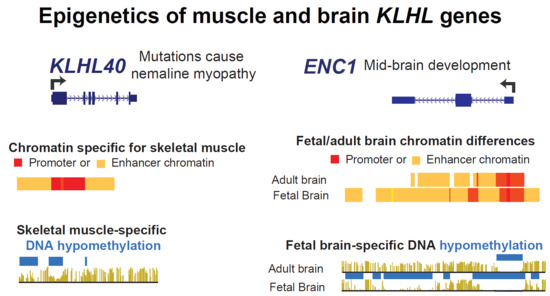
Epigenetics of Muscle- and Brain-Specific Expression of KLHL Family Genes
Abstract
1. Introduction
2. Results
2.1. Many KLHL and KBTBD Genes Are Expressed Preferentially in SkM or Brain
2.2. Extensive Intragenic Promoter Chromatin and Overlapping DNA Hypomethylation Correlates with the Extremely High Expression of KLHL41 in Skeletal Muscle
2.3. Intragenic and Intergenic Enhancer Chromatin at KLHL40 or Its Neighbor, HHATL May Upregulate Both Genes or One or the Other Gene Depending on the Tissue
2.4. Skeletal Muscle-Specific Expression, Promoter Chromatin, and Enhancer Chromatin in KLHL30 and in KLHL38, a Paralog That Contains a Retrogene from KLHL30
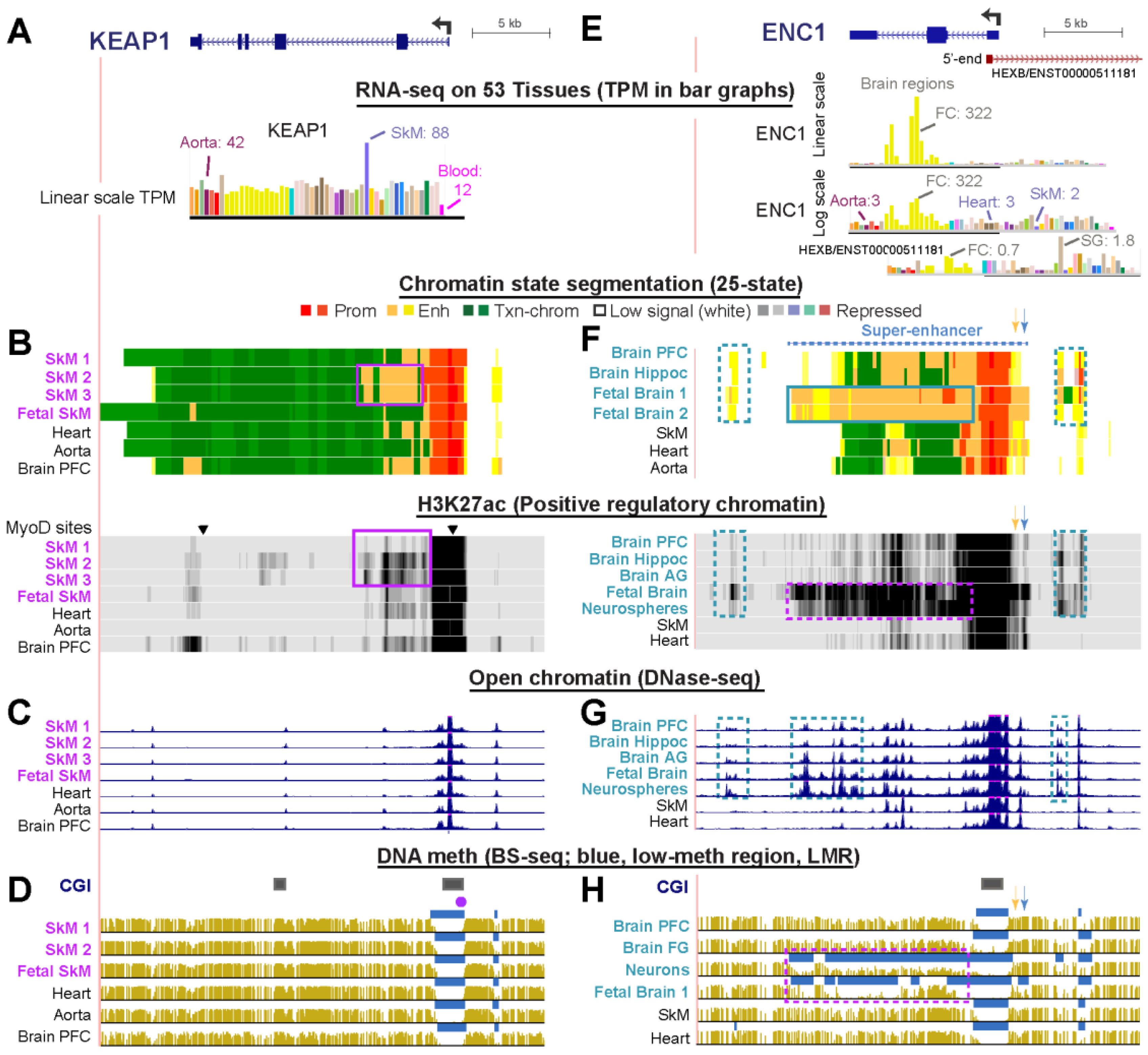
2.5. Only Minor Skeletal Muscle-Associated Epigenetic Differences Occur in KEAP1 Consistent with Its Highest Expression in SkM But Otherwise Broad Tissue Expression Profile
2.6. The Neurogenesis-Associated ENC1 Gene Exhibited Much More Enhancer Chromatin and DNA Hypomethylation in Fetal than in Adult Brain
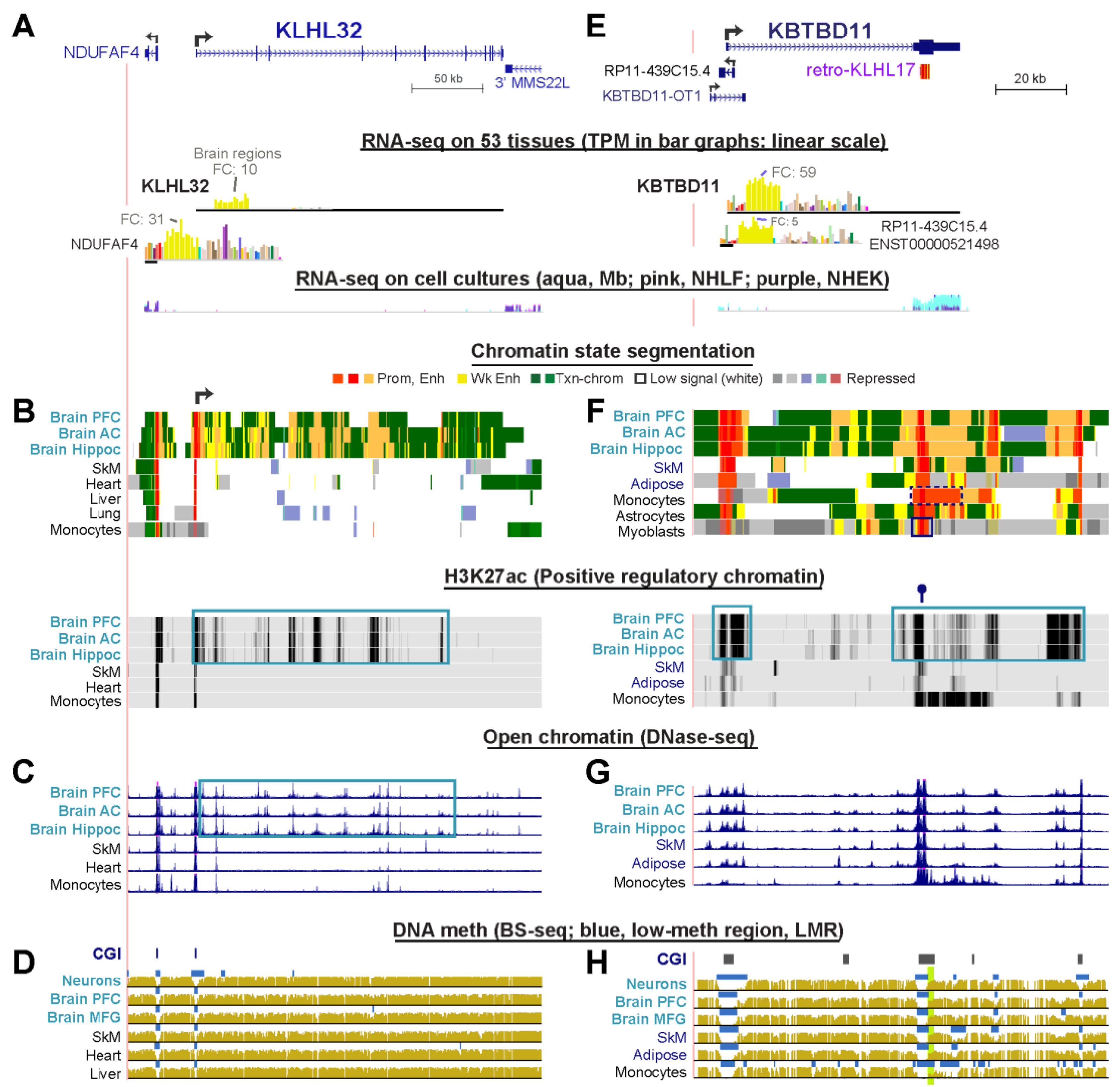
2.7. The Strong Brain-Specificity of KLHL32 Is Mirrored by Clusters of Intragenic Enhancer Chromatin Seen Only in Brain
2.8. KBTBD11, a Brain-Specific Gene, Has a Retrogene Overlaying a 3′ Promoter for a Novel Noncoding RNA Gene
2.9. Overview of Epigenetic Features Associated with the Studied KLHF Family Genes
3. Discussion
4. Methods
4.1. RNA-Seq for Tissues and Cells
4.2. Databases and Analyses Used for Epigenetics Studies
4.3. Alignments and Phylogenetic Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| BS | bisulfite |
| CAGE | 5′ Cap Analysis of Gene Expression |
| CGI | CpG island |
| ChIP-seq | chromatin immunoprecipitation with next-gen sequencing |
| CTCF | CCCTC-binding factor |
| Enh | enhancer |
| FPKM | fragments per kilobase of exon per million reads mapped |
| H3K27ac | Histone H3 lysine 27 acetylation |
| H3K4me1 | histone H3 lysine 4 monomethylation |
| H3K4me3 | histone H3 lysine 4 trimethylation |
| LCL | lymphoblastoid cell line |
| LMR | low methylated region |
| Micro-C | a high-resolution type of chromatin capture analysis |
| ncRNA | non-coding RNA |
| NHLF | normal human lung fibroblasts |
| ORF | open reading frame |
| PBMC | peripheral blood mononuclear cells |
| Prom | promoter |
| SkM | skeletal muscle |
| TAD | topologically associating domain |
| TPM | transcripts per kilobase million |
| TSS | transcription start site |
| Txn | transcription |
References
- Guo, Y.; Yu, S.; Zhang, C.; Kong, A.N. Epigenetic regulation of Keap1-Nrf2 signaling. Free Radic. Biol. Med. 2015, 88, 337–349. [Google Scholar] [CrossRef]
- Varshavsky, A. The Ubiquitin System, Autophagy, and Regulated Protein Degradation. Annu. Rev. Biochem. 2017, 86, 123–128. [Google Scholar] [CrossRef]
- Mumtaz, P.T.; Taban, Q.; Dar, M.A.; Mir, S.; Haq, Z.U.; Zargar, S.M.; Shah, R.A.; Ahmad, S.M. Deep Insights in Circular RNAs: From biogenesis to therapeutics. Biol. Proced. Online 2020, 22, 10. [Google Scholar] [CrossRef]
