Tumor-Cell–Macrophage Fusion Cells as Liquid Biomarkers and Tumor Enhancers in Cancer
Abstract
1. Introduction
2. Cell Fusion as a Physiological and Pathological Mechanism
3. Genetic Evidence for Presence of Fusion Cells in Cancer Patients
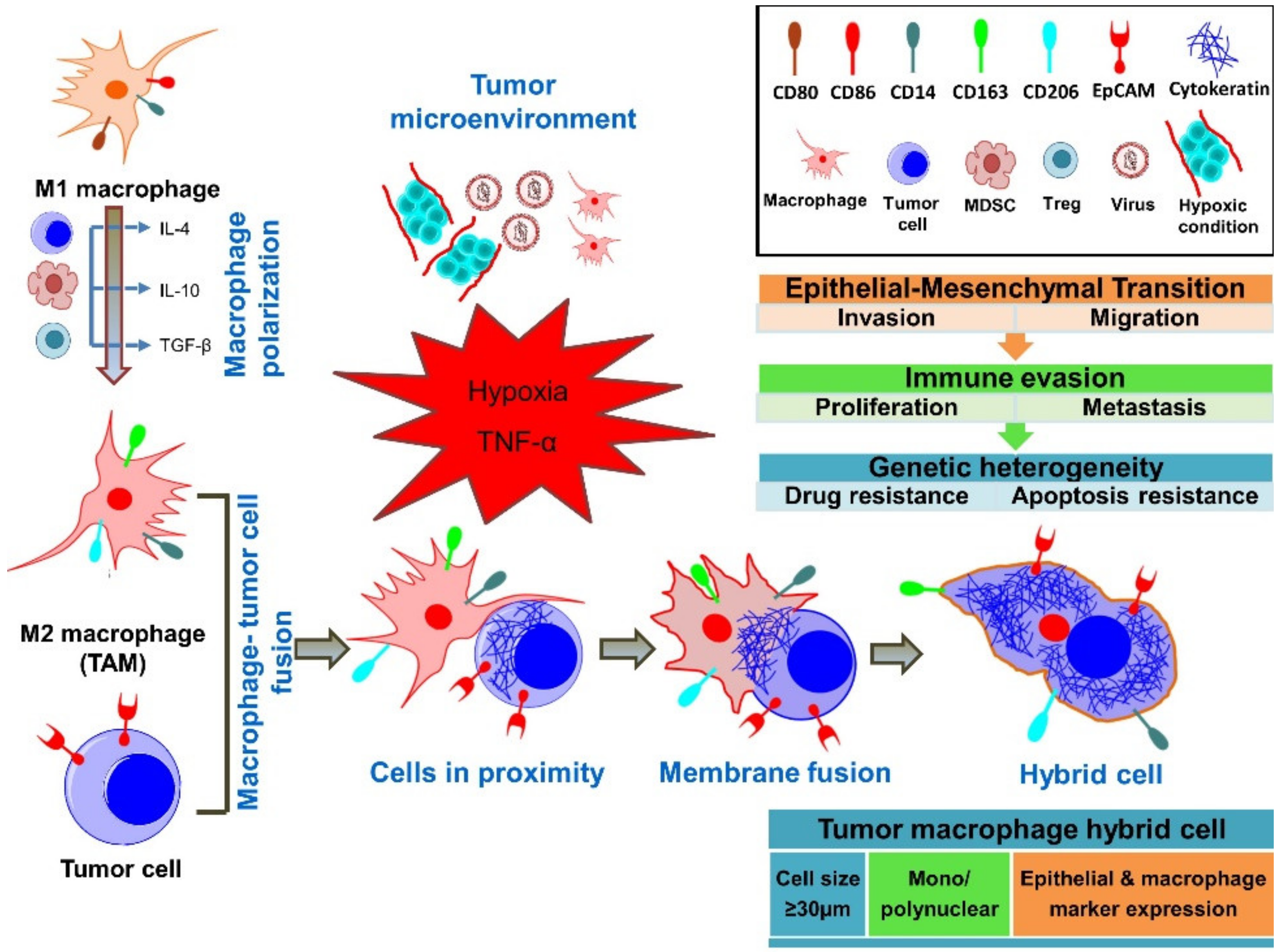
4. Fusion of Tumor Cells with Macrophages
5. Fusion Cell Detection in the Peripheral Blood of Cancer Patients
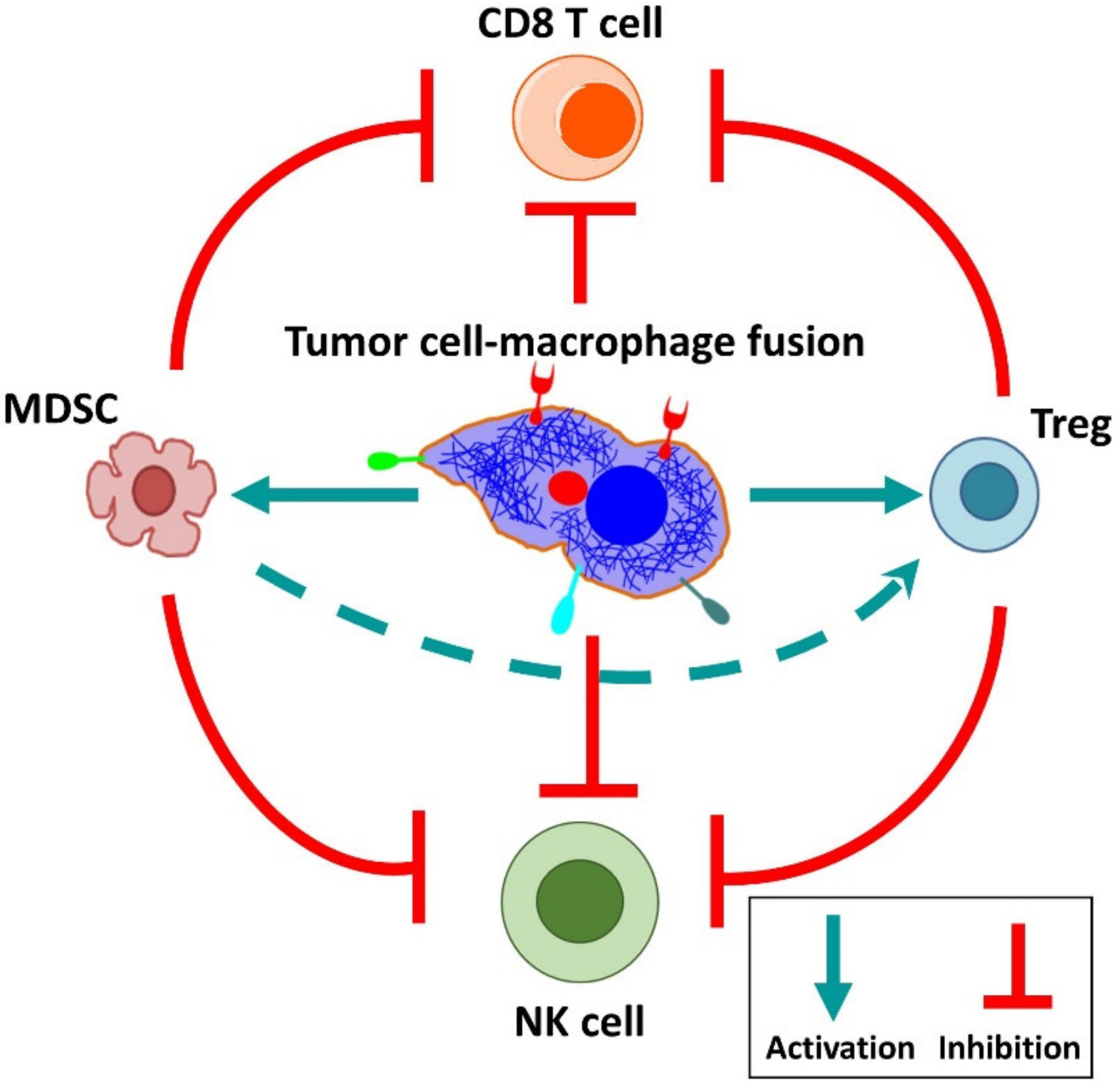
6. Cancer-Associated Properties of Fusion Cells
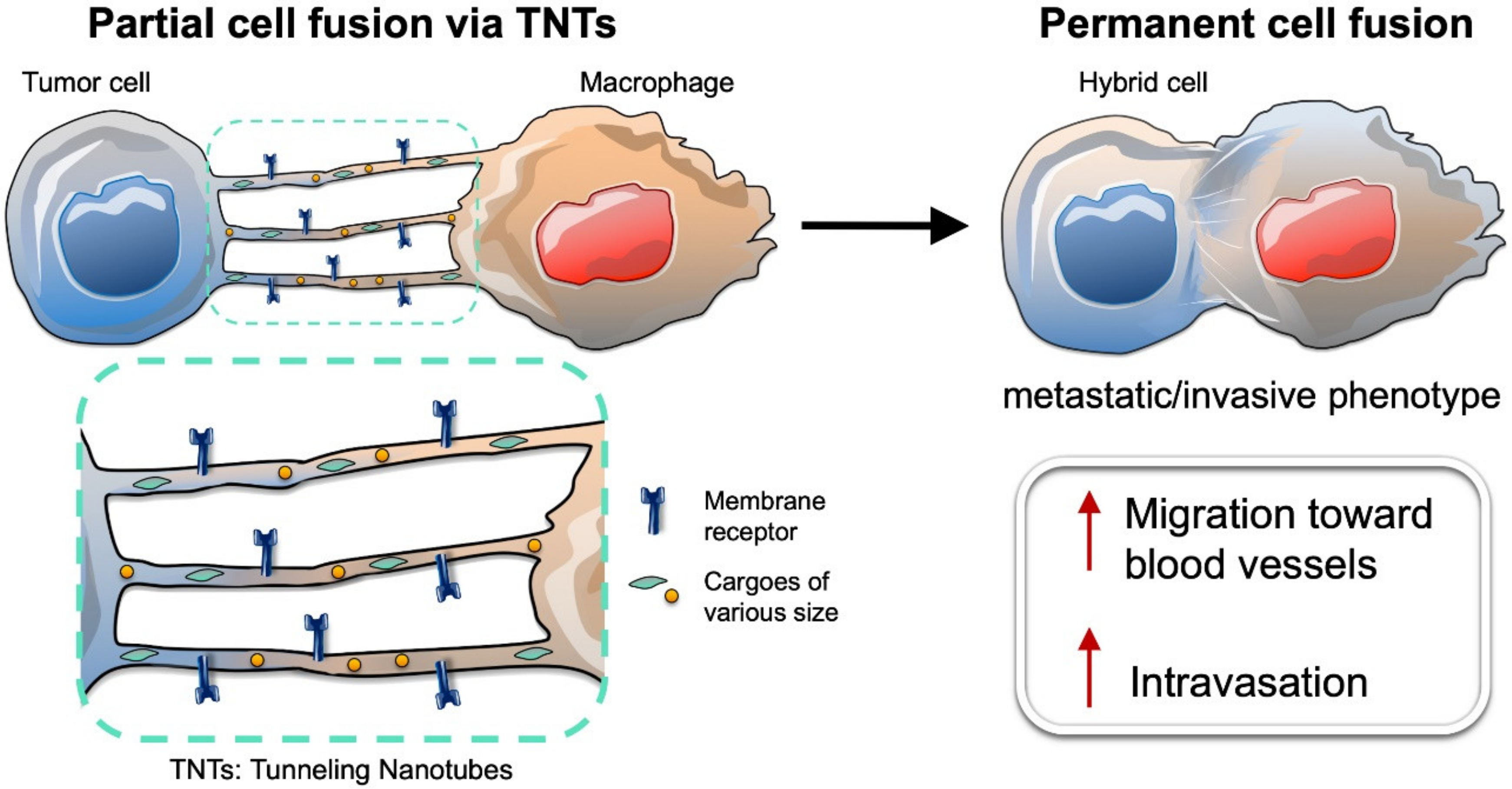
7. Intercellular Connections between Tumor Cells and Macrophages Lead to Partial Cell Fusion, and Favor Cell Migration and Invasion
8. Tumor Cell Fusion Leads to Tumor Heterogeneity and Chemoresistance
9. Summary and Perspective for Precision Medicine on Fusion Cells in Cancer
Author Contributions
Funding
Conflicts of Interest
References
- Stephens, P.J.; Tarpey, P.S.; Davies, H.; Van Loo, P.; Greenman, C.; Wedge, D.C.; Nik-Zainal, S.; Martin, S.; Varela, I.; Bignell, G.R.; et al. The landscape of cancer genes and mutational processes in breast cancer. Nature 2012, 486, 400–404. [Google Scholar]
- Pantel, K.; Alix-Panabieres, C. Liquid biopsy and minimal residual disease—Latest advances and implications for cure. Nat. Rev. Clin. Oncol. 2019, 16, 409–424. [Google Scholar]
- Racila, E.; Euhus, D.; Weiss, A.J.; Rao, C.; McConnell, J.; Terstappen, L.W.; Uhr, J.W. Detection and characterization of carcinoma cells in the blood. Proc. Natl. Acad. Sci. USA 1998, 95, 4589–4594. [Google Scholar]
- De Wit, S.; van Dalum, G.; Lenferink, A.T.; Tibbe, A.G.; Hiltermann, T.J.; Groen, H.J.; van Rijn, C.J.; Terstappen, L.W. The detection of EpCAM(+) and EpCAM(-) circulating tumor cells. Sci. Rep. 2015, 5, 12270. [Google Scholar]
- Kaifi, J.T.; Li, G.; Clawson, G.; Kimchi, E.T.; Staveley-O’Carroll, K.F. Perioperative circulating tumor cell detection: Current perspectives. Cancer Biol. Ther. 2016, 17, 859–869. [Google Scholar]
- Yu, M.; Bardia, A.; Wittner, B.S.; Stott, S.L.; Smas, M.E.; Ting, D.T.; Isakoff, S.J.; Ciciliano, J.C.; Wells, M.N.; Shah, A.M.; et al. Circulating breast tumor cells exhibit dynamic changes in epithelial and mesenchymal composition. Science 2013, 339, 580–584. [Google Scholar]
- Manjunath, Y.; Upparahalli, S.V.; Avella, D.M.; Deroche, C.B.; Kimchi, E.T.; Staveley-O’Carroll, K.F.; Smith, C.J.; Li, G.; Kaifi, J.T. PD-L1 Expression with Epithelial Mesenchymal Transition of Circulating Tumor Cells Is Associated with Poor Survival in Curatively Resected Non-Small Cell Lung Cancer. Cancers (Basel) 2019, 11, 806. [Google Scholar]
- Harouaka, R.A.; Zhou, M.D.; Yeh, Y.T.; Khan, W.J.; Das, A.; Liu, X.; Christ, C.C.; Dicker, D.T.; Baney, T.S.; Kaifi, J.T.; et al. Flexible micro spring array device for high-throughput enrichment of viable circulating tumor cells. Clin. Chem. 2014, 60, 323–333. [Google Scholar]
- Aichel, O. Ueber Zellverschmelzung mit qualitativ abnormer Chromosomenverteilung als Ursache der Geschwulstbildung. Heft XIII Herausgegebenen von Wilhelm Roux (Verlag von Wilhelm Engelmann, Leipzig) 1911, XIII, 92–111. [Google Scholar]
- Fernandes, C.; Prabhu, P.; Juvale, K.; Suares, D.; Yc, M. Cancer cell fusion: A potential target to tackle drug-resistant and metastatic cancer cells. Drug Discov. Today 2019, 24, 1836–1844. [Google Scholar]
- Laberge, G.S.; Duvall, E.; Haedicke, K.; Pawelek, J. Leukocyte(-)Cancer Cell Fusion-Genesis of a Deadly Journey. Cells 2019, 8, 170. [Google Scholar]
- Pawelek, J.M.; Chakraborty, A.K. The cancer cell—Leukocyte fusion theory of metastasis. Adv. Cancer Res. 2008, 101, 397–444. [Google Scholar]
- Noubissi, F.K.; Harkness, T.; Alexander, C.M.; Ogle, B.M. Apoptosis-induced cancer cell fusion: A mechanism of breast cancer metastasis. FASEB J. 2015, 29, 4036–4045. [Google Scholar]
- Kemeny, L.V.; Kurgyis, Z.; Buknicz, T.; Groma, G.; Jakab, A.; Zanker, K.; Dittmar, T.; Kemeny, L.; Nemeth, I.B. Melanoma Cells Can Adopt the Phenotype of Stromal Fibroblasts and Macrophages by Spontaneous Cell Fusion in Vitro. Int. J. Mol. Sci. 2016, 17, 826. [Google Scholar]
- Clawson, G.A. Cancer. Fusion for moving. Science 2013, 342, 699–700. [Google Scholar]
- Ramakrishnan, M.; Mathur, S.R.; Mukhopadhyay, A. Fusion-derived epithelial cancer cells express hematopoietic markers and contribute to stem cell and migratory phenotype in ovarian carcinoma. Cancer Res. 2013, 73, 5360–5370. [Google Scholar]
- Lazova, R.; Laberge, G.S.; Duvall, E.; Spoelstra, N.; Klump, V.; Sznol, M.; Cooper, D.; Spritz, R.A.; Chang, J.T.; Pawelek, J.M. A Melanoma Brain Metastasis with a Donor-Patient Hybrid Genome following Bone Marrow Transplantation: First Evidence for Fusion in Human Cancer. PLoS ONE 2013, 8, e66731. [Google Scholar]
- LaBerge, G.S.; Duvall, E.; Grasmick, Z.; Haedicke, K.; Pawelek, J. A Melanoma Lymph Node Metastasis with a Donor-Patient Hybrid Genome following Bone Marrow Transplantation: A Second Case of Leucocyte-Tumor Cell Hybridization in Cancer Metastasis. PLoS ONE 2017, 12, e0168581. [Google Scholar]
- Duelli, D.M.; Padilla-Nash, H.M.; Berman, D.; Murphy, K.M.; Ried, T.; Lazebnik, Y. A virus causes cancer by inducing massive chromosomal instability through cell fusion. Curr. Biol. 2007, 17, 431–437. [Google Scholar]
- Shabo, I.; Midtbo, K.; Andersson, H.; Akerlund, E.; Olsson, H.; Wegman, P.; Gunnarsson, C.; Lindstrom, A. Macrophage traits in cancer cells are induced by macrophage-cancer cell fusion and cannot be explained by cellular interaction. BMC Cancer 2015, 15, 922. [Google Scholar]
- Clawson, G.A.; Matters, G.L.; Xin, P.; McGovern, C.; Wafula, E.; dePamphilis, C.; Meckley, M.; Wong, J.; Stewart, L.; D’Jamoos, C.; et al. “Stealth dissemination” of macrophage-tumor cell fusions cultured from blood of patients with pancreatic ductal adenocarcinoma. PLoS ONE 2017, 12, e0184451. [Google Scholar]
- Clawson, G.A.; Matters, G.L.; Xin, P.; Imamura-Kawasawa, Y.; Du, Z.; Thiboutot, D.M.; Helm, K.F.; Neves, R.I.; Abraham, T. Macrophage-tumor cell fusions from peripheral blood of melanoma patients. PLoS ONE 2015, 10, e0134320. [Google Scholar]
- Aceto, N.; Bardia, A.; Miyamoto, D.T.; Donaldson, M.C.; Wittner, B.S.; Spencer, J.A.; Yu, M.; Pely, A.; Engstrom, A.; Zhu, H.; et al. Circulating tumor cell clusters are oligoclonal precursors of breast cancer metastasis. Cell 2014, 158, 1110–1122. [Google Scholar]
- Scholtens, T.M.; Schreuder, F.; Ligthart, S.T.; Swennenhuis, J.F.; Greve, J.; Terstappen, L.W. Automated identification of circulating tumor cells by image cytometry. Cytometry A 2012, 81, 138–148. [Google Scholar]
- Kasimir-Bauer, S.; Hoffmann, O.; Wallwiener, D.; Kimmig, R.; Fehm, T. Expression of stem cell and epithelial–mesenchymal transition markers in primary breast cancer patients with circulating tumor cells. Breast Cancer Res. 2012, 14, R15. [Google Scholar]
- Iinuma, H.; Watanabe, T.; Mimori, K.; Adachi, M.; Hayashi, N.; Tamura, J.; Matsuda, K.; Fukushima, R.; Okinaga, K.; Sasako, M.; et al. Clinical significance of circulating tumor cells, including cancer stem-like cells, in peripheral blood for recurrence and prognosis in patients with Dukes’ stage B and C colorectal cancer. J. Clin. Oncol. 2011, 29, 1547–1555. [Google Scholar]
- Clawson, G.A.; Kimchi, E.; Patrick, S.D.; Xin, P.; Harouaka, R.; Zheng, S.; Berg, A.; Schell, T.; Staveley-O’Carroll, K.F.; Neves, R.I.; et al. Circulating tumor cells in melanoma patients. PLoS ONE 2012, 7, e41052. [Google Scholar]
- Kaifi, J.T.; Kunkel, M.; Das, A.; Harouaka, R.A.; Dicker, D.T.; Li, G.; Zhu, J.; Clawson, G.A.; Yang, Z.; Reed, M.F.; et al. Circulating tumor cell isolation during resection of colorectal cancer lung and liver metastases: A prospective trial with different detection techniques. Cancer Biol. Ther. 2015, 16, 699–708. [Google Scholar]
- Adams, D.L.; Martin, S.S.; Alpaugh, R.K.; Charpentier, M.; Tsai, S.; Bergan, R.C.; Ogden, I.M.; Catalona, W.; Chumsri, S.; Tang, C.M.; et al. Circulating giant macrophages as a potential biomarker of solid tumors. Proc. Natl. Acad. Sci. USA 2014, 111, 3514–3519. [Google Scholar]
- Kohler, G.; Milstein, C. Continuous cultures of fused cells secreting antibody of predefined specificity. Nature 1975, 256, 495–497. [Google Scholar]
- Aguilar, P.S.; Baylies, M.K.; Fleissner, A.; Helming, L.; Inoue, N.; Podbilewicz, B.; Wang, H.; Wong, M. Genetic basis of cell-cell fusion mechanisms. Trends Genet 2013, 29, 427–437. [Google Scholar]
- Vignery, A. Macrophage fusion: The making of osteoclasts and giant cells. J. Exp. Med. 2005, 202, 337–340. [Google Scholar]
- Dittmar, T.; Nagler, C.; Niggemann, B.; Zanker, K.S. The dark side of stem cells: Triggering cancer progression by cell fusion. Curr. Mol. Med. 2013, 13, 735–750. [Google Scholar]
- Pawelek, J.M. Cancer-cell fusion with migratory bone-marrow-derived cells as an explanation for metastasis: New therapeutic paradigms. Future Oncol. 2008, 4, 449–452. [Google Scholar]
- Mi, S.; Lee, X.; Li, X.; Veldman, G.M.; Finnerty, H.; Racie, L.; LaVallie, E.; Tang, X.Y.; Edouard, P.; Howes, S.; et al. Syncytin is a captive retroviral envelope protein involved in human placental morphogenesis. Nature 2000, 403, 785–789. [Google Scholar]
- Mohler, W.A.; Shemer, G.; del Campo, J.J.; Valansi, C.; Opoku-Serebuoh, E.; Scranton, V.; Assaf, N.; White, J.G.; Podbilewicz, B. The type I membrane protein EFF-1 is essential for developmental cell fusion. Dev. Cell 2002, 2, 355–362. [Google Scholar]
- Lu, X.; Kang, Y. Efficient acquisition of dual metastasis organotropism to bone and lung through stable spontaneous fusion between MDA-MB-231 variants. Proc. Natl. Acad. Sci. USA 2009, 106, 9385–9390. [Google Scholar]
- Miller, F.R.; McInerney, D.; Rogers, C.; Miller, B.E. Spontaneous fusion between metastatic mammary tumor subpopulations. J. Cell Biochem. 1988, 36, 129–136. [Google Scholar]
- Kurgyis, Z.; Kemeny, L.V.; Buknicz, T.; Groma, G.; Olah, J.; Jakab, A.; Polyanka, H.; Zanker, K.; Dittmar, T.; Kemeny, L.; et al. Melanoma-Derived BRAF(V600E) Mutation in Peritumoral Stromal Cells: Implications for in Vivo Cell Fusion. Int. J. Mol. Sci. 2016, 17, 980. [Google Scholar]
- Wang, R.; Chen, S.; Li, C.; Ng, K.T.; Kong, C.W.; Cheng, J.; Cheng, S.H.; Li, R.A.; Lo, C.M.; Man, K.; et al. Fusion with stem cell makes the hepatocellular carcinoma cells similar to liver tumor-initiating cells. BMC Cancer 2016, 16, 56. [Google Scholar]
- Weiler, J.; Dittmar, T. Cell Fusion in Human Cancer: The Dark Matter Hypothesis. Cells 2019, 8, 132. [Google Scholar]
- Teissie, J.; Ramos, C. Correlation between electric field pulse induced long-lived permeabilization and fusogenicity in cell membranes. Biophys. J. 1998, 74, 1889–1898. [Google Scholar]
- Wojcieszyn, J.W.; Schlegel, R.A.; Lumley-Sapanski, K.; Jacobson, K.A. Studies on the mechanism of polyethylene glycol-mediated cell fusion using fluorescent membrane and cytoplasmic probes. J. Cell Biol. 1983, 96, 151–159. [Google Scholar]
- Sakai, H.; Okafuji, I.; Nishikomori, R.; Abe, J.; Izawa, K.; Kambe, N.; Yasumi, T.; Nakahata, T.; Heike, T. The CD40-CD40L axis and IFN-gamma play critical roles in Langhans giant cell formation. Int. Immunol. 2012, 24, 5–15. [Google Scholar]
- Hochreiter-Hufford, A.E.; Lee, C.S.; Kinchen, J.M.; Sokolowski, J.D.; Arandjelovic, S.; Call, J.A.; Klibanov, A.L.; Yan, Z.; Mandell, J.W.; Ravichandran, K.S. Phosphatidylserine receptor BAI1 and apoptotic cells as new promoters of myoblast fusion. Nature 2013, 497, 263–267. [Google Scholar]
- Lu, M.; Ravichandran, K.S. Dock180-ELMO cooperation in Rac activation. Methods Enzymol. 2006, 406, 388–402. [Google Scholar]
- Pajcini, K.V.; Pomerantz, J.H.; Alkan, O.; Doyonnas, R.; Blau, H.M. Myoblasts and macrophages share molecular components that contribute to cell-cell fusion. J. Cell Biol. 2008, 180, 1005–1019. [Google Scholar]
- Kim, J.H.; Ren, Y.; Ng, W.P.; Li, S.; Son, S.; Kee, Y.S.; Zhang, S.; Zhang, G.; Fletcher, D.A.; Robinson, D.N.; et al. Mechanical tension drives cell membrane fusion. Dev. Cell 2015, 32, 561–573. [Google Scholar]
- Berndt, B.; Zanker, K.S.; Dittmar, T. Cell fusion is a potent inducer of aneuploidy and drug resistance in tumor cell/normal cell hybrids. Crit. Rev. Oncog. 2013, 18, 97–113. [Google Scholar]
- Mortensen, K.; Lichtenberg, J.; Thomsen, P.D.; Larsson, L.I. Spontaneous fusion between cancer cells and endothelial cells. Cell Mol. Life Sci. 2004, 61, 2125–2131. [Google Scholar]
- Xu, M.H.; Gao, X.; Luo, D.; Zhou, X.D.; Xiong, W.; Liu, G.X. EMT and acquisition of stem cell-like properties are involved in spontaneous formation of tumorigenic hybrids between lung cancer and bone marrow-derived mesenchymal stem cells. PLoS ONE 2014, 9, e87893. [Google Scholar]
- Li, H.; Feng, Z.; Tsang, T.C.; Tang, T.; Jia, X.; He, X.; Pennington, M.E.; Badowski, M.S.; Liu, A.K.; Chen, D.; et al. Fusion of HepG2 cells with mesenchymal stem cells increases cancerassociated and malignant properties: An in vivo metastasis model. Oncol Rep. 2014, 32, 539–547. [Google Scholar]
- Yilmaz, Y.; Lazova, R.; Qumsiyeh, M.; Cooper, D.; Pawelek, J. Donor Y chromosome in renal carcinoma cells of a female BMT recipient: Visualization of putative BMT-tumor hybrids by FISH. Bone Marrow Transplant 2005, 35, 1021–1024. [Google Scholar]
- Chakraborty, A.; Lazova, R.; Davies, S.; Backvall, H.; Ponten, F.; Brash, D.; Pawelek, J. Donor DNA in a renal cell carcinoma metastasis from a bone marrow transplant recipient. Bone Marrow Transplant 2004, 34, 183–186. [Google Scholar]
- Gast, C.E.; Silk, A.D.; Zarour, L.; Riegler, L.; Burkhart, J.G.; Gustafson, K.T.; Parappilly, M.S.; Roh-Johnson, M.; Goodman, J.R.; Olson, B.; et al. Cell fusion potentiates tumor heterogeneity and reveals circulating hybrid cells that correlate with stage and survival. Sci. Adv. 2018, 4, eaat7828. [Google Scholar]
- Rappa, G.; Mercapide, J.; Lorico, A. Spontaneous formation of tumorigenic hybrids between breast cancer and multipotent stromal cells is a source of tumor heterogeneity. Am J. Pathol. 2012, 180, 2504–2515. [Google Scholar]
- Najafi, M.; Hashemi Goradel, N.; Farhood, B.; Salehi, E.; Nashtaei, M.S.; Khanlarkhani, N.; Khezri, Z.; Majidpoor, J.; Abouzaripour, M.; Habibi, M.; et al. Macrophage polarity in cancer: A review. J. Cell Biochem. 2019, 120, 2756–2765. [Google Scholar]
- Ding, J.; Jin, W.; Chen, C.; Shao, Z.; Wu, J. Tumor associated macrophage x cancer cell hybrids may acquire cancer stem cell properties in breast cancer. PLoS ONE 2012, 7, e41942. [Google Scholar]
- Blomberg, O.S.; Spagnuolo, L.; de Visser, K.E. Immune regulation of metastasis: Mechanistic insights and therapeutic opportunities. Dis. Model Mech. 2018. [Google Scholar] [CrossRef]
- Sainz, B., Jr.; Carron, E.; Vallespinos, M.; Machado, H.L. Cancer Stem Cells and Macrophages: Implications in Tumor Biology and Therapeutic Strategies. Mediators Inflamm. 2016, 2016, 9012369. [Google Scholar]
- Svensson, J.; Jenmalm, M.C.; Matussek, A.; Geffers, R.; Berg, G.; Ernerudh, J. Macrophages at the fetal-maternal interface express markers of alternative activation and are induced by M-CSF and IL-10. J. Immunol. 2011, 187, 3671–3682. [Google Scholar]
- Mitchem, J.B.; Brennan, D.J.; Knolhoff, B.L.; Belt, B.A.; Zhu, Y.; Sanford, D.E.; Belaygorod, L.; Carpenter, D.; Collins, L.; Piwnica-Worms, D.; et al. Targeting tumor-infiltrating macrophages decreases tumor-initiating cells, relieves immunosuppression, and improves chemotherapeutic responses. Cancer Res. 2013, 73, 1128–1141. [Google Scholar]
- Kuwada, K.; Kagawa, S.; Yoshida, R.; Sakamoto, S.; Ito, A.; Watanabe, M.; Ieda, T.; Kuroda, S.; Kikuchi, S.; Tazawa, H.; et al. The epithelial-to-mesenchymal transition induced by tumor-associated macrophages confers chemoresistance in peritoneally disseminated pancreatic cancer. J. Exp. Clin. Cancer Res. 2018, 37, 307. [Google Scholar]
- Chen, E.H.; Olson, E.N. Unveiling the mechanisms of cell-cell fusion. Science 2005, 308, 369–373. [Google Scholar]
- Chakraborty, A.K.; Sodi, S.; Rachkovsky, M.; Kolesnikova, N.; Platt, J.T.; Bolognia, J.L.; Pawelek, J.M. A spontaneous murine melanoma lung metastasis comprised of host x tumor hybrids. Cancer Res 2000, 60, 2512–2519. [Google Scholar]
- Shabo, I.; Olsson, H.; Stal, O.; Svanvik, J. Breast cancer expression of DAP12 is associated with skeletal and liver metastases and poor survival. Clin. Breast Cancer 2013, 13, 371–377. [Google Scholar]
- Shabo, I.; Svanvik, J. Expression of macrophage antigens by tumor cells. Adv. Exp. Med. Biol. 2011, 714, 141–150. [Google Scholar]
- Joosse, S.A.; Pantel, K. Biologic challenges in the detection of circulating tumor cells. Cancer Res. 2013, 73, 8–11. [Google Scholar]
- Andree, K.C.; van Dalum, G.; Terstappen, L.W. Challenges in circulating tumor cell detection by the CellSearch system. Mol. Oncol. 2016, 10, 395–407. [Google Scholar]
- Konigsberg, R.; Gneist, M.; Jahn-Kuch, D.; Pfeiler, G.; Hager, G.; Hudec, M.; Dittrich, C.; Zeillinger, R. Circulating tumor cells in metastatic colorectal cancer: Efficacy and feasibility of different enrichment methods. Cancer Lett. 2010, 293, 117–123. [Google Scholar]
- Zhou, L.; Dicker, D.T.; Matthew, E.; El-Deiry, W.S.; Alpaugh, R.K. Circulating tumor cells: Silent predictors of metastasis. F1000Res 2017, 6, F1000. [Google Scholar]
- O’Flaherty, L.; Wikman, H.; Pantel, K. Biology and clinical significance of circulating tumor cell subpopulations in lung cancer. Transl Lung Cancer Res. 2017, 6, 431–443. [Google Scholar]
- Adams, D.L.; Adams, D.K.; Alpaugh, R.K.; Cristofanilli, M.; Martin, S.S.; Chumsri, S.; Tang, C.M.; Marks, J.R. Circulating Cancer-Associated Macrophage-Like Cells Differentiate Malignant Breast Cancer and Benign Breast Conditions. Cancer Epidemiol. Biomarkers Prev. 2016, 25, 1037–1042. [Google Scholar]
- Clawson, G. The Fate of Fusions. Cells 2018, 8, 13. [Google Scholar]
- Adams, D.L.; Stefansson, S.; Haudenschild, C.; Martin, S.S.; Charpentier, M.; Chumsri, S.; Cristofanilli, M.; Tang, C.M.; Alpaugh, R.K. Cytometric characterization of circulating tumor cells captured by microfiltration and their correlation to the CellSearch((R)) CTC test. Cytometry A 2015, 87, 137–144. [Google Scholar]
- Tang, C.M.; Zhu, P.; Li, S.; Makarova, O.V.; Amstutz, P.T.; Adams, D.L. Blood-based biopsies-clinical utility beyond circulating tumor cells. Cytometry A 2018, 93, 1246–1250. [Google Scholar]
- Shabo, I.; Olsson, H.; Sun, X.F.; Svanvik, J. Expression of the macrophage antigen CD163 in rectal cancer cells is associated with early local recurrence and reduced survival time. Int. J. Cancer 2009, 125, 1826–1831. [Google Scholar]
- Andersen, T.L.; Boissy, P.; Sondergaard, T.E.; Kupisiewicz, K.; Plesner, T.; Rasmussen, T.; Haaber, J.; Kolvraa, S.; Delaisse, J.M. Osteoclast nuclei of myeloma patients show chromosome translocations specific for the myeloma cell clone: A new type of cancer-host partnership? J. Pathol. 2007, 211, 10–17. [Google Scholar]
- Mu, Z.; Wang, C.; Ye, Z.; Rossi, G.; Sun, C.; Li, L.; Zhu, Z.; Yang, H.; Cristofanilli, M. Prognostic values of cancer associated macrophage-like cells (CAML) enumeration in metastatic breast cancer. Breast Cancer Res. Treat 2017, 165, 733–741. [Google Scholar]
- Kallergi, G.; Papadaki, M.A.; Politaki, E.; Mavroudis, D.; Georgoulias, V.; Agelaki, S. Epithelial to mesenchymal transition markers expressed in circulating tumour cells of early and metastatic breast cancer patients. Breast Cancer Res. 2011, 13, R59. [Google Scholar]
- Yang, J.; Weinberg, R.A. Epithelial–mesenchymal transition: At the crossroads of development and tumor metastasis. Dev. Cell 2008, 14, 818–829. [Google Scholar]
- Paget, S. The distribution of secondary growths in cancer of the breast. Lancet 1889, 133, 571–573. [Google Scholar]
- Fidler, I.J. The pathogenesis of cancer metastasis: The ‘seed and soil’ hypothesis revisited. Nat. Rev. Cancer 2003, 3, 453–458. [Google Scholar]
- Itakura, E.; Huang, R.R.; Wen, D.R.; Cochran, A.J. “Stealth” melanoma cells in histology-negative sentinel lymph nodes. Am J. Surg. Pathol. 2011, 35, 1657–1665. [Google Scholar]
- Berndt, B.; Haverkampf, S.; Reith, G.; Keil, S.; Niggemann, B.; Zanker, K.S.; Dittmar, T. Fusion of CCL21 non-migratory active breast epithelial and breast cancer cells give rise to CCL21 migratory active tumor hybrid cell lines. PLoS ONE 2013, 8, e63711. [Google Scholar]
- McArdle, T.J.; Ogle, B.M.; Noubissi, F.K. An In Vitro Inverted Vertical Invasion Assay to Avoid Manipulation of Rare or Sensitive Cell Types. J. Cancer 2016, 7, 2333–2340. [Google Scholar]
- Zhang, L.N.; Huang, Y.H.; Zhao, L. Fusion of macrophages promotes breast cancer cell proliferation, migration and invasion through activating epithelial–mesenchymal transition and Wnt/beta-catenin signaling pathway. Arch. Biochem. Biophys. 2019, 676, 108137. [Google Scholar]
- Varley, K.E.; Mutch, D.G.; Edmonston, T.B.; Goodfellow, P.J.; Mitra, R.D. Intra-tumor heterogeneity of MLH1 promoter methylation revealed by deep single molecule bisulfite sequencing. Nucleic Acids Res. 2009, 37, 4603–4612. [Google Scholar]
- Yang, J.Y.; Ha, S.A.; Yang, Y.S.; Kim, J.W. p-Glycoprotein ABCB5 and YB-1 expression plays a role in increased heterogeneity of breast cancer cells: Correlations with cell fusion and doxorubicin resistance. BMC Cancer 2010, 10, 388. [Google Scholar]
- Ozel, C.; Seidel, J.; Meyer-Staeckling, S.; Brandt, B.H.; Niggemann, B.; Zanker, K.S.; Dittmar, T. Hybrid cells derived from breast epithelial cell/breast cancer cell fusion events show a differential RAF-AKT crosstalk. Cell Commun. Signal 2012, 10, 10. [Google Scholar]
- Nagler, C.; Zanker, K.S.; Dittmar, T. Cell Fusion, Drug Resistance and Recurrence CSCs. Adv. Exp. Med. Biol. 2011, 714, 173–182. [Google Scholar]
- Sergeant, G.; van Eijsden, R.; Roskams, T.; Van Duppen, V.; Topal, B. Pancreatic cancer circulating tumour cells express a cell motility gene signature that predicts survival after surgery. BMC Cancer 2012, 12, 527. [Google Scholar]
- Kaifi, J.T.; Yekebas, E.F.; Schurr, P.; Obonyo, D.; Wachowiak, R.; Busch, P.; Heinecke, A.; Pantel, K.; Izbicki, J.R. Tumor-cell homing to lymph nodes and bone marrow and CXCR4 expression in esophageal cancer. J. Natl. Cancer Inst. 2005, 97, 1840–1847. [Google Scholar]
- Chakraborty, A.K.; Pawelek, J.; Ikeda, Y.; Miyoshi, E.; Kolesnikova, N.; Funasaka, Y.; Ichihashi, M.; Taniguchi, N. Fusion hybrids with macrophage and melanoma cells up-regulate N-acetylglucosaminyltransferase V, beta1-6 branching, and metastasis. Cell Growth Differ. 2001, 12, 623–630. [Google Scholar]
