1. Introduction
Two-pore domain potassium (K
2P) channel family members, also known as leak K
+ channels, play a pivotal role in maintaining background conductance at the plasma membrane stabilizing the resting membrane potential [
1]. Fifteen mammalian genes encoding K
2P channels were identified, which are classified into six subfamilies, according to their sequence identity and functional properties [
2]: TWIK, THIK, TREK, TASK, TRESK, and TALK. K
2P channels exhibit one of the most versatile patterns of regulation known for any class of ion channels [
3,
4]. They respond to a wide range of regulatory factors, which include extra- and intracellular pH (pH
o and pH
i) [
5,
6,
7], temperature [
8], membrane stretch [
9], polyunsaturated fatty acids [
10,
11], and pharmacological compounds (e.g., volatile anesthetics [
12] or antidepressants [
13,
14]). Currently, the crystallization of 20 different structures of these channels from the TREK, TWIK, and TASK subfamilies has considerably increased our comprehension of the structure and function of K
2P channels (
Table S1).
The K
2P channels assemble as dimers with two pore-forming domains (PD1 and PD2) and four transmembrane helices (TM1–TM4) per subunit. The inner pore-lining helices TM2 and TM4 are in quasi-oblique position below the selectivity filter (SF) with respect to the membrane, while the outer helices TM1 and TM3 are located vertically. Two side-openings, called fenestrations, were resolved in K
2P channel crystallographic structures [
15]. These portals connect the lipid membrane with the central cavity below the SF. In the TWIK-1 structure (Protein Data Bank code (PDB): 3UKM;
Table S1), Miller et al. [
15] suggested that these fenestrations can be occupied by lipid acyl chains, which would interrupt the ion flux at the central cavity. Subsequently, a high-resolution TRAAK structure [
16] (PDB: 4I9W;
Table S1) showed that the fenestrations could coexist in open or closed states, depending on the TM4 down or up movements, respectively. Brohawn et al. [
16,
17] postulated that the TM4 helix in up-state corresponds to the conductive state in TRAAK. In contrast, the down-state allows lipids protruding from the fenestration into the central cavity blocking the passage of the ions. The TREK-2 structure (PDB: 4XDK;
Table S1) co-crystallized with the inhibitor norfluoxetine (NFX) by Dong et al. [
14] is consistent with Brohawn’s hypothesis, where the down conformation primarily corresponds to the closed channel. Regarding the SF in TREK-2, the ion occupancy decreased from four to three potassium when the TM4 helix was found in the down-state, suggesting that gating may occur at the SF level.
Eight of the 15 mammalian K
2P channels respond prominently to changes in extra- or intracellular H
+ concentration. They are related with numerous physiological functions, from hormone secretion to central respiratory adjustment [
6,
18]. TASK-2 or TWIK-related acid-sensitive K
+ channel 2, belongs to the TALK subfamily sharing between 33% and 36% sequence identity with other TALK members (TALK-1 and TALK-2/TASK-4). It was identified in the human kidney as an outward rectifying K
+ channel at physiological potassium concentrations [
19]. TASK-2 plays important roles in cell volume regulation, bicarbonate reabsorption in the proximal tubule of the nephron, and CO
2 chemosensing in neurons of the retrotrapezoid nucleus [
20]. More recently, roles for TASK-2 in K
+ recycling in cochlear outer sulcus cells [
21] and in the process of anion secretion initiated by cyclic adenosine monophosphate (cAMP) in intestinal epithelium [
22] were also reported.
