Recent Advances of Functional Proteomics in Gastrointestinal Cancers- a Path towards the Identification of Candidate Diagnostic, Prognostic, and Therapeutic Molecular Biomarkers
Abstract
1. Introduction
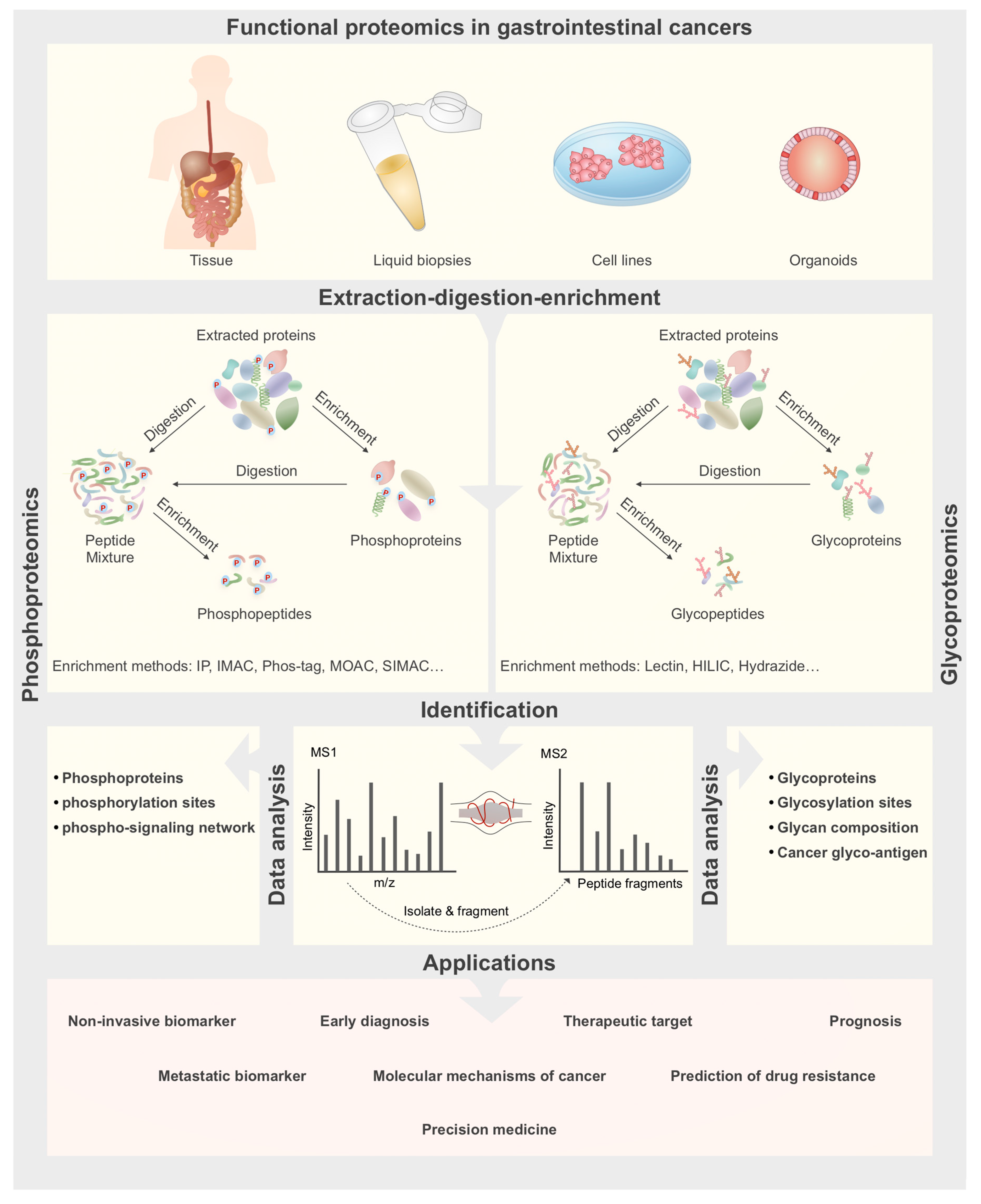
2. Quantitative Phosphoproteomics and Glycoproteomics Strategies
2.1. Phosphoproteomics
2.2. Glycoproteomics
3. Phosphoproteomics in GI Cancers
3.1. Colorectal Cancer
3.2. Gastric Cancer
3.3. Pancreatic Cancer
3.4. Hepatocellular Carcinoma (HCC)
4. Glycoproteomics in GI Cancers
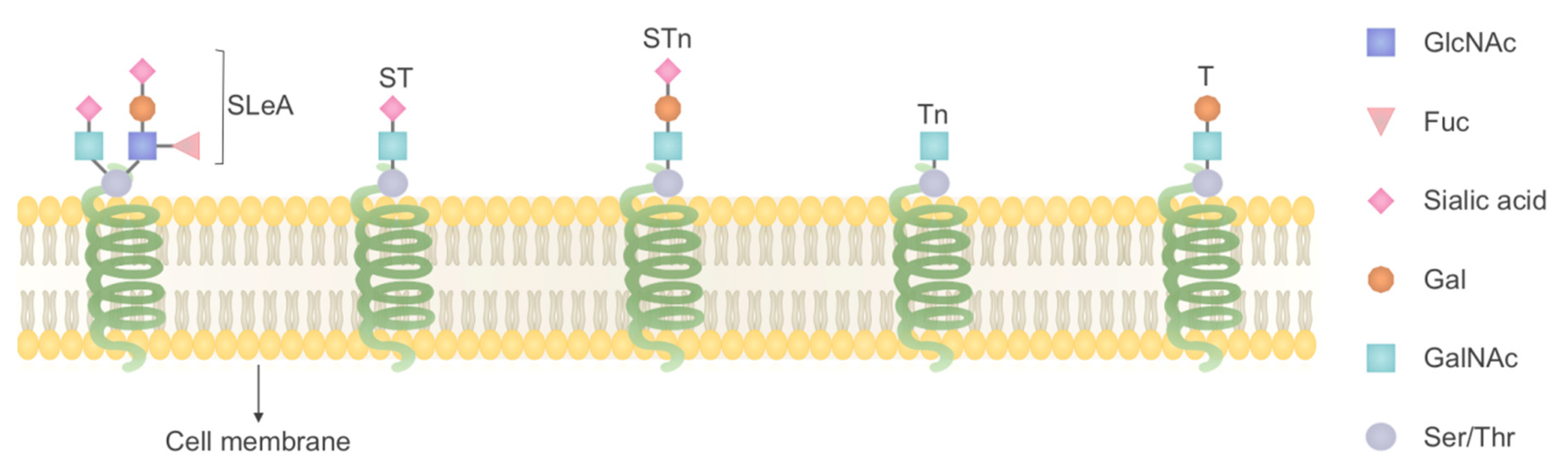
4.1. O-Linked Glycosylation
4.2. N-Linked Glycosylation
4.3. Whole Glycoproteome
5. Conclusions
Funding
Conflicts of Interest
References
- Bray, F.; Me, J.F.; Soerjomataram, I.; Siegel, R.L.; Torre, L.A.; Jemal, A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 2018, 68, 394–424. [Google Scholar] [CrossRef] [PubMed]
- Arnold, M.; Abnet, C.C.; Neale, R.E.; Vignat, J.; Giovannucci, E.L.; McGlynn, K.A.; Bray, F. Global Burden of 5 Major Types of Gastrointestinal Cancer. Gastroenterology 2020, 159, 335–349. [Google Scholar] [CrossRef] [PubMed]
- Somi, M.H.; Dolatkhah, R.; Sepahi, S.; Belalzadeh, M.; Naghashi, S.; Jafarabadi, M.A. A 12-year trend analysis of the incidence of gastrointestinal cancers in East Azerbaijan: Last updated results of an ongoing population-based cancer registry. BMC Cancer 2019, 19, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Mereiter, S.; Balmaña, M.; Gomes, J.; Magalhães, A.; Reis, C.A. Glycomic Approaches for the Discovery of Targets in Gastrointestinal Cancer. Front. Oncol. 2016, 6, 55. [Google Scholar] [CrossRef] [PubMed]
- Ebert, M.P.; Yu, J.; Lordick, F.; Röcken, C. Proteomics in gastrointestinal cancer. Ann. Oncol. 2006, 17, x252–x258. [Google Scholar] [CrossRef]
- Takahashi, T.; ElZawahry, A.; Mimaki, S.; Furukawa, E.; Nakatsuka, R.; Nakamura, H.; Nishigaki, T.; Serada, S.; Naka, T.; Hirota, S.; et al. Genomic and transcriptomic analysis of imatinib resistance in gastrointestinal stromal tumors. Genes Chromosom. Cancer 2017, 56, 303–313. [Google Scholar] [CrossRef]
- Yu, J.; Zhao, J.; Zhang, M.; Guo, J.; Liu, X.; Liu, L. Metabolomics studies in gastrointestinal cancer: A systematic review. Expert Rev. Gastroenterol. Hepatol. 2020, 14, 9–25. [Google Scholar] [CrossRef]
- Monti, M.; Orrù, S.; Pagnozzi, D.; Pucci, P. Functional proteomics. Clin. Chim. Acta 2005, 357, 140–150. [Google Scholar] [CrossRef]
- Darling, A.L.; Uversky, V.N. Intrinsic Disorder and Posttranslational Modifications: The Darker Side of the Biological Dark Matter. Front. Genet. 2018, 9, 158. [Google Scholar] [CrossRef]
- Abe, Y.; Hirano, H.; Shoji, H.; Tada, A.; Isoyama, J.; Kakudo, A.; Gunji, D.; Honda, K.; Boku, N.; Adachi, J.; et al. Comprehensive characterization of the phosphoproteome of gastric cancer from endoscopic biopsy specimens. Theranostics 2020, 10, 2115. [Google Scholar] [CrossRef]
- Chang, T.-T.; Cheng, J.; Tsai, H.; Young, K.; Hsieh, S.-Y.; Ho, C.-H. Plasma proteome plus site-specific N-glycoprofiling for hepatobiliary carcinomas. J. Pathol. Clin. Res. 2019, 5, 199–212. [Google Scholar] [CrossRef] [PubMed]
- Zhu, D.; Hou, L.; Hu, B.; Zhao, H.; Sun, J.; Wang, J.; Meng, X. Crosstalk among proteome, acetylome and succinylome in colon cancer HCT116 cell treated with sodium dichloroacetate. Sci. Rep. 2016, 6, 37478. [Google Scholar] [CrossRef] [PubMed]
- Zhou, J.; Wang, W.; Xie, Y.; Zhao, Y.; Chen, X.; Xu, W.; Wang, Y.; Guan, Z. Proteomics-Based Identification and Analysis of Proteins Associated with Helicobacter pylori in Gastric Cancer. PLoS ONE 2016, 11, e0146521. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Chen, C.; Yu, T.; Chen, T. Proteomic Analysis of Protein Ubiquitination Events in Human Primary and Metastatic Colon Adenocarcinoma Tissues. Front. Oncol. 2020, 10. [Google Scholar] [CrossRef] [PubMed]
- Ardito, F.; Giuliani, M.; Perrone, D.; Troiano, G.; Muzio, L.L. The crucial role of protein phosphorylation in cell signaling and its use as targeted therapy. Int. J. Mol. Med. 2017, 40, 271–280. [Google Scholar] [CrossRef]
- Harsha, H.; Pandey, A. Phosphoproteomics in cancer. Mol. Oncol. 2010, 4, 482–495. [Google Scholar] [CrossRef]
- Moremen, K.W.; Tiemeyer, M.; Nairn, A.V. Vertebrate protein glycosylation: Diversity, synthesis and function. Nat. Rev. Mol. Cell Biol. 2012, 13, 448–462. [Google Scholar] [CrossRef]
- Cantin, G.T.; Shock, T.R.; Park, S.K.; Madhani, A.H.D.; Yates, J.R. Optimizing TiO2-Based Phosphopeptide Enrichment for Automated Multidimensional Liquid Chromatography Coupled to Tandem Mass Spectrometry. Anal. Chem. 2007, 79, 4666–4673. [Google Scholar] [CrossRef] [PubMed]
