BacSJ—Another Bacteriocin with Distinct Spectrum of Activity that Targets Man-PTS
Abstract
:1. Introduction
2. Results
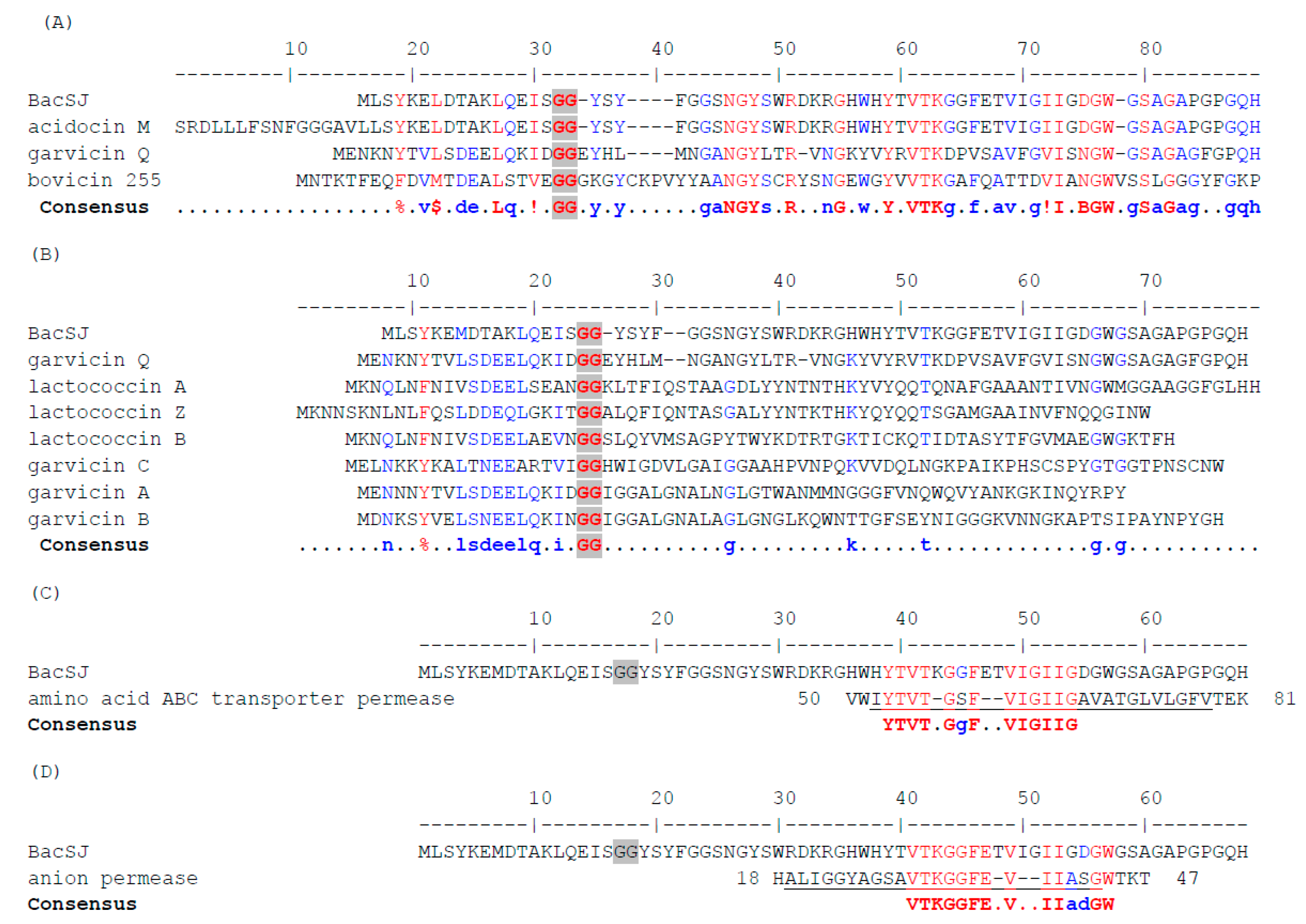
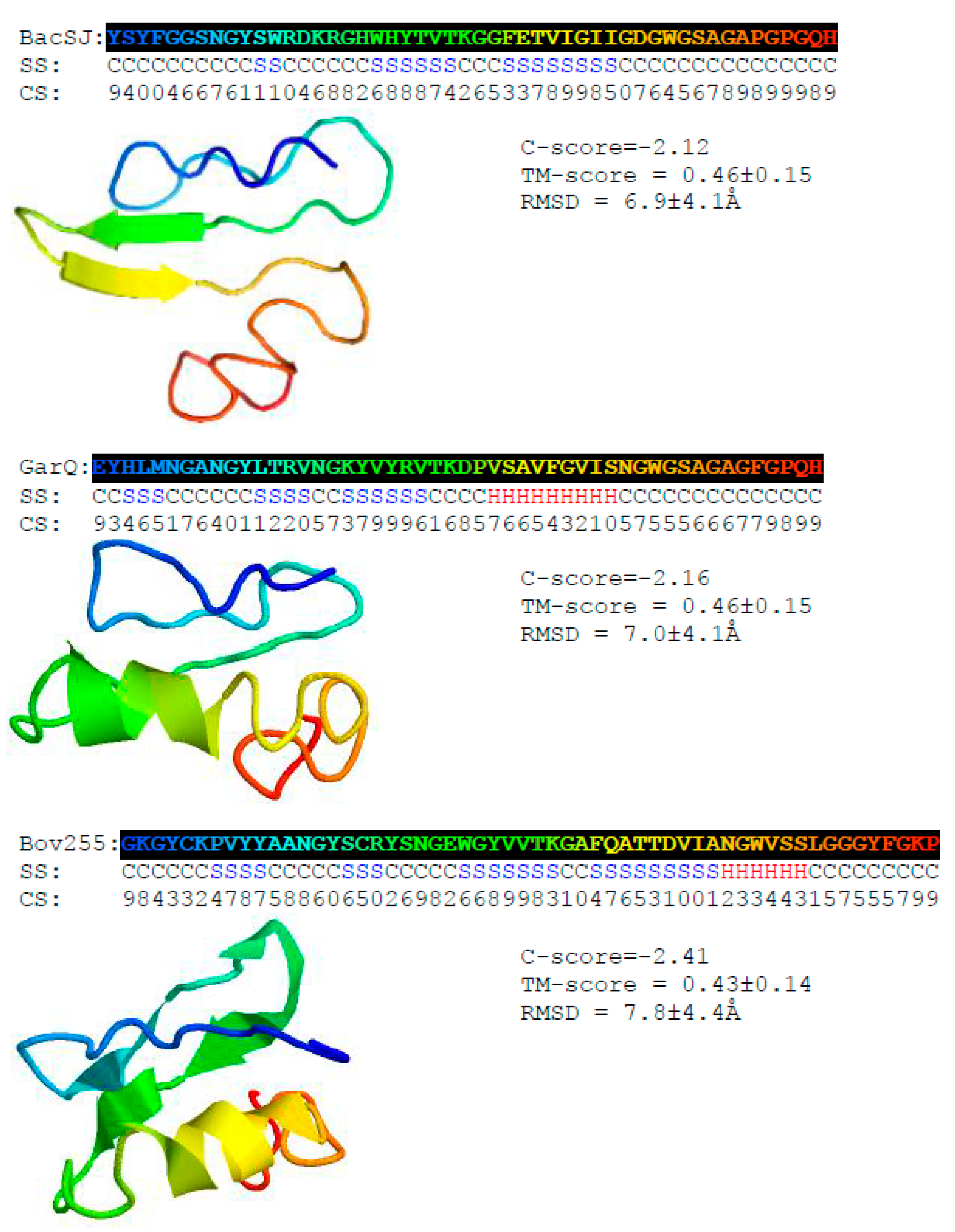
2.1. BacSJ Homologs are Encoded in Some Lactobacillales
2.2. BacSJ Has a Moderate Spectrum of Activity
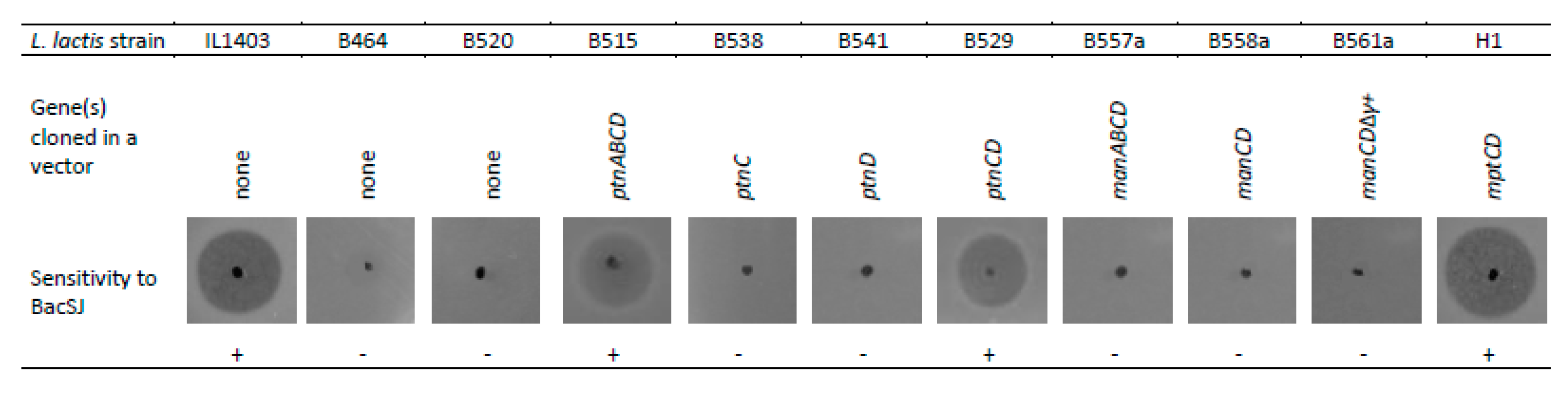
2.3. Man-PTS Subunits IIC and IID are Necessary for BacSJ Activity
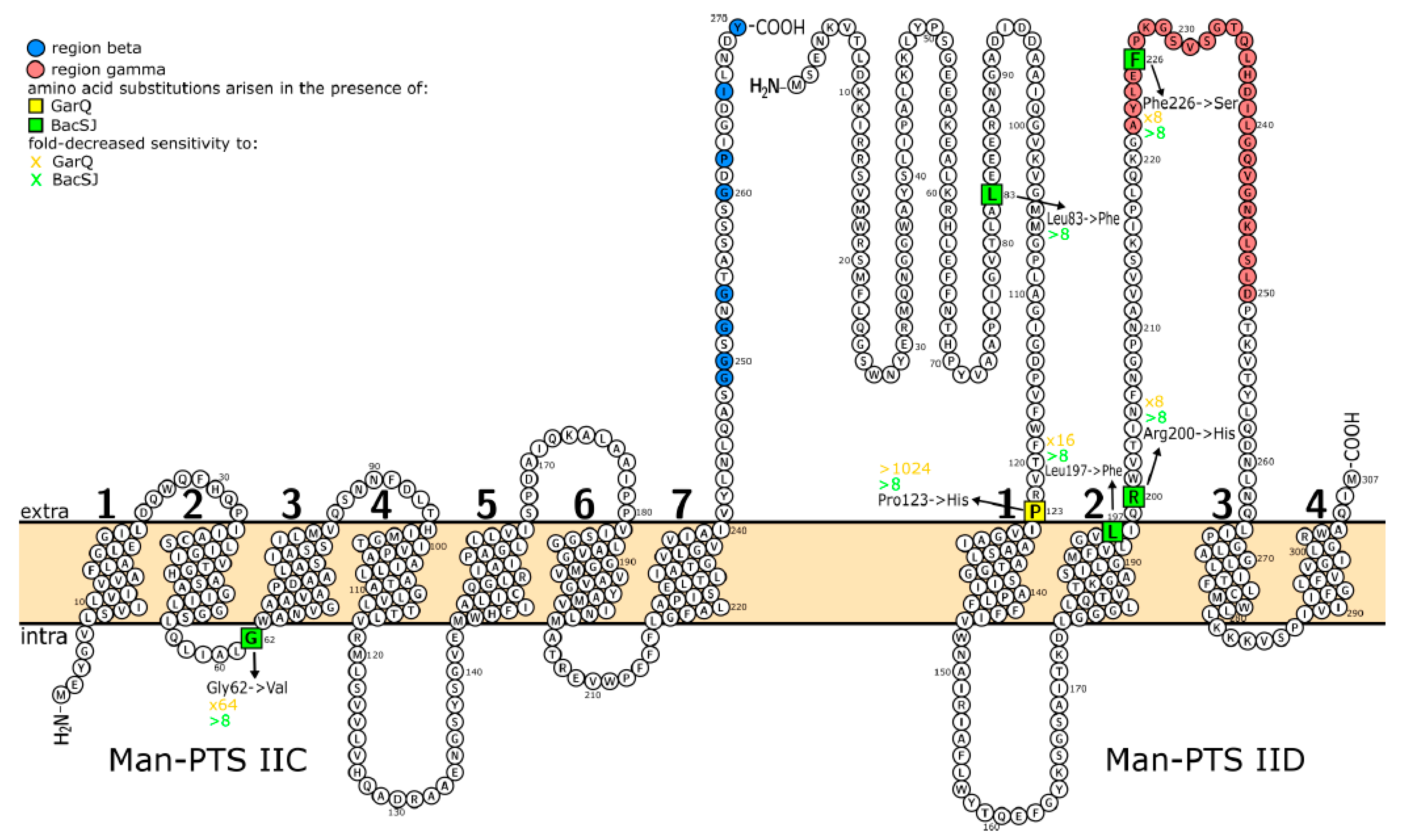
2.4. Targeted Mutagenesis Allows Identification of Man-PTS Amino Acids Needed for BacSJ Activity
3. Discussion
4. Materials and Methods
4.1. Bacterial and Yeast Strains and Growth Conditions
4.2. Bacteriocin Preparation
4.3. Activity Spectrum Assay and Selection of Resistant Mutants
4.4. DNA Isolation and Manipulation
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Yang, S.-C.; Lin, C.-H.; Sung, C.T.; Fang, J.-Y. Antibacterial activities of bacteriocins: Application in foods and pharmaceuticals. Front. Microbiol. 2014, 5, 241. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cleveland, J.; Montville, T.J.; Nes, I.F.; Chikindas, M.L. Bacteriocins: Safe, natural antimicrobials for food preservation. Int. J. Food Microbiol. 2001, 71, 1–20. [Google Scholar] [CrossRef]
- Silva, C.C.G.; Silva, S.P.M.; Ribeiro, S.C. Application of bacteriocins and protective cultures in dairy food preservation. Front. Microbiol. 2018, 9, 594. [Google Scholar] [CrossRef] [PubMed]
- Cotter, P.D.; Ross, R.P.; Hill, C. Bacteriocins—A viable alternative to antibiotics? Nat. Rev. Genet. 2013, 11, 95–105. [Google Scholar] [CrossRef]
- Hegarty, J.W.; Guinane, C.M.; Ross, R.P.; Hill, C.; Cotter, P.D. Bacteriocin production: A relatively unharnessed probiotic trait? F1000Research 2016, 5, 2587. [Google Scholar] [CrossRef]
- Baindara, P.; Korpole, S.; Grover, V. Bacteriocins: Perspective for the development of novel anticancer drugs. Appl. Microbiol. Biotechnol. 2018, 102, 10393–10408. [Google Scholar] [CrossRef]
- Perez, R.H.; Zendo, T.; Sonomoto, K. Novel bacteriocins from lactic acid bacteria (LAB): Various structures and applications. Microb. Cell Factories 2014, 13, S3. [Google Scholar] [CrossRef] [Green Version]
- Havarstein, L.S.; Diep, D.B.; Nes, I.F. A family of bacteriocin ABC transporters carry out proteolytic processing of their substrates concomitant with export. Mol. Microbiol. 1995, 16, 229–240. [Google Scholar] [CrossRef]
