Abstract
The mitochondrial genome of the nematode Romanomermis culicivorax encodes for miniaturized hairpin-like tRNA molecules that lack D- as well as T-arms, strongly deviating from the consensus cloverleaf. The single tRNA nucleotidyltransferase of this organism is fully active on armless tRNAs, while the human counterpart is not able to add a complete CCA-end. Transplanting single regions of the Romanomermis enzyme into the human counterpart, we identified a beta-turn element of the catalytic core that—when inserted into the human enzyme—confers full CCA-adding activity on armless tRNAs. This region, originally identified to position the 3′-end of the tRNA primer in the catalytic core, dramatically increases the enzyme’s substrate affinity. While conventional tRNA substrates bind to the enzyme by interactions with the T-arm, this is not possible in the case of armless tRNAs, and the strong contribution of the beta-turn compensates for an otherwise too weak interaction required for the addition of a complete CCA-terminus. This compensation demonstrates the remarkable evolutionary plasticity of the catalytic core elements of this enzyme to adapt to unconventional tRNA substrates.
1. Introduction
tRNAs are the essential adaptor molecules which enable the decoding of the nucleic acid code into the amino acid sequence during the translational process [1]. To fulfill this function, they need to undergo several maturation steps and interact with the translational machinery [2,3,4,5]. In eukaryotes, this also includes the corresponding enzymes and proteins of mitochondria and chloroplasts [6,7]. For the efficient recognition by a wide range of processing enzymes, translation factors as well as ribosomes, tRNAs fold into a conserved cloverleaf-like secondary structure consisting of acceptor stem, anticodon arm as well as D- and T-arm that adopts an equally conserved three-dimensional L-shape [8,9,10]. The 3′-terminal CCA-triplet of the acceptor stem is a prerequisite for aminoacylation and the correct positioning of the charged tRNA in the ribosome [11,12]. In eukaryotes, this triplet is not genomically encoded but is added post-transcriptionally by tRNA nucleotidyltransferase (CCA-adding enzyme) [13,14,15].
tRNA nucleotidyltransferases represent essential enzymes and are ubiquitously found in all domains of life. Representing members of the polymerase β superfamily, they split up into two classes, based on the composition of their catalytic core [16]. Archaeal CCA-adding enzymes represent class I, while their bacterial and eukaryotic counterparts belong to class II [16]. The overall sequence identity among both tRNA nucleotidyltransferase classes is rather low, although they catalyze the same reaction [17]. The catalytic core motif common in both classes consists of two aspartate residues DxD (x, any amino acid) that coordinate the catalytically important metal ions [18,19,20]. In class II enzymes, the DxD sequence belongs to motif A, one of the five conserved motifs A to E located in the N-terminal part of this tRNA nucleotidyltransferase type (Figure 1) [21]. Motif A binds two Mg2+ ions required for nucleotide transfer onto the tRNA substrate via the general two metal ion mechanism of polymerases [19]. Motif B is involved in ribose binding [21], while motif C is a flexible element which coordinates interdomain movements, contributing to the proper orientation of the substrates within the active center [22,23]. Motif D represents an amino acid-based template, where arginine and aspartate residues form Watson–Crick-like hydrogen bonds with the incoming nucleotides, and their orientation in the catalytic core determines the specificity for CTP or ATP, respectively [21]. Lastly, motif E stabilizes a helix-turn structure in motif D and is discussed to interact with the tRNA primer strand in the catalytic core [21,24].

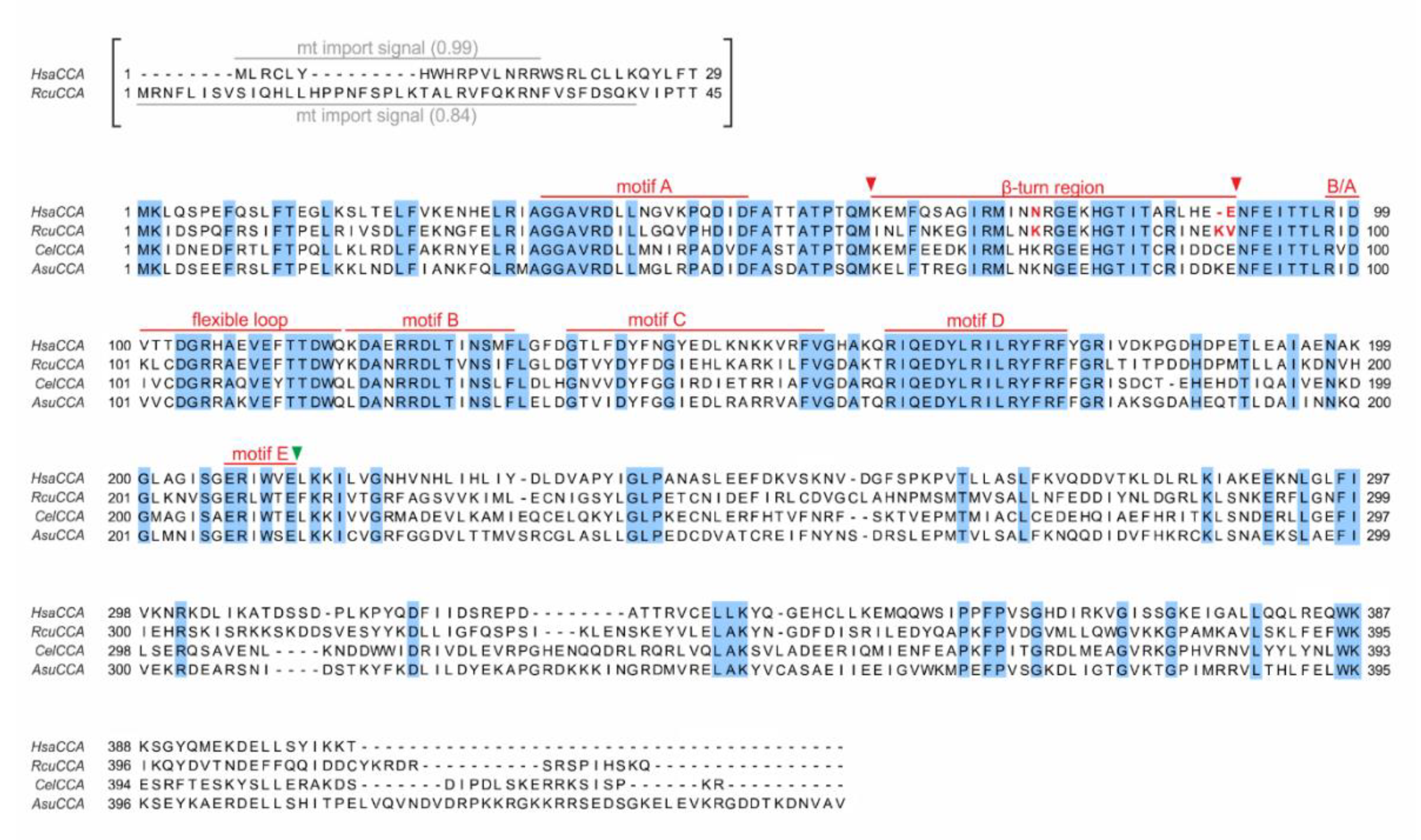
Figure 1.
Sequence alignment of CCA-adding enzymes from Homo sapiens, Romanomermis culicivorax, Caenorhabditis elegans, and Ascaris suum. Light blue positions indicate identical residues. The predicted mitochondrial import signals for H. sapiens CCA (HsaCCA) and the R. culicivorax CCA-adding enzyme (RcuCCA) (grey bars, import probability is given in brackets) are shown in brackets and were excluded from the cloned open reading frames. Catalytically important elements are labeled in red. Fusion position of reciprocal chimeras A and B are indicated by a green arrowhead. Fusion positions of chimera E (β-turn element) are indicated by red arrowheads (K/I61–E/V90). Mutations K74N and K89Δ/V90E introduced in RcuCCA and N74K and K89ins/E90V in HsaCCA are indicated in red.
Another catalytically important region is a flexible loop consisting of 10–20 amino acids that is located immediately upstream of motif B. While it is not conserved at the sequence level [25], its interaction with the amino acid template of motif D is required for the specificity switch from C to A incorporation, where it acts as a lever to accommodate the ATP in the nucleotide binding pocket [22,26]. Between motif A and the flexible loop, a β-turn element was identified that is also involved in A-addition, as it binds and positions the priming 3′-end of the growing CCA terminus in the catalytic core [22]. Similar β-turn regions are present in many different polymerases, ranging from both classes of tRNA nucleotidyltransferases to poly(A) polymerases and DNA polymerases, underscoring the catalytically important function of this element [27,28,29,30]. For a structural view of the reaction mechanism, we refer to Tomita and Yamashita, 2014 [31].
All these conserved motifs build up the active site in the N-terminal part of the enzyme. The C-terminus, in contrast, is much less conserved. Yet, it is of functional importance, as it is involved in tRNA binding, where it anchors the TΨC loop of the L-shaped tRNA substrate during nucleotide addition [24,32,33,34,35]. Correspondingly, artificial CCA-adding substrates like mini- or microhelices are recognized and accepted for CCA incorporation at a much lower efficiency [36,37]. Yet, metazoan mitochondria carry tRNA molecules that deviate from the cloverleaf structure, lacking either the D- or the T-arm [38,39,40]. As an example, the mammalian mt-tRNASer(AGY) lacks the complete D-arm, so that D- and T-arm interactions do not exist [41,42]. In the mitochondrial genomes of nematodes, acari and arachnids, this situation comes to an extreme. These genomes are rich in genes for tRNAs that lack either the D- or the T-arm or even both [38,43,44,45,46]. In the mermithid Romanomermis culicivorax, mt-tRNA molecules with the most dramatic truncations were identified, resulting in miniaturized hairpin-like tRNAs with a length of down to 45 nts, in contrast to the standard average tRNA size of 76 nts [47,48]. Such extremely truncated tRNAs fold into a three-dimensional boomerang-like shape that deviates from the consensus L-form [47]. Yet, these organisms encode for a single CCA-adding enzyme that has to act on both cytosolic as well as mitochondrial tRNA pools [49,50,51,52,53], and it was shown for the corresponding enzyme of Caenorhabditis elegans that it recognizes mt-tRNAs lacking D- or T-arm [52]. Since in R. culicivorax nine mt-tRNAs lack both arms, representing the strongest deviation from the consensus structure [45], we investigated the co-evolution and substrate adaptation of its CCA-adding enzyme. In a comparative analysis, we identified the β-turn element as a major adaptation to the hairpin-like tRNA substrates. This adaptation is based on an increased substrate affinity of the enzyme. Hence, while the conventional substrate binding based on interactions between the enzyme’s C-terminus and the TΨC loop/T-arm is not possible with such tRNA hairpins, a different part of the enzyme took over this function to assure a sufficiently strong substrate binding for CCA-addition, demonstrating a surprising evolutionary plasticity of these enzymes.
2. Results
2.1. RcuCCA Adds a Complete CCA-Triplet to Armless and Canonical tRNAs
To investigate the catalytic activity and substrate specificity of the R. culicivorax CCA-adding enzyme (RcuCCA), we identified a corresponding singular open reading frame in the R. culicivorax genome assembly available at https://parasite.wormbase.org/Romanomermis_culicivorax_prjeb1358/Info/Index/ [54]. The sequence conservation was investigated in an alignment with the corresponding human enzyme (HsaCCA). As it was reported that the CCA-adding enzyme of Caenorhabditis elegans is adapted to the bizarre mitochondrial tRNAs of this organism [52], the sequence of this enzyme as well as that of Ascaris suum (another nematode with mt-tRNAs lacking D- or T-arm [38,39,43]) were included (Figure 1). At the amino acid level, the overall sequence identity and similarity between RcuCCA and HsaCCA is 48% and 66%, respectively. Carrying the complete active site as well as a putative mitochondrial import sequence as predicted [55], the N-terminus shows a comparatively higher conservation (78% sequence similarity), as it is true for the corresponding enzymes of C. elegans and A. suum (Figure 1). In the less conserved C-terminal part, RcuCCA and the other nematode enzymes carry a short insertion and a terminal extension.
