Differential Defense Responses of Upland and Lowland Switchgrass Cultivars to a Cereal Aphid Pest
Abstract
1. Introduction
2. Results
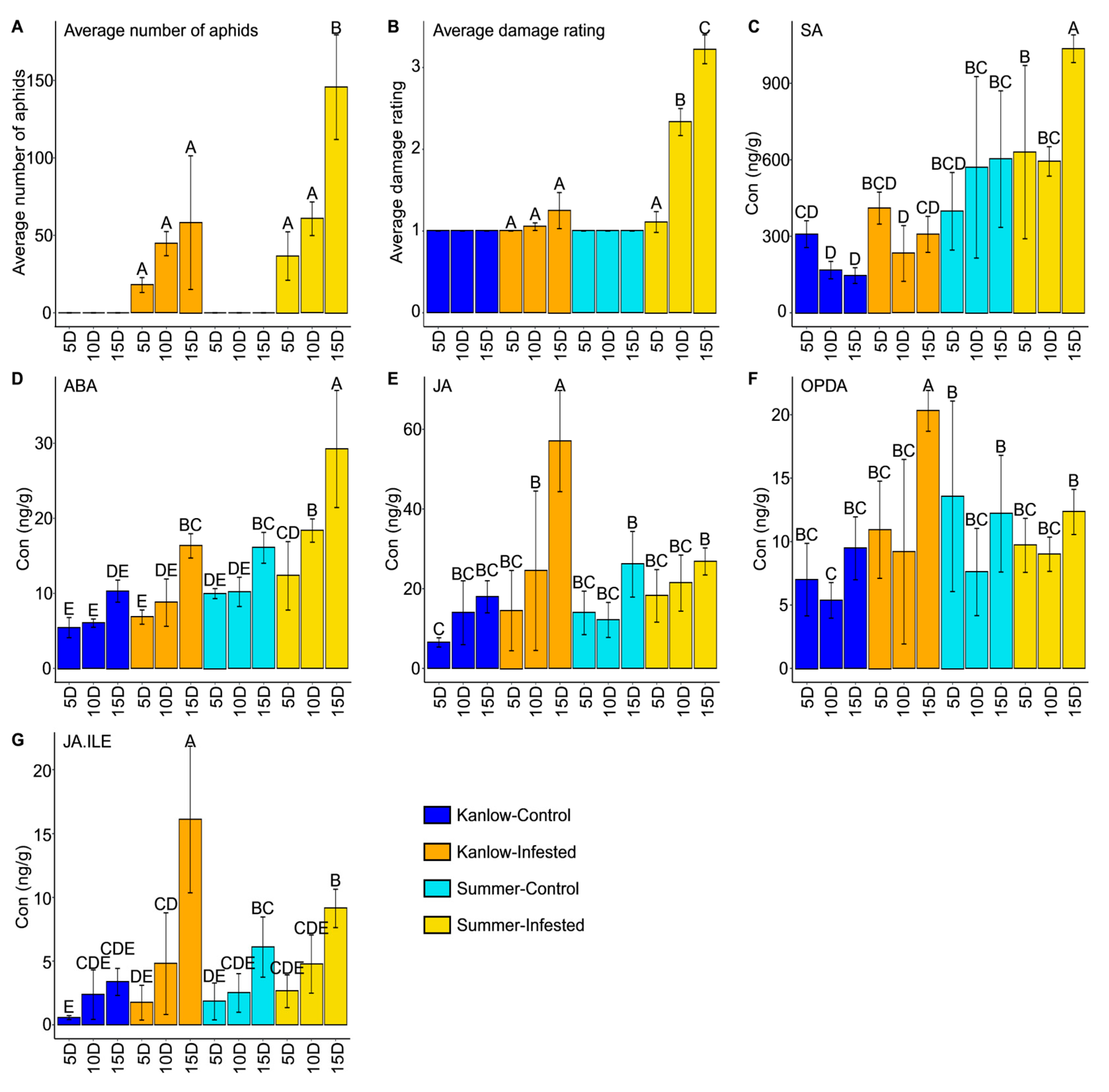
2.1. YSA Colonized Both Cultivars with Different Damage Outcomes
2.2. Hormones and Metabolites Were Affected by YSA Infestation
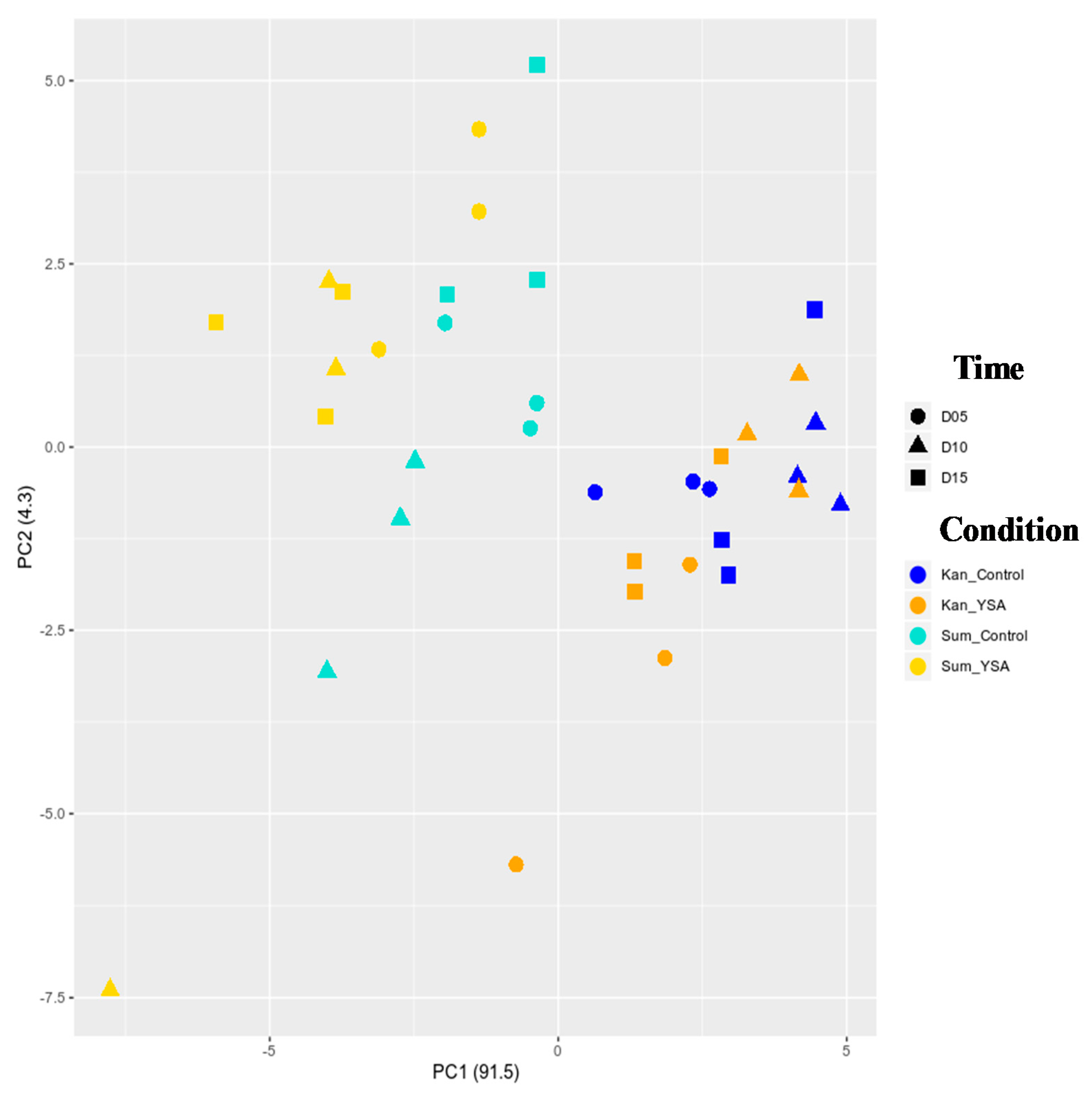
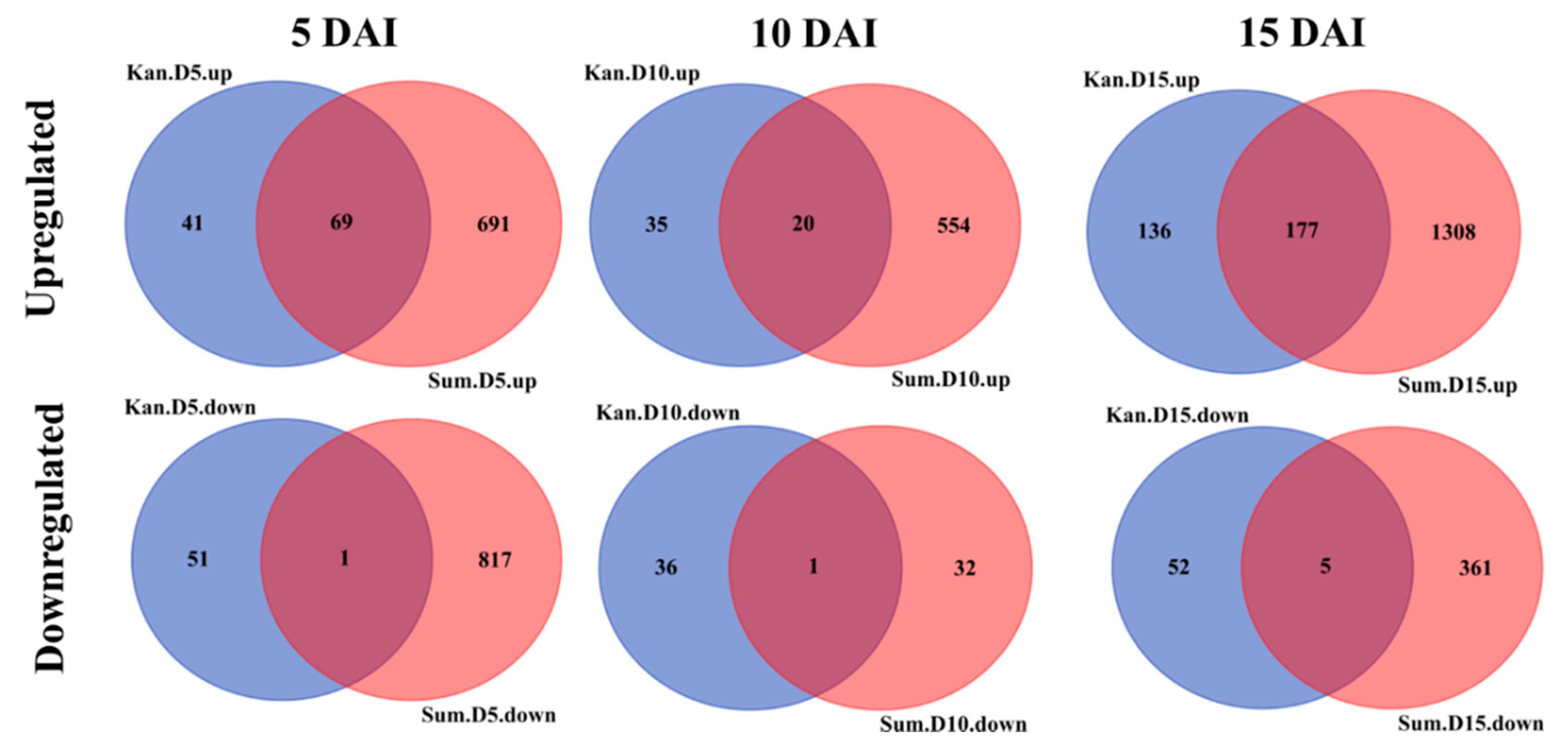
2.3. Transcriptomic Responses to YSA Infestation
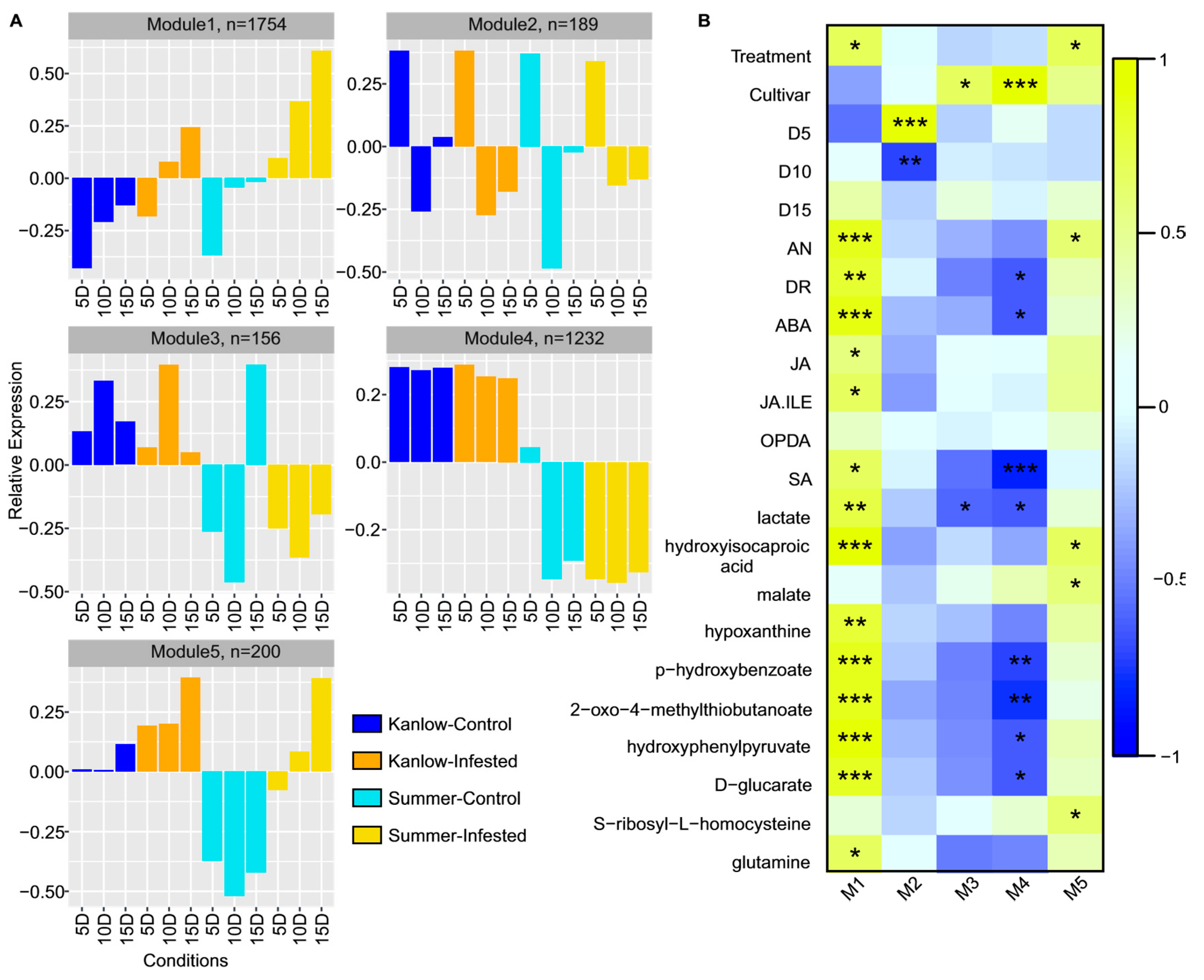
2.4. Temporal Gene Expression Varied in Response to YSA Infestation
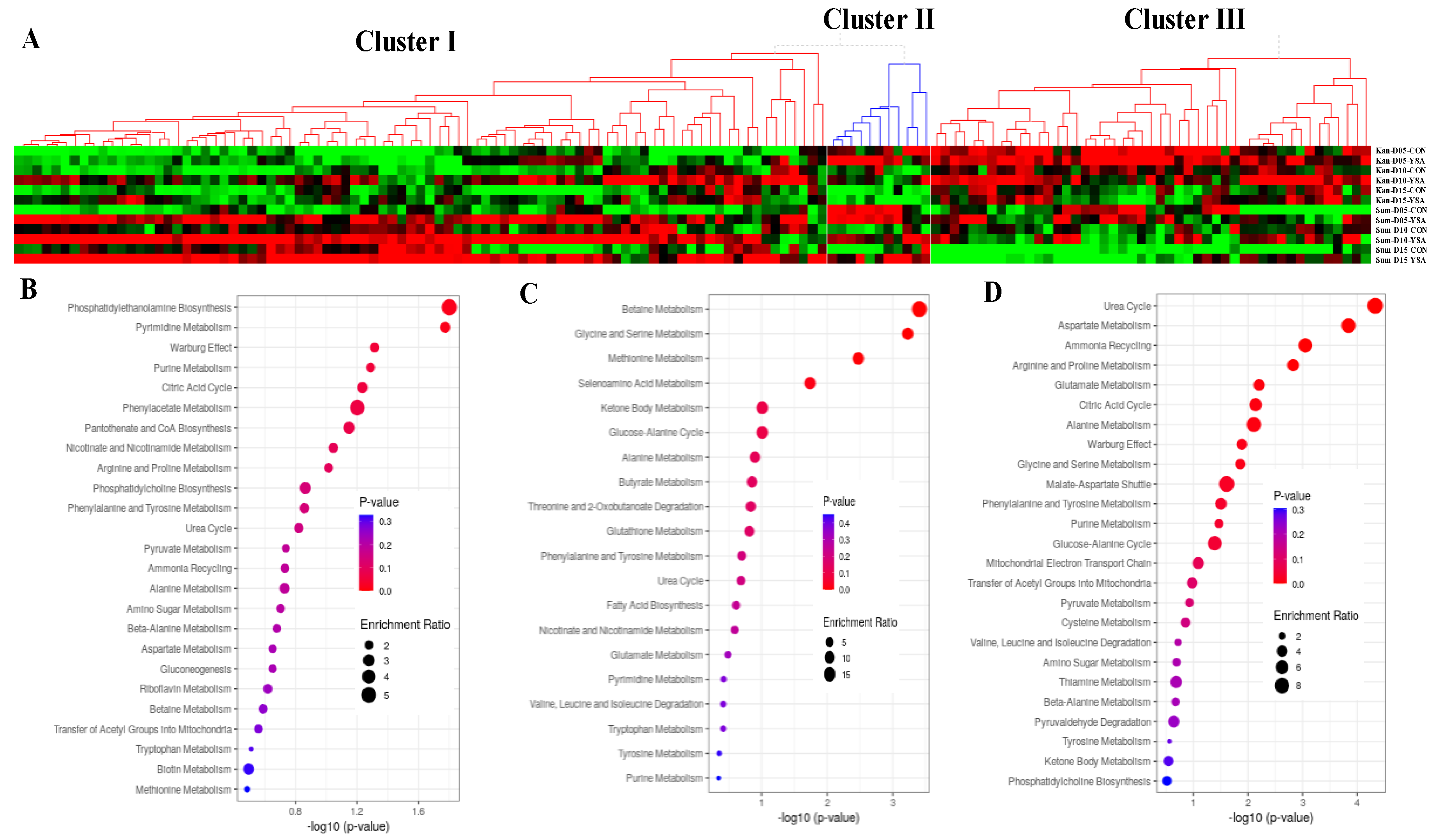
2.5. Functional Mechanisms in Response to YSA Infestation
2.6. Cultivar Specific Functional Mechanisms in Response to YSA Infestation
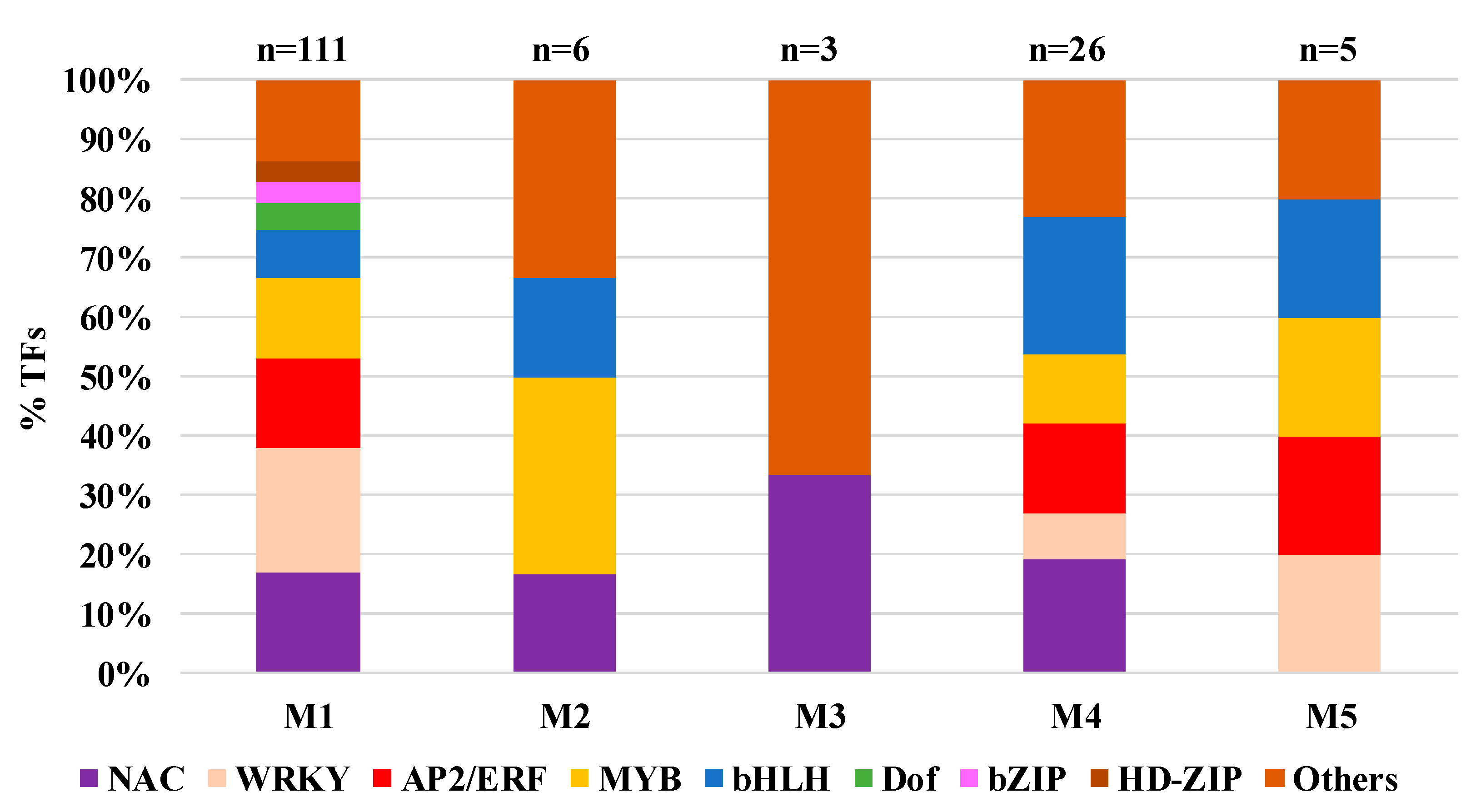
2.7. Transcription Factors’ Role in Transcriptome Response to YSA Infestation
3. Discussion
3.1. Commonalities in Defense Response between Both Cultivars
3.2. Rate in the Change of Plant Damage and Gene Expression/Metabolomics Variation over the Time
3.3. Evidence for Differential Priming of Aphid Defense Mechanisms
4. Materials and Methods
4.1. Plant Growth Conditions and Treatments
4.2. RNA Extraction and Sequencing