- Cintas, L.M.; Casaus, P.; Håvarstein, L.S.; Hernández, P.; Nes, I.F. Biochemical and genetic characterization of enterocin P, a novel sec-dependent bacteriocin from Enterococcus faecium P13 with a broad antimicrobial spectrum. Appl. Environ. Microbiol. 1997, 63, 4321–4330. [Google Scholar] [CrossRef] [Green Version]
- Alvarez-Sieiro, P.; Montalbán-López, M.; Mu, D.; Kuipers, O.P. Bacteriocins of lactic acid bacteria: Extending the family. Appl. Microbiol. Biotechnol. 2016, 100, 2939–2951. [Google Scholar] [CrossRef] [Green Version]
- Perez, R.H.; Zendo, T.; Sonomoto, K. Circular and leaderless bacteriocins: Biosynthesis, mode of action, applications, and prospects. Front. Microbiol. 2018, 9, 2085. [Google Scholar] [CrossRef] [PubMed]
- Cotter, P.D. An ‘Upp’-turn in bacteriocin receptor identification. Mol. Microbiol. 2014, 92, 1159–1163. [Google Scholar] [CrossRef] [PubMed]
- Martínez, B.; Böttiger, T.; Schneider, T.; Rodríguez, A.; Sahl, H.-G.; Wiedemann, I. Specific interaction of the unmodified bacteriocin lactococcin 972 with the cell wall precursor lipid II. Appl. Environ. Microbiol. 2008, 74, 4666–4670. [Google Scholar] [CrossRef] [Green Version]
- Oppegård, C.; Kjos, M.; Veening, J.-W.; Nissen-Meyer, J.; Kristensen, T. A putative amino acid transporter determines sensitivity to the two-peptide bacteriocin plantaricin JK. MicrobiologyOpen 2016, 5, 700–708. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ovchinnikov, K.V.; Kristiansen, P.E.; Straume, D.; Jensen, M.S.; Aleksandrzak-Piekarczyk, T.; Nes, I.F.; Diep, D.B. The leaderless bacteriocin enterocin K1 is highly potent against Enterococcus faecium: A study on structure, target spectrum and receptor. Front. Microbiol. 2017, 8, 774. [Google Scholar] [CrossRef] [Green Version]
- Tymoszewska, A.; Diep, D.B.; Wirtek, P.; Aleksandrzak-Piekarczyk, T. The non-lantibiotic bacteriocin garvicin Q targets Man-PTS in a broad spectrum of sensitive bacterial genera. Sci. Rep. 2017, 7, 8359. [Google Scholar] [CrossRef] [Green Version]
- Tymoszewska, A.; Diep, D.B.; Aleksandrzak-Piekarczyk, T. The extracellular loop of Man-PTS subunit IID is responsible for the sensitivity of Lactococcus garvieae to garvicins A, B and C. Sci. Rep. 2018, 8, 15790. [Google Scholar] [CrossRef] [Green Version]
- Daba, G.M.; Ishibashi, N.; Gong, X.; Taki, H.; Yamashiro, K.; Lim, Y.Y.; Zendo, T.; Sonomoto, K. Characterisation of the action mechanism of a Lactococcus-specific bacteriocin, lactococcin Z. J. Biosci. Bioeng. 2018, 126, 603–610. [Google Scholar] [CrossRef]