The TASK-2 channel is activated by extracellular alkalinization [
19]. The pH
o sensor in TASK-2 is an arginine residue (R224) located at the outermost portion of the TM4 helix and near to PD2. This pH
o sensor is protonated at acidic pH
o, affecting the electrostatic potential of the SF, the ion occupancy and, therefore, the gating [
23]. The basic residues in this position are conserved within the TALK subfamily, suggesting a unified alkaline pH
o-gating mechanism for the TALK clade [
23]. TASK-2 can be also opened by intracellular alkalization, mediated by a neutralization of the residue K245 [
7]. This pH
i sensor is positioned at the end of the TM4 helix. Mutation in K245 by alanine or cysteine remove the dependence of pH
i in the TASK-2 channel [
7]. Our group built a molecular model of TASK-2 based on the TRAAK structure [
1], in order to estimate the relation between the Brohawn’s hypothesis [
16,
17] and the pH
i mechanism. Based on this model, we proposed that a neutral K245 would favor the proximity of TM2 and TM4 by hydrophobic interactions that close the TASK-2 fenestrations, leading to channel opening. On the other hand, a protonated K245 at acidic pH
i would favor a TM4 change from up- to down-state, opening the side fenestrations and closing the channel. Intra- and extracellular pH-gated mechanisms in TASK-2 are independent [
7]. The pH
o modulation is extracellular K
+- and voltage-dependent; instead, pH
i gating is not affected by either factor [
7]. The pH
o dependence controls the gate of TASK-2 in a mechanism reminiscent to C-type gating at the SF level, whereas the pH
i mechanism remains unclear.
The main goal of this investigation was to study the molecular blockage mechanism that protonated K245 (K245+) exerts in TASK-2 gating and its relationship to the side fenestrations, using computational methodologies. The tridimensional topologies of K2P channels are quite similar, supporting the premise that the crystallographic K2P data can be used to build molecular models of the TASK-2 channel to accomplish our research goal.
Our results show that, firstly, K245+ in an up-state might disrupt the TASK-2 channel function, altering the SF. Secondly, during molecular dynamics (MD) simulations, K245+ precluded closure of the fenestrations and the transformation of the channel to the up-state. Thirdly, although closure of TASK-2 channel fenestrations was only observed by MD simulations, we validated the existence of the fenestrations by blocking the TASK-2 channel with NFX (half maximal inhibitory concentration (IC50) = 17 ± 5 µM). Fourthly, NFX can occupy a similar position in the fenestrations of the TASK-2 channel as in TREK-2 structure. Fifthly, protonation/deprotonation events of K245 take place in the down-state. Sixthly, once the fenestrations are opened and K245 is protonated, the current block of the TASK-2 channel could occur via a positive electrostatic potential effect of K245+. In addition, K245+ could induce a higher number of lipid molecules protruding trough the side fenestrations. Seventhly, the TASK-2 channel could exist in three states in terms of the intracellular pH sensing mechanism: (i) in the up-state and K245 residue neutral (K2450); (ii) in the down-state and K245 residue neutral; and (iii) in the down-state and K245 protonated. Lastly, the most conductive state of TASK-2 is the first, whereas the second state is about 1 kcal/mol more favorable than the third. We expect that these results might contribute to the rational design of molecules to interact with K245 to modulate TASK-2 channel function.
2. Results
2.1. Homology Modeling of TASK-2: Model Relaxation and Equilibration
Here, we aimed to study the effects on the conduction properties of the TASK-2 channel as a function of the aperture or closure of its fenestrations and as a function of the protonation state of K245. Homology models of the TASK-2 channel were obtained from known crystallographic structures in different fenestration states (see
Section 4 for details). For each subset of templates, 150 independent models were built with Modeller [
24].
Six crystal structures with both fenestrations open (OO) selected from TREK-2 (PDB identifiers 4XDK, 4XDJ, and 4XDL) and TASK-1 (PDB: 6RV2, 6RV3, and 6RV4) channels (
Figure S1A) were used to build TASK-2 homology models with OO fenestrations. The final TASK-2 homology model using TREK-2 structures as templates was named T2Tre2OO, while the TASK-2 model using TASK-1 structures as templates was named T2Tas1OO.
Models of the TASK-2 channel with both fenestrations closed (CC) were built from templates obtained from TREK-1 structures (4TWK, 6CQ8, 6CQ6, 6CQ9) and TRAAK (4WFE, 4WFG). The subset of models built using TREK-1 structures was named T2Tre1CC. The subset of models built using TRAAK structures was named T2TraaCC.