- Dunn, J.D.; Reid, G.E.; Bruening, M.L. Techniques for phosphopeptide enrichment prior to analysis by mass spectrometry. Mass Spectrom. Rev. 2010, 29, 29–54. [Google Scholar] [CrossRef] [PubMed]
- Fíla, J.; Honys, D. Enrichment techniques employed in phosphoproteomics. Amino Acids 2012, 43, 1025–1047. [Google Scholar] [CrossRef] [PubMed]
- Ashman, K.; Villar, E.L. Phosphoproteomics and cancer research. Clin. Transl. Oncol. 2009, 11, 356–362. [Google Scholar] [CrossRef] [PubMed]
- Mann, M.; Ong, S.-E.; Grønborg, M.; Steen, H.; Jensen, O.N.; Pandey, A. Analysis of protein phosphorylation using mass spectrometry: Deciphering the phosphoproteome. Trends Biotechnol. 2002, 20, 261–268. [Google Scholar] [CrossRef]
- Marcus, K.; Immler, D.; Sternberger, J.; Meyer, H.E. Identification of platelet proteins separated by two-dimensional gel electrophoresis and analyzed by matrix assisted laser desorption/ionization-time of flight-mass spectrometry and detection of tyrosine-phosphorylated proteins. Electrophor. Int. J. 2000, 21, 2622–2636. [Google Scholar] [CrossRef]
- Thingholm, T.E.; Jensen, O.N.; Larsen, M.R. Analytical strategies for phosphoproteomics. Proteomics 2009, 9, 1451–1468. [Google Scholar] [CrossRef] [PubMed]
- Tsai, C.-F.; Wang, Y.-T.; Chen, Y.-R.; Lai, C.-Y.; Lin, P.-Y.; Pan, K.-T.; Chen, J.-Y.; Khoo, K.-H.; Chen, Y.-J. Immobilized Metal Affinity Chromatography Revisited: pH/Acid Control toward High Selectivity in Phosphoproteomics. J. Proteome Res. 2008, 7, 4058–4069. [Google Scholar] [CrossRef] [PubMed]
- Karve, T.M.; Cheema, A.K. Small Changes Huge Impact: The Role of Protein Posttranslational Modifications in Cellular Homeostasis and Disease. J. Amino Acids 2011, 2011, 1–13. [Google Scholar] [CrossRef]
- Thingholm, T.E.; Jensen, O.N.; Robinson, P.J.; Larsen, M.R. SIMAC (Sequential Elution from IMAC), a Phosphoproteomics Strategy for the Rapid Separation of Monophosphorylated from Multiply Phosphorylated Peptides. Mol. Cell. Proteom. 2008, 7, 661–671. [Google Scholar] [CrossRef]
- Tan, Y.-J.; Sui, D.; Wang, W.-H.; Kuo, M.-H.; Reid, G.E.; Bruening, M.L. Phosphopeptide Enrichment with TiO2-Modified Membranes and Investigation of Tau Protein Phosphorylation. Anal. Chem. 2013, 85, 5699–5706. [Google Scholar] [CrossRef]
- Ong, S.-E.; Blagoev, B.; Kratchmarova, I.; Kristensen, D.B.; Steen, H.; Pandey, A.; Mann, M. Stable Isotope Labeling by Amino Acids in Cell Culture, SILAC, as a Simple and Accurate Approach to Expression Proteomics. Mol. Cell. Proteom. 2002, 1, 376–386. [Google Scholar] [CrossRef]
- Blagoev, B.; Ong, S.-E.; Kratchmarova, I.; Mann, M. Temporal analysis of phosphotyrosine-dependent signaling networks by quantitative proteomics. Nat. Biotechnol. 2004, 22, 1139–1145. [Google Scholar] [CrossRef]
- Ross, P.L.; Huang, Y.N.; Marchese, J.N.; Williamson, B.; Parker, K.; Hattan, S.; Khainovski, N.; Pillai, S.; Dey, S.; Daniels, S.; et al. Multiplexed Protein Quantitation inSaccharomyces cerevisiaeUsing Amine-reactive Isobaric Tagging Reagents. Mol. Cell. Proteom. 2004, 3, 1154–1169. [Google Scholar] [CrossRef] [PubMed]
- Hsu, J.-L.; Huang, S.-Y.; Chow, N.-H.; Chen, S.-H. Stable-Isotope Dimethyl Labeling for Quantitative Proteomics. Anal. Chem. 2003, 75, 6843–6852. [Google Scholar] [CrossRef] [PubMed]
- Song, C.; Wang, F.; Ye, M.; Cheng, K.; Chen, R.; Zhu, J.; Tan, Y.; Wang, H.; Figeys, D.; Zou, H. Improvement of the Quantification Accuracy and Throughput for Phosphoproteome Analysis by a Pseudo Triplex Stable Isotope Dimethyl Labeling Approach. Anal. Chem. 2011, 83, 7755–7762. [Google Scholar] [CrossRef] [PubMed]
- Lin, Y.-T.; Chien, K.-Y.; Wu, C.-C.; Chang, W.-Y.; Chu, L.J.; Chen, M.-C.; Yeh, C.-T.; Yu, J.-S. Super-SILAC mix coupled with SIM/AIMS assays for targeted verification of phosphopeptides discovered in a large-scale phosphoproteome analysis of hepatocellular carcinoma. J. Proteom. 2017, 157, 40–51. [Google Scholar] [CrossRef] [PubMed]
- Zhou, H.; Liu, Y.; Chui, J.; Guo, K.; Shun, Q.; Lu, W.; Jin, H.; Wei, L.; Yang, P. Investigation on glycosylation patterns of proteins from human liver cancer cell lines based on the multiplexed proteomics technology. Arch. Biochem. Biophys. 2007, 459, 70–78. [Google Scholar] [CrossRef]
- Cao, J.; Shen, C.; Wang, H.; Shen, H.; Chen, Y.; Nie, A.; Yan, G.; Lu, H.; Liu, Y.; Yang, P. Identification of N-Glycosylation Sites on Secreted Proteins of Human Hepatocellular Carcinoma Cells with a Complementary Proteomics Approach. J. Proteome Res. 2009, 8, 662–672. [Google Scholar] [CrossRef]
