Antineoplastic Activity of Chrysin against Human Hepatocellular Carcinoma: New Insight on GPC3/SULF2 Axis and lncRNA-AF085935 Expression
Abstract
1. Introduction
2. Results
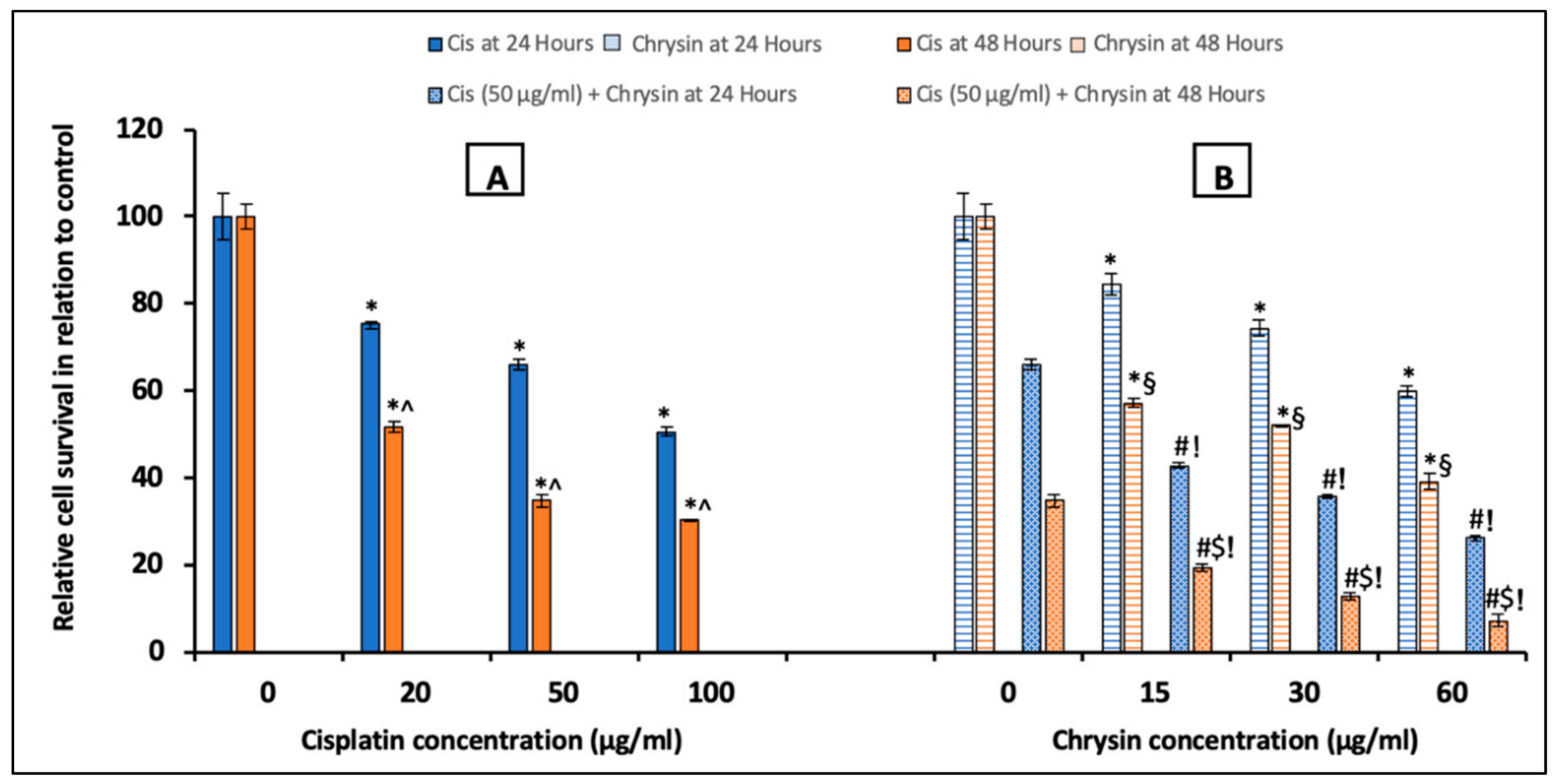
2.1. Chrysin Combined with Cisplatin Decreased Cell Survival in HepG2 Cells
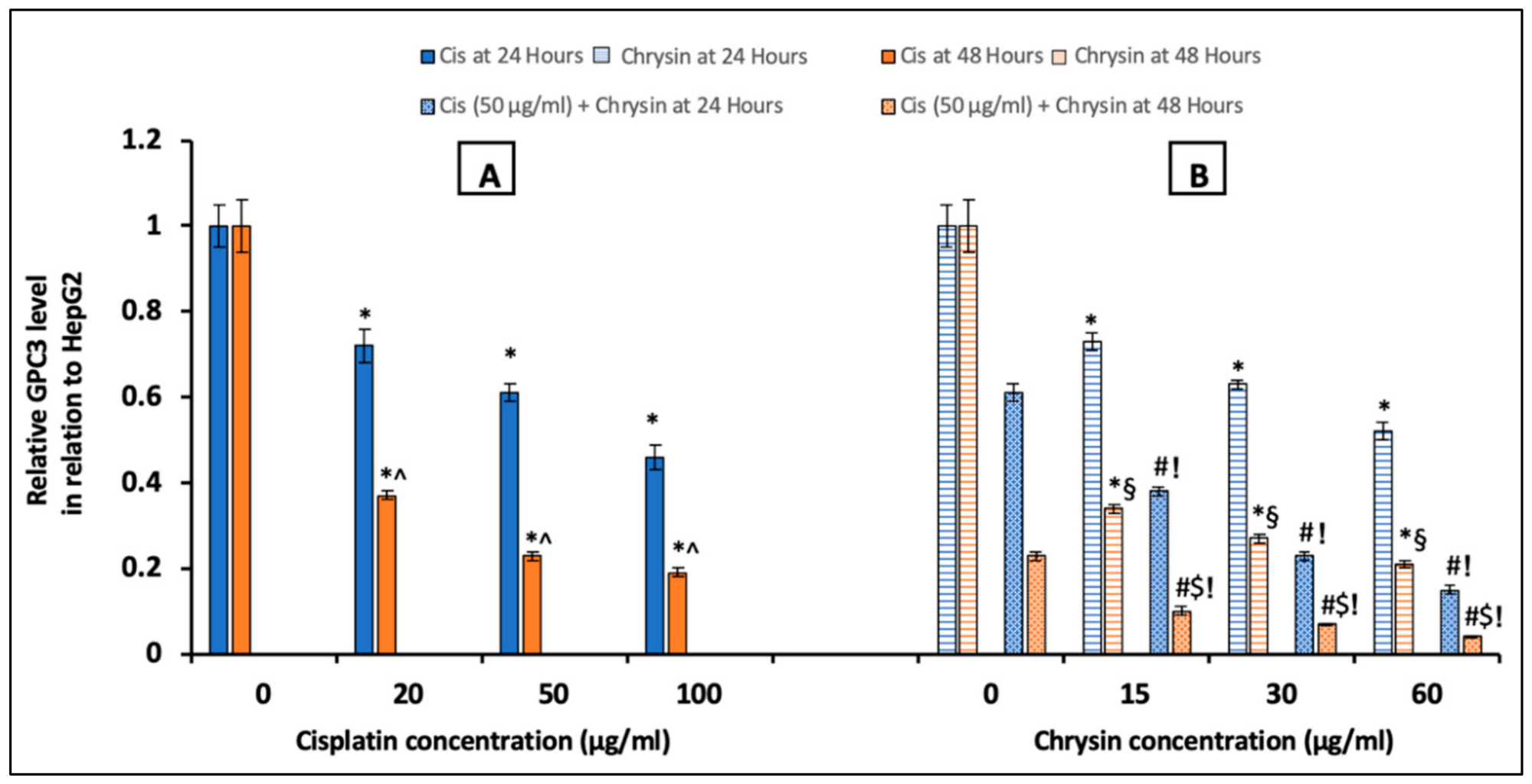
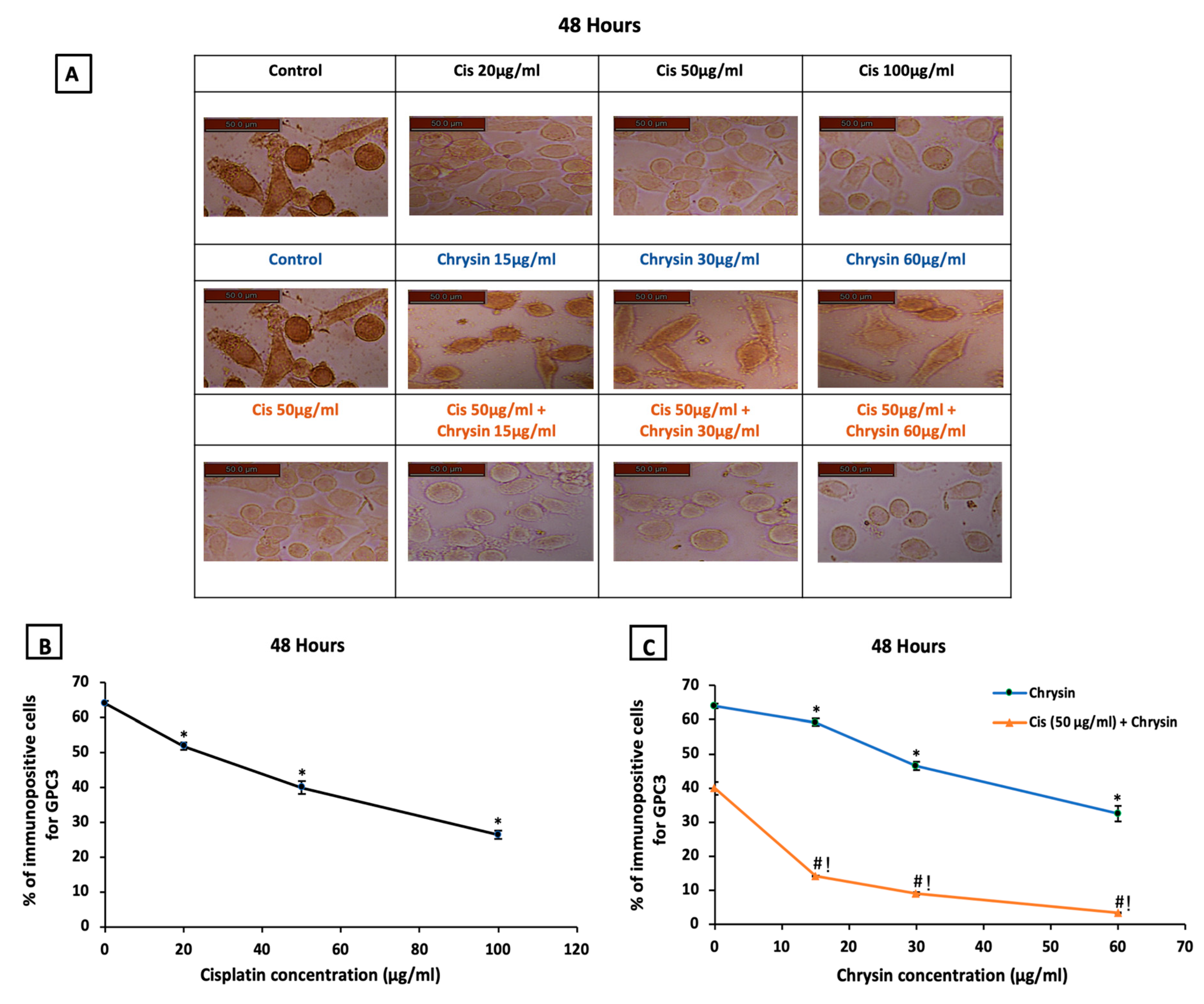
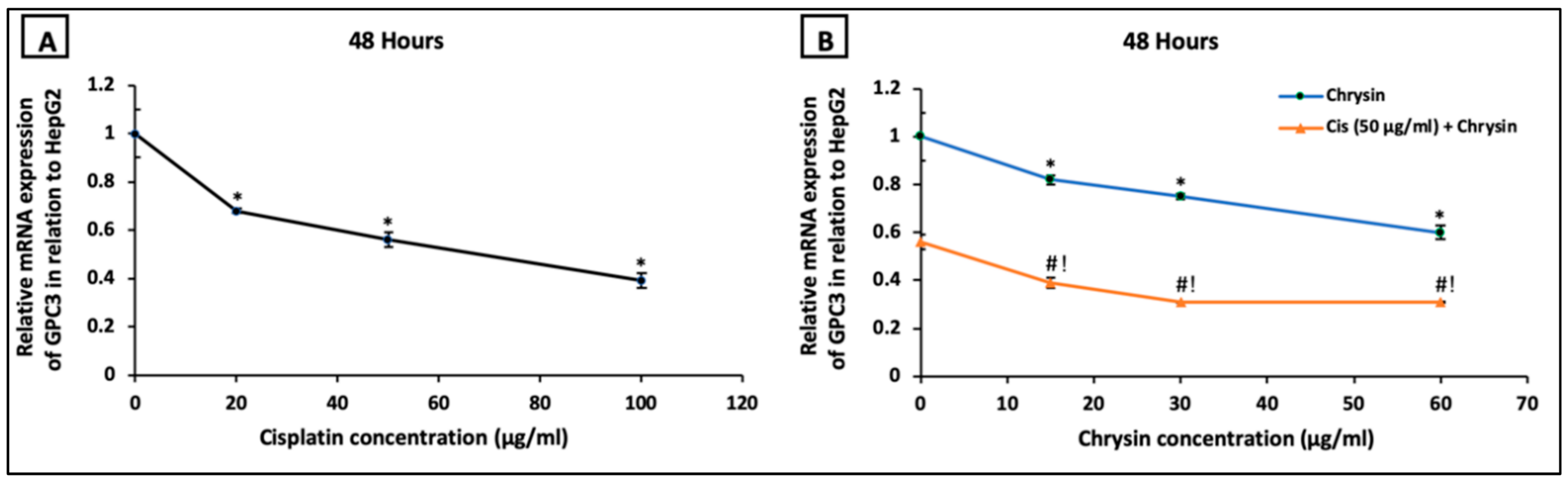
2.2. Chrysin Combined with Cisplatin Decreased Glypican-3 (GPC3) Protein and mRNA Expressions in HepG2 Cells
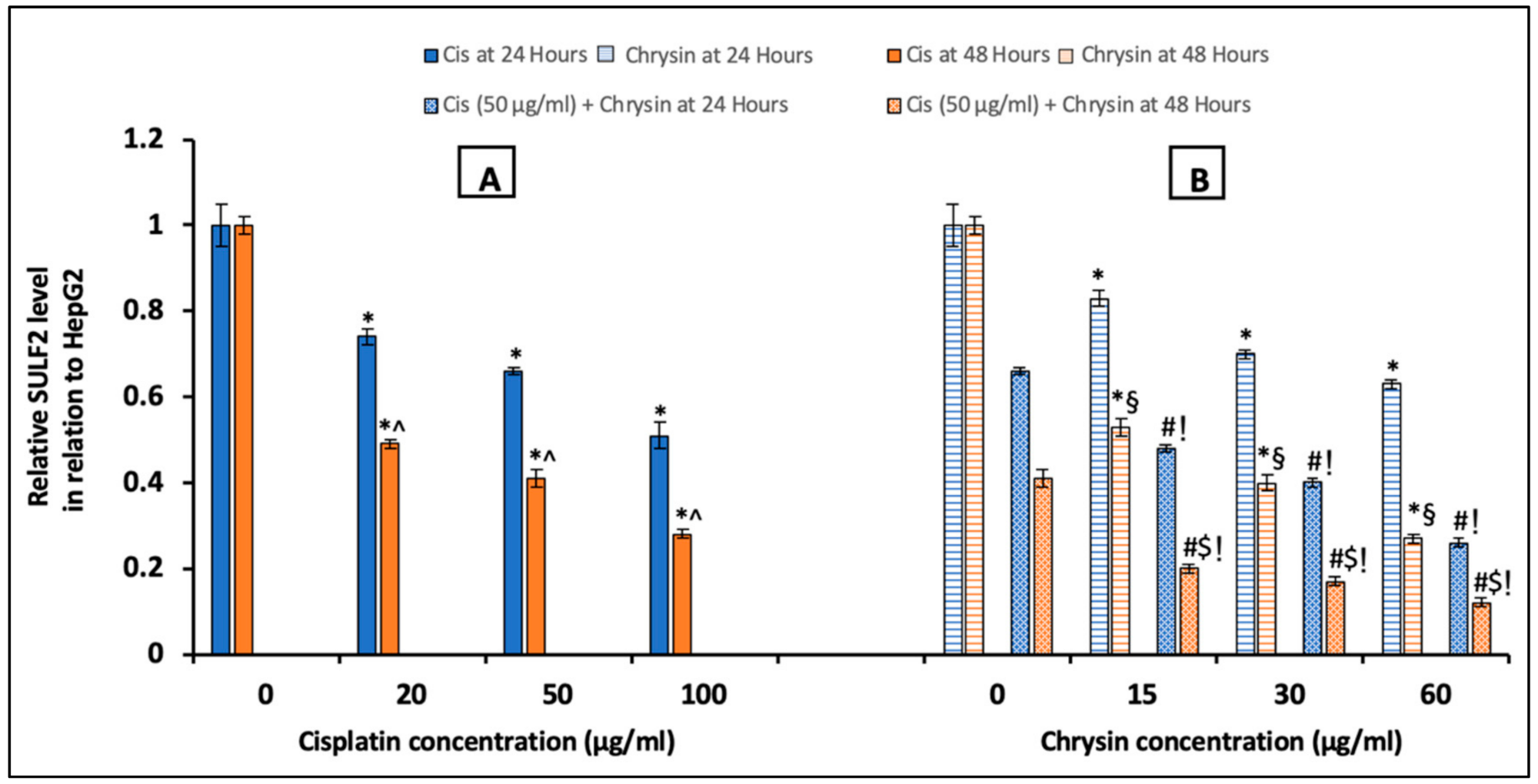
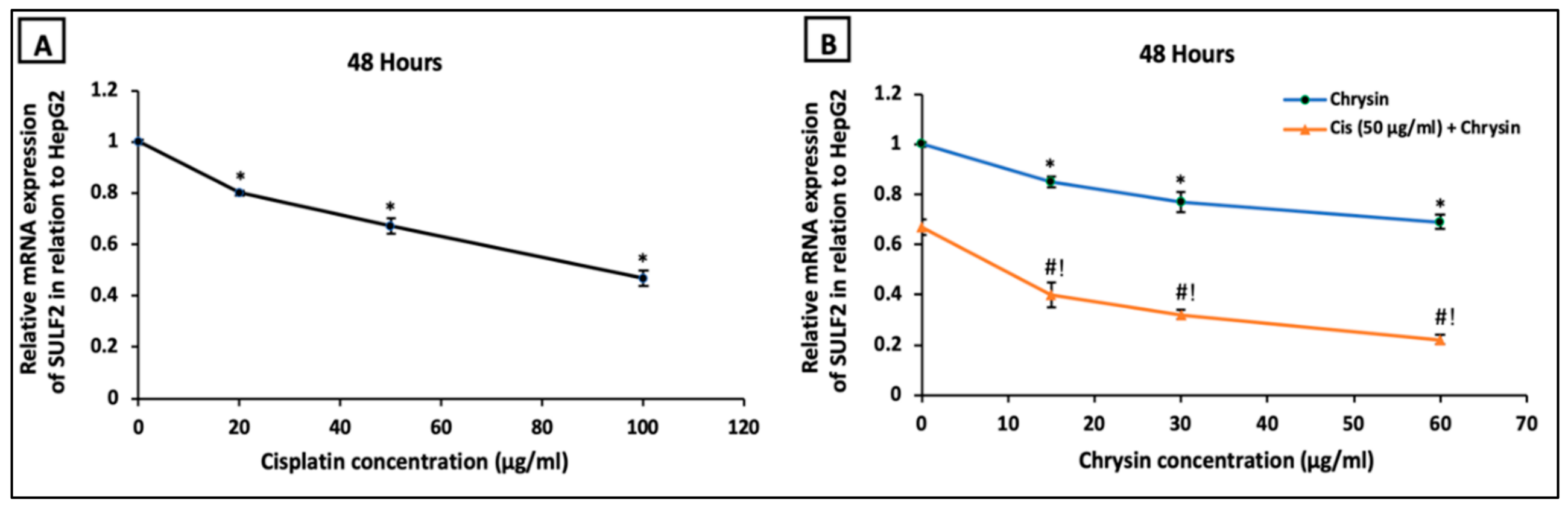
2.3. Chrysin Combined with Cisplatin Decreased Sulfatase-2 (SULF2) Protein and mRNA Expressions in HepG2 Cells
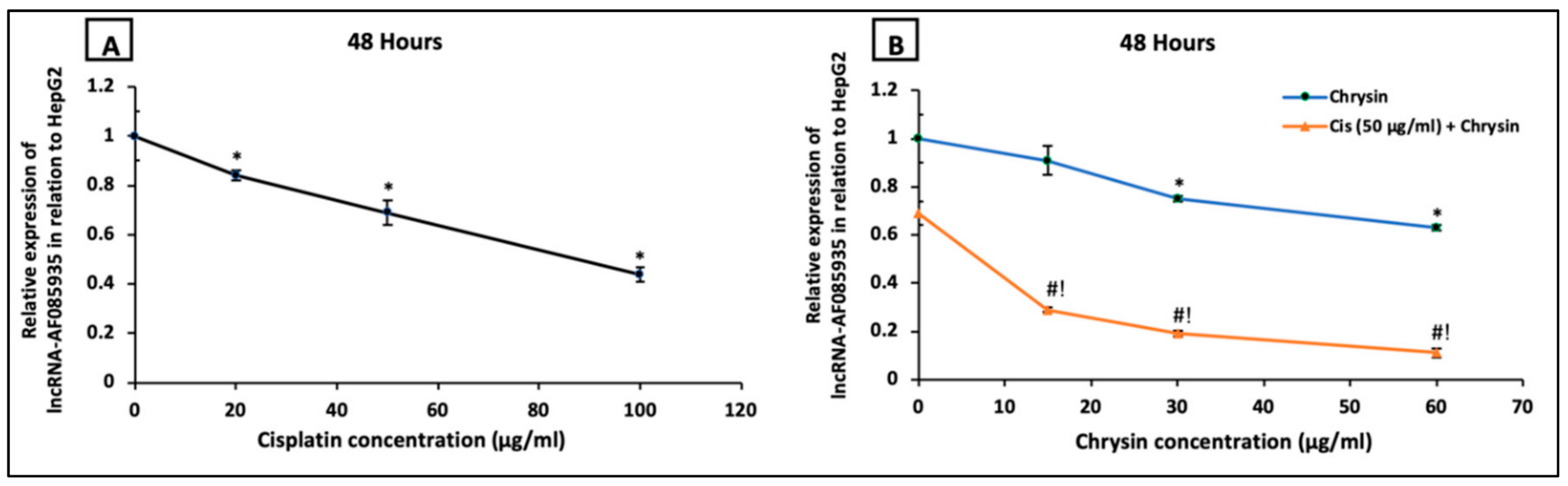
2.4. Chrysin Combined with Cisplatin Decreased lncRNA-AF085935 Expression in HepG2 Cells
3. Discussion
4. Materials and methods
4.1. Drugs
4.2. Cell Culture
4.3. MTT Assay for Cell Viability
4.4. Immunocytochemistry (ICC) for the Detection of GPC3 Protein Expression
4.5. Enzyme-Linked Immunosorbent Assay (ELISA) for the Determination of GPC3/SULF2 Protein Levels
4.6. Quantitative Real-Time Polymerase Chain Reaction (qPCR) for the Estimation of GPC3/SULF2 mRNA Expression and lncRNA-AF085935 Expression
4.7. Data Analysis
Author Contributions
Funding
Conflicts of Interest
References
- Zhu, R.X.; Seto, W.-K.; Lai, C.-L.; Yuen, M.-F. Epidemiology of hepatocellular carcinoma in the Asia-Pacific region. Gut Liver 2016, 10, 332. [Google Scholar] [CrossRef]
- Li, H.; Jiang, D.; Zhang, L.; Wu, J. Inhibition of tumor growth of human hepatocellular carcinoma HepG2 cells in a nude mouse xenograft model by the total flavonoids from Arachniodes exilis. Evid. Based Complement. Altern. Med. 2017. [Google Scholar] [CrossRef]
- Wu, Y.; Liu, H.; Ding, H. GPC-3 in hepatocellular carcinoma: Current perspectives. J. Hepatocell. Carcinoma 2016, 3, 63. [Google Scholar] [CrossRef] [PubMed]
- Zhong, C.; Qiu, S.; Li, J.; Shen, J.; Zu, Y.; Shi, J.; Sui, G. Ellagic acid synergistically potentiates inhibitory activities of chemotherapeutic agents to human hepatocellular carcinoma. Phytomedicine 2019, 59, 152921. [Google Scholar] [CrossRef] [PubMed]
