Genetic Characterization of Endometriosis Patients: Review of the Literature and a Prospective Cohort Study on a Mediterranean Population
Abstract
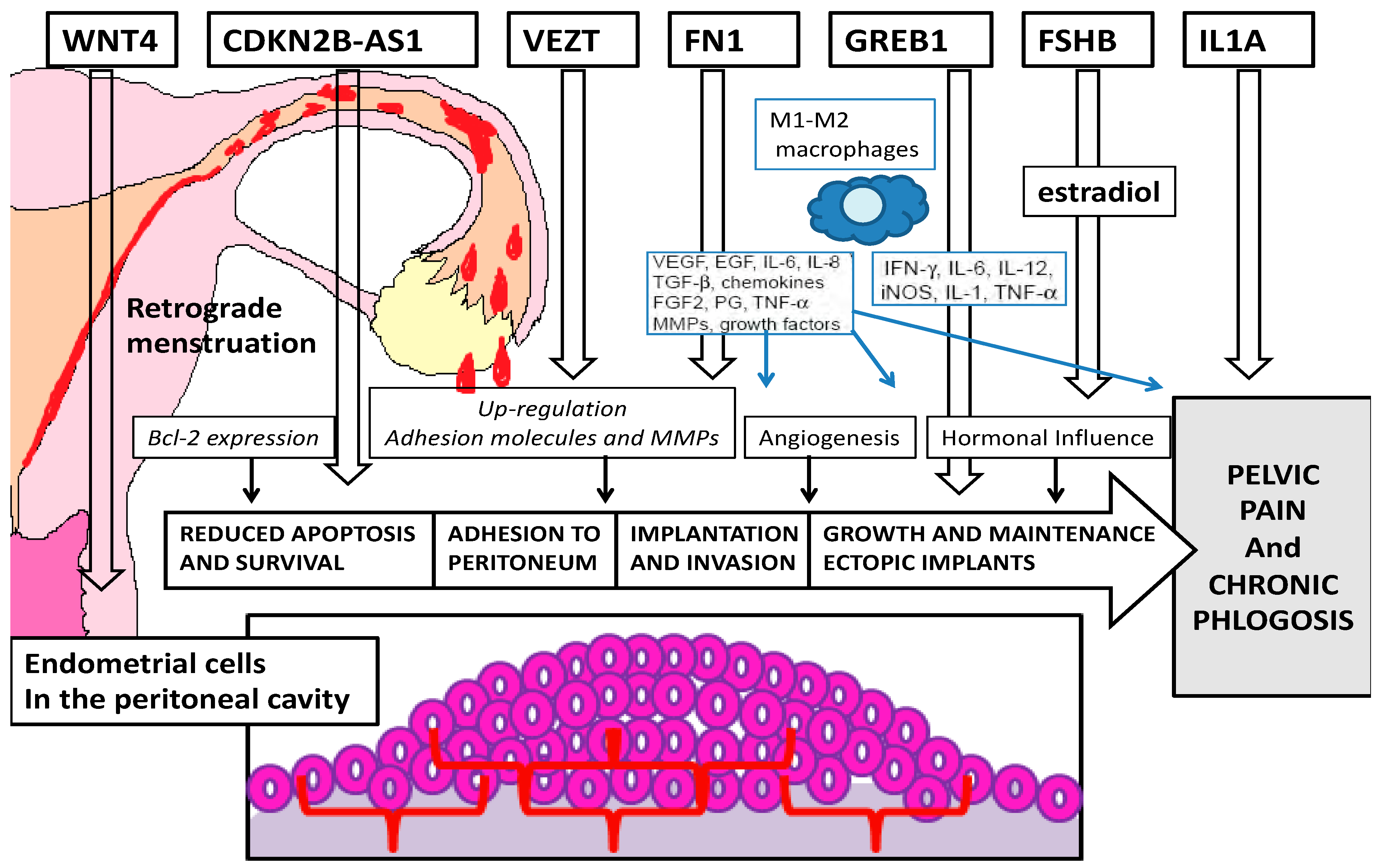
1. Introduction
1.1. Review of the Literature
1.2. Cohort Study
Patients and Study Design
2. Results
2.1. Review of the Literature
2.1.1. Gene Polymorphisms
2.1.2. Genes Involved in Stereidogenesis and Receptorial Activity of Sex Hormones
2.1.3. Genes Involved in Inflammation and Immune Response
2.1.4. Genes Involved in the Processes of Tissue Remodeling and Neoangiogenesis
2.1.5. Genes Regulating Metabolism
2.1.6. Genes Related to the Process of DNA Repair
2.1.7. Genome-Wide Association Study (GWAS) and Next-Generation Sequencing (NGS)
2.1.8. Epigenetics
2.2. Cohort Study
3. Discussion
4. Materials and Methods
4.1. Sampling and Genotyping
4.2. Statistical Analysis
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| SNP | Single nucleotide polymorphism |
| PCR | Polymerase chain reaction |
| UTR | Untranslated region |
| AR | Androgen receptor |
| ER | Estrogenic receptor |
| PR | Progesterone receptor |
| A | Adenine |
| T | Thymine |
| C | Cytosine |
| G | Guanine |
| TGF-β | Transforming growth factor β |
| IL | Interleukin |
| VEGF | Vascular endothelial growth factor |
| TNF | Tumour necrosis factor |
| HLA | Human leukocyte antigen |
| MHC | Major histocompatibility complex |
| EGFR | Endothelial growth factor receptor |
| mRNA | Messenger ribonucleic acid |
| MMP | Matrix metalloproteinase |
| GSTM1 | Glutathione S-transferase mu, M1 |
| GSTP1 | Glutathione S-transferase P1 |
| GSTT1 | Glutathione S-transferase theta, T1 |
| NAT | N-acetyltransferase |
| CYP | Cytochrome P450 |
| XRCC | X-ray repair cross-complementing |
| ERCC | Excision repair cross-complementing |
| GWAS | Genome-wide association study |
| NGS | Next-generation sequencing |
| KRAS | Kirsten rat sarcoma viral oncogene homolog |
| eQTLs | Expression quantitative traits loci |
| MiRNA | Micro RNA |
| NcRNA | Non-coding RNA |
References
- Alio, L.; Angioni, S.; Arena, S.; Bartiromo, L.; Bergamini, N.; Berlanda, N.; Bonanni, V.; Bonin, C.; Buggio, L.; Candiani, M.; et al. Endometriosis: Seeking optimal management in women approaching menopause. Climacteric 2019, 22, 329–338. [Google Scholar] [CrossRef] [PubMed]
- Deiana, D.; Gessa, S.; Anardu, M.; Danilidis, A.; Nappi, L.; D’Alterio, M.N.; Pontis, A.; Angioni, S. Genetic of endometriosis: A comprehensive review. Gynecol. Endocrinol. 2019, 35, 553–558. [Google Scholar] [CrossRef] [PubMed]
- Melis, I.; Agus, M.; Pluchino, N.; Sardo, A.D.S.; Litta, P.; Melis, G.B.; Angioni, S. Alexithymia in women with deep endometriosis? A pilot. Study. J. Endometr. Pelvic. Pain Dis. 2014, 6, 26–33. [Google Scholar]
- Angioni, S. New insights on endometriosis. Minerva Ginecol. 2017, 69, 438–439. [Google Scholar] [PubMed]
- Locci, R.; Nisolle, M.; Angioni, S.; Foidart, J.M.; Munat, C. Expression of the gamma 2 chain of laminin-332 in eutopic and ectopic endometrium of patients with endometriosis. Rep. Biol. Endocrinol. 2013, 11, 94. [Google Scholar] [CrossRef] [PubMed]
- Simpson, J.L.; Elias, S.; Malinak, L.R.; Buttram, V.C., Jr. Heritable aspects of endometriosis. Genetic studies. Am. J. Obstet. Gynecol. 1980, 137, 327–331. [Google Scholar] [CrossRef]
- Treloar, S.A.; O’Connor, D.T.; O’Connor, V.M.; Martin, N.G. Genetic influences on endometriosis in an Australian twin sample. Fertil. Steril. 1999, 71, 701–710. [Google Scholar] [CrossRef]
