Transcriptome Analysis Reveals the Different Response to Toxic Stress in Rootstock Grafted and Non-Grafted Cucumber Seedlings
Abstract
1. Introduction
2. Results
2.1. Plant Morphology and Biomass
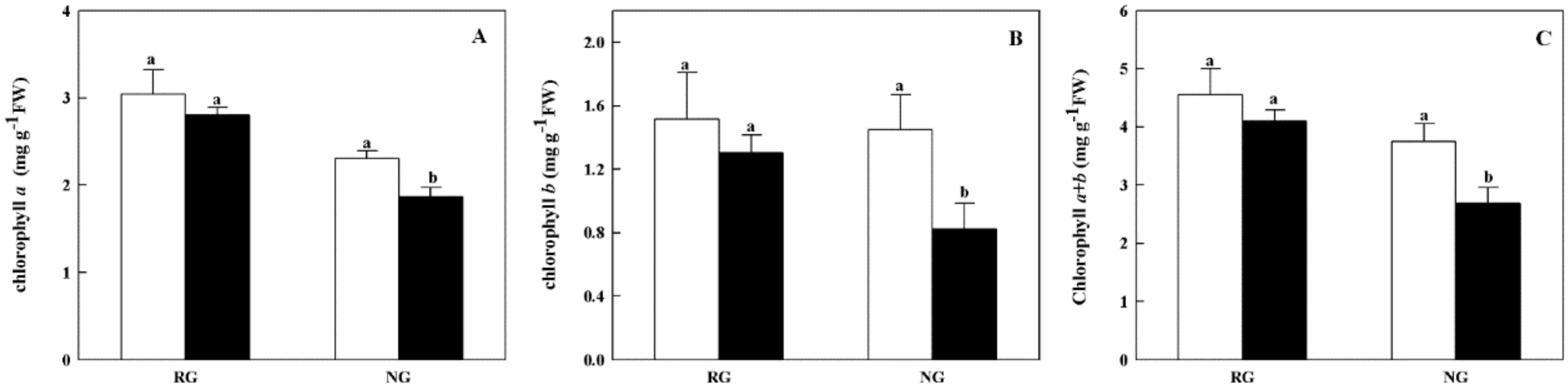
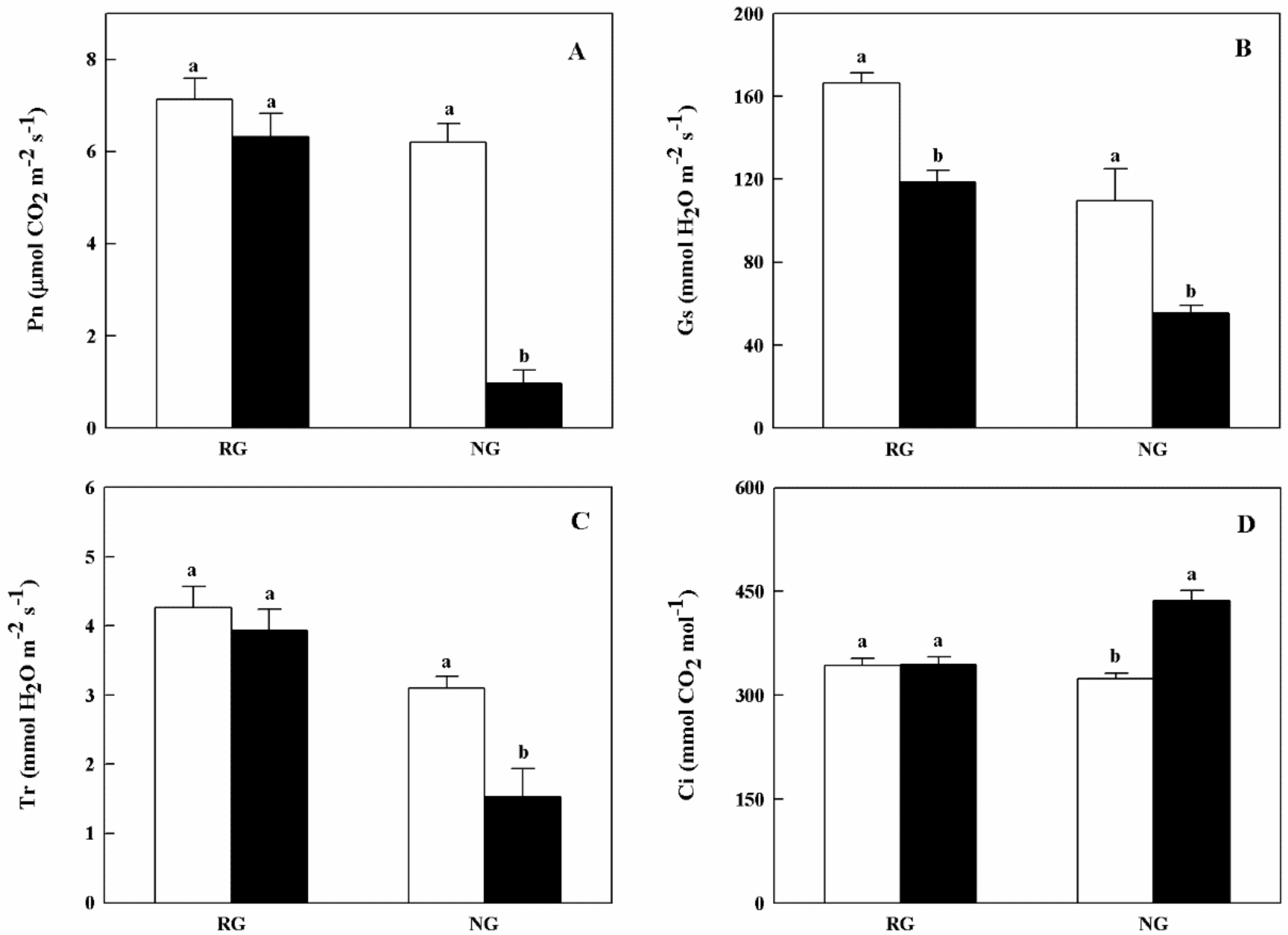
2.2. Chlorophyll Contents and Gas Exchange Parameters
2.3. Summary Analysis of mRNA Sequencing
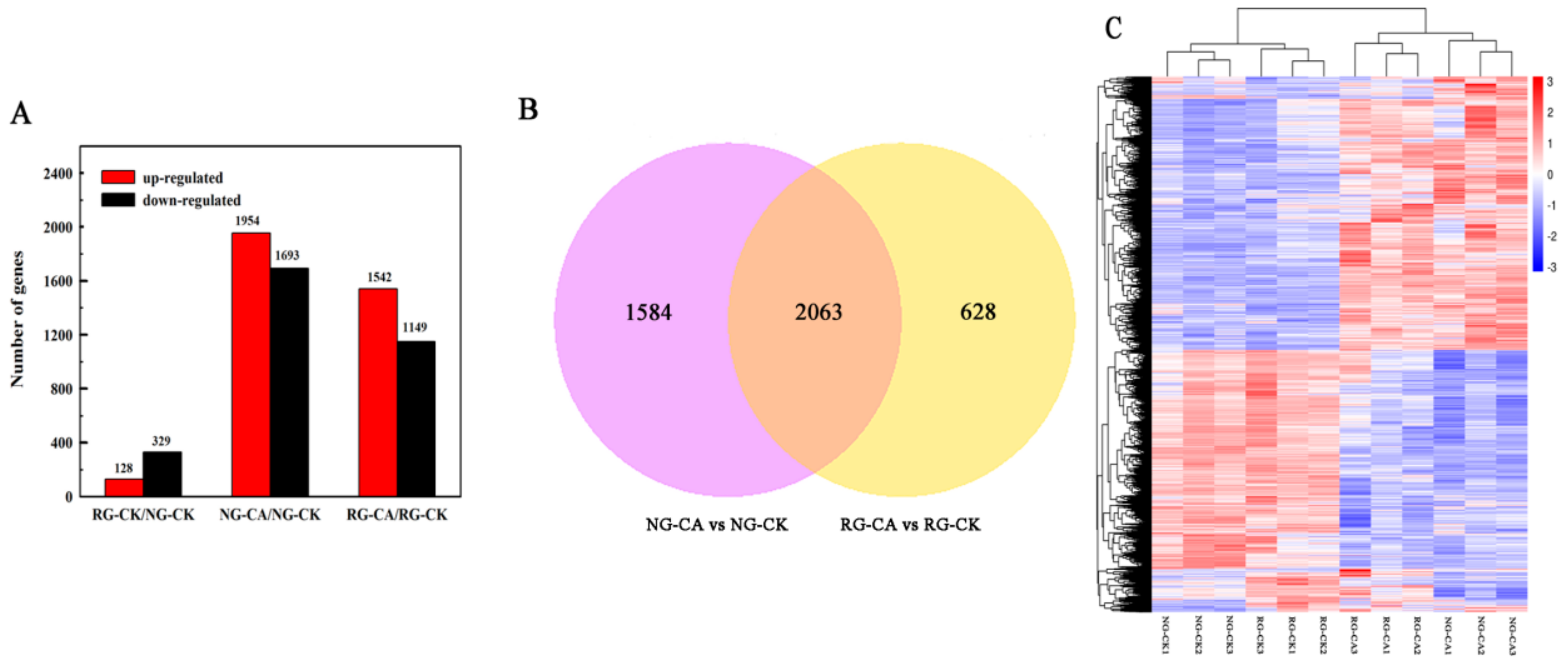
2.4. Identification of Differentially Expressed Genes (DEGs)
2.5. Functional Classifications of DEGs Response to Grafting
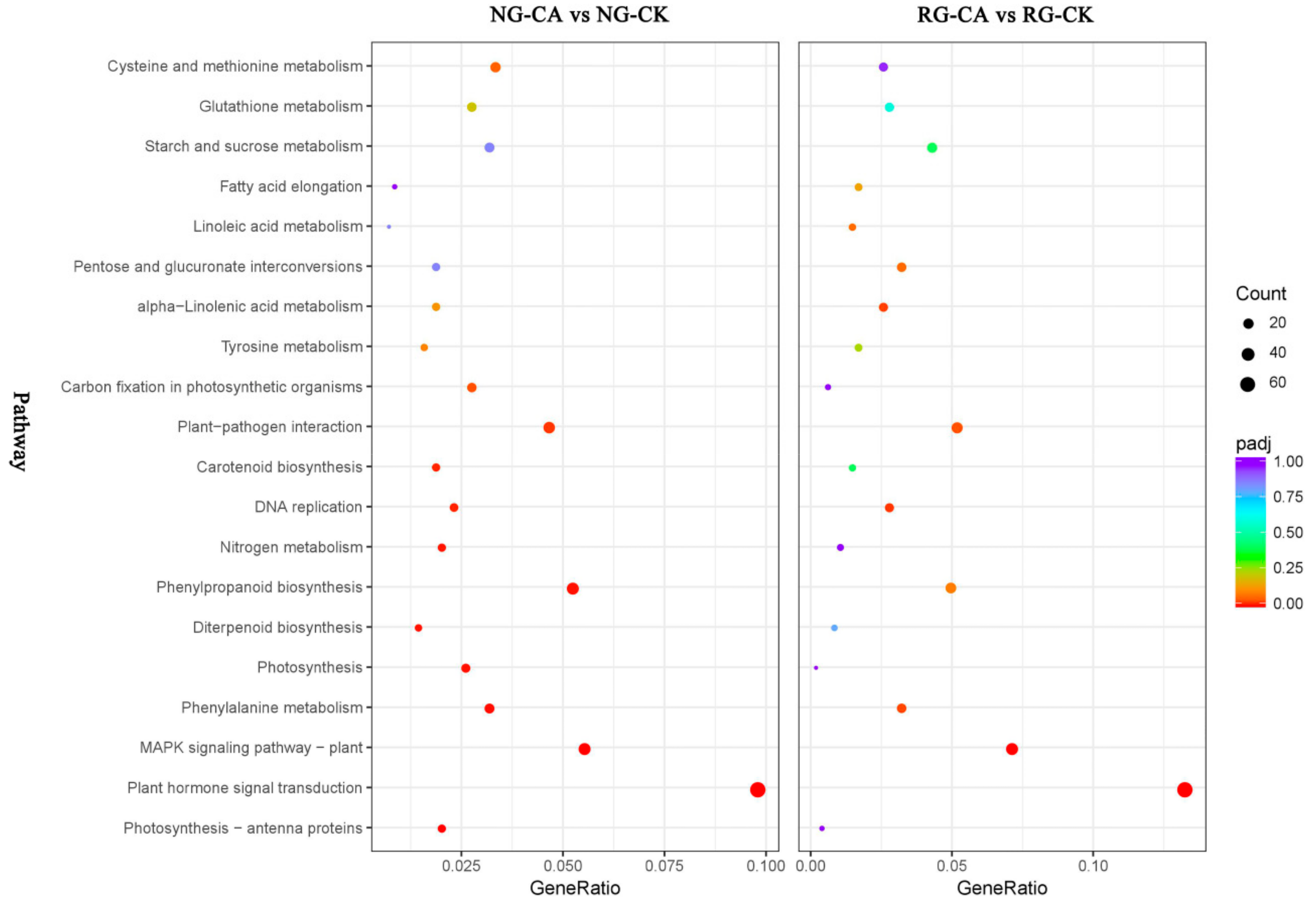
2.6. Functional Classifications of DEGs Response to CA Stress
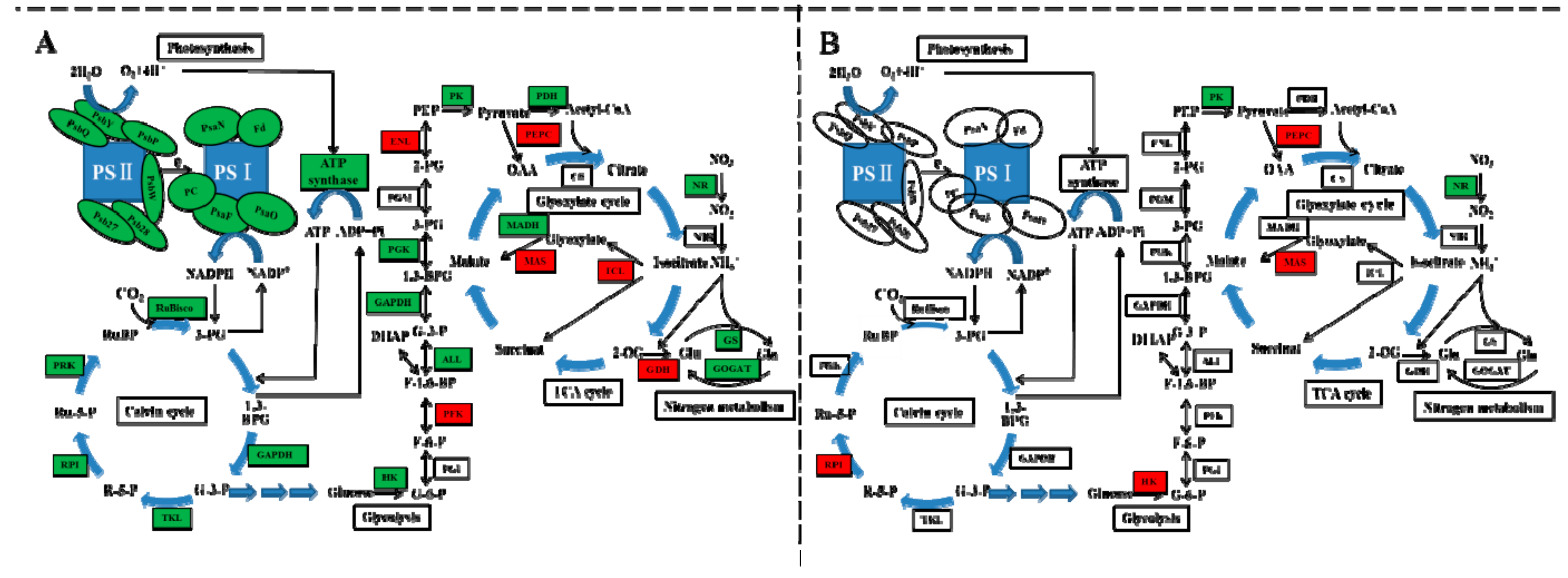
2.7. DEGs Involved in Photosynthesis and Carbon Metabolism
2.8. Transcription Factors Encoded by DEGs
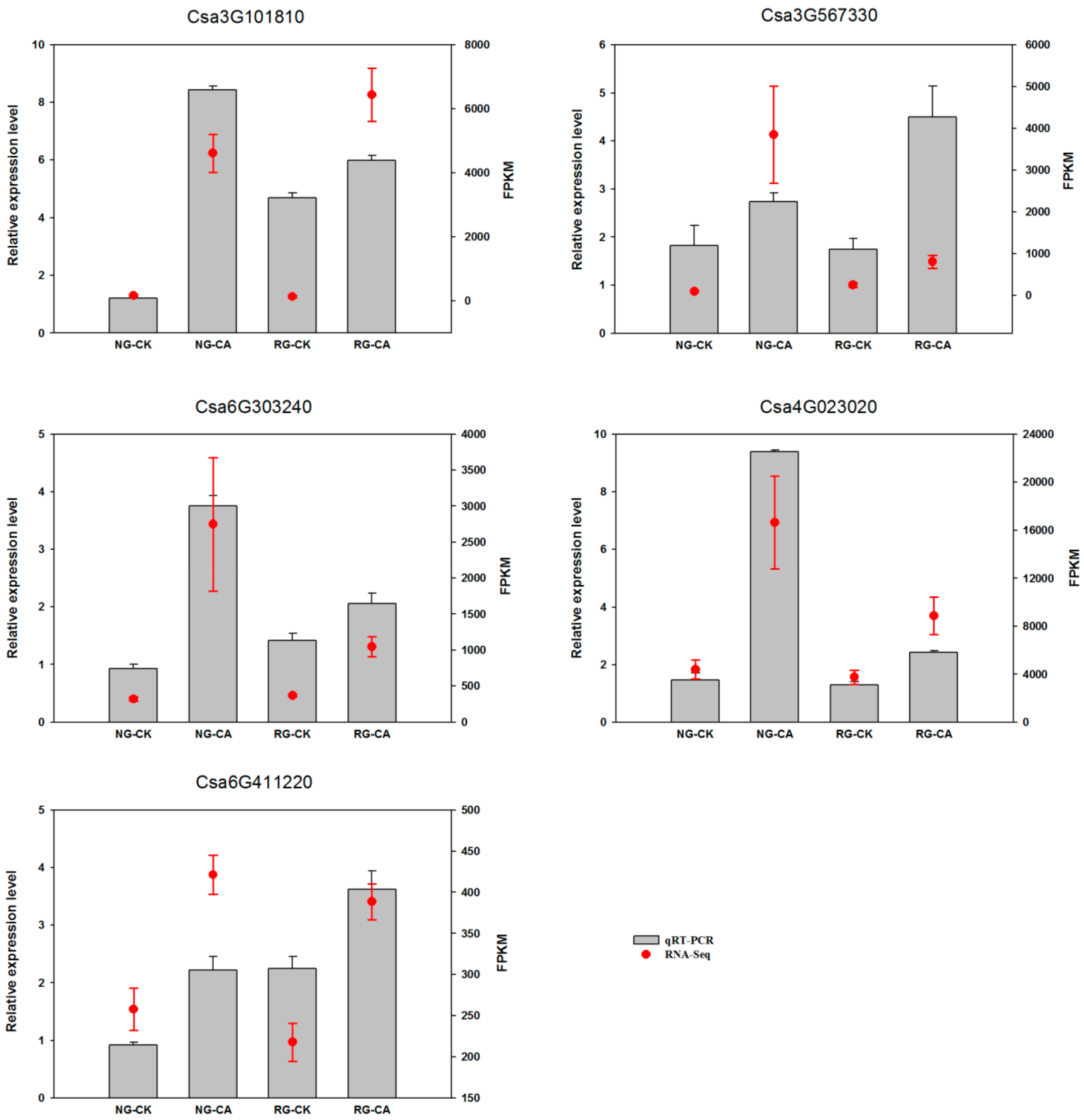
2.9. Validation of mRNA-Seq by qRT-PCR
3. Discussion
3.1. Overall Analysis of Gene Expression Based on RNA-seq
3.2. Discrepancy of Morphology and Photosynthesis between NG and RG Cucumber Seedlings
3.3. Carbon and Nitrogen Metabolism Analysis
3.4. Transcription Factor Analysis
4. Materials and Methods
4.1. Plant Materials and Cinnamic acid Treatments
4.2. Plant Morphology and Biomass
4.3. Leaf Photosynthesis
4.4. RNA-Seq Library Construction and Sequencing
4.5. Transcriptomic Analysis
4.6. Validation of mRNA-Seq by Quantitative Real-Time PCR (qRT-PCR)
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Ali, A.; Imran Ghani, M.; Li, Y.; Ding, H.; Meng, H.; Cheng, Z. Hiseq base molecular characterization of soil microbial community, diversity structure, and predictive functional profiling in continuous cucumber planted soil affected by diverse cropping systems in an intensive greenhouse region of northern China. Int. J. Mol. Sci. 2019, 20, 2619. [Google Scholar] [CrossRef]
- Tian, Y.; Feng, F.; Zhang, B.; Li, M.; Wang, F.; Gu, L.; Chen, A.; Li, Z.; Shan, W.; Wang, X.; et al. Transcriptome analysis reveals metabolic alteration due to consecutive monoculture and abiotic stress stimuli in Rehamannia glutinosa Libosch. Plant. Cell Rep. 2017, 36, 859–875. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Y.; Wu, L.; Chu, L.; Yang, Y.; Li, Z.; Azeem, S.; Zhang, Z.; Fang, C.; Lin, W. Interaction of Pseudostellaria heterophylla with Fusarium oxysporum f. sp. heterophylla mediated by its root exudates in a consecutive monoculture system. Sci. Rep. 2015, 5, 8197. [Google Scholar] [CrossRef]