- Roadmap Epigenomics Consortium; Kundaje, A.; Meuleman, W.; Ernst, J.; Bilenky, M.; Yen, A.; Heravi-Moussavi, A.; Kheradpour, P.; Zhang, Z.; Wang, J.; et al. Integrative analysis of 111 reference human epigenomes. Nature 2015, 518, 317–330. [Google Scholar] [CrossRef]
- Dhanoa, B.S.; Cogliati, T.; Satish, A.G.; Bruford, E.A.; Friedman, J.S. Update on the Kelch-like (KLHL) gene family. Hum. Genom. 2013, 7, 13. [Google Scholar] [CrossRef] [PubMed]
- Shi, X.; Xiang, S.; Cao, J.; Zhu, H.; Yang, B.; He, Q.; Ying, M. Kelch-like proteins: Physiological functions and relationships with diseases. Pharm. Res. 2019, 148, 104404. [Google Scholar] [CrossRef]
- Gupta, V.A.; Beggs, A.H. Kelch proteins: Emerging roles in skeletal muscle development and diseases. Skelet. Muscle 2014, 4, 11. [Google Scholar] [CrossRef]
- Elshaer, M.; ElManawy, A.I.; Hammad, A.; Namani, A.; Wang, X.J.; Tang, X. Integrated data analysis reveals significant associations of KEAP1 mutations with DNA methylation alterations in lung adenocarcinomas. Aging (Albany NY) 2020, 12, 7183–7206. [Google Scholar] [CrossRef]
- Sewry, C.A.; Laitila, J.M.; Wallgren-Pettersson, C. Nemaline myopathies: A current view. J. Muscle Res. Cell Motil. 2019, 40, 111–126. [Google Scholar] [CrossRef] [PubMed]
- Jirka, C.; Pak, J.H.; Grosgogeat, C.A.; Marchetii, M.M.; Gupta, V.A. Dysregulation of NRAP degradation by KLHL41 contributes to pathophysiology in Nemaline Myopathy. Hum. Mol. Genet. 2019, 28, 2549–2560. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Z.; Turer, E.; Li, X.; Zhan, X.; Choi, M.; Tang, M.; Press, A.; Smith, S.R.; Divoux, A.; Moresco, E.M.; et al. Insulin resistance and diabetes caused by genetic or diet-induced KBTBD2 deficiency in mice. Proc. Natl. Acad. Sci. USA 2016, 113, E6418–E6426. [Google Scholar] [CrossRef]
- Yoshida, S.; Araki, Y.; Mori, T.; Sasaki, E.; Kasagi, Y.; Isobe, K.; Susa, K.; Inoue, Y.; Bomont, P.; Okado, T.; et al. Decreased KLHL3 expression is involved in the pathogenesis of pseudohypoaldosteronism type II caused by cullin 3 mutation in vivo. Clin. Exp. Nephrol. 2018, 22, 1251–1257. [Google Scholar] [CrossRef] [PubMed]
- Hedberg-Oldfors, C.; Abramsson, A.; Osborn, D.P.S.; Danielsson, O.; Fazlinezhad, A.; Nilipour, Y.; Hubbert, L.; Nennesmo, I.; Visuttijai, K.; Bharj, J.; et al. Cardiomyopathy with lethal arrhythmias associated with inactivation of KLHL24. Hum. Mol. Genet. 2019, 28, 1919–1929. [Google Scholar] [CrossRef]
- Deaton, A.M.; Bird, A. CpG islands and the regulation of transcription. Genes Dev. 2011, 25, 1010–1022. [Google Scholar] [CrossRef]
- Ehrlich, K.C.; Paterson, H.L.; Lacey, M.; Ehrlich, M. DNA hypomethylation in intragenic and intergenic enhancer chromatin of muscle-specific genes usually correlates with their expression. Yale J. Biol. Med. 2016, 89, 441–455. [Google Scholar]
- Ehrlich, K.C.; Lacey, M.; Ehrlich, M. Epigenetics of Skeletal Muscle-Associated Genes in the ASB, LRRC, TMEM, and OSBPL Gene Families. Epigenomes 2020, 4, 1. [Google Scholar] [CrossRef]