- Sun, Z.; Sun, D.; Wang, F.; Cheng, K.; Zhang, Z.; Xu, B.; Xuan, X.; Wang, L.; Zou, H. Differential analysis of N-glycoproteome between hepatocellular carcinoma and normal human liver tissues by combination of multiple protease digestion and solid phase based labeling. Clin. Proteom. 2014, 11, 26. [Google Scholar] [CrossRef]
- Jiang, B.; Huang, J.; Yu, Z.; Wu, M.; Liu, M.; Yao, J.; Zhao, H.; Yan, G.; Ying, W.; Cao, W.; et al. A multi-parallel N-glycopeptide enrichment strategy for high-throughput and in-depth mapping of the N-glycoproteome in metastatic human hepatocellular carcinoma cell lines. Talanta 2019, 199, 254–261. [Google Scholar] [CrossRef]
- Mertins, P.; Yang, F.; Liu, T.; Mani, D.R.; Petyuk, V.A.; Gillette, M.A.; Clauser, K.R.; Qiao, J.W.; Gritsenko, M.A.; Moore, R.J.; et al. Ischemia in Tumors Induces Early and Sustained Phosphorylation Changes in Stress Kinase Pathways but Does Not Affect Global Protein Levels. Mol. Cell. Proteom. 2014, 13, 1690–1704. [Google Scholar] [CrossRef]
- Apweiler, R.; Hermjakob, H.; Sharon, N. On the frequency of protein glycosylation, as deduced from analysis of the SWISS-PROT database11Dedicated to Prof. Akira Kobata and Prof. Harry Schachter on the occasion of their 65th birthdays. Biochim. Biophys. Acta (BBA)-Gen. Subj. 1999, 1473, 4–8. [Google Scholar] [CrossRef]
- Hart, G.W.; Copeland, R.J. Glycomics Hits the Big Time. Cell 2010, 143, 672–676. [Google Scholar] [CrossRef] [PubMed]
- Helenius, A. Intracellular Functions of N-Linked Glycans. Science 2001, 291, 2364–2369. [Google Scholar] [CrossRef] [PubMed]
- Rudd, P.M.; Woods, R.J.; Wormald, M.R.; Opdenakker, G.; Downing, A.; Campbell, I.D.; Dwek, R.A. The effects of variable glycosylation on the functional activities of ribonuclease, plasminogen and tissue plasminogen activator. Biochim. Biophys. Acta 1995, 1248, 1–10. [Google Scholar] [CrossRef]
- Rudd, P.M.; Wormald, M.R.; Stanfield, R.L.; Huang, M.; Mattsson, N.; Speir, J.A.; DiGennaro, J.A.; Fetrow, J.S.; Dwek, R.A.; Wilson, I.A. Roles for glycosylation of cell surface receptors involved in cellular immune recognition. J. Mol. Biol. 1999, 293, 351–366. [Google Scholar] [CrossRef]
- Varki, A. Biological roles of oligosaccharides: All of the theories are correct. Glycobiology 1993, 3, 97–130. [Google Scholar] [CrossRef]
- Veillon, L.; Huang, Y.; Peng, W.; Dong, X.; Cho, B.G.; Mechref, Y. Characterization of isomeric glycan structures by LC-MS/MS. Electrophoresis 2017, 38, 2100–2114. [Google Scholar] [CrossRef]
- Wooding, K.M.; Peng, W.; Mechref, Y. Characterization of Pharmaceutical IgG and Biosimilars Using Miniaturized Platforms and LC-MS/MS. Curr. Pharm. Biotechnol. 2016, 17, 788–801. [Google Scholar] [CrossRef]
- Dwek, R.A. Glycobiology: Toward Understanding the Function of Sugars. Chem. Rev. 1996, 96, 683–720. [Google Scholar] [CrossRef]
- Dennis, J.W.; Granovsky, M.; Warren, C.E. Protein glycosylation in development and disease. Bioessays 1999, 21, 412–421. [Google Scholar] [CrossRef]
- Lowe, J.B.; Marth, J.D. A Genetic Approach to Mammalian Glycan Function. Annu. Rev. Biochem. 2003, 72, 643–691. [Google Scholar] [CrossRef]
- Xia, B.; Royall, J.A.; Damera, G.; Sachdev, G.P.; Cummings, R.D. Altered O-glycosylation and sulfation of airway mucins associated with cystic fibrosis. Glycobiology 2005, 15, 747–775. [Google Scholar] [CrossRef] [PubMed]
- Chace, K.V.; Flux, M.; Sachdev, G.P. Comparison of physicochemical properties of purified mucus glycoproteins isolated from respiratory secretions of cystic fibrosis and asthmatic patients. Biochemistry 1985, 24, 7334–7341. [Google Scholar] [CrossRef] [PubMed]
- Davril, M.; DeGroote, S.; Humbert, P.; Galabert, C.; Dumur, V.; Lafitte, J.-J.; Lamblin, G.; Roussel, P. The sialylation of bronchial mucins secreted by patients suffering from cystic fibrosis or from chronic bronchitis is related to the severity of airway infection. Glycobiology 1999, 9, 311–321. [Google Scholar] [CrossRef] [PubMed]
- Mechref, Y.; Hu, Y.; Garcia, A.; Hussein, A. Identifying cancer biomarkers by mass spectrometry-based glycomics. Electrophoresis 2012, 33, 1755–1767. [Google Scholar] [CrossRef] [PubMed]
- Dell, A. Glycoprotein Structure Determination by Mass Spectrometry. Science 2001, 291, 2351–2356. [Google Scholar] [CrossRef] [PubMed]
- Mechref, Y.; Novotny, M.V. Structural Investigations of Glycoconjugates at High Sensitivity. Chem. Rev. 2002, 102, 321–370. [Google Scholar] [CrossRef]
- Novotny, M.V.; Mechref, Y. New hyphenated methodologies in high-sensitivity glycoprotein analysis. J. Sep. Sci. 2005, 28, 1956–1968. [Google Scholar] [CrossRef]
- Banazadeh, A.; Peng, W.; Veillon, L.; Mechref, Y. Carbon Nanoparticles and Graphene Nanosheets as MALDI Matrices in Glycomics: A New Approach to Improve Glycan Profiling in Biological Samples. J. Am. Soc. Mass Spectrom. 2018, 29, 1892–1900. [Google Scholar] [CrossRef]
- Tholey, A.; Becker, A. Top-down proteomics for the analysis of proteolytic events—Methods, applications and perspectives. Biochim. Biophys. Acta 2017, 1864, 2191–2199. [Google Scholar] [CrossRef]
- Reily, C.; Stewart, T.J.; Renfrow, M.B.; Novak, J. Glycosylation in health and disease. Nat. Rev. Nephrol. 2019, 15, 346–366. [Google Scholar] [CrossRef]
- Kay Li, Q.; Gabrielson, E.; Zhang, H. Application of glycoproteomics for the discovery of biomarkers in lung cancer. Proteom. Clin. Appl. 2012, 6, 244–256. [Google Scholar] [CrossRef] [PubMed]
- Tian, Y.; Zhang, H. Glycoproteomics and clinical applications. Proteom. Clin. Appl. 2010, 4, 124–132. [Google Scholar] [CrossRef] [PubMed]
- Chen, R.; Jiang, X.; Sun, D.; Han, G.; Wang, F.; Ye, M.; Wang, L.; Zou, H. Glycoproteomics Analysis of Human Liver Tissue by Combination of Multiple Enzyme Digestion and Hydrazide Chemistry. J. Proteome Res. 2009, 8, 651–661. [Google Scholar] [CrossRef] [PubMed]
- Bie, Z.; Chen, Y.; Ye, J.; Wang, S.; Liu, Z. Boronate-Affinity Glycan-Oriented Surface Imprinting: A New Strategy to Mimic Lectins for the Recognition of an Intact Glycoprotein and Its Characteristic Fragments. Angew. Chem. Int. Ed. 2015, 54, 10211–10215. [Google Scholar] [CrossRef]
- Zhang, C.; Ye, Z.; Xue, P.; Shu, Q.; Zhou, Y.; Ji, Y.; Fu, Y.; Wang, J.; Yang, F. Evaluation of Different N-Glycopeptide Enrichment Methods for N-Glycosylation Sites Mapping in Mouse Brain. J. Proteome Res. 2016, 15, 2960–2968. [Google Scholar] [CrossRef]
- Dong, X.; Qin, H.; Mao, J.; Yu, D.; Li, X.; Shen, A.; Yan, J.; Yu, L.; Guo, Z.; Xuan, X.; et al. In-Depth Analysis of Glycoprotein Sialylation in Serum Using a Dual-Functional Material with Superior Hydrophilicity and Switchable Surface Charge. Anal. Chem. 2017, 89, 3966–3972. [Google Scholar] [CrossRef]
- Dubois, F.; Leroy, C.; Simon, V.; Benistant, C.; Roche, S. YES oncogenic activity is specified by its SH4 domain and regulates RAS/MAPK signaling in colon carcinoma cells. Am. J. Cancer Res. 2015, 5, 1972–1987. [Google Scholar]
- Vasaikar, S.; Huang, C.; Wang, X.; Petyuk, V.A.; Savage, S.R.; Wen, B.; Dou, Y.; Zhang, Y.; Shi, Z.; Arshad, O.A.; et al. Proteogenomic Analysis of Human Colon Cancer Reveals New Therapeutic Opportunities. Cell 2019, 177, 1035–1049.e19. [Google Scholar] [CrossRef]
- Lin, P.-C.; Yang, Y.-F.; Tyan, Y.-C.; Hsiao, E.S.L.; Chu, P.-C.; Lee, C.-T.; Lee, J.-C.; Chen, Y.-M.A.; Liao, P.-C. Identification of Phosphorylated Cyclin-Dependent Kinase 1 Associated with Colorectal Cancer Survival Using Label-Free Quantitative Analyses. PLoS ONE 2016, 11, e0158844. [Google Scholar] [CrossRef]
- Rasmussen, M.H.; Lyskjær, I.; Jersie-Christensen, R.R.; Tarpgaard, L.S.; Primdal-Bengtson, B.; Nielsen, M.M.; Pedersen, J.S.; Hansen, T.P.; Hansen, F.; Olsen, J.V.; et al. miR-625-3p regulates oxaliplatin resistance by targeting MAP2K6-p38 signalling in human colorectal adenocarcinoma cells. Nat. Commun. 2016, 7, 12436. [Google Scholar] [CrossRef]
- Abe, Y.; Nagano, M.; Kuga, T.; Tada, A.; Isoyama, J.; Adachi, J.; Tomonaga, T. Deep Phospho- and Phosphotyrosine Proteomics Identified Active Kinases and Phosphorylation Networks in Colorectal Cancer Cell Lines Resistant to Cetuximab. Sci. Rep. 2017, 7, 10463. [Google Scholar] [CrossRef] [PubMed]
- Demircioglu, F.; Wang, J.; Candido, J.; Costa, A.S.H.; Casado, P.; Delgado, B.D.L.; Reynolds, L.E.; Gomez-Escudero, J.; Newport, E.; Rajeeve, V.; et al. Cancer associated fibroblast FAK regulates malignant cell metabolism. Nat. Commun. 2020, 11, 1–18. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.-T.; Liao, L.-Z.; Lu, C.-H.; Huang, Y.-H.; Lin, Y.-K.; Lin, J.-H.; Chow, L.-P. Quantitative phosphoproteomic analysis identifies the potential therapeutic target EphA2 for overcoming sorafenib resistance in hepatocellular carcinoma cells. Exp. Mol. Med. 2020, 52, 497–513. [Google Scholar] [CrossRef] [PubMed]
- Peiris, D.; Ossondo, M.; Fry, S.; Loizidou, M.; Smith-Ravin, J.; Dwek, M.V. Identification of O-Linked Glycoproteins Binding to the Lectin Helix pomatia Agglutinin as Markers of Metastatic Colorectal Cancer. PLoS ONE 2015, 10, e0138345. [Google Scholar] [CrossRef]
- Lavrsen, K.; Dabelsteen, S.; Vakhrushev, S.Y.; Levann, A.M.R.; Haue, A.D.; Dylander, A.; Mandel, U.; Hansen, L.; Frödin, M.; Bennett, E.P.; et al. De novo expression of human polypeptide N-acetylgalactosaminyltransferase 6 (GalNAc-T6) in colon adenocarcinoma inhibits the differentiation of colonic epithelium. J. Biol. Chem. 2018, 293, 1298–1314. [Google Scholar] [CrossRef]
- Campos, D.; Freitas, D.; Gomes, J.; Magalhães, A.; Steentoft, C.; Gomes, C.; Vester-Christensen, M.B.; Ferreira, J.A.; Afonso, L.P.; Santos, L.L.; et al. Probing the O-glycoproteome of gastric cancer cell lines for biomarker discovery. Mol. Cell. Proteom. 2015, 14, 1616–1629. [Google Scholar] [CrossRef]
- Fernandes, E.; Freitas, R.; Ferreira, D.; Soares, J.; Azevedo, R.; Gaiteiro, C.; Peixoto, A.; Oliveira, S.; Cotton, S.; Relvas-Santos, M.; et al. Nucleolin-Sle A Glycoforms as E-Selectin Ligands and Potentially Targetable Biomarkers at the Cell Surface of Gastric Cancer Cells. Cancers 2020, 12, 861. [Google Scholar] [CrossRef]
- Zhang, X.-F.; Wang, J.; Jia, H.-L.; Zhu, W.-W.; Lu, L.; Ye, Q.-H.; Nelson, P.J.; Qin, Y.; Gao, D.-M.; Zhou, H.; et al. Core fucosylated glycan-dependent inhibitory effect of QSOX1-S on invasion and metastasis of hepatocellular carcinoma. Cell Death Discov. 2019, 5, 84. [Google Scholar] [CrossRef]
- Block, T.M.; Comunale, M.A.; Lowman, M.; Steel, L.F.; Romano, P.R.; Fimmel, C.; Tennant, B.C.; London, W.T.; Evans, A.A.; Blumberg, B.S.; et al. Use of targeted glycoproteomics to identify serum glycoproteins that correlate with liver cancer in woodchucks and humans. Proc. Natl. Acad. Sci. USA 2005, 102, 779–784. [Google Scholar] [CrossRef]
- Ang, I.L.; Poon, T.C.W.; Lai, P.B.S.; Chan, A.T.C.; Ngai, S.-M.; Hui, A.Y.; Johnson, P.J.; Sung, J.J.Y. Study of Serum Haptoglobin and Its Glycoforms in the Diagnosis of Hepatocellular Carcinoma: A Glycoproteomic Approach. J. Proteome Res. 2006, 5, 2691–2700. [Google Scholar] [CrossRef]
- Arnold, M.; Sierra, M.S.; Laversanne, M.; Soerjomataram, I.; Jemal, A.; Bray, F. Global patterns and trends in colorectal cancer incidence and mortality. Gut 2017, 66, 683–691. [Google Scholar] [CrossRef] [PubMed]
- Vatandoust, S.; Price, T.J.; Karapetis, C.S. Colorectal cancer: Metastases to a single organ. World J. Gastroenterol. 2015, 21, 11767–11776. [Google Scholar] [CrossRef] [PubMed]
- Haggar, F.A.; Boushey, R.P. Colorectal Cancer Epidemiology: Incidence, Mortality, Survival, and Risk Factors. Clin. Colon Rectal Surg. 2009, 22, 191. [Google Scholar] [CrossRef] [PubMed]
- Amata, I.; Maffei, M.; Pons, M. Phosphorylation of unique domains of Src family kinases. Front. Genet. 2014, 5, 181. [Google Scholar] [CrossRef] [PubMed]
- Sancier, F.; Dumont, A.; Sirvent, A.; De Plater, L.P.; Edmonds, T.; David, G.; Jan, M.; De Montrion, C.; Cogé, F.; Leonce, S.; et al. Specific Oncogenic Activity of the Src-Family Tyrosine Kinase c-Yes in Colon Carcinoma Cells. PLoS ONE 2011, 6, e17237. [Google Scholar] [CrossRef]
- De Roock, W.; Jonker, D.J.; Di Nicolantonio, F.; Sartore-Bianchi, A.; Tu, D.; Siena, S.; Lamba, S.; Arena, S.; Frattini, M.; Piessevaux, H.; et al. Association of KRAS p.G13D Mutation With Outcome in Patients With Chemotherapy-Refractory Metastatic Colorectal Cancer Treated With Cetuximab. JAMA 2010, 304, 1812–1820. [Google Scholar] [CrossRef]
- Hammond, D.E.; Mageean, C.J.; Rusilowicz, E.V.; Wickenden, J.A.; Clague, M.J.; Prior, I.A. Differential Reprogramming of Isogenic Colorectal Cancer Cells by Distinct Activating KRAS Mutations. J. Proteome Res. 2015, 14, 1535–1546. [Google Scholar] [CrossRef]
- Valastyan, S.; Weinberg, R.A. Tumor Metastasis: Molecular Insights and Evolving Paradigms. Cell 2011, 147, 275–292. [Google Scholar] [CrossRef]
- Schunter, A.J.; Yue, X.; Hummon, A.B. Phosphoproteomics of colon cancer metastasis: Comparative mass spectrometric analysis of the isogenic primary and metastatic cell lines SW480 and SW620. Anal. Bioanal. Chem. 2017, 409, 1749–1763. [Google Scholar] [CrossRef]
- Gherman, A.; Balacescu, L.; Gheorghe-Cetean, S.; Vlad, C.; Balacescu, O.; Irimie, A.; Lisencu, C. Current and New Predictors for Treatment Response in Metastatic Colorectal Cancer. The Role of Circulating miRNAs as Biomarkers. Int. J. Mol. Sci. 2020, 21, 2089. [Google Scholar] [CrossRef]
- Iida, M.; Brand, T.M.; Campbell, D.A.; Li, C.; Wheeler, D.L. Yes and Lyn play a role in nuclear translocation of the epidermal growth factor receptor. Oncogene 2013, 32, 759–767. [Google Scholar] [CrossRef]
- Kubiniok, P.; Lavoie, H.; Therrien, M.; Thibault, P. Time-resolved Phosphoproteome Analysis of Paradoxical RAF Activation Reveals Novel Targets of ERK. Mol. Cell. Proteom. 2017, 16, 663–679. [Google Scholar] [CrossRef] [PubMed]
- Taniguchi, H.; Baba, Y.; Sagiya, Y.; Gotou, M.; Nakamura, K.; Sawada, H.; Yamanaka, K.; Sakakibara, Y.; Mori, I.; Hikichi, Y.; et al. Biologic Response of Colorectal Cancer Xenograft Tumors to Sequential Treatment with Panitumumab and Bevacizumab. Neoplasia 2018, 20, 668–677. [Google Scholar] [CrossRef] [PubMed]
- Stewart, A.; Coker, E.A.; Pölsterl, S.; Georgiou, A.; Minchom, A.; Carreira, S.; Cunningham, D.; O’Brien, M.; Raynaud, F.I.; De Bono, J.S.; et al. Differences in signaling patterns on PI3K inhibition reveal context specificity in KRAS mutant cancers. Mol. Cancer Ther. 2019, 18, 1396–1404. [Google Scholar] [CrossRef] [PubMed]
- Rawla, P.; Barsouk, A. Epidemiology of gastric cancer: Global trends, risk factors and prevention. Gastroenterol. Rev. 2019, 14, 26. [Google Scholar] [CrossRef]
- Pucułek, M.; Machlowska, J.; Wierzbicki, R.; Baj, J.; Maciejewski, R.; Sitarz, R. Helicobacter pylori associated factors in the development of gastric cancer with special reference to the early-onset subtype. Oncotarget 2018, 9, 31146. [Google Scholar] [CrossRef]
- Wroblewski, L.E.; Peek, R.M.; Wilson, K.T. Helicobacter pylori and Gastric Cancer: Factors That Modulate Disease Risk. Clin. Microbiol. Rev. 2010, 23, 713–739. [Google Scholar] [CrossRef]
- Keller, S.; Kneissl, J.; Grabher-Meier, V.; Heindl, S.; Hasenauer, J.; Maier, D.; Mattes, J.; Ewinter, P.; Luber, B. Evaluation of epidermal growth factor receptor signaling effects in gastric cancer cell lines by detailed motility-focused phenotypic characterization linked with molecular analysis. BMC Cancer 2017, 17, 845. [Google Scholar] [CrossRef]
- Chaturvedi, R.; Asim, M.; Romero–Gallo, J.; Barry, D.P.; Hoge, S.; De Sablet, T.; Delgado, A.G.; Wroblewski, L.E.; Piazuelo, M.B.; Yan, F.; et al. Spermine Oxidase Mediates the Gastric Cancer Risk Associated With Helicobacter pylori CagA. Gastroenterology 2011, 141, 1696–1708.e2. [Google Scholar] [CrossRef]
- Chaturvedi, R.; Asim, M.; Piazuelo, M.B.; Yan, F.; Barry, D.P.; Sierra, J.C.; Delgado, A.G.; Hill, S.; Casero, R.A.; Bravo, L.E.; et al. Activation of EGFR and ERBB2 by Helicobacter pylori Results in Survival of Gastric Epithelial Cells With DNA Damage. Gastroenterology 2014, 146, 1739–1751.e14. [Google Scholar] [CrossRef]
- Ansari, S.; Gantuya, B.; Tuan, V.P.; Yamaoka, Y. Diffuse Gastric Cancer: A Summary of Analogous Contributing Factors for Its Molecular Pathogenicity. Int. J. Mol. Sci. 2018, 19, 2424. [Google Scholar] [CrossRef] [PubMed]
- Tong, M.; Yu, C.; Shi, J.; Huang, W.; Ge, S.; Liu, M.; Song, L.; Zhan, D.; Xia, X.; Liu, W.; et al. Phosphoproteomics Enables Molecular Subtyping and Nomination of Kinase Candidates for Individual Patients of Diffuse-Type Gastric Cancer. iScience 2019, 22, 44–57. [Google Scholar] [CrossRef] [PubMed]
- Lee, Y.Y.; Kim, H.-P.; Kang, M.J.; Cho, B.-K.; Han, S.-W.; Kim, T.-Y.; Yi, E.C. Phosphoproteomic analysis identifies activated MET-axis PI3K/AKT and MAPK/ERK in lapatinib-resistant cancer cell line. Exp. Mol. Med. 2013, 45, e64. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Rawla, P.; Sunkara, T.; Gaduputi, V. Epidemiology of Pancreatic Cancer: Global Trends, Etiology and Risk Factors. World J. Oncol. 2019, 10, 10–27. [Google Scholar] [CrossRef] [PubMed]
- Bian, Y.; Teper, Y.; Griner, L.A.M.; Aiken, T.; Shukla, V.; Guha, R.; Shinn, P.; Xin, H.-W.; Pflicke, H.; Powers, A.; et al. Target Deconvolution of a Multikinase Inhibitor with Antimetastatic Properties Identifies TAOK3 as a Key Contributor to a Cancer Stem Cell–Like Phenotype. Mol. Cancer Ther. 2019, 18, 2097–2110. [Google Scholar] [CrossRef]
- Yachida, S.; A Iacobuzio-Donahue, C. The pathology and genetics of metastatic pancreatic cancer. Arch. Pathol. Lab. Med. 2009, 133, 413–422. [Google Scholar] [PubMed]
- Kim, M.-S.; Zhong, Y.; Yachida, S.; RajeshKumar, N.V.; Abel, M.L.; Marimuthu, A.; Mudgal, K.; Hruban, R.H.; Poling, J.S.; Tyner, J.W.; et al. Heterogeneity of Pancreatic Cancer Metastases in a Single Patient Revealed by Quantitative Proteomics. Mol. Cell. Proteom. 2014, 13, 2803–2811. [Google Scholar] [CrossRef]
- Tape, C.J.; Ling, S.; Dimitriadi, M.; McMahon, K.M.; Worboys, J.D.; Leong, H.S.; Norrie, I.C.; Miller, C.J.; Poulogiannis, G.; Lauffenburger, D.A.; et al. Oncogenic KRAS regulates tumor cell signaling via stromal reciprocation. Cell 2016, 165, 910–920. [Google Scholar] [CrossRef]
- Chen, P.-Y.; Muzumdar, M.D.; Dorans, K.J.; Robbins, R.; Bhutkar, A.; Del Rosario, A.M.; Mertins, P.; Qiao, J.; Schafer, A.C.; Gertler, F.B.; et al. Adaptive and Reversible Resistance to Kras Inhibition in Pancreatic Cancer Cells. Cancer Res. 2017, 78, 985–1002. [Google Scholar] [CrossRef]
- Yang, J.D.; Hainaut, P.; Gores, G.J.; Amadou, A.; Plymoth, A.; Roberts, L.R. A global view of hepatocellular carcinoma: Trends, risk, prevention and management. Nat. Rev. Gastroenterol. Hepatol. 2019, 16, 589–604. [Google Scholar] [CrossRef]
- Dhanasekaran, R.; Limaye, A.; Cabrera, R. Hepatocellular carcinoma: Current trends in worldwide epidemiology, risk factors, diagnosis, and therapeutics. Hepat. Med. 2012, 4, 19. [Google Scholar] [PubMed]
- Le Grazie, M.; Biagini, M.R.; Tarocchi, M.; Polvani, S.; Galli, A. Chemotherapy for hepatocellular carcinoma: The present and the future. World J. Hepatol. 2017, 9, 907. [Google Scholar] [CrossRef] [PubMed]
- Ren, L.; Li, C.; Wang, Y.; Teng, Y.; Sun, H.; Xing, B.; Yang, X.; Jiang, Y.; He, F. In Vivo Phosphoproteome Analysis Reveals Kinome Reprogramming in Hepatocellular Carcinoma. Mol. Cell. Proteom. 2018, 17, 1067–1083. [Google Scholar] [CrossRef] [PubMed]
- El-Serag, H.B. Epidemiology of Viral Hepatitis and Hepatocellular Carcinoma. Gastroenterology 2012, 142, 1264–1273.e1. [Google Scholar] [CrossRef]
- Lu, N.T.; Liu, N.M.; Vu, J.Q.; Patel, D.; Cohn, W.; Capri, J.; Ziegler, M.; Patel, N.; Tramontano, A.; Williams, R.; et al. Phospho-Network Analysis Identifies and Quantifies Hepatitis C Virus (HCV)-induced Hepatocellular Carcinoma (HCC) Proteins Regulating Viral-mediated Tumor Growth. Cancer Genom. Proteom. 2016, 13, 339–357. [Google Scholar]
- Abdallah, C.; Lejamtel, C.; Benzoubir, N.; Battaglia, S.; SidAhmed-Adrar, N.; Desterke, C.; Lemasson, M.; Rosenberg, A.R.; Samuel, D.; Brechot, C.; et al. Hepatitis C virus core protein targets 4E-BP1 expression and phosphorylation and potentiates Myc-induced liver carcinogenesis in transgenic mice. Oncotarget 2017, 8, 56228. [Google Scholar] [CrossRef]
- Jiang, Y.; Sun, A.; Zhao, Y.; Ying, W.; Sun, H.; Yang, X.; Xing, B.; Sun, W.; Ren, L.; Chinese Human Proteome Project (CNHPP) Consortium; et al. Proteomics identifies new therapeutic targets of early-stage hepatocellular carcinoma. Nat. Cell Biol. 2019, 567, 257–261. [Google Scholar] [CrossRef]
- Liu, Z.; Wang, Y.; Yao, Y.; Fang, Z.; Miao, Q.R.; Ye, M. Quantitative proteomic and phosphoproteomic studies reveal novel 5-fluorouracil resistant targets in hepatocellular carcinoma. J. Proteom. 2019, 208, 103501. [Google Scholar] [CrossRef]
- Melas, I.N.; Lauffenburger, U.A.; Alexopoulos, L.G. Identification of signaling pathways related to drug efficacy in hepatocellular carcinoma via integration of phosphoproteomic, genomic and clinical data. In Proceedings of the 13th IEEE International Conference on BioInformatics and BioEngineering, Chania, Greece, 10–13 November 2013. [Google Scholar]
- Fernández-Varo, G.; Perramon, M.; Carvajal, S.; Oró, D.; Casals, E.; Boix, L.; Oller, L.; Macías-Muñoz, L.; Marfà, S.; Casals, G.; et al. Bespoken nanoceria: A new effective treatment in experimental hepatocellular carcinoma. Hepatology 2020. [Google Scholar] [CrossRef]
- Kailemia, M.J.; Park, D.; Lebrilla, C.B. Glycans and glycoproteins as specific biomarkers for cancer. Anal. Bioanal. Chem. 2017, 409, 395–410. [Google Scholar] [CrossRef]
- Peixoto, A.; Relvas-Santos, M.; Azevedo, R.; Santos, L.L.; Ferreira, J.A. Protein Glycosylation and Tumor Microenvironment Alterations Driving Cancer Hallmarks. Front. Oncol. 2019, 9, 380. [Google Scholar] [CrossRef] [PubMed]
- Pan, S.; Chen, R.; Tamura, Y.; Crispin, D.A.; Lai, L.A.; May, D.H.; McIntosh, M.W.; Goodlett, D.R.; Brentnall, T.A. Quantitative Glycoproteomics Analysis Reveals Changes in N-Glycosylation Level Associated with Pancreatic Ductal Adenocarcinoma. J. Proteome Res. 2014, 13, 1293–1306. [Google Scholar] [CrossRef] [PubMed]
- Gao, H.-J.; Chen, Y.-J.; Zuo, D.; Xiao, M.-M.; Li, Y.; Guo, H.; Zhang, N.; Chen, R.-B. Quantitative proteomic analysis for high-throughput screening of differential glycoproteins in hepatocellular carcinoma serum. Cancer Biol. Med. 2015, 12, 246. [Google Scholar] [PubMed]
- Barefoot, M.E.; Varghese, R.S.; Zhou, Y.; Di Poto, C.; Ferrarini, A.; Ressom, H.W. Multi-omic Pathway and Network Analysis to Identify Biomarkers for Hepatocellular Carcinoma. In Proceedings of the 2019 41st Anual International Conference of the IEEE Engineering in Medicine and Biology Society (EMBC), Berlin, Germany, 23–27 July 2019. [Google Scholar]
| Study | Developed Strategy | Sample | Results | References |
|---|---|---|---|---|
| Phosphoproteomic studies | ||||
| Song et al. 2011 | Pseudo triplex stable isotope dimethyl labeling approach coupled with on line RP-SCX-RP LC-MS/MS | Hepatocellular carcinoma (HCC) and normal human liver tissues | 1934 phosphopeptides from 1033 phosphoproteins | [33] |
