EGFR-Binding Peptides: From Computational Design towards Tumor-Targeting of Adeno-Associated Virus Capsids
Abstract
1. Introduction
2. Results
2.1. Computational Design of β-Hairpin Peptides Competitively Inhibiting Dimerization of EGFR
2.2. Synthesis and Cyclization Of Peptides
2.3. Biophysical Characterization of Peptides
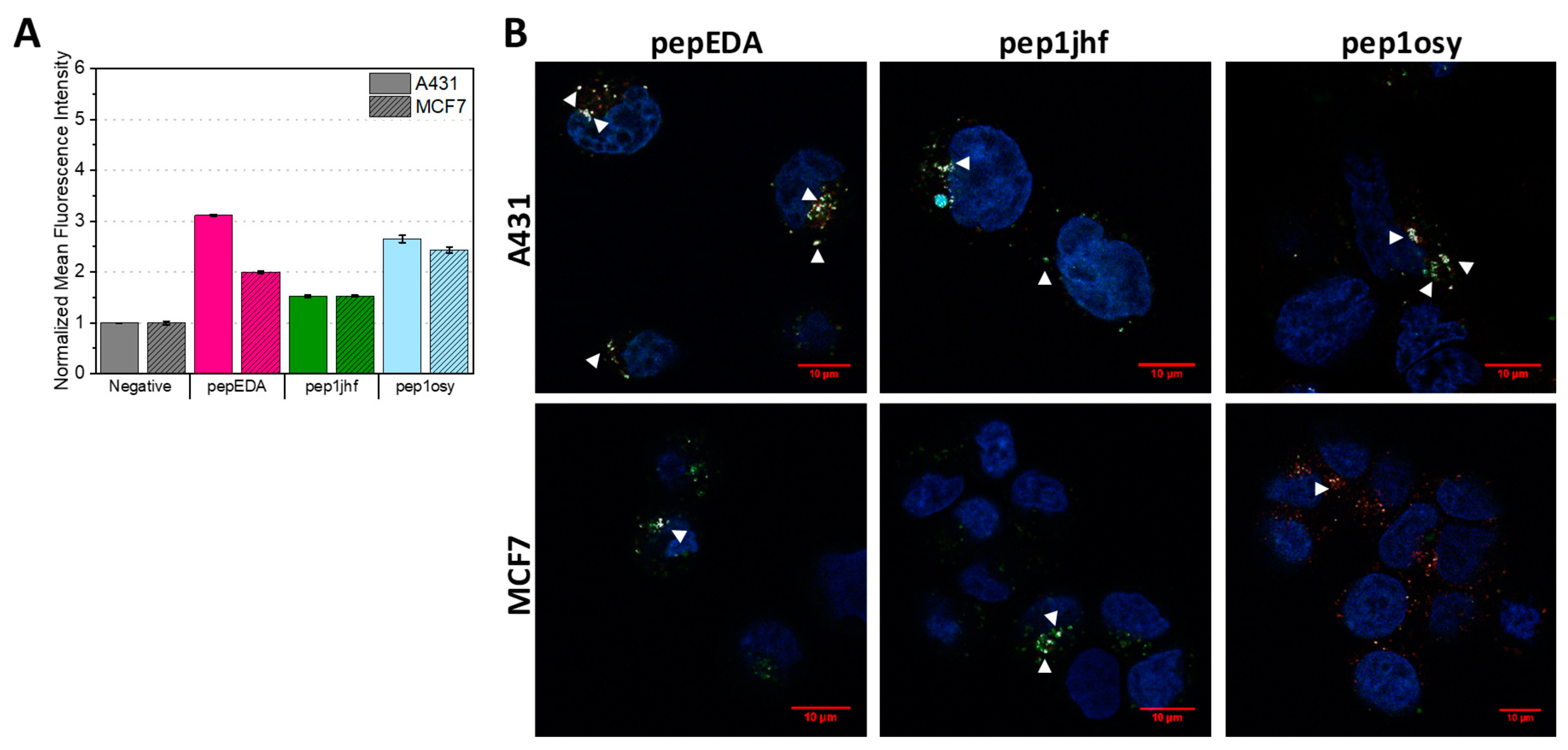
2.4. Cyclic Peptides Show Binding to Cells Expressing EGFR at High and Low Levels
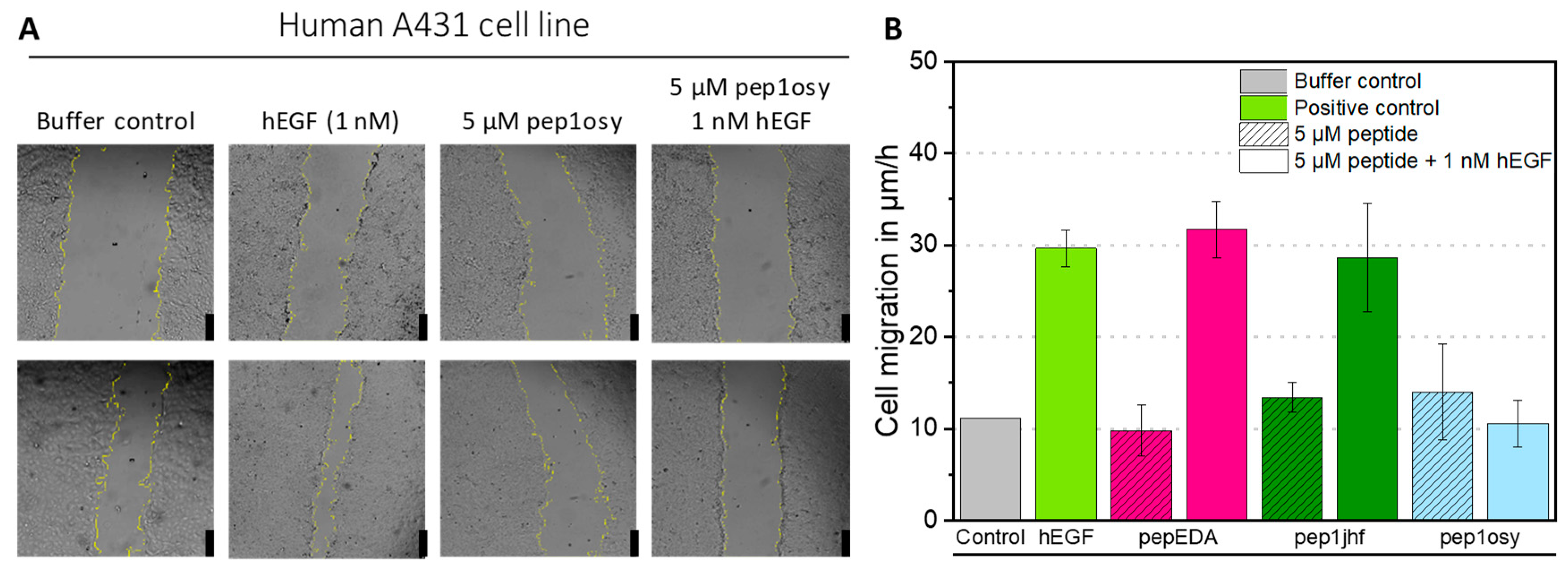
2.5. Cyclic Peptide Pep1osy Shows Inhibitory Properties in Wound Healing Assays
2.6. Peptides Grafted on the Recombinant Adeno-Associated Virus 2 Capsid
2.7. Insertion of Designed EGFR-Targeting Peptides Changes the Tropism of rAAV2 towards EGFR Overexpressing Cells
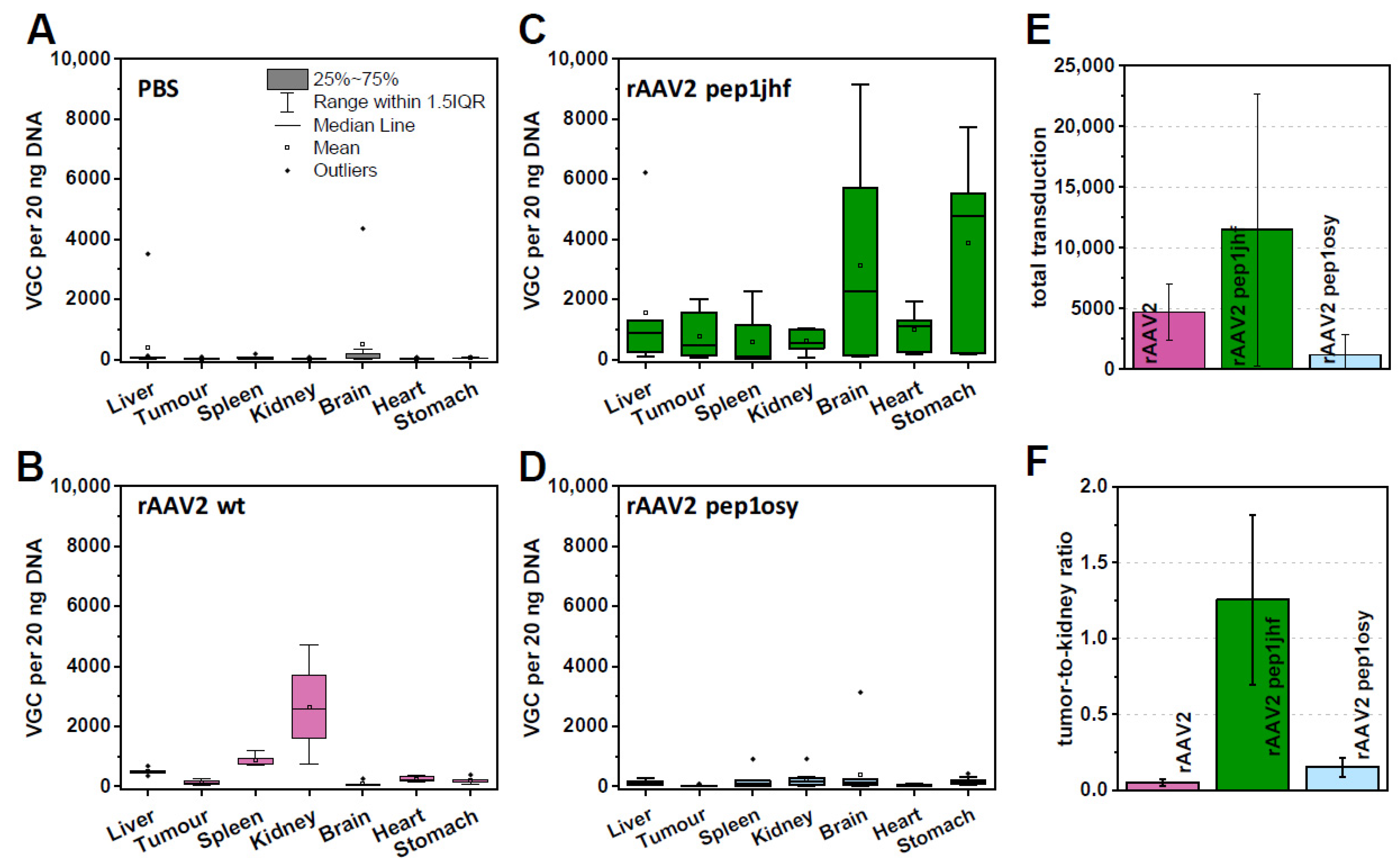
2.8. rAAV2 587 Pep1jhf Demonstrates Efficacy in Egg Xenografts
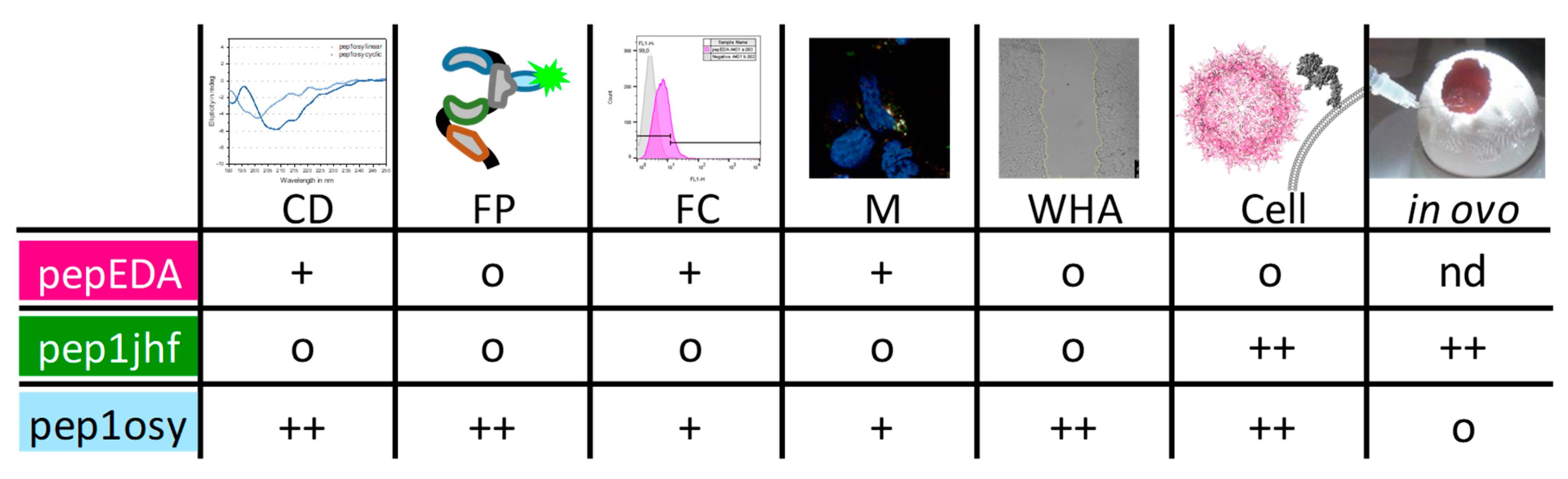
3. Discussion
4. Materials and Methods
4.1. Structural Modeling
4.2. General Procedure
4.3. Peptide Synthesis
4.4. N-Terminal Fluorescein Labeling
4.5. Peptide Cleavage
4.6. Cyclization of Peptides
4.7. Cell Culture
4.8. Wound Healing Assay
4.9. Confocal Fluorescence Microscopy
4.10. Circular Dichroism Spectrometry
4.11. Fluorescence Polarization
4.12. Viral Particle Production
4.13. Purification of Viral Particles
4.14. SDS-PAGE and Western Blot Analysis
4.15. Determination of Genomic Titers
4.16. Transduction Assay
4.17. Chorioallantoic Membrane (CAM) Assay
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| CAM | Chorioallantoic membrane assay |
| CD | Circular dichroism |
| DMF | Dimethylformamide |
| EGFR | Epidermal growth factor receptor |
| LC-MS | Liquid chromatography–mass spectroscopy |
| MALDI | Matrix-assisted laser desorption ionization |
| PBS | Phosphate buffered saline |
| rAAV | Recombinant adeno-associated virus |
| RP-HPLC | Reversed-phase high-pressure liquid chromatography |
| VGC | Vector genome copies |
References
- Lemmon, M.A.; Schlessinger, J. Cell signaling by receptor tyrosine kinases. Cell 2010, 141, 1117–1134. [Google Scholar] [CrossRef]