- Heberle, E.; Bardet, A.F. Sensitivity of transcription factors to DNA methylation. Essays Biochem. 2019, 63, 727–741. [Google Scholar] [CrossRef] [PubMed]
- Lazaris, C.; Aifantis, I.; Tsirigos, A. On Epigenetic Plasticity and Genome Topology. Trends Cancer 2020, 6, 177–180. [Google Scholar] [CrossRef] [PubMed]
- GTEx_Consortium. Human genomics. The Genotype-Tissue Expression (GTEx) pilot analysis: Multitissue gene regulation in humans. Science 2015, 348, 648–660. [Google Scholar] [CrossRef]
- Illingworth, R.S.; Gruenewald-Schneider, U.; De Sousa, D.; Webb, S.; Merusi, C.; Kerr, A.R.; James, K.D.; Smith, C.; Walker, R.; Andrews, R.; et al. Inter-individual variability contrasts with regional homogeneity in the human brain DNA methylome. Nucleic Acids Res. 2015, 43, 732–744. [Google Scholar] [CrossRef]
- Dayalan Naidu, S.; Dinkova-Kostova, A.T. KEAP1, a cysteine-based sensor and a drug target for the prevention and treatment of chronic disease. Open Biol. 2020, 10, 200105. [Google Scholar] [CrossRef] [PubMed]
- Pirinen, M.; Lappalainen, T.; Zaitlen, N.A.; GTEx_Consortium; Dermitzakis, E.T.; Donnelly, P.; McCarthy, M.I.; Rivas, M.A. Assessing allele-specific expression across multiple tissues from RNA-seq read data. Bioinformatics 2015, 31, 2497–2504. [Google Scholar] [CrossRef]
- Searle, B.C.; Gittelman, R.M.; Manor, O.; Akey, J.M. Detecting Sources of Transcriptional Heterogeneity in Large-Scale RNA-Seq Data Sets. Genetics 2016, 204, 1391–1396. [Google Scholar] [CrossRef][Green Version]
- Haeussler, M.; Zweig, A.S.; Tyner, C.; Speir, M.L.; Rosenbloom, K.R.; Raney, B.J.; Lee, C.M.; Lee, B.T.; Hinrichs, A.S.; Gonzalez, J.N.; et al. The UCSC Genome Browser database: 2019 update. Nucleic Acids Res. 2019, 47, D853–D858. [Google Scholar] [CrossRef] [PubMed]
- Gheorghe, M.; Sandve, G.K.; Khan, A.; Cheneby, J.; Ballester, B.; Mathelier, A. A map of direct TF-DNA interactions in the human genome. Nucleic Acids Res. 2019, 47, 7715. [Google Scholar] [CrossRef] [PubMed]
- Xi, H.; Shulha, H.P.; Lin, J.M.; Vales, T.R.; Fu, Y.; Bodine, D.M.; McKay, R.D.; Chenoweth, J.G.; Tesar, P.J.; Furey, T.S.; et al. Identification and characterization of cell type-specific and ubiquitous chromatin regulatory structures in the human genome. PLoS Genet. 2007, 3, e136. [Google Scholar] [CrossRef] [PubMed]
- Whyte, W.A.; Orlando, D.A.; Hnisz, D.; Abraham, B.J.; Lin, C.Y.; Kagey, M.H.; Rahl, P.B.; Lee, T.I.; Young, R.A. Master transcription factors and mediator establish super-enhancers at key cell identity genes. Cell 2013, 153, 307–319. [Google Scholar] [CrossRef]
- Khan, A.; Zhang, X. dbSUPER: A database of super-enhancers in mouse and human genome. Nucleic Acids Res. 2016, 44, D164–D171. [Google Scholar] [CrossRef]
- Blondelle, J.; Tallapaka, K.; Seto, J.T.; Ghassemian, M.; Clark, M.; Laitila, J.M.; Bournazos, A.; Singer, J.D.; Lange, S. Cullin-3 dependent deregulation of ACTN1 represents a new pathogenic mechanism in nemaline myopathy. JCI Insight 2019, 4, 1–19. [Google Scholar] [CrossRef]