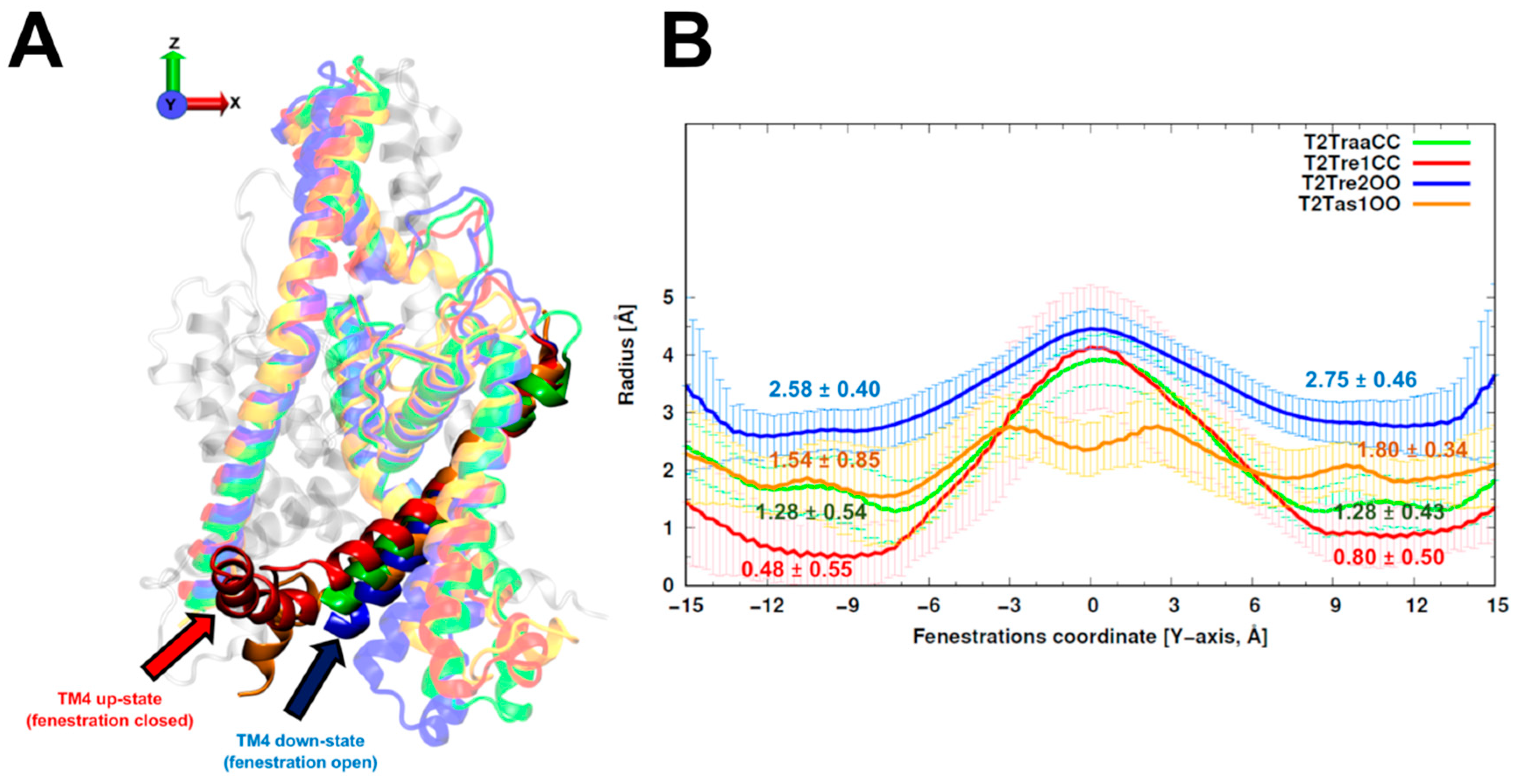
The average and standard deviations of fenestration radii for the 150 models obtained with different template sets are shown in
Figure 1. The TASK-2 models based on the recently solved TASK-1 structures (T2Tas1OO) were barely over the cutoff (1.6 Å) proposed by Jorgensen et al. [
25] to consider the fenestrations open. As expected, the radius of the fenestrations in the model of the open state (T2Tre2OO) was significantly larger than that for the models of the closed fenestration states (T2Tre1CC and T2TraaCC). Indeed, the bottleneck radii of T2Tre2OO models were three- and five-fold (2.75 and 2.58 Å) greater than those of the T2Tre1CC models (0.80 and 0.48 Å). The T2TraaCC and T2Tas1OO models showed an intermediate state of closure/opening for both fenestrations compared with the other models. The models representing the extremes of aperture and closure of the fenestrations (T2Tre1CC and T2Tre2OO) were selected for further analysis through molecular dynamics (MD) simulations in different K245 protonation states.