- Ullrich, A.; Coussens, L.; Hayflick, J.; Dull, T.; Gray, A.; Tam, A.; Lee, J.; Yarden, Y.; Libermann, T.; Schlessinger, J. Human epidermal growth factor receptor cDNA sequence and aberrant expression of the amplified gene in A431 epidermoid carcinoma cells. Nature 1984, 309, 418–425. [Google Scholar] [CrossRef]
- Ogiso, H.; Ishitani, R.; Nureki, O.; Fukai, S.; Yamanaka, M.; Kim, J.H.; Saito, K.; Sakamoto, A.; Inoue, M.; Shirouzu, M.; et al. Crystal structure of the complex of human epidermal growth factor and receptor extracellular domains. Cell 2002, 110, 775–787. [Google Scholar] [CrossRef]
- Arkhipov, A.; Shan, Y.; Das, R.; Endres, N.F.F.F.; Eastwood, M.P.P.P.; Wemmer, D.E.E.E.; Kuriyan, J.; Shaw, D.E.E.E. Architecture and Membrane Interactions of the EGF Receptor. Cell 2013, 152, 557–569. [Google Scholar] [CrossRef] [PubMed]
- Bessman, N.J.; Bagchi, A.; Ferguson, K.M.; Lemmon, M.A. Complex Relationship between Ligand Binding and Dimerization in the Epidermal Growth Factor Receptor. Cell Rep. 2014, 9, 1306–1317. [Google Scholar] [CrossRef] [PubMed]
- Burgess, A.W.; Cho, H.; Eigenbrot, C.; Ferguson, K.M.; Garrett, T.P.J.; Leahy, D.J.; Lemmon, M.A.; Sliwkowski, M.X.; Ward, C.W.; Yokoyama, S. An open-and-shut case? Recent insights into the activation of EGF/ErbB receptors. Mol. Cell 2003, 12, 541–552. [Google Scholar] [CrossRef]
- Roskoski, R. The ErbB/HER family of protein-tyrosine kinases and cancer. Pharmacol. Res. 2014, 79, 34–74. [Google Scholar] [CrossRef]
- Yarden, Y.; Sliwkowski, M.X. Untangling the ErbB signalling network. Nat. Rev. Mol. Cell Biol. 2001, 2, 127–137. [Google Scholar] [CrossRef]
- Schlessinger, J. Ligand-induced, receptor-mediated dimerization and activation of EGF receptor. Cell 2002, 110, 669–672. [Google Scholar] [CrossRef]
- Harris, R.C.; Chung, E.; Coffey, R.J. EGF receptor ligands. Exp. Cell Res. 2003, 284, 2–13. [Google Scholar] [CrossRef]
- Wilson, K.J.; Mill, C.; Lambert, S.; Buchman, J.; Wilson, T.R.; Hernandez-Gordillo, V.; Gallo, R.M.; Ades, L.M.C.; Settleman, J.; Riese, D.J. EGFR ligands exhibit functional differences in models of paracrine and autocrine signaling. Growth Factors 2012, 30, 107–116. [Google Scholar] [CrossRef]