- Bowlin, K.M.; Embree, L.J.; Garry, M.G.; Garry, D.J.; Shi, X. Kbtbd5 is regulated by MyoD and restricted to the myogenic lineage. Differentiation 2013, 86, 184–191. [Google Scholar] [CrossRef]
- Cao, Y.; Yao, Z.; Sarkar, D.; Lawrence, M.; Sanchez, G.J.; Parker, M.H.; MacQuarrie, K.L.; Davison, J.; Morgan, M.T.; Ruzzo, W.L.; et al. Genome-wide MyoD binding in skeletal muscle cells: A potential for broad cellular reprogramming. Dev. Cell 2010, 18, 662–674. [Google Scholar] [CrossRef]
- Hsieh, T.S.; Fudenberg, G.; Goloborodko, A.; Rando, O.J. Micro-C XL: Assaying chromosome conformation from the nucleosome to the entire genome. Nat. Methods 2016, 13, 1009–1011. [Google Scholar] [CrossRef]
- Van, B.; Nishi, M.; Komazaki, S.; Ichimura, A.; Kakizawa, S.; Nakanaga, K.; Aoki, J.; Park, K.H.; Ma, J.; Ueyama, T.; et al. Mitsugumin 56 (hedgehog acyltransferase-like) is a sarcoplasmic reticulum-resident protein essential for postnatal muscle maturation. FEBS Lett. 2015, 589, 1095–1104. [Google Scholar] [CrossRef] [PubMed]
- Zhang, P.; Zhang, L.; Li, Y.; Zhu, S.; Zhao, M.; Ding, S.; Li, J. Quantitative Proteomic Analysis To Identify Differentially Expressed Proteins in Myocardium of Epilepsy Using iTRAQ Coupled with Nano-LC-MS/MS. J. Proteome Res. 2018, 17, 305–314. [Google Scholar] [CrossRef]
- de Winter, J.M.; Molenaar, J.P.; Yuen, M.; van der Pijl, R.; Shen, S.; Conijn, S.; van de Locht, M.; Willigenburg, M.; Bogaards, S.J.; van Kleef, E.S.; et al. KBTBD13 is an actin-binding protein that modulates muscle kinetics. J. Clin. Investig. 2020, 130, 754–767. [Google Scholar] [CrossRef]
- Sartorelli, V.; Lauberth, S.M. Enhancer RNAs are an important regulatory layer of the epigenome. Nat. Struct. Mol. Biol. 2020, 27, 521–528. [Google Scholar] [CrossRef] [PubMed]
- Bodine, S.C.; Baehr, L.M. Skeletal muscle atrophy and the E3 ubiquitin ligases MuRF1 and MAFbx/atrogin-1. Am. J. Physiol. Endocrinol. Metab. 2014, 307, E469–E484. [Google Scholar] [CrossRef]
- de O’Coelho, P.; Guarnier, F.A.; Figueiredo, L.B.; Zaramela, L.S.; Pacini, E.S.A.; Godinho, R.O.; Gomes, M.D. Identification of potential target genes associated with the reversion of androgen-dependent skeletal muscle atrophy. Arch. Biochem. Biophys. 2019, 663, 173–182. [Google Scholar] [CrossRef]
- Baertsch, R.; Diekhans, M.; Kent, W.J.; Haussler, D.; Brosius, J. Retrocopy contributions to the evolution of the human genome. BMC Genom. 2008, 9, 466. [Google Scholar] [CrossRef]
- Sukari, A.; Muqbil, I.; Mohammad, R.M.; Philip, P.A.; Azmi, A.S. F-BOX proteins in cancer cachexia and muscle wasting: Emerging regulators and therapeutic opportunities. Semin. Cancer Biol. 2016, 36, 95–104. [Google Scholar] [CrossRef]
- Bellezza, I.; Giambanco, I.; Minelli, A.; Donato, R. Nrf2-Keap1 signaling in oxidative and reductive stress. Biochim. Biophys. Acta Mol. Cell Res. 2018, 1865, 721–733. [Google Scholar] [CrossRef]
- van Rooij, E.; Quiat, D.; Johnson, B.A.; Sutherland, L.B.; Qi, X.; Richardson, J.A.; Kelm, R.J., Jr.; Olson, E.N. A family of microRNAs encoded by myosin genes governs myosin expression and muscle performance. Dev. Cell 2009, 17, 662–673. [Google Scholar] [CrossRef]
- Wang, X.J.; Zhang, D.D. Ectodermal-neural cortex 1 down-regulates Nrf2 at the translational level. PLoS ONE 2009, 4, e5492. [Google Scholar] [CrossRef]
- Mesman, S.; Kruse, S.J.; Smidt, M.P. Expression analyzes of early factors in midbrain differentiation programs. Gene Expr. Patterns 2018, 27, 8–15. [Google Scholar] [CrossRef] [PubMed]
- Lister, R.; Mukamel, E.A.; Nery, J.R.; Urich, M.; Puddifoot, C.A.; Johnson, N.D.; Lucero, J.; Huang, Y.; Dwork, A.J.; Schultz, M.D.; et al. Global epigenomic reconfiguration during mammalian brain development. Science 2013, 341, 1237905. [Google Scholar] [CrossRef]
- White, C.C.; Yang, H.S.; Yu, L.; Chibnik, L.B.; Dawe, R.J.; Yang, J.; Klein, H.U.; Felsky, D.; Ramos-Miguel, A.; Arfanakis, K.; et al. Identification of genes associated with dissociation of cognitive performance and neuropathological burden: Multistep analysis of genetic, epigenetic, and transcriptional data. PLoS Med. 2017, 14, e1002287. [Google Scholar] [CrossRef]
- van den Oord, E.J.; Clark, S.L.; Xie, L.Y.; Shabalin, A.A.; Dozmorov, M.G.; Kumar, G.; Swedish Schizophrenia, C.; Vladimirov, V.I.; Magnusson, P.K.; Aberg, K.A. A Whole Methylome CpG-SNP Association Study of Psychosis in Blood and Brain Tissue. Schizophr. Bull. 2016, 42, 1018–1026. [Google Scholar] [CrossRef] [PubMed]
- Terragni, J.; Zhang, G.; Sun, Z.; Pradhan, S.; Song, L.; Crawford, G.E.; Lacey, M.; Ehrlich, M. Notch signaling genes: Myogenic DNA hypomethylation and 5-hydroxymethylcytosine. Epigenetics 2014, 9, 842–850. [Google Scholar] [CrossRef] [PubMed]
- Do, C.; Lang, C.F.; Lin, J.; Darbary, H.; Krupska, I.; Gaba, A.; Petukhova, L.; Vonsattel, J.P.; Gallagher, M.P.; Goland, R.S.; et al. Mechanisms and Disease Associations of Haplotype-Dependent Allele-Specific DNA Methylation. Am. J. Hum. Genet. 2016, 98, 934–955. [Google Scholar] [CrossRef]
- Worton, L.E.; Shi, Y.C.; Smith, E.J.; Barry, S.C.; Gonda, T.J.; Whitehead, J.P.; Gardiner, E.M. Ectodermal-Neural Cortex 1 Isoforms Have Contrasting Effects on MC3T3-E1 Osteoblast Mineralization and Gene Expression. J. Cell Biochem. 2017, 118, 2141–2150. [Google Scholar] [CrossRef]
- Watanabe, K.; Yokota, K.; Yoshida, K.; Matsumoto, A.; Iwamoto, S. Kbtbd11 contributes to adipocyte homeostasis through the activation of upstream stimulatory factor 1. Heliyon 2019, 5, e02777. [Google Scholar] [CrossRef] [PubMed]
- Kachroo, P.; Szymczak, S.; Heinsen, F.A.; Forster, M.; Bethune, J.; Hemmrich-Stanisak, G.; Baker, L.; Schrappe, M.; Stanulla, M.; Franke, A. NGS-based methylation profiling differentiates TCF3-HLF and TCF3-PBX1 positive B-cell acute lymphoblastic leukemia. Epigenomics 2018, 10, 133–147. [Google Scholar] [CrossRef] [PubMed]
- Szabo, Q.; Bantignies, F.; Cavalli, G. Principles of genome folding into topologically associating domains. Sci. Adv. 2019, 5, eaaw1668. [Google Scholar] [CrossRef]
- Andersson, R.; Sandelin, A. Determinants of enhancer and promoter activities of regulatory elements. Nat. Rev. Genet. 2020, 21, 71–87. [Google Scholar] [CrossRef] [PubMed]
- Chen, H.; Xiao, J.; Shao, T.; Wang, L.; Bai, J.; Lin, X.; Ding, N.; Qu, Y.; Tian, Y.; Chen, X.; et al. Landscape of Enhancer-Enhancer Cooperative Regulation during Human Cardiac Commitment. Mol. Ther. Nucleic Acids 2019, 17, 840–851. [Google Scholar] [CrossRef]
- Chen, Q.; Li, C.; Gong, Z.; Chun Yong Chan, E.; Snyder, S.A.; Lam, S.H. Common deregulated gene expression profiles and morphological changes in developing zebrafish larvae exposed to environmental-relevant high to low concentrations of glucocorticoids. Chemosphere 2017, 172, 429–439. [Google Scholar] [CrossRef]
- Papizan, J.B.; Garry, G.A.; Brezprozvannaya, S.; McAnally, J.R.; Bassel-Duby, R.; Liu, N.; Olson, E.N. Deficiency in Kelch protein Klhl31 causes congenital myopathy in mice. J. Clin. Investig. 2017, 127, 3730–3740. [Google Scholar] [CrossRef]
- Wang, X.; Cairns, M.J.; Yan, J. Super-enhancers in transcriptional regulation and genome organization. Nucleic Acids Res. 2019, 47, 11481–11496. [Google Scholar] [CrossRef]
- Bell, E.; Curry, E.W.; Megchelenbrink, W.; Jouneau, L.; Brochard, V.; Tomaz, R.A.; Mau, K.H.T.; Atlasi, Y.; de Souza, R.A.; Marks, H.; et al. Dynamic CpG methylation delineates subregions within super-enhancers selectively decommissioned at the exit from naive pluripotency. Nat. Commun. 2020, 11, 1112. [Google Scholar] [CrossRef]
- Jeziorska, D.M.; Murray, R.J.S.; De Gobbi, M.; Gaentzsch, R.; Garrick, D.; Ayyub, H.; Chen, T.; Li, E.; Telenius, J.; Lynch, M.; et al. DNA methylation of intragenic CpG islands depends on their transcriptional activity during differentiation and disease. Proc. Natl. Acad. Sci. USA 2017, 114, E7526–E7535. [Google Scholar] [CrossRef]
- Ponnaluri, V.K.; Ehrlich, K.C.; Zhang, G.; Lacey, M.; Johnston, D.; Pradhan, S.; Ehrlich, M. Association of 5-hydroxymethylation and 5-methylation of DNA cytosine with tissue-specific gene expression. Epigenetics 2017, 12, 123–138. [Google Scholar] [CrossRef]
- Ehrlich, M. DNA hypermethylation in disease: Mechanisms and clinical relevance. Epigenetics 2019, 14, 1141–1163. [Google Scholar] [CrossRef]
- Paxton, C.W.; Cosgrove, R.A.; Drozd, A.C.; Wiggins, E.L.; Woodhouse, S.; Watson, R.A.; Spence, H.J.; Ozanne, B.W.; Pell, J.M. BTB-Kelch protein Krp1 regulates proliferation and differentiation of myoblasts. Am. J. Physiol. Cell Physiol. 2011, 300, C1345–C1355. [Google Scholar] [CrossRef]
- Xian, S.; Li, J.; Zhang, Z. miR-26b inhibits isoproterenol-induced cardiac fibrosis via the Keap1/Nrf2 signaling pathway. Exp. Ther. Med. 2020, 19, 2067–2074. [Google Scholar] [CrossRef]
- Zhou, Y.; Tang, X.; Niu, L.; Liu, Y.; Wang, B.; He, J. Ectodermal-neural cortex 1 as a novel biomarker predicts poor prognosis and induces metastasis in breast cancer by promoting Wnt/beta-catenin pathway. J. Cell Mol. Med. 2020. [Google Scholar] [CrossRef]
- Watanabe, K.; Yoshida, K.; Iwamoto, S. Kbtbd11 gene expression in adipose tissue increases in response to feeding and affects adipocyte differentiation. J. Diabetes Investig. 2019, 10, 925–932. [Google Scholar] [CrossRef]
- Li, W.; Pung, D.; Su, Z.Y.; Guo, Y.; Zhang, C.; Yang, A.Y.; Zheng, X.; Du, Z.Y.; Zhang, K.; Kong, A.N. Epigenetics Reactivation of Nrf2 in Prostate TRAMP C1 Cells by Curcumin Analogue FN1. Chem. Res. Toxicol. 2016, 29, 694–703. [Google Scholar] [CrossRef]
- Fabrizio, F.P.; Sparaneo, A.; Centra, F.; Trombetta, D.; Storlazzi, C.T.; Graziano, P.; Maiello, E.; Fazio, V.M.; Muscarella, L.A. Methylation Density Pattern of KEAP1 Gene in Lung Cancer Cell Lines Detected by Quantitative Methylation Specific PCR and Pyrosequencing. Int. J. Mol. Sci. 2019, 20, 2697. [Google Scholar] [CrossRef]
- Trapnell, C.; Roberts, A.; Goff, L.; Pertea, G.; Kim, D.; Kelley, D.R.; Pimentel, H.; Salzberg, S.L.; Rinn, J.L.; Pachter, L. Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat. Protoc. 2012, 7, 562–578. [Google Scholar] [CrossRef]
- Tsumagari, K.; Baribault, C.; Terragni, J.; Varley, K.E.; Gertz, J.; Pradhan, S.; Badoo, M.; Crain, C.M.; Song, L.; Crawford, G.E.; et al. Early de novo DNA methylation and prolonged demethylation in the muscle lineage. Epigenetics 2013, 8, 317–332. [Google Scholar] [CrossRef]
- Chen, P.H.; Smith, T.J.; Wu, J.; Siesser, P.F.; Bisnett, B.J.; Khan, F.; Hogue, M.; Soderblom, E.; Tang, F.; Marks, J.R.; et al. Glycosylation of KEAP1 links nutrient sensing to redox stress signaling. EMBO J. 2017, 36, 2233–2250. [Google Scholar] [CrossRef]
- Song, Q.; Decato, B.; Hong, E.E.; Zhou, M.; Fang, F.; Qu, J.; Garvin, T.; Kessler, M.; Zhou, J.; Smith, A.D. A reference methylome database and analysis pipeline to facilitate integrative and comparative epigenomics. PLoS ONE 2013, 8, e81148. [Google Scholar] [CrossRef]
- Krietenstein, N.; Abraham, S.; Venev, S.V.; Abdennur, N.; Gibcus, J.; Hsieh, T.S.; Parsi, K.M.; Yang, L.; Maehr, R.; Mirny, L.A.; et al. Ultrastructural Details of Mammalian Chromosome Architecture. Mol. Cell 2020, 78, 554–565.e557. [Google Scholar] [CrossRef] [PubMed]
- Papadopoulos, J.S.; Agarwala, R. COBALT: Constraint-based alignment tool for multiple protein sequences. Bioinformatics 2007, 23, 1073–1079. [Google Scholar] [CrossRef]


| Description | TPM in SkM b | TPM Ratio: SkM to Median of Other Tissues | FPKM in Myoblasts | FPKM Ratio: Myoblasts to Median of Heterologous Cell Cultures c | FPKM Ratio: Myotubes to Myoblasts |
|---|---|---|---|---|---|
| KBTBD12 (Figure S4) | 15 | 13 | 0.3 | 0 | 2.8 |
| KBTBD13d (Figure S6) | 1.5 | 25 | 0 | 0 | 0 |
| KEAP1 (Figure 3) | 88 | 2.4 | 32 | 0.7 | 1.0 |
| KLHL21 (Figure S10) | 84 | 3.3 | 31 | 0.6 | 1.4 |
| KLHL30 (Figure 2) | 110 | 145 | 16 | 18.0 | 2.8 |
| KLHL31 (Figure S2) | 18 | 59 | 5.6 | 5.8 | 872 |
