1. Introduction
Light is an essential energy source for all plant life and is used to generate a usable carbon-based energy source for growth via photosynthesis, however, when light intensities exceed the plant’s saturated photosynthesis requirements it has the potential to cause damage to the plant [
1]. This potential damage is driven by an overproduction of reactive oxygen species (ROS). The oxygen concentration in chloroplasts is governed by the activity of photosystem II (PSII) and the generation of ROS from the oxygen generated by PSII, e.g., H
2O
2, superoxide(O
2−), hydroxyl radicals, and
1O
2* in chloroplasts is closely associated with exposure to excess light [
2]. High-light intensities lead to an increase in ROS which, if not scavenged, will result in photo-oxidative damage and photoinhibition [
3]. Photoinhibition in turn results in the degradation of carotenoids, bleaching of chlorophylls, and an increase in lipid peroxidation driven by the reactive-oxygen derivatives [
4]. Photosynthetic organisms must thus find a balance between the requirements for efficient light harvesting and the danger of accumulating light-induced damage when light absorption exceeds the photosynthesis capacity.
Bryophytes, as one of the earliest diverging lineages of the extant land plants, are believed to have faced tremendous challenges in occupying a variety of ecological niches many of which were characterized by water deficits, high-light intensities, and increased exposure to UV radiation. In meeting these challenges bryophytes have evolved an efficient photosynthetic apparatus and photoprotection mechanisms. The drought-tolerant mosses,
Rhytidium rugosum and
Ceratodon purpureus, have demonstrably improved photosystem II (PSII) chlorophyll fluorescence quenching when dehydrated that is hypothesized to provide a means to dissipate excess excitation energy as heat [
5]. The desert moss,
Pterygoneurum lamellatum, exhibits damage to the photosynthetic apparatus that directly correlates to the rate of desiccation; F
v/F
mvalues measured for gametophores dried slowly were similar to undried controls but were significantly reduced in plants dried quickly [
6].
Syntrichia caninervis and
Bryum argenteum, as predominant moss species of the Gobi Desert [
7], rapidly reassemble thylakoid proteins during rehydration after a desiccation event, presumably as a means to recover photosynthesis and growth [
8]. These mosses also quickly increase transcript abundance for genes involved in the light reactions, photorespiration, and other photosynthesis-related genes [
9,
10], which presumably speeds the recovery of photosynthesis and carbon assimilation [
11]. In
Syntrichia caninervis, the increase in transcript abundance for genes involved in photosynthesis accompanies the rapid restoration and reorganization of PSII and increased chlorophyll synthesis [
12].
To maintain normal function under high-light stress conditions plants induce light-stress proteins, the Early Light-Inducible Proteins (ELIPs), as part of the protective response [
13]. ELIPs were initially described as proteins that were transiently expressed during greening of etiolated seedlings [
14,
15]. ELIPs are members of a closely related group of proteins that share highly conserved regions of homology: The three-helix
ELIPs, the two-helix
SEPs (stress-enhanced proteins), and the one-helix proteins (
OHPs) along with
HLIPs (high-light induced proteins), and
SCPs (small Cab-like proteins) [
16]. The
Arabidopsis genome contains two
ELIP genes,
Elip1 and
Elip2, both of which, along with light-induced greening, have been implicated in the control of seed germination, particularly when germination occurs under abiotic stress conditions [
17]. ELIPs were also induced in mature green plants exposed to light stress in
Arabidopsis [
18] and are thought to play a role in photoprotection by transiently binding free chlorophyll preventing photo oxidative damage and facilitating energy dissipation that protects the PS-reaction center from photoinhibition [
19].
The environmental responsiveness of ELIPs, as measured by transcript abundance, has been explored in many plants, primarily in angiosperms.
ELIP cDNA clones have been constructed from RNA isolated from the alga
D. salina [
20], the bryophytes
S. ruralis [
21], and
Physcomitrella patens [
22], as well as a number of tracheophytes, including pea [
23,
24], barley [
25],
Arabidopsis [
26], wheat [
27], and the resurrection plant,
Craterostigma plantagineum [
28]. In addition to cases mentioned earlier, high-light stress induced accumulation of
ELIP transcripts has been reported for pea [
23,
24], wheat [
27], barley [
25,
29], grapevine [
30], and
Arabidopsis [
26]. Low temperature can also induce
ELIP gene transcript accumulation in pea [
24], barely [
25], cabbage [
31], and
Medicago sativa [
32]. Salt stress can induce
ELIP gene transcript accumulation in cabbage [
31],
Dunaliella salina [
20], and
Syntrichia ruralis [
21]. In
Madicago sativa [
32],
Arabidopsis [
18], and
S. ruralis [
21] the
ELIP transcript accumulates during dehydration. In all cases, the implication is that ELIPs play a major role in abiotic stress tolerance. This suggested role of ELIPs in abiotic stress tolerance mechanisms was recently strengthened by the observation that tandem proliferation of ELIP genes, coupled with the elevation of ELIP transcript abundance, is a distinctive feature of the genomes of resurrection plants [
33].
In this report, we investigated ELIP transcript abundance in S. caninervis, a desiccation-tolerant moss, exposed to different light conditions. The aim was to provide further insights into the ability of ScELIPs to photo-protect photosynthetic capacity under light-stress conditions and to determine if light quality also impacted ELIP transcript abundance. In this study, we also investigated the functional ability of the ScELIPs to protect the photosynthetic machinery using an ectopic expression approach. The cDNA sequences were used to generate transgenic Arabidopsis Atelip mutants (lacking one of the AtELIP genes (AtELIP2) constitutively expressing the ScELIPs transcripts with the aim of assessing the ability of the ScELIPs to complement the Atelip mutant phenotype under different light treatments in different growth phases.