- Funamizu, N.; Hu, C.; Lacy, C.; Schetter, A.; Zhang, G.; He, P.; Gaedcke, J.; Ghadimi, M.B.; Ried, T.; Yfantis, H.G.; et al. Macrophage migration inhibitory factor induces epithelial to mesenchymal transition, enhances tumor aggressiveness and predicts clinical outcome in resected pancreatic ductal adenocarcinoma. Int. J. Cancer 2013, 132, 785–794. [Google Scholar]
- Cayrefourcq, L.; Mazard, T.; Joosse, S.; Solassol, J.; Ramos, J.; Assenat, E.; Schumacher, U.; Costes, V.; Maudelonde, T.; Pantel, K.; et al. Establishment and characterization of a cell line from human circulating colon cancer cells. Cancer Res. 2015, 75, 892–901. [Google Scholar]
- Donnenberg, V.S.; Huber, A.; Basse, P.; Rubin, J.P.; Donnenberg, A.D. Neither epithelial nor mesenchymal circulating tumor cells isolated from breast cancer patients are tumorigenic in NOD-scid Il2rg(null) mice. NPJ Breast Cancer 2016, 2, 16004. [Google Scholar]
- Yamashita, Y.M.; Inaba, M.; Buszczak, M. Specialized Intercellular Communications via Cytonemes and Nanotubes. Annu. Rev. Cell Dev. Biol. 2018, 34, 59–84. [Google Scholar]
- Sahu, P.; Jena, S.R.; Samanta, L. Tunneling Nanotubes: A Versatile Target for Cancer Therapy. Curr. Cancer Drug Targets 2018, 18, 514–521. [Google Scholar]
- Rustom, A.; Saffrich, R.; Markovic, I.; Walther, P.; Gerdes, H.H. Nanotubular highways for intercellular organelle transport. Science 2004, 303, 1007–1010. [Google Scholar]
- Hsiung, F.; Ramirez-Weber, F.A.; Iwaki, D.D.; Kornberg, T.B. Dependence of Drosophila wing imaginal disc cytonemes on Decapentaplegic. Nature 2005, 437, 560–563. [Google Scholar]
- Burtey, A.; Wagner, M.; Hodneland, E.; Skaftnesmo, K.O.; Schoelermann, J.; Mondragon, I.R.; Espedal, H.; Golebiewska, A.; Niclou, S.P.; Bjerkvig, R.; et al. Intercellular transfer of transferrin receptor by a contact-, Rab8-dependent mechanism involving tunneling nanotubes. FASEB J. 2015, 29, 4695–4712. [Google Scholar]
- Benard, M.; Schapman, D.; Lebon, A.; Monterroso, B.; Bellenger, M.; Le Foll, F.; Pasquier, J.; Vaudry, H.; Vaudry, D.; Galas, L. Structural and functional analysis of tunneling nanotubes (TnTs) using gCW STED and gconfocal approaches. Biol. Cell 2015, 107, 419–425. [Google Scholar]
- Wang, Y.; Cui, J.; Sun, X.; Zhang, Y. Tunneling-nanotube development in astrocytes depends on p53 activation. Cell Death Differ. 2011, 18, 732–742. [Google Scholar]
- Lou, E.; Fujisawa, S.; Morozov, A.; Barlas, A.; Romin, Y.; Dogan, Y.; Gholami, S.; Moreira, A.L.; Manova-Todorova, K.; Moore, M.A. Tunneling nanotubes provide a unique conduit for intercellular transfer of cellular contents in human malignant pleural mesothelioma. PLoS ONE 2012, 7, e33093. [Google Scholar]
- Osswald, M.; Jung, E.; Sahm, F.; Solecki, G.; Venkataramani, V.; Blaes, J.; Weil, S.; Horstmann, H.; Wiestler, B.; Syed, M.; et al. Brain tumour cells interconnect to a functional and resistant network. Nature 2015, 528, 93–98. [Google Scholar]
- Desir, S.; O’Hare, P.; Vogel, R.I.; Sperduto, W.; Sarkari, A.; Dickson, E.L.; Wong, P.; Nelson, A.C.; Fong, Y.; Steer, C.J.; et al. Chemotherapy-Induced Tunneling Nanotubes Mediate Intercellular Drug Efflux in Pancreatic Cancer. Sci. Rep. 2018, 8, 9484. [Google Scholar]
- Sherer, N.M.; Mothes, W. Cytonemes and tunneling nanotubules in cell-cell communication and viral pathogenesis. Trends Cell Biol. 2008, 18, 414–420. [Google Scholar]
- Mattes, B.; Scholpp, S. Emerging role of contact-mediated cell communication in tissue development and diseases. Histochem Cell Biol. 2018, 150, 431–442. [Google Scholar]
- Abounit, S.; Zurzolo, C. Wiring through tunneling nanotubes—From electrical signals to organelle transfer. J. Cell Sci. 2012, 125, 1089–1098. [Google Scholar]
- Dupont, M.; Souriant, S.; Lugo-Villarino, G.; Maridonneau-Parini, I.; Verollet, C. Tunneling Nanotubes: Intimate Communication between Myeloid Cells. Front Immunol. 2018, 9, 43. [Google Scholar]
- Ariazi, J.; Benowitz, A.; De Biasi, V.; Den Boer, M.L.; Cherqui, S.; Cui, H.; Douillet, N.; Eugenin, E.A.; Favre, D.; Goodman, S.; et al. Tunneling Nanotubes and Gap Junctions-Their Role in Long-Range Intercellular Communication during Development, Health, and Disease Conditions. Front. Mol. Neurosci. 2017, 10, 333. [Google Scholar]
- Onfelt, B.; Nedvetzki, S.; Benninger, R.K.; Purbhoo, M.A.; Sowinski, S.; Hume, A.N.; Seabra, M.C.; Neil, M.A.; French, P.M.; Davis, D.M. Structurally distinct membrane nanotubes between human macrophages support long-distance vesicular traffic or surfing of bacteria. J. Immunol. 2006, 177, 8476–8483. [Google Scholar]
- Acquistapace, A.; Bru, T.; Lesault, P.F.; Figeac, F.; Coudert, A.E.; le Coz, O.; Christov, C.; Baudin, X.; Auber, F.; Yiou, R.; et al. Human mesenchymal stem cells reprogram adult cardiomyocytes toward a progenitor-like state through partial cell fusion and mitochondria transfer. Stem Cells 2011, 29, 812–824. [Google Scholar]
- Smith, I.F.; Shuai, J.; Parker, I. Active generation and propagation of Ca2+ signals within tunneling membrane nanotubes. Biophys. J. 2011, 100, L37–L39. [Google Scholar]
- Wang, X.; Gerdes, H.H. Transfer of mitochondria via tunneling nanotubes rescues apoptotic PC12 cells. Cell Death Differ. 2015, 22, 1181–1191. [Google Scholar]
- Sowinski, S.; Jolly, C.; Berninghausen, O.; Purbhoo, M.A.; Chauveau, A.; Kohler, K.; Oddos, S.; Eissmann, P.; Brodsky, F.M.; Hopkins, C.; et al. Membrane nanotubes physically connect T cells over long distances presenting a novel route for HIV-1 transmission. Nat. Cell Biol. 2008, 10, 211–219. [Google Scholar]
- Delage, E.; Cervantes, D.C.; Penard, E.; Schmitt, C.; Syan, S.; Disanza, A.; Scita, G.; Zurzolo, C. Differential identity of Filopodia and Tunneling Nanotubes revealed by the opposite functions of actin regulatory complexes. Sci. Rep. 2016, 6, 39632. [Google Scholar]
- Carter, K.P.; Hanna, S.; Genna, A.; Lewis, D.; Segall, J.E.; Cox, D. Macrophages enhance 3D invasion in a breast cancer cell line by induction of tumor cell tunneling nanotubes. Cancer Rep. 2019, 2, e1213. [Google Scholar]
- Petrova, V.; Annicchiarico-Petruzzelli, M.; Melino, G.; Amelio, I. The hypoxic tumour microenvironment. Oncogenesis 2018, 7, 10. [Google Scholar]
- Zhang, M.; Chen, J.; Wang, L.; Ji, X.; Yang, L.; Sheng, Y.; Liu, H.; Wang, H.; Wang, A.; Dai, X.; et al. Preliminary analysis of cellular sociology of co-cultured glioma initiating cells and macrophages in vitro. Transl. Neurosci. Clin. 2016, 2, 10. [Google Scholar]
- Hanna, S.J.; McCoy-Simandle, K.; Leung, E.; Genna, A.; Condeelis, J.; Cox, D. Tunneling nanotubes, a novel mode of tumor cell-macrophage communication in tumor cell invasion. J. Cell Sci. 2019, 132, jcs223321. [Google Scholar]
- Goswami, S.; Sahai, E.; Wyckoff, J.B.; Cammer, M.; Cox, D.; Pixley, F.J.; Stanley, E.R.; Segall, J.E.; Condeelis, J.S. Macrophages promote the invasion of breast carcinoma cells via a colony-stimulating factor-1/epidermal growth factor paracrine loop. Cancer Res. 2005, 65, 5278–5283. [Google Scholar]
- Dwyer, A.R.; Greenland, E.L.; Pixley, F.J. Promotion of Tumor Invasion by Tumor-Associated Macrophages: The Role of CSF-1-Activated Phosphatidylinositol 3 Kinase and Src Family Kinase Motility Signaling. Cancers (Basel) 2017, 9, 68. [Google Scholar]
- Mohr, M.; Zaenker, K.S.; Dittmar, T. Fusion in cancer: An explanatory model for aneuploidy, metastasis formation, and drug resistance. Methods Mol. Biol. 2015, 1313, 21–40. [Google Scholar]
- Zhou, X.; Merchak, K.; Lee, W.; Grande, J.P.; Cascalho, M.; Platt, J.L. Cell Fusion Connects Oncogenesis with Tumor Evolution. Am. J. Pathol. 2015, 185, 2049–2060. [Google Scholar]
- Jackson, C.M.; Choi, J.; Lim, M. Mechanisms of immunotherapy resistance: Lessons from glioblastoma. Nat. Immunol. 2019, 20, 1100–1109. [Google Scholar]
- Harkness, T.; Weaver, B.A.; Alexander, C.M.; Ogle, B.M. Cell fusion in tumor development: Accelerated genetic evolution. Crit. Rev. Oncog. 2013, 18, 19–42. [Google Scholar]
| Description | Phenotype (+: Expression; -: No Expression) |
|---|---|
| Traditional CTCs [3] | EpCAM+/(cytokeratin)CK+/CD45-, intact DAPI+ nucleus, ≥8 µm |
| CTC clusters [23] | ≥2 EpCAM+/CK+/CD45-/DAPI+ CTCs in aggregate |
| Apoptotic CTCs [24] | DAPI fluorescence intensity significantly less compared to a CTC Dotted cytoplasmic CK staining pattern (in contrast to filamentous) |
| CTC debris [24] | EpCAM+/CK+, CD45-, DAPI, <4 µm |
| EMT CTCs [6,7] | EpCAM+/CK+/EMT+ (e.g., vimentin, N-Cadherin) |
| Stem-cell CTCs [25,26] | EpCAM+/CK+/CD133+/CD44+/CD24-/ALDH1+ |
| Macrophage-tumor fusion cells (MTFs) [21,22,27,28] Cancer-associated macrophage-like cells (CAMLs) [29] | EpCAM+/CK+/CD14+/CD45+, ≥30 µm Diffuse cytoplasmic CK staining pattern ≥1 DAPI+ nucleus Polymorph cell shape |
| Cancer Type | Fusion | Marker Co-Expression (Detection Method) | References |
|---|---|---|---|
| Breast cancer | Tumor cell-macrophage | CK, CD163, MAC387, DAP12 CD14, CD45 (IHC) | Shabo et al. [20,66,67], Tang et al. [76], Adams et al. [29] |
| Colorectal cancer | Tumor cell-macrophage | CK, CD14, CD45 (IHC) | Clawson et al. [27], Kaifi et al. [28] |
| Esophageal cancer | Tumor cell-macrophage | CK, CD14, CD45 (IHC) | Tang et al. [76] |
| Malignant melanoma | Tumor cell-macrophage | EpCAM, CK, CD14, CD45, CD163, CD204, CD206, ALCAM, MLANA (IHC/IF), BRAF mutations (PCR) Short tandem repeat analysis (PCR) | Clawson et al. [22], Lazova et al. [17] |
| Multiple myeloma | Tumor cell-osteoclasts | TRACP (IF), specific translocations (FISH) | Andersen et al. [78] |
| Non-small cell lung cancer | Tumor cell-macrophage | CK, CD14, CD45 (IHC) | Tang et al. [76] |
| Ovarian cancer | Tumor cell-bone marrow-derived cell | EpCAM, CD45, CA125 (IHC/IF); EpCAM, CD14, CD34, CD44, CD68, CD117, CD133, CD163, CD204, CD206, CA125, CXCR4 (FC) | Ramakrishnan et al. [16] |
| Pancreatic ductal adenocarcinoma | Tumor cell-macrophage | EpCAM, MIF, ALDH1A1, CD44, CD68, CD163, CD204/MSR1, CD206, CXCR4, S100PBP, Pan-keratin, ZG16B (IHC/IF) CK, CD14, CD45 (IHC) | Clawson et al. [21], Tang et al. [76], Adams et al. [29] |
| Prostate cancer | Tumor cell-macrophage | CK, CD14, CD45 (IHC) | Tang et al. [76], Adams et al. [29] |
| Renal cell carcinoma | Tumor cell-bone marrow-derived cell, Tumor cell-macrophage | PCR and blood group alleles, FISH analysis and Y chromosome detection, CK, CD14, CD45 (IHC) | Chakraborty et al. [54], Yilmaz et al. [53], Tang et al. [76] |
| Features | Cancer Type | References |
|---|---|---|
| Enhanced migration | Breast cancer Ovarian cancer | Berndt et al. (2013) [85] Ramakrishnan et al. (2013) [16] |
| Increased invasive and migratory potential | Breast cancer Breast cancer | Noubissi et al. (2015) [13] McArdle et al. (2016) [86] |
| Metastatic potential after xenografting | Non-small cell lung cancer Malignant melanoma Malignant melanoma Pancreatic ductal adenocarcinoma Hepatocellular carcinoma | Xu et al. (2014) [51] Chakraborty et al. (2000) [65] Clawson et al. (2015) [22] Clawson et al. (2017) [21] Li et al. (2014) [52] |
| EMT properties | Non-small cell lung cancer Breast cancer | Xu et al. (2014) [51] Zhang et al. (2019) [87] |
| Stem cell properties | Ovarian Non-small cell lung cancer | Ramakrishnan et al. (2013) [16] Xu et al. (2014) [51] |
| Genetic evidence of fusion events | Malignant melanoma Endometrial cancer Multiple myeloma Renal cell cancer | Lazova et al. (2013) [17] Varley et al. (2009) [88] Anderson et al. (2007) [78] Yilmaz et al. (2005) [53] |
| Chemoresistance | Breast cancer Breast cancer Breast cancer Hepatocellular cancer | Yang et al. (2010) [89] Ozel et al. (2012) [90] Nagler et al. (2011) [91] Wang et al. (2015) [40] |
| Patients’ survival and cancer recurrences | Pancreatic ductal adenocarcinoma Pooled analysis of solid cancers: breast/esophagus/non-small cell lung/prostate/renal cell cancer, pancreatic ductal adenocarcinoma (≥6 fusion cells/7.5 mL whole blood; ≥50 µm cell size) | Gast et al. (2018) [55] Tang et al. (2018) [76] |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Manjunath, Y.; Porciani, D.; Mitchem, J.B.; Suvilesh, K.N.; Avella, D.M.; Kimchi, E.T.; Staveley-O’Carroll, K.F.; Burke, D.H.; Li, G.; Kaifi, J.T. Tumor-Cell–Macrophage Fusion Cells as Liquid Biomarkers and Tumor Enhancers in Cancer. Int. J. Mol. Sci. 2020, 21, 1872. https://doi.org/10.3390/ijms21051872
Manjunath Y, Porciani D, Mitchem JB, Suvilesh KN, Avella DM, Kimchi ET, Staveley-O’Carroll KF, Burke DH, Li G, Kaifi JT. Tumor-Cell–Macrophage Fusion Cells as Liquid Biomarkers and Tumor Enhancers in Cancer. International Journal of Molecular Sciences. 2020; 21(5):1872. https://doi.org/10.3390/ijms21051872
Chicago/Turabian StyleManjunath, Yariswamy, David Porciani, Jonathan B. Mitchem, Kanve N. Suvilesh, Diego M. Avella, Eric T. Kimchi, Kevin F. Staveley-O’Carroll, Donald H. Burke, Guangfu Li, and Jussuf T. Kaifi. 2020. "Tumor-Cell–Macrophage Fusion Cells as Liquid Biomarkers and Tumor Enhancers in Cancer" International Journal of Molecular Sciences 21, no. 5: 1872. https://doi.org/10.3390/ijms21051872
APA StyleManjunath, Y., Porciani, D., Mitchem, J. B., Suvilesh, K. N., Avella, D. M., Kimchi, E. T., Staveley-O’Carroll, K. F., Burke, D. H., Li, G., & Kaifi, J. T. (2020). Tumor-Cell–Macrophage Fusion Cells as Liquid Biomarkers and Tumor Enhancers in Cancer. International Journal of Molecular Sciences, 21(5), 1872. https://doi.org/10.3390/ijms21051872