Initially, 100-ns MD simulations were performed in triplicate for each system (T2Tre1CC and T2Tre2OO) and with the two possible protonation states of K245. The models were named T2Tre1CC-K245+ and T2Tre1CC-K2450 for protonated and neutral K245 states of the channel with the closed fenestrations, and T2Tre2OO-K245+ and T2Tre2OO-K2450 for the pronated and neutral K245 states of the channel with open fenestrations.
The overall variability of the models along the 100-ns MD simulations was evaluated by computing the root-mean-square deviations (RMSDs) relative to the initial structures. The two models with open fenestrations (T2Tre2OO-K245
+ and T2Tre2OO-K245
0) and the model of the neutral closed fenestration state (T2Tre1CC-K245
0) displayed RMSDs systematically smaller than about 3 Å, indicating that the structures were highly preserved (
Figure S2A). The model of the closed fenestration state and K245 protonated (T2Tre1CC-K245
+), on the other hand, displayed larger fluctuations and, although the RMSDs smaller than 5 Å did not suggest major structural changes, some fluctuations not observed for the other models were present.
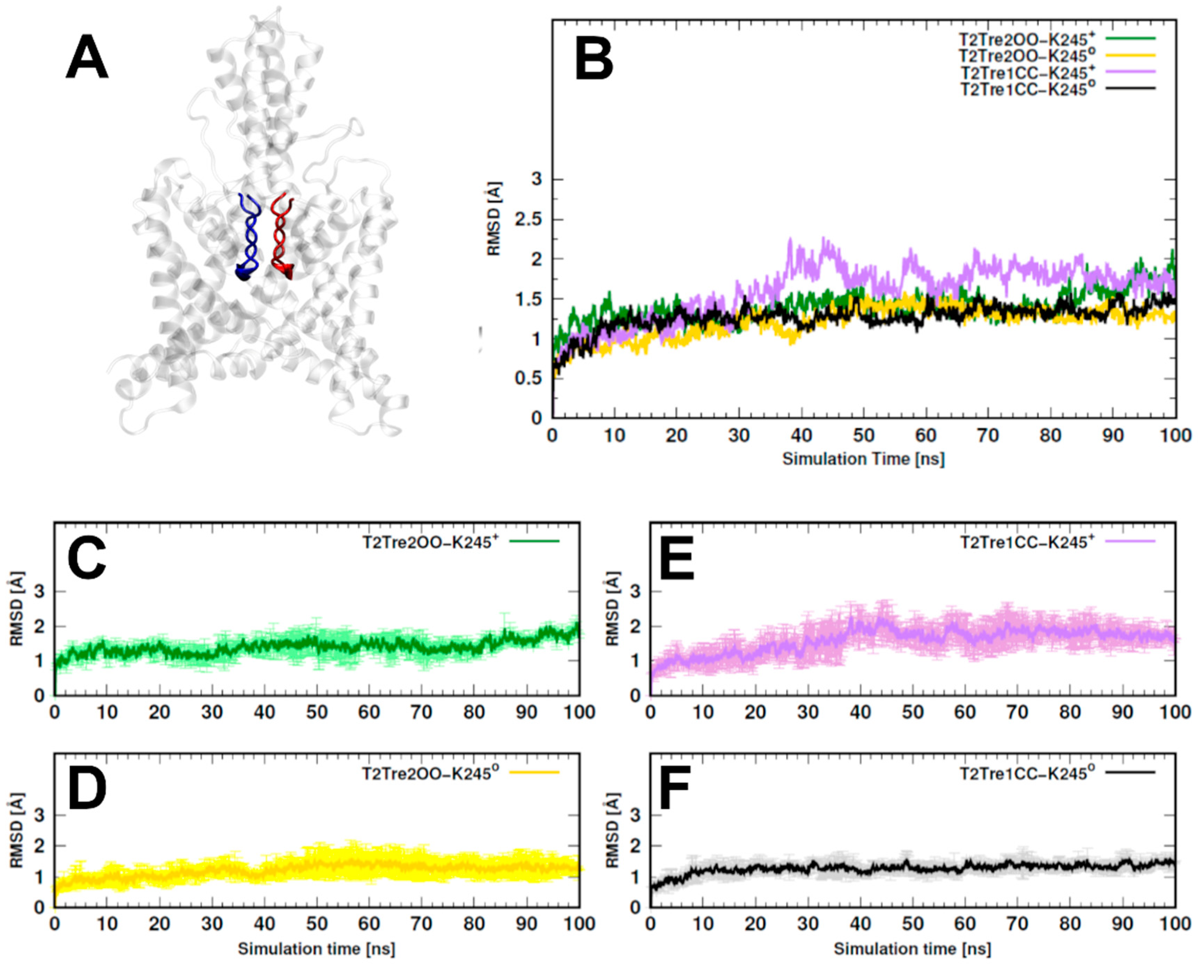
In
Figure 2, we show the RMSDs of the residues of the selectivity filter (residues 97–103 and 203–208, as shown in
Figure 2A), which displayed a similar trend as observed for the entire structures, but with smaller deviations. Effectively, for the two open states and for the neutral closed state, the channel residues preserved the initial structure within 2 Å RMSD (
Figure 2B–D,F), while, for T2Tre1CC-K245
+-, larger structural deviations were observed, and the RMSD variability between MD replicas was also greater for this model (
Figure 2E). To avoid biases associated with the initial models used, analyses were performed in all cases for the last 50 ns of simulations.
The greater structural variability of the T2Tre1CC-K245
