Overexpression of PtrMYB121 Positively Regulates the Formation of Secondary Cell Wall in Arabidopsis thaliana
Abstract
1. Introduction
2. Results
2.1. Phylogenetic Analysis and Amino Acid Sequence Alignment of PtrMYB121
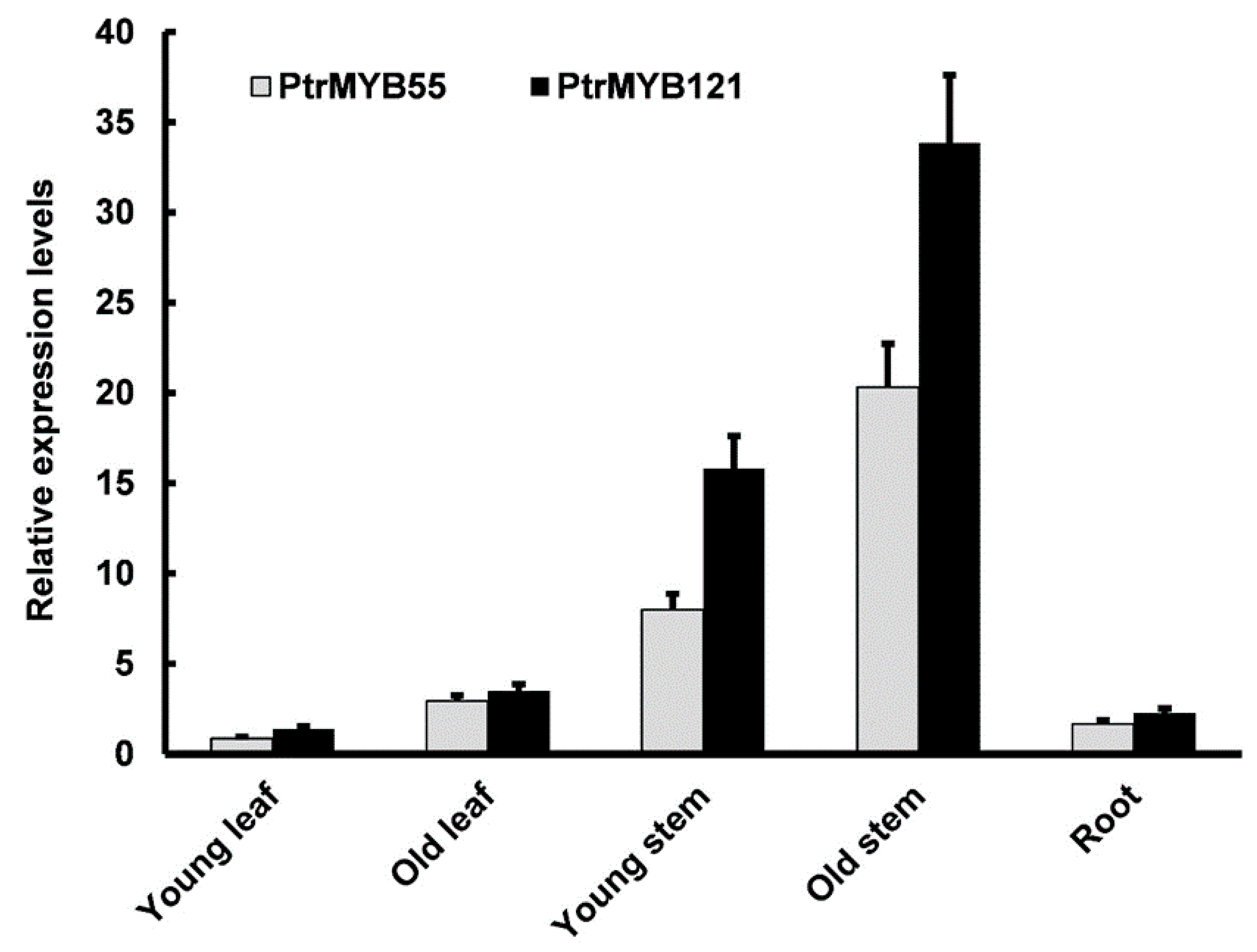
2.2. Tissue Expression Pattern Analysis of PtrMYB55 and PtrMYB121
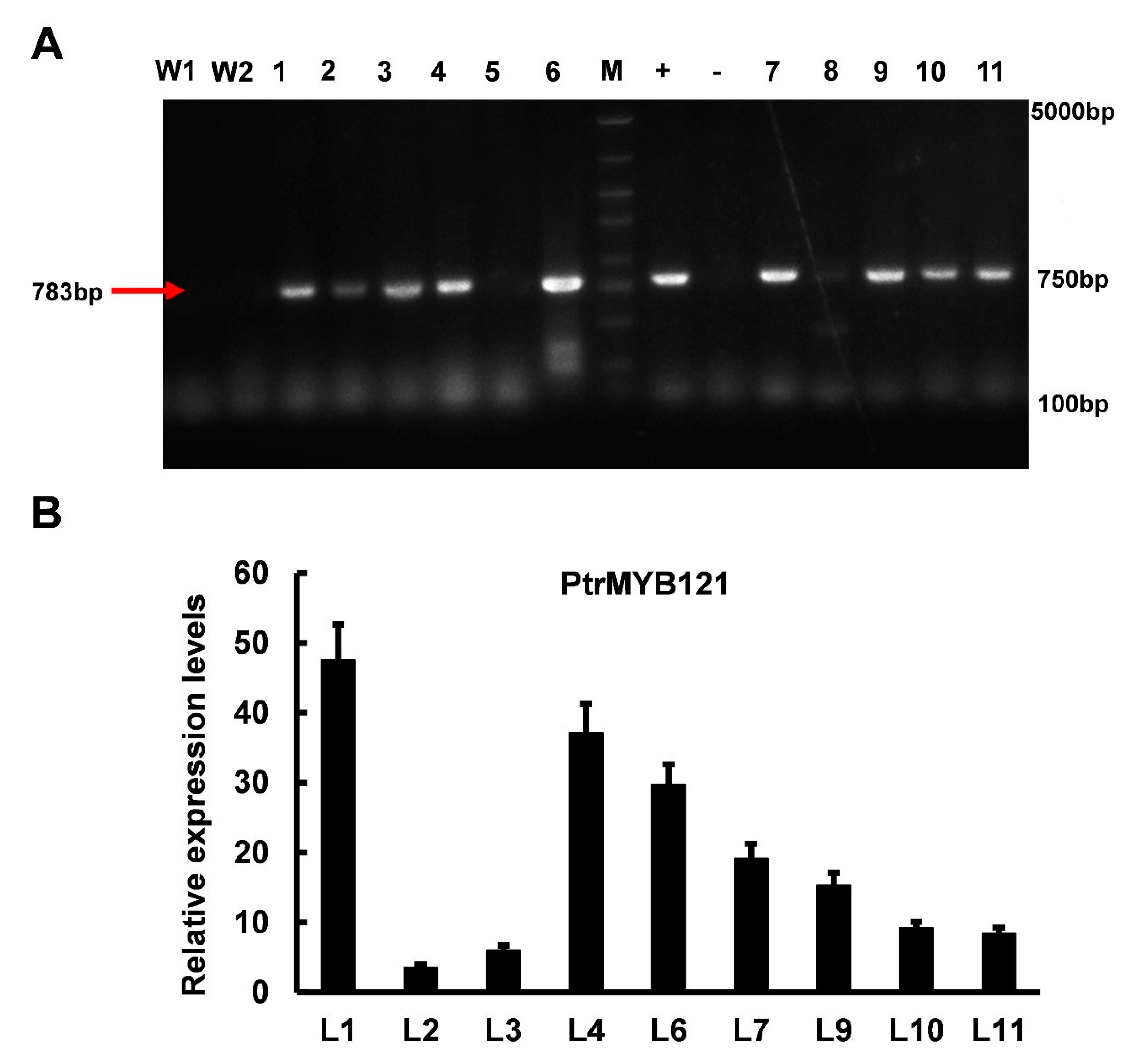
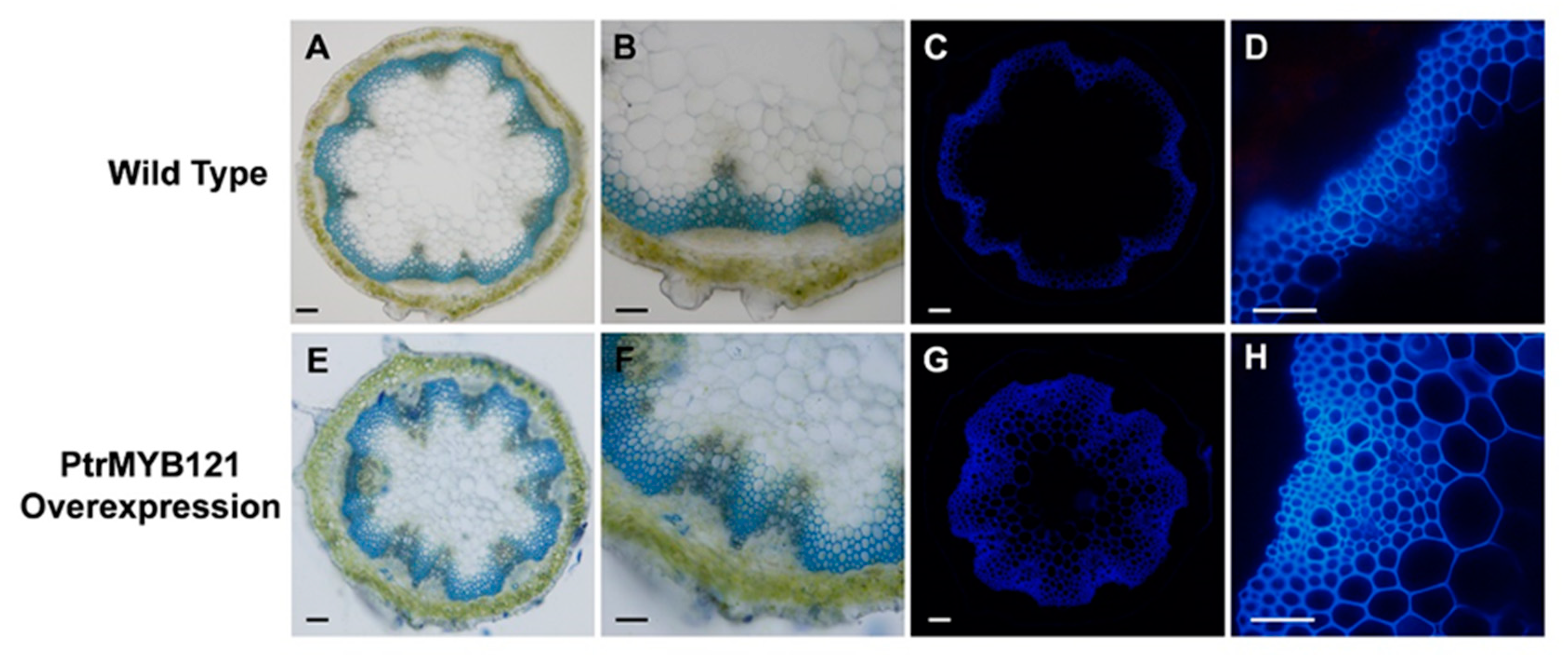
2.3. Overexpression of PtrMYB121 in A. thaliana
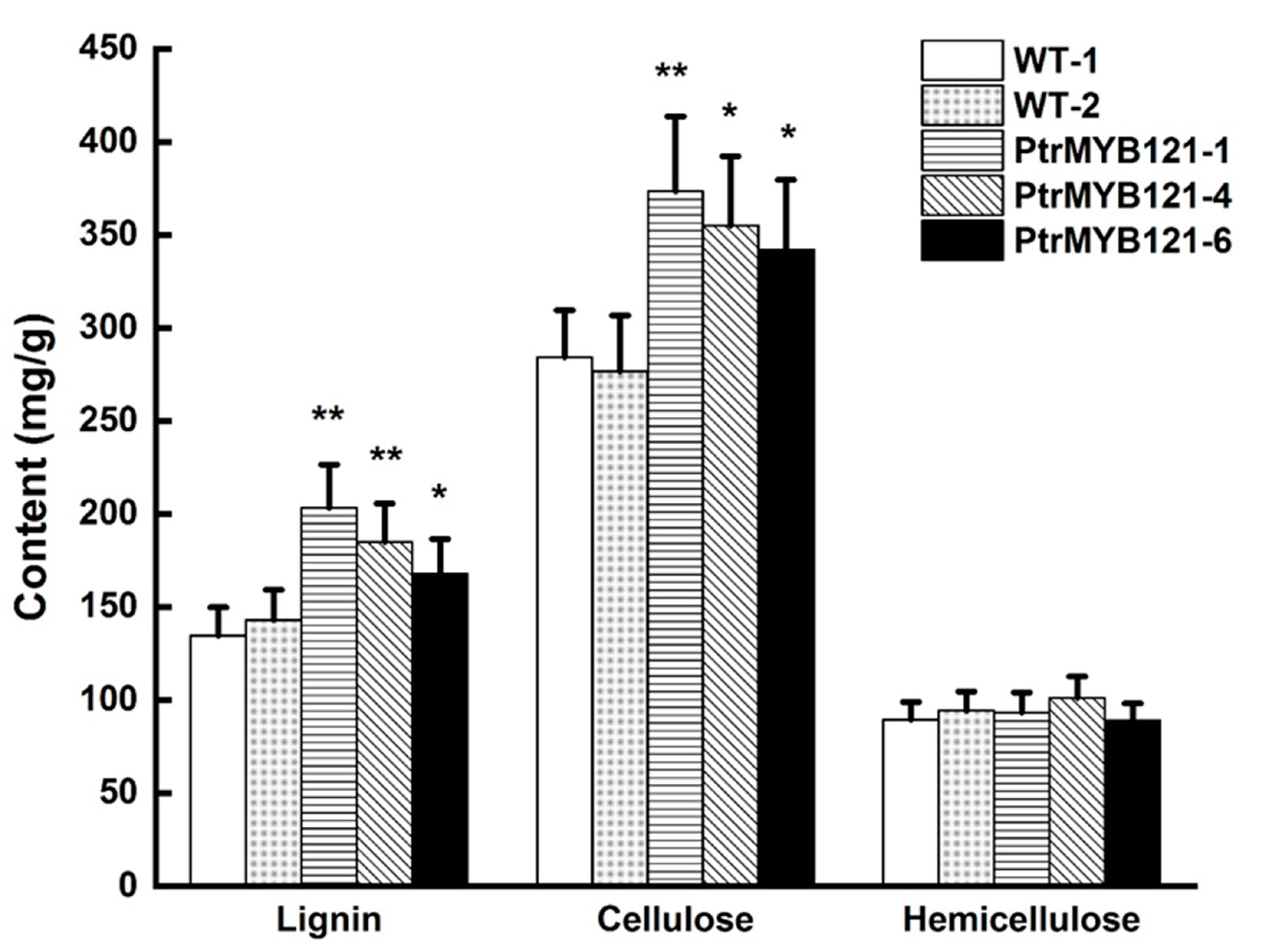
2.4. Overexpression of PtrMYB121 Promotes the Accumulation of Lignin and Cellulose
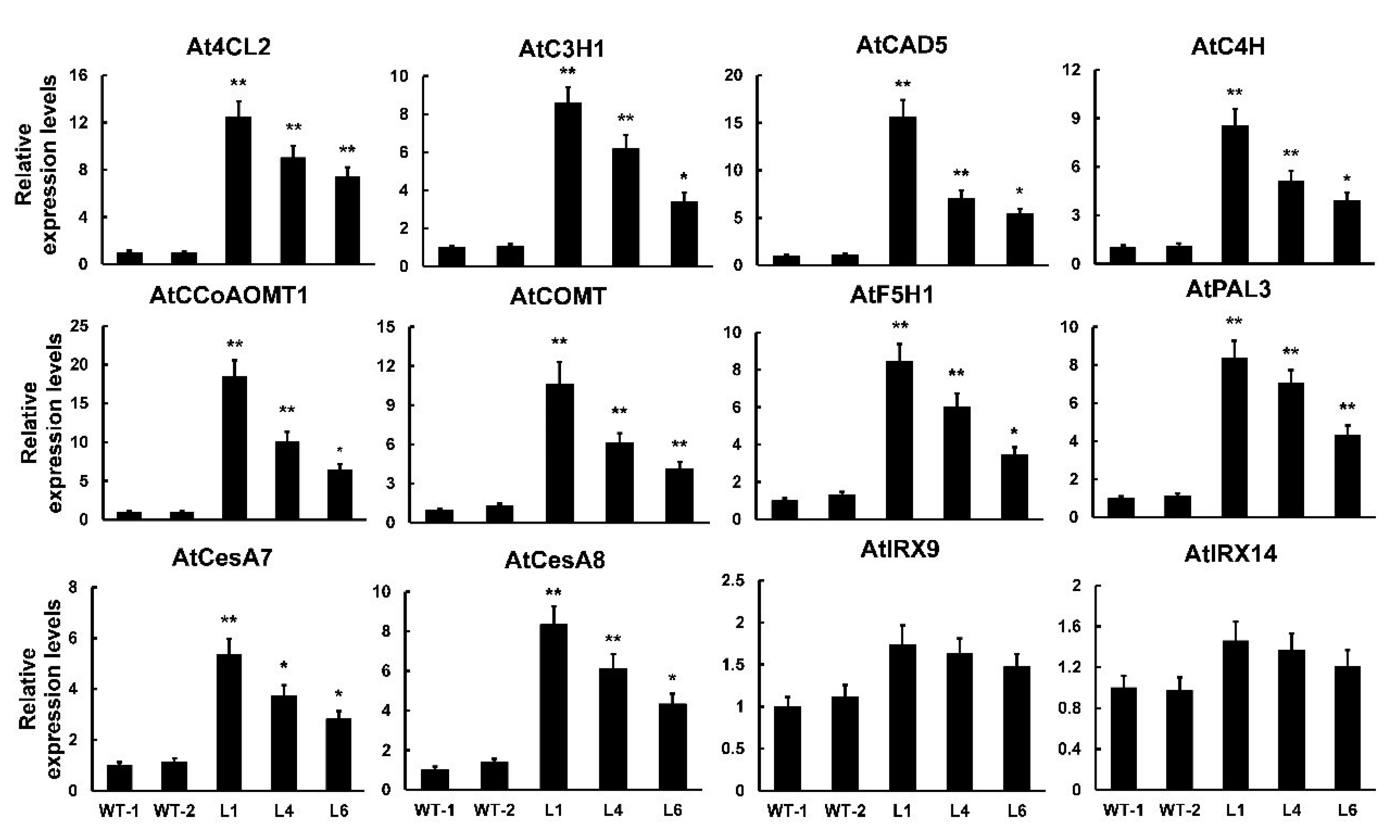
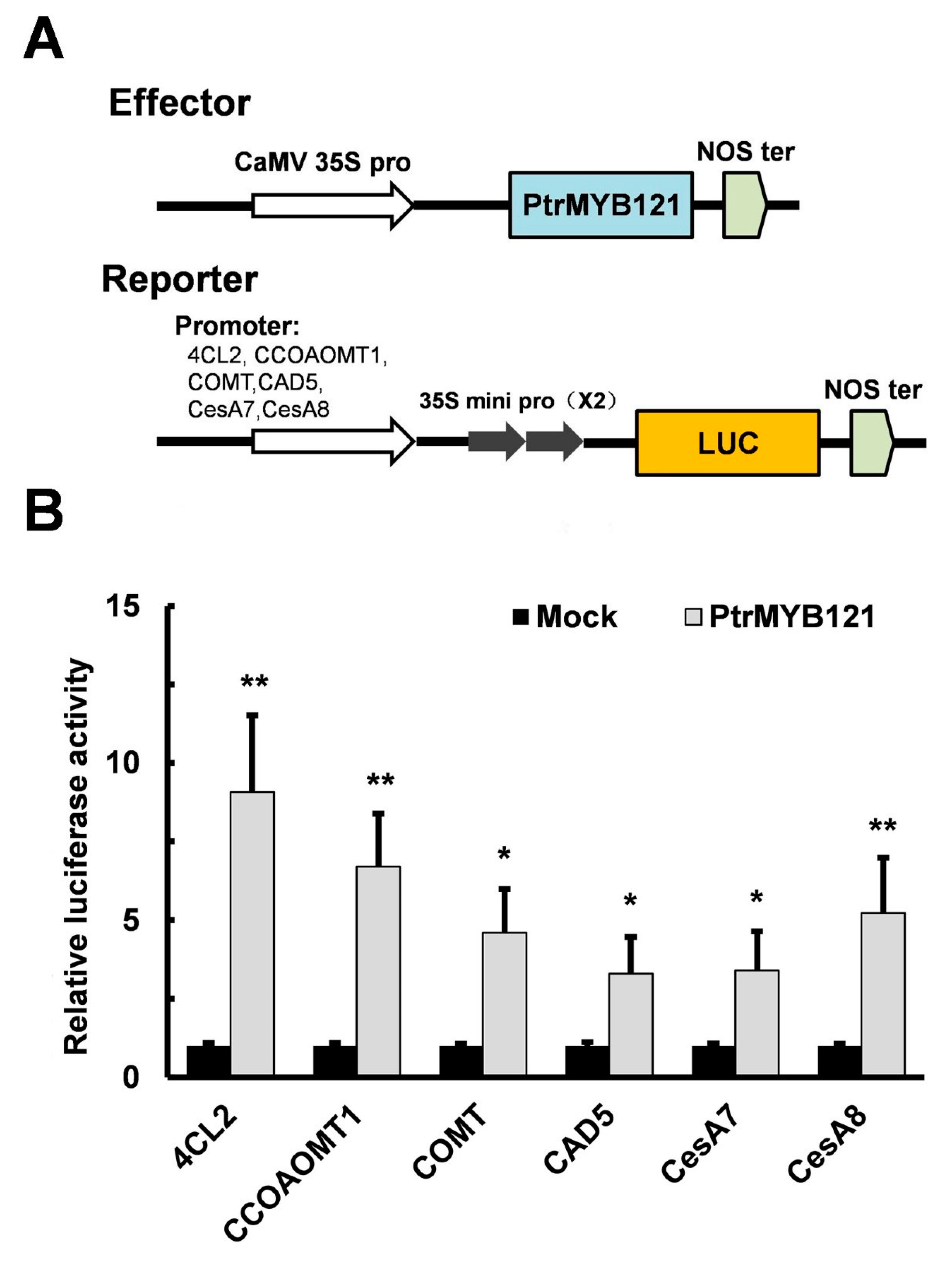
2.5. Overexpression of PtrMYB121 Activates the Expression of SCW Biosynthetic Genes
3. Discussion
4. Materials and Methods
4.1. Plant Materials
4.2. Phylogenetic Analysis
4.3. RNA Isolation and qRT-PCR Analysis
4.4. Cloning of Full-Length PtrMYB121 Coding Sequence (CDS)
4.5. Arabidopsis Transformation
4.6. Phenotypic Observation of PtrMYB121-Overexpressing Lines in Arabidopsis
4.7. Measurement of Lignin, Cellulose, and Hemicellulose Components in Arabidopsis
4.8. LUC Activity Assay
4.9. Statistical Analysis
4.10. GenBank Accession Numbers of Genes Used in this Study
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Carroll, A.; Somerville, C. Cellulosic biofuels. Annu. Rev. Plant Biol. 2009, 60, 165–182. [Google Scholar] [CrossRef] [PubMed]
- Plomion, C.; Leprovost, G.; Stokes, A. Wood formation in trees. Plant Physiol. 2001, 127, 1513–1523. [Google Scholar] [CrossRef] [PubMed]
- Zhong, R.; Cui, D.; Ye, Z.H. Secondary cell wall biosynthesis. New Phytol. 2019, 221, 1703–1723. [Google Scholar] [CrossRef]
- Richmond, T.A.; Somerville, C.R. The cellulose synthase superfamily. Plant Physiol. 2000, 124, 495–498. [Google Scholar] [CrossRef]
- Suzuki, S.; Li, L.; Sun, Y.-H.; Chiang, V.L. The Cellulose Synthase Gene Superfamily and Biochemical Functions of Xylem-Specific Cellulose Synthase-Like Genes in Populus trichocarpa. Plant Physiol. 2006, 142, 1233–1245. [Google Scholar] [CrossRef]
- Tanaka, K.; Murata, K.; Yamazaki, M.; Onosato, K.; Miyao, A.; Hirochika, H. Three distinct rice cellulose synthase catalytic subunit genes required for cellulose synthesis in the secondary wall. Plant Physiol. 2003, 133, 73–83. [Google Scholar] [CrossRef]
