; Yano, R.; Seo, M.; Kamiya, Y. Abscisic acid and the control of seed dormancy and germination. Seed Sci. Res. 2010, 20, 55. [Google Scholar] [CrossRef]
- Dekkers, B.J.W.; Bentsink, L. Regulation of seed dormancy by abscisic acid and delay of germination 1. Seed Sci. Res. 2015, 25, 82–98. [Google Scholar] [CrossRef]
- Nishimura, N.; Tsuchiya, W.; Moresco, J.J.; Hayashi, Y.; Satoh, K.; Kaiwa, N.; Irisa, T.; Kinoshita, T.; Schroeder, J.I.; Yates, J.R.; et al. Control of seed dormancy and germination by DOG1-AHG1 PP2C phosphatase complex via binding to heme. Nat. Commun. 2018, 9, 2132. [Google Scholar] [CrossRef]
- Nakabayashi, K.; Bartsch, M.; Xiang, Y.; Miatton, E.; Pellengahr, S.; Yano, R.; Seo, M.; Soppe, W.J.J. The time required for dormancy release in Arabidopsis is determined by DELAY OF GERMINATION1 protein levels in freshly harvested seeds. Plant Cell 2012, 24, 2826–2838. [Google Scholar] [CrossRef] [PubMed]
- El-Maarouf-Bouteau, H.; Meimoun, P.; Job, C.; Job, D.; Bailly, C. Role of protein and mRNA oxidation in seed dormancy and germination. Front. Plant Sci. 2013, 4, 77. [Google Scholar] [CrossRef]
- Bailly, C. Active oxygen species and antioxidants in seed biology. Seed Sci. Res. 2004, 14, 93–107. [Google Scholar] [CrossRef]
- Frolov, A.; Mamontova, T.; Ihling, C.; Lukasheva, E.; Bankin, M.; Chantseva, V.; Vikhnina, M.; Soboleva, A.; Shumilina, J.; Mavropolo-Stolyarenko, G.; et al. Mining seed proteome: From protein dynamics to modification profiles. Biol. Commun. 2018, 63, 43–58. [Google Scholar] [CrossRef]
- Antonova, K.; Vikhnina, M.; Soboleva, A.; Mehmood, T.; Heymich, M.; Leonova, T.; Bankin, M.; Lukasheva, E.; Gensberger-Reigl, S.; Medvedev, S.; et al. Analysis of chemically labile glycation adducts in seed proteins: Case study of methylglyoxal-derived hydroimidazolone 1 (MG-H1). Int. J. Mol. Sci. 2019, 20, 3659. [Google Scholar] [CrossRef]
- Shumilina, J.; Kusnetsova, A.; Tsarev, A.; Janse van Rensburg, H.C.; Medvedev, S.; Demidchik, V.; Van den Ende, W.; Frolov, A. Glycation of plant proteins: Regulatory roles and interplay with sugar signalling? Int. J. Mol. Sci. 2019, 20, 2366. [Google Scholar] [CrossRef]
- Kranner, I.; Minibayeva, F.V.; Beckett, R.P.; Seal, C.E. What is stress? Concepts, definitions and applications in seed science. New Phytol. 2010, 188, 655–673. [Google Scholar] [CrossRef]
- Veselova, T.V.; Veselovsky, V.A.; Obroucheva, N.V. Deterioration mechanisms in air-dry pea seeds during early aging. Plant Physiol. Biochem. 2015, 87, 133–139. [Google Scholar] [CrossRef] [PubMed]
- Geigenberger, P.; Fernie, A.R. Metabolic control of redox and redox control of metabolism in plants. Antioxid. Redox Signal. 2014, 21, 1389–1421. [Google Scholar] [CrossRef] [PubMed]
- Pradedova, E.V.; Nimaeva, O.D.; Salyaev, R.K. Redox processes in biological systems. Russ. J. Plant Physiol. 2017, 64, 822–832. [Google Scholar] [CrossRef]
- Demidchik, V. Reactive oxygen species and their role in plant oxidative stress. In Plant Stress Physiology; Shabala, S., Ed.; CABI: Wallingford, UK, 2017; ISBN 9781780647296. [Google Scholar]