4.3. RNA-Seq Analysis
4.4. Gene Ontology (GO) Analysis
4.5. Plant Hormone and Metabolite Analyses
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Casler, M.D.; Vogel, K.P.; Lee, D.K.; Mitchell, R.B.; Adler, P.R.; Sulc, R.M.; Johnson, K.D.; Kallenbach, R.L.; Boe, A.R.; Mathison, R.D.; et al. 30 years of progress toward increased biomass yield of switchgrass and big bluestem. Crop Sci. 2018, 58, 1242–1254. [Google Scholar] [CrossRef]
- Evans, J.; Sanciangco, M.D.; Lau, K.H.; Crisovan, E.; Barry, K.; Daum, C.; Hundley, H.; Jenkins, J.; Kennedy, M.; Kunde-Ramamoorthy, G.; et al. Extensive genetic diversity is present within North American switchgrass germplasm. Plant Genome 2018, 11, 170055. [Google Scholar] [CrossRef] [PubMed]
- Stewart, C.L.; Pyle, J.D.; Jochum, C.C.; Vogel, K.P.; Yuen, G.Y.; Scholthof, K.-B.G. Multi-year pathogen survey of biofuel switchgrass breeding plots reveals high prevalence of infections by panicum mosaic virus and its satellite virus. Phytopathology 2015, 105, 1146–1154. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Koch, K.; Donze-Reiner, T.; Baird, L.; Louis, J.; Amundsen, K.; Sarath, G.; Bradshaw, J.; Heng-Moss, T. Evaluation of greenbug and yellow sugarcane aphid feeding behavior on resistant and susceptible switchgrass cultivars. Bioenergy Res. 2018, 11, 480–490. [Google Scholar] [CrossRef]
- Muchlinski, A.; Chen, X.; Lovell, J.T.; Köllner, T.G.; Pelot, K.A.; Zerbe, P.; Ruggiero, M.; Callaway, L.I.; Laliberte, S.; Chen, F.; et al. Biosynthesis and emission of stress-induced volatile terpenes in roots and leaves of switchgrass (Panicum virgatum L.). Front. Plant Sci. 2019, 10. [Google Scholar] [CrossRef]
- Poudel, H.P.; Sanciangco, M.D.; Kaeppler, S.M.; Buell, C.R.; Casler, M.D. Quantitative trait loci for freezing tolerance in a lowland x upland switchgrass population. Front. Plant Sci. 2019, 10. [Google Scholar] [CrossRef]
- VanWallendael, A.; Bonnette, J.; Juenger, T.E.; Fritschi, F.B.; Fay, P.A.; Mitchell, R.B.; Lloyd-Reilley, J.; Rouquette, F.M.; Bergstrom, G.C.; Lowry, D.B. Geographic variation in the genetic basis of resistance to leaf rust between locally adapted ecotypes of the biofuel crop switchgrass (Panicum virgatum). New Phytol. 2020, 227, 1696–1708. [Google Scholar] [CrossRef]
- Koch, K.G.; Fithian, R.; Heng-Moss, T.M.; Bradshaw, J.D.; Sarath, G.; Spilker, C. Evaluation of tetraploid switchgrass (Poales: Poaceae) populations for host suitability and differential resistance to four cereal aphids. J. Econ. Entomol. 2014, 107, 424–431. [Google Scholar] [CrossRef][Green Version]
- Palmer, N.A.; Basu, S.; Heng-Moss, T.; Bradshaw, J.D.; Sarath, G.; Louis, J. Fall armyworm (Spodoptera frugiperda Smith) feeding elicits differential defense responses in upland and lowland switchgrass. PLoS ONE 2019, 14, e0218352. [Google Scholar] [CrossRef]
- Schuh, M.K.; Bahlai, C.A.; Malmstrom, C.M.; Landis, D.A. Effect of switchgrass ecotype and cultivar on establishment, feeding, and development of fall armyworm (Lepidoptera: Noctuidae). J. Econ. Entomol. 2019, 112, 440–449. [Google Scholar] [CrossRef]
- Koch, K.G.; Bradshaw, J.D.; Heng-Moss, T.M.; Sarath, G. Categories of resistance to greenbug and yellow sugarcane aphid (Hemiptera: Aphididae) in three tetraploid switchgrass populations. Bioenergy Res. 2014, 7, 909–918. [Google Scholar] [CrossRef]
- Alderson, J.; Sharp, W.C.; States, U. Grass Varieties in the United States; U.S. Department of Agriculture: Washington, DC, USA, 1994; pp. 1–308.
- Sarath, G.; Baird, L.M.; Mitchell, R.B. Senescence, dormancy and tillering in perennial C4 grasses. Plant Sci. 2014, 217–218, 140–151. [Google Scholar] [CrossRef]
- Betsiashvili, M.; Ahern, K.R.; Jander, G. Additive effects of two quantitative trait loci that confer Rhopalosiphum maidis (corn leaf aphid) resistance in maize inbred line Mo17. J. Exp. Bot. 2015, 66, 571–578. [Google Scholar] [CrossRef]
- Boissot, N.; Thomas, S.; Chovelon, V.; Lecoq, H. NBS-LRR-mediated resistance triggered by aphids: Viruses do not adapt; aphids adapt via different mechanisms. BMC Plant Biol. 2016, 16, 25. [Google Scholar] [CrossRef] [PubMed]
- Züst, T.; Agrawal, A.A. Mechanisms and evolution of plant resistance to aphids. Nat. Plants 2016, 2, 15206. [Google Scholar] [CrossRef]
- Nalam, V.; Louis, J.; Shah, J. Plant defense against aphids, the pest extraordinaire. Plant Sci. 2019, 279, 96–107. [Google Scholar] [CrossRef] [PubMed]
- Varsani, S.; Grover, S.; Zhou, S.; Koch, K.G.; Huang, P.-C.; Kolomiets, M.V.; Williams, W.P.; Heng-Moss, T.; Sarath, G.; Luthe, D.S.; et al. 12-Oxo-phytodienoic acid acts as a regulator of maize defense against corn leaf aphid. Plant Physiol. 2019, 179, 1402–1415. [Google Scholar] [CrossRef] [PubMed]
- Tzin, V.; Fernandez-Pozo, N.; Richter, A.; Schmelz, E.A.; Schoettner, M.; Schäfer, M.; Ahern, K.R.; Meihls, L.N.; Kaur, H.; Huffaker, A.; et al. Dynamic maize responses to aphid feeding are revealed by a time series of transcriptomic and metabolomic assays. Plant Physiol. 2015, 169, 1727–1743. [Google Scholar] [CrossRef] [PubMed]
- Foyer, C.H.; Rasool, B.; Davey, J.W.; Hancock, R.D. Cross-tolerance to biotic and abiotic stresses in plants: A focus on resistance to aphid infestation. J. Exp. Bot. 2016, 67, 2025–2037. [Google Scholar] [CrossRef]
- Koch, K.G.; Chapman, K.; Louis, J.; Heng-Moss, T.; Sarath, G. Plant Tolerance: A unique approach to control hemipteran pests. Front. Plant Sci. 2016, 7. [Google Scholar] [CrossRef]
- Shoala, T.; Edwards, M.G.; Knight, M.R.; Gatehouse, A.M.R. OXI1 kinase plays a key role in resistance of Arabidopsis towards aphids (Myzus persicae). Transgenic Res. 2018, 27, 355–366. [Google Scholar] [CrossRef] [PubMed]
- Erb, M.; Reymond, P. Molecular interactions between plants and insect herbivores. Annu. Rev. Plant Biol. 2019, 70, 527–557. [Google Scholar] [CrossRef]
- Xiao, L.; Carrillo, J.; Siemann, E.; Ding, J. Herbivore-specific induction of indirect and direct defensive responses in leaves and roots. AoB Plants 2019, 11. [Google Scholar] [CrossRef]
- Painter, R. Insect Resistance in Crop Plants; Macmillan: New York, NY, USA, 1951. [Google Scholar]
- Smith, C.M.; Boyko, E.V. The molecular bases of plant resistance and defense responses to aphid feeding: Current status. Entomol. Exp. Appl. 2007, 122, 1–16. [Google Scholar] [CrossRef]
- Smith, C.M.; Chuang, W.-P. Plant resistance to aphid feeding: Behavioral, physiological, genetic and molecular cues regulate aphid host selection and feeding. Pest Manag. Sci. 2014, 70, 528–540. [Google Scholar] [CrossRef]
- Gutsche, A.; Heng-Moss, T.; Sarath, G.; Twigg, P.; Xia, Y.; Lu, G.; Mornhinweg, D. Gene expression profiling of tolerant barley in response to Diuraphis noxia (Hemiptera: Aphididae) feeding. Bull. Entomol. Res. 2009, 99, 163–173. [Google Scholar] [CrossRef]
- Donze-Reiner, T.; Palmer, N.A.; Scully, E.D.; Prochaska, T.J.; Koch, K.G.; Heng-Moss, T.; Bradshaw, J.D.; Twigg, P.; Amundsen, K.; Sattler, S.E.; et al. Transcriptional analysis of defense mechanisms in upland tetraploid switchgrass to greenbugs. BMC Plant Biol. 2017, 17, 46. [Google Scholar] [CrossRef]
- Koch, K.G.; Palmer, N.A.; Donze-Reiner, T.; Scully, E.D.; Seravalli, J.; Amundsen, K.; Twigg, P.; Louis, J.; Bradshaw, J.D.; Heng-Moss, T.M.; et al. Aphid-responsive defense networks in hybrid switchgrass. Front. Plant Sci. 2020, 11. [Google Scholar] [CrossRef]
- Martinez-Reyna, J.M.; Vogel, K.P. Heterosis in switchgrass: Spaced plants. Crop Sci. 2008, 48, 1312–1320. [Google Scholar] [CrossRef]
- War, A.R.; Paulraj, M.G.; Ahmad, T.; Buhroo, A.A.; Hussain, B.; Ignacimuthu, S.; Sharma, H.C. Mechanisms of plant defense against insect herbivores. Plant Signal. Behav. 2012, 7, 1306–1320. [Google Scholar] [CrossRef] [PubMed]
- Ma, F.; Yang, X.; Shi, Z.; Miao, X. Novel crosstalk between ethylene- and jasmonic acid-pathway responses to a piercing-sucking insect in rice. New Phytol. 2020, 225, 474–487. [Google Scholar] [CrossRef]
- Hanley, M.E.; Lamont, B.B.; Fairbanks, M.M.; Rafferty, C.M. Plant structural traits and their role in anti-herbivore defence. Perspect. Plant Ecol. Evol. Syst. 2007, 8, 157–178. [Google Scholar] [CrossRef]
- Dao, T.T.H.; Linthorst, H.J.M.; Verpoorte, R. Chalcone synthase and its functions in plant resistance. Phytochem. Rev. 2011, 10, 397–412. [Google Scholar] [CrossRef] [PubMed]
- Le Hir, R.; Bellini, C. The plant-specific Dof transcription factors family: New players involved in vascular system development and functioning in Arabidopsis. Front. Plant Sci. 2013, 4. [Google Scholar] [CrossRef] [PubMed]
- Dombrecht, B.; Xue, G.P.; Sprague, S.J.; Kirkegaard, J.A.; Ross, J.J.; Reid, J.B.; Fitt, G.P.; Sewelam, N.; Schenk, P.M.; Manners, J.M.; et al. MYC2 differentially modulates diverse jasmonate-dependent functions in Arabidopsis. Plant Cell 2007, 19, 2225–2245. [Google Scholar] [CrossRef] [PubMed]
- Kazan, K.; Manners, J.M. MYC2: The master in action. Mol. Plant 2013, 6, 686–703. [Google Scholar] [CrossRef] [PubMed]
- Niu, Y.; Figueroa, P.; Browse, J. Characterization of JAZ-interacting bHLH transcription factors that regulate jasmonate responses in Arabidopsis. J. Exp. Bot. 2011, 62, 2143–2154. [Google Scholar] [CrossRef]
- Fernández-Calvo, P.; Chini, A.; Fernández-Barbero, G.; Chico, J.-M.; Gimenez-Ibanez, S.; Geerinck, J.; Eeckhout, D.; Schweizer, F.; Godoy, M.; Franco-Zorrilla, J.M.; et al. The Arabidopsis bHLH transcription factors MYC3 and MYC4 are targets of JAZ repressors and act additively with MYC2 in the activation of jasmonate responses. Plant Cell 2011, 23, 701–715. [Google Scholar] [CrossRef]
- Lorenzo, O.; Chico, J.M.; Sánchez-Serrano, J.J.; Solano, R. JASMONATE-INSENSITIVE1 encodes a MYC transcription factor essential to discriminate between different jasmonate-regulated defense responses in Arabidopsis. Plant Cell 2004, 16, 1938–1950. [Google Scholar] [CrossRef]
- Verhage, A.; Vlaardingerbroek, I.; Raaymakers, C.; Van Dam, N.M.; Dicke, M.; Van Wees, S.C.M.; Pieterse, C.M.J. Rewiring of the jasmonate signaling pathway in Arabidopsis during insect herbivory. Front. Plant Sci. 2011, 2, 47. [Google Scholar] [CrossRef]
- Huibers, R.P.; de Jong, M.; Dekter, R.W.; Van den Ackerveken, G. Disease-specific expression of host genes during downy mildew infection of Arabidopsis. Mol. Plant Microbe Interact. 2009, 22, 1104–1115. [Google Scholar] [CrossRef] [PubMed]
- Gao, Q.-M.; Venugopal, S.; Navarre, D.; Kachroo, A. Low oleic acid-derived repression of jasmonic acid-inducible defense responses requires the WRKY50 and WRKY51 Proteins. Plant Physiol. 2011, 155, 464–476. [Google Scholar] [CrossRef]
- Zheng, Z.; Qamar, S.A.; Chen, Z.; Mengiste, T. Arabidopsis WRKY33 transcription factor is required for resistance to necrotrophic fungal pathogens. Plant J. 2006, 48. [Google Scholar] [CrossRef] [PubMed]
- Yue, R.; Lu, C.; Sun, T.; Peng, T.; Han, X.; Qi, J.; Yan, S.; Tie, S. Identification and expression profiling analysis of calmodulin-binding transcription activator genes in maize (Zea mays L.) under abiotic and biotic stresses. Front. Plant Sci. 2015, 6. [Google Scholar] [CrossRef]
- Kim, Y.S.; An, C.; Park, S.; Gilmour, S.J.; Wang, L.; Renna, L.; Brandizzi, F.; Grumet, R.; Thomashow, M.F. CAMTA-mediated regulation of salicylic acid immunity pathway genes in Arabidopsis exposed to low temperature and pathogen infection. Plant Cell 2017, 29, 2465–2477. [Google Scholar] [CrossRef] [PubMed]
- Wang, W.; Tang, W.; Ma, T.; Niu, D.; Jin, J.B.; Wang, H.; Lin, R. A pair of light signaling factors FHY3 and FAR1 regulates plant immunity by modulating chlorophyll biosynthesis. J. Integr. Plant Biol. 2016, 58, 91–103. [Google Scholar] [CrossRef]
- Kiani, M.; Szczepaniec, A. Effects of sugarcane aphid herbivory on transcriptional responses of resistant and susceptible sorghum. BMC Genom. 2018, 19, 774. [Google Scholar] [CrossRef]
- Niu, L.; Pan, L.; Zeng, W.; Lu, Z.; Cui, G.; Fan, M.; Xu, Q.; Wang, Z.; Li, G. Dynamic transcriptomes of resistant and susceptible peach lines after infestation by green peach aphids (Myzus persicae Sülzer) reveal defence responses controlled by the Rm3 locus. BMC Genom. 2018, 19. [Google Scholar] [CrossRef]
- Wang, Y.; Zeng, X.; Iyer, N.J.; Bryant, D.W.; Mockler, T.C.; Mahalingam, R. Exploring the switchgrass transcriptome using second-generation sequencing technology. PLoS ONE 2012, 7, e34225. [Google Scholar] [CrossRef]
- Vogel, K.P.; Mitchell, R.B.; Casler, M.D.; Sarath, G. Registration of ‘Liberty’ switchgrass. J. Plant Regist. 2014, 8, 242–247. [Google Scholar] [CrossRef]
- Edmé, S.; Mitchell, R.; Sarath, G. Genetic parameters and prediction of breeding values in switchgrass bred for bioenergy. Crop Sci. 2017, 57, 1464–1474. [Google Scholar] [CrossRef]
- Fiedler, J.D.; Lanzatella, C.; Edmé, S.J.; Palmer, N.A.; Sarath, G.; Mitchell, R.; Tobias, C.M. Genomic prediction accuracy for switchgrass traits related to bioenergy within differentiated populations. BMC Plant Biol. 2018, 18, 142. [Google Scholar] [CrossRef] [PubMed]
- Silva-Sanzana, C.; Celiz-Balboa, J.; Garzo, E.; Marcus, S.E.; Parra-Rojas, J.P.; Rojas, B.; Olmedo, P.; Rubilar, M.A.; Rios, I.; Chorbadjian, R.A.; et al. Pectin methylesterases modulate plant homogalacturonan status in defenses against the aphid Myzus persicae. Plant Cell 2019, 31, 1913–1929. [Google Scholar] [CrossRef]
- Zhang, Y.; Fu, Y.; Fan, J.; Li, Q.; Francis, F.; Chen, J. Comparative transcriptome and histological analyses of wheat in response to phytotoxic aphid Schizaphis graminum and non-phytotoxic aphid Sitobion avenae feeding. BMC Plant Biol. 2019, 19, 547. [Google Scholar] [CrossRef]
- Batyrshina, Z.S.; Yaakov, B.; Shavit, R.; Singh, A.; Tzin, V. Comparative transcriptomic and metabolic analysis of wild and domesticated wheat genotypes reveals differences in chemical and physical defense responses against aphids. BMC Plant Biol. 2020, 20, 19. [Google Scholar] [CrossRef] [PubMed]
- de Jesus Correa, L.; Maciel, O.V.B.; Bücker-Neto, L.; Pilati, L.; Morozini, A.M.; Faria, M.V.; Da-Silva, P.R. A Comprehensive analysis of wheat resistance to Rhopalosiphum padi (Hemiptera: Aphididae) in Brazilian wheat cultivars. J. Econ. Entomol. 2020, 113, 1493–1503. [Google Scholar] [CrossRef]
- Escudero-Martinez, C.; Rodriguez, P.A.; Liu, S.; Santos, P.A.; Stephens, J.; Bos, J.I.B. An aphid effector promotes barley susceptibility through suppression of defence gene expression. J. Exp. Bot. 2020, 71, 2796–2807. [Google Scholar] [CrossRef]
- Yates-Stewart, A.D.; Pekarcik, A.; Michel, A.; Blakeslee, J.J. Jasmonic acid-Isoleucine (JA-Ile) is involved in the host-plant resistance mechanism against the soybean aphid (Hemiptera: Aphididae). J. Econ. Entomol. 2020. [Google Scholar] [CrossRef]
- Zhang, Y.; Bouwmeester, H.J.; Kappers, I.F. Combined transcriptome and metabolome analysis identifies defence responses in spider mite-infested pepper (Capsicum annuum). J. Exp. Bot. 2020, 71, 330–343. [Google Scholar] [CrossRef]
- Kloth, K.J.; Kormelink, R. Defenses against Virus and Vector: A phloem-biological perspective on RTM- and SLI1-mediated resistance to potyviruses and aphids. Viruses 2020, 12, 129. [Google Scholar] [CrossRef]
- Mitchell, C.; Brennan, R.M.; Graham, J.; Karley, A.J. Plant defense against herbivorous pests: Exploiting resistance and tolerance traits for sustainable crop protection. Front. Plant Sci. 2016, 7. [Google Scholar] [CrossRef] [PubMed]
- War, A.R.; Taggar, G.K.; Hussain, B.; Taggar, M.S.; Nair, R.M.; Sharma, H.C. Plant defence against herbivory and insect adaptations. AoB Plants 2018, 10. [Google Scholar] [CrossRef]
- Zogli, P.; Pingault, L.; Grover, S.; Louis, J. Ento(o)mics: The intersection of ‘omic’ approaches to decipher plant defense against sap-sucking insect pests. Curr. Opin. Plant Biol. 2020, 56, 153–161. [Google Scholar] [CrossRef]
- Jaouannet, M.; Morris, J.A.; Hedley, P.E.; Bos, J.I.B. Characterization of Arabidopsis transcriptional responses to different aphid species reveals genes that contribute to host susceptibility and non-host resistance. PLoS Pathog. 2015, 11, e1004918. [Google Scholar] [CrossRef] [PubMed]