3. Discussion
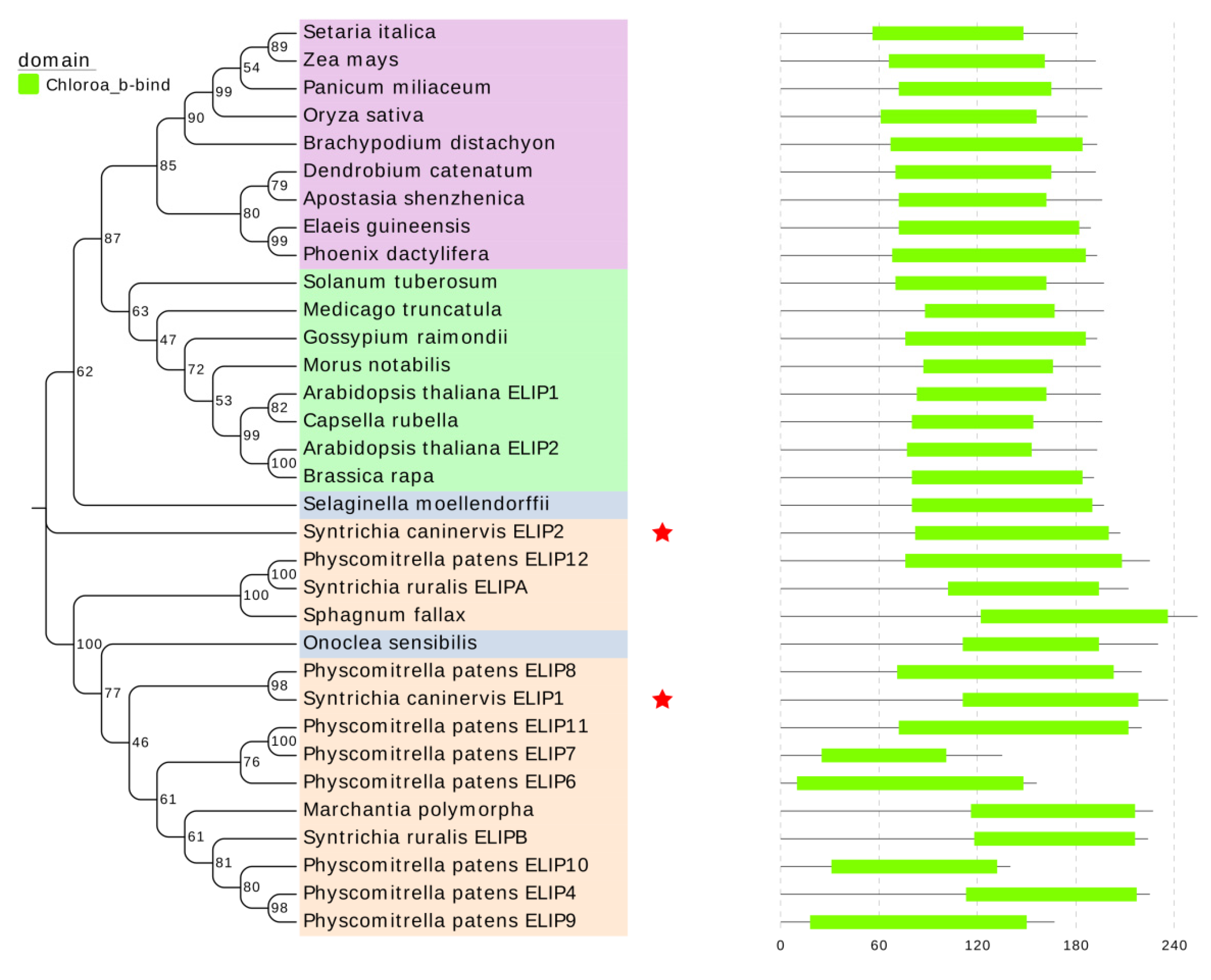
In this study, we identified and characterized two
ELIP genes from the
S. caninervis transcriptome.
ScELIP1 has close primary amino acid sequence similarity to the
ELIPs of the non-vascular plants
Physcomitrella patens,
Sphagnum fallax, and
Syntrichia ruralis [
21] excepting one kind of fern (
Onoclea sensibilis).
ScELIP2 that is more similar to the
ELIPs of vascular plants such as
Arabidopsis [
26],
Zea mays [
35], etc. (
Figure 1). According to the amino acid sequence similarity of
ELIPs, bryophytes ELIPs were clearly separated from monocots and eudicots
ELIPs, because bryophytes are a special evolutionary group. Neither of the
ScELIP sequences fully corresponds to either of the two
Arabidopsis ELIP genes which, by ectopically expressing the
ScELIP genes independently in the
Arabidopsis Atelip mutant background, allowed us to evaluate both the possible activity of the
ScELIP transcripts in photoprotection of chloroplasts in situ but also to determine if the moss ELIPs could rescue the
Atelip phenotype resulting from the loss of
AtELIP2.In
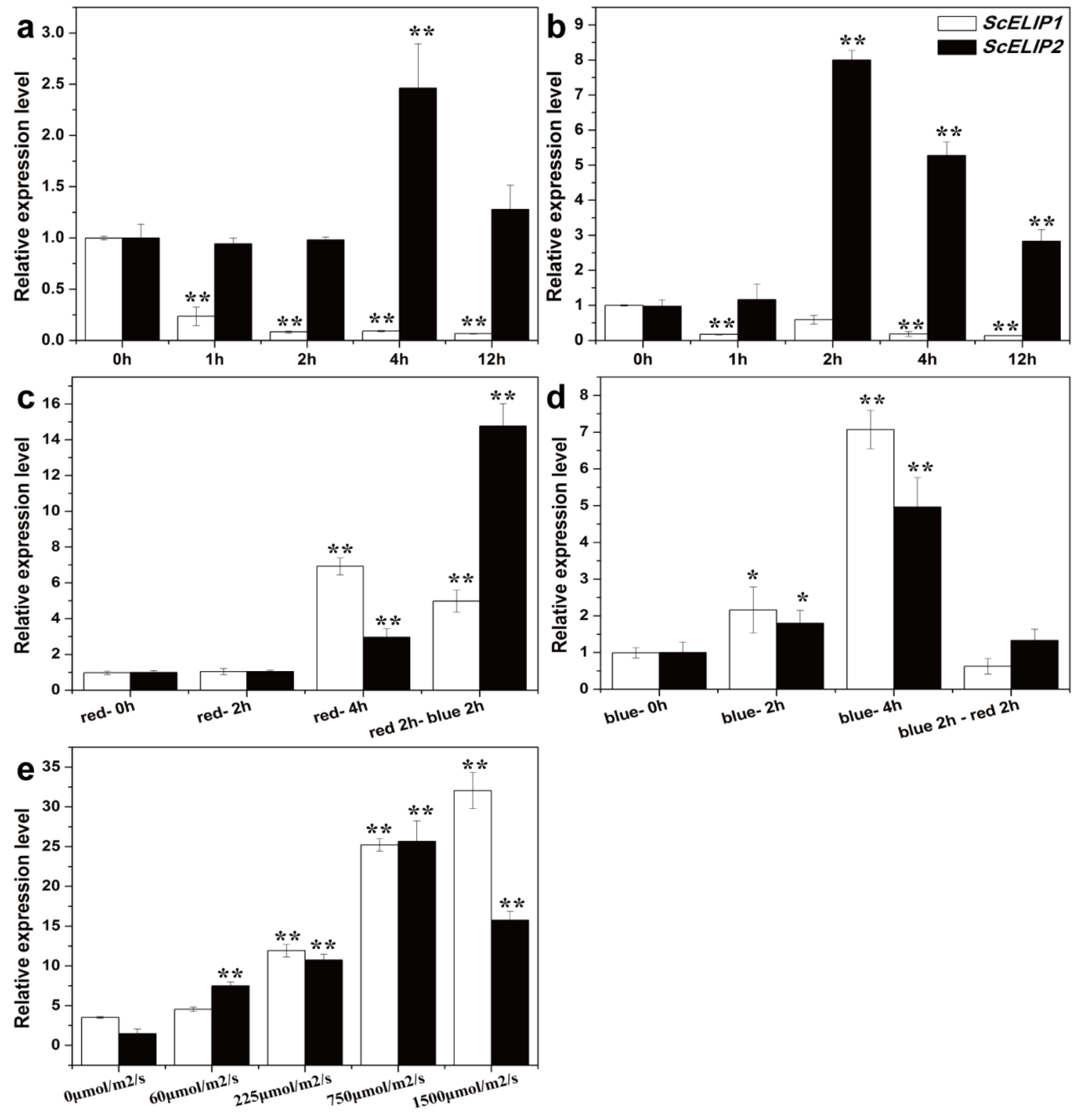
Syntrichia caninervis ScELIP1 and
ScELIP2 transcripts abundance were elevated by different light stresses (
Figure 2).