- Waszczak, C.; Carmody, M.; Kangasjärvi, J. Reactive oxygen species in plant signaling. Annu. Rev. Plant Biol. 2018, 69, 209–236. [Google Scholar] [CrossRef]
- Bailly, C. The signalling role of ROS in the regulation of seed germination and dormancy. Biochem. J. 2019, 476, 3019–3032. [Google Scholar] [CrossRef]
- Mittler, R. Oxidative stress, antioxidants and stress tolerance. Trends Plant Sci. 2002, 7, 405–410. [Google Scholar] [CrossRef]
- Jajic, I.; Sarna, T.; Strzalka, K. Senescence, stress, and reactive oxygen species. Plants 2015, 4, 393–411. [Google Scholar] [CrossRef]
- Mignolet-Spruyt, L.; Xu, E.; Idänheimo, N.; Hoeberichts, F.A.; Mühlenbock, P.; Brosché, M.; Van Breusegem, F.; Kangasjärvi, J. Spreading the news: Subcellular and organellar reactive oxygen species production and signalling. J. Exp. Bot. 2016, 67, 3831–3844. [Google Scholar] [CrossRef]
- Basudhar, D.; Ridnour, L.A.; Cheng, R.; Kesarwala, A.H.; Heinecke, J.; Wink, D.A. Biological signaling by small inorganic molecules. Coord. Chem. Rev. 2016, 306, 708–723. [Google Scholar] [CrossRef]
- Corpas, F.J.; González-Gordo, S.; Cañas, A.; Palma, J.M. Nitric oxide and hydrogen sulfide in plants: Which comes first? J. Exp. Bot. 2019, 70, 4391–4404. [Google Scholar] [CrossRef]
- Mano, J.; Kanameda, S.; Kuramitsu, R.; Matsuura, N.; Yamauchi, Y. Detoxification of reactive carbonyl species by glutathione transferase tau isozymes. Front. Plant Sci. 2019, 10, 487. [Google Scholar] [CrossRef] [PubMed]
- Bailly, C.; El-Maarouf-Bouteau, H.; Corbineau, F. From intracellular signaling networks to cell death: The dual role of reactive oxygen species in seed physiology. Comptes Rendus Biol. 2008, 331, 806–814. [Google Scholar] [CrossRef] [PubMed]
- Del Río, L.A. ROS and RNS in plant physiology: An overview. J. Exp. Bot. 2015, 66, 2827–2837. [Google Scholar] [CrossRef] [PubMed]
- Mittler, R. ROS are good. Trends Plant Sci. 2017, 22, 11–19. [Google Scholar] [CrossRef]
- Frolov, A.; Bilova, T.; Paudel, G.; Berger, R.; Balcke, G.U.; Birkemeyer, C.; Wessjohann, L.A. Early responses of mature Arabidopsis thaliana plants to reduced water potential in the agar-based polyethylene glycol infusion drought model. J. Plant Physiol. 2017, 208, 70–83. [Google Scholar] [CrossRef]
- Goldstein, S.; Meyerstein, D.; Czapski, G. The Fenton reagents. Free Radic. Biol. Med. 1993, 15, 435–445. [Google Scholar] [CrossRef]
- Hajiboland, R. Reactive oxygen species and photosynthesis. In Oxidative Damage to Plants; Elsevier Inc.: Amsterdam, The Netherlands, 2014; pp. 1–63. ISBN 9780127999630. [Google Scholar]
- Sharma, P.; Jha, A.B.; Dubey, R.S.; Pessarakli, M. Reactive oxygen species, oxidative damage, and antioxidative defense mechanism in plants under stressful conditions. J. Bot. 2012, 2012, 1–26. [Google Scholar] [CrossRef]
- Mittler, R.; Vanderauwera, S.; Suzuki, N.; Miller, G.; Tognetti, V.B.; Vandepoele, K.; Gollery, M.; Shulaev, V.; Van Breusegem, F. ROS signaling: The new wave? Trends Plant Sci. 2011, 16, 300–309. [Google Scholar] [CrossRef]
- Kreslavski, V.D.; Los, D.A.; Allakhverdiev, S.I.; Kuznetsov, V.V. Signaling role of reactive oxygen species in plants under stress. Russ. J. Plant Physiol. 2012, 59, 141–154. [Google Scholar] [CrossRef]
- Kalemba, E.M.; Pukacka, S. Carbonylated proteins accumulated as vitality decreases during long-term storage of beech (Fagus sylvatica L.) seeds. Trees 2014, 28, 503–515. [Google Scholar] [CrossRef]
- Osmolovskaya, N.; Shumilina, J.; Kim, A.; Didio, A.; Grishina, T.; Bilova, T.; Keltsieva, O.A.; Zhukov, V.; Tikhonovich, I.; Tarakhovskaya, E.; et al. Methodology of drought stress research: Experimental setup and physiological characterization. Int. J. Mol. Sci. 2018, 19, 4089. [Google Scholar] [CrossRef] [PubMed]
- Mullineaux, P.M.; Baker, N.R. Oxidative stress: Antagonistic signaling for acclimation or cell death? Plant Physiol. 2010, 154, 521–525. [Google Scholar] [CrossRef] [PubMed]
- Tommasi, F.; Paciolla, C.; de Pinto, M.C.; Gara, L. De A comparative study of glutathione and ascorbate metabolism during germination of Pinus pinea L. seeds. J. Exp. Bot. 2001, 52, 1647–1654. [Google Scholar] [CrossRef] [PubMed]
- Noctor, G.; Foyer, C.H. Ascorbate and glutathione: Keeping active oxygen under control. Annu. Rev. Plant Physiol. Plant Mol. Biol. 1998, 49, 249–279. [Google Scholar] [CrossRef]
- Roach, T.; Nagel, M.; Börner, A.; Eberle, C.; Kranner, I. Changes in tocochromanols and glutathione reveal differences in the mechanisms of seed ageing under seedbank conditions and controlled deterioration in barley. Environ. Exp. Bot. 2018, 156, 8–15. [Google Scholar] [CrossRef]
- Kranner, I.; Birtić, S.; Anderson, K.M.; Pritchard, H.W. Glutathione half-cell reduction potential: A universal stress marker and modulator of programmed cell death? Free Radic. Biol. Med. 2006, 40, 2155–2165. [Google Scholar] [CrossRef]
- Morscher, F.; Kranner, I.; Arc, E.; Bailly, C.; Roach, T. Glutathione redox state, tocochromanols, fatty acids, antioxidant enzymes and protein carbonylation in sunflower seed embryos associated with after-ripening and ageing. Ann. Bot. 2015, 116, 669–678. [Google Scholar] [CrossRef]
- Begara-Morales, J.C.; Sánchez-Calvo, B.; Chaki, M.; Valderrama, R.; Mata-Pérez, C.; Padilla, M.N.; Corpas, F.J.; Barroso, J.B. Antioxidant systems are regulated by nitric oxide-mediated post-translational modifications (NO-PTMs). Front. Plant Sci. 2016, 7, 152. [Google Scholar] [CrossRef]
- Mamaeva, A.S.; Fomenkov, A.A.; Nosov, A.V.; Moshkov, I.E.; Mur, L.A.J.; Hall, M.A.; Novikova, G.V. Regulatory role of nitric oxide in plants. Russ. J. Plant Physiol. 2015, 62, 427–440. [Google Scholar] [CrossRef]
- Arasimowicz-Jelonek, M.; Floryszak-Wieczorek, J. A physiological perspective on targets of nitration in NO-based signaling networks in plants. J. Exp. Bot. 2019, 70, 4379–4389. [Google Scholar] [CrossRef]
- Khan, M.N.; Mobin, M.; Mohammad, F.; Corpas, F.J. (Eds.) Nitric Oxide in Plants: Metabolism and Role in Stress Physiology; Springer International Publishing: Cham, Switzerland, 2014; ISBN 978-3-319-06709-4. [Google Scholar]
- Wendehenne, D.; Hancock, J.T. New frontiers in nitric oxide biology in plant. Plant Sci. 2011, 181, 507–508. [Google Scholar] [CrossRef]
- Yu, M.; Lamattina, L.; Spoel, S.H.; Loake, G.J. Nitric oxide function in plant biology: A redox cue in deconvolution. New Phytol. 2014, 202, 1142–1156. [Google Scholar] [CrossRef]
- Scheler, C.; Durner, J.; Astier, J. Nitric oxide and reactive oxygen species in plant biotic interactions. Curr. Opin. Plant Biol. 2013, 16, 534–539. [Google Scholar] [CrossRef] [PubMed]
- Bethke, P.C.; Libourel, I.G.L.; Jones, R.L. Nitric oxide reduces seed dormancy in Arabidopsis. J. Exp. Bot. 2006, 57, 517–526. [Google Scholar] [CrossRef] [PubMed]
- Libourel, I.G.L.; Bethke, P.C.; De Michele, R.; Jones, R.L. Nitric oxide gas stimulates germination of dormant Arabidopsis seeds: Use of a flow-through apparatus for delivery of nitric oxide. Planta 2006, 223, 813–820. [Google Scholar] [CrossRef] [PubMed]
- Gniazdowska, A.; Dobrzyńska, U.; Babańczyk, T.; Bogatek, R. Breaking the apple embryo dormancy by nitric oxide involves the stimulation of ethylene production. Planta 2007, 225, 1051–1057. [Google Scholar] [CrossRef]
- Signorelli, S.; Considine, M.J. Nitric oxide enables germination by a four-pronged attack on ABA-induced seed dormancy. Front. Plant Sci. 2018, 9, 296. [Google Scholar] [CrossRef]
- Prakash, V.; Singh, V.P.; Tripathi, D.K.; Sharma, S.; Corpas, F.J. Crosstalk between nitric oxide (NO) and abscisic acid (ABA) signalling molecules in higher plants. Environ. Exp. Bot. 2019, 161, 41–49. [Google Scholar] [CrossRef]
- Ogé, L.; Bourdais, G.; Bove, J.; Collet, B.; Godin, B.; Granier, F.; Boutin, J.P.; Job, D.; Jullien, M.; Grappin, P. Protein repair L-Isoaspartyl methyltransferasel is involved in both seed longevity and germination vigor in arabidopsis. Plant Cell 2008, 20, 3022–3037. [Google Scholar] [CrossRef]
- Waterworth, W.M.; Bray, C.M.; West, C.E. Seeds and the art of genome maintenance. Front. Plant Sci. 2019, 10, 1–11. [Google Scholar] [CrossRef]
- Faria, J.M.R.; Buitink, J.; van Lammeren, A.A.M.; Hilhorst, H.W.M. Changes in DNA and microtubules during loss and re-establishment of desiccation tolerance in germinating Medicago truncatula seeds. J. Exp. Bot. 2005, 56, 2119–2130. [Google Scholar] [CrossRef] [PubMed]
- Ventura, L.; Donà, M.; Macovei, A.; Carbonera, D.; Buttafava, A.; Mondoni, A.; Rossi, G.; Balestrazzi, A. Understanding the molecular pathways associated with seed vigor. Plant Physiol. Biochem. 2012, 60, 196–206. [Google Scholar] [CrossRef] [PubMed]
- Chen, H.; Chu, P.; Zhou, Y.; Li, Y.; Liu, J.; Ding, Y.; Tsang, E.W.T.; Jiang, L.; Wu, K.; Huang, S. Overexpression of AtOGG1, a DNA glycosylase/AP lyase, enhances seed longevity and abiotic stress tolerance in Arabidopsis. J. Exp. Bot. 2012, 63, 4107–4121. [Google Scholar] [CrossRef]
- Roldán-Arjona, T.; Ariza, R.R.; Córdoba-Cañero, D. DNA base excision repair in plants: An unfolding story with familiar and novel characters. Front. Plant Sci. 2019, 10, 1055. [Google Scholar] [CrossRef] [PubMed]
- Cõrdoba-Cañero, D.; Roldán-Arjona, T.; Ariza, R.R. Arabidopsis ZDP DNA 3’-phosphatase and ARP endonuclease function in 8-oxoG repair initiated by FPG and OGG1 DNA glycosylases. Plant J. 2014, 79, 824–834. [Google Scholar] [CrossRef]
- Lamb, R.S.; Citarelli, M.; Teotia, S. Functions of the poly(ADP-ribose) polymerase superfamily in plants. Cell. Mol. Life Sci. 2012, 69, 175–189. [Google Scholar] [CrossRef]
- Amé, J.-C.; Spenlehauer, C.; de Murcia, G. The PARP superfamily. BioEssays 2004, 26, 882–893. [Google Scholar] [CrossRef]
- Feng, B.; Ma, S.; Chen, S.; Zhu, N.; Zhang, S.; Yu, B.; Yu, Y.; Le, B.; Chen, X.; Dinesh-Kumar, S.P.; et al. PARylation of the forkhead-associated domain protein DAWDLE regulates plant immunity. EMBO Rep. 2016, 17, 1799–1813. [Google Scholar] [CrossRef]
- Rissel, D.; Heym, P.P.; Thor, K.; Brandt, W.; Wessjohann, L.A.; Peiter, E. No silver bullet—Canonical poly(ADP-ribose) polymerases (PARPs) are no universal factors of abiotic and biotic stress resistance of Arabidopsis thaliana. Front. Plant Sci. 2017, 8, 59. [Google Scholar] [CrossRef]
- Fedorova, M.; Frolov, A.; Hoffmann, R. Fragmentation behavior of Amadori-peptides obtained by non-enzymatic glycosylation of lysine residues with ADP-ribose in tandem mass spectrometry. J. Mass Spectrom. 2010, 45, 664–669. [Google Scholar] [CrossRef]
- Zheng, Q.; Omans, N.D.; Leicher, R.; Osunsade, A.; Agustinus, A.S.; Finkin-Groner, E.; D’Ambrosio, H.; Liu, B.; Chandarlapaty, S.; Liu, S.; et al. Reversible histone glycation is associated with disease-related changes in chromatin architecture. Nat. Commun. 2019, 10, 1289. [Google Scholar] [CrossRef] [PubMed]
- Bai, B.; Peviani, A.; van der Horst, S.; Gamm, M.; Snel, B.; Bentsink, L.; Hanson, J. Extensive translational regulation during seed germination revealed by polysomal profiling. New Phytol. 2017, 214, 233–244. [Google Scholar] [CrossRef] [PubMed]
- Sano, N.; Rajjou, L.; North, H.M. Lost in translation: Physiological roles of stored mRNAs in seed germination. Plants 2020, 9, 347. [Google Scholar] [CrossRef] [PubMed]
- Dirk, L.M.A.; Downie, A.B. An examination of Job’s rule: Protection and repair of the proteins of the translational apparatus in seeds. Seed Sci. Res. 2018, 28, 168–181. [Google Scholar] [CrossRef]
- Sano, N.; Ono, H.; Murata, K.; Yamada, T.; Hirasawa, T.; Kanekatsu, M. Accumulation of long-lived mRNAs associated with germination in embryos during seed development of rice. J. Exp. Bot. 2015, 66, 4035–4046. [Google Scholar] [CrossRef]
- Rajjou, L.; Debeaujon, I. Seed longevity: Survival and maintenance of high germination ability of dry seeds. C. R. Biol. 2008, 331, 796–805. [Google Scholar] [CrossRef] [PubMed]
- Kranner, I.; Chen, H.; Pritchard, H.W.; Pearce, S.R.; Birtić, S. Inter-nucleosomal DNA fragmentation and loss of RNA integrity during seed ageing. Plant Growth Regul. 2011, 63, 63–72. [Google Scholar] [CrossRef]
- Rajjou, L.; Lovigny, Y.; Groot, S.P.C.; Belghazi, M.; Job, C.; Job, D. Proteome-wide characterization of seed aging in Arabidopsis: A comparison between artificial and natural aging protocols. Plant Physiol. 2008, 148, 620–641. [Google Scholar] [CrossRef]
- Burroughs, A.M.; Aravind, L. RNA damage in biological conflicts and the diversity of responding RNA repair systems. Nucleic Acids Res. 2016, 44, 8525–8555. [Google Scholar] [CrossRef]
- Châtelain, E.; Satour, P.; Laugier, E.; Vu, B.L.; Payet, N.; Rey, P.; Montrichard, F. Evidence for participation of the methionine sulfoxide reductase repair system in plant seed longevity. Proc. Natl. Acad. Sci. USA 2013, 110, 3633–3638. [Google Scholar] [CrossRef]