- Faraji, M.; Fonseca, L.L.; Escamilla-Treviño, L.; Dixon, R.A.; Voit, E.O. Computational inference of the structure and regulation of the lignin pathway in Panicum virgatum. Biotechnol. Biofuels 2015, 8, 151. [Google Scholar] [CrossRef]
- Zhao, Q.; Dixon, R.A. Transcriptional networks for lignin biosynthesis: More complex than we thought? Trends Plant Sci. 2011, 16, 227–233. [Google Scholar] [CrossRef] [PubMed]
- Geshi, N.; Harholt, J.; Sakuragi, Y.; Jensen, J.K.; Scheller, H.V. Authors. In Annual Plant Reviews; Ulvskov, P., Ed.; Wiley-Blackwell: West Sussex, UK, 2010; Volume 41, pp. 265–283. [Google Scholar] [CrossRef]
- Ade, C.P.; Bemm, F.; Dickson, J.M.; Walter, C.; Harris, P.J. Family 34 glycosyltransferase (GT34) genes and proteins in Pinus radiata (radiata pine) and Pinus taeda (loblolly pine). Plant J. 2014, 78, 305–318. [Google Scholar] [CrossRef] [PubMed]
- Ohtani, M.; Demura, T. The quest for transcriptional hubs of lignin biosynthesis: Beyond the NAC-MYB-gene regulatory network model. Curr. Opin. Biotechnol. 2019, 56, 82–87. [Google Scholar] [CrossRef]
- Kubo, M.; Udagawa, M.; Nishikubo, N.; Horiguchi, G.; Yamaguchi, M.; Ito, J.; Mimura, T.; Fukuda, H.; Demura, T. Transcription switches for protoxylem and metaxylem vessel formation. Genes Dev. 2005, 19, 1855–1860. [Google Scholar] [CrossRef] [PubMed]
- Reusche, M.; Thole, K.; Janz, D.; Truskina, J.; Rindfleisch, S.; Drübert, C.; Polle, A.; Lipka, V.; Teichmann, T. Verticillium infection triggers VASCULAR-RELATED NAC DOMAIN7-dependent de novo xylem formation and enhances drought tolerance in Arabidopsis. Plant Cell 2012, 24, 3823–3837. [Google Scholar] [CrossRef] [PubMed]
- Ohashi-Ito, K.; Oda, Y.; Fukuda, H. Arabidopsis VASCULAR-RELATED NAC-DOMAIN6 directly regulates the genes that govern programmed cell death and secondary wall formation during xylem differentiation. Plant Cell 2010, 22, 3461–3473. [Google Scholar] [CrossRef]
- Ohtani, M.; Nishikubo, N.; Xu, B.; Yamaguchi, M.; Mitsuda, N.; Goué, N.; Shi, F.; Ohme-Takagi, M.; Demura, T. A NAC domain protein family contributing to the regulation of wood formation in poplar. Plant J. 2011, 67, 499–512. [Google Scholar] [CrossRef] [PubMed]
- Zhong, R.; McCarthy, R.L.; Lee, C.; Ye, Z.H. Dissection of the transcriptional program regulating secondary wall biosynthesis during wood formation in poplar. Plant Physiol. 2011, 157, 1452–1468. [Google Scholar] [CrossRef]