ScELIP1 and
ScELIP2 transcripts exhibit similar responses, both exhibit greater accumulation in response to blue and high-light exposure and both exhibit transcript accumulation only after a 4 h exposure to red-light, more evident for
ScELIP1. Our understanding of how or why
ELIPs respond to different light qualities is still very limited, but in pea seedlings
ELIPs transcripts are specifically induced by blue-light but not by red-light, and the accumulation of blue-light induced
ScELIPs transcripts are significantly repressed by low intensity red-light [
23]. These findings are consistent with the
S. caninervis data presented here with the exception that prolonged exposure to red-light alone does result in an elevation
ScELIP transcript abundance. The two
Arabidopsis ELIP genes can be induced in dark grown
Arabidopsis exposed to illumination by high intensities of both red and blue-light [
18] which again are consistent with observations. In
Arabidopsis,
ELIP transcript abundance was elevated by UVA but not by UVB [
36]. In
S. caninervis,
ScELIP1 transcripts declined in abundance in response to UVA and
ScELIP2 transcript levels only exhibited a response to UVA after a 4 h exposure (
Figure 2a). UVB exposure caused a general decline in
ScELIP1 transcripts but generated an increase in abundance of ScELIP2 transcripts after 2 h of exposure. Clearly, the
S. caninervis ELIP response to UV light differs from that seen in
Arabidopsis, perhaps indicating a more complex role in UV protection mechanisms in the moss. However, this possibility must be tempered by differences in experimental protocols and by the distinct possibility that the moss genome contains more than two ELIP genes as would be suggested by the observation that the genomes of desiccation tolerant plants are rich in ELIP genes [
33].
It is clear from our analyses that ELIP transcript abundance in
S. caninervis is altered by light quality (UVA and B, red and blue-light), suggesting that the promoters that control the transcription of
ELIP genes contain elements that respond to light signal transduction pathways. At the present time, the promoter sequences for the
S. caninervis ELIP genes are unavailable but ELIP promoters from other species have been investigated. In