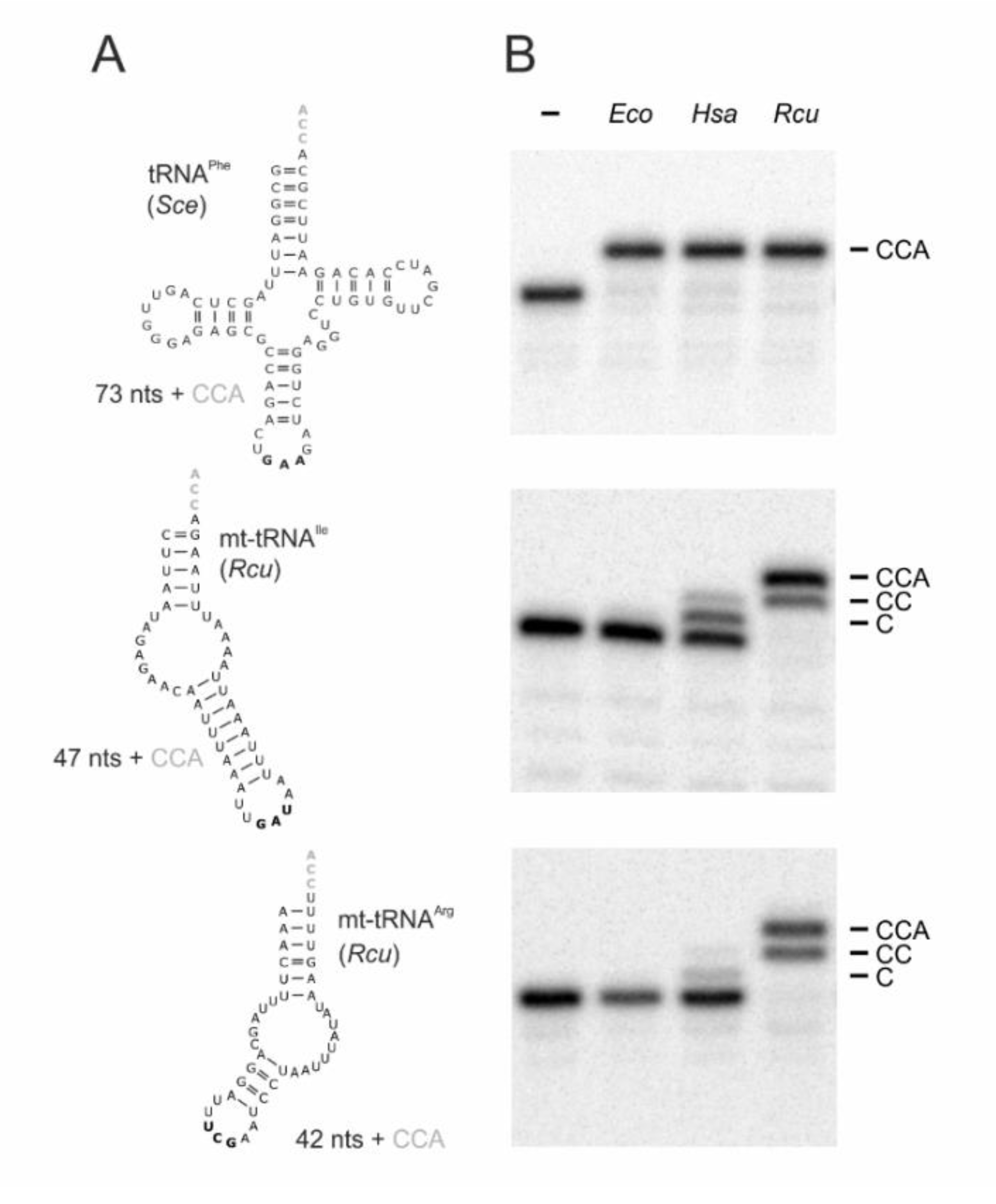
A conserved methionine residue downstream of the mt import signal was chosen as the N-terminus of the expressed open reading frame (labeled as position 1 in Figure 1). In previous experiments on the human enzyme, this position was successfully used and the absence of the mt import signal had no effect on its catalytic activity [37,50,56,57]. The open reading frame was synthesized as a codon-optimized DNA sequence and recombinantly expressed in Escherichia coli. Together with the corresponding enzymes from E. coli (EcoCCA; an organism exclusively carrying conventional cloverleaf-like tRNAs) and Homo sapiens (HsaCCA; an organism carrying conventional cytosolic as well as moderately reduced mt-tRNAs), the purified enzyme was tested in vitro for activity. As substrates, three different radioactively labeled tRNA transcripts were generated by in vitro transcription (Figure 2A), as it is well established that tRNA nucleotidyltransferases from all kingdoms readily accept in vitro transcripts lacking base modifications [25,58,59,60]. tRNAPhe from Saccharomyces cerevisiae is one of the best characterized tRNAs and represents a standard substrate for many tRNA-interacting enzymes [58,61,62], since the unmodified in vitro transcript folds into a structure almost identical to the native tRNA [63,64]. Furthermore, two armless mt-tRNAs from R. culicivorax were generated. With a length of 42 nts and 47 nts, respectively, mt-tRNAArg and mt-tRNAIle represent the shortest tRNAs identified so far, and the in vitro transcripts fold into hairpin-like structures with two single-stranded connector elements replacing D- and T-arm (Figure 2A) [47]. On the standard tRNAPhe, all enzymes added a complete CCA-triplet, indicating highly active enzyme preparations (Figure 2B). On the armless mt-tRNA substrates, however, the bacterial enzyme EcoCCA was completely inactive and did not add any nucleotides. The human enzyme HsaCCA that has to recognize the human D-arm-lacking mt-tRNASer(AGY) (Figure S1) catalyzed a moderate incorporation of two residues on the armless Rcu mt-tRNAIle, but was almost inactive on the even smaller Rcu mt-tRNAArg. In contrast, RcuCCA added complete CCA-triplets, regardless whether the substrate represented a conventional cloverleaf-structured tRNA or an armless hairpin-like tRNA, indicating an efficient adaptation to these miniaturized substrates (Figure 2B).
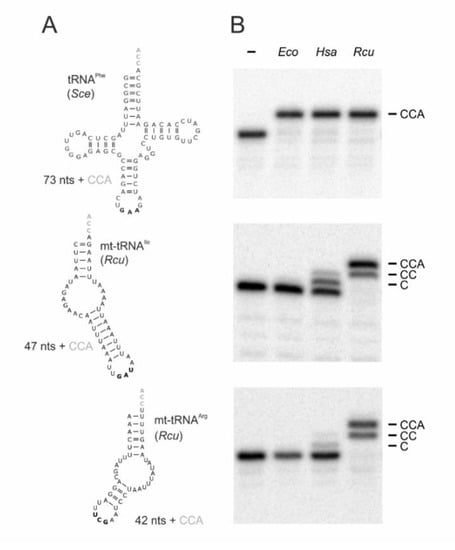
Figure 2.
CCA-addition on conventional and hairpin-like tRNA substrates. (A) tRNAPhe from Saccharomyces cerevisiae (Sce) represents a conventionally structured tRNA substrate of standard size (73 nts without CCA), while the mitochondrial tRNAs for isoleucine and arginine from R. culicivorax (Rcu) considerably deviate in size (47 and 42 nts, respectively; both without 3′-terminal CCA-triplet) and structure, lacking both D- and T-arms. Anticodons are indicated in bold. (B) CCA-addition on radioactively labeled tRNA transcripts catalyzed by the corresponding enzymes (20 ng each) of Escherichia coli (Eco), H. sapiens (Hsa), and R. culicivorax (Rcu). Incubation without enzymes represent negative controls (−). All enzymes completely convert the canonical tRNAPhe from S. cerevisiae into a mature transcript with CCA-end. On armless mt-tRNAs, the E. coli enzyme shows no activity at all, while the human enzyme adds only one or two C residues to mt-tRNAArg and mt-tRNAIle, respectively. In contrast, the enzyme of R. culicivorax readily synthesizes a complete CCA-end on both transcripts, although the time of incubation was not sufficient for 100% A-addition. The experiment was done in three independent replicates. The panel shows a representative autoradiogram.
To investigate the substrate preferences of HsaCCA and RcuCCA in more detail, a Michaelis–Menten kinetics analysis was performed. Due to the limited RNA solubility, excessive saturating amounts of tRNA cannot be used, and the obtained parameters represent apparent values [24,25,30,58,65]. As EcoCCA showed no activity on armless tRNAs, this enzyme was excluded from further analysis. To discriminate between C- and A-addition, assays were performed on tRNAs lacking the CCA-end in the presence of either α-32P-CTP or α-32P-ATP and unlabeled NTPs [23,37,66]. As shown in Table 1, the kinetic parameters of both enzymes on tRNAPhe are rather similar for CC-incorporation, and RcuCCA is somewhat less efficient (0.5-fold) in adding the complete CCA terminus (CCA*). On the armless mt-tRNAs for isoleucine and arginine, HsaCCA and RcuCCA are almost equally active in CC-addition, with somewhat higher values for RcuCCA (1.2 to 1.7x). However, when the complete CCA-incorporation is investigated (CCA*), the Romanomermis enzyme is much more efficient on mt-tRNAIle and mt-tRNAArg (3.3 to 10x). As the obtained kcat values for HsaCCA in this reaction are at the detection limit, the actual efficiency of RcuCCA relative to HsaCCA is probably much higher.

Table 1.
Kinetic parameters of HsaCCA and RcuCCA for CC- and A-addition. As indicated by the relative activity of RcuCCA compared to HsaCCA (change), CC-addition on standard and armless tRNA substrates is similar. In terminal A-addition on the armless tRNAs, RcuCCA shows a 3.3 to 10-fold increase, a clear indication of its adaptation to these substrates. HsaCCA is strongly affected in this reaction, and the obtained low values likely represent an overestimation, as they are close to the detection limit. The actual values are probably much lower. For each analysis, three independent experiments were performed.
Taken together, both enzymes prefer a conventionally structured tRNA as substrate. On the hairpin-like tRNAs, RcuCCA still adds complete CCA-ends, although at a lower efficiency for mt-tRNAArg. The human enzyme, however, strongly prefers the conventional tRNA and is severely affected in A-addition on armless tRNAs, resulting in incomplete and hence non-functional tRNA molecules as already seen in CCA-addition assay (Figure 2B).