- Yu, J.Q.; Matsui, Y. Phytotoxic substances in root exudates of cucumber (Cucumis sativus L.). J. Chem. Ecol. 1994, 20, 21–31. [Google Scholar] [CrossRef] [PubMed]
- Yu, J.Q.; Ye, S.F.; Zhang, M.F.; Hu, W.H. Effects of root exudates and aqueous root extracts of cucumber (Cucumis sativus) and allelochemicals, on photosynthesis and antioxidant enzymes in cucumber. Biochem. Syst. Ecol. 2003, 31, 129–139. [Google Scholar] [CrossRef]
- Ye, S.F.; Zhou, Y.H.; Sun, Y.; Zou, L.Y.; Yu, J.Q. Cinnamic acid causes oxidative stress in cucumber roots, and promotes incidence of Fusarium wilt. Environ. Exp. Bot. 2006, 56, 255–262. [Google Scholar] [CrossRef]
- Wu, Y.; Zhang, X.; Gao, L.; Yuan, L.; Wei, W. Effect of different cultivaiton patterns on continuous cropping soil environment and cucumber growth. In XXVII International Horticultural Congress-IHC2006: International Symposium on Advances in Environmental Control, Automation; International Society for Horticultural Science (ISHS): Leuven, Belgium, 2007; pp. 547–554. [Google Scholar]
- Yang, W.; Yan, H.; Zhang, J.; Meng, Y.; Wang, X.; Ji, L.; Luo, Y. Response of rhizosphere microbial diversity and soil physico-chemical properties in a rotation of cucumber with Volvariella volvacea. Biocontrol. Sci. Technol. 2017, 27, 311–323. [Google Scholar] [CrossRef]
- Xiao, X.; Cheng, Z.; Meng, H.; Liu, L.; Li, H.; Dong, Y. Intercropping of green garlic (Allium sativum L.) induces nutrient concentration changes in the soil and plants in continuously cropped cucumber (Cucumis sativus L.) in a plastic tunnel. PLoS ONE 2013, 8, e62173. [Google Scholar] [CrossRef]
- Xiao, X.; Cheng, Z.; Lv, J.; Xie, J.; Ma, N.; Yu, J. A green garlic (Allium sativum L.) based intercropping system reduces the strain of continuous monocropping in cucumber (Cucumis sativus L.) by adjusting the micro-ecological environment of soil. PeerJ 2019, 7, e7267. [Google Scholar] [CrossRef]