- Bodenhausen, N.; Reymond, P. Signaling pathways controlling induced resistance to insect herbivores in Arabidopsis. Mol. Plant Microbe Interact. 2007, 20, 1406–1420. [Google Scholar] [CrossRef] [PubMed]
- Rojas, C.M.; Senthil-Kumar, M.; Tzin, V.; Mysore, K. Regulation of primary plant metabolism during plant-pathogen interactions and its contribution to plant defense. Front. Plant Sci. 2014, 5. [Google Scholar] [CrossRef] [PubMed]
- Isah, T. Stress and defense responses in plant secondary metabolites production. Biol. Res. 2019, 52. [Google Scholar] [CrossRef]
- Shinya, T.; Hojo, Y.; Desaki, Y.; Christeller, J.T.; Okada, K.; Shibuya, N.; Galis, I. Modulation of plant defense responses to herbivores by simultaneous recognition of different herbivore-associated elicitors in rice. Sci. Rep. 2016, 6, 32537. [Google Scholar] [CrossRef]
- Bansal, R.; Mian, M.; Mittapalli, O.; Michel, A.P. RNA-Seq reveals a xenobiotic stress response in the soybean aphid, Aphis glycines, when fed aphid-resistant soybean. BMC Genom. 2014, 15, 972. [Google Scholar] [CrossRef]
- Hohenstein, J.D.; Studham, M.E.; Klein, A.; Kovinich, N.; Barry, K.; Lee, Y.-J.; MacIntosh, G.C. Transcriptional and chemical changes in soybean leaves in response to long-term aphid colonization. Front. Plant Sci. 2019, 10. [Google Scholar] [CrossRef]
- Silva-Sanzana, C.; Estevez, J.M.; Blanco-Herrera, F. Influence of cell wall polymers and their modifying enzymes during plant–aphid interactions. J. Exp. Bot. 2020, 71, 3854–3864. [Google Scholar] [CrossRef] [PubMed]
- Ben-Issa, R.; Gomez, L.; Gautier, H. Companion plants for aphid pest management. Insects 2017, 8, 112. [Google Scholar] [CrossRef]
- Hori, M. Repellency of rosemary oil against Myzus persicae in a laboratory and in a screenhouse. J. Chem. Ecol. 1998, 24, 1425–1432. [Google Scholar] [CrossRef]
- Ahn, I.-P.; Kim, S.; Lee, Y.-H. Vitamin B1 functions as an activator of plant disease resistance. Plant Physiol. 2005, 138, 1505–1515. [Google Scholar] [CrossRef]
- Hanson, A.D.; Beaudoin, G.A.; McCarty, D.R.; Gregory, J.F. Does abiotic stress cause functional B Vitamin deficiency in plants? Plant Physiol. 2016, 172, 2082–2097. [Google Scholar] [CrossRef] [PubMed]
- Rapala-Kozik, M.; Wolak, N.; Kujda, M.; Banas, A.K. The upregulation of thiamine (vitamin B1) biosynthesis in Arabidopsis thaliana seedlings under salt and osmotic stress conditions is mediated by abscisic acid at the early stages of this stress response. BMC Plant Biol. 2012, 12, 2. [Google Scholar] [CrossRef]
- Shi, H.; Xiong, L.; Stevenson, B.; Lu, T.; Zhu, J.-K. The Arabidopsis salt overly sensitive 4 mutants uncover a critical role for vitamin B6 in plant salt tolerance. Plant Cell 2002, 14, 575–588. [Google Scholar] [CrossRef]
- Creelman, R.A.; Tierney, M.L.; Mullet, J.E. Jasmonic acid/methyl jasmonate accumulate in wounded soybean hypocotyls and modulate wound gene expression. Proc. Natl. Acad. Sci. USA 1992, 89, 4938–4941. [Google Scholar] [CrossRef]
- Gonzáles, W.L.; Ramírez, C.C.; Olea, N.; Niemeyer, H.M. Host plant changes produced by the aphid Sipha flava: Consequences for aphid feeding behaviour and growth. Entomol. Exp. Appl. 2002, 103, 107–113. [Google Scholar] [CrossRef]
- Innes, R. The positives and negatives of NPR: A unifying model for salicylic acid signaling in plants. Cell 2018, 173, 1314–1315. [Google Scholar] [CrossRef]
- Ding, Y.; Sun, T.; Ao, K.; Peng, Y.; Zhang, Y.; Li, X.; Zhang, Y. Opposite roles of salicylic acid receptors NPR1 and NPR3/NPR4 in transcriptional regulation of plant immunity. Cell 2018, 173, 1454–1467.e15. [Google Scholar] [CrossRef]
- Morkunas, I.; Mai, V.C.; Gabryś, B. Phytohormonal signaling in plant responses to aphid feeding. Acta Physiol. Plant 2011, 33, 2057–2073. [Google Scholar] [CrossRef]
- Hillwig, M.S.; Chiozza, M.; Casteel, C.L.; Lau, S.T.; Hohenstein, J.; Hernández, E.; Jander, G.; MacIntosh, G.C. Abscisic acid deficiency increases defence responses against Myzus persicae in Arabidopsis. Mol. Plant Pathol. 2016, 17, 225–235. [Google Scholar] [CrossRef] [PubMed]
- Lee, S.; Cassone, B.J.; Wijeratne, A.; Jun, T.-H.; Michel, A.P.; Mian, M.A.R. Transcriptomic dynamics in soybean near-isogenic lines differing in alleles for an aphid resistance gene, following infestation by soybean aphid biotype 2. BMC Genom. 2017, 18, 472. [Google Scholar] [CrossRef] [PubMed]
- Palmer, N.A.; Chowda-Reddy, R.V.; Muhle, A.A.; Tatineni, S.; Yuen, G.; Edmé, S.J.; Mitchell, R.B.; Sarath, G. Transcriptome divergence during leaf development in two contrasting switchgrass (Panicum virgatum L.) cultivars. PLoS ONE 2019, 14, e222080. [Google Scholar] [CrossRef]
- Domergue, F.; Vishwanath, S.J.; Joubès, J.; Ono, J.; Lee, J.A.; Bourdon, M.; Alhattab, R.; Lowe, C.; Pascal, S.; Lessire, R.; et al. Three Arabidopsis fatty acyl-coenzyme A reductases, FAR1, FAR4, and FAR5, generate primary fatty alcohols associated with suberin deposition. Plant Physiol. 2010, 153, 1539–1554. [Google Scholar] [CrossRef]
- Molina, I.; Li-Beisson, Y.; Beisson, F.; Ohlrogge, J.B.; Pollard, M. Identification of an Arabidopsis feruloyl-coenzyme a transferase required for suberin synthesis. Plant Physiol. 2009, 151, 1317–1328. [Google Scholar] [CrossRef]
- Anderson, J.P.; Badruzsaufari, E.; Schenk, P.M.; Manners, J.M.; Desmond, O.J.; Ehlert, C.; Maclean, D.J.; Ebert, P.R.; Kazan, K. Antagonistic interaction between abscisic acid and jasmonate-ethylene signaling pathways modulates defense gene expression and disease resistance in Arabidopsis. Plant Cell 2004, 16, 3460–3479. [Google Scholar] [CrossRef]
- Escudero-Martinez, C.M.; Morris, J.A.; Hedley, P.E.; Bos, J.I.B. Barley transcriptome analyses upon interaction with different aphid species identify thionins contributing to resistance. Plant Cell Environ. 2017, 40, 2628–2643. [Google Scholar] [CrossRef]
- Neupane, S.; Varenhorst, A.J.; Nepal, M.P. Transcriptome profiling of induced susceptibility effects on soybean–soybean aphid (Hemiptera: Aphididae) interaction. BMC Res. Notes 2019, 12, 325. [Google Scholar] [CrossRef]
- Duhlian, L.; Koramutla, M.K.; Subramanian, S.; Chamola, R.; Bhattacharya, R. Comparative transcriptomics revealed differential regulation of defense related genes in Brassica juncea leading to successful and unsuccessful infestation by aphid species. Sci. Rep. 2020, 10, 10583. [Google Scholar] [CrossRef] [PubMed]