- Lettre, G.; Hirschhorn, J.N. Small island, big genetic discoveries. Nat. Genet. 2015, 47, 1224–1225. [Google Scholar] [CrossRef]
- Mafra, F.; Catto, M.; Bianco, B.; Barbosa, C.P.; Christofolini, D. Association of WNT4 polymorphisms with endometriosis in infertile patients. J. Assist. Reprod. Genet. 2015, 32, 1359–1364. [Google Scholar] [CrossRef]
- Pagliardini, L.; Gentilini, D.; Vigano’, P.; Panina-Bordignon, P.; Busacca, M.; Candiani, M.; Di Blasio, A.M. An Italian association study and meta-analysis with previous GWAS confirm WNT4, CDKN2BAS and FN1 as the first identified susceptibility loci for endometriosis. J. Med. Genet. 2013, 50, 43–46. [Google Scholar] [CrossRef]
- Vainio, S.; Heikkilä, M.; Kispert, A.; Chin, N.; McMahon, A.P. Female development in mammals is regulated by Wnt-4 signalling. Nature 1999, 397, 405–409. [Google Scholar] [CrossRef] [PubMed]
- Gaetje, R.; Holtrich, U.; Engels, K.; Kissler, S.; Rody, A.; Karn, T.; Kaufmann, M. Endometriosis may be generated by mimicking the ontogenetic development of the female genital tract. Fertil. Steril. 2007, 87, 651–656. [Google Scholar] [CrossRef] [PubMed]
- Sundqvist, J.; Xu, H.; Vodolazkaia, A.; Fassbender, A.; Kyama, C.; Bokor, A.; Gemzell-Danielsson, K.; D’Hooghe, T.M.; Falconer, H. Replication of endometriosis-associated single-nucleotide polymorphisms from genome-wide association studies in a Caucasian population. Hum. Reprod. 2013, 28, 835–839. [Google Scholar] [CrossRef] [PubMed]
- Wu, Z.; Yuan, M.; Li, Y.; Fu, F.; Ma, W.; Li, H.; Wang, W.; Wang, S. Analysis of WNT4 polymorphism in Chinese Han women with endometriosis. Reprod. Biomed. Online 2015, 30, 415–420. [Google Scholar] [CrossRef] [PubMed]
- Jordan, B.K.; Mohammed, M.; Ching, S.T.; Délot, E.; Chen, X.N.; Dewing, P.; Swain, A.; Rao, P.N.; Elejalde, B.R.; Vilain, E. Up-regulation ofWNT-4 signaling and dosage-sensitive sex reversal in humans. Am. J. Hum. Genet. 2001, 68, 1102–1109. [Google Scholar] [CrossRef]
- Rahmioglu, N.; Nyholt, D.R.; Morris, A.P.; Missmer, S.A.; Montgomery, G.W.; Zondervan, K.T. Genetic variants underlying risk of endometriosis: Insights from meta-analysis of eight genome-wide association and replication datasets. Hum. Reprod. Update 2014, 20, 702–716. [Google Scholar] [CrossRef]
- Nyholt, D.R.; Low, S.K.; Anderson, C.A.; Painter, J.N.; Uno, S.; Morris, A.P.; MacGregor, S.; Gordon, S.D.; Henders, A.K.; Martin, N.G.; et al. Genome-wide association meta-analysis identified new endometriosis risk loci. Nat. Genet. 2012, 44, 1355–1359. [Google Scholar] [CrossRef]
- Uno, S.; Zembutsu, H.; Hirasawa, A.; Takahashi, A.; Kubo, M.; Akahane, T.; Aoki, D.; Kamatani, N.; Hirata, K.; Nakamura, Y. A genome-wide association study identifies genetic variants in the CDKN2BAS locus associated with endometriosis in Japanese. Nat. Genet. 2010, 42, 707–710. [Google Scholar] [CrossRef]
- Painter, J.N.; Anderson, C.A.; Nyholt, D.R.; Macgregor, S.; Lin, J.; Lee, S.H.; Lambert, A.; Zhao, Z.Z.; Roseman, F.; Guo, Q.; et al. Genome-wide association study identifies a locus at 7p15.2 associated with the development of endometriosis. Nat. Genet. 2011, 43, 51–54. [Google Scholar] [CrossRef]
- Haldsworth-Carson, S.J.; Fung, J.N.; Luong, H.T.; Sapkota, Y.; Bowdler, L.M.; Wallace, L.; Teh, W.T.; Powell, J.E.; Girling, J.E.; Healey, M.; et al. Endometrial vezatin and it association with endometriosis risk. Hum. Reprod. 2016, 31, 999–1013. [Google Scholar] [CrossRef]