- Chen, L.; Yang, X.; Raza, W.; Li, J.; Liu, Y.; Qiu, M.; Zhang, F.; Shen, Q. Trichoderma harzianum SQR-T037 rapidly degrades allelochemicals in rhizospheres of continuously cropped cucumbers. Appl. Microbiol. Biotechnol. 2011, 89, 1653–1663. [Google Scholar] [CrossRef]
- Xie, X.-G.; Dai, C.-C. Biodegradation of a model allelochemical cinnamic acid by a novel endophytic fungus Phomopsis liquidambari. Int. Biodeterior. Biodegrad. 2015, 104, 498–507. [Google Scholar] [CrossRef]
- Monisha, T.R.; Ismailsab, M.; Masarbo, R.; Nayak, A.S.; Karegoudar, T.B. Degradation of cinnamic acid by a newly isolated bacterium Stenotrophomonas sp. TRMK2. 3 Biotech. 2018, 8, 368. [Google Scholar] [CrossRef] [PubMed]
- Ling, N.; Zhang, W.; Wang, D.; Mao, J.; Huang, Q.; Guo, S.; Shen, Q. Root exudates from grafted-root watermelon showed a certain contribution in inhibiting Fusarium oxysporum f. sp. niveum. PLoS ONE 2013, 8, e63383. [Google Scholar] [CrossRef] [PubMed]
- Chen, S.; Zhou, B.; Lin, S.; Li, X.; Xi, H.; Yin, Y.; Ye, X. Allelopathic effects of cinnamic acid and vanillin on soil microbes, soil enzymes activities and growth of grafted eggplants. Allelopath. J. 2011, 28, 29–40. [Google Scholar]
- Ding, J.; Sun, Y.; Xiao, C.L.; Shi, K.; Zhou, Y.H.; Yu, J.Q. Physiological basis of different allelopathic reactions of cucumber and figleaf gourd plants to cinnamic acid. J. Exp. Bot. 2007, 58, 3765–3773. [Google Scholar] [CrossRef] [PubMed]
- Xu, J.; Zhang, M.; Liu, G.; Yang, X.; Hou, X. Comparative transcriptome profiling of chilling stress responsiveness in grafted watermelon seedlings. Plant Physiol. Biochem. 2016, 109, 561–570. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Z.; Cao, B.; Li, N.; Chen, Z.; Xu, K. Comparative transcriptome analysis of the regulation of ABA signaling genes in different rootstock grafted tomato seedlings under drought stress. Environ. Exp. Bot. 2019, 166, 103814. [Google Scholar] [CrossRef]
- Kim, D.; Langmead, B.; Salzberg, S.L. HISAT: A fast spliced aligner with low memory requirements. Nat. Methods 2015, 12, 357. [Google Scholar] [CrossRef]
- Li, Z.; Zhang, Z.; Yan, P.; Huang, S.; Fei, Z.; Lin, K. RNA-Seq improves annotation of protein-coding genes in the cucumber genome. BMC Genom. 2011, 12, 540. [Google Scholar] [CrossRef]