- Corrales, A.-R.; Nebauer, S.G.; Carrillo, L.; Fernández-Nohales, P.; Marqués, J.; Renau-Morata, B.; Granell, A.; Pollmann, S.; Vicente-Carbajosa, J.; Molina, R.-V.; et al. Characterization of tomato Cycling Dof Factors reveals conserved and new functions in the control of flowering time and abiotic stress responses. J. Exp. Bot. 2014, 65, 995–1012. [Google Scholar] [CrossRef] [PubMed]
- Ma, J.; Li, M.-Y.; Wang, F.; Tang, J.; Xiong, A.-S. Genome-wide analysis of Dof family transcription factors and their responses to abiotic stresses in Chinese cabbage. BMC Genom. 2015, 16, 33. [Google Scholar] [CrossRef] [PubMed]
- Martínez, M.; Rubio-Somoza, I.; Fuentes, R.; Lara, P.; Carbonero, P.; Díaz, I. The barley cystatin gene (Icy) is regulated by DOF transcription factors in aleurone cells upon germination. J. Exp. Bot. 2005, 56, 547–556. [Google Scholar] [CrossRef]
- Wen, C.; Cheng, Q.; Zhao, L.; Mao, A.; Yang, J.; Yu, S.; Weng, Y.; Xu, Y. Identification and characterisation of Dof transcription factors in the cucumber genome. Sci. Rep. 2016, 6, 1–11. [Google Scholar] [CrossRef]
- White, C.; Eigenbrode, S.D. Effects of surface wax variation in Pisum sativum on herbivorous and entomophagous insects in the field. Environ. Entomol. 2000, 29, 773–780. [Google Scholar] [CrossRef]
- Wójcicka, A. Surface waxes as a plant defense barrier towards grain aphid. Acta Biol. Crac. Ser. Bot. 2015, 57, 1. [Google Scholar] [CrossRef]
- Wójcicka, A. Changes in pigment content of triticale genotypes infested with grain aphid Sitobion avenae (Fabricius) (Homoptera: Aphididae). Acta Biol. Crac. Ser. Bot. 2014, 56, 121–127. [Google Scholar] [CrossRef]
- Bragg, J.; Tomasi, P.; Zhang, L.; Williams, T.; Wood, D.; Lovell, J.T.; Healey, A.; Schmutz, J.; Bonnette, J.E.; Cheng, P.; et al. Environmentally responsive QTL controlling surface wax load in switchgrass. Appl. Genet. 2020. [Google Scholar] [CrossRef]
- Heng-Moss, T.M.; Baxendale, F.P.; Riordan, T.P.; Foster, J.E. Evaluation of buffalograss germplasm for resistance to Blissus occiduus (Hemiptera: Lygaeidae). J. Econ. Entomol. 2002, 95, 1054–1058. [Google Scholar] [CrossRef][Green Version]
- Palmer, N.A.; Donze-Reiner, T.; Horvath, D.; Heng-Moss, T.; Waters, B.; Tobias, C.; Sarath, G. Switchgrass (Panicum virgatum L) flag leaf transcriptomes reveal molecular signatures of leaf development, senescence, and mineral dynamics. Funct. Integr. Genom. 2015, 15, 1–16. [Google Scholar] [CrossRef] [PubMed]
- Goodstein, D.M.; Shu, S.; Howson, R.; Neupane, R.; Hayes, R.D.; Fazo, J.; Mitros, T.; Dirks, W.; Hellsten, U.; Putnam, N.; et al. Phytozome: A comparative platform for green plant genomics. Nucleic Acids Res. 2012, 40, D1178–D1186. [Google Scholar] [CrossRef]
- Dobin, A.; Davis, C.A.; Schlesinger, F.; Drenkow, J.; Zaleski, C.; Jha, S.; Batut, P.; Chaisson, M.; Gingeras, T.R. STAR: Ultrafast universal RNA-seq aligner. Bioinformatics 2013, 29, 15–21. [Google Scholar] [CrossRef]
- Liao, Y.; Smyth, G.K.; Shi, W. featureCounts: An efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 2014, 30, 923–930. [Google Scholar] [CrossRef]
- Love, M.I.; Huber, W.; Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014, 15, 550. [Google Scholar] [CrossRef]
- Langfelder, P.; Horvath, S. WGCNA: An R package for weighted correlation network analysis. BMC Bioinform. 2008, 9, 559. [Google Scholar] [CrossRef] [PubMed]
- Lin, W.-D.; Chen, Y.-C.; Ho, J.; Hsiao, C. GOBU: Toward an integration interface for biological objects. J. Inf. Sci. Eng. 2006, 22, 19–29. [Google Scholar]
- Pan, X.; Welti, R.; Wang, X. Simultaneous quantification of major phytohormones and related compounds in crude plant extracts by liquid chromatography-electrospray tandem mass spectrometry. Phytochemistry 2008, 69, 1773–1781. [Google Scholar] [CrossRef]
- Schmitz, A.J.; Begcy, K.; Sarath, G.; Walia, H. Rice Ovate Family Protein 2 (OFP2) alters hormonal homeostasis and vasculature development. Plant Sci. 2015, 241, 177–188. [Google Scholar] [CrossRef]
- Yuan, M.; Breitkopf, S.B.; Yang, X.; Asara, J.M. A positive/negative ion-switching, targeted mass spectrometry-based metabolomics platform for bodily fluids, cells, and fresh and fixed tissue. Nat. Protoc. 2012, 7, 872–881. [Google Scholar] [CrossRef]
- Tetreault, H.M.; Scully, E.D.; Gries, T.; Palmer, N.A.; Funnell-Harris, D.L.; Baird, L.; Seravalli, J.; Dien, B.S.; Sarath, G.; Clemente, T.E.; et al. Overexpression of the Sorghum bicolor SbCCoAOMT alters cell wall associated hydroxycinnamoyl groups. PLoS ONE 2018, 13, e204153. [Google Scholar] [CrossRef]
- Kanehisa, M.; Sato, Y.; Kawashima, M.; Furumichi, M.; Tanabe, M. KEGG as a reference resource for gene and protein annotation. Nucleic Acids Res. 2016, 44, D457–D462. [Google Scholar] [CrossRef] [PubMed]

| Conditions | Up | Down | Gene Number | |
|---|---|---|---|---|
| Kanlow | 5 DAI | 110 | 52 | 162 |
| 10 DAI | 55 | 37 | 92 | |
| 15 DAI | 313 | 57 | 370 | |
| Summer | 5 DAI | 760 | 818 | 1578 |
| 10 DAI | 574 | 33 | 607 | |
| 15 DAI | 1485 | 366 | 1851 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Pingault, L.; Palmer, N.A.; Koch, K.G.; Heng-Moss, T.; Bradshaw, J.D.; Seravalli, J.; Twigg, P.; Louis, J.; Sarath, G. Differential Defense Responses of Upland and Lowland Switchgrass Cultivars to a Cereal Aphid Pest. Int. J. Mol. Sci. 2020, 21, 7966. https://doi.org/10.3390/ijms21217966
Pingault L, Palmer NA, Koch KG, Heng-Moss T, Bradshaw JD, Seravalli J, Twigg P, Louis J, Sarath G. Differential Defense Responses of Upland and Lowland Switchgrass Cultivars to a Cereal Aphid Pest. International Journal of Molecular Sciences. 2020; 21(21):7966. https://doi.org/10.3390/ijms21217966
Chicago/Turabian StylePingault, Lise, Nathan A. Palmer, Kyle G. Koch, Tiffany Heng-Moss, Jeffrey D. Bradshaw, Javier Seravalli, Paul Twigg, Joe Louis, and Gautam Sarath. 2020. "Differential Defense Responses of Upland and Lowland Switchgrass Cultivars to a Cereal Aphid Pest" International Journal of Molecular Sciences 21, no. 21: 7966. https://doi.org/10.3390/ijms21217966
APA StylePingault, L., Palmer, N. A., Koch, K. G., Heng-Moss, T., Bradshaw, J. D., Seravalli, J., Twigg, P., Louis, J., & Sarath, G. (2020). Differential Defense Responses of Upland and Lowland Switchgrass Cultivars to a Cereal Aphid Pest. International Journal of Molecular Sciences, 21(21), 7966. https://doi.org/10.3390/ijms21217966