- Heeney, D.D.; Yarov-Yarovoy, V.; Marco, M.L. Sensitivity to the two peptide bacteriocin plantaricin EF is dependent on CorC, a membrane-bound, magnesium/cobalt efflux protein. Microbiol. Open 2019, 8, e827. [Google Scholar] [CrossRef] [Green Version]
- Jeckelmann, J.-M.; Erni, B. The mannose phosphotransferase system (Man-PTS)-mannose transporter and receptor for bacteriocins and bacteriophages. Biochim. Biophys. Acta (BBA) Biomembr. 2020, 1862, 183412. [Google Scholar] [CrossRef]
- Ramnath, M.; Beukes, M.; Tamura, K.; Hastings, J.W. Absence of a putative mannose-specific phosphotransferase system enzyme IIAB component in a leucocin A-resistant strain of Listeria monocytogenes, as shown by two-dimensional sodium dodecyl sulfate-polyacrylamide gel electrophoresis. Appl. Environ. Microbiol. 2000, 66, 3098–3101. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dalet, K.; Cenatiempo, Y.; Cossart, P.; Héchard, Y. European Listeria Genome Consortium A sigma(54)-dependent PTS permease of the mannose family is responsible for sensitivity of Listeria monocytogenes to mesentericin Y105. Microbiol. Read. Engl. 2001, 147, 3263–3269. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ramnath, M.; Arous, S.; Gravesen, A.; Hastings, J.W.; Hechard, Y. Expression of mptC of Listeria monocytogenes induces sensitivity to class IIa bacteriocins in Lactococcus lactis. Microbiol. Read. Engl. 2004, 150, 2663–2668. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Diep, D.B.; Skaugen, M.; Salehian, Z.; Holo, H.; Nes, I.F. Common mechanisms of target cell recognition and immunity for class II bacteriocins. Proc. Natl. Acad. Sci. USA 2007, 104, 2384–2389. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kjos, M.; Salehian, Z.; Nes, I.F.; Diep, D.B. An extracellular loop of the mannose phosphotransferase system component IIC is responsible for specific targeting by class IIa bacteriocins. J. Bacteriol. 2010, 192, 5906–5913. [Google Scholar] [CrossRef] [Green Version]
- Chikindas, M.L.; García-Garcerá, M.J.; Driessen, A.J.; Ledeboer, A.M.; Nissen-Meyer, J.; Nes, I.F.; Abee, T.; Konings, W.N.; Venema, G. Pediocin PA-1, a bacteriocin from Pediococcus acidilactici PAC1.0, forms hydrophilic pores in the cytoplasmic membrane of target cells. Appl. Environ. Microbiol. 1993, 59, 3577–3584. [Google Scholar] [CrossRef] [Green Version]
- Minahk, C.J.; Farãas, M.E.; Sesma, F.; Morero, R.D.; Farías, M.E. Effect of enterocin CRL35 on Listeria monocytogenes cell membrane. FEMS Microbiol. Lett. 2000, 192, 79–83. [Google Scholar] [CrossRef] [Green Version]
- Van Belkum, M.J.; Kok, J.; Venema, G.; Holo, H.; Nes, I.F.; Konings, W.N.; Abee, T. The bacteriocin lactococcin A specifically increases permeability of lactococcal cytoplasmic membranes in a voltage-independent, protein-mediated manner. J. Bacteriol. 1991, 173, 7934–7941. [Google Scholar] [CrossRef] [Green Version]
- Venema, K.; Abee, T.; Haandrikman, A.J.; Leenhouts, K.J.; Kok, J.; Konings, W.N.; Venema, G. Mode of action of lactococcin B, a thiol-activated bacteriocin from Lactococcus lactis. Appl. Environ. Microbiol. 1993, 59, 1041–1048. [Google Scholar] [CrossRef] [Green Version]
- Colombo, N.S.R.; Chalón, M.C.; Navarro, S.A.; Bellomio, A. Pediocin-like bacteriocins: New perspectives on mechanism of action and immunity. Curr. Genet. 2018, 64, 345–351. [Google Scholar] [CrossRef]
- Barraza, D.E.; Colombo, N.S.R.; Galván, A.E.; Acuña, L.; Minahk, C.J.; Bellomio, A.; Chalón, M.C. New insights into enterocin CRL35: Mechanism of action and immunity revealed by heterologous expression in Escherichia coli. Mol. Microbiol. 2017, 105, 922–933. [Google Scholar] [CrossRef] [Green Version]
- Maldonado-Barragán, A.; Cárdenas, N.; Martínez, B.; Ruiz-Barba, J.L.; Fernández-Garayzábal, J.F.; Rodríguez, J.M.; Gibello, A. Garvicin A, a novel class IId bacteriocin from Lactococcus garvieae that inhibits septum formation in L. garvieae strains. Appl. Environ. Microbiol. 2013, 79, 4336–4346. [Google Scholar] [CrossRef] [Green Version]
- Lozo, J.; Jovcic, B.; Kojic, M.; Dalgalarrondo, M.; Chobert, J.-M.; Haertlé, T.; Topisirovic, L. Molecular characterization of a novel bacteriocin and an unusually large aggregation factor of Lactobacillus paracasei subsp. paracasei BGSJ2-8, a natural isolate from homemade cheese. Curr. Microbiol. 2007, 55, 266–271. [Google Scholar] [CrossRef]
- Kojic, M.; Lozo, J.; Jovcic, B.; Strahinic, I.; Fira, D.; Topisirovic, L. Construction of a new shuttle vector and its use for cloning and expression of two plasmid-encoded bacteriocins from Lactobacillus paracasei subsp. paracasei BGSJ2-8. Int. J. Food Microbiol. 2010, 140, 117–124. [Google Scholar] [CrossRef]
- Tosukhowong, A.; Zendo, T.; Visessanguan, W.; Roytrakul, S.; Pumpuang, L.; Jaresitthikunchai, J.; Sonomoto, K. Garvieacin Q, a novel class II bacteriocin from Lactococcus garvieae BCC 43578. Appl. Environ. Microbiol. 2012, 78, 1619–1623. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Whitford, M.F.; McPherson, M.A.; Forster, R.J.; Teather, R.M. Identification of bacteriocin-like inhibitors from rumen Streptococcus spp. and isolation and characterization of bovicin 255. Appl. Environ. Microbiol. 2001, 67, 569–574. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kanatani, K.; Tahara, T.; Oshimura, M.; Sano, K.; Umezawa, C. Cloning and nucleotide sequence of the gene for acidocin 8912, a bacteriocin from Lactobacillus acidophilus TK8912. Lett. Appl. Microbiol. 1995, 21, 384–386. [Google Scholar] [CrossRef] [PubMed]
- Watanabe, K.; Suzuki, T. Universal genetic code and its natural variations. In eLS; John Wiley &Sons, Ltd.: Chichester, UK, 2008. [Google Scholar] [CrossRef]
- Tahara, T.; Kanatani, K.; Yoshida, K.; Miura, H.; Sakamoto, M.; Oshimura, M. Purification and some properties of acidocin 8912, a novel bacteriocin produced by Lactobacillus acidophilus TK8912. Biosci. Biotechnol. Biochem. 1992, 56, 1212–1215. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Thakur, M.; Asrani, R.K.; Patial, V. Listeria monocytogenes: A food-borne pathogen In Foodborne Diseases; Elsevier: Amsterdam, The Netherlands, 2018; pp. 157–192. [Google Scholar]
- Kaiser, J.C.; Heinrichs, D.E. Branching out: Alterations in bacterial physiology and virulence due to branched-chain amino acid deprivation. mBio 2018, 9, e01188-18. [Google Scholar] [CrossRef] [Green Version]
- Mansilla, M.C.; De Mendoza, D. The Bacillus subtilis cysP gene encodes a novel sulphate permease related to the inorganic phosphate transporter (Pit) family. Microbiology 2000, 146, 815–821. [Google Scholar] [CrossRef] [Green Version]
- Raya, R.; Bardowski, J.; Andersen, P.S.; Ehrlich, S.D.; Chopin, A. Multiple transcriptional control of the Lactococcus lactis trp operon. J. Bacteriol. 1998, 180, 3174–3180. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sanger, F.; Nicklen, S.; Coulson, A.R. DNA sequencing with chain-terminating inhibitors. Proc. Natl. Acad. Sci. USA 1977, 74, 5463–5467. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gasteiger, E.; Gattiker, A.; Hoogland, C.; Ivanyi, I.; Appel, R.D.; Bairoch, A. ExPASy: ExPASy: The proteomics server for in-depth protein knowledge and analysis. Nucleic Acids Res. 2003, 31, 3784–3788. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Corpet, F. Multiple sequence alignment with hierarchical clustering. Nucleic Acids Res. 1988, 16, 10881–10890. [Google Scholar] [CrossRef]
- Tusnady, G.E.; Simon, I. The HMMTOP transmembrane topology prediction server. Bioinformatics 2001, 17, 849–850. [Google Scholar] [CrossRef]
- Omasits, U.; Ahrens, C.H.; Müller, S.; Wollscheid, B. Protter: Interactive protein feature visualization and integration with experimental proteomic data. Bioinformatics 2014, 30, 884–886. [Google Scholar] [CrossRef] [Green Version]
- Yang, J.; Yan, R.; Roy, A.; Xu, D.; Poisson, J.; Zhang, Y. The I-TASSER suite: Protein structure and function prediction. Nat. Methods 2015, 12, 7–8. [Google Scholar] [CrossRef] [Green Version]
| Group | Indicator Strain | BacSJ Activity |
|---|---|---|
| Fungi | Candida albicans CAI-4 | − |
| G− | Campylobacter jejuni 12 | − |
| G− | Campylobacter jejuni 480 | − |
| G− | Campylobacter jejuni 81176 | − |
| G− | Campylobacter coli 23/1 | − |
| G− | Escherichia coli EC1000 | − |
| G− | Escherichia coli TG1 | − |
| G− | Pseudomonas aeruginosa ATCC 9027 | − |
| G− | Salmonella typhimurium TT622 | − |
| G+ | Bacillus cereus IBB3390 | − |
| G+ | Bacillus subtilis BSB1 | − |
| G+ | Carnobacterium maltaromaticum IBB3447 | + |
| G+ | Enterococcus durans IBB3441 | + |
| G+ | Enterococcus faecalis IBB3439 | + |
| G+ | Enterococcus faecalis IBB3444 | + |
| G+ | Enterococcus faecium LMGT 2783 | + |
| G+ | Enterococcus faecium LMGT 2787 | + |
| G+ | Lactobacillus johnsonii IBB3155 | +/− |
| G+ | Lactobacillus kunkeei AH1 | − |
| G+ | Lactobacillus kunkeei AH38 | − |
| G+ | Lactobacillus kunkeei AH119 | − |
| G+ | Lactobacillus paracasei IBB3418 | + |
| G+ | Lactobacillus paracasei IBB3425 | + |
| G+ | Lactobacillus paracasei IBB3426 | + |
| G+ | Lactobacillus paracasei IBB3427 | + |
| G+ | Lactobacillus paracasei IBB3428 | + |
| G+ | Lactobacillus paracasei LOCK 0919 | + |
| G+ | Lactobacillus paracasei subsp. paracasei IBB3423 | + |
| G+ | Lactobacillus paraplantarum IBB3438 | + |
| G+ | Lactobacillus plantarum NC8 | + |
| G+ | Lactobacillus plantarum WCSF1 | + |
| G+ | Lactobacillus plantarum IBB3433 | + |
| G+ | Lactobacillus plantarum IBB3436 | + |
| G+ | Lactobacillus plantarum subsp. plantarum IBB3434 | + |
| G+ | Lactobacillus rhamnosus GG | + |
| G+ | Lactobacillus rhamnosus IBB3429 | + |
| G+ | Lactobacillus rhamnosus LOCK 0900 | ++ |
| G+ | Lactobacillus rhamnosus LOCK 0908 | + |
| G+ | Lactobacillus salivarius IBB3154 | + |
| G+ | Lactococcus garvieae IBB3403 | − |
| G+ | Lactococcus garvieae IBB66 | − |
| G+ | Lactococcus lactis IBB3404 | + |
| G+ | Lactococcus lactis IBB3411 | + |
| G+ | Lactococcus lactis QU5 LMGT 3419 | + |
| G+ | Lactococcus lactis subsp. cremoris IBB3409 | + |
| G+ | Lactococcus lactis subsp. lactis IBB3407 | + |
| G+ | Lactococcus lactis subsp. lactis IL1403 | + |
| G+ | Lactococcus raffinolactis IBB91 | + |
| G+ | Lactococcus lactis IBB3446 | + |
| G+ | Leuconostoc mesenteroides IBB3442 | + |
| G+ | Leuconostoc mesenteroides IBB3443 | + |
| G+ | Listeria monocytogenes EDG-e | + |
| G+ | Pediococcus acidilactici LMGT 2002 | − |
| G+ | Pediococcus parvulus IBB3448 | − |
| G+ | Pediococcus pentosaceus IBB3369 | + |
| G+ | Staphylococcus aureus ATCC 6538 | − |
| G+ | Staphylococcus caprae DSM-20608 | − |
| G+ | Staphylococcus delphini DSM-20771 | − |
| G+ | Staphylococcus epidermidis DSM-20044 | − |
| G+ | Staphylococcus hyicus DSM-20459 | − |
| G+ | Staphylococcus intermedius DSM-20373 | − |
| G+ | Staphylococcus lugdunensis DSM-4804 | − |
| G+ | Staphylococcus pseudintermedius DSM-21284 | − |
| G+ | Staphylococcus saprophyticus DSM-18669 | − |
| G+ | Staphylococcus schleiferi DSM-6628 | − |
| G+ | Streptococcus agalactiae IBB123 | − |
| G+ | Streptococcuss agalactiae IBB130 | − |
| G+ | Streptococcus mitis IBB3449 | − |
| G+ | Streptococcus parauberis IBB272 | − |
| G+ | Streptococcus sobrinus IBB3450 | − |
| Mutant | Mutation | Amino Acid Change | Sensitivity to (Fold-Decreased Relative to WT) | Position in the Cell Membrane of Substituted Man-PTS Amino Acid | |
|---|---|---|---|---|---|
| GarQ | BacSJ | ||||
| GarQ-resistant mutant | |||||
| LLN1 | C368→ A in ptnD | Pro123 → His | >1024× | >8× | outside |
| BacSJ-resistant mutants | |||||
| M3, M8, M9 | G185 → T in ptnC | Gly62→Val | 64× | >8× | inside |
| M6, M33 | G599→ A in ptnD | Arg200 → His | 8× | >8× | outside |
| M16 | C247→T in ptnD | Leu83→Phe | 0× | >8× | outside |
| M19 | T677→C in ptnD | Phe226→Ser | 8× | >8× | outside |
| M30 | G591→ T in ptnD | Leu197→Phe | 16× | >8× | transmembrane |
| Strain | IIC | IID | ||||
|---|---|---|---|---|---|---|
| BacSJ sensitive | L. lactis IL1403 | Gly62 i | Leu83 o | Leu197 t | Arg200 o | Phe226 o |
| L. lactis IBB3407 | Gly62 i | Leu83 o | Leu197 t | Arg200 o | Phe226 o | |
| L. lactis IBB3409 | Gly62 i | Leu83 o | Leu197 t | Arg200 o | Phe226 o | |
| L. lactis IBB2955 | Gly62 i | Leu83 o | Leu197 t | Arg200 o | Phe226 o | |
| L. lactis LMGT3419 | Gly62 i | Leu83 o | Leu197 t | Arg200 o | Phe226 o | |
| L. paracasei IBB3418 | Gly62 i | Leu82 o | Leu196 o | Arg199 o | Trp222 o | |
| L. paracasei IBB3424 | Gly62 i | Leu82o | Leu196o | Arg199o | Trp222o | |
| L. paracasei IBB3427 | Gly62 i | Leu82 o | Leu196 o | Arg199 o | Trp222 o | |
| L. paracasei LOCK 0919 | Gly62 i | Leu82 o | Leu196 o | Arg199 o | Trp222 o | |
| L. paraplantarum IBB3438 | Gly62 i | Leu83 o | Leu197 t | Arg200 o | Trp224 o | |
| L. plantarum IBB3036 | Gly62 i | Leu88 o | Leu202 o | Arg205 o | Trp229 o | |
| L. plantarum IBB3434 | Gly62 i | Leu88 o | Leu202 t | Arg205 o | Trp229 o | |
| L. plantarum IBB3436 | Gly62 i | Leu83 o | Leu197 t | Arg200 o | Trp224 o | |
| L. plantarum NC8 | Gly62 i | Leu88 o | Leu202 o | Arg205 o | Trp229 o | |
| L. plantarum WCFS1 | Gly62 i | Leu83 o | Leu197 t | Arg200 o | Trp224 o | |
| L. rhamnosus GG | Gly62 i | Leu82 o | Leu196 o | Arg199 o | Trp222 o | |
| L. rhamnosus IBB3429 | Gly62 i | Leu82 o | Leu196 o | Arg199 o | Trp222 o | |
| L. rhamnosus LOCK 0900 | Gly62 i | Leu82 o | Leu196 o | Arg199 o | Trp222 o | |
| L. rhamnosus LOCK 0908 | Gly62 i | Leu82 o | Leu196 o | Arg199 o | Trp222 o | |
| L. salivarius IBB3154 | Gly60 i | Leu83 o | Leu197 o | Arg200 o | Trp223 o | |
| E. faecium LMGT2783 | Gly60 i | Leu83 o | Leu197 o | Arg200 o | Trp223 o | |
| L. monocytogenes EGD-e | Gly60 i | Leu82 o | Leu196 t | Arg199 o | Trp222 o | |
| BacSJ resistant | L. garvieae IBB66 | Gly60 i | Leu86 o | Leu200 t | Arg203 o | Trp230 o |
| L. garvieae IBB3403 | Gly60 i | Leu86 o | Leu200 t | Arg203 o | Trp230 o | |
| * | * | * | * | : | ||
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Tymoszewska, A.; Walczak, P.; Aleksandrzak-Piekarczyk, T. BacSJ—Another Bacteriocin with Distinct Spectrum of Activity that Targets Man-PTS. Int. J. Mol. Sci. 2020, 21, 7860. https://doi.org/10.3390/ijms21217860
Tymoszewska A, Walczak P, Aleksandrzak-Piekarczyk T. BacSJ—Another Bacteriocin with Distinct Spectrum of Activity that Targets Man-PTS. International Journal of Molecular Sciences. 2020; 21(21):7860. https://doi.org/10.3390/ijms21217860
Chicago/Turabian StyleTymoszewska, Aleksandra, Piotr Walczak, and Tamara Aleksandrzak-Piekarczyk. 2020. "BacSJ—Another Bacteriocin with Distinct Spectrum of Activity that Targets Man-PTS" International Journal of Molecular Sciences 21, no. 21: 7860. https://doi.org/10.3390/ijms21217860