- Ronan, T.; Macdonald-Obermann, J.L.; Huelsmann, L.; Bessman, N.J.; Naegle, K.M.; Pike, L.J. Different Epidermal Growth Factor Receptor (EGFR) agonists produce unique signatures for the recruitment of downstream signaling proteins. J. Biol. Chem. 2016, 291, 5528–5540. [Google Scholar] [CrossRef] [PubMed]
- Dawson, J.P.; Berger, M.B.; Lin, C.; Schlessinger, J.; Lemmon, M.A.; Kathryn, M.; Ferguson, K.M. Epidermal Growth Factor Receptor Dimerization and Activation Require Ligand-Induced Conformational Changes in the Dimer Interface Epidermal Growth Factor Receptor Dimerization and Activation Require Ligand-Induced Conformational Changes in the Dimer Inter. Mol. Cell. Biol. 2005, 25, 7734–7742. [Google Scholar] [CrossRef] [PubMed]
- Friedländer, E.; Barok, M.; Szöllosi, J.; Vereb, G. ErbB-directed immunotherapy: Antibodies in current practice and promising new agents. Immunol. Lett. 2008, 116, 126–140. [Google Scholar] [CrossRef] [PubMed]
- Salomon, D.S.; Brandt, R.; Ciardiello, F.; Normanno, N. Epidermal growth factor-related peptides and their receptors in human malignancies. Crit. Rev. Oncol. Hematol. 1995, 19, 183–232. [Google Scholar] [CrossRef]
- Richard, J.; Sainsbury, C.; Needham, G.K.; Farndon, J.R.; Malcolm, A.J.; Harris, A.L. Epidermal-Growth-Factor Receptor Status As Predictor of Early Recurrence of and Death From Breast Cancer. Lancet 1987, 329, 1398–1402. [Google Scholar] [CrossRef]
- Pao, W.; Chmielecki, J. Rational, biologically based treatment of EGFR-mutant non- small-cell lung cancer. Nat. Rev. Cancer 2010, 10, 760–774. [Google Scholar] [CrossRef]
- Maxwell, S.A.; Sacks, P.G.; Gutterman, J.U.; Gallick, G.E. Epidermal growth factor receptor protein-tyrosine kinase activity in human cell lines established from squamous carcinomas of the head and neck. Cancer Res. 1989, 49, 1130–1137. [Google Scholar]
- Masui, H.; Kawamoto, T.; Sato, J.D.; Wolf, B.; Sato, G.; Mendelsohn, J. Growth inhibition of human tumor cells in athymic mice by anti- epidermal growth factor receptor monoclonal antibodies. Cancer Res. 1984, 44, 1002–1007. [Google Scholar] [CrossRef]
- National Cancer Institute Description of Cetuximab. Available online: http://www.cancer.gov/about-cancer/treatment/drugs/cetuximab (accessed on 13 August 2019).