Ginkgo biloba, high-light defense and stress responsive elements were delineated in the promoter region of
GbELIP [
37]. In
Arabidopsis,
ELIP1 expression is enhanced by red, far-red, and blue-light and that phytochrome A and B are involved in the signaling pathway along with the transcription factor HY5 [
18]. The
Arabidopsis ELIP2 promoter region was later demonstrated to contain two regulatory elements, that form a regulatory unit, that regulate
ELIP2 expression in response to UVB, high-light, and cold stress [
38]. If similar elements appear, the
ScELIP promoter might add to our understanding of the evolution of light response transduction pathways in plants.
The significant and rapid elevation in the
ScELIPs transcript abundances when gametophytes are transferred from the dark to high-light is consistent with similar observations in
Arabidopsis [
39],
Hordeum vulgare [
25],
Pisum sativum [
23],
D. salina [
20], and
Syntrichia ruralis [
21] and supportive of the hypothesis that ELIPs are useful indicators of high-light stress and the initiation of photoprotection mechanisms during the early phases of photodamage [
40].
Light stress has been demonstrated to exert influence on seed germination and photomorphogenesis during the initial developmental stages of seedling growth [
17]. Our results supported the observation that light quality alters photomorphogenesis following germination (
Supplementary Figure S8) such that the hypocotyl was shortened and the cytoledons remained closed following germination under UVB light [
41]. In a recent report, UVB photoreceptor UVR8 (UV resistance locus 8) promotes rapid PIF (phytochrome interacting factors) degradation and reduces PIF abundance, which lead to UVB inhibits shade avoidance, finally, UVB inhibits stem elongation of Sharma et al. [
42]. While this phenotype was not altered by the expression of the
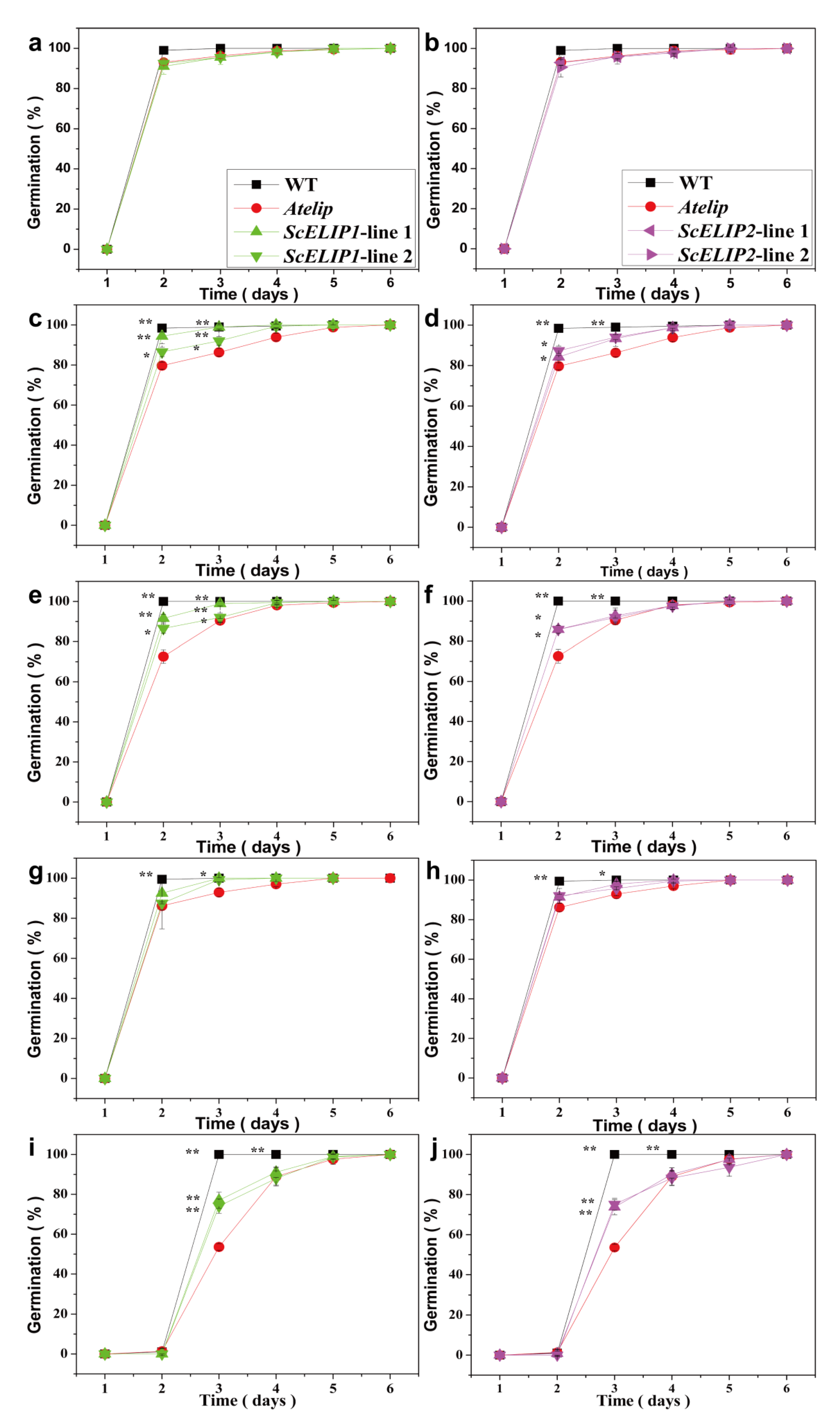
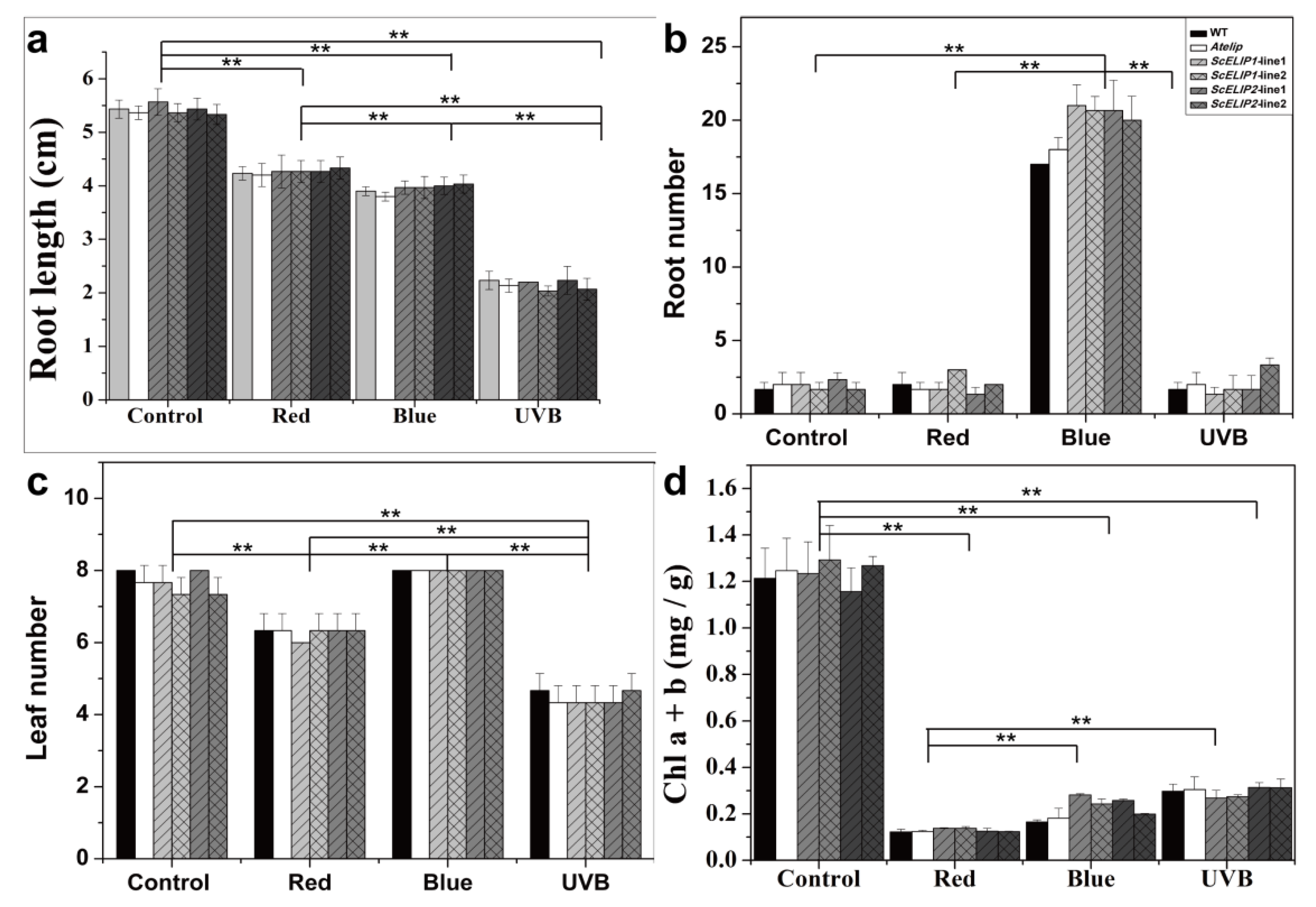
35S-ScELIPs in the transgenic seeds. The addition of the
35S-ScELIPs genes into the
Atelip mutant line significantly improved the germination rate above that of the mutant alone without altering leaf number, root number, or root length with the exception of an increase in root number in the
35S-ScELIP transgenics under blue-light (
Figure 4).