| KLHL33 (Figure S1) | 11 | 24 | 0 | 0 | 0 |
| KLHL34 (Figure S3) | 8.5 | 113 | 0 | 0 | 0 |
| KLHL38 (Figure 2) | 90 | 405 | 0.1 | 0 | 19 |
| KLHL40 (Figure 1; Figure S5) | 303 | 3373 | 7.7 | #DIV/0! | 17 |
| KLHL41 (Figure 1) | 3420 | 1946 | 104 | 1613 | 254 |
| Description | Brain Tissue (Median TPM) | Ratio of Brain Tissue TPM to Median TPM of Non-Brain Tissues b | ||||
|---|---|---|---|---|---|---|
| Cortex c | Hippoc. c | Cerebel. c | Cortex | Hippoc. | Cerebel. | |
| ENC1 (Figure 3) | 322 | 87 | 4.3 | 56 | 15 | 0.7 |
| KBTBD11 (Figure 4) | 59 | 27 | 46 | 17 | 7.2 | 12 |
| KLHL2 (Figure S9) | 47 | 47 | 21 | 3.2 | 45 | 39 |
| KLHL32 (Figure 4) | 10 | 12 | 11 | 37 | 3.9 | 6.0 |
| KLHL35d (Figure S9) | 3.3 | 1.6 | 2.5 | 8.0 | 13 | 2.9 |
| KLHL4 (Figure S9) | 3.6 | 4.5 | 1.0 | 11 | 15 | 0.7 |
| Epigenetic Profile Specifically Associated with SkM or Brain | Genes Preferentially Expressed in SkM (Some Also in Heart) | Genes Preferentially Expressed in Brain |
|---|---|---|
| Promoter chromatin upstream of TSS b | KLHL31, 33, 34 | None |
| Promoter chromatin dnstrm of TSS b | KLHL30, 31, 33, 38, 41 | KLHL32, 35, KBTBD11 |
| Broadening of a constitutively unmethylated region at the TSS | KLHL21, 33, 34, 40; KBTBD12 | KLHL2, 4, 32 (neurons c); ENC1, KBTBD11 (neurons/fetal brain) |
| DNA hypomethylation upstream of TSS b | KLHL30, 31, 33, 34, 38, 40, KBTBD13 | KLHL32 (neurons) |
| DNA hypomethylation dnstrm of TSS b | KLHL21, 30, 31, 33, 41; KBTBD12 | KLHL2, 4, 32 (neurons) |
| Super-enhancer | KLHL 21, 30, 31, 38, 40, 41 | ENC1 |
| Intragenic enhancer chromatin | KLHL21, 30, 31, 32, 33, 34, 38, 40, 41; KEAP1; KBTBD12, 13 | KLHL2, 35; KBTBD11, ENC1 |
| Gene-upstream enhancer chromatin | KLHL21, 30, 31, 33, 34, 38, 40, 41 | KLHL32, KBTBD11, ENC1 (neurons/fetal brain) |
| Gene-downstream enhancer chromatin | KLHL21, 38, KBTBD12 | KBTBD11, ENC1 (fetal brain) |
| DNA hypomethylation in enhancer chromatin | KLHL21, 30, 31, 33, 34, 38, 40, 41, KBTBD12, 13 | KLHL2 (hippocampus, anterior caudate) |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ehrlich, K.C.; Baribault, C.; Ehrlich, M. Epigenetics of Muscle- and Brain-Specific Expression of KLHL Family Genes. Int. J. Mol. Sci. 2020, 21, 8394. https://doi.org/10.3390/ijms21218394
Ehrlich KC, Baribault C, Ehrlich M. Epigenetics of Muscle- and Brain-Specific Expression of KLHL Family Genes. International Journal of Molecular Sciences. 2020; 21(21):8394. https://doi.org/10.3390/ijms21218394
Chicago/Turabian StyleEhrlich, Kenneth C., Carl Baribault, and Melanie Ehrlich. 2020. "Epigenetics of Muscle- and Brain-Specific Expression of KLHL Family Genes" International Journal of Molecular Sciences 21, no. 21: 8394. https://doi.org/10.3390/ijms21218394
APA StyleEhrlich, K. C., Baribault, C., & Ehrlich, M. (2020). Epigenetics of Muscle- and Brain-Specific Expression of KLHL Family Genes. International Journal of Molecular Sciences, 21(21), 8394. https://doi.org/10.3390/ijms21218394