- National Cancer Institute Description of Panitumumab. Available online: http://www.cancer.gov/about-cancer/treatment/drugs/panitumumab (accessed on 13 August 2019).
- National Cancer Institute Description of Necitumumab. Available online: https://www.cancer.gov/about-cancer/treatment/drugs/necitumumab (accessed on 13 August 2019).
- Franklin, M.C.; Carey, K.D.; Vajdos, F.F.; Leahy, D.J.; De Vos, A.M.; Sliwkowski, M.X. Insights into ErbB signaling from the structure of the ErbB2-pertuzumab complex. Cancer Cell 2004, 5, 317–328. [Google Scholar] [CrossRef]
- Purba, E.; Saita, E.; Maruyama, I. Activation of the EGF Receptor by Ligand Binding and Oncogenic Mutations: The “Rotation Model”. Cells 2017, 6, 13. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.; Zhao, R.; Wu, X.; Sun, Y.; Yao, M.; Li, J.; Xu, Y.; Gu, J. Identification and characterization of a novel peptide ligand of epidermal growth factor receptor for targeted delivery of therapeutics. FASEB J. 2005, 19, 1978–1985. [Google Scholar] [CrossRef] [PubMed]
- Song, S.; Liu, D.; Peng, J.; Deng, H.; Guo, Y.; Xu, L.X.; Miller, A.D.; Xu, Y. Novel peptide ligand directs liposomes toward EGF-R high-expressing cancer cells in vitro and in vivo. FASEB J. 2009, 23, 1396–1404. [Google Scholar] [CrossRef] [PubMed]
- Master, A.M.; Gupta, A. Sen EGF receptor-targeted nanocarriers for enhanced cancer treatment. Nanomedicine 2012, 7, 1895–1906. [Google Scholar] [CrossRef] [PubMed]
- Hong, S.; Leroueil, P.R.; Majoros, I.J.; Orr, B.G.; Baker, J.R.; Banaszak Holl, M.M. The Binding Avidity of a Nanoparticle-Based Multivalent Targeted Drug Delivery Platform. Chem. Biol. 2007, 14, 107–115. [Google Scholar] [CrossRef] [PubMed]
- Mizuguchi, T.; Uchimura, H.; Kakizawa, T.; Kimura, T.; Yokoyama, S.; Kiso, Y.; Saito, K. Inhibitory effect of a dimerization-arm-mimetic peptide on EGF receptor activation. Bioorg. Med. Chem. Lett. 2009, 19, 3279–3282. [Google Scholar] [CrossRef] [PubMed]
- Hanold, L.E.; Oruganty, K.; Ton, N.T.; Beedle, A.M.; Kannan, N.; Kennedy, E.J. Inhibiting EGFR Dimerization Using Triazolyl-Bridged Dimerization Arm Mimics. PLoS ONE 2015, 10, e0118796. [Google Scholar] [CrossRef]
- Toyama, K.; Mizuguchi, T.; Nomura, W.; Tamamura, H. Functional evaluation of fluorescein-labeled derivatives of a peptide inhibitor of the EGF receptor dimerization. Bioorg. Med. Chem. 2016, 24, 3406–3412. [Google Scholar] [CrossRef]
- Daya, S.; Berns, K.I. Gene Therapy Using Adeno-Associated Virus Vectors. Clin. Microbiol. Rev. 2008, 21, 583–593. [Google Scholar] [CrossRef]
- Hagen, S.; Baumann, T.; Wagner, H.J.; Morath, V.; Kaufmann, B.; Fischer, A.; Bergmann, S.; Schindler, P.; Arndt, K.M.; Müller, K.M. Modular adeno-associated virus (rAAV) vectors used for cellular virus-directed enzyme prodrug therapy. Sci. Rep. 2014, 4, 3759. [Google Scholar] [CrossRef]
- Büning, H.; Srivastava, A. Capsid Modifications for Targeting and Improving the Efficacy of AAV Vectors. Mol. Ther. Methods Clin. Dev. 2019, 12, 248–265. [Google Scholar] [CrossRef] [PubMed]
- Summerford, C.; Samulski, R.J. Membrane-associated heparan sulfate proteoglycan is a receptor for adeno-associated virus type 2 virions. J. Virol. 1998, 72, 1438–1445. [Google Scholar] [CrossRef] [PubMed]
- Opie, S.R.; Warrington, K.H., Jr.; Agbandje-McKenna, M.; Zolotukhin, S.; Muzyczka, N. Identification of amino acid residues in the capsid proteins of adeno-associated virus type 2 that contribute to heparan sulfate proteoglycan binding. J. Virol. 2003, 77, 6995–7006. [Google Scholar] [CrossRef] [PubMed]
- Ros, C.; Baltzer, C.; Mani, B.; Kempf, C. Parvovirus uncoating in vitro reveals a mechanism of DNA release without capsid disassembly and striking differences in encapsidated DNA stability. Virology 2006, 345, 137–147. [Google Scholar] [CrossRef]
- Leaver-Fay, A.; Tyka, M.; Lewis, S.M.; Lange, O.F.; Thompson, J.; Jacak, R.; Kaufman, K.; Renfrew, P.D.; Smith, C.A.; Sheffler, W.; et al. ROSETTA3: An object-oriented software suite for the simulation and design of macromolecules. Methods Enzymol. 2011, 487, 545–574. [Google Scholar]
- Greenfield, N.; Fasman, G.D. Computed Circular Dichroism Spectra for the Evaluation of Protein Conformation. Biochemistry 1969, 8, 4108–4116. [Google Scholar] [CrossRef]