- Tayel, A.; El Galil, K.H.A.; Ebrahim, M.A.; Ibrahim, A.S.; El-Gayar, A.M.; Al-Gayyar, M.M. Suramin inhibits hepatic tissue damage in hepatocellular carcinoma through deactivation of heparanase enzyme. Eur. J. Pharmacol. 2014, 728, 151–160. [Google Scholar] [CrossRef] [PubMed]
- Roy, G.; Guan, S.; Liu, H.; Zhang, L. Rotundic acid induces DNA damage and cell death in hepatocellular carcinoma through AKT/mTOR and MAPK pathways. Front. Oncol. 2019, 9, 545. [Google Scholar] [CrossRef]
- Sherif, I.O.; Al-Gayyar, M.M. Oleuropein potentiates anti-tumor activity of cisplatin against HepG2 through affecting proNGF/NGF balance. Life Sci. 2018, 198, 87–93. [Google Scholar] [CrossRef]
- Zhang, L.; Jiang, G.; Yao, F.; He, Y.; Liang, G.; Zhang, Y.; Hu, B.; Wu, Y.; Li, Y.; Liu, H. Growth inhibition and apoptosis induced by osthole, a natural coumarin, in hepatocellular carcinoma. PLoS ONE 2012, 7, e37865. [Google Scholar] [CrossRef]
- Liu, Z.; Peng, Q.; Li, Y.; Gao, Y. Resveratrol enhances cisplatin-induced apoptosis in human hepatoma cells via glutamine metabolism inhibition. BMB Rep. 2018, 51, 474. [Google Scholar] [CrossRef]
- Wu, Z.; Wu, J.; Fang, P.; Kan, S. Puerarin increases the chemosensitivity of hepatocellular carcinoma cells. Oncol. Lett. 2017, 14, 3006–3010. [Google Scholar] [CrossRef]
- Li, M.; Chen, J.; Yu, X.; Xu, S.; Li, D.; Zheng, Q.; Yin, Y. Myricetin suppresses the propagation of hepatocellular carcinoma via down-regulating expression of YAP. Cells 2019, 8, 358. [Google Scholar] [CrossRef] [PubMed]
- Xu, D.; Jin, J.; Yu, H.; Zhao, Z.; Ma, D.; Zhang, C.; Jiang, H. Chrysin inhibited tumor glycolysis and induced apoptosis in hepatocellular carcinoma by targeting hexokinase-2. J. Exp. Clin. Cancer Res. 2017, 36, 44. [Google Scholar] [CrossRef] [PubMed]
- Oliveira, G.; Ferraz, E.; Souza, A.; Lourenco, R.; Oliveira, D.P.; Dorta, D.J. Evaluation of the mutagenic activity of chrysin, a flavonoid inhibitor of the aromatization process. J. Toxicol. Environ. Health Part A 2012, 75, 1000–1011. [Google Scholar] [CrossRef] [PubMed]
- Gao, A.-M.; Ke, Z.-P.; Shi, F.; Sun, G.-C.; Chen, H. Chrysin enhances sensitivity of BEL-7402/ADM cells to doxorubicin by suppressing PI3K/Akt/Nrf2 and ERK/Nrf2 pathway. Chem. Biol. Interact. 2013, 206, 100–108. [Google Scholar] [CrossRef]
- Mohammadian, F.; Pilehvar-Soltanahmadi, Y.; Alipour, S.; Dadashpour, M.; Zarghami, N. Chrysin alters microRNAs expression levels in gastric cancer cells: Possible molecular mechanism. Drug Res. 2017, 67, 509–514. [Google Scholar] [CrossRef]
- Samarghandian, S.; Azimi-Nezhad, M.; Borji, A.; Hasanzadeh, M.; Jabbari, F.; Farkhondeh, T.; Samini, M. Inhibitory and cytotoxic activities of chrysin on human breast adenocarcinoma cells by induction of apoptosis. Pharmacogn. Mag. 2016, 12 (Suppl. S4), S436. [Google Scholar]
- Lin, Y.-M.; Chen, C.-I.; Hsiang, Y.-P.; Hsu, Y.-C.; Cheng, K.-C.; Chien, P.-H.; Pan, H.-L.; Lu, C.-C.; Chen, Y.-J. Chrysin attenuates cell viability of human colorectal cancer cells through autophagy induction unlike 5-fluorouracil/oxaliplatin. Int. J. Mol. Sci. 2018, 19, 1763. [Google Scholar] [CrossRef]
- Ryu, S.; Lim, W.; Bazer, F.W.; Song, G. Chrysin induces death of prostate cancer cells by inducing ROS and ER stress. J. Cell. Physiol. 2017, 232, 3786–3797. [Google Scholar] [CrossRef] [PubMed]
- Kasala, E.R.; Bodduluru, L.N.; Madana, R.M.; Gogoi, R.; Barua, C.C. Chemopreventive and therapeutic potential of chrysin in cancer: Mechanistic perspectives. Toxicol. Lett. 2015, 233, 214–225. [Google Scholar] [CrossRef]
- Zhang, Q.; Ma, S.; Liu, B.; Liu, J.; Zhu, R.; Li, M. Chrysin induces cell apoptosis via activation of the p53/Bcl-2/caspase-9 pathway in hepatocellular carcinoma cells. Exp. Ther. Med. 2016, 12, 469–474. [Google Scholar] [CrossRef] [PubMed]
- Rehman, M.U.; Ali, N.; Rashid, S.; Jain, T.; Nafees, S.; Tahir, M.; Khan, A.Q.; Lateef, A.; Khan, R.; Hamiza, O.O. Alleviation of hepatic injury by chrysin in cisplatin administered rats: Probable role of oxidative and inflammatory markers. Pharmacol. Rep. 2014, 66, 1050–1059. [Google Scholar] [CrossRef] [PubMed]
- Khan, R.; Khan, A.Q.; Qamar, W.; Lateef, A.; Tahir, M.; Rehman, M.U.; Ali, F.; Sultana, S. Chrysin protects against cisplatin-induced colon. toxicity via amelioration of oxidative stress and apoptosis: Probable role of p38MAPK and p53. Toxicol. Appl. Pharmacol. 2012, 258, 315–329. [Google Scholar] [CrossRef]
- Wu, X.Z.; Chen, D.; Xie, G.R. Extracellular matrix remodeling in hepatocellular carcinoma: Effects of soil on seed? Med Hypotheses 2006, 66, 1115–1120. [Google Scholar] [CrossRef] [PubMed]
- Lin, X. Functions of heparan sulfate proteoglycans in cell signaling during development. Development 2004, 131, 6009–6021. [Google Scholar] [CrossRef] [PubMed]
- Alyoussef, A.; Al-Gayyar, M.M. Cytotoxic and partial hepatoprotective activity of sodium ascorbate against hepatocellular carcinoma through inhibition of sulfatase-2 in vivo and in vitro. Biomed. Pharmacother. 2018, 103, 362–372. [Google Scholar] [CrossRef]
- Nishida, T.; Kataoka, H. Glypican 3-Targeted Therapy in Hepatocellular Carcinoma. Cancers 2019, 11, 1339. [Google Scholar] [CrossRef]
- Qiao, S.-S.; Cui, Z.; Gong, L.; Han, H.; Chen, P.-C.; Guo, L.-M.; Yu, X.; Wei, Y.-H.; Ha, S.-A.; Kim, J.W. Simultaneous measurements of serum AFP, GPC-3 and HCCR for diagnosing hepatocellular carcinoma. Hepato Gastroenterol. 2011, 58, 1718–1724. [Google Scholar] [CrossRef]
- Carr, R.M.; Duran, P.A.R.; Tolosa, E.J.; Ma, C.; Oseini, A.M.; Moser, C.D.; Banini, B.A.; Huang, J.; Asumda, F.; Dhanasekaran, R. The extracellular sulfatase SULF2 promotes liver tumorigenesis by stimulating assembly of a promoter-looping GLI1-STAT3 transcriptional complex. J. Biol. Chem. 2020, 295, 2698–2712. [Google Scholar] [CrossRef]
- Zaghloul, R.A.; El-Shishtawy, M.M.; El Galil, K.H.A.; Ebrahim, M.A.; Metwaly, A.A.; Al-Gayyar, M.M. Evaluation of antiglypican-3 therapy as a promising target for amelioration of hepatic tissue damage in hepatocellular carcinoma. Eur. J. Pharmacol. 2015, 746, 353–362. [Google Scholar] [CrossRef]