The complementation of the
35S-ScELIP genes to the
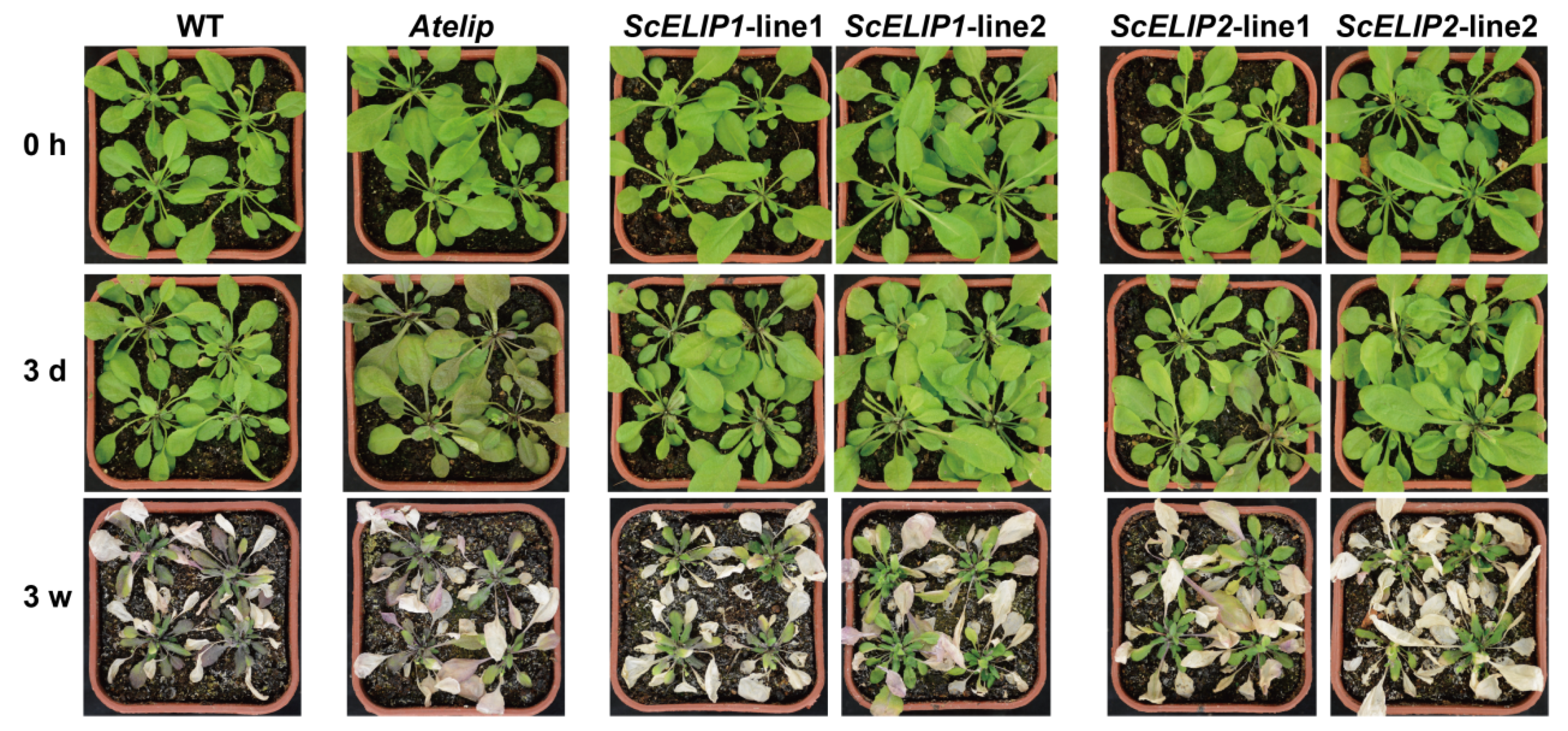
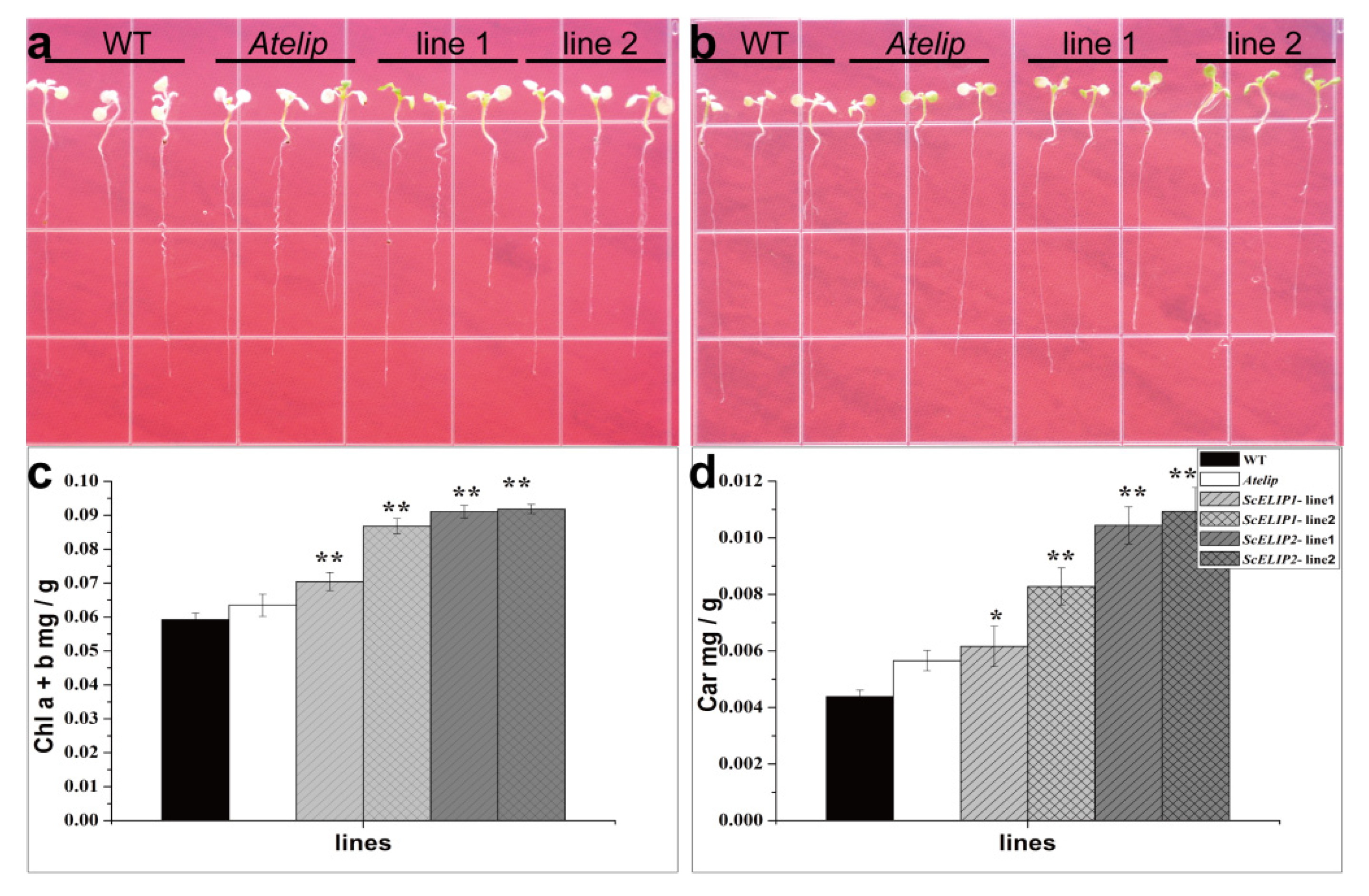
Atelip mutant line enhanced the survival rate of seedlings under high-light stress in part by minimizing chlorophyll degradation. The transgenic lines, by protecting the chlorophyll, were greener in appearance than even the Col-0 WT lines after 24 h of recovery following exposure to high-light conditions and also had a higher percent survival (
Figure 5). This was also evident in the more mature plant, where the transgenic lines exhibited significantly less photodamage under prolonged exposure to high-light intensities than the mutant line (
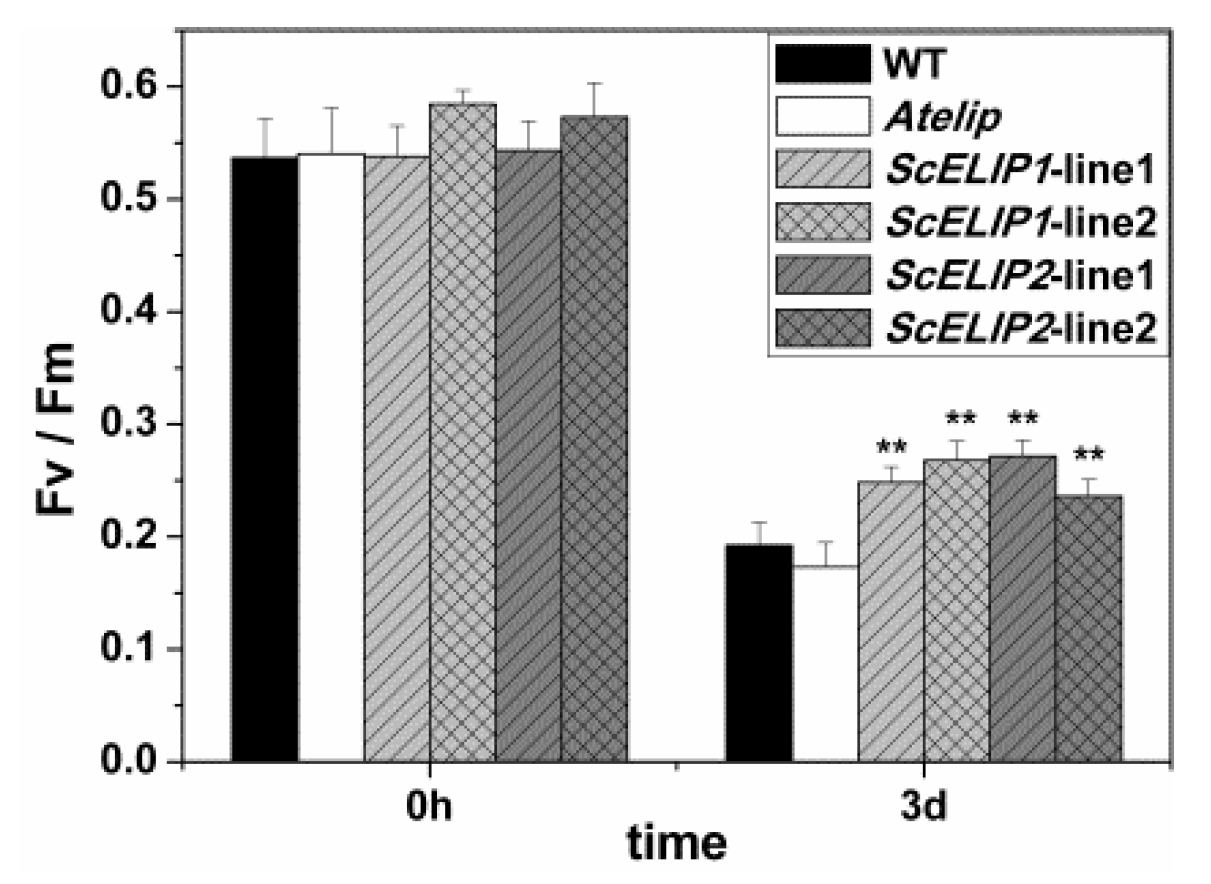
Figure 6). The improved protection to photodamage observed in the
35S-ScELIP transgenics was reflected in their improved photosynthetic performance measures compared to the
Atelip mutant increased F
v/F
m (
Figure 7). These data suggest that the
ScELIPs can protect PSII functionality of
Arabidopsis plantlets under high-light conditions. ELIP accumulation under high-light stress has been correlated with photoinhibition of PS, degradation of the D1 protein, and changes in the level of pigments in the mature plant [
36,
43]. These conclusions are consistent with our data indicating enhanced PSII protection in the
35S-ScELIP transgenics.
The results of this study indicate that the physiological function of ELIPs are related to the protection of chlorophyll, and thus enhance the protection of the chloroplasts from photooxidative stress. The
35S-ScELIP1 and
35S-ScELIP2 lines also have improved carotenoid levels than those recorded for the
Atelip mutant following high-light exposure and thus it is possible that
ScELIPs may also play a role in stabilizing the photosynthetic apparatus and is consistent with the idea that ELIP acts as a photoprotectant for PSII in the thylakoid membrane and as such would play a major role in the ability of
Syntrichia caninervis to survive in extreme conditions, as suggested by VanBuren et al. [
33], and that it regularly encounters in the Gobi Desert.
The
35S-ScELIP1 and
35S-ScELIP2 lines have a significant response to high-light (
Figure 8a,b) even though their transcription is controlled by the