- Zhang, E.P.; Zhang, S.H.; Zhang, W.B.; Li, L.L.; Li, T.L. Effects of exogenic benzoic acid and cinnamic acid on the root oxidative damage of tomato seedlings J. Hortic. For. 2010, 2, 22–29. [Google Scholar]
- Wu, H.S.; Raza, W.; Fan, J.Q.; Sun, Y.G.; Bao, W.; Shen, Q.R. Cinnamic acid inhibits growth but stimulates production of pathogenesis factors by in vitro cultures of Fusarium oxysporum f.sp. niveum. J. Agric. Food Chem. 2008, 56, 1316–1321. [Google Scholar] [CrossRef] [PubMed]
- Guan, W.J.; Zhao, X.; Hassell, R.; Thies, J. Defense mechanisms involved in disease resistance of grafted vegetables. HortScience 2012, 47, 164–170. [Google Scholar] [CrossRef]
- Lee, J.-M.; Kubota, C.; Tsao, S.J.; Bie, Z.; Echevarria, P.H.; Morra, L.; Oda, M. Current status of vegetable grafting: Diffusion, grafting techniques, automation. Sci. Hortic. 2010, 127, 93–105. [Google Scholar] [CrossRef]
- Wang, Z.; Gerstein, M.; Snyder, M. RNA-Seq: A revolutionary tool for transcriptomics. Nat. Rev. Genet. 2009, 10, 57. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.; Bie, Z.; He, S.; Hua, B.; Zhen, A.; Liu, Z. Improving cucumber tolerance to major nutrients induced salinity by grafting onto Cucurbita ficifolia. Environ. Exp. Bot. 2010, 69, 32–38. [Google Scholar] [CrossRef]
- Zhou, Y.; Zhou, J.; Huang, L.; Ding, X.; Shi, K.; Yu, J. Grafting of Cucumis sativus onto Cucurbita ficifolia leads to improved plant growth, increased light utilization and reduced accumulation of reactive oxygen species in chilled plants. J. Plant Res. 2009, 122, 529–540. [Google Scholar] [CrossRef]
- Kumar, P.; Khapte, P.S.; Saxena, A.; Singh, A.; Parwar, N.R.; Kumar, P. Intergeneric grafting for enhanced growth, yield and nutrient acquisition in greenhouse cucumber during winter. J. Environ. Biol. 2019, 40, 295–301. [Google Scholar] [CrossRef]
- Pramanik, M.; Asao, T.; Yamamoto, T.; Matsui, Y. Sensitive bioassay to evaluate toxicity of aromatic acids to cucumber seedlings. Allelopath. J. 2001, 8, 161–169. [Google Scholar]
- Liu, X.; Zhang, J.W.; Guo, L.X.; Liu, Y.Z.; Jin, L.F.; Hussain, S.B.; Du, W.; Deng, Z.; Peng, S.A. Transcriptome changes associated with boron deficiency in leaves of two citrus scion-rootstock combinations. Front. Plant Sci. 2017, 8, 317. [Google Scholar] [CrossRef]
- Gao, M.; Hu, L.; Li, Y.; Weng, Y. The chlorophyll-deficient golden leaf mutation in cucumber is due to a single nucleotide substitution in CsChlI for magnesium chelatase I subunit. Theor. Appl. Genet. 2016, 129, 1961–1973. [Google Scholar] [CrossRef]
- Zima, J.; Šesták, Z. Carbon fixation pathways, their enzymes and products during leaf ontogeny. In Photosynthesis during Leaf Development; Šestăk, Z., Ed.; Springer: Dordrecht, The Netherlands, 1985; pp. 145–156. [Google Scholar]
- Bradbeer, J.W.; Ireland, H.M.M.; Smith, J.W.; Rest, J.; Edge, H.J.W. Plastid development in primary leaves of Phaseolus vulgaris VII. Development during growth in continuous darkness. New Phytol. 1974, 73, 263–270. [Google Scholar] [CrossRef]
- Zhang, N.; Zhang, H.J.; Sun, Q.Q.; Cao, Y.Y.; Li, X.; Zhao, B.; Wu, P.; Guo, Y.D. Proteomic analysis reveals a role of melatonin in promoting cucumber seed germination under high salinity by regulating energy production. Sci. Rep. 2017, 7, 503. [Google Scholar] [CrossRef] [PubMed]
- De Bellis, L.; Nishimura, M. Development of enzymes of the glyoxylate cycle during senescence of pumpkin cotyledons. Plant Cell Physiol. 1991, 32, 555–561. [Google Scholar] [CrossRef]
- Yuenyong, W.; Sirikantaramas, S.; Qu, L.J.; Buaboocha, T. Isocitrate lyase plays important roles in plant salt tolerance. BMC Plant Biol. 2019, 19, 472. [Google Scholar] [CrossRef]
- O’Leary, B.; Park, J.; Plaxton, W.C. The remarkable diversity of plant PEPC (phosphoenolpyruvate carboxylase): Recent insights into the physiological functions and post-translational controls of non-photosynthetic PEPCs. Biochem. J. 2011, 436, 15–34. [Google Scholar] [CrossRef]
- Liu, A.; Xiao, Z.; Li, M.W.; Wong, F.L.; Yung, W.S.; Ku, Y.S.; Wang, Q.; Wang, X.; Xie, M.; Yim, A.K.; et al. Transcriptomic reprogramming in soybean seedlings under salt stress. Plant. Cell Environ. 2019, 42, 98–114. [Google Scholar] [CrossRef]
- Meng, S.; Zhang, C.; Su, L.; Li, Y.; Zhao, Z. Nitrogen uptake and metabolism of Populus simonii in response to PEG-induced drought stress. Environ. Exp. Bot. 2016, 123, 78–87. [Google Scholar] [CrossRef]
- Shu, S.; Tang, Y.; Yuan, Y.; Sun, J.; Zhong, M.; Guo, S. The role of 24-epibrassinolide in the regulation of photosynthetic characteristics and nitrogen metabolism of tomato seedlings under a combined low temperature and weak light stress. Plant Physiol. Biochem. 2016, 107, 344–353. [Google Scholar] [CrossRef]
- Mizoi, J.; Shinozaki, K.; Yamaguchi-Shinozaki, K. AP2/ERF family transcription factors in plant abiotic stress responses. Biochim. Biophys. Acta 2012, 1819, 86–96. [Google Scholar] [CrossRef]
- Vadassery, J.; Jisha, V.; Kappara, S.; Mithöfer, A.; Dampanaboina, L.; Ramanan, R. Overexpression of an AP2/ERF type transcription factor OsEREBP1 confers biotic and abiotic stress tolerance in rice. Public Library of Science: San Francisco, CA, USA. PLoS ONE 2015, 10, e0127831. [Google Scholar] [CrossRef]
- Tang, Y.; Liu, K.; Zhang, J.; Li, X.; Xu, K.; Zhang, Y.; Qi, J.; Yu, D.; Wang, J.; Li, C. JcDREB2, a physic nut AP2/ERF gene, alters plant growth and salinity stress responses in transgenic rice. Front. Plant Sci. 2017, 8, 306. [Google Scholar] [CrossRef] [PubMed]
- Vannozzi, A.; Boss, P.K.; Walker, A.R.; Lucchin, M.; Dry, I.B. Identification of an R2r3 Myb transcription factor involved in the regulation of the stilbene synthase pathway in grapevine. Acta Hortic. 2014, 1046, 57–64. [Google Scholar] [CrossRef]
- Cui, M.H.; Yoo, K.S.; Hyoung, S.; Nguyen, H.T.; Kim, Y.Y.; Kim, H.J.; Ok, S.H.; Yoo, S.D.; Shin, J.S. An Arabidopsis R2R3-MYB transcription factor, AtMYB20, negatively regulates type 2C serine/threonine protein phosphatases to enhance salt tolerance. FEBS Lett. 2013, 587, 1773–1778. [Google Scholar] [CrossRef] [PubMed]
- Liao, Y.; Zou, H.-F.; Wang, H.-W.; Zhang, W.-K.; Ma, B.; Zhang, J.-S.; Chen, S.-Y. Soybean GmMYB76, GmMYB92, and GmMYB177 genes confer stress tolerance in transgenic Arabidopsis plants. Cell Res. 2008, 18, 1047. [Google Scholar] [CrossRef] [PubMed]
- Wang, R.K.; Cao, Z.H.; Hao, Y.J. Overexpression of a R2R3 MYB gene MdSIMYB1 increases tolerance to multiple stresses in transgenic tobacco and apples. Physiol. Plant. 2014, 150, 76–87. [Google Scholar] [CrossRef] [PubMed]
- Stracke, R.; Ishihara, H.; Huep, G.; Barsch, A.; Mehrtens, F.; Niehaus, K.; Weisshaar, B. Differential regulation of closely related R2R3-MYB transcription factors controls flavonol accumulation in different parts of the Arabidopsis thaliana seedling. Plant J. 2007, 50, 660–677. [Google Scholar] [CrossRef] [PubMed]
- Xi, Y.; Zhang, Z.; Xie, X.; Gao, Q. Cloning and expression analysis of one related to autotoxicity MYB transcription factor in melon. Chin. J. Trop. Crop. 2015, 36, 296–301. [Google Scholar]
- Davis, A.R.; Perkins-Veazie, P.; Sakata, Y.; López-Galarza, S.; Maroto, J.V.; Lee, S.-G.; Huh, Y.-C.; Sun, Z.; Miguel, A.; King, S.R. Cucurbit grafting. Crit. Rev. Plant Sci. 2008, 27, 50–74. [Google Scholar] [CrossRef]
- Yamazaki, K. Yōeki Saibai Zenpen; Hakuyūsha: Tokyo, Japan, 1978. [Google Scholar]
- Arnon, D.I. Copper enzymes in isolated chloroplasts. Polyphenoloxidase in Beta vulgaris. Plant Physiol. 1949, 24, 1. [Google Scholar] [CrossRef]
- Love, M.I.; Huber, W.; Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014, 15, 550. [Google Scholar] [CrossRef]
- Tarazona, S.; García-Alcalde, F.; Dopazo, J.; Ferrer, A.; Conesa, A. Differential expression in RNA-seq: A matter of depth. Genome Res. 2011, 21, 2213–2223. [Google Scholar] [CrossRef] [PubMed]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]

| Treatment | Plant Height (cm) | Stem Diameter (mm) | Leaves Area (cm2) | Shoot | Root | ||
|---|---|---|---|---|---|---|---|
| Fresh Weight (g) | Dry Weight (g) | Fresh Weight (g) | Dry Weight (g) | ||||
| RG-CK | 14.67 ± 1.76 a | 7.96 ± 0.13 a | 432.81 ± 2.61 a | 12.13 ± 0.09 a | 1.11 ± 0.010 a | 2.54 ± 0.23 a | 0.13 ± 0.010 a |
| RG-CA | 15.17 ± 1.04 a | 7.86 ± 0.25 a | 427.86 ± 10.44 a | 12.02 ± 0.17 a | 1.10 ± 0.021 a | 2.12 ± 0.16 a | 0.12 ± 0.012 a |
| NG-CK | 14.17 ± 2.08 a | 7.75 ± 0.46 a | 420.68 ± 3.28 a | 9.83 ± 0.20 a | 0.73 ± 0.026 a | 1.82 ± 0.06 a | 0.07 ± 0.006 a |
| NG-CA | 9.50 ± 0.50 b | 7.37 ± 0.25 a | 380.15 ± 5.72 b | 8.16 ± 0.22 b | 0.69 ± 0.017 a | 1.45 ± 0.07 b | 0.04 ± 0.006 b |
| Sample Name | Raw Reads | Clean Reads | Total Mapped Ratio (%) | Unique Mapped Ratio (%) | Q30 Percentage (%) | Expressed Genes | Expressed Genes Ratio (% of Annotated Genes) |
|---|---|---|---|---|---|---|---|
| RG-CK-1 | 60065280 | 58347004 | 93.64% | 91.81% | 89.84 | 17434 | 74.99 |
| RG-CK-2 | 52976972 | 51443526 | 93.71% | 91.92% | 90.03 | 17386 | 74.78 |
| RG-CK-3 | 52288760 | 51039256 | 94.32% | 92.45% | 90.83 | 17453 | 75.07 |
| RG-CA-1 | 46296420 | 45246686 | 93.23% | 91.53% | 88.34 | 17629 | 75.83 |
| RG-CA-2 | 49512360 | 48120914 | 93.17% | 91.27% | 89.82 | 17564 | 75.55 |
| RG-CA-3 | 48778530 | 47442262 | 92.92% | 90.96% | 90.11 | 17279 | 74.32 |
| NG-CK-1 | 50039680 | 48367266 | 93.51% | 91.83% | 89.96 | 17686 | 76.08 |
| NG-CK-2 | 59498560 | 58473188 | 94.13% | 92.32% | 90.53 | 17484 | 75.21 |
| NG-CK-3 | 44090680 | 43071056 | 94.1% | 92.35% | 90.56 | 17430 | 74.97 |
| NG-CA-1 | 48323566 | 47469736 | 93.98% | 92.28% | 90.43 | 17467 | 75.13 |
| NG-CA-2 | 54922480 | 53952812 | 94.08% | 92.27% | 90.81 | 17626 | 75.82 |
| NG-CA-3 | 50581658 | 49314228 | 94.35% | 92.58% | 90.72 | 17417 | 74.92 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Xiao, X.; Lv, J.; Xie, J.; Feng, Z.; Ma, N.; Li, J.; Yu, J.; Calderón-Urrea, A. Transcriptome Analysis Reveals the Different Response to Toxic Stress in Rootstock Grafted and Non-Grafted Cucumber Seedlings. Int. J. Mol. Sci. 2020, 21, 774. https://doi.org/10.3390/ijms21030774
Xiao X, Lv J, Xie J, Feng Z, Ma N, Li J, Yu J, Calderón-Urrea A. Transcriptome Analysis Reveals the Different Response to Toxic Stress in Rootstock Grafted and Non-Grafted Cucumber Seedlings. International Journal of Molecular Sciences. 2020; 21(3):774. https://doi.org/10.3390/ijms21030774
Chicago/Turabian StyleXiao, Xuemei, Jian Lv, Jianming Xie, Zhi Feng, Ning Ma, Ju Li, Jihua Yu, and Alejandro Calderón-Urrea. 2020. "Transcriptome Analysis Reveals the Different Response to Toxic Stress in Rootstock Grafted and Non-Grafted Cucumber Seedlings" International Journal of Molecular Sciences 21, no. 3: 774. https://doi.org/10.3390/ijms21030774
APA StyleXiao, X., Lv, J., Xie, J., Feng, Z., Ma, N., Li, J., Yu, J., & Calderón-Urrea, A. (2020). Transcriptome Analysis Reveals the Different Response to Toxic Stress in Rootstock Grafted and Non-Grafted Cucumber Seedlings. International Journal of Molecular Sciences, 21(3), 774. https://doi.org/10.3390/ijms21030774