| Lin et al. 2017 | Stable isotope dimethylation labeling coupled with online 3D SCX-TiO2/RP LC-MS/MS and super-SILAC mix coupled with SIM/AIMS | Human HCC tissue | 7868 phosphopeptides | [34] |
| Abe et al. 2020 | Fe3+ IMAC-TMT labeling-SCX LC-MS/MS | Gastric cancer patients endoscopic biopsy specimens | 4034 phosphoproteins | [10] |
| Glycoproteomic studies | ||||
| Zhou et al. 2007 | Two-dimensional gel electrophoresis (2-DE) followed by the fluorescence staining of glycoprotein-MALDI-TOF-MS/MS | Three Human HCC cell lines | 80 glycoproteins | [35] |
| Cao et al. 2009 | Glycopeptide enrichment methods; hydrophilic affinity (HA) and hydrazide chemistry (HC), were used complementarily-LC-MS/MS | Human HCC cells | 300 glycosylation sites within 194 glycoproteins | [36] |
| Sun et al. 2014 | HC- multiple protease digestion-dimethyl labeling -SCX-RP LC-MS/MS | Human HCC and healthy liver tissues | 2329 N-glycosites on 1052 N-glycoproteins | [37] |
| Jiang et al. 2019 | A multi-parallel enrichment strategy based on the optimized ZIC-HILIC enrichment method assisted by a filter-coated 96-well plate-MALDI-TOF MS | Three HCC cell lines | 5466 N-glycosites in 2383 glycoproteins | [38] |
| Protein Name | Sample Type | Method | Clinical Significance | References |
|---|---|---|---|---|
| Phosphoproteomics studies | ||||
| Phosphorylated YES (Src family) | CRC cell line | SILAC-HPLC-ESI-MS/MS | Therapeutic target | [67] |
| Rb phosphorylation | Colon cancer tumor tissues | IMAC–LC/MS/MS | Therapeutic target | [68] |
| CDK1 pTyr15 | CRC cell line | TiO2-LC-MS/MS | Prognostic biomarker | [69] |
| miR-625-3p | CRC cell line | SILAC-TiO2-LC-MS/MS | predictive biomarker for oxPt-resistance | [70] |
| Src family | CRC cell line | IMAC-TMT-LC-MS/MS | Therapeutic targets in cetuximab-resistant CRC | [71] |
| FAK | Mouse model of PC | TiO2-LC-MS/MS | Survival rate | [72] |
| EphA2 | PDAC cell line | SILAC-TiO2-LC-MS/MS | Therapeutic target in sorafenib resistant CRC | [73] |
| Glycoproteomics studies | ||||
| Annexin 4 | lymph node positive CRC tissue samples | Lectin HPA- 2DE - MALDI-MS | Survival rate | [74] |
| GalNAc-T6 | colon cancer cell line | VVA LWAC-nLC- nES-MS/MS | Cancer development | [75] |
| CD44 and GalNAc-T5, with O-glycan STn | cell line and sear of patient with gastric cancer | VVA LWAC- nLC- nES -MS/MS | Diagnostic biomarker | [76] |
| Nucleolin (NCL)-SLeA glycoforms | Gastric cancer cell models | anti-SLeA -nLC-nES-MS/MS | Worst prognosis | [77] |
| C3 with Man5, Man6, or Man7 glycoform at Asn85 | HCC patient plasma samples | LC-MS/MS | Indicator of HCC tumor grade | [11] |
| cf-QSOX1 | HCC patient serum samples | LAC- nLC-ESI-MS/MS | Biomarker for postoperative recurrence of HCC | [78] |
| GP73 | animal models (woodchucks) of HCC | LAC-2DE-HPLC-MS/MS | Diagnostic biomarker | [79] |
| Hp | HCC patient serum samples | LAC-2DE- MALDI- MS/MS | Diagnostic biomarker | [80] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Abyadeh, M.; Meyfour, A.; Gupta, V.; Zabet Moghaddam, M.; Fitzhenry, M.J.; Shahbazian, S.; Hosseini Salekdeh, G.; Mirzaei, M. Recent Advances of Functional Proteomics in Gastrointestinal Cancers- a Path towards the Identification of Candidate Diagnostic, Prognostic, and Therapeutic Molecular Biomarkers. Int. J. Mol. Sci. 2020, 21, 8532. https://doi.org/10.3390/ijms21228532
Abyadeh M, Meyfour A, Gupta V, Zabet Moghaddam M, Fitzhenry MJ, Shahbazian S, Hosseini Salekdeh G, Mirzaei M. Recent Advances of Functional Proteomics in Gastrointestinal Cancers- a Path towards the Identification of Candidate Diagnostic, Prognostic, and Therapeutic Molecular Biomarkers. International Journal of Molecular Sciences. 2020; 21(22):8532. https://doi.org/10.3390/ijms21228532
Chicago/Turabian StyleAbyadeh, Morteza, Anna Meyfour, Vivek Gupta, Masoud Zabet Moghaddam, Matthew J. Fitzhenry, Shila Shahbazian, Ghasem Hosseini Salekdeh, and Mehdi Mirzaei. 2020. "Recent Advances of Functional Proteomics in Gastrointestinal Cancers- a Path towards the Identification of Candidate Diagnostic, Prognostic, and Therapeutic Molecular Biomarkers" International Journal of Molecular Sciences 21, no. 22: 8532. https://doi.org/10.3390/ijms21228532
APA StyleAbyadeh, M., Meyfour, A., Gupta, V., Zabet Moghaddam, M., Fitzhenry, M. J., Shahbazian, S., Hosseini Salekdeh, G., & Mirzaei, M. (2020). Recent Advances of Functional Proteomics in Gastrointestinal Cancers- a Path towards the Identification of Candidate Diagnostic, Prognostic, and Therapeutic Molecular Biomarkers. International Journal of Molecular Sciences, 21(22), 8532. https://doi.org/10.3390/ijms21228532