+ model could be associated with disfavored hydrophobic interactions that close the fenestrations. The protonation of K245 led to increased fluctuations of the TM4 helix in both T2Tre2OO-K245
+ and T2Tre1CC-K245
+ (
Figure S3A) and, in the case of the closed fenestration model, increased fluctuations were observed also in the TM2 helix. The TM2 helix interacts with TM4 (K245 included) only if the fenestrations are closed. Therefore, the simulations show that the protonation of K245 increases the structural fluctuations of TM4, and that these fluctuations propagate to TM2 if the fenestrations are closed. Nevertheless, we could not identify hydrogen bonds or salt bridges systematically involving the K245 residue in any system, such that these protonation effects on the fluctuations of TM2 might impede the hydrophobic interactions that close the fenestrations.
The radii of the fenestrations within the simulations are shown in
Figure 3. The bottlenecks of closed states varied around 1 Å radius and were similar in both protonation states. A slightly smaller central cavity was found in T2TreCC-K245
+ relative to T2Tre1CC-K245
0 (
Figure 3A). The bottlenecks for open states varied between averages of 0.88 Å and 2.41 Å. A constriction of the fenestration was observed for the neutral K245 model, mainly at the right-side fenestration, regarding the initial configuration (~2 Å in T2Tre2OO-K245
0—
Figure 3B versus
Figure 1). Thus, the neutral K245 led to a partial closure of the fenestrations in the simulations starting with the open states.
The rotamers of the side chain of K245 could be classified into two distinguishable clusters, as shown in
Figure S4A: (1) the ε-amino group in a pseudo-orthogonal position regarding the conduction pore, orienting the side chain toward the lipidic membrane (in cyan color), and (2) the ε-amino group in a quasi-parallel position with respect to the pore (in yellow color). Both conformers were identified in all but the T2Tre1CC-K245
+ system, for which the K245 side chain was only found in the pseudo-orthogonal conformation (
Figure S4B).