2.2. In the Romanomermis Enzyme, Especially the Catalytic Core is Adapted to Armless tRNA Substrates
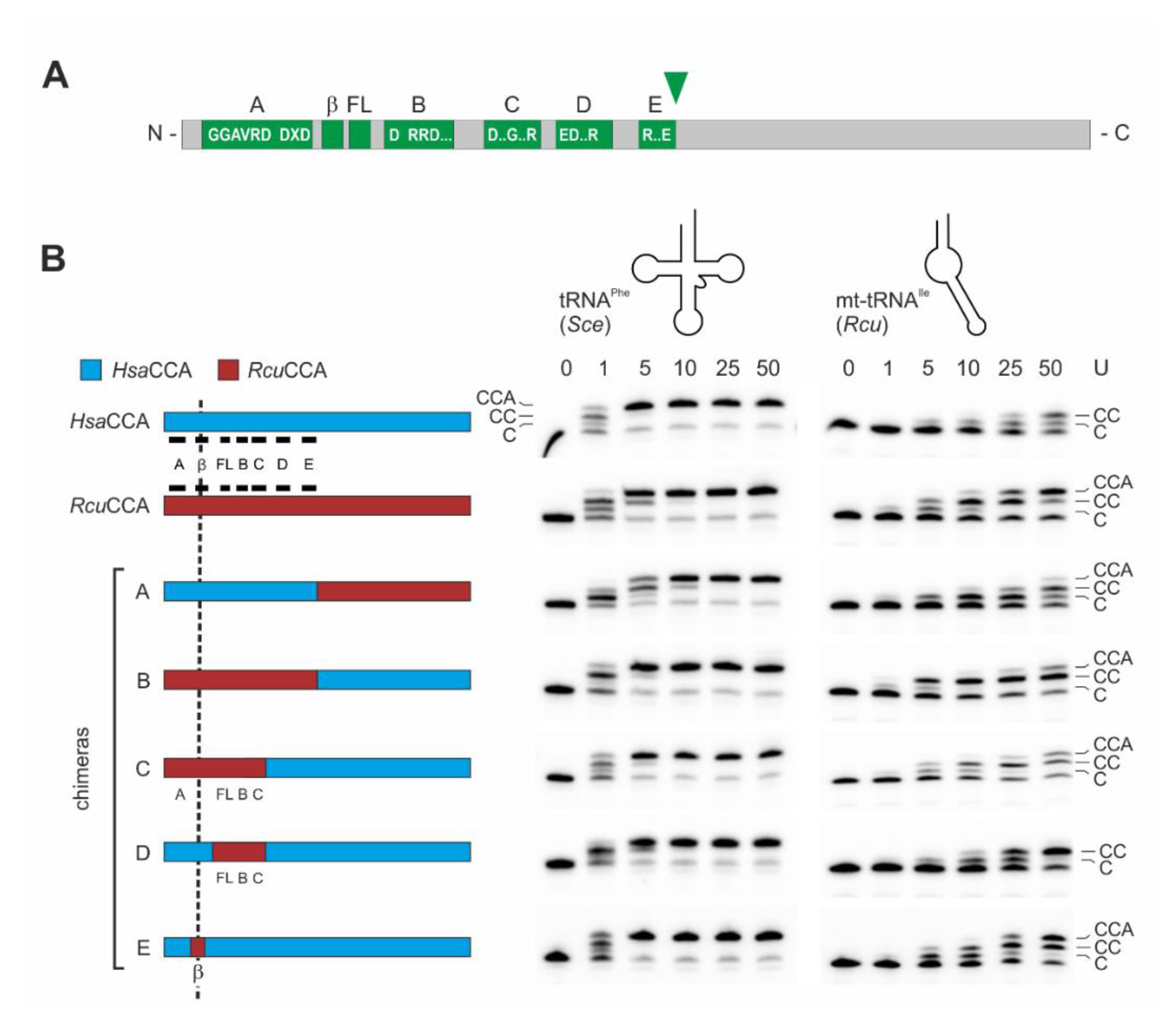
To identify the contribution of individual enzyme regions to the recognition of armless tRNAs as substrates for CCA-addition, we followed a strategy that we successfully applied to investigate several CCA-adding enzymes concerning their enzymatic features [33,58,67]. We reciprocally exchanged N- and C-termini of HsaCCA and RcuCCA, carrying the complete catalytic core and the region involved in tRNA binding, respectively. Based on the sequence alignment shown in Figure 1, we selected a glutamate at position 212 (HsaCCA) and 213 (RcuCCA), representing the last invariant residue of motif E [68] as fusion position (Figure 3A). To allow for a direct comparison of enzymatic activities of the resulting proteins, we adjusted the efficiency of CCA-addition on the canonically structured tRNAPhe substrate and defined an arbitrary unit as the amount of enzyme required for 50% substrate turnover, ranging between 0.3 ng for wild type (wt) enzymes and 0.5 to 1.3 ng for chimeras. Incubation of the standard substrate tRNAPhe with 1 to 50 arbitrary units of both wt enzymes as well as chimera A (N-terminal catalytic core of RcuCCA, C-terminus of HsaCCA) and the reciprocal chimera B indicate that all enzymes are fully active and efficiently synthesize a complete CCA-terminus (Figure 3B, left gel panel). On the minimalized mt-tRNAIle (Figure 3B, right gel panel), even 50 units of the human wt enzyme added only two C residues. In contrast, the same amount of Romanomermis enzyme added a complete CCA-end. Surprisingly, chimera B (with the catalytic core of RcuCCA) synthesized a complete CCA-end at considerable efficiency, while identical units of chimera A, carrying the tRNA-binding C-terminus of RcuCCA, catalyzed the full CCA-addition to a lesser extent.
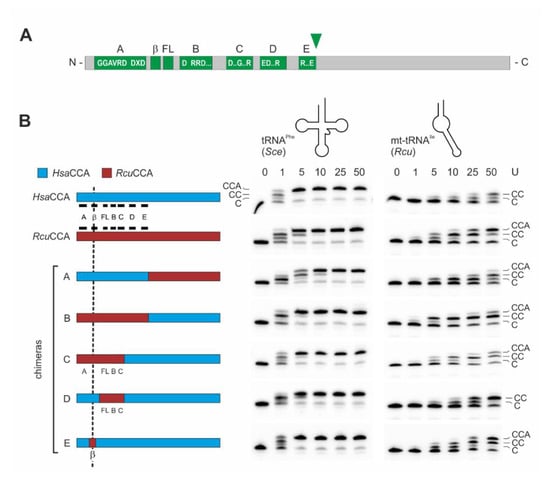
Figure 3.
Catalytic activity of wild-type and chimeric enzymes on tRNAPhe and mt-tRNAIle. (A) Bar representation of class II CCA-adding enzymes. The N-terminal region contains the catalytic core consisting of five motifs A to E, a β-turn element (β) and a flexible loop (FL). The green arrowhead indicates the fusion position for enzyme chimeras A and B. (B) Left: Bar representation of tested enzyme chimeras consisting of HsaCCA (cyan) and RcuCCA (red) regions. Catalytic core elements are indicated in black. The reciprocal chimeras A and B are fused after position E213, downstream of motif E. The replaced β-turn element (β) in chimera E is located between motif A and the flexible loop (FL). Gel panels: CCA-addition on the conventional tRNAPhe from yeast (Sce, left) and the armless mt-tRNAIle from R. culicivorax (Rcu, right) with increasing amounts of enzymes indicated as arbitrary units (U). All enzymes catalyze an efficient CCA-incorporation on the conventional tRNA. On the armless tRNAIle, the Romanomermis wt enzyme synthesizes a complete CCA-end, while the corresponding human enzyme adds only two C-residues. Chimeras A and B also add the terminal A, but at a somewhat reduced level. On this tRNA substrate, chimera B, carrying the catalytic core of RcuCCA, is more efficient than chimera A, where A-incorporation is only visible at the highest enzyme concentration. Chimera C still shows full CCA-addition, whereas no terminal A-incorporation is observed for chimera D. Chimera E shows an efficiency comparable to that of the RcuCCA wt enzyme, indicating the importance of the β-turn region in the reaction on armless tRNAs. The fact that chimera E is more active than chimera B likely reflects differences in the compatibility of the chosen fusion positions in these chimeras, resulting in different protein folding and/or catalytic efficiency. For each construct, up to four independent experiments were performed. For each tRNA substrate, a representative gel is shown.
While these reactions indicate that both enzyme parts participate in the adaptation to the armless tRNA substrates, the contribution of the catalytic core seems to have a greater impact on the acceptance of these substrates. Based on these results, we generated a series of chimeric enzymes carrying parts of the RcuCCA catalytic core in the context of the human enzyme (Figure 3B, Table S1). For chimera C, the human C-terminal part was extended to include motifs D and E. Comparable to chimera B, this construct added the full CCA-end to Rcu mt-tRNAIle. As the flexible loop (FL) and motif C are involved in the enzymes’ specificity switch to A-addition [22,23,26], chimera D was constructed, consisting of HsaCCA where the region spanning these two elements was replaced by the corresponding part from RcuCCA. While this enzyme was fully active on tRNAPhe, the loop and motif C replacement did not result in A-addition on mt-tRNAIle. These results led to the conclusion that an element upstream of the flexible loop must be responsible for A-addition on armless tRNA substrates. As this region contains a small β-turn element that was described to contribute to tRNA-CC primer positioning and, as a consequence, A-addition [22], chimera E was constructed, where a stretch of 30 amino acid residues (positions 61–91) carrying this element was replaced in the backbone of the human enzyme. The resulting enzyme catalyzed CCA-addition on both tRNAPhe and mt-tRNAIle at efficiencies comparable to the wt RcuCCA enzyme (Figure 3B). Hence, this divide-and-conquer approach allowed us to identify the β-turn as an important element involved in the adaptation of RcuCCA to the armless tRNA substrates.
2.3. The β-Turn of the R. culicivorax CCA-Adding Enzyme Strongly Contributes to Substrate Binding and CCA Incorporation on Armless tRNAs
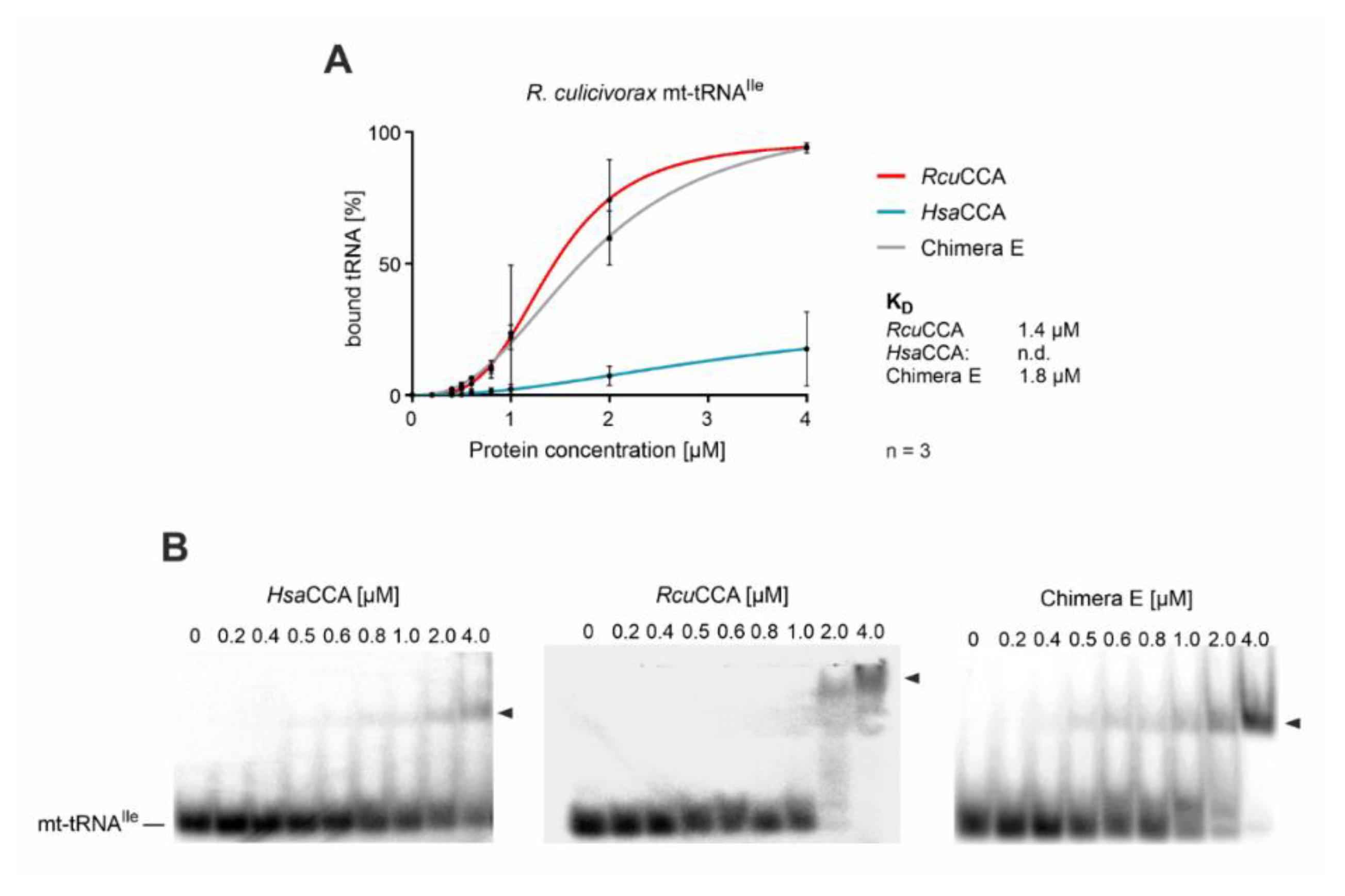
To investigate whether the β-turn of RcuCCA contributes to an increased efficiency in binding of armless tRNA substrates, we performed electrophoretic mobility shift experiments on chimera E and the parental enzymes HsaCCA and RcuCCA with an armless tRNA transcript. Radioactively labeled mt-tRNAIle lacking the CCA terminus was incubated with increasing amounts of recombinantly expressed enzymes and separated by native polyacrylamide gel electrophoresis (Figure 4). Enzyme-bound and free substrates were visualized and binding parameters were determined by nonlinear regression. The Romanomermis enzyme showed an efficient and robust binding to mt-tRNAIle, resulting in a Kd value of 1.4 µM, while for the human enzyme, no significant binding could be detected, as previously reported for this enzyme class [23,33,69]. Interestingly, and concurrent with its wt-like activity on mt-tRNAIle (Figure 3B), chimera E also showed a high affinity for mt-tRNAIle at a Kd of 1.8 µM.