- Buchanan, B.B.; Balmer, Y. Redox regulation: A broadening horizon. Annu. Rev. Plant Biol. 2005, 56, 187–220. [Google Scholar] [CrossRef] [PubMed]
- Hägglund, P.; Finnie, C.; Yano, H.; Shahpiri, A.; Buchanan, B.B.; Henriksen, A.; Svensson, B. Seed thioredoxin h. Biochim. Biophys. Acta Proteins Proteomics 2016, 1864, 974–982. [Google Scholar] [CrossRef]
- Soboleva, A.; Vikhnina, M.; Grishina, T.; Frolov, A. Probing protein glycation by chromatography and mass spectrometry: Analysis of glycation adducts. Int. J. Mol. Sci. 2017, 18, 2557. [Google Scholar] [CrossRef] [PubMed]
- Zinsmeister, J.; Berriri, S.; Basso, D.P.; Ly-Vu, B.; Dang, T.; Lalanne, D.; Silva, E.A.A.; Leprince, O.; Buitink, J. The seed-specific heat shock factor A9 regulates the depth of dormancy in Medicago truncatula seeds via ABA signaling. Plant Cell Environ. 2020, 43, 2508–2522. [Google Scholar] [CrossRef] [PubMed]
- Job, C.; Rajjou, L.; Lovigny, Y.; Belghazi, M.; Job, D. Patterns of protein oxidation in Arabidopsis seeds and during germination. Plant Physiol. 2005, 138, 790–802. [Google Scholar] [CrossRef]
- Nisarga, K.N.; Vemanna, R.S.; Chandrashekar, B.K.; Rao, H.; Vennapusa, A.R.; Narasimaha, A.; Makarla, U.; Basavaiah, M.R. Aldo-ketoreductase 1 (AKR1) improves seed longevity in tobacco and rice by detoxifying reactive cytotoxic compounds generated during ageing. Rice 2017, 10, 1–12. [Google Scholar]
- Obroucheva, N.V.; Sinkevich, I.A.; Lityagina, S.V.; Novikova, G.V. Water relations in germinating seeds. Russ. J. Plant Physiol. 2017, 64, 625–633. [Google Scholar] [CrossRef]
- Dekkers, B.J.W.; Costa, M.C.D.; Maia, J.; Bentsink, L.; Ligterink, W.; Hilhorst, H.W.M. Acquisition and loss of desiccation tolerance in seeds: From experimental model to biological relevance. Planta 2015, 241, 563–577. [Google Scholar] [CrossRef]
- Zha, P.; Liu, S.; Li, Y.; Ma, T.; Yang, L.; Jing, Y.; Lin, R. The evening complex and the chromatin-remodeling factor PICKLE coordinately control seed dormancy by directly repressing DOG1 in Arabidopsis. Plant Commun. 2020, 1, 100011. [Google Scholar] [CrossRef]

Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Smolikova, G.; Leonova, T.; Vashurina, N.; Frolov, A.; Medvedev, S. Desiccation Tolerance as the Basis of Long-Term Seed Viability. Int. J. Mol. Sci. 2021, 22, 101. https://doi.org/10.3390/ijms22010101
Smolikova G, Leonova T, Vashurina N, Frolov A, Medvedev S. Desiccation Tolerance as the Basis of Long-Term Seed Viability. International Journal of Molecular Sciences. 2021; 22(1):101. https://doi.org/10.3390/ijms22010101
Chicago/Turabian StyleSmolikova, Galina, Tatiana Leonova, Natalia Vashurina, Andrej Frolov, and Sergei Medvedev. 2021. "Desiccation Tolerance as the Basis of Long-Term Seed Viability" International Journal of Molecular Sciences 22, no. 1: 101. https://doi.org/10.3390/ijms22010101
APA StyleSmolikova, G., Leonova, T., Vashurina, N., Frolov, A., & Medvedev, S. (2021). Desiccation Tolerance as the Basis of Long-Term Seed Viability. International Journal of Molecular Sciences, 22(1), 101. https://doi.org/10.3390/ijms22010101