- Duval, I.; Lachance, D.; Giguère, I.; Bomal, C.; Morency, M.J.; Pelletier, G.; Boyle, B.; MacKay, J.J.; Séguin, A. Large-scale screening of transcription factor-promoter interactions in spruce reveals a transcriptional network involved in vascular development. J. Exp. Bot. 2014, 65, 2319–2333. [Google Scholar] [CrossRef]
- Zhong, R.; Lee, C.; McCarthy, R.L.; Reeves, C.K.; Jones, E.G.; Ye, Z.H. Transcriptional activation of secondary wall biosynthesis by rice and maize NAC and MYB transcription factors. Plant Cell Physiol. 2011, 52, 1856–1871. [Google Scholar] [CrossRef]
- Zhao, Q.; Gallego-Giraldo, L.; Wang, H.; Zeng, Y.; Ding, S.Y.; Chen, F.; Dixon, R.A. An NAC transcription factor orchestrates multiple features of cell wall development in Medicago truncatula. Plant J. 2010, 63, 100–114. [Google Scholar] [CrossRef]
- Valdivia, E.R.; Herrera, M.T.; Gianzo, C.; Fidalgo, J.; Revilla, G.; Zarra, I.; Sampedro, J. Regulation of secondary wall synthesis and cell death by NAC transcription factors in the monocot Brachypodium distachyon. J. Exp. Bot. 2013, 64, 1333–1343. [Google Scholar] [CrossRef]
- Ko, J.-H.; Kim, W.-C.; Kim, J.Y.; Ahn, S.-J.; Han, K.-H. MYB46-Mediated Transcriptional Regulation of Secondary Wall Biosynthesis. Mol. Plant 2012, 5, 961–963. [Google Scholar] [CrossRef]
- McCarthy, R.L.; Zhong, R.; Ye, Z.H. MYB83 is a direct target of SND1 and acts redundantly with MYB46 in the regulation of secondary cell wall biosynthesis in Arabidopsis. Plant Cell Physiol. 2009, 50, 1950–1964. [Google Scholar] [CrossRef] [PubMed]
- Zhong, R.; Ye, Z.H. Regulation of cell wall biosynthesis. Curr. Opin. Plant Biol. 2007, 10, 564–572. [Google Scholar] [CrossRef] [PubMed]
- Baxter, H.; Poovaiah, C.; Yee, K.; Mazarei, M.; Rodriguez, M.; Thompson, O.; Shen, H.; Turner, G.; Decker, S.; Sykes, R.; et al. Field Evaluation of Transgenic Switchgrass Plants Overexpressing PvMYB4 for Reduced Biomass Recalcitrance. BioEnergy Res. 2015, 8, 910–921. [Google Scholar] [CrossRef]
- Shen, H.; He, X.; Poovaiah, C.R.; Wuddineh, W.A.; Ma, J.; Mann, D.G.; Wang, H.; Jackson, L.; Tang, Y.; Stewart, C.N., Jr.; et al. Functional characterization of the switchgrass (Panicum virgatum) R2R3-MYB transcription factor PvMYB4 for improvement of lignocellulosic feedstocks. New Phytol. 2012, 193, 121–136. [Google Scholar] [CrossRef]
- Romano, J.M.; Dubos, C.; Prouse, M.B.; Wilkins, O.; Hong, H.; Poole, M.; Kang, K.Y.; Li, E.; Douglas, C.J.; Western, T.L.; et al. AtMYB61, an R2R3-MYB transcription factor, functions as a pleiotropic regulator via a small gene network. New Phytol. 2012, 195, 774–786. [Google Scholar] [CrossRef] [PubMed]