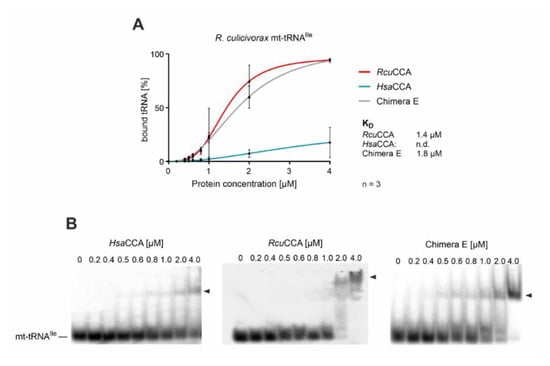
Figure 4.
Binding of wt and chimeric CCA-adding enzymes to an armless tRNA. Quantitative analysis of enzyme binding to the armless mt-tRNAIle determined by electrophoretic mobility shifts. (A) While the tRNA interaction of HsaCCA over the whole concentration range (0–4 µM) is too weak to calculate dissociation constants, RcuCCA as well as chimera E exhibit a strong affinity to this substrate, resulting in dissociation constants of 1.4 and 1.8 µM, respectively. Data are means ± SD; n = 3. (B) Images of representative gel shift assays on HsaCCA, RcuCCA, and chimera E with mt-tRNAIle as substrate.
When transcripts of mt-tRNAIle with different 3′-ends (mt-tRNAIle, mt-tRNAIle-C, mt-tRNAIle-CC, mt-tRNAIle-CCA) were offered to RcuCCA, the enzyme bound all substrates with similar affinity, indicating that the composition of the tRNA 3′-end does not affect this interaction (Figure S2A). The human enzyme, in contrast, did not show efficient binding to any of these substrates. Gel shift experiments on the conventional substrate tRNAPhe indicate that this high affinity of the RcuCCA enzyme is not restricted to armless tRNA substrates (Figure S2B). With a binding constant of 1.3 µM, the enzyme exhibits a similar binding behavior to the cloverleaf-shaped tRNA as it does on the armless mt-tRNAIle (Kd = 1.4 µM), while the human enzyme again shows almost no interaction. Hence, the adaptation of the Romanomermis culicivorax CCA-adding enzyme to process both cloverleaf-structured cytosolic as well as armless mitochondrial tRNAs is obviously achieved by a general exceptional tight interaction with its substrates, regardless whether they represent canonical or minimalized tRNAs.
While the β-turn itself (GEKH) is identical in RcuCCA and HsaCCA, the flanking sequences in the RcuCCA enzyme differ in 13 positions from the counterpart, and some of them likely contribute to the extended substrate specificity of this enzyme. In these regions, the Romanomermis enzyme carries several lysine residues that are not present in HsaCCA (Figure 5). K74 is also found in the corresponding enzymes of C. elegans and A. suum, while the human enzyme carries an asparagine at this position (Figure 1). As the basic lysine might enhance the enzymes’ binding to negatively charged tRNA substrates, it represents a top candidate for such a substrate adaptation. At positions 89 and 90, RcuCCA carries a further lysine residue followed by a valine position that might also be involved in tRNA primer binding, although these residues are not present in CelCCA and AsuCCA. Hence, we replaced these positions in RcuCCA by the corresponding residues of the human enzyme, resulting in RcuCCA K74N, K89Δ/V90E, and the combination thereof (K74N/K89Δ/V90E). The CCA-adding activity of the recombinantly expressed enzyme variants was determined on yeast tRNAPhe and adjusted to arbitrary units (1 U corresponding to 0.3 ng protein; Figure 5D,E). On this conventional tRNA, all enzymes showed a comparable activity, with a slight reduction in activity of RcuCCA K74N (Figure 5D, upper panel). On mt-tRNAIle, however, the variants carrying the K74N replacement were considerably affected in incorporating the terminal A residue. RcuCCA K89Δ/V90E, in contrast, added a full CCA-end on this tRNA, although it seems to be somewhat less efficient than the wt enzyme. The reciprocal amino acid exchanges and insertions introduced in the human enzyme support these results (Figure 5E). HsaCCA N74K (for reasons of clarity, numbering is according to RcuCCA positions) shows an efficient addition of a complete CCA-end on mt-tRNAIle. Intriguingly, also HsaCCA 89Kins/E90V and the combination N74K/K89ins/E90V exhibit improved CCA-addition on this armless substrate. As a conclusion, K74, and to a certain extent also K89/V90, represent important positions to accept armless tRNA substrates and probably contribute to a stable interaction with the tRNA and the correct positioning of its 3′-end as a primer.

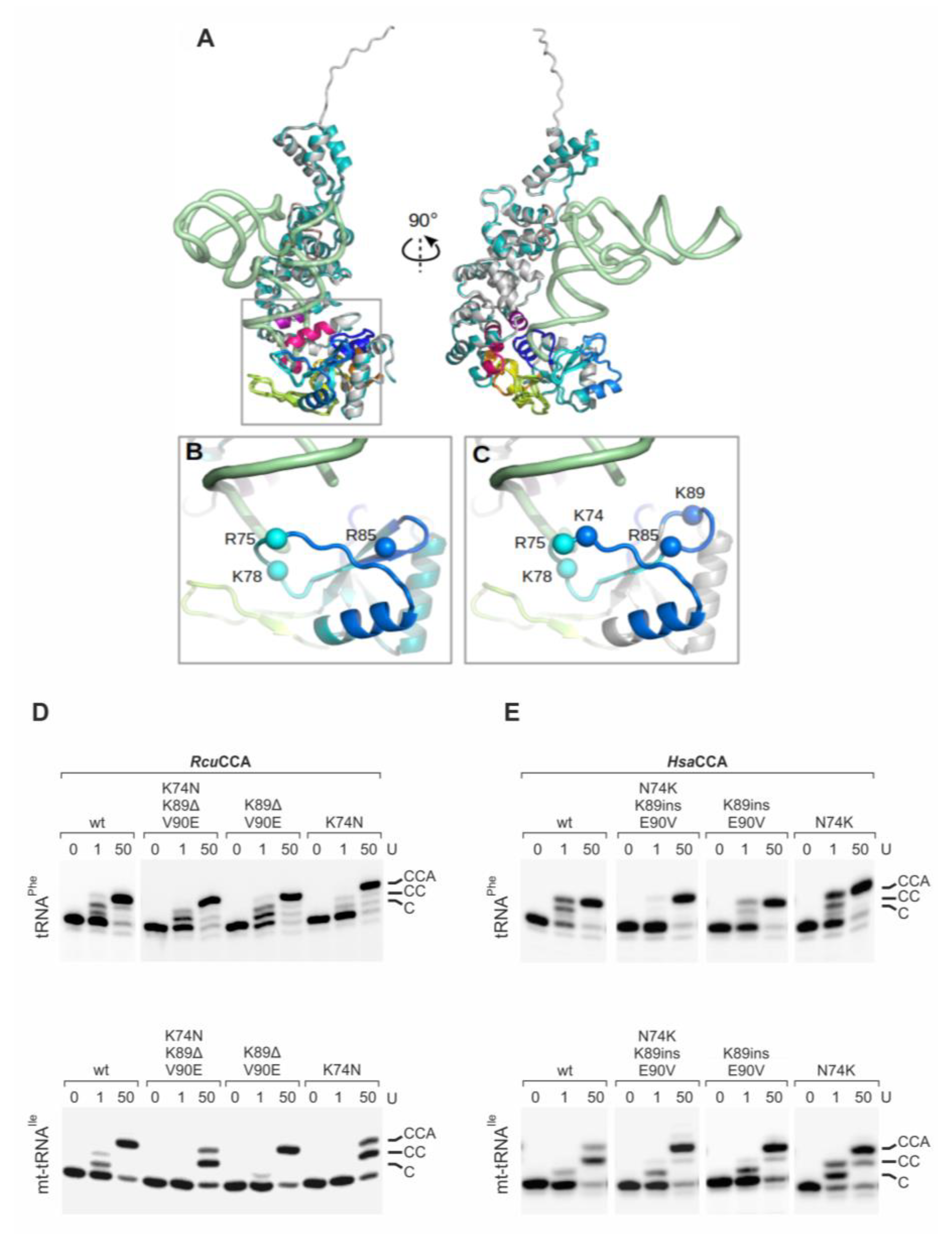
Figure 5.
The β-turn in HsaCCA and RcuCCA enzymes. (A) Superimposed full-length models of HsaCCA (cyan) and RcuCCA (light gray) with the backbone of a bound tRNA (green) in two perpendicular views. The tRNA position was obtained by superimposing the A-adding enzyme:tRNA-CC complex from Aquifex aeolicus onto the human enzyme [34,35,70]. Motif A (dark blue), β-turn region (medium blue), β-turn (light blue), B/A motif and flexible loop (green), motif B (yellow), motif C (orange), motif D (red) and motif E (violet) are indicated. (B,C). Zoom into the β-turn region and the tRNA 3′-end (corresponding to the squared region in (A) of HsaCCA (B) and RcuCCA (C). Spheres represent the Cα positions of positively charged residues (K and R). RcuCCA carries two additional lysines at positions 74 and 89 that might contribute to tRNA binding and primer positioning. Model is based on the crystal structure of the A-adding enzyme from Aquifex aeolicus [34]. (D) Enzymatic activity of RcuCCA carrying mutations K74N, K89Δ/V90E and K74N/K89Δ/V90E. 0, 1, and 50 arbitrary units of enzyme variants were incubated with yeast tRNAPhe and the armless mt-tRNAIle. RcuCCA wt accepts both tRNAs for CCA-addition, while RcuCCA K74N is less active on mt-tRNAIle, resulting in a considerably reduced A-addition. In contrast, RcuCCA K89Δ/V90E catalyzes full CCA-addition on the conventional (comparable to wt activity) as well as on the armless tRNA. The triple variant RcuCCA K74N/K89Δ/V90E shows the same activity as RcuCCA K74N. (E) The introduction of the corresponding amino acids of RcuCCA into HsaCCA enables this enzyme to add a complete CCA-end on mt-tRNAIle, in contrast to the wildtype situation. These results indicate that especially position K74 of the β-turn, but to a certain extent also K89/V90, contribute to the substrate adaptation of RcuCCA.