The pKa values for the TASK-2 K245 residues were estimated from the molecular environment observed in the MD simulations and are shown in
Table 1. The protonation led to fluctuations in the molecular environment of K245 that, as expected, implied a greater pKa (the variations of the molecular environment in the presence of the proton should stabilize its presence, thus increasing the basicity required to remove it). This stabilization of the protonated state was stronger in the open fenestration states (pKa ≈ 8.65 and 8.98) than in the closed fenestrations (pKa ≈ 7.95 and 7.75). Therefore, the protonation of K245 is more stable with the fenestrations open. The average pKa for K245 in open-state simulations (T2Tre2OO-K245
+ and T2Tre2OO-K245
0) was 7.93 ± 0.98, and that in closed-state simulations (T2Tre1CC-K245
+ and T2Tre1CC-K245
0) was 7.34 ± 0.63. Both pKas were, within the fluctuations, consistent with the experimental pK
1/2 of 8.0, but the data suggests that the experimental pKa is closely associated with the protonation events in the open fenestrations.
In summary, structural models were built for TASK-2 with open and closed fenestrations, and MD simulations were performed for the final models in the two possible K245 protonation states. The models displayed fenestration radii consistent with the expected aperture of the channels, but the closure of one of the fenestrations was observed in a K245 neutral state starting from an open state. The overall structures were mostly stable within the timescales of