- Newman, L.J.; Perazza, D.E.; Juda, L.; Campbell, M.M. Involvement of the R2R3-MYB, AtMYB61, in the ectopic lignification and dark-photomorphogenic components of the det3 mutant phenotype. Plant J. 2004, 37, 239–250. [Google Scholar] [CrossRef]
- Matías-Hernández, L.; Jiang, W.; Yang, K.; Tang, K.; Brodelius, P.E.; Pelaz, S. AaMYB1 and its orthologue AtMYB61 affect terpene metabolism and trichome development in Artemisia annua and Arabidopsis thaliana. Plant J. 2017, 90, 520–534. [Google Scholar] [CrossRef]
- Liang, Y.K.; Dubos, C.; Dodd, I.C.; Holroyd, G.H.; Hetherington, A.M.; Campbell, M.M. AtMYB61, an R2R3-MYB transcription factor controlling stomatal aperture in Arabidopsis thaliana. Curr. Biol. 2005, 15, 1201–1206. [Google Scholar] [CrossRef]
- Penfield, S.; Meissner, R.C.; Shoue, D.A.; Carpita, N.C.; Bevan, M.W. MYB61 is required for mucilage deposition and extrusion in the Arabidopsis seed coat. Plant Cell 2001, 13, 2777–2791. [Google Scholar] [CrossRef]
- Tuskan, G.A.; Difazio, S.; Jansson, S.; Bohlmann, J.; Grigoriev, I.; Hellsten, U.; Putnam, N.; Ralph, S.; Rombauts, S.; Salamov, A.; et al. The genome of black cottonwood, Populus trichocarpa (Torr. & Gray). Science 2006, 313, 1596–1604. [Google Scholar] [CrossRef]
- Ma, T.; Wang, J.; Zhou, G.; Yue, Z.; Hu, Q.; Chen, Y.; Liu, B.; Qiu, Q.; Wang, Z.; Zhang, J.; et al. Genomic insights into salt adaptation in a desert poplar. Nat. Commun. 2013, 4, 2797. [Google Scholar] [CrossRef] [PubMed]
- Ma, T.; Wang, K.; Hu, Q.; Xi, Z.; Wan, D.; Wang, Q.; Feng, J.; Jiang, D.; Ahani, H.; Abbott, R.J.; et al. Ancient polymorphisms and divergence hitchhiking contribute to genomic islands of divergence within a poplar species complex. Proc. Natl. Acad. Sci. USA 2018, 115, E236–E243. [Google Scholar] [CrossRef]
- Chai, G.; Wang, Z.; Tang, X.; Yu, L.; Qi, G.; Wang, D.; Yan, X.; Kong, Y.; Zhou, G. R2R3-MYB gene pairs in Populus: Evolution and contribution to secondary wall formation and flowering time. J. Exp. Bot. 2014, 65, 4255–4269. [Google Scholar] [CrossRef] [PubMed]
- Xu, C.; Fu, X.; Liu, R.; Guo, L.; Ran, L.; Li, C.; Tian, Q.; Jiao, B.; Wang, B.; Luo, K. PtoMYB170 positively regulates lignin deposition during wood formation in poplar and confers drought tolerance in transgenic Arabidopsis. Tree Physiol. 2017, 37, 1713–1726. [Google Scholar] [CrossRef]
- Tian, Q.; Wang, X.; Li, C.; Lu, W.; Yang, L.; Jiang, Y.; Luo, K. Functional Characterization of the Poplar R2R3-MYB Transcription Factor PtoMYB216 Involved in the Regulation of Lignin Biosynthesis during Wood Formation. PLoS ONE 2013, 8, e76369. [Google Scholar] [CrossRef] [PubMed]
- Iiyama, K.; Wallis, A.F.A. An improved acetyl bromide procedure for determining lignin in woods and wood pulps. Wood Sci. Technol. 1988, 22, 271–280. [Google Scholar] [CrossRef]
- Van Soest, P.J.; Wine, R.H. Use of detergents in the analysis of fibrous feeds. IV. Determination of plant cell-wall constituents. J. Assoc. Off. Anal. Chem. 1967, 50, 50–55. [Google Scholar] [CrossRef]
- Zhong, R.; McCarthy, R.L.; Haghighat, M.; Ye, Z.H. The poplar MYB master switches bind to the SMRE site and activate the secondary wall biosynthetic program during wood formation. PLoS ONE 2013, 8, e69219. [Google Scholar] [CrossRef]
- Li, Z.; Omranian, N.; Neumetzler, L.; Wang, T.; Herter, T.; Usadel, B.; Demura, T.; Giavalisco, P.; Nikoloski, Z.; Persson, S. A Transcriptional and Metabolic Framework for Secondary Wall Formation in Arabidopsis. Plant Physiol. 2016, 172, 1334–1351. [Google Scholar] [CrossRef]
- Lepelley, M.; Cheminade, G.; Tremillon, N.; Simkin, A.; Caillet, V.; McCarthy, J. Chlorogenic acid synthesis in coffee: An analysis of CGA content and real-time RT-PCR expression of HCT, HQT, C3H1, and CCoAOMT1 genes during grain development in C. canephora. Plant Sci. 2007, 172, 978–996. [Google Scholar] [CrossRef]
- Perdereau, A.C.; Douglas, G.C.; Hodkinson, T.R.; Kelleher, C.T. High levels of variation in Salix lignocellulose genes revealed using poplar genomic resources. Biotechnol. Biofuels 2013, 6, 114. [Google Scholar] [CrossRef]
- Hill, J.L., Jr.; Hammudi, M.B.; Tien, M. The Arabidopsis cellulose synthase complex: A proposed hexamer of CESA trimers in an equimolar stoichiometry. Plant Cell 2014, 26, 4834–4842. [Google Scholar] [CrossRef] [PubMed]
- Wu, A.M.; Hörnblad, E.; Voxeur, A.; Gerber, L.; Rihouey, C.; Lerouge, P.; Marchant, A. Analysis of the Arabidopsis IRX9/IRX9-L and IRX14/IRX14-L Pairs of Glycosyltransferase Genes Reveals Critical Contributions to Biosynthesis of the Hemicellulose Glucuronoxylan. Plant Physiol. 2010, 153, 542–554. [Google Scholar] [CrossRef] [PubMed]
- Zhou, J.; Lee, C.; Zhong, R.; Ye, Z.H. MYB58 and MYB63 are transcriptional activators of the lignin biosynthetic pathway during secondary cell wall formation in Arabidopsis. Plant Cell 2009, 21, 248–266. [Google Scholar] [CrossRef]