3. Discussion
3.1. A Specific Adaptation within the Catalytic Core Enables CCA-Addition to Minimalized tRNA Substrates
As the tRNA genes of most organisms do not encode the 3′-terminal CCA-triplet, this essential feature has to be added post-transcriptionally by the CCA-adding enzyme. In eukaryotes, a single enzyme is responsible for this maturation step in both cytosolic as well as organellar tRNA pools [50,51,71]. A second function of CCA-adding enzyme is to monitor the intactness of its tRNA substrate, so that only undamaged molecules are accepted for CCA-incorporation [72,73,74,75,76]. The CCA-adding enzyme does not recognize a specific sequence or base pair in its substrates, but relies on common elements of the overall tRNA cloverleaf structure, as it is also observed for RNase P, tRNase Z, and some tRNA modifying enzymes [77,78,79,80,81]. Metazoan mitochondria, however, encode for tRNAs with deviations from the cloverleaf, where D- or T-arms are reduced or lacking [38,41,48]. As these tRNAs still carry a conventional acceptor stem, they are correctly processed by RNase P, since this enzyme predominantly recognizes this structural feature [78,79]. The CCA-adding enzyme, in contrast, specifically interacts with the tRNA elbow region, and especially with the T-loop region [32,69,82,83,84]. Hence, a bacterial enzyme like the E. coli version, evolved for conventional tRNAs, is strongly impaired on truncated tRNAs [52]. The most extreme truncations are observed in nematodes like R. culicivorax, in acari and in arachnids, where hairpin-like tRNAs are found that require a specific co-evolution of the corresponding enzymes. On such substrates, the E. coli enzyme is completely inactive (Figure 2B). However, the human enzyme co-evolved to accept the D-armless tRNASerAGY found in human mitochondria (Figure S1) [41]. Hence, this enzyme accepts the armless tRNAs to a certain extent and adds two C-residues, but not the terminal A, indicating that A incorporation requires a specific adaptation to such extreme substrates (Figure 2B, Figure 3, Table 1). In contrast, the R. culicivorax enzyme is adapted to these tRNA structures and efficiently adds the complete CCA-triplet. For HsaCCA and RcuCCA, the kinetic parameters for CC-incorporation on standard as well as armless tRNAs are quite similar and correspond to published values [37]. However, when it comes to A-addition, only RcuCCA is adapted to the hairpin-like tRNAs (Figure 2B, Figure 3), although this reaction step is less efficient than C-addition (Table 1).
In the transition from C- towards A-addition, CCA-adding enzymes undergo domain rearrangements to accommodate the growing tRNA 3′-end in the catalytic core and to switch the specificity of the amino acid template in the nucleotide binding pocket from CTP towards ATP recognition [21,22,23,85]. During this structural rearrangement, the tRNA substrate has to remain bound to the enzyme, and this is usually accomplished by specific interactions of the tRNA 3′-end in the catalytic core and of the elbow region (T- and D-loop) with the C-terminal region of the CCA-adding enzyme [22,24]. For CCA-adding enzymes adapted to conventional tRNAs or tRNAs lacking only one arm, this interaction is not very tight, as no Kd values could be determined [23,33,69] — yet it is sufficient for a complete synthesis of the CCA-end. However, for armless tRNAs, these interactions seem to be insufficient, and the observed high substrate affinity of RcuCCA (in contrast to the human enzyme; Figure 4, Figure S2) corroborates this hypothesis. Obviously, the Romanomermis enzyme is able to bind its substrate very tightly, and since this is also the case for armless tRNAs, this interaction cannot involve the conventional contacts between C-terminus of the enzyme and T-loop of the tRNA but must be located elsewhere, although the C-terminus contributes to the terminal A-addition to a certain extent, as shown by the Rcu/HsaCCA chimera A (Figure 4).
In the detailed analysis of enzyme chimeras between RcuCCA and HsaCCA, an adaptation of both enzymes to different tRNA substrates is obvious. In HsaCCA and chimeras B, C, D, and E, the tRNA-binding C-terminus helps to confer an efficient CCA-addition on the conventional tRNAPhe (Figure 3B). As HsaCCA shows a rather weak interaction with tRNA substrates (Figure 4, Figure S2), this C-terminal part obviously contributes to an efficient CCA-synthesis by enabling the final product release [85]. In contrast, in RcuCCA, an adaptation to armless tRNA substrates in the N-terminus is obvious. Here, two elements of the catalytic core are described to play an important role in the specificity switch from C- to A-addition. The flexible loop acts as a lever that adjusts the templating amino acids for correct ATP binding [22,25,26,66], and motif C represents a springy hinge that supports the domain rearrangements involved in this reaction step [22,23]. Accordingly, one could expect that both elements are adapted to the CCA-addition on armless tRNAs. Yet, chimera D (human enzyme with flexible loop and motif C of RcuCCA) does not show any A-addition on the armless tRNA, while it is fully active on a standard substrate (Figure 3), excluding a specific adaptation of these two elements.
A detailed inspection of the residual N-terminal RcuCCA components in the chimeras revealed that constructs carrying a β-turn element located between strands 3 and 4 of the β-sheet in the catalytic core are able to add the terminal A, indicating that this element represents a major adaptation to armless tRNAs (Figure 3B, chimeras B, C, and E)
3.2. The ß-Turn Element Impacts the Substrate Affinity of R. culicivorax CCA-Adding Enzyme
In chimera E, the inserted β-turn element consists of 30 amino acids located between motif A and a conserved stretch upstream of the flexible loop (Figure 1). Toh et al., showed that in CCA-adding enzymes for conventional tRNAs, this region plays an important role in binding the growing CC-end of the tRNA and adjusting it in the catalytic core for the nucleophilic attack initiating A-addition [22]. Similar β-turn elements are found for the archaeal class I CCA-adding enzymes [29,30,86], poly(A) polymerases as well as for DNA- and RNA-polymerases, where they are described to position the 3′-hydroxyl of the primer 3′-end in close vicinity of the catalytically important metal ions located in the active site [68]. In RcuCCA, however, the β-turn element has an additional function, as it dramatically contributes to tRNA binding. Here, especially the lysine residue at position 74 (K74) seems to be involved, as its replacement by asparagine, the corresponding position in HsaCCA, affects A-addition on an armless tRNA (Figure 5D). The introduction of K74 into HsaCCA supports the importance of this lysine residue, as it enables a strongly improved CCA-addition on mt-tRNAIle by this enzyme. Intriguingly, also the HsaCCA K89ins/E90V variant shows an enhanced CCA-addition on mt-tRNAIle. This could be a general effect of the positively charged lysine side chain that keeps the tRNA in close proximity during polymerization, enabling a more efficient nucleotide transfer. In RcuCCA, these positions seem to be less important, as their replacement by the human residues have only very subtle effects on CCA-addition. It is conceivable that other positions not directly involved in tRNA binding might structurally support the substrate adaptation of RcuCCA by folding the loop region into a conformation that optimally positions K74 (and other interacting residues) for primer binding during polymerization. Corresponding crystal structures of RcuCCA in complex with an armless tRNA are needed to clarify this point.
The presented results on RcuCCA, HsaCCA, and the corresponding chimeras support the following hypothesis. As described above, the binding of the tRNA’s T-loop to its C-terminus represents one of the major substrate interactions of the CCA-adding enzyme, ensuring that the tRNA primer remains correctly located for A-addition during the domain rearrangements inducing the specificity switch [34,35]. As a consequence, product release seems to be a limiting factor in the reaction [85]. As this interaction is not possible with armless tRNAs, the human enzyme loses contact after adding the C residues, while a conventional tRNA remains correctly bound in the T-loop/C-terminus interaction and gets elongated by a complete CCA-terminus. In R. culicivorax, the CCA-adding enzyme had to adapt to armless tRNAs and evolved a different mode of tRNA interaction, as a binding of T-loop and C-terminus is not possible anymore. Here, the function of the β-turn region evolved from simple primer positioning for A-incorporation into an enhanced high-affinity substrate binding compensating for the loss of the conventional C-terminal tRNA interaction. As this strong interaction is visible at all stages of CCA-addition on canonical as well as armless tRNAs (Figure S2A), the function of the β-turn region in RcuCCA is not limited to bind a tRNA-CC intermediate but generally contributes to an efficient substrate binding during the complete CCA-incorporation reaction. This ensures that the enzyme’s interaction with armless tRNAs is sufficiently strong to survive the structural rearrangements during polymerization. A drawback of this tight binding of RcuCCA might be a reduced product release, and the slightly less efficient CCA-addition on the conventional tRNAPhe could be an indication of this (Table 1).
3.3. An Orthogonal Translation System?
Swapping domains between proteins is a useful method to generate proteins with new functions—an approach that is frequently used by nature as well as researchers [87,88,89]. The fact that the replacement of a small β-turn element converts the human tRNA nucleotidyltransferase into an enzyme that accepts armless tRNAs as substrates raises the question whether it is possible to generate an orthogonal system for the incorporation of non-natural amino acids in a host cell system. Representing a fascinating idea, this is highly unlikely, as the armless tRNAs are so different to their conventional counterparts. It is known that many mt-tRNAs are not recognized and charged by cytosolic aminoacyl-tRNA synthetases but require specifically adapted mitochondrial enzymes [90]. Similarly, mitochondrial ribosomes have undergone a specific co-evolution to compensate for the unusual structural features of mt-tRNAs [91]. While metazoan mt-rRNAs are usually shorter than their cytosolic counterparts, the mitochondrial ribosomal proteins are generally enlarged [92,93,94,95]. Again, nematodes represent the extreme case, having ribosomes strongly enriched in protein content but with reduced rRNA components [96,97,98]. Due to this intricate co-evolution of the mitochondrial protein synthesis components, it is probably not feasible to use armless tRNAs as an orthogonal tool in synthetic biology.
Taken together, the CCA-adding enzyme of R. culicivorax shows a remarkable adaptation to hairpin-like tRNA where the loss of substrate interactions with the C-terminus is compensated by enhanced tRNA binding of a different enzyme region. The evolutionary plasticity of enzymes is described for the composition of active site residues, where amino acids with identical catalytic roles are located at different positions in the primary sequence [99]. The catalytic core, however, remains unchanged and structurally almost identical. In contrast, the β-turn element of RcuCCA is not a mimicry of the T-loop/C-terminus interaction but recognizes a very different region of the tRNA (the 3′-end) and is still part of the catalytic core. Hence, its specific adaptation to the miniaturized tRNAs add a new layer of evolutionary plasticity. The structural resolution of this enzyme in complex with its tRNA substrate is expected to shed more light into this unusual and fascinating substrate adaptation.
4. Materials and Methods
4.1. Construction of Recombinant Enzymes
Open reading frames of CCA-adding enzymes from Escherichia coli and Homo sapiens were cloned into pET30 Ek/LIC plasmid with an N-terminal His6-Tag. The mt target signals were not included, and in both enzymes, the cloned coding regions started at the following conserved methionine residue as described [37,50,56,57]. For the CCA-adding enzyme of Romanomermis culicivorax, the coding sequence was identified in the R. culicivorax genome assembly available at https://parasite.wormbase.org/Romanomermis_culicivorax_prjeb1358/Info/Index/, codon-optimized for expression in E. coli and synthesized in pET28a by GenScript®® (Piscataway, NJ, USA). All alignments were done using Jalview 2 [100]. Point mutations were introduced as described [23].
4.2. Cloning of Chimeric Enzymes
Chimeric enzymes of CCA-adding enzymes from Homo sapiens and Romanomermis culicivorax were generated via site-directed mutagenesis in pET30-Ek/LIC or pET28a plasmids, respectively. All chimeras were cloned with an N-terminal His6-tag. The fusion positions of all chimeras are shown in Table S1 and Figure S3.