- Küssel-Andermann, P.; El-Amraoui, A.; Safieddine, S.; Nouaille, S.; Perfettini, I.; Lecuit, M.; Cossart, P.; Wolfrum, U.; Petit, C. Vezatin, a novel transmembrane protein, bridges myosin VIIA to the cadherin-catenins complex. Embo J. 2000, 19, 6020–6029. [Google Scholar] [CrossRef] [PubMed]
- Blaschuk, O.W.; Rowlands, T.M. Plasma membrane components of adherens junctions. Mol. Membr. Biol. 2002, 19, 75–80. [Google Scholar] [CrossRef] [PubMed]
- Luong, H.T.T.; Painter, J.N.; Sapkota, Y.; Niholt, D.R.; Rogers, P.A.; Montgomery, G.W. Identifying the functional role of VEZT Gene for endometriosis risk. Ann. Transl. Med. 2015, 3, AB028. [Google Scholar]
- Pagliardini, L.; Gentilini, D.; Sanchez, A.M.; Candiani, M.; Viganò, P.; Di Blasio, A.M. Replication and meta-analysis of previous genome-wide association studies confirm vezatin as the locus with the strongest evidence for association with endometriosis. Hum. Reprod. 2015, 30, 987–993. [Google Scholar] [CrossRef] [PubMed]
- Meola, J.; Rosa, E.; Silva, J.C.; Dentillo, D.B.; Da Silva, W.A., Jr.; Veiga-Castelli, L.C.; Bernardes, L.A.; Ferriani, R.A.; De Paz, C.C.; Giuliatti, S.; et al. Differentially expressed genes in eutopic and ectopic endometrium of women with endometriosis. Fertil. Steril. 2010, 93, 1750–1773. [Google Scholar] [CrossRef]
- Guo, S.-W. Nuclear factor-kappab (NF-κB): An unsuspected major culprit in the pathogenesis of endometriosis that is still at large? Gynecol. Obstet. Investig. 2007, 63, 71–97. [Google Scholar] [CrossRef]
- Sapkota, Y.; Steinthorsdottir, V.; Morris, A.P.; Fassbender, A.; Rahmioglu, N.; De Vivo, I.; Buring, J.E.; Zhang, F.; Edwards, T.L.; Jones, S.; et al. Meta-analysis identifies five novel loci associated with endometriosis highlighting key genes involved in hormone metabolism. Nat. Commun. 2017, 8, 15539. [Google Scholar] [CrossRef]
- Ruth, K.S.; Beaumont, R.N.; Tyrrell, J.; Jones, S.E.; Tuke, M.A.; Yaghootkar, H.; Wood, A.R.; Freathy, R.M.; Weedon, M.N.; Frayling, T.M.; et al. Genetic evidence that lower circulating FSH levels lengthen menstrual cycle, increase age at menopause and impact female reproductive health. Hum. Reprod. 2016, 31, 473–481. [Google Scholar] [CrossRef]
- Matalliotakis, M.; Zervou, M.I.; Matalliotaki, C.; Ramhioglu, N.; Koumantakis, G.; Kalogiannidis, I.; Prapas, I.; Zondervan, K.; Spandidos, D.A.; Matalliotakis, I.; et al. The role of gene polymorphisms in endometriosis. Molec. Med. Rep. 2017, 16, 5881–5886. [Google Scholar] [CrossRef]
- Revised American Society for Reproductive Medicine classification of endometriosis: 1996. Fertil. Steril. 1997, 67, 817–821. [CrossRef]
- Vercellini, P.; Viganò, P.; Somigliana, E.; Fedele, L. Endometriosis: Pathogenesis and treatment. Nat. Rev. Endocrinol. 2014, 10, 261–275. [Google Scholar] [CrossRef] [PubMed]
- Laganà, A.S.; Garzon, S.; Gotte, M.; Viganò, P.; Franchi, M.; Ghezzi, F.; Martin, D.C. Pathogenesis of Endometriosis: Molecular and Cell Biology Insights. Int. J. Mol. Sci. 2019, 20, 5615. [Google Scholar] [CrossRef] [PubMed]
- Angioni, S.; Cofelice, V.; Pontis, A.; Tinelli, R.; Socolov, R. New trends of progestins treatment of endometriosis. Gynecol. Endocrinol. 2014, 30, 769–773. [Google Scholar] [CrossRef] [PubMed]
- Angioni, S.; Cofelice, V.; Sedda, F.; Stochino Loi, E.; Multinu, F.; Pontis, A.; Melis, G.B. Progestins for symptomatic endometriosis: Results of clinical studies. Curr. Drug. Ther. 2015, 10, 91–104. [Google Scholar] [CrossRef]
- Wang, Z.; Yoshida, S.; Negoro, K.; Kennedy, S.; Barlow, D.; Maruo, T. Polymorphisms in the estrogen receptor beta gene but not estrogen receptor alpha gene affect the risk of developing endometriosis in a Japanese population. Fertil. Steril. 2004, 81, 1650–1656. [Google Scholar] [CrossRef]