35S promoter. It is possible that the overall transcription rates affected by high-light or transcript abundance under high-light is partially controlled by transcript stability. The former seems unlikely given the variability in transcript abundance for other genes in response to high-light (
Figure 8). It is possible that transcript abundance was controlled in part by the altered stability but this possibility was not investigated. It is also possible that the 35S promoter used in these constructs is affected by exposure to high-light but there are no reports of this as a possible explanation. It is intriguing that the transcript abundance profile for
ScELIP1 is more similar to the native
Arabidopsis SEP2 and the transcript abundance profile for
ScELIP2 is more similar to
OHP2 than to other photosynthetic related transcripts. If the 35S promoter, which is active in the untreated controls, is not high-light inducible then it could suggest the presence of a post-transcriptional mechanism that may control the abundance of the transcripts of three
ELIP families of protective proteins. The transcript abundance profiles in response to both short and long exposure to high-light for all of the photosynthetic related transcripts in the genotypes we investigated were complex. However, it was evident that the
Atelip mutant was, with the exception of the
SEP2 transcript which was elevated above control levels (
Figure 8f), deficient in proteins that function to protect PSII. The ectopic expression of the
ScELIP transcripts in the
Atelip mutant appears to provide the necessary protective proteins (at least the transcripts) which overcomes the high-light phenotype of the mutant. In addition, the transcript abundance profiles for all genes and genotypes indicate either a recovery of or elevation in transcript abundance after 3 weeks of exposure to high-light which may be indicative of the ability of
Arabidopsis to acclimate to this environmental stress.
4. Materials and Methods
4.1. Plant Material and Growth Conditions
Dry Syntrichia caninervis gametophytes were collected from the Gurbantunggut Desert in the Xinjiang Uygur Autonomous Region, China (44° 32′ 30″ N, 88° 6′ 42″ E) and kept in the dark in a paper sack at room temperature. Dry gametophytes were fully hydrated on filter paper saturated with MINIQ-filtered water (8 mL) in glass petri dishes for 24 h at 25 °C, with light at a photosynthetic photon flux density (PPFD) of 50 μmol/m2/s, prior to the described light treatments.