4.3. Expression and Purification of Recombinant Enzymes
E. coli BL21 (DE3) cca::cam lacking the endogenous CCA-adding enzyme were transformed with plasmids encoding the CCA-adding enzymes from H. sapiens (HsaCCA), R. culicivorax (RcuCCA), or chimeric enzymes. For CCA-adding enzyme from E. coli (EcoCCA), E. coli BL21 (DE3) was used. Cells were grown in 400 ml LB or TB with 50 µg/ml kanamycin and 35 µg/ml chloramphenicol (only for cca::cam strains) at 30 °C. Expression was induced at OD600 = 1.5 by adding 400 ml ice-cold LB or TB containing both antibiotics and IPTG to a final concentration of 1 mM. Cultures were incubated over night at 16 °C and then harvested at 6340 g for 15 min.
Pellets were resuspended in 8 ml ice-cold lysis-buffer (25 mM Tris/HCl pH 7.6, 500 mM NaCl, 1 mM DTT for EcoCCA and HsaCCA, 100 mM phosphate buffer pH 7.0, 500 mM NaCl, 10% glycerol, 0.2% NP-40, 1 mM DTT for RcuCCA) and disrupted with 5 g Zirconia beads and Fastprep-24 homogenizer (6 m/s, 30 s). Cell lysates were centrifuged at 30,600 g, 30 min, 4 °C, sterile filtrated and loaded onto a HisTrap FF 1 ml or 5 ml column (GE Healthcare). Column wash was performed with 4–10 column volumes of binding buffer (25 mM Tris/HCl pH 7.6, 500 mM NaCl for EcoCCA and HsaCCA, 100 mM phosphate buffer pH 7.0, 500 mM NaCl, 10% glycerol for RcuCCA) with 50 mM imidazole. His-tagged proteins were eluted with 3–8 column volumes of elution buffer (binding buffer with 500 mM imidazole). If necessary, protein containing fractions were further purified by size exclusion chromatography on a HiLoad 16/60 Superdex 75 pg column in binding-buffer containing 200 mM NaCl. Protein-containing fractions were combined and concentrated on Vivaspin 6 columns (15–30 kDa MWCO, GE Healthcare). Proteins were stored in 40% glycerol (v/v) at −80 °C. Protein concentration was determined according to Bradford [101].
4.4. tRNA Preparation
Armless mitochondrial tRNAs for isoleucine and arginine from R. culicivorax [48] and canonical cytosolic tRNAPhe from Saccharomyces cerevisiae were generated as in vitro transcripts lacking the CCA-end in the presence of α32P-ATP (3000 Ci/mmol). Homogeneous 5′- and 3′-ends of the transcripts were generated as described [102]. For kinetic analyses, tRNAs were transcribed without α32P-ATP.
4.5. Electrophoretic Mobility Shift Assay (EMSA)
0.5 pmol α32P-ATP-labeled tRNA substrates were heated for 2 min at 90 °C, cooled to room temperature and incubated with 0 to 4 µM of enzyme in HEPES/KOH (pH 7.6), 30 mM KCl and 6 mM MgCl2 at 20 °C for 10 min. After addition of glycerol to a final concentration of 18.5%, tRNAs were separated by 5% native polyacrylamide gel electrophoresis. For visualization of enzyme-bound and free substrates, a Typhoon 9410 scanner (Cytiva, Freiburg, Germany) was used. Dissociation constants were determined in three independent experiments by nonlinear regression using GraphPad Prism 7.
4.6. Activity Test and Determination of Arbitrary Units
Initial activity tests for CCA-addition were performed in 30 mM HEPES/KOH pH 7.6, 30 mM KCl, 6 mM MgCl2, 2 mM DTT, 0.5 mM NTPs, 5 pmol tRNA, and 20 ng of enzyme in a reaction volume of 20 µl at 20 °C.
For comparative analysis of CCA-addition on armless tRNAs (Figure 3), activity of all enzyme preparations was normalized using canonical tRNAPhe as a substrate. CCA-addition was performed in in the same buffer as described above. For calculation of arbitrary units, 5 pmol of tRNA were incubated with increasing amounts of enzymes for 30 min at 20 °C. Reactions were ethanol-precipitated and analyzed on 10% or 12.5% polyacrylamide gels by autoradiography. Enzyme amounts leading to 50% substrate-turnover were defined as 1 arbitrary unit. Nucleotide incorporation assays were performed as mentioned above. 5 pmol of tRNA were incubated with 1, 5, 10, 25, and 50 arbitrary units of each enzyme preparation for 30 min at 20 °C and analyzed as described [25,58].
4.7. Kinetic Analysis
Steady-state Michaelis–Menten kinetics were performed as described [103]. Each reaction contained 1 mM NTPs and 3 µCi of α32P-CTP or α32P-ATP (3000 Ci/mmol) and 30–35 ng HsaCCA or 30–75 ng RcuCCA. Non-labeled tRNA transcripts without CCA-end were titrated from 1-10 µM and incubated for 15–20 min at 20 °C. Determination of incorporated radioactivity was performed as described [25,66] and kinetic parameters of three independent experiments were calculated by non-linear regression Michaelis–Menten kinetic (GraphPad Prism). As the tRNA transcripts are not soluble at excessive saturating conditions, the calculated kinetic parameters represent apparent values, as frequently used for this type of enzymes [24,25,30,58,65].
4.8. Enzyme Modeling
A homology model of RcuCCA enzyme was built using Modeller [104] based on a sequence alignment with the human enzyme and on the human structure (PDB id: 4x4w) [70]. A model of the human enzyme was also built to include all loops that are not visible in the crystal structure. Models were superimposed to the structure of A-adding enzyme:tRNA complex for (PDB id: 4wc2) [34] in PyMOL (v2.4.0, Schrödinger) to position a tRNA in the active site and visualize loops in the vicinity of the tRNA 3′-tail.
Supplementary Materials
The following are available online at https://www.mdpi.com/1422-0067/21/23/9047/s1.
Author Contributions
H.B., C.S. and M.M. designed the experiments; S.P., O.H., T.K., S.B., K.R., and T.J. performed the experiments; H.B., C.S., M.M., S.P., K.R., and O.H. analyzed the data; O.H., S.P., H.B., and M.M. wrote the manuscript. All authors have read and agreed to the published version of the manuscript.
Funding
This research was funded by the Deutsche Forschungsgemeinschaft, grant number Mo 634/10-1. We acknowledge support from the Deutsche Forschungsgemeinschaft (DFG) and Leipzig University within the program of Open Access Publishing.
Acknowledgments
We thank Philipp Schiffer (University of Cologne) for help with the annotation of the CCA-adding enzyme from Romanomermis culicivorax and Barbara Klüver for expert technical assistance.
Conflicts of Interest
The authors declare no conflict of interest. The funders had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript, or in the decision to publish the results.
References
- Rak, R.; Dahan, O.; Pilpel, Y. Repertoires of tRNAs: The Couplers of Genomics and Proteomics. Annu. Rev. Cell Dev. Biol. 2018, 34, 239–264. [Google Scholar] [CrossRef] [PubMed]
- Wolin, S.L.; Matera, A.G. The trials and travels of tRNA. Genes Dev. 1999, 13, 1–10. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Hopper, A.K. Transfer RNA post-transcriptional processing, turnover, and subcellular dynamics in the yeast Saccharomyces cerevisiae. Genetics 2013, 194, 43–67. [Google Scholar] [CrossRef]
- Caetano-Anollés, G.; Sun, F.-J. The natural history of transfer RNA and its interactions with the ribosome. Front. Genet. 2014, 5, 127. [Google Scholar] [CrossRef] [PubMed]
- Hopper, A.K.; Nostramo, R.T. tRNA Processing and Subcellular Trafficking Proteins Multitask in Pathways for Other RNAs. Front. Genet. 2019, 10, 96. [Google Scholar] [CrossRef] [PubMed]
- D’Souza, A.R.; Minczuk, M. Mitochondrial transcription and translation: Overview. Essays Biochem. 2018, 62, 309–320. [Google Scholar] [CrossRef]
- Zoschke, R.; Bock, R. Chloroplast Translation: Structural and Functional Organization, Operational Control, and Regulation. Plant Cell 2018, 30, 745–770. [Google Scholar] [CrossRef]
- Kim, S.H.; Suddath, F.L.; Quigley, G.J.; McPherson, A.; Sussman, J.L.; Wang, A.H.; Seeman, N.C.; Rich, A. Three-dimensional tertiary structure of yeast phenylalanine transfer RNA. Science 1974, 185, 435–440. [Google Scholar] [CrossRef]
- Jühling, F.; Mörl, M.; Hartmann, R.K.; Sprinzl, M.; Stadler, P.F.; Pütz, J. tRNAdb 2009: Compilation of tRNA sequences and tRNA genes. Nucleic Acids Res. 2009, 37, D159–D162. [Google Scholar] [CrossRef]
- Giegé, R.; Jühling, F.; Pütz, J.; Stadler, P.; Sauter, C.; Florentz, C. Structure of transfer RNAs: Similarity and variability. Wiley Interdiscip. Rev. RNA 2012, 3, 37–61. [Google Scholar] [CrossRef]
- Sprinzl, M.; Cramer, F. The -C-C-A end of tRNA and its role in protein biosynthesis. Prog. Nucleic Acid Res. Mol. Biol. 1979, 22, 1–69. [Google Scholar] [PubMed]
- Green, R.; Noller, H.F. Ribosomes and translation. Annu. Rev. Biochem. 1997, 66, 679–716. [Google Scholar] [CrossRef] [PubMed]
- Weiner, A.M. tRNA maturation: RNA polymerization without a nucleic acid template. Curr. Biol. 2004, 14, R883–R885. [Google Scholar] [CrossRef] [PubMed]
- Xiong, Y.; Steitz, T.A. A story with a good ending: tRNA 3’-end maturation by CCA-adding enzymes. Curr. Opin. Struct. Biol. 2006, 16, 12–17. [Google Scholar] [CrossRef]
- Betat, H.; Rammelt, C.; Mörl, M. tRNA nucleotidyltransferases: Ancient catalysts with an unusual mechanism of polymerization. Cell. Mol. Life Sci. 2010, 67, 1447–1463. [Google Scholar] [CrossRef]
- Yue, D.; Maizels, N.; Weiner, A.M. CCA-adding enzymes and poly(A) polymerases are all members of the same nucleotidyltransferase superfamily: Characterization of the CCA-adding enzyme from the archaeal hyperthermophile Sulfolobus shibatae. RNA 1996, 2, 895–908. [Google Scholar]
- Xiong, Y.; Li, F.; Wang, J.; Weiner, A.M.; Steitz, T.A. Crystal structures of an archaeal class I CCA-adding enzyme and its nucleotide complexes. Mol. Cell 2003, 12, 1165–1172. [Google Scholar] [CrossRef]