- Zheng, X.; Gai, X.; Han, S.; Moser, C.D.; Hu, C.; Shire, A.M.; Floyd, R.A.; Roberts, L.R. The human sulfatase 2 inhibitor 2, 4-disulfonylphenyl-tert-butylnitrone (OKN-007) has an antitumor effect in hepatocellular carcinoma mediated via suppression of TGFB1/SMAD2 and Hedgehog/GLI1 signaling. Genes Chromosom. Cancer 2013, 52, 225–236. [Google Scholar] [CrossRef]
- Bolha, L.; Ravnik-Glavač, M.; Glavač, D. Long noncoding RNAs as biomarkers in cancer. Dis. Markers 2017, 2017. [Google Scholar] [CrossRef]
- Bao, H.; Su, H. Long noncoding RNAs act as novel biomarkers for hepatocellular carcinoma: Progress and prospects. Biomed Res. Int. 2017, 2017. [Google Scholar] [CrossRef]
- Sabry, D.; Abdelaleem, O.O.; Ali, A.M.; Mohammed, R.A.; Abdel-Hameed, N.D.; Hassouna, A.; Khalifa, W.A. Anti-proliferative and anti-apoptotic potential effects of epigallocatechin-3-gallate and/or metformin on hepatocellular carcinoma cells: In vitro study. Mol. Biol. Rep. 2019, 46, 2039–2047. [Google Scholar] [CrossRef]
- Au, J.S.; Frenette, C.T. Management of hepatocellular carcinoma: Current status and future directions. Gut Liver 2015, 9, 437. [Google Scholar] [CrossRef]
- Hanahan, D.; Weinberg, R.A. Hallmarks of cancer: The next generation. Cell 2011, 144, 646–674. [Google Scholar] [CrossRef]
- Lim, W.; Ryu, S.; Bazer, F.W.; Kim, S.M.; Song, G. Chrysin attenuates progression of ovarian cancer cells by regulating signaling cascades and mitochondrial dysfunction. J. Cell. Physiol. 2018, 233, 3129–3140. [Google Scholar] [CrossRef]
- Zhong, X.; Liu, D.; Jiang, Z.; Li, C.; Chen, L.; Xia, Y.; Liu, D.; Yao, Q.; Wang, D. Chrysin Induced Cell Apoptosis and Inhibited Invasion Through Regulation of TET1 Expression in Gastric Cancer Cells. Oncotargets Ther. 2020, 13, 3277. [Google Scholar] [CrossRef]
- Kolluri, A.; Ho, M. The role of glypican-3 in regulating Wnt, YAP and hedgehog in liver cancer. Front. Oncol. 2019, 9, 708. [Google Scholar] [CrossRef]
- Wang, S.; Chen, N.; Chen, Y.; Sun, L.; Li, L.; Liu, H. Elevated GPC3 level promotes cell proliferation in liver cancer. Oncol. Lett. 2018, 16, 970–976. [Google Scholar] [CrossRef]