Seeds of
Arabidopsis thaliana L. cv. Columbia and the
Arabidopsis mutant
Atelip were germinated on MS medium [
44] after surface sterilization with 70% ethanol for 30 s followed by 3% sodium hypochlorite for 7 min and washed 5 times with sterile water. Seedlings were transferred to 5 cm diameter pots at 4–6 leaf stage containing autoclaved peat substrate (Pindstrup, Mosebrug) and grown under controlled conditions at 22 ± 2 °C with a 14 h light/10 h dark cycle, 60–75% relative humidity, and light at PPFD of 100 μmol/m
2/s.
A. thaliana (Col-0) and the mutant
Atelip (NASC ID: N544166)
, a T-DNA insertion mutant in the promoter region of
ELIP2 [
45], were obtained from The European Arabidopsis Stock Centre.
4.2. Bioinformatic Analysis of Syntrichia Caninervis ELIP Sequences
Publicly available
ELIP sequences of
A. thaliana and
Physcomitrella patens were used to screen the transcriptome of
S. caninervis [
10] for single copy transcripts (designated as unigenes) that were putative
ELIP coding sequences. We translated each unigene to obtain the amino acid sequence using ORF finder. All generated amino acid sequences were investigated using NCBI-blastp. The final sequences used to construct the
ScELIP clones were consistent with annotated
ELIP genes in the NCBI database. A molecular phylogeny was constructed using MEGA7.0 with the neighbor-joining method and 500 bootstrap replicates [
46] utilizing the eudicots, monocots, ferns, and bryophytes ELIP sequences obtained from public resources and presented in additional data (
Table S2). All of ELIP genes domain prediction was conducted with the Pfam database (
http://pfam.xfam.org/search). Further physiological and biochemical properties prediction for two ScELIPs was obtained on ExPASy (
https://www.expasy.org/).
4.3. Extraction of RNA and RT-qPCR Based Expression Analysis
Total RNA was isolated from 0.1 g of
S. caninervis gametophytes using the TRIzol reagent (Qiagen, Berlin, Germany) according to the manufacturer’s protocol. The cDNA was synthesized from 1 μg of total RNA using random hexamer primers with the PrimeScript RT reagent kit including gDNA Eraser (Real time; Takara, Japan) according to the manufacturer’s protocol and stored at -20 °C until use. The cDNA was diluted 10-fold with H
2O to serve as a template for Real-time PCR. Each 20 μL real-time PCR reaction contained 2 μL cDNA, 0.5 μL each of forward and reverse primers as indicated below (10 μM for each primer), and 10 μL of SYBR Premix Ex Taq
TM (Takara Biomedical Technology, Takara, Japan). Real-time quantitative PCR was performed in 96-well plates with the CFX96 Real-Time PCR Detection System (Bio-Rad, New York, NY, USA) according to the manufacturer’s instructions [
47], with three technical replicates and two biological replicates. The RT-qPCR protocol was as follows: 30 s initial denaturation at 95 °C, 40 cycles of 94 °C for 5 s, and 60 °C for 30 s. Parallel reactions to amplify
α-TUB2 were used to normalize the amount of template. The forward and reverse primers used for
ScELIP1 were 5′ GGCTATGCTGGGATTCGTGT 3′ and 5′ TTGATCTGGTTCCTGGCGTC 3′, the primers for
ScELIP2 were 5′GCAATGATCGGACTTGTCGC3′ and 5′ ACCTGCTGTGAACAATCCGT 3′. All PCR efficiencies were >95%. Transcript abundance data were collected using Bio-Rad CFX Manager.
Hydrated S. caninervis gametophytes were exposed to the following environmental treatments: A progressive series of PPFD, Ultraviolet A (UVA), Ultraviolet B (UVB), red-light, and blue-light. Fully hydrated S. caninervis gametophytes were transferred to MINIQ-filtered water saturated filter paper in clean petri dishes for all treatments. The light series treatment consisted of a 2 h exposure to either 0 μmol/m2/s, 60 μmol/m2/s, 225 μmol/m2/s, 750μmol/m2/s, or 1500 μmol/m2/s provided by a cool-white fluorescent light lamp (Philips lighting). For the ultraviolet radiation treatment gametophytes were exposed to either UVA (380 nm) or UVB (308 nm) irradiation for 0 h, 1 h, 2 h, 4 h, 12 h under a fluorescent light lamp (Philips lighting) that delivered a PPFD of 10 μmol/m2/s. In the red-light treatment, gametophytes were either exposed to red-light using a monochromatic light-emitting diode (LED, Philips lighting, Amsterdam, Netherlands) at 655 nm, and PPFD of 100 μmol/m2/s for 0, 2, 4 h, or exposed to red-light for 2 h and then transferred to blue-light for 2 h. In the blue-light treatment gametophytes were either exposed to blue-light using a monochromatic light-emitting diode (LED, Philips lighting) at 450 nm and PPFD of 100 μmol/m2/s for 0 h, 2 h, 4 h, or exposed to blue-light for 2 h and then transferred to red-light for 2 h. Each treatment consisted of three biological replicates.
4.4. Cloning of ScELIP Genes and Establishment of Transgenic Lines
The transcripts
ScELIP1 (Unigene 40121) and
ScELIP2 (Unigene 68225) were chosen from the
Syntrichia caninervis transcriptome database [
10]. The full-length coding sequence for each gene product was obtained by RT-PCR using the cDNA previously constructed for the RT-qPCR based expression analysis. The RT-PCR was performed using gene-specific primers: For
ScELIP1 the forward and reverse primers were 5′ GGGGTACCC ATGGCAGCGATGGCG 3′ and 5′ CGAGATCTCGCTAGACAGGGAAGCG 3′, and these primers include inserted
KpnI and
XbalI enzyme sequences. For
ScELIP2, the forward and reverse primers were 5′ CGGGATCCATGGCGATGACTTT 3′ and 5′ CGAGATCTCGTTATACAAGTGGGC 3′, and these primers include
Bam HI and
XbalI site sequences. The PCR product was extracted from agarose gel electrophoresis (0.8% agarose, 5 v/cm voltage for 20 min) using the Gel Extraction Kit (OMEGA, St. Louis, MI, USA), and 4 μL PCR extract production, 1 μL pMD 19-T vector, 1 μL T4 DNA ligase, 2 μL 10× ligation buffer, and 12 μL H
2O for 2 h (TaKaRa, Japan), linking the product with pMD 19-T vector. The sequences were validated by sanger sequencing at the Beijing Genomics Institute (BGI) to ensure that there were no PCR introduced mutations into the coding sequences.