- Zhong, R.; Lee, C.; Zhou, J.; McCarthy, R.L.; Ye, Z.H. A battery of transcription factors involved in the regulation of secondary cell wall biosynthesis in Arabidopsis. Plant Cell 2008, 20, 2763–2782. [Google Scholar] [CrossRef]
- Xie, M.; Zhang, J.; Tschaplinski, T.; Tuskan, G.; Chen, J.-G.; Muchero, W. Regulation of Lignin Biosynthesis and Its Role in Growth-Defense Tradeoffs. Front. Plant Sci. 2018, 9, 1427. [Google Scholar] [CrossRef]
- Legay, S.; Sivadon, P.; Blervacq, A.S.; Pavy, N.; Baghdady, A.; Tremblay, L.; Levasseur, C.; Ladouce, N.; Lapierre, C.; Séguin, A.; et al. EgMYB1, an R2R3 MYB transcription factor from eucalyptus negatively regulates secondary cell wall formation in Arabidopsis and poplar. New Phytol. 2010, 188, 774–786. [Google Scholar] [CrossRef] [PubMed]
- Zhao, K.; Bartley, L. Comparative genomic analysis of the R2R3 MYB secondary cell wall regulators of Arabidopsis, poplar, rice, maize, and switchgrass. BMC Plant Biol. 2014, 14, 135. [Google Scholar] [CrossRef] [PubMed]
- Chen, H.; Wang, J.; Liu, H.; Li, H.; Lin, Y.-C.; Shi, R.; Yang, C.; Gao, J.; Zhou, C.; Li, Q.; et al. Hierarchical Transcription-Factor and Chromatin Binding Network for Wood Formation in Populus trichocarpa. Plant Cell 2019, 31, 602–626. [Google Scholar] [CrossRef] [PubMed]
- Scully, E.D.; Gries, T.; Sarath, G.; Palmer, N.A.; Baird, L.; Serapiglia, M.J.; Dien, B.S.; Boateng, A.A.; Ge, Z.; Funnell-Harris, D.L.; et al. Overexpression of SbMyb60 impacts phenylpropanoid biosynthesis and alters secondary cell wall composition in Sorghum bicolor. Plant J. 2016, 85, 378–395. [Google Scholar] [CrossRef] [PubMed]
- Ye, Y.; Liu, B.; Zhao, M.; Wu, K.; Cheng, W.; Chen, X.; Liu, Q.; Liu, Z.; Fu, X.; Wu, Y. CEF1/OsMYB103L is involved in GA-mediated regulation of secondary wall biosynthesis in rice. Plant Mol. Biol. 2015, 89, 385–401. [Google Scholar] [CrossRef] [PubMed]
- Handakumbura, P.; Brow, K.; Whitney, I.; Zhao, K.; Sanguinet, K.; Lee, S.; Olins, J.; Romero-Gamboa, S.; Harrington, M.; Bascom, C.; et al. SECONDARY WALL ASSOCIATED MYB1 is a positive regulator of secondary cell wall thickening in Brachypodium distachyon and is not found in the Brassicaceae. Plant J. 2018, 96, 532–545. [Google Scholar] [CrossRef]
- McCarthy, R.L.; Zhong, R.; Fowler, S.; Lyskowski, D.; Piyasena, H.; Carleton, K.; Spicer, C.; Ye, Z.H. The poplar MYB transcription factors, PtrMYB3 and PtrMYB20, are involved in the regulation of secondary wall biosynthesis. Plant Cell Physiol. 2010, 51, 1084–1090. [Google Scholar] [CrossRef]
- Li, C.; Wang, X.; Ran, L.; Tian, Q.; Fan, D.; Luo, K. PtoMYB92 is a Transcriptional Activator of the Lignin Biosynthetic Pathway During Secondary Cell Wall Formation in Populus tomentosa. Plant Cell Physiol. 2015, 56, 2436–2446. [Google Scholar] [CrossRef] [PubMed]
- Yang, L.; Zhao, X.; Ran, L.; Li, C.; Fan, D.; Luo, K. PtoMYB156 is involved in negative regulation of phenylpropanoid metabolism and secondary cell wall biosynthesis during wood formation in poplar. Sci. Rep. 2017, 7, 41209. [Google Scholar] [CrossRef] [PubMed]
- Tang, X.; Zhuang, Y.; Qi, G.; Wang, D.; Liu, H.; Wang, K.; Chai, G.; Zhou, G. Poplar PdMYB221 is involved in the direct and indirect regulation of secondary wall biosynthesis during wood formation. Sci. Rep. 2015, 5, 12240. [Google Scholar] [CrossRef]
- Rueda-López, M.; Pascual, M.B.; Pallero, M.; Henao, L.M.; Lasa, B.; Jauregui, I.; Aparicio-Tejo, P.M.; Cánovas, F.M.; Ávila, C. Overexpression of a pine Dof transcription factor in hybrid poplars: A comparative study in trees growing under controlled and natural conditions. PLoS ONE 2017, 12, e0174748. [Google Scholar] [CrossRef]
- Ye, Q.; Liu, X.; Bian, W.; Zhang, Z.; Zhang, H. Over-expression of transcription factor ARK1 gene leads to down-regulation of lignin synthesis related genes in hybrid poplar ‘717’. Sci. Rep. 2020, 10, 8549. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Y.; Xueqin, S.; Zhou, H.; Wei, K.; Jiang, C.; Wang, J.; Cao, Y.; Tang, F.; Zhao, S.; Lu, M.-Z. KNAT2/6b, a class I KNOX gene, impedes xylem differentiation by regulating NAC domain transcription factors in poplar. New Phytol. 2019, 225, 1531–1544. [Google Scholar] [CrossRef]
- Li, S.; Zhang, Y.; Xin, X.; Ding, C.; Lv, F.; Mo, W.; Xia, Y.; Wang, S.; Cai, J.; Sun, L. The Osmotin-Like Protein Gene PdOLP1 Is Involved in Secondary Cell Wall Biosynthesis during Wood Formation in Poplar. Int. J. Mol. Sci. 2020, 21, 3993. [Google Scholar] [CrossRef]
- Yang, J.; Wang, H. Molecular Mechanisms for Vascular Development and Secondary Cell Wall Formation. Front. Plant Sci. 2016, 7, 356. [Google Scholar] [CrossRef] [PubMed]
- Li, C.; Ma, X.; Yu, H.; Fu, Y.; Luo, K. Ectopic Expression of PtoMYB74 in Poplar and Arabidopsis Promotes Secondary Cell Wall Formation. Front. Plant Sci. 2018, 9, 1262. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular Evolutionary Genetics Analysis Version 7.0 for Bigger Datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef] [PubMed]
- Altschul, S.F.; Gish, W.; Miller, W.; Myers, E.W.; Lipman, D.J. Basic local alignment search tool. J. Mol. Biol. 1990, 215, 403–410. [Google Scholar] [CrossRef]
- Xiuren, Z.; Henriques, R.; Lin, S.-S.; Niu, Q.-W.; Nam-Hai, C. Agrobacterium-mediated transformation of Arabidopsis thaliana using the floral dip method. Nat. Protoc. 2006, 2, 641. [Google Scholar] [CrossRef]
- Sparkes, I.A.; Runions, J.; Kearns, A.; Hawes, C. Rapid, transient expression of fluorescent fusion proteins in tobacco plants and generation of stably transformed plants. Nat. Protoc. 2006, 1, 2019–2025. [Google Scholar] [CrossRef] [PubMed]

| Line | Samples | Cell Layer (n) | Radial Width (μm) |
|---|---|---|---|
| WT-1 | 1 | 6.78 ± 0.81 a | 439.72 ± 28.79 a |
| 2 | 7.11 ± 0.94 a | 461.77 ± 33.47 a | |
| 3 | 6.43 ± 0.87 a | 422.36 ± 30.16 a | |
| PtrMYB121-L1 | 1 | 13.76 ± 1.57 b | 657.42 ± 53.34 b |
| 2 | 14.20 ± 1.72 b | 693.17 ± 58.38 b | |
| 3 | 13.49 ± 1.55 b | 638.84 ± 49.67 b |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Liu, Y.; Man, J.; Wang, Y.; Yuan, C.; Shi, Y.; Liu, B.; Hu, X.; Wu, S.; Zhang, T.; Lian, C. Overexpression of PtrMYB121 Positively Regulates the Formation of Secondary Cell Wall in Arabidopsis thaliana. Int. J. Mol. Sci. 2020, 21, 7734. https://doi.org/10.3390/ijms21207734
Liu Y, Man J, Wang Y, Yuan C, Shi Y, Liu B, Hu X, Wu S, Zhang T, Lian C. Overexpression of PtrMYB121 Positively Regulates the Formation of Secondary Cell Wall in Arabidopsis thaliana. International Journal of Molecular Sciences. 2020; 21(20):7734. https://doi.org/10.3390/ijms21207734
Chicago/Turabian StyleLiu, Ying, Jiayin Man, Yinghao Wang, Chao Yuan, Yuyu Shi, Bobin Liu, Xia Hu, Songqing Wu, Taoxiang Zhang, and Chunlan Lian. 2020. "Overexpression of PtrMYB121 Positively Regulates the Formation of Secondary Cell Wall in Arabidopsis thaliana" International Journal of Molecular Sciences 21, no. 20: 7734. https://doi.org/10.3390/ijms21207734
APA StyleLiu, Y., Man, J., Wang, Y., Yuan, C., Shi, Y., Liu, B., Hu, X., Wu, S., Zhang, T., & Lian, C. (2020). Overexpression of PtrMYB121 Positively Regulates the Formation of Secondary Cell Wall in Arabidopsis thaliana. International Journal of Molecular Sciences, 21(20), 7734. https://doi.org/10.3390/ijms21207734