- Gao, W.; Ho, M. The role of glypican-3 in regulating Wnt in hepatocellular carcinomas. Cancer Rep. 2011, 1, 14. [Google Scholar]
- Guo, M.; Zhang, H.; Zheng, J.; Liu, Y. Glypican-3: A New Target for Diagnosis and Treatment of Hepatocellular Carcinoma. J. Cancer 2020, 11, 2008. [Google Scholar] [CrossRef]
- Capurro, M.I.; Xiang, Y.-Y.; Lobe, C.; Filmus, J. Glypican-3 promotes the growth of hepatocellular carcinoma by stimulating canonical Wnt signaling. Cancer Res. 2005, 65, 6245–6254. [Google Scholar] [CrossRef]
- Gong, T.; Ning, X.; Deng, Z.; Liu, M.; Zhou, B.; Chen, X.; Huang, S.; Xu, Y.; Chen, Z.; Luo, R. Propofol-induced miR-219-5p inhibits growth and invasion of hepatocellular carcinoma through suppression of GPC3-mediated Wnt/β-catenin signalling activation. J. Cell. Biochem. 2019, 120, 16934–16945. [Google Scholar] [CrossRef]
- Lai, J.P.; Sandhu, D.S.; Yu, C.; Han, T.; Moser, C.D.; Jackson, K.K.; Guerrero, R.B.; Aderca, I.; Isomoto, H.; Garrity-Park, M.M. Sulfatase 2 up-regulates glypican 3, promotes fibroblast growth factor signaling, and decreases survival in hepatocellular carcinoma. Hepatology 2008, 47, 1211–1222. [Google Scholar] [CrossRef]
- Yang, J.D.; Sun, Z.; Hu, C.; Lai, J.; Dove, R.; Nakamura, I.; Lee, J.S.; Thorgeirsson, S.S.; Kang, K.J.; Chu, I.S. Sulfatase 1 and sulfatase 2 in hepatocellular carcinoma: Associated signaling pathways, tumor phenotypes, and survival. Genes Chromosomes Cancer 2011, 50, 122–135. [Google Scholar] [CrossRef]
- Lai, J.P.; Oseini, A.M.; Moser, C.D.; Yu, C.; Elsawa, S.F.; Hu, C.; Nakamura, I.; Han, T.; Aderca, I.; Isomoto, H. The oncogenic effect of sulfatase 2 in human hepatocellular carcinoma is mediated in part by glypican 3–dependent Wnt activation. Hepatology 2010, 52, 1680–1689. [Google Scholar] [CrossRef]
- Al-Gayyar, M.M.; Abbas, A.; Hamdan, A.M. Chemopreventive and hepatoprotective roles of adiponectin (SULF2 inhibitor) in hepatocelluar carcinoma. Biol. Chem. 2016, 397, 257–267. [Google Scholar] [CrossRef]
- Lu, J.; Xie, F.; Geng, L.; Shen, W.; Sui, C.; Yang, J. Investigation of serum lncRNA-uc003wbd and lncRNA-AF085935 expression profile in patients with hepatocellular carcinoma and HBV. Tumor Biol. 2015, 36, 3231–3236. [Google Scholar] [CrossRef]
- Motawi, T.M.; El-Maraghy, S.A.; Sabry, D.; Mehana, N.A. The expression of long non coding RNA genes is associated with expression with polymorphisms of HULC rs7763881 and MALAT1 rs619586 in hepatocellular carcinoma and HBV Egyptian patients. J. Cell. Biochem. 2019, 120, 14645–14656. [Google Scholar] [CrossRef]
- Yuan, S.-X.; Tao, Q.-F.; Wang, J.; Yang, F.; Liu, L.; Wang, L.-L.; Zhang, J.; Yang, Y.; Liu, H.; Wang, F. Antisense long non-coding RNA PCNA-AS1 promotes tumor growth by regulating proliferating cell nuclear antigen in hepatocellular carcinoma. Cancer Lett. 2014, 349, 87–94. [Google Scholar] [CrossRef]
- Zhu, X.T.; Yuan, J.H.; Zhu, T.T.; Li, Y.Y.; Cheng, X.Y. Long noncoding RNA glypican 3 (GPC3) antisense transcript 1 promotes hepatocellular carcinoma progression via epigenetically activating GPC3. FEBS J. 2016, 283, 3739–3754. [Google Scholar] [CrossRef]
- Galijatovic, A.; Otake, Y.; Walle, U.; Walle, T. Extensive metabolism of the flavonoid chrysin by human Caco-2 and Hep G2 cells. Xenobiotica 1999, 29, 1241–1256. [Google Scholar] [CrossRef]
- Chen, H.Y.; Jiang, Y.W.; Kuo, C.L.; Way, T.D.; Chou, Y.C.; Chang, Y.S.; Chung, J.G. Chrysin inhibit human melanoma A375. S2 cell migration and invasion via affecting MAPK signaling and NF-κB signaling pathway in vitro. Environ. Toxicol. 2019, 34, 434–442. [Google Scholar] [CrossRef] [PubMed]
- Mohammadian, F.; Abhari, A.; Dariushnejad, H.; Zarghami, F.; Nikanfar, A.; Pilehvar-Soltanahmadi, Y.; Zarghami, N. Upregulation of Mir-34a in AGS gastric cancer cells by a PLGA-PEG-PLGA chrysin nano formulation. Asian Pac. J. Cancer Prev. 2016, 16, 8259–8263. [Google Scholar] [CrossRef] [PubMed]
- Ghavami, G.; Sardari, S. Synergistic Effect of Vitamin C with Cisplatin for Inhibiting Proliferation of Gastric Cancer Cells. Iran. Biomed. J. 2020, 24, 119–127. [Google Scholar] [CrossRef] [PubMed]