4.5. Transformation of A. Thaliana
The
ScELIP1 ORF was removed from
ScELIP1-19T by digestion with
KpnI and
XbalI and cloned into
KpnI and
XbalI—digested pCAMBIA 1301 to construct a 35S-
ScELIP1 gene (
Figure S2). Likewise, the
ScELIP2 ORF was removed from
ScELIP1-19T by digestion with
Bam HI and
XbalI and cloned into
BamHI and
XbalI—digested pCAMBIA 1301 to construct a 35S-
ScELIP2 gene (
Figure S3). The p
ScELIP1-CAMBIA 1301 and p
ScELIP2-CAMBIA 1301 were introduced separately into an
Agrobacterium tumefaciens strain EHA105 by triparental mating as described by Hoekema et al. [
48] and introduced into 6-week old
Arabidopsis ELIP mutant
Atelip plants by the floral dip method [
49]. T1 transgenic plants were selected by germination on MS agar plates containing Hygromycin (80 mg/L). After 7 days, the majority of the seedlings turn yellow or white and die. After 10 days, those seedlings that continued to stay green and grow were considered as stable transformants. The stable transformants were verified by PCR by using the same gene-specific primers used to isolate the coding sequences for each
ScELIP. Primary transformants were shelfed to generate T2 homozygous transgenic seed stocks for production of T3 homozygous plants used in the phenotypic analysis.
4.6. Seed Germination Assay
Petri plates containing 1/2 MS agar media (1%) were divided into four quadrants and approximately 50 seeds were sown of each genotype into separate quadrants: Arabidopsis Col-0 wildtype (WT), the Atelip mutant, and two transgenic Arabidopsis lines of ScELIP1 or ScELIP2. Seed germination, when the radicle protruded through the seed coat, was scored from the first day after sowing. The seed germination assay was conducted under standard conditions (22 ± 2 °C, 14 h light/ 10 h dark cycle, 60–75% relative humidity, and PPFD of 100 μmol/m2/s) as a control or under the following treatments; irradiated by red, blue, UVB, or high-light (1000 μmol/m2/s) as their single light source (as mentioned in 2.3 Extraction of RNA and RT-qPCR based expression analysis) at 22 °C 14 h light/10 h dark cycle for 6 days.
4.7. Phenotypic Analysis of Transgenic A. Thaliana
Sterile WT, Atelip mutant, and ScELIPs transgenic Arabidopsis seeds were vernalized for 2 d and germinated on 1/2 MS agar medium under control conditions of 22 °C and 14 h light/10 h dark cycle and PPFD of 100 μmol/m2/s. Seven-day old seedlings, of similar size, were chosen for exposure to experimental treatments. WT, Atelip mutant, and transgenic ScELIPs lines, were cultured on 1/2 MS medium for 7 days at 22 °C 14 h light/10 h dark cycle. Individual light treatments were administered for 7 days at 22 °C 14 h light/10 h dark cycle and PPFD of 100 μmol/m2/s under the appropriate light source of either red, blue, or UVB as described for the gene expression study. After all treatments the number of leaves, roots, and the length of the longest root were recorded.
For the high-light treatment, seven-day-old seedling, grown under control conditions of 22 °C and 14 h light/10 h dark cycle and PPFD of 100 μmol/m2/s, were briefly exposed to light at an intensity of 1500 μmol/m2/s for 2 h and subsequently transferred to normal conditions for 24 h to recover. Following recovery, the seedlings were assessed for chlorophyll content as described in a later section.
4.8. High-Light Treatment of Plantlets
Seedlings of WT, Atelip mutant, and ScELIPs transgenic lines of Arabidopsis were grown in 5 cm diameter pots in a peat substrate (Pindstrup, Mosebrug, Ryomgård, Denmark) and grown under well-watered controlled conditions at 22 ± 2 °C with a 14 h light/ 10 h dark cycle, PPFD of 100 μmol/m2/s, and 60–75% relative humidity. After four weeks plants were irradiated with high-light (1000 μmol/m2/s) at 22 °C 14 h light/10 h dark cycle under well-watered conditions. Fv/Fm was measured before treatment (0 h), after exposure to high-light for 3 days and then continuously for 3 weeks. Leaf samples were taken for chlorophyll and carotenoid analyses before and after 3 days and 3 weeks of high-light treatment.
The relative transcript abundance was assessed for
ScELIP1,
ScELIP2, One-Helix Protein (
OHP),
OHP2, two-helix protein (
SEP2), PSII CP29 antenna protein (
LHCB4.2), PSI antenna protein (
LHCA2), PSII CP22 protein (
PSBS), and PSII D2 protein (
PSBD), after exposure to high-light for 0 h and 2 h, 3 days and 3 weeks. Each treatment consisted of 3 biological replicates. The method of extraction of RNA and RT-qPCR based expression analysis was as stated previously. To compare relative expression levels, Col-0 WT and all
ScELIPs transgenic lines of
Arabidopsis were compared to the
Atelip mutant, and
Atα-TUB was used as a reference gene; primer sequences are listed in
Table S3.
4.9. Pigment Analysis
Chlorophyll content was measured according to the methods described by Ritchie 2006 [
50]. Leaf samples (50 mg Fwt) were collected from three separate plants for each line and at each treatment phase. Pigments were extracted by incubation of the leaf samples in 2 mL of 96% ethanol (room temperature ~25 °C) for 4 h in darkness with constant agitation. The extracts were centrifuged at 10,000 rpm for 2 min and the supernatants removed for analysis. Supernatants were analyzed spectrophotometrically at wavelengths of 470, 649, and 665 nm using a UV–Visible spectrophotometer (Biomate 3S, Thermo Fisher Scientific, Waltham, MA, USA). The concentration of the chlorophyll a, b, and total carotenoids were determined using the following equations: Chl a = 13.95 × OD
665 − 6.88 × OD
649, Chl b = 24.96 × OD
649 − 7.32 × OD
665, and Car = (1000 × OD
470-2.05 × Chl a-114.8 × Chl b)/245. The total pigment content in mg/g =N × C × V/W, “N” represents dilution ratio, “C” represents pigments concentration (mg/mL), “V” represents the volume of extracting solution (mL), “W” represents sample fresh weight (g).
4.10. Fluorometric Assessment of Photosynthetic Performance
Photosynthetic performance of
S. caninervis at a different light intensity and times were assessed by pulse amplitude modulated fluorometry using a portable chlorophyll fluorometer (PAM 2500) (Heinz, Walz, Berlin, Germany). Measurements of chlorophyll fluorescence were recorded in situ on the mid portion of the uppermost fully mature leaf. The saturation pulse method was used to calculate the F
v/F
m, the parameter settings were based on the recommendations of Zhang et al. [
51]. All parameters were measured on 6 individual plants and the values were averaged for each genotype.
4.11. Statistical Analyses
All statistical analyses were performed using Statistical Product and Service Solutions (SPSS) 16.0 software (SPSS Inc., Chicago, IL, USA). Data were compared using one-way ANOVA, Dunnett’s T3 was used to examine the difference in significance of ANOVA results, and values were considered as significantly statistically different at p < 0.05, or distinctly statistically different at p < 0.01. Error bars represent standard deviations.