Genome-Wide Identification and Analysis on YUCCA Gene Family in Isatis indigotica Fort. and IiYUCCA6-1 Functional Exploration
Abstract
1. Introduction
2. Results
2.1. IiYUCCA Gene Family Analysis in I. indigotica
2.2. Classification and Structural Analysis of IiYUCCA Genes
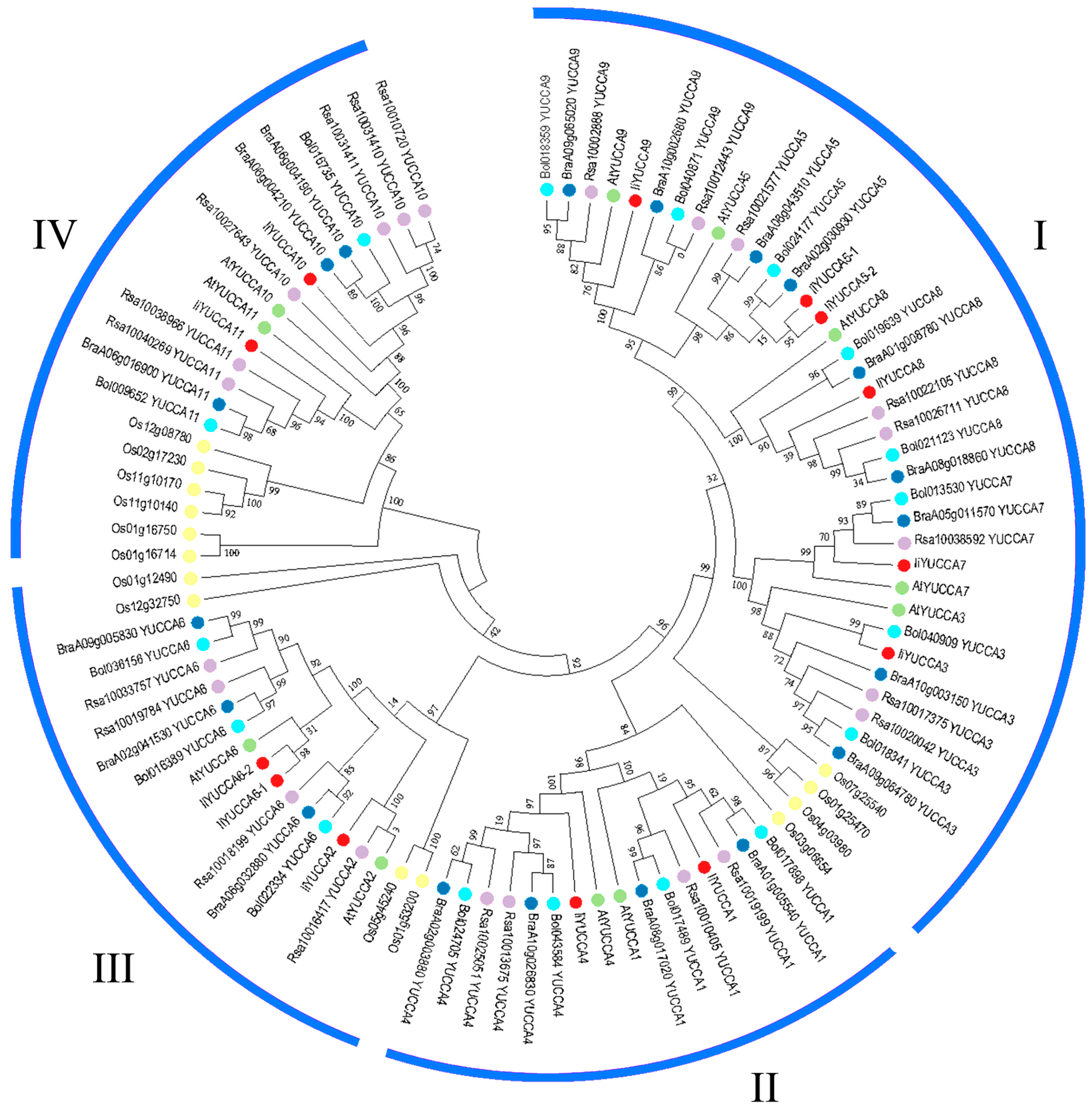
2.3. Phylogenetic Analysis of YUCCA Gene Family
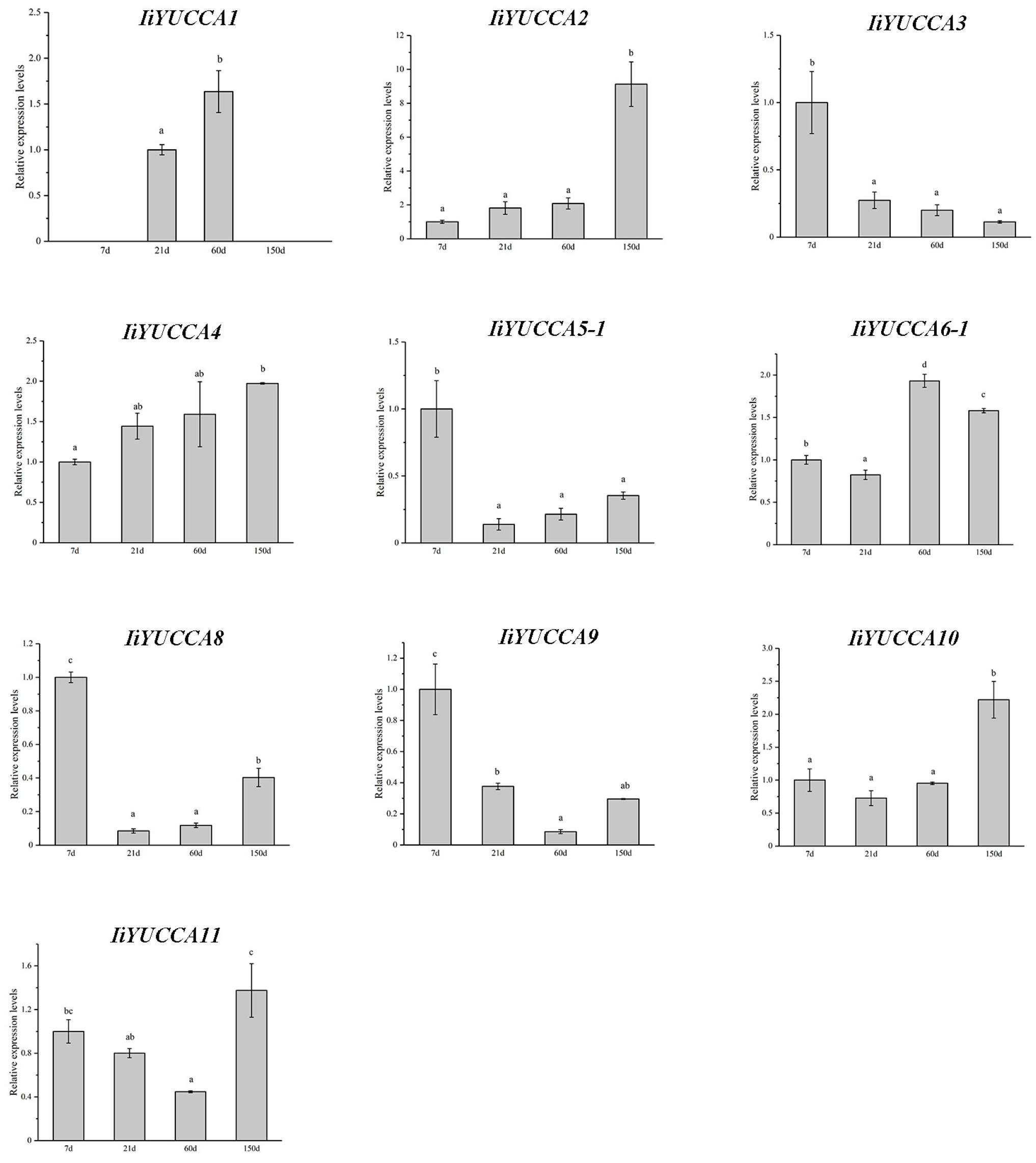
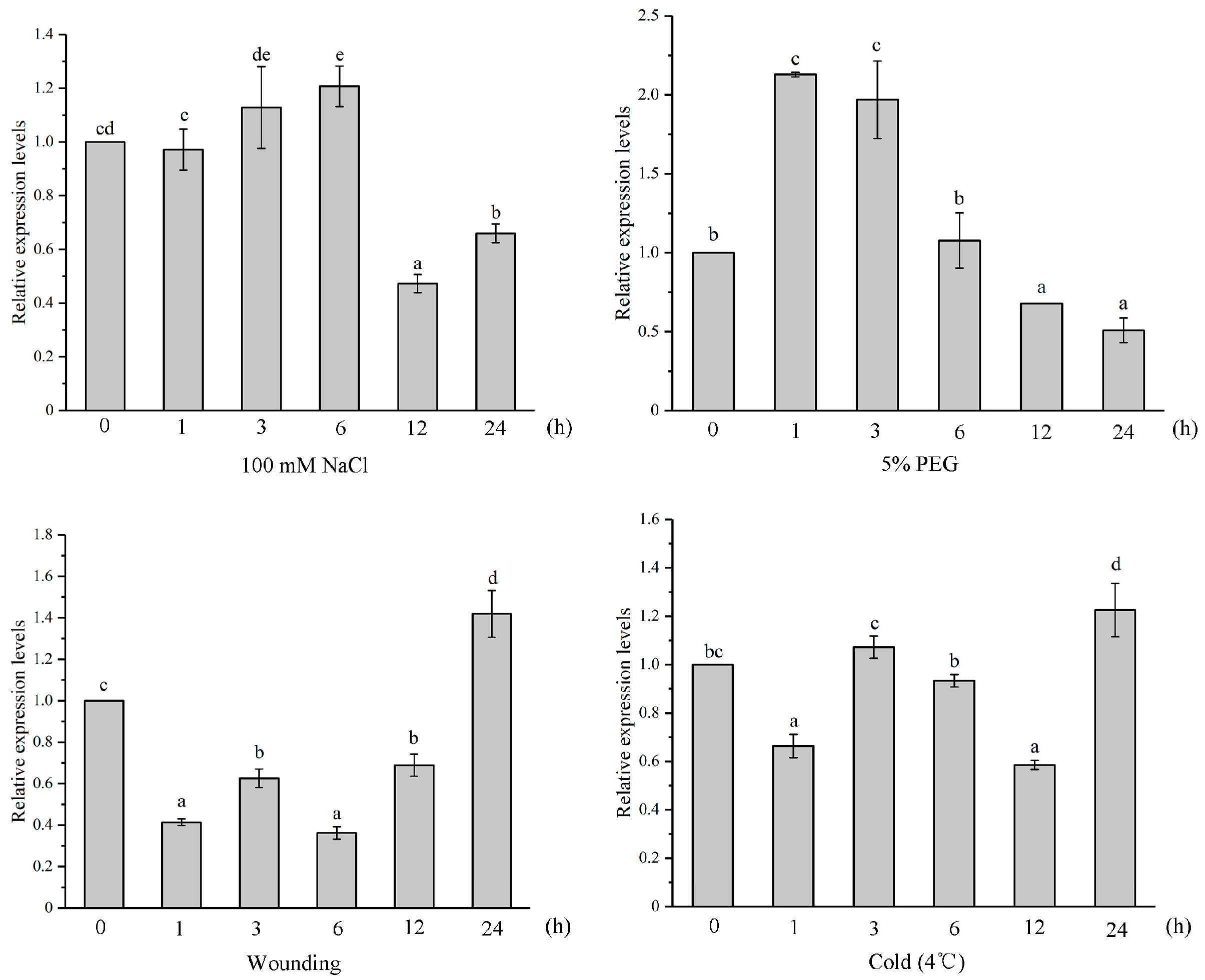
2.4. Expression Pattern Analysis on IiYUCCA Genes
2.5. Heterologous Expression of IiYUCCA6-1 in Nicotiana benthamiana
2.6. IiYUCCA6-1 Could Delay the Senescence Process
3. Discussion
3.1. YUCCA Numbers, Structures, and Phylogenetic Analysis in I. indigotica
3.2. YUCCA Expression Patterns in I. indigotica
3.3. Functional Analysis on IiYUCCA6-1 Ectopically Expressed in Tobacco
4. Materials and Methods
4.1. Isolation and Sequence Analysis of YUCCA Gene Family in I. indigotica
4.2. Plant Materials and Treatments
4.3. Gene Expression Analysis
4.4. Generation of Transgenic N. benthamiana
4.5. Heterologous Transformation of IiYUCCA6-1 to N. benthamiana
4.6. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Feng, H.Q.; Li, C. Research advances of auxin signal transduction. Biotechnol. Bull. 2018, 34, 24–30. [Google Scholar]
- Ellis, C.M.; Nagpal, P.; Young, J.C.; Hagen, G.; Guilfoyle, T.J.; Reed, J.W. Auxin response factor1 and auxin response factor2 regulate senescence and floral organ abscission in Arabidopsis thaliana. Development 2005, 132, 4563–4574. [Google Scholar] [CrossRef] [PubMed]
- Péret, B.; De Rybel, B.; Casimiro, I.; Benková, E.; Swarup, R.; Laplaze, L.; Beeckman, T.; Bennett, M.J. Arabidopsis lateral root development: An emerging story. Trends Plant Sci. 2009, 14, 399–408. [Google Scholar] [CrossRef] [PubMed]