- Thévenet, P.; Shen, Y.; Maupetit, J.; Guyon, F.; Derreumaux, P.; Tufféry, P. PEP-FOLD: An updated de novo structure prediction server for both linear and disulfide bonded cyclic peptides. Nucleic Acids Res. 2012, 40, W288–W293. [Google Scholar] [CrossRef]
- Kuo, W.-T.; Lin, W.-C.; Chang, K.-C.; Huang, J.-Y.; Yen, K.-C.; Young, I.-C.; Sun, Y.-J.; Lin, F.-H. Quantitative Analysis of Ligand-EGFR Interactions: A Platform for Screening Targeting Molecules. PLoS ONE 2015, 10, e0116610. [Google Scholar] [CrossRef]
- Reilly, R.M.; Kiarash, R.; Sandhu, J.; Lee, Y.W.; Cameron, R.G.; Hendler, A.; Vallis, K.; Gariépy, J. A comparison of EGF and MAb 528 labeled with 111In for imaging human breast cancer. J. Nucl. Med. 2000, 41, 903–911. [Google Scholar]
- Haigler, H.; Ash, J.F.; Singer, S.J.; Cohen, S. Visualization by fluorescence of the binding and internalization of epidermal growth factor in human carcinoma cells A-431. Proc. Natl. Acad. Sci. USA 1978, 75, 3317–3321. [Google Scholar] [CrossRef]
- Feiner, R.C.; Pennè, I.; Müller, B.; Müller, K.M. EGF-mCherry Fusion Protein Expressed in E. coli Shows Product Heterogeneity but a High Biological Activity. Biochemistry 2019, 58, 1043–1047. [Google Scholar] [CrossRef] [PubMed]
- Vieira, A.V.; Lamaze, C.; Schmid, S.L. Control of EGF receptor signaling by clathrin-mediated endocytosis. Science 1996, 274, 2086–2089. [Google Scholar] [CrossRef] [PubMed]
- Roepstorff, K.; Grandal, M.V.; Henriksen, L.; Knudsen, S.L.J.; Lerdrup, M.; Grøvdal, L.; Willumsen, B.M.; van Deurs, B. Differential effects of EGFR ligands on endocytic sorting of the receptor. Traffic 2009, 10, 1115–1127. [Google Scholar] [CrossRef] [PubMed]
- Ferguson, K.M.; Berger, M.B.; Mendrola, J.M.; Cho, H.; Leahy, D.J.; Lemmon, M.A.; Biophysics, M. EGF Activates Its Receptor by Removing Interactions that Autoinhibit Ectodomain Dimerization. Mol. Cell 2003, 11, 507–517. [Google Scholar] [CrossRef]
- Martin, M.M.; Lindqvist, L. The pH dependence of fluorescein fluorescence. J. Lumin. 1975, 10, 381–390. [Google Scholar] [CrossRef]
- Girod; Ried; Wobus; Lahm; Leike; Kleinschmidt; Deléage; Hallek Genetic capsid modifications allow efficient re-targeting of adeno-associated virus type 2. Nat. Med. 1999, 5, 1438.
- Shi, W.; Bartlett, J.S. RGD inclusion in VP3 provides adeno-associated virus type 2 (AAV2)-based vectors with a heparan sulfate-independent cell entry mechanism. Mol. Ther. 2003, 7, 515–525. [Google Scholar] [CrossRef]
- Lokman, N.A.; Elder, A.S.F.; Ricciardelli, C.; Oehler, M.K. Chick chorioallantoic membrane (CAM) assay as an in vivo model to study the effect of newly identified molecules on ovarian cancer invasion and metastasis. Int. J. Mol. Sci. 2012, 13, 9959–9970. [Google Scholar] [CrossRef]
- Durand, J.-O.; Jimenez, C.M.; Tamanoi, F.; Khashab, N.; Croissant, J.; Ratliff, L.; Shahin, S.A.; Vu, B.T.; Yik, T.; Raehm, L.; et al. Chick chorioallantoic membrane assay as an in vivo model to study the effect of nanoparticle-based anticancer drugs in ovarian cancer. Sci. Rep. 2018, 8, 8524. [Google Scholar]
- Zincarelli, C.; Soltys, S.; Rengo, G.; Rabinowitz, J.E. Analysis of AAV serotypes 1-9 mediated gene expression and tropism in mice after systemic injection. Mol. Ther. 2008, 16, 1073–1080. [Google Scholar] [CrossRef]
- Toyama, K.; Kobayakawa, T.; Nomura, W.; Tamamura, H. Inhibition of EGFR Activation by Bivalent Ligands Based on a Cyclic Peptide Mimicking the Dimerization Arm Structure of EGFR. Chem. Pharm. Bull. 2018, 66, 1083–1089. [Google Scholar] [CrossRef]
- Toyama, K.; Nomura, W.; Kobayakawa, T.; Tamamura, H. Delivery of a Proapoptotic Peptide to EGFR-Positive Cancer Cells by a Cyclic Peptide Mimicking the Dimerization Arm Structure of EGFR. Bioconjug. Chem. 2018, 29, 2050–2057. [Google Scholar] [CrossRef] [PubMed]
- Börner, K.; Kienle, E.; Huang, L.Y.; Weinmann, J.; Sacher, A.; Bayer, P.; Stüllein, C.; Fakhiri, J.; Zimmermann, L.; Westhaus, A.; et al. Pre-arrayed Pan-AAV Peptide Display Libraries for Rapid Single-Round Screening. Mol. Ther. 2020, 28, 1016–1032. [Google Scholar] [CrossRef] [PubMed]
- Krutzke, L.; Allmendinger, E.; Hirt, K.; Kochanek, S. Chorioallantoic Membrane Tumor Model for Evaluating Oncolytic Viruses (Preprint). Hum. Gene Ther. 2020, 31, 1100–1113. [Google Scholar] [CrossRef] [PubMed]
- Takeda, J.; Tabata, H.; Fukuda, K.; Yasugi, S. Involvement of the signal transduction pathway mediated by epidermal growth factor receptor in the differentiation of chicken glandular stomach. Dev. Growth Differ. 2002, 44, 501–508. [Google Scholar] [CrossRef]
- Weber-Adrian, D.; Heinen, S.; Silburt, J.; Noroozian, Z.; Aubert, I. The human brain endothelial barrier: Transcytosis of AAV9, transduction by AAV2. J. Neurochem. 2016, 140, 192–194. [Google Scholar] [CrossRef]
- Wechsler, W. Die Entwicklung der Gefäße und pervivasculären Gewebsräume im Zentralnervensystem von Hühnern. Z. Anat. Entwicklungsgesch. 1965, 124, 367–395. [Google Scholar] [CrossRef]
- Shen, Y.; Maupetit, J.; Derreumaux, P.; Tufféry, P. Improved PEP-FOLD approach for peptide and miniprotein structure prediction. J. Chem. Theory Comput. 2014, 10, 4745–4758. [Google Scholar] [CrossRef]
- Waterhouse, A.; Bertoni, M.; Bienert, S.; Studer, G.; Tauriello, G.; Gumienny, R.; Heer, F.T.; De Beer, T.A.P.; Rempfer, C.; Bordoli, L.; et al. SWISS-MODEL: Homology modelling of protein structures and complexes. Nucleic Acids Res. 2018, 46, W296–W303. [Google Scholar] [CrossRef]
- Stahl, P.J.; Cruz, J.C.; Li, Y.; Michael Yu, S.; Hristova, K. On-the-resin N-terminal modification of long synthetic peptides. Anal. Biochem. 2012, 424, 137–139. [Google Scholar] [CrossRef]
- Pohl, M.; Ambrosius, D.; Grötzinger, J.; Kretzschmar, T.; Saunders, D.; Wollmer, A.; Brandenburg, D.; Bitter-Suermann, D.; Höcker, H. Cyclic disulfide analogues of the complement component C3a. Synthesis and conformational investigations. Int. J. Pept. Protein Res. 1993, 41, 362–375. [Google Scholar] [CrossRef] [PubMed]
- Schindelin, J.; Rueden, C.T.; Hiner, M.C.; Eliceiri, K.W. The ImageJ ecosystem: An open platform for biomedical image analysis. Mol. Reprod. Dev. 2017, 82, 518–529. [Google Scholar] [CrossRef] [PubMed]
- Roehrl, M.H.A.; Wang, J.Y.; Wagner, G. A general framework for development and data analysis of competitive high-throughput screens for small-molecule inhibitors of protein-protein interactions by fluorescence polarization. Biochemistry 2004, 43, 16056–16066. [Google Scholar] [CrossRef] [PubMed]
- Agilent Technologies AAV Helper-Free System Instruction Manual. Available online: https://www.agilent.com/cs/library/usermanuals/Public/240071.pdf (accessed on 13 August 2019).
- Zolotukhin, S.; Byrne, B.J.; Mason, E.; Zolotukhin, I.; Potter, M.; Chesnut, K.; Summerford, C.; Samulski, R.J.; Muzyczka, N. Recombinant adeno-associated virus purification using novel methods improves infectious titer and yield. Gene Ther. 1999, 6, 973–985. [Google Scholar] [CrossRef] [PubMed]
- Maniatis, T.; Fritsch, E.F.; Sambrook, J. Molecular Cloning: A Laboratory Manual; 9. Print; Cold Spring Harbor Laboratory Press: Cold Spring Harbor, NY, USA, 1984; ISBN 0-87969-136-0. [Google Scholar]


| Channel | Color | Laser line | Detector Range |
|---|---|---|---|
| NucBlue (Hoechst 33342) | Blue | 405 nm | 414–417 nm (Ch1) |
| 5(6)-Carboxyfluorescein | Green | 488 nm (argon) | 499–553 nm (ChS1) |
| Lysotracker-DND99 | Red | 594 nm | 609–700 nm (Ch2) |
| Name | Sequence 5′-3′ |
|---|---|
| βActin (gallus gallus) for | ATTGCCCCACCTGAGCGCAA |
| β-Actin (gallus gallus) rev | CATCTGCTGGAAGGTGGACA |
| β-Actin (human) for | GCTCCTCCTGAGCGCAAG |
| β-Actin (human) rev | CATCTGCTGGAAGGTGGACA |
| mVenus for | AAACTGATTTGCACCACCGG |
| mVenus rev | GCAAAGCATTGCAGGCCATA |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Feiner, R.C.; Kemker, I.; Krutzke, L.; Allmendinger, E.; Mandell, D.J.; Sewald, N.; Kochanek, S.; Müller, K.M. EGFR-Binding Peptides: From Computational Design towards Tumor-Targeting of Adeno-Associated Virus Capsids. Int. J. Mol. Sci. 2020, 21, 9535. https://doi.org/10.3390/ijms21249535
Feiner RC, Kemker I, Krutzke L, Allmendinger E, Mandell DJ, Sewald N, Kochanek S, Müller KM. EGFR-Binding Peptides: From Computational Design towards Tumor-Targeting of Adeno-Associated Virus Capsids. International Journal of Molecular Sciences. 2020; 21(24):9535. https://doi.org/10.3390/ijms21249535
Chicago/Turabian StyleFeiner, Rebecca C., Isabell Kemker, Lea Krutzke, Ellen Allmendinger, Daniel J. Mandell, Norbert Sewald, Stefan Kochanek, and Kristian M. Müller. 2020. "EGFR-Binding Peptides: From Computational Design towards Tumor-Targeting of Adeno-Associated Virus Capsids" International Journal of Molecular Sciences 21, no. 24: 9535. https://doi.org/10.3390/ijms21249535
APA StyleFeiner, R. C., Kemker, I., Krutzke, L., Allmendinger, E., Mandell, D. J., Sewald, N., Kochanek, S., & Müller, K. M. (2020). EGFR-Binding Peptides: From Computational Design towards Tumor-Targeting of Adeno-Associated Virus Capsids. International Journal of Molecular Sciences, 21(24), 9535. https://doi.org/10.3390/ijms21249535