the simulations, except for an apparent instability of the TM2 helix in the closed protonated state. The simulations suggest, then, that the neutral open state and the protonated closed state of the fenestrations are somewhat unstable, and that there is a coupling between the aperture of the fenestrations and the protonation of the K245 residue.
2.2. Fenestration Aperture and Channel Conductivity
The TREK-2 channel conductance is inhibited by NFX, the active metabolite of fluoxetine (Prozac). A crystallographic structure of the NFX–TREK-2 complex was described (PDB 4XDK), with NFX located in the open fenestrations [
14] (
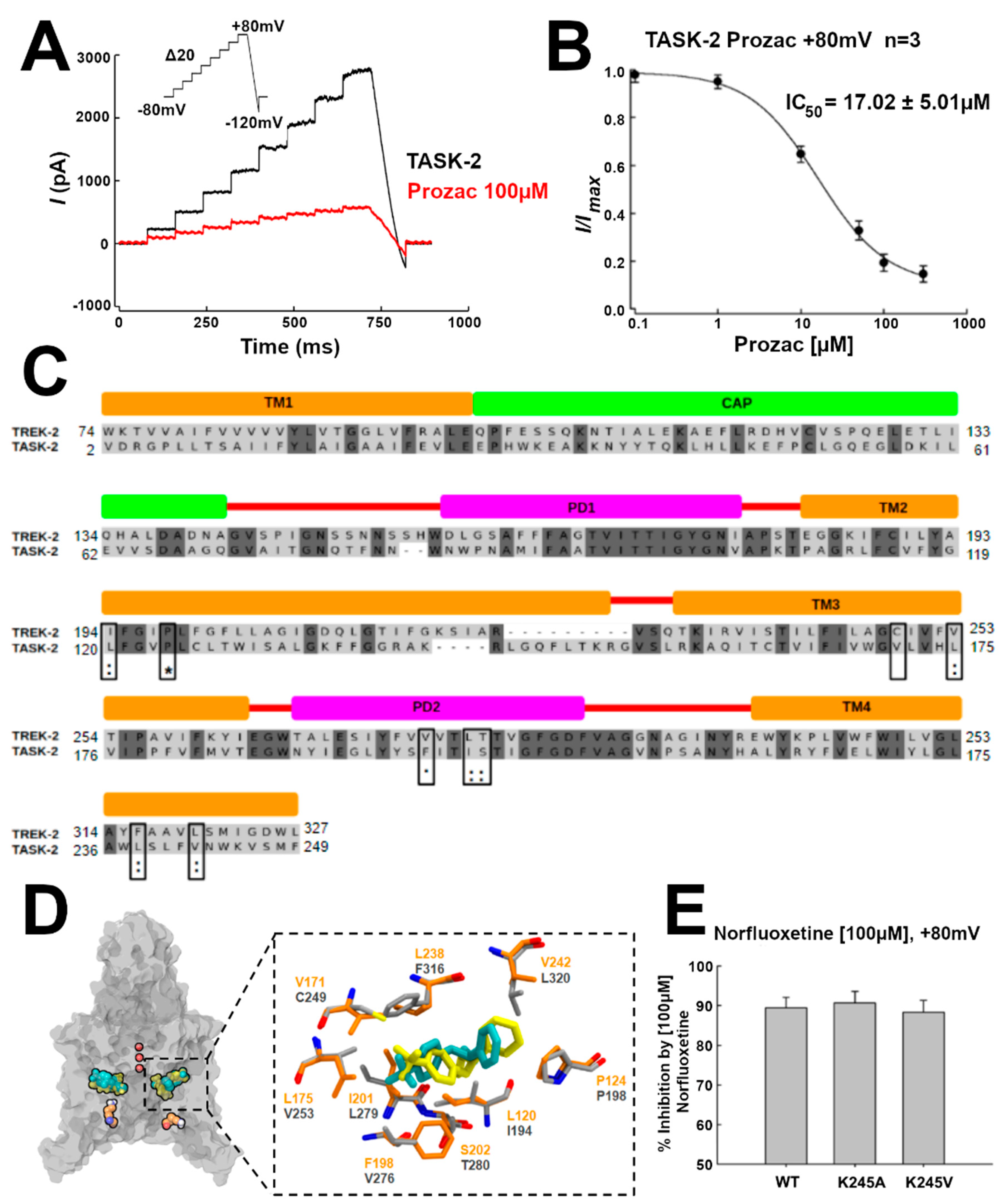
Table S1). We hypothesized that, if TASK-2 can also display open fenestrations, NFX should also block the TASK-2 channel. Indeed, as shown by current traces and the IC
50 in
Figure 4A,B, respectively, NFX blocked the conductivity of TASK-2 (at pH = 7.4, where we expect that K245 is mostly protonated) [
7]. Additionally, we docked NFX in the TASK-2 model with open fenestrations (T2TreOO), and found that NFX can occupy a similar position to that reported in the TREK-2 structure (
Figure 4C). The norfluoxetine binding site [
14] is conserved in TASK-2 except for a cysteine (C249) in TREK-2 which is replaced by V171 in TASK-2 (
Figure 4C,D). Thus, a stabilization of an open fenestration state induced by NFX would inhibit the channel, consistent with the concept that a conducting structure is that with fenestrations closed.
The inhibition of K