- Woodward, A.W. Auxin: Regulation, action, and interaction. Ann. Bot. 2005, 95, 707–735. [Google Scholar] [CrossRef] [PubMed]
- Andrea, G. The role of auxin in shaping shoot architecture. J. Exp. Bot. 2013, 64, 2593–2608. [Google Scholar]
- Robert, H.S.; Park, C.; Gutièrrez, C.L.; Wójcikowska, B.; Pěnčík, A.; Novák, O.; Chen, J.; Grunewald, W.; Dresselhaus, T.; Friml, J.; et al. Maternal auxin supply contributes to early embryo patterning in Arabidopsis. Nat. Plants 2018, 4, 548–553. [Google Scholar] [CrossRef]
- Yuan, H.Z.; Zhao, M.Z.; Wu, W.M.; Yu, H.M.; Qian, Y.M.; Wang, Z.W.; Wang, X.C. Genome-wide identification and expression analysis of auxin-related gene families in grape. Yi Chuan 2015, 37, 720–730. [Google Scholar]
- Wang, S.; Bai, Y.; Shen, C.; Wu, Y.; Zhang, S.; Jiang, D.; Guilfoyle, T.J.; Chen, M.; Qi, Y. Auxin-related gene families in abiotic stress response in Sorghum bicolor. Funct. Integr. Genomics 2010, 10, 533–546. [Google Scholar] [CrossRef]
- Jain, M.; Khurana, J.P. Transcript profiling reveals diverse roles of auxin-responsive genes during reproductive development and abiotic stress in rice. FEBS J. 2009, 276, 3148–3162. [Google Scholar] [CrossRef]
- Zhang, S.W.; Li, C.H.; Cao, J.; Zhang, Y.C.; Zhang, S.Q.; Xia, Y.F.; Sun, D.Y.; Sun, Y. Altered architecture and enhanced drought tolerance in rice via the down-regulation of indole-3-acetic acid by TLD1/OsGH3.13 activation. Plant Physiol. 2009, 151, 1889–1901. [Google Scholar] [CrossRef]
- Shibasaki, K.; Uemura, M.; Tsurumi, S.; Rahman, A. Auxin response in Arabidopsis under cold stress: Underlying molecular mechanisms. Plant Cell 2009, 21, 3823–3838. [Google Scholar] [CrossRef] [PubMed]
- Mashiguchi, K.; Tanaka, K.; Sakai, T.; Sugawara, S.; Kawaide, H.; Natsume, M.; Hanada, A.; Yaeno, T.; Shirasu, K.; Yao, H.; et al. The main auxin biosynthesis pathway in Arabidopsis. Proc. Natl. Acad. Sci. USA 2011, 108, 18512–18517. [Google Scholar] [CrossRef] [PubMed]
- Cheng, Y.; Dai, X.; Zhao, Y. Auxin biosynthesis by the YUCCA flavin monooxygenases controls the formation of floral organs and vascular tissues in Arabidopsis. Genes Dev. 2006, 20, 1790–1799. [Google Scholar] [CrossRef] [PubMed]
- Stepanova, A.N.; Robertson-Hoyt, J.; Yun, J.; Benavente, L.M.; Xie, D.Y.; Doležal, K.; Schlereth, A.; Jürgens, G.; Alonso, J.M. TAA1-mediated auxin biosynthesis is essential for hormone crosstalk and plant development. Cell 2008, 133, 177–191. [Google Scholar] [CrossRef]
- Won, C.; Shen, X.; Mashiguchi, K.; Zheng, Z.; Dai, X.; Cheng, Y.; Kasahara, H.; Kamiya, Y.; Chory, J.; Zhao, Y. Conversion of tryptophan to indole-3-acetic acid by tryptophan aminotransferases of Arabidopsis and YUCCAs in Arabidopsis. Proc. Natl. Acad. Sci. USA 2011, 108, 18518–18523. [Google Scholar] [CrossRef]
- Kim, J.I.; Murphy, A.S.; Baek, D.; Lee, S.W.; Yun, D.; Bressan, R.A.; Narasimhan, M.L. YUCCA6 over-expression demonstrates auxin function in delaying leaf senescence in Arabidopsis thaliana. J. Exp. Bot. 2011, 62, 3981–3992. [Google Scholar] [CrossRef]
- Lee, M.; Jung, J.H.; Han, D.Y.; Seo, P.J.; Park, W.J.; Park, C.M. Activation of a flavin monooxygenase gene YUCCA7 enhances drought resistance in Arabidopsis. Planta 2012, 235, 923–938. [Google Scholar] [CrossRef]
- Li, L.; Ljung, K.; Breton, G.; Schmitz, R.J.; Pruneda-Paz, J.; Cowing-Zitron, C.; Cole, B.J.; Ivans, L.J.; Pedmale, U.V.; Jung, H.S.; et al. Linking photoreceptor excitation to changes in plant architecture. Genes Dev. 2012, 26, 785–790. [Google Scholar] [CrossRef]
- Woodward, C.; Bemis, S.M.; Hill, E.J.; Sawa, S.; Koshiba, T.; Torii, K.U. Interaction of auxin and erecta in elaborating Arabidopsis inflorescence architecture revealed by the activation tagging of a new member of the YUCCA family putative flavin monooxygenases. Plant Physiol. 2005, 139, 192–203. [Google Scholar] [CrossRef]
- Kim, J.I.; Sharkhuu, A.; Jin, J.B.; Li, P.; Jeong, J.C.; Baek, D.; Lee, S.Y.; Blakeslee, J.J.; Murphy, A.S.; Bohnert, H.J.; et al. yucca6, a dominant mutation in Arabidopsis, affects auxin accumulation and auxin-related phenotypes. Plant Physiol. 2007, 145, 722–735. [Google Scholar] [CrossRef]
- Park, S.C.; Kim, H.S.; Lee, H.U.; Kim, Y.H.; Kwak, S.S. Overexpression of Arabidopsis YUCCA6 enhances environment stress tolerance and inhibits storage root formation in sweet potato. Plant Biotechnol. Rep. 2019, 13, 345–352. [Google Scholar] [CrossRef]
- Nana, L. Effect of overexpression of IAA synthetic gene YUCCA1 on the ripening process of strawberry fruit. J. Xinjiang Agric. Univ. 2018, 41, 267–275. [Google Scholar]
- Ye, X.; Kang, B.G.; Osburn, L.D.; Li, Y.; Max, C.Z.M. Identification of the flavin-dependent monooxygenase-encoding YUCCA gene family in Populus trichocarpa and their expression in vegetative tissues and in response to hormone and environmental stresses. Plant Cell Tiss. Organ Cult. 2009, 97, 271–283. [Google Scholar] [CrossRef]
- Wang, Y.; Liu, H.; Wang, S.; Li, H. Genome-wide identification and expression analysis of the YUCCA gene family in soybean (Glycine max L.). Plant Growth Regul. 2016, 81, 265–275. [Google Scholar] [CrossRef]
- Qian, Z.Q.; Wei, T.S.; Jie, Z.; Ping, W.Y.; He, S.H.; Yi, R.; Gui, G.S.; Yi, G.G.; Yong, X.; Ying, Z.H. Identification of YUCCA gene family and expression analysis during watermelon fruit ripening process. China Veg. 2019, 3, 21–29. [Google Scholar]
- Yamamoto, Y.; Kamiya, N.; Morinaka, Y.; Matsuoka, M.; Sazuka, T. Auxin biosynthesis by the YUCCA genes in rice. Plant Physiol. 2007, 143, 1362–1371. [Google Scholar] [CrossRef]
- Li, W.; Zhao, X.; Zhang, X. Genome-wide analysis and expression patterns of the YUCCA genes in Maize. J. Genet. Genom. 2015, 42, 707–710. [Google Scholar] [CrossRef]
- Li, N.; Yin, N.; Niu, Z.; Hui, W.; Song, J.; Huang, C.; Wang, H.; Kong, L.; Feng, D. Isolation and characterization of three TaYUC10 genes from wheat. Gene 2014, 546, 187–194. [Google Scholar] [CrossRef]
- Yan, S.; Che, G.; Ding, L.; Chen, Z.; Liu, X.; Wang, H.; Zhao, W.; Ning, K.; Zhao, J.; Tesfamichael, K.; et al. Different cucumber CsYUC genes regulate response to abiotic stresses and flower development. Sci. Rep. 2016, 6, 20760. [Google Scholar] [CrossRef]
- Poulet, A.; Kriechbaumer, V. Bioinformatics analysis of phylogeny and transcription of TAA/YUC auxin biosynthetic genes. Int. J. Mol. Sci. 2017, 18, 1791. [Google Scholar] [CrossRef]
- Liu, H.; Xie, W.F.; Zhang, L.; Victoriano, V.; Ye, Z.W.; Gao, Q.H.; Duan, K. Auxin biosynthesis by the YUCCA6 flavin monooxygenase gene in woodland strawberry. J. Integr. Plant Biol. 2014, 56, 350–363. [Google Scholar]
- Cao, X.; Yang, H.L.; Shang, C.Q.; Ma, S.; Liu, L.; Cheng, J.L. The roles of auxin biosynthesis YUCCA gene family in plants. Int. J. Mol. Sci. 2019, 20, 6343. [Google Scholar] [CrossRef] [PubMed]
- Xiao, S.S.; Jin, Y.; Sun, Y.Q. Recent progress in the studies of chemical con-stituents, pharmacological effects and quality control methods on the roots of Isatis indigotica. J. Shenyang Pharm. Univ. 2003, 20, 455–459. [Google Scholar]
- Kang, M.; Wu, H.; Yang, Q.; Huang, L.; Hu, Q.; Ma, T.; Li, Z.; Liu, J. A chromosome-scale genome assembly of Isatis indigotica, an important medicinal plant used in traditional Chinese medicine: An Isatis genome. Hortic. Res. 2020, 7, 18. [Google Scholar] [CrossRef]
- Hurst, L.D. The Ka/Ks ratio: Diagnosing the form of sequence evolution. Trends Genet. 2002, 18, 486. [Google Scholar] [CrossRef]
- Tobena-Santamaria, R. FLOOZY of petunia is a flavin mono-oxygenase-like protein required for the specification of leaf and flower architecture. Genes Dev. 2002, 16, 753–763. [Google Scholar] [CrossRef]
- Zhao, Y.; Christensen, S.K.; Fankhauser, C.; Cashman, J.R.; Cohen, J.D.; Weigel, D.; Chory, J. A role for flavin monooxygenase-like enzymes in auxin biosynthesis. Science 2001, 291, 306–309. [Google Scholar] [CrossRef]
- Gallavotti, A.; Barazesh, S.; Malcomber, S.; Hall, D.; Jackson, D.; Schmidt, R.J.; McSteen, P. Sparse inflorescence1 encodes a monocot-specific YUCCA-like gene required for vegetative and reproductive development in maize. Proc. Natl. Acad. Sci. USA 2008, 105, 15196–15201. [Google Scholar] [CrossRef]
- Exposito-Rodriguez, M.; Borges, A.A.; Borges-Perez, A.; Hernandez, M.; Perez, J.A. Cloning and biochemical characterization of ToFZY, a tomato gene encoding a flavin monooxygenase involved in a tryptophan-dependent auxin biosynthesis pathway. J. Plant Growth Regul. 2007, 26, 329–340. [Google Scholar] [CrossRef]
- Exposito-Rodriguez, M.; Borges, A.A.; Borges-Perez, A.; Perez, J.A. Gene structure and spatiotemporal expression profile of tomato genes encoding YUCCA-like flavin monooxygenases: The ToFZY gene family. Plant Physiol. Biochem. 2011, 49, 782–791. [Google Scholar] [CrossRef]
- Woo, Y.M.; Park, H.J.; Su’udi, M.; Yang, J.I.; Park, J.J.; Back, K.; Park, Y.M.; An, G. Constitutively wilted 1, a member of the rice YUCCA gene family, is required for maintaining water homeostasis and an appropriate root to shoot ratio. Plant Mol. Biol. 2007, 65, 125–136. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.; Ying, Y.Y.; Zhang, L.; Gao, Q.H.; Li, J.; Zhang, Z.; Fang, J.G.; Duan, K. Isolation and characterization of two YUCCA flavin monooxygenase genes from cultivated strawberry (Fragaria × ananassa Duch.). Plant Cell Rep. 2012, 31, 1425–1435. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.I.; Baek, D.; Park, H.C.; Chun, H.J.; Oh, D.H.; Lee, M.K.; Cha, J.Y.; Kim, W.Y.; Kim, M.C.; Chung, W.S.; et al. Overexpression of Arabidopsis YUCCA6 in potato results in high-auxin developmental phenotypes and enhanced resistance to water deficit. Mol. Plant 2013, 6, 337–349. [Google Scholar] [CrossRef] [PubMed]
- Li, T.; Wang, J.; Lu, M.; Zhang, T.; Qu, X.; Wang, Z. Selection and validation of appropriate reference genes for qRT-PCR analysis in Isatis indigotica Fort. Front. Plant Sci. 2017, 8, 1139. [Google Scholar] [CrossRef]






| Gene Name | Gene ID | Chromosome Location | CDS Length (bp) | Exons | Protein Length (aa) | MW (Da) | pI | GRAVY | Instability Index |
|---|---|---|---|---|---|---|---|---|---|
| IiYUCCA1 | EVM0033712 | Chr06: 30576212–30578748 | 1317 | 5 | 438 | 48,658.38 | 9.58 | −0.218 | 40.24 |
| IiYUCCA2 | EVM0000868 | Chr03: 15976139–15977849 | 1248 | 4 | 415 | 46,528.12 | 8.36 | −0.116 | 45.62 |
| IiYUCCA3 | EVM0030530 | Chr06: 30914630–30916362 | 1269 | 4 | 422 | 47,162.38 | 9.15 | −0.243 | 35.7 |
| IiYUCCA4 | EVM0029692 | Chr02: 3979445–3983464 | 1236 | 5 | 411 | 45,732.06 | 9.39 | −0.123 | 33.44 |
| IiYUCCA5-1 | EVM0003921 | Chr01: 14186499–14187976 | 1287 | 3 | 428 | 47,701.11 | 9 | −0.18 | 51.45 |
| IiYUCCA5-2 | EVM0010459 | Chr01: 14011394–14012852 | 1203 | 3 | 400 | 44,461.14 | 8.91 | −0.255 | 50.61 |
| IiYUCCA6-1 | EVM0022538 | contig1043: 115626–118449 | 1335 | 4 | 444 | 49,773.53 | 8.71 | −0.201 | 39.95 |
| IiYUCCA6-2 | Pseudogene492 | contig1042: 131146–133992 | 456 | - | - | - | - | - | - |
| IiYUCCA7 | EVM0024488 | Chr07: 21875662–21877324 | 1311 | 3 | 436 | 48,814.38 | 9.07 | −0.207 | 42.26 |
| IiYUCCA8 | EVM0005748 | Chr03: 28640430–28641928 | 1287 | 3 | 428 | 48,092.61 | 8.95 | −0.225 | 48.86 |
| IiYUCCA9 | EVM0000072 | Chr06: 31112478–31113966 | 1245 | 3 | 414 | 46,277.41 | 9.32 | −0.243 | 47.12 |
| IiYUCCA10 | EVM0002314 | Chr06: 3442231–3444777 | 1044 | 3 | 347 | 38,643.58 | 8.53 | −0.122 | 26.13 |
| IiYUCCA11 | EVM0031632 | Chr06: 23649486–23650852 | 1176 | 3 | 391 | 43,467.14 | 9.44 | −0.154 | 42.9 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Qin, M.; Wang, J.; Zhang, T.; Hu, X.; Liu, R.; Gao, T.; Zhao, S.; Yuan, Y.; Zheng, J.; Wang, Z.; et al. Genome-Wide Identification and Analysis on YUCCA Gene Family in Isatis indigotica Fort. and IiYUCCA6-1 Functional Exploration. Int. J. Mol. Sci. 2020, 21, 2188. https://doi.org/10.3390/ijms21062188
Qin M, Wang J, Zhang T, Hu X, Liu R, Gao T, Zhao S, Yuan Y, Zheng J, Wang Z, et al. Genome-Wide Identification and Analysis on YUCCA Gene Family in Isatis indigotica Fort. and IiYUCCA6-1 Functional Exploration. International Journal of Molecular Sciences. 2020; 21(6):2188. https://doi.org/10.3390/ijms21062188
Chicago/Turabian StyleQin, Miaomiao, Jing Wang, Tianyi Zhang, Xiangyang Hu, Rui Liu, Tian’e Gao, Shuaijing Zhao, Yilin Yuan, Jinyu Zheng, Zirong Wang, and et al. 2020. "Genome-Wide Identification and Analysis on YUCCA Gene Family in Isatis indigotica Fort. and IiYUCCA6-1 Functional Exploration" International Journal of Molecular Sciences 21, no. 6: 2188. https://doi.org/10.3390/ijms21062188
APA StyleQin, M., Wang, J., Zhang, T., Hu, X., Liu, R., Gao, T., Zhao, S., Yuan, Y., Zheng, J., Wang, Z., Wei, X., & Li, T. (2020). Genome-Wide Identification and Analysis on YUCCA Gene Family in Isatis indigotica Fort. and IiYUCCA6-1 Functional Exploration. International Journal of Molecular Sciences, 21(6), 2188. https://doi.org/10.3390/ijms21062188