- Lee, G.H.; Kim, S.H.; Choi, Y.M.; Suh, C.S.; Kim, J.G.; Moon, S.Y. Estrogen receptor beta gene +1730 G/A polymorphism in women with endometriosis. Fertil. Steril. 2007, 88, 785–788. [Google Scholar] [CrossRef]
- Lattuada, D.; Somigliana, E.; Vigano, P.; Candiani, M.; Pardi, G.; Di Biasio, A.M. Genetics of endometriosis: A role for the progesterone receptor gene polymorphism PROGINS? Clin. Endocrinol. 2004, 61, 190–194. [Google Scholar] [CrossRef]
- Wieser, F.; Schneeberger, C.; Tong, D.; Tempfer, C.; Huber, J.C.; Wenzl, R. PROGINS receptor gene polymorphism is associated with endometriosis. Fertil. Steril. 2002, 77, 309–312. [Google Scholar] [CrossRef]
- De Carvalho, C.V.; Nogueira-De-Souza, N.C.; Costa, A.M.; Baracat, E.C.; Girao, M.J.; D’Amora, P.; Schor, E.; da Silva, I.D. Genetic polymorphisms of cytochrome P450cl7alpha (CYP17) and progesterone receptor gene (PROGINS) in the assessment of endometriosis risk. Gynecol. Endocrinol. 2007, 23, 29–33. [Google Scholar] [CrossRef]
- Govindan, S.; Ahmad, S.N.; Vedicherla, B.; Kodati, V.; Jahan, P.; Rao, K.P.; Ahuja, Y.R.; Hasan, Q. Association of progesterone receptor gene (PROGINS) with endometriosis, uterine fibroids and breast cancer. Cancer Biomark. 2007, 3, 73–77. [Google Scholar] [CrossRef]
- Treloar, S.A.; Zhao, Z.Z.; Armitage, T.; Duffy, D.L.; Wicks, J.; O’Connor, D.T.; Martin, N.G.; Montgomery, G.W. Association between polymorphisms in the progesterone receptor gene and endometriosis. Mol. Hum. Reprod. 2005, 11, 641–647. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Van Kaam, K.J.; Romano, A.; Schouten, J.P.; Dunselman, G.A.; Groothuis, P.G. Progesterone receptor polymorphism +331G/A is associated with a decreased risk of deep infiltrating endometriosis. Hum. Reprod. 2007, 22, 129–135. [Google Scholar] [CrossRef] [PubMed]
- Slayden, O.D.; Nayak, N.R.; Burton, K.A.; Chwalisz, K.; Cameron, S.T.; Critchley, H.O.; Baird, D.T.; Brenner, R.M. Progesterone antagonist increase androgen receptor expression in the rhesus macaque and human endometrium. J. Clin. Endocrinol. Metab. 2001, 86, 2668–2679. [Google Scholar] [CrossRef] [PubMed]
- Mertens, H.J.; Heinemann, M.J.; Theunissen, P.H.; de Jong, F.H.; Evers, J.L. Androgen, estrogen and progesterone receptor expression in the human uterus during the mestrual cycle. Eur. J. Obstet. Gynecol. Reprod. Biol. 2001, 98, 58–65. [Google Scholar] [CrossRef]
- Tuckermann, E.M.; Okon, M.A.; Li, T.; Laird, S.M. Do androgens have a direct effect on endometrial function? An in vitro study. Fertil. Steril. 2000, 74, 771–779. [Google Scholar] [CrossRef]
- Carneiro, M.M.; Morsch, D.M.; Camargos, A.F.; Reis, F.M.; Spritzer, P.M. Androgen receptor and 5α-reductase are expressed in pelvic endometriosis. Int. J. Obstet. Gynaecol. 2008, 115, 113–117. [Google Scholar] [CrossRef]
- Gelmann, P.E. Molecular Biology of the AR. J. Clin. Oncol. 2002, 20, 3001–3015. [Google Scholar] [CrossRef]
- Rajender, S.; Singh, L.; Thangaraj, K. Phenotypic heterogeneity of mutations in androgen receptor gene. Asian J. Androl. 2007, 9, 147–179. [Google Scholar]
- Shaik, N.A.; Govindan, S.; Kodati, V.; Rao, K.P.; Hasan, Q. Polymorphic (CAG) repeats in the androgen receptor gene: A risk marker for endometriosis and uterine leiomyomas. Hematol. Oncol. Stem Cel. Ther. 2009, 2, 289–293. [Google Scholar] [CrossRef]
- Hsieh, Y.Y.; Chang, C.C.; Tsai, F.J.; Wu, J.Y.; Tsai, C.H.; Tsai, H.D. Androgen receptor trinucleotide polymorphism in endometriosis. Fertil. Steril. 2001, 76, 412–413. [Google Scholar] [CrossRef]
- Lattuada, D.; Viganò, P.; Somigliana, E.; Odorizzi, M.P.; Vignali, M.; Di Biasio, A.M. Androgen receptor gene cytosine, adenine, and guanine trinucleotide repeats in patients with endometriosis. J. Soc. Gynecol. Investig. 2004, 11, 237–240. [Google Scholar] [CrossRef] [PubMed]
- Hsieh, Y.Y.; Chang, C.C.; Tsai, F.J.; Hsu, Y.; Tsai, H.D.; Tsai, C.H. Polymorphisms for interleukin-4 (IL-4)—590 promoter, IL-4 intron3, and tumor necrosis factor alpha -308 promoter: Non-association with endometriosis. J. Clin. Lab. Anal. 2002, 16, 121–126. [Google Scholar] [CrossRef] [PubMed]
- Yu, Y.X.; Xiu, Y.L.; Chen, X.; Li, Y.L. Transforming Growth Factor-beta 1 involved in the pathogenesis of endometriosis through regulating expression of vascular endothelial growth factor under hypoxia. Chin. Med. J. 2017, 130, 950–956. [Google Scholar] [CrossRef] [PubMed]
- Wieser, F.; Fabjani, G.; Tempfer, C.; Schneeberger, C.; Zeillinger, R.; Huber, J.C.; Wenzl, R. Tumor necrosis factor–alpha promoter polymorphisms and endometriosis. J. Soc. Gynecol. Investig. 2002, 9, 313–318. [Google Scholar] [CrossRef]
- Somigliana, E.; Viganò, P.; Vignali, M. Endometriosis and unexplained recurrent spontaneous abortion: Pathological states resulting from aberrant modulation of natural killer cell function? Hum. Reprod. Update 1999, 5, 41–52. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Lin, Q.; Guo, S. Study on polymorphism of human leukocyte antigen I in patients with endometriosis. Zhanghua Fu Chan Ke Za Zhi 2001, 36, 150–152. [Google Scholar]
- Ishii, K.; Takakuwa, K.; Kashima, K.; Tamura, M.; Tanaka, K. Associations between patients with endometriosis and HLA class II; the analysis of HLA-DQB1 and HLA- DPB1 genotypes. Hum. Reprod. 2003, 18, 985–989. [Google Scholar] [CrossRef]
- Pabalan, N.; Jarjanazi, H.; Christofolini, D.M.; Bianco, B.; Barbosa, C.P. Association of the protein tyrosine phosphatase non receptor 22 polymorphism (PNTN22) with endometriosis: A meta-analysis. Rev. Artic. Einstein 2017, 15, 105–111. [Google Scholar] [CrossRef]
- Perini, J.A.; Cardoso, J.V.; Berardo, P.T.; Vianna-Jorge, R.; Nasciutti, L.E.; Bellodi-Privato, M.; Machado, D.E.; Abrao, M.S. Role of vascular endothelial growth factor polymorphisms (−2578C > A, −460T > C, −1154G > A, +405G > C and + 936C > T) in endometriosis: A case-control study with Brazilians. BMC Womens Health 2014, 14, 117. [Google Scholar] [CrossRef]
- Hsieh, Y.Y.; Chang, C.C.; Tsai, F.J.; Lin, C.C.; Tsai, C.H. T homozygote and allele of epidermal growth factor receptor 2073 gene polymorphism are associated with higher susceptibility to endometriosis and leiomyomas. Fertil. Steril. 2005, 83, 796–799. [Google Scholar] [CrossRef]
- Guo, S.W. Glutathione S-transferases M1/T1 gene polymorphism and endometriosis: A meta-analysis of genetic association studies. Mol. Hum. Reprod. 2005, 11, 729–743. [Google Scholar] [CrossRef] [PubMed]
- Arvanatis, D.A.; Koumantakis, G.E.; Gouemenou, A.G.; Metalliokatis, E.E.; Spandidos, D.A. CYP1A1, CYP19, and GSTM1 polymorphisms increase the risk of endometriosis. Fertil. Steril. 2003, 79, 707–709. [Google Scholar] [CrossRef]
- Hsieh, Y.Y.; Chang, C.C.; Tsai, F.J.; Lin, C.C.; Tsai, C.H. Cytochrome P450c17alpha 5’- untranslated region *T/C polymorphism in endometriosis. J. Genet. 2004, 83, 189–192. [Google Scholar] [CrossRef] [PubMed]
- Lambrinoudaki, I.V.; Augoulea, A.; Christodoulakos, G.E.; Economou, E.V.; Kaparos, G.; Kontoravdis, A.; Papadias, C.; Creatsas, G. Measurable serum markers of oxidative stress response in women with endometriosis. Fertil. Steril. 2009, 91, 46–50. [Google Scholar] [CrossRef]
- Attar, R.; Cacina, C.; Sozen, S.; Attar, E.; Agachan, B. DNA repair genes in endometriosis. Gen. Mol. Res. 2010, 9, 629–636. [Google Scholar] [CrossRef]
- Rahmioglu, N.; Montgomery, G.W.; Zondervan, K.T. Genetics of endometriosis. Womens Health 2015, 11, 577–586. [Google Scholar] [CrossRef]
- Treloar, S.A.; Wicks, J.; Nyholt, D.R.; Montgomery, G.W.; Bahlo, M.; Smith, V.; Dawson, G.; Mackay, I.J.; Weeks, D.E.; Bennett, S.T.; et al. Genome-wide Linkage Study in 1,176 affected sister pair families identifies a significant susceptibility locus for endometriosis on chromosome 10q26. Am. J. Hum. Genet. 2005, 77, 365–376. [Google Scholar] [CrossRef]
- Painter, J.N.; Nyholt, D.R.; Morris, A.; Zhao, Z.Z.; Henders, A.K.; Lambert, A.; Wallace, L.; Martin, N.G.; Kennedy, S.H.; Treloar, S.A.; et al. High-density fine-mapping of a chromosome 10q26 linkage peak suggests association between endometriosis and variants close to CYP2C19. Fertil. Steril. 2011, 95, 2236–2240. [Google Scholar] [CrossRef]
- Cribb, A.E.; Knight, M.J.; Dryer, D.; Guernsey, J.; Hender, K.; Tesch, M.; Saleh, T.M. Role of a polymorphic human CYP450 enzymes in estrone oxidation. Cancer Epidemiol. Biomark. Prev. 2006, 15, 551–558. [Google Scholar] [CrossRef]
- Martini, M.; Ciccarone, M.; Garganese, G.; Maggiore, C.; Evangelista, A.; Rahimi, S.; Zannoni, G.; Vittori, G.; Larocca, L.M. Possible involvement of hMLH1, p16INK4a and PTEN in the malignant transformation of endometriosis. Int. J. Cancer 2002, 102, 398–406. [Google Scholar] [CrossRef]
- Powell, J.E.; Fung, J.N.; Shakhbazov, K.; Sapkota, Y.; Cloonan, N.; Hemani, G.; Hillman, K.M.; Kaufmann, S.; Luong, H.T.; Bowdler, L.; et al. Endometriosis risk alleles at 1p36.12 act through inverse regulation of CDC42 and LINC00339. Hum. Mol. Genet. 2016, 25, 5046–5058. [Google Scholar] [PubMed]
- Grechukhina, O.; Petracco, R.; Popkhadze, S.; Massasa, E.; Paranjape, T.; Chan, E.; Flores, I.; Weidhaas, J.B.; Taylor, H.S. A polymorphism in a let-7 microRNA binding site of KRAS in women with endometriosis. EMBO Mol. Med. 2012, 4, 202–217. [Google Scholar] [CrossRef] [PubMed]
- Luong, H.T.; Nyholt, D.R.; Painter, J.N.; Chapman, B.; Kennedy, S.; Treloar, S.A.; Zondervan, K.T.; Montgomery, G.W. No evidence for genetic association with the let-7 microRNA-binding site or the common KRAS variants in risk of endometriosis. Hum. Reprod. 2012, 12, 3616–3621. [Google Scholar] [CrossRef] [PubMed]
- Chettier, R.; Albertsen, H.M.; Ward, K. Next Generation Sequencing of families with endometriosis identifies new genomic regions likely to contribute to heritability. Fertil. Steril. 2014, 102, 76–77. [Google Scholar] [CrossRef]
- Er, T.K.; Su, Y.F.; Wu, C.C.; Chen, C.C.; Wang, J.; Hsieh, T.H.; Herreros-Villanueva, M.; Chen, W.T.; Chen, Y.T.; Liu, T.C.; et al. Targeted next-generation sequencing for molecular diagnosis of endometriosis-associated ovarian cancer. J. Mol. Med. 2016, 94, 835–847. [Google Scholar] [CrossRef]
- Sapkota, Y.; Vivo, I.; Steinthorsdottir, V.; Fassbender, A.; Bowdler, L.; Buring, J.E.; Edwards, T.L.; Jones, S.; Dorien, O.; Peterse, D.; et al. Analysis of potential protein-modifying variants in 9000 endometriosis patients and 150000 controls of European ancestry. Sci. Rep. 2017, 7, 11380. [Google Scholar] [CrossRef]
- Fung, J.N.; Montgomery, G.W. Genetics of endometriosis: State of art on genetic risk factors for endometriosis. Best. Pract. Res. Clin. Obstetr. Gynecol. 2018, 50, 61–71. [Google Scholar] [CrossRef]
- Freedman, M.L.; Monteiro, A.N.; Gayther, S.A.; Coetzee, G.A.; Risch, A.; Plass, C.; Casey, G.; De Biasi, M.; Carlson, C.; Duggan, D.; et al. Principles for the post- GWAS functional characterization of cancer risk. Nat. Genet. 2011, 43, 513–518. [Google Scholar] [CrossRef]
- Westra, H.J.; Peters, M.J.; Esko, T.; Yaghootkar, H.; Schurmann, C.; Kettunen, J.; Christiansen, M.W.; Fairfax, B.P.; Schramm, K.; Powell, J.E.; et al. Systematic identification of trans eQTLs as putative drivers of known disease associations. Nat. Genet. 2013, 45, 1238–1243. [Google Scholar] [CrossRef]
- Guo, S.W. Epigenetics of endometriosis. Mol. Hum. Reprod. 2009, 15, 587–607. [Google Scholar] [CrossRef]
- Wu, Y.; Starzinski-Powitz, A.; Guo, S.W. Trichostatin A, a histone deacetylase inhibitor, attenuates invasiveness and reactivates E-cadherin expression in immortalized endometriotic cells. Reprod. Sci. 2007, 14, 374–382. [Google Scholar] [CrossRef] [PubMed]
- Jenuwein, T.; Allis, C.D. Translating the histone code. Science 2001, 293, 1074–1080. [Google Scholar] [CrossRef] [PubMed]
- Bernstein, B.E.; Meissner, A.; Lander, E.S. The mammalian epigenome. Cell 2007, 128, 669–681. [Google Scholar] [CrossRef] [PubMed]
- Sun, H.S.; Hsiao, K.Y.; Hsu, C.C.; Wu, M.H.; Tsai, S.J. Transactivation of steroidogenic acute regulatory protein in human endometriotic stromalcells is mediated by the prostaglandin EP2 receptor. Endocrinology 2003, 144, 3934–3942. [Google Scholar] [CrossRef] [PubMed]
| Clinical and Personal Data of the Study Population | |||
|---|---|---|---|
| Patients | Total (n = 72) | Endometriosis (n = 41) | No Endometriosis (n = 31) |
| Age (median, interval) | 35 (20–68) | 33 (20–45) | 36 (26–68) |
| BMI (median, interval) | 21 (15–30) | 21 (17–30) | 22 (15–30) |
| Comorbidity | 18 | 9 | 9 |
| Parity | 0.25 | 2.3 | |
| Stage | I | II | III | IV |
|---|---|---|---|---|
| Number of patients | 6 | 5 | 14 | 16 |
| Age (median) | 35 | 22 | 37 | 31 |
| Parity (number of patients with at least one pregnancy) | 0 | 1 | 5 | 2 |
| BMI (median) | 21 | 20 | 20 | 20 |
| Number of patients with previous surgical intervention for endometriosis | 0 | 2 | 10 | 14 |
| Dysmenorrhea | ||||
| Mild | 2 | 0 | 0 | 1 |
| Moderate | 1 | 2 | 6 | 1 |
| Severe | 2 | 3 | 7 | 14 |
| Chronic pelvic pain | ||||
| Mild | 0 | 1 | 2 | 0 |
| Moderate | 2 | 2 | 3 | 6 |
| Severe | 0 | 0 | 3 | 6 |
| Dyspareunia | ||||
| Mild | 2 | 1 | 4 | 2 |
| Moderate | 1 | 2 | 2 | 6 |
| Severe | 0 | 1 | 3 | 6 |
| Intestinal symptoms | 2 | 0 | 7 | 10 |
| Urinary symptoms | 2 | 0 | 3 | 3 |
| Severe symptomatology percentage * | 40% | 60% | 64% | 94% |
| SNP | TEST | A1 Minor Allele | A2 Major Allele | GENO | Observed (HET) | Expected (HET) | P |
|---|---|---|---|---|---|---|---|
| SNP1(FSHB)G>A | ALL | A | G | 1/7/62 | 0.1 | 0.1203 | 0.2396 |
| SNP2(VEZT)A>C | ALL | C | A | 8/27/32 | 0.403 | 0.4358 | 0.5766 |
| SNP3(WNT4)C>A | ALL | A | C | 0/10/55 | 0.1538 | 0.142 | 1 |
| SNP | TEST | A1 Minor Allele | A2 Major Allele | GENO | Observed (HET) | Expected (HET) | P |
|---|---|---|---|---|---|---|---|
| SNP1(FSHB)G>A | AFF | A | G | 1/4/34 | 0.1026 | 0.142 | 0.187 |
| SNP1(FSHB)G>A | UNAFF | A | G | 0/3/28 | 0.09677 | 0.09209 | 1 |
| SNP2(VEZT)A>C | AFF | C | A | 1/19/18 | 0.5 | 0.3999 | 0.2255 |
| SNP2(VEZT)A>C | UNAFF | C | A | 7/8/14 | 0.2759 | 0.4709 | 0.0435 |