| Group | Culture Description |
|---|---|
| Control | Untreated HepG2 cells cultured for 24 and 48 h. |
| Cisplatin | HepG2 cells treated with three concentrations of cisplatin (Cis; 20, 50, and 100 µg/mL) for 24 and 48 h. |
| Chrysin | HepG2 cells treated with three concentrations of chrysin (15, 30, and 60 µg/mL) for 24 and 48 h. |
| Combination | HepG2 cells treated with combined Cis 50 µg/mL and three concentrations of chrysin (15, 30, and 60 µg/mL) for 24 and 48 h. |
| Genes | Forward (5′–3′) | Reverse (5′–3′) |
|---|---|---|
| β-actin | ATGCTCTCCCTCACGCCATC | CAGGATTCCATACCCAAGA |
| GPC3 | GTCCCTGAACGCGACTATTT | AGCTTGTGCCAGCTCTTT |
| SULF2 | CTGAATCCCCACATCGTCCTC | GTCCACCTTGTCATTGTCTCTCTTGT |
| lncRNA AF085935 | CAGGGCAGCAAGGTGTTTTC | TTGGTGGGT TGCCTGATACC |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sherif, I.O.; Al-Mutabagani, L.A.; Sabry, D.; Elsherbiny, N.M. Antineoplastic Activity of Chrysin against Human Hepatocellular Carcinoma: New Insight on GPC3/SULF2 Axis and lncRNA-AF085935 Expression. Int. J. Mol. Sci. 2020, 21, 7642. https://doi.org/10.3390/ijms21207642
Sherif IO, Al-Mutabagani LA, Sabry D, Elsherbiny NM. Antineoplastic Activity of Chrysin against Human Hepatocellular Carcinoma: New Insight on GPC3/SULF2 Axis and lncRNA-AF085935 Expression. International Journal of Molecular Sciences. 2020; 21(20):7642. https://doi.org/10.3390/ijms21207642
Chicago/Turabian StyleSherif, Iman O., Laila A. Al-Mutabagani, Dina Sabry, and Nehal M. Elsherbiny. 2020. "Antineoplastic Activity of Chrysin against Human Hepatocellular Carcinoma: New Insight on GPC3/SULF2 Axis and lncRNA-AF085935 Expression" International Journal of Molecular Sciences 21, no. 20: 7642. https://doi.org/10.3390/ijms21207642
APA StyleSherif, I. O., Al-Mutabagani, L. A., Sabry, D., & Elsherbiny, N. M. (2020). Antineoplastic Activity of Chrysin against Human Hepatocellular Carcinoma: New Insight on GPC3/SULF2 Axis and lncRNA-AF085935 Expression. International Journal of Molecular Sciences, 21(20), 7642. https://doi.org/10.3390/ijms21207642