- Holm, L.; Sander, C. DNA polymerase beta belongs to an ancient nucleotidyltransferase superfamily. Trends Biochem. Sci. 1995, 20, 345–347. [Google Scholar] [CrossRef]
- Steitz, T.A. A mechanism for all polymerases. Nature 1998, 391, 231–232. [Google Scholar] [CrossRef]
- Aravind, L.; Koonin, E.V. DNA polymerase beta-like nucleotidyltransferase superfamily: Identification of three new families, classification and evolutionary history. Nucleic Acids Res. 1999, 27, 1609–1618. [Google Scholar] [CrossRef]
- Li, F.; Xiong, Y.; Wang, J.; Cho, H.D.; Tomita, K.; Weiner, A.M.; Steitz, T.A. Crystal structures of the Bacillus stearothermophilus CCA-adding enzyme and its complexes with ATP or CTP. Cell 2002, 111, 815–824. [Google Scholar] [CrossRef]
- Toh, Y.; Takeshita, D.; Numata, T.; Fukai, S.; Nureki, O.; Tomita, K. Mechanism for the definition of elongation and termination by the class II CCA-adding enzyme. EMBO J. 2009, 28, 3353–3365. [Google Scholar] [CrossRef] [PubMed]
- Ernst, F.G.M.; Rickert, C.; Bluschke, A.; Betat, H.; Steinhoff, H.-J.; Mörl, M. Domain movements during CCA-addition: A new function for motif C in the catalytic core of the human tRNA nucleotidyltransferases. RNA Biol. 2015, 12, 435–446. [Google Scholar] [CrossRef] [PubMed]
- Tomita, K.; Fukai, S.; Ishitani, R.; Ueda, T.; Takeuchi, N.; Vassylyev, D.G.; Nureki, O. Structural basis for template-independent RNA polymerization. Nature 2004, 430, 700–704. [Google Scholar] [CrossRef]
- Hoffmeier, A.; Betat, H.; Bluschke, A.; Gunther, R.; Junghanns, S.; Hofmann, H.-J.; Mörl, M. Unusual evolution of a catalytic core element in CCA-adding enzymes. Nucleic Acids Res. 2010, 38, 4436–4447. [Google Scholar] [CrossRef]
- Neuenfeldt, A.; Just, A.; Betat, H.; Mörl, M. Evolution of tRNA nucleotidyltransferases: A small deletion generated CC-adding enzymes. Proc. Natl. Acad. Sci. USA 2008, 105, 7953–7958. [Google Scholar] [CrossRef]
- Davies, J.F.; Almassy, R.J.; Hostomska, Z.; Ferre, R.A.; Hostomsky, Z. 2.3 Å crystal structure of the catalytic domain of DNA polymerase β. Cell 1994, 76, 1123–1133. [Google Scholar] [CrossRef]
- Sawaya, M.R.; Pelletier, H.; Kumar, A.; Wilson, S.H.; Kraut, J. Crystal structure of rat DNA polymerase beta: Evidence for a common polymerase mechanism. Science 1994, 264, 1930–1935. [Google Scholar] [CrossRef]
- Cho, H.D.; Verlinde, C.L.; Weiner, A.M. Archaeal CCA-adding enzymes: Central role of a highly conserved beta-turn motif in RNA polymerization without translocation. J. Biol. Chem. 2005, 280, 9555–9566. [Google Scholar] [CrossRef]
- Tomita, K.; Ishitani, R.; Fukai, S.; Nureki, O. Complete crystallographic analysis of the dynamics of CCA sequence addition. Nature 2006, 443, 956–960. [Google Scholar] [CrossRef]
- Tomita, K.; Yamashita, S. Molecular mechanisms of template-independent RNA polymerization by tRNA nucleotidyltransferases. Front. Genet. 2014, 5, 36. [Google Scholar] [CrossRef]
- Li, Z.; Gillis, K.A.; Hegg, L.A.; Zhang, J.; Thurlow, D.L. Effects of nucleotide substitutions within the T-loop of precursor tRNAs on interaction with ATP/CTP:tRNA nucleotidyltransferases from Escherichia coli and yeast. Biochem. J. 1996, 314 Pt 1, 49–53. [Google Scholar] [CrossRef]
- Tretbar, S.; Neuenfeldt, A.; Betat, H.; Mörl, M. An inhibitory C-terminal region dictates the specificity of A-adding enzymes. Proc. Natl. Acad. Sci. USA 2011, 108, 21040–21045. [Google Scholar] [CrossRef] [PubMed]
- Yamashita, S.; Martinez, A.; Tomita, K. Measurement of Acceptor-TΨC Helix Length of tRNA for Terminal A76-Addition by A-Adding Enzyme. Structure 2015, 23, 830–842. [Google Scholar] [CrossRef]
- Yamashita, S.; Takeshita, D.; Tomita, K. Translocation and rotation of tRNA during template-independent RNA polymerization by tRNA nucleotidyltransferase. Structure 2014, 22, 315–325. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.; Sun, Y.; Thurlow, D.L. RNA minihelices as model substrates for ATP/CTP:tRNA nucleotidyltransferase. Biochem. J. 1997, 327 Pt 3, 847–851. [Google Scholar] [CrossRef]
- Lizano, E.; Scheibe, M.; Rammelt, C.; Betat, H.; Mörl, M. A comparative analysis of CCA-adding enzymes from human and E. coli: Differences in CCA addition and tRNA 3’-end repair. Biochimie 2008, 90, 762–772. [Google Scholar] [CrossRef] [PubMed]
- Wolstenholme, D.R.; Macfarlane, J.L.; Okimoto, R.; Clary, D.O.; Wahleithner, J.A. Bizarre tRNAs inferred from DNA sequences of mitochondrial genomes of nematode worms. Proc. Natl. Acad. Sci. USA 1987, 84, 1324–1328. [Google Scholar] [CrossRef]
- Okimoto, R.; Wolstenholme, D.R. A set of tRNAs that lack either the T psi C arm or the dihydrouridine arm: Towards a minimal tRNA adaptor. EMBO J. 1990, 9, 3405–3411. [Google Scholar] [CrossRef]
- Watanabe, Y.-I.; Suematsu, T.; Ohtsuki, T. Losing the stem-loop structure from metazoan mitochondrial tRNAs and co-evolution of interacting factors. Front. Genet. 2014, 5, 109. [Google Scholar] [CrossRef]
- de Bruijn, M.H.; Schreier, P.H.; Eperon, I.C.; Barrell, B.G.; Chen, E.Y.; Armstrong, P.W.; Wong, J.F.; Roe, B.A. A mammalian mitochondrial serine transfer RNA lacking the “dihydrouridine” loop and stem. Nucleic Acids Res. 1980, 8, 5213–5222. [Google Scholar] [CrossRef] [PubMed]
- Hunter, S.E.; Spremulli, L.L. Interaction of mitochondrial elongation factor Tu with aminoacyl-tRNAs. Mitochondrion 2004, 4, 21–29. [Google Scholar] [CrossRef] [PubMed]
- Watanabe, Y.; Tsurui, H.; Ueda, T.; Furushima, R.; Takamiya, S.; Kita, K.; Nishikawa, K.; Watanabe, K. Primary and higher order structures of nematode (Ascaris suum) mitochondrial tRNAs lacking either the T or D stem. J. Biol. Chem. 1994, 269, 22902–22906. [Google Scholar] [PubMed]
- Klimov, P.B.; Oconnor, B.M. Improved tRNA prediction in the American house dust mite reveals widespread occurrence of extremely short minimal tRNAs in acariform mites. BMC Genom. 2009, 10, 598. [Google Scholar] [CrossRef]
- Jühling, F.; Pütz, J.; Florentz, C.; Stadler, P.F. Armless mitochondrial tRNAs in Enoplea (Nematoda). RNA Biol. 2012, 9, 1161–1166. [Google Scholar] [CrossRef] [PubMed]
- Pons, J.; Bover, P.; Bidegaray-Batista, L.; Arnedo, M.A. Arm-less mitochondrial tRNAs conserved for over 30 millions of years in spiders. BMC Genom. 2019, 20, 665. [Google Scholar] [CrossRef] [PubMed]
- Jühling, T.; Duchardt-Ferner, E.; Bonin, S.; Wöhnert, J.; Pütz, J.; Florentz, C.; Betat, H.; Sauter, C.; Mörl, M. Small but large enough: Structural properties of armless mitochondrial tRNAs from the nematode Romanomermis culicivorax. Nucleic Acids Res. 2018, 46, 9170–9180. [Google Scholar] [CrossRef]
- Wende, S.; Platzer, E.G.; Jühling, F.; Pütz, J.; Florentz, C.; Stadler, P.F.; Mörl, M. Biological evidence for the world’s smallest tRNAs. Biochimie 2014, 100, 151–158. [Google Scholar] [CrossRef] [PubMed]
- Wolfe, C.L.; Lou, Y.C.; Hopper, A.K.; Martin, N.C. Interplay of heterogeneous transcriptional start sites and translational selection of AUGs dictate the production of mitochondrial and cytosolic/nuclear tRNA nucleotidyltransferase from the same gene in yeast. J. Biol. Chem. 1994, 269, 13361–13366. [Google Scholar]
- Reichert, A.S.; Thurlow, D.L.; Mörl, M. A eubacterial origin for the human tRNA nucleotidyltransferase? Biol. Chem. 2001, 382, 1431–1438. [Google Scholar] [CrossRef]
- Nagaike, T.; Suzuki, T.; Tomari, Y.; Takemoto-Hori, C.; Negayama, F.; Watanabe, K.; Ueda, T. Identification and characterization of mammalian mitochondrial tRNA nucleotidyltransferases. J. Biol. Chem. 2001, 276, 40041–40049. [Google Scholar] [CrossRef] [PubMed]
- Tomari, Y.; Suzuki, T.; Ueda, T. tRNA recognition by CCA-adding enzyme. Nucleic Acids Res. Suppl. 2002, 2, 77–78. [Google Scholar] [CrossRef] [PubMed]
- Shikha, S.; Schneider, A. The single CCA-adding enzyme of T. brucei has distinct functions in the cytosol and in mitochondria. J. Biol. Chem. 2020, 295, 6138–6150. [Google Scholar] [CrossRef] [PubMed]
- Schiffer, P.H.; Kroiher, M.; Kraus, C.; Koutsovoulos, G.D.; Kumar, S.; Camps, J.I.R.; Nsah, N.A.; Stappert, D.; Morris, K.; Heger, P.; et al. The genome of Romanomermis culicivorax: Revealing fundamental changes in the core developmental genetic toolkit in Nematoda. BMC Genom. 2013, 14, 923. [Google Scholar] [CrossRef]
- Claros, M.G.; Vincens, P. Computational method to predict mitochondrially imported proteins and their targeting sequences. Eur. J. Biochem. 1996, 241, 779–786. [Google Scholar] [CrossRef]
- Augustin, M.A.; Reichert, A.S.; Betat, H.; Huber, R.; Mörl, M.; Steegborn, C. Crystal structure of the human CCA-adding enzyme: Insights into template-independent polymerization. J. Mol. Biol. 2003, 328, 985–994. [Google Scholar] [CrossRef]
- Lizano, E.; Schuster, J.; Müller, M.; Kelso, J.; Mörl, M. A splice variant of the human CCA-adding enzyme with modified activity. J. Mol. Biol. 2007, 366, 1258–1265. [Google Scholar] [CrossRef]
- Ernst, F.G.M.; Erber, L.; Sammler, J.; Jühling, F.; Betat, H.; Mörl, M. Cold adaptation of tRNA nucleotidyltransferases: A tradeoff in activity, stability and fidelity. RNA Biol. 2018, 15, 144–155. [Google Scholar] [CrossRef]
- Okabe, M.; Tomita, K.; Ishitani, R.; Ishii, R.; Takeuchi, N.; Arisaka, F.; Nureki, O.; Yokoyama, S. Divergent evolutions of trinucleotide polymerization revealed by an archaeal CCA-adding enzyme structure. EMBO J. 2003, 22, 5918–5927. [Google Scholar] [CrossRef]
- Erber, L.; Franz, P.; Betat, H.; Prohaska, S.; Mörl, M. Divergent Evolution of Eukaryotic CC- and A-Adding Enzymes. Int. J. Mol. Sci. 2020, 21, 462. [Google Scholar] [CrossRef]
- Oommen, A.; Li, X.Q.; Gegenheimer, P. Cleavage specificity of chloroplast and nuclear tRNA 3’-processing nucleases. Mol. Cell. Biol. 1992, 12, 865–875. [Google Scholar] [CrossRef] [PubMed]
- Loria, A.; Pan, T. The 3’ substrate determinants for the catalytic efficiency of the Bacillus subtilis RNase P holoenzyme suggest autolytic processing of the RNase P RNA in vivo. RNA 2000, 6, 1413–1422. [Google Scholar] [CrossRef]
- Shi, H.; Moore, P.B. The crystal structure of yeast phenylalanine tRNA at 1.93 A resolution: A classic structure revisited. RNA 2000, 6, 1091–1105. [Google Scholar] [CrossRef]
- Byrne, R.T.; Konevega, A.L.; Rodnina, M.V.; Antson, A.A. The crystal structure of unmodified tRNAPhe from Escherichia coli. Nucleic Acids Res. 2010, 38, 4154–4162. [Google Scholar] [CrossRef] [PubMed]
- Wende, S.; Bonin, S.; Götze, O.; Betat, H.; Mörl, M. The identity of the discriminator base has an impact on CCA addition. Nucleic Acids Res. 2015, 43, 5617–5629. [Google Scholar] [CrossRef] [PubMed]
- Just, A.; Butter, F.; Trenkmann, M.; Heitkam, T.; Mörl, M.; Betat, H. A comparative analysis of two conserved motifs in bacterial poly(A) polymerase and CCA-adding enzyme. Nucleic Acids Res. 2008, 36, 5212–5220. [Google Scholar] [CrossRef] [PubMed]
- Betat, H.; Rammelt, C.; Martin, G.; Mörl, M. Exchange of regions between bacterial poly(A) polymerase and the CCA-adding enzyme generates altered specificities. Mol. Cell 2004, 15, 389–398. [Google Scholar] [CrossRef]
- Martin, G.; Keller, W. Sequence motifs that distinguish ATP(CTP): tRNA nucleotidyl transferases from eubacterial poly(A) polymerases. RNA 2004, 10, 899–906. [Google Scholar] [CrossRef]
- Shi, P.Y.; Maizels, N.; Weiner, A.M. CCA addition by tRNA nucleotidyltransferase: Polymerization without translocation? EMBO J. 1998, 17, 3197–3206. [Google Scholar] [CrossRef]
- Kuhn, C.-D.; Wilusz, J.E.; Zheng, Y.; Beal, P.A.; Joshua-Tor, L. On-Enzyme Refolding Permits Small RNA and tRNA Surveillance by the CCA-Adding Enzyme. Cell 2015, 160, 644–658. [Google Scholar] [CrossRef]
- Chen, J.Y.; Joyce, P.B.; Wolfe, C.L.; Steffen, M.C.; Martin, N.C. Cytoplasmic and mitochondrial tRNA nucleotidyltransferase activities are derived from the same gene in the yeast Saccharomyces cerevisiae. J. Biol. Chem. 1992, 267, 14879–14883. [Google Scholar] [PubMed]
- Dupasquier, M.; Kim, S.; Halkidis, K.; Gamper, H.; Hou, Y.-M. tRNA integrity is a prerequisite for rapid CCA addition: Implication for quality control. J. Mol. Biol 2008, 379, 579–588. [Google Scholar] [CrossRef] [PubMed]
- Hou, Y.-M. CCA addition to tRNA: Implications for tRNA quality control. IUBMB life 2010, 62, 251–260. [Google Scholar] [CrossRef] [PubMed]
- Wellner, K.; Betat, H.; Mörl, M. A tRNA’s fate is decided at its 3’ end: Collaborative actions of CCA-adding enzyme and RNases involved in tRNA processing and degradation. Biochim. Biophys. Acta Gene Regul. Mech. 2018, 1861, 433–441. [Google Scholar] [CrossRef] [PubMed]
- Wellner, K.; Czech, A.; Ignatova, Z.; Betat, H.; Mörl, M. Examining tRNA 3’-ends in Escherichia coli: Teamwork between CCA-adding enzyme, RNase T, and RNase R. RNA 2018, 24, 361–370. [Google Scholar] [CrossRef] [PubMed]
- Wilusz, J.E.; Whipple, J.M.; Phizicky, E.M.; Sharp, P.A. tRNAs marked with CCACCA are targeted for degradation. Science 2011, 334, 817–821. [Google Scholar] [CrossRef] [PubMed]
- Levinger, L.; Bourne, R.; Kolla, S.; Cylin, E.; Russell, K.; Wang, X.; Mohan, A. Matrices of paired substitutions show the effects of tRNA D/T loop sequence on Drosophila RNase P and 3’-tRNase processing. J. Biol. Chem. 1998, 273, 1015–1025. [Google Scholar] [CrossRef]
- Zahler, N.H.; Christian, E.L.; Harris, M.E. Recognition of the 5’ leader of pre-tRNA substrates by the active site of ribonuclease P. RNA 2003, 9, 734–745. [Google Scholar] [CrossRef]
- Kirsebom, L.A. RNase P RNA mediated cleavage: Substrate recognition and catalysis. Biochimie 2007, 89, 1183–1194. [Google Scholar] [CrossRef]
- Levinger, L.; Hopkinson, A.; Desetty, R.; Wilson, C. Effect of changes in the flexible arm on tRNase Z processing kinetics. J. Biol. Chem. 2009, 284, 15685–15691. [Google Scholar] [CrossRef]
- McKenney, K.M.; Rubio, M.A.T.; Alfonzo, J.D. The Evolution of Substrate Specificity by tRNA Modification Enzymes. Enzymes 2017, 41, 51–88. [Google Scholar] [CrossRef] [PubMed]
- Spacciapoli, P.; Doviken, L.; Mulero, J.J.; Thurlow, D.L. Recognition of tRNA by the enzyme ATP/CTP:tRNA nucleotidyltransferase. Interference by nucleotides modified with diethyl pyrocarbonate or hydrazine. J. Biol. Chem. 1989, 264, 3799–3805. [Google Scholar] [PubMed]
- Spacciapoli, P.; Thurlow, D.L. Purines in tRNAs required for recognition by ATP/CTP:tRNA nucleotidyltransferase from rabbit liver. J. Mol. Recognit. 1990, 3, 149–155. [Google Scholar] [CrossRef] [PubMed]
- Hegg, L.A.; Thurlow, D.L. Cytidines in tRNAs that are required intact by ATP/CTP:tRNA nucleotidyltransferases from Escherichia coli and Saccharomyces cerevisiae. Nucleic Acids Res. 1990, 18, 5975–5979. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.; Liu, C.; Halkidis, K.; Gamper, H.B.; Hou, Y.-M. Distinct kinetic determinants for the stepwise CCA addition to tRNA. RNA 2009, 15, 1827–1836. [Google Scholar] [CrossRef] [PubMed]
- Xiong, Y.; Steitz, T.A. Mechanism of transfer RNA maturation by CCA-adding enzyme without using an oligonucleotide template. Nature 2004, 430, 640–645. [Google Scholar] [CrossRef] [PubMed]
- Kries, H.; Niquille, D.L.; Hilvert, D. A subdomain swap strategy for reengineering nonribosomal peptides. Chem. Biol. 2015, 22, 640–648. [Google Scholar] [CrossRef] [PubMed]
- Ostermeier, M.; Benkovic, S.J. Evolution of protein function by Domain swapping. In Evolutionary Protein Design; Arnold, F.H., Ed.; Academic Press: San Diego, CA, USA, 2001; pp. 29–77. ISBN 9780120342556. [Google Scholar]
- Truman, A.W.; Dias, M.V.B.; Wu, S.; Blundell, T.L.; Huang, F.; Spencer, J.B. Chimeric glycosyltransferases for the generation of hybrid glycopeptides. Chem. Biol. 2009, 16, 676–685. [Google Scholar] [CrossRef][Green Version]
- Tzagoloff, A.; Gatti, D.; Gampel, A. Mitochondrial aminoacyl-tRNA synthetases. Prog. Nucleic Acid Res. Mol. Biol 1990, 39, 129–158. [Google Scholar]
- Salinas-Giegé, T.; Giegé, R.; Giegé, P. tRNA biology in mitochondria. Int. J. Mol. Sci. 2015, 16, 4518–4559. [Google Scholar] [CrossRef]
- Greber, B.J.; Bieri, P.; Leibundgut, M.; Leitner, A.; Aebersold, R.; Boehringer, D.; Ban, N. Ribosome. The complete structure of the 55S mammalian mitochondrial ribosome. Science 2015, 348, 303–308. [Google Scholar] [CrossRef] [PubMed]
- Koc, E.C.; Burkhart, W.; Blackburn, K.; Moyer, M.B.; Schlatzer, D.M.; Moseley, A.; Spremulli, L.L. The large subunit of the mammalian mitochondrial ribosome. Analysis of the complement of ribosomal proteins present. J. Biol. Chem. 2001, 276, 43958–43969. [Google Scholar] [CrossRef] [PubMed]
- Sharma, M.R.; Koc, E.C.; Datta, P.P.; Booth, T.M.; Spremulli, L.L.; Agrawal, R.K. Structure of the Mammalian Mitochondrial Ribosome Reveals an Expanded Functional Role for Its Component Proteins. Cell 2003, 115, 97–108. [Google Scholar] [CrossRef]
- Suzuki, T.; Terasaki, M.; Takemoto-Hori, C.; Hanada, T.; Ueda, T.; Wada, A.; Watanabe, K. Structural compensation for the deficit of rRNA with proteins in the mammalian mitochondrial ribosome. Systematic analysis of protein components of the large ribosomal subunit from mammalian mitochondria. J. Biol. Chem. 2001, 276, 21724–21736. [Google Scholar] [CrossRef]
- Okimoto, R.; Macfarlane, J.L.; Wolstenholme, D.R. The mitochondrial ribosomal RNA genes of the nematodes Caenorhabditis elegans and Ascaris suum: Consensus secondary-structure models and conserved nucleotide sets for phylogenetic analysis. J. Mol. Evol. 1994, 39, 598–613. [Google Scholar] [CrossRef]
- Zhao, F.; Ohtsuki, T.; Yamada, K.; Yoshinari, S.; Kita, K.; Watanabe, Y.-I.; Watanabe, K. Isolation and physiochemical properties of protein-rich nematode mitochondrial ribosomes. Biochemistry 2005, 44, 9232–9237. [Google Scholar] [CrossRef]
- Mears, J.A.; Cannone, J.J.; Stagg, S.M.; Gutell, R.R.; Agrawal, R.K.; Harvey, S.C. Modeling a Minimal Ribosome Based on Comparative Sequence Analysis. J. Mol. Biol 2002, 321, 215–234. [Google Scholar] [CrossRef]
- Todd, A.E.; Orengo, C.A.; Thornton, J.M. Plasticity of enzyme active sites. Trends Biochem. Sci. 2002, 27, 419–426. [Google Scholar] [CrossRef]
- Waterhouse, A.M.; Procter, J.B.; Martin, D.M.A.; Clamp, M.; Barton, G.J. Jalview Version 2—A multiple sequence alignment editor and analysis workbench. Bioinformatics 2009, 25, 1189–1191. [Google Scholar] [CrossRef]
- Bradford, M.M. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal. Biochem. 1976, 72, 248–254. [Google Scholar] [CrossRef]
- Schürer, H.; Lang, K.; Schuster, J.; Mörl, M. A universal method to produce in vitro transcripts with homogeneous 3’ ends. Nucleic Acids Res. 2002, 30, e56. [Google Scholar] [CrossRef] [PubMed]
- Scheibe, M.; Bonin, S.; Hajnsdorf, E.; Betat, H.; Mörl, M. Hfq stimulates the activity of the CCA-adding enzyme. BMC Mol. Biol. 2007, 8, 92. [Google Scholar] [CrossRef] [PubMed]
- Webb, B.; Sali, A. Comparative Protein Structure Modeling Using MODELLER. Curr. Protoc. Bioinform. 2016, 54, 5.6.1–5.6.37. [Google Scholar] [CrossRef] [PubMed]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).