| SNP3(WNT4)C>A | AFF | A | C | 0/7/27 | 0.2059 | 0.1847 | 1 |
| SNP3(WNT4)C>A | UNAFF | A | C | 0/3/28 | 0.09677 | 0.09209 | 1 |
| Genotipic Test | Controls | Cases | ||||||
| N° | % | N° | % | P Value | Odds Ratio | 95% CI | ||
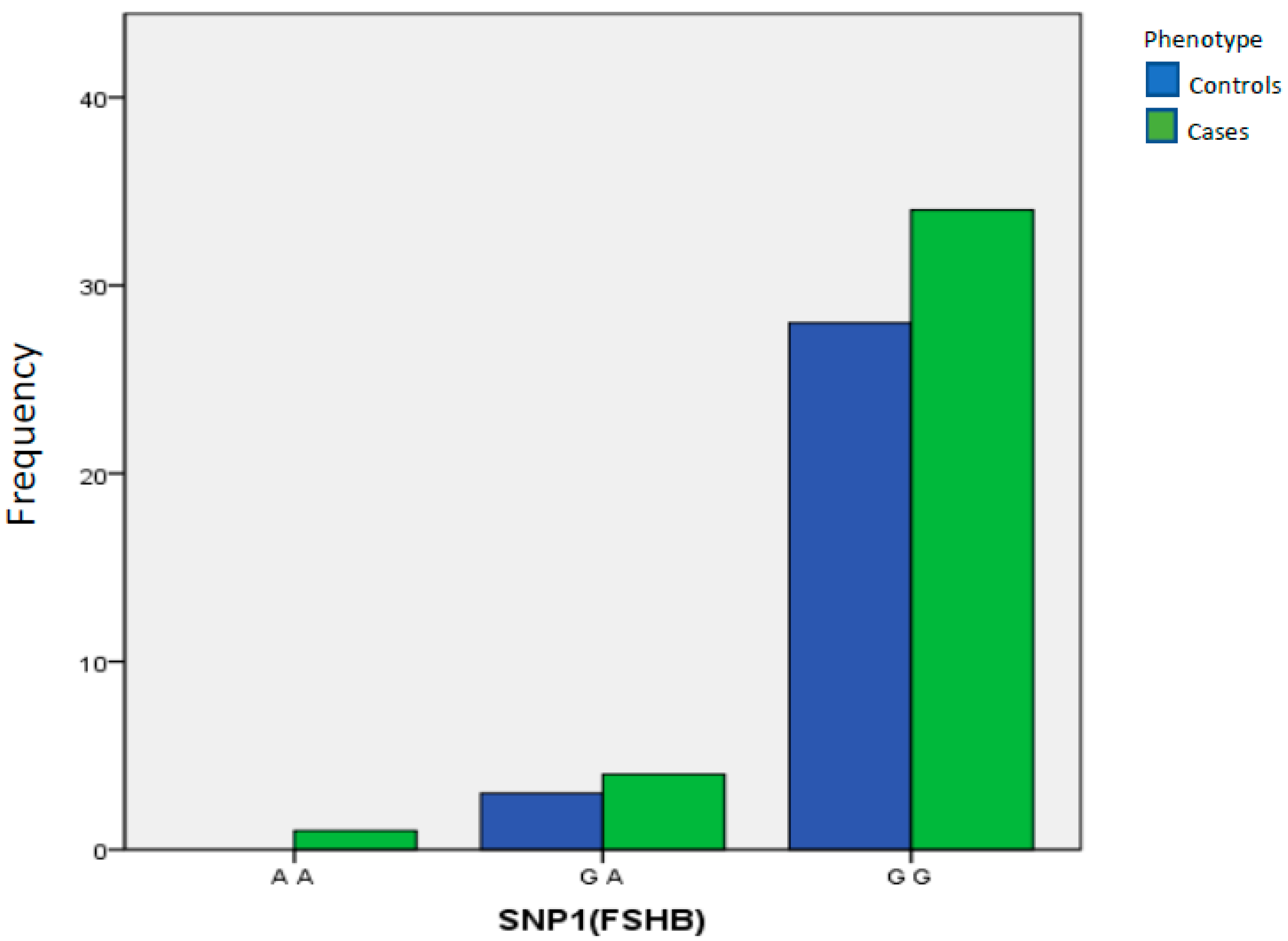
| SNP1(FSHB) | A A | 0 | 0.000% | 1 | 2.56% | 1.00 Reference | ||
| G A | 3 | 9.68% | 4 | 10.26% | >0.9999 | 0.000 | 0.000 to 15.000 | |
| G G | 28 | 90.32% | 34 | 87.18% | >0.9999 | 0.91 | 0.215 to 3.631 | |
| Total | 31 | 100.00% | 39 | 100.00% | ||||
| Allelic Test | Controls | Cases | ||||||
| N° | % | N° | % | P Value | Odds Ratio | 95% CI | ||
| SNP1(FSHB) | A | 3 | 4.84% | 6 | 7.69% | 0.7311 | 1639 | 0.393 to 683 |
| G | 59 | 95.16% | 72 | 92.30% | ||||
| Total | 62 | 100.00% | 78 | 100.00% | ||||
| Genotipic Test | Controls | Cases | ||||||
| N° | % | N° | % | P Value | Odds Ratio | 95% CI | ||
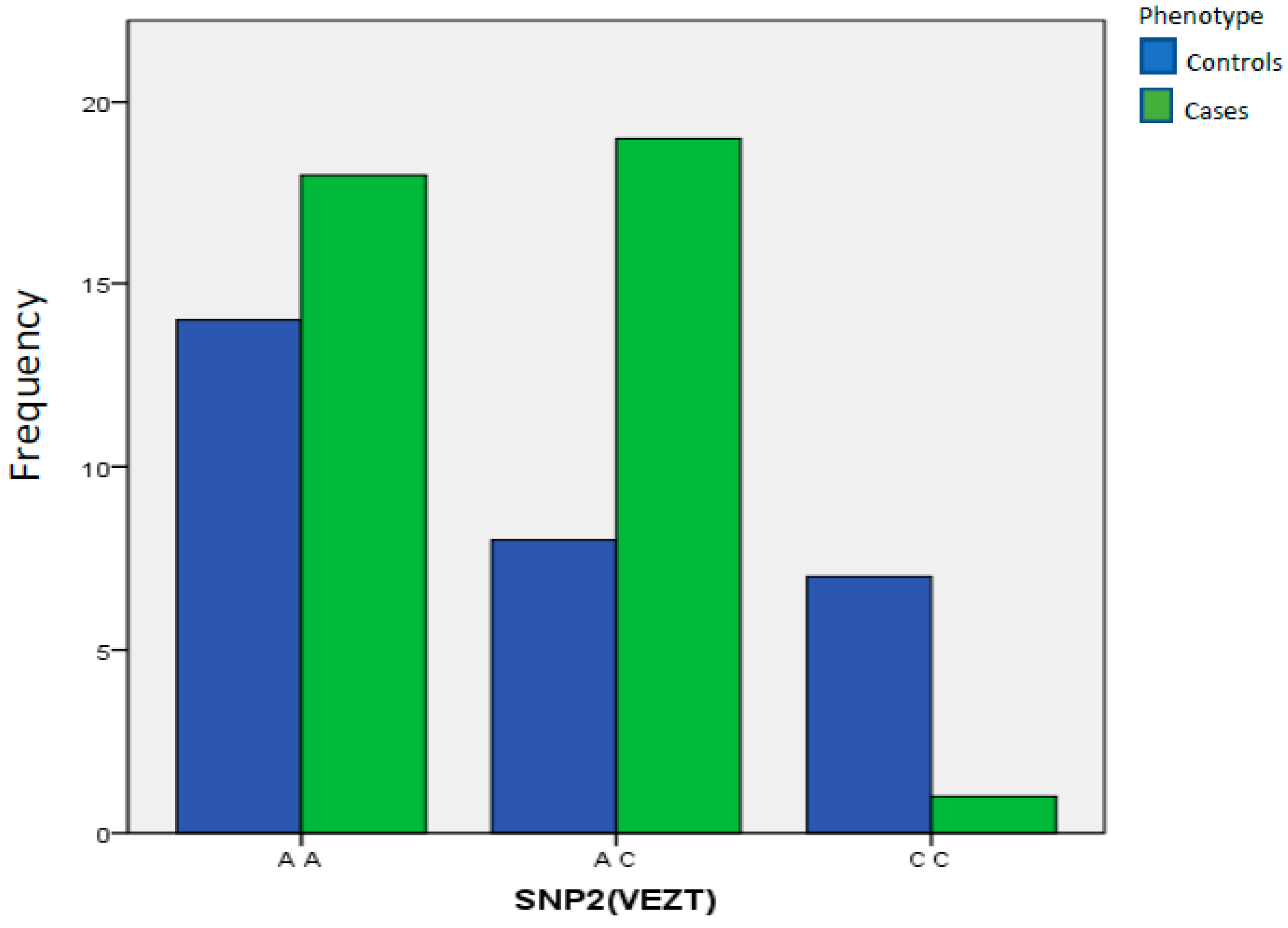
| SNP2(VEZT) | A A | 14 | 48.28% | 18 | 47.37% | 1.00 Reference | ||
| A C | 8 | 27.59% | 19 | 50.00% | 0.2931 | 1847.000 | 0.662 to 5.08 | |
| C C | 7 | 24.14% | 1 | 2.63% | 0.0111 | 0.06015 | 0.005 to 0.501 | |
| Total | 29 | 100.00% | 38 | 100.00% | ||||
| Allelic Test | Controls | Cases | ||||||
| N° | % | N° | % | P Value | Odds Ratio | 95% CI | ||
| SNP2(VEZT) | C | 22 | 37.93% | 21 | 27.63% | 0.2627 | 0.6248 | 0.300 to 129 |
| A | 36 | 62.07% | 55 | 72.36% | ||||
| Total | 58 | 100.00% | 76 | 100.00% | ||||
| Genotipic Test | Controls | Cases | ||||||
| N° | % | N° | % | P Value | Odds Ratio | 95% CI | ||
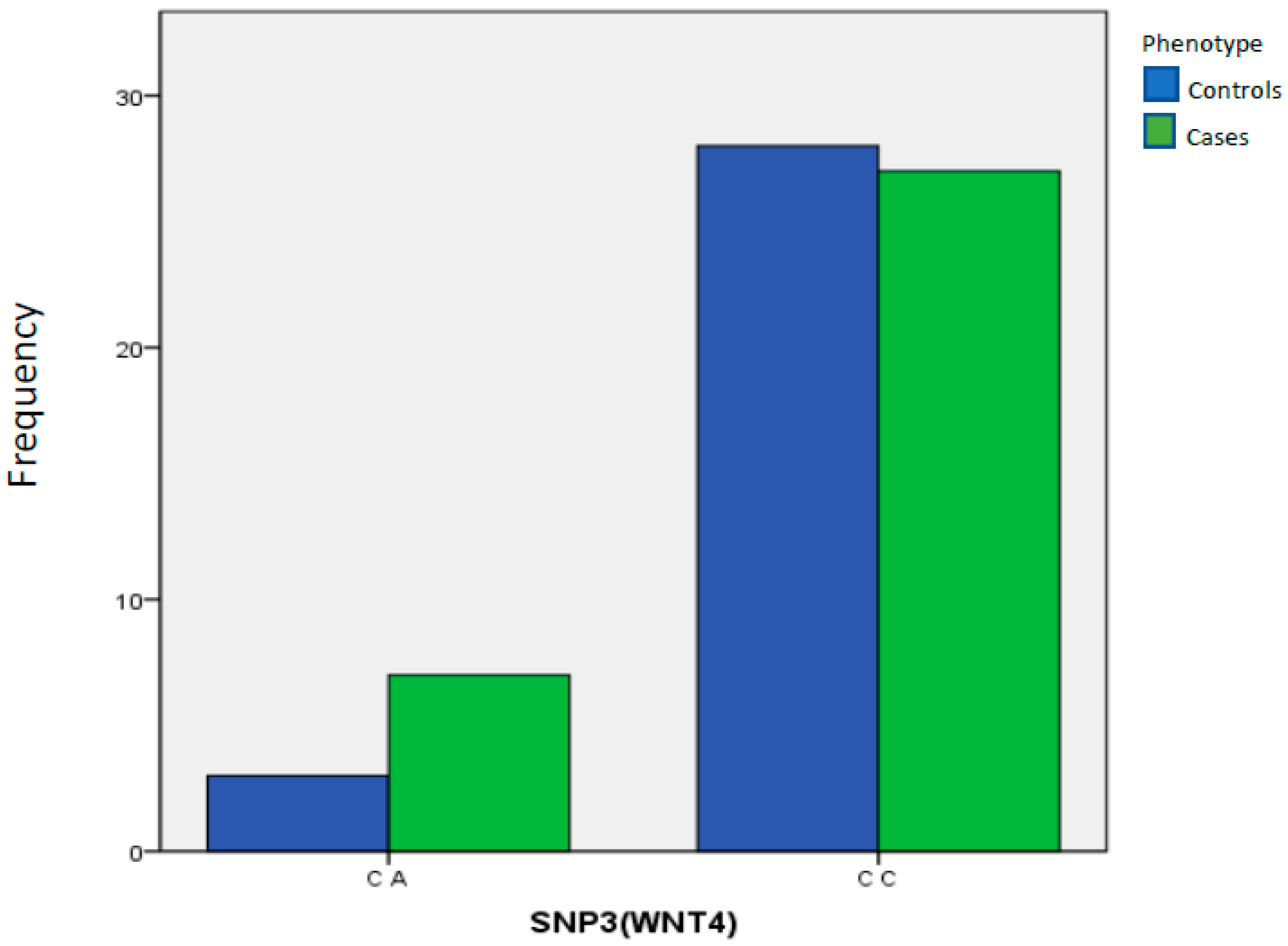
| SNP3(WNT4) | C A | 3 | 9.68% | 7 | 20.59% | 1.00 Reference | ||
| C C | 28 | 90.32% | 27 | 79.41% | 0.3088 | 0.4133 | 0.108 to 1.811 | |
| Total | 31 | 100.00% | 34 | 100.00% | ||||
| Allelic Test | Controls | Cases | ||||||
| N° | % | N° | % | P Value | Odds Ratio | 95% CI | ||
| SNP3(WNT4) | A | 3 | 4.84% | 7 | 10.29% | 0.3297 | 2257 | 0.557 to 914 |
| C | 59 | 95.16% | 61 | 89.71% | ||||
| Total | 62 | 100.00% | 68 | 100.00% | ||||
| Gene | SNP | MAF Sardinia |
|---|---|---|
| WNT4 | rs7521902 | 0.0971 |
| VEZT | rs10859871 | 0.407 |
| FSHB | rs11031006 | 0.106 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Angioni, S.; D’Alterio, M.N.; Coiana, A.; Anni, F.; Gessa, S.; Deiana, D. Genetic Characterization of Endometriosis Patients: Review of the Literature and a Prospective Cohort Study on a Mediterranean Population. Int. J. Mol. Sci. 2020, 21, 1765. https://doi.org/10.3390/ijms21051765
Angioni S, D’Alterio MN, Coiana A, Anni F, Gessa S, Deiana D. Genetic Characterization of Endometriosis Patients: Review of the Literature and a Prospective Cohort Study on a Mediterranean Population. International Journal of Molecular Sciences. 2020; 21(5):1765. https://doi.org/10.3390/ijms21051765
Chicago/Turabian StyleAngioni, Stefano, Maurizio Nicola D’Alterio, Alessandra Coiana, Franco Anni, Stefano Gessa, and Danilo Deiana. 2020. "Genetic Characterization of Endometriosis Patients: Review of the Literature and a Prospective Cohort Study on a Mediterranean Population" International Journal of Molecular Sciences 21, no. 5: 1765. https://doi.org/10.3390/ijms21051765
APA StyleAngioni, S., D’Alterio, M. N., Coiana, A., Anni, F., Gessa, S., & Deiana, D. (2020). Genetic Characterization of Endometriosis Patients: Review of the Literature and a Prospective Cohort Study on a Mediterranean Population. International Journal of Molecular Sciences, 21(5), 1765. https://doi.org/10.3390/ijms21051765