2P channels by the opening of the fenestrations was suggested to be associated with the blocking of the passage of ions as a consequence of the penetration of the lipids into the channel [
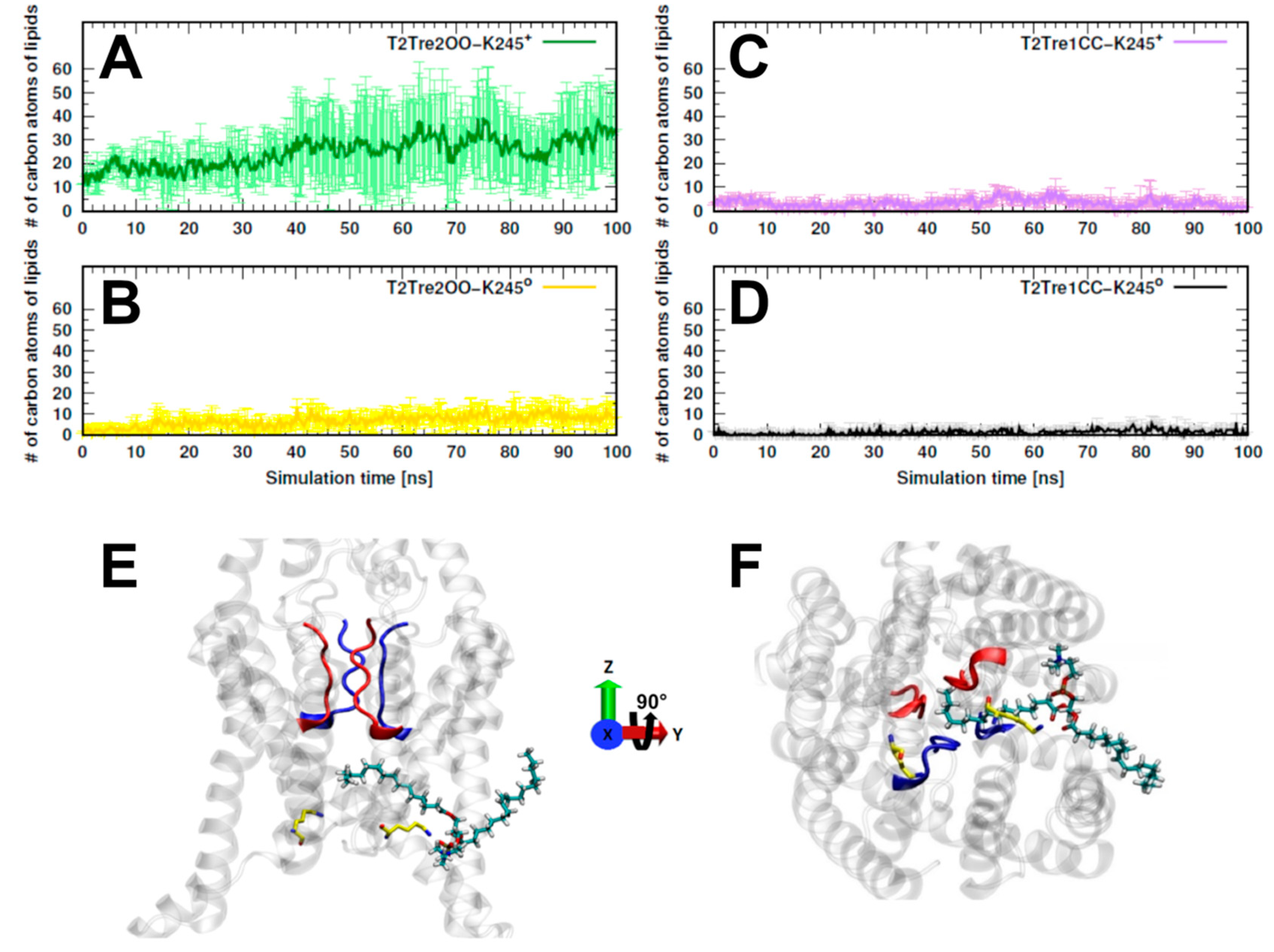
17]. We computed, then, the number of carbon atoms belonging to lipids that were found inside the fenestrations along the entire MD simulations. In
Figure 5A, we see that, for T2Tre2OO-K245
+, about 30 to 40 lipid atoms penetrated the fenestrations. For the simulation initiated with the open channel but with neutral K245 (T2Tre2OO-K245
0), there was some insertion of the lipids (at most about 10 C atoms,
Figure 5B), which was already inhibited by the partial constriction of the fenestrations (
Figure 3B). For the closed fenestration simulations (T2Tre1CC-K245
+ and T2Tre1CC-K245
0,
Figure 5C,D), the lipids could not penetrate the channel.
Figure 5E,F show the lipids in the conduction pore, crossing with one hydrophobic tail through the fenestrations, and reaching the central cavity below the SF in T2Tre2OO-K245
+ simulations. Thus, these results are consistent with a mechanism of channel blocking by lipid insertion into the open fenestrations.
2.3. K245 Protonation State and Channel Conductivity
The protonation of K245 appears to be coupled with the aperture of the fenestrations. In Reference [
7], Sepúlveda and co-workers showed that the channel conductance is mostly unaffected by mutating this lysine residue to alanine. This indicates that the channel can exist in the closed state, which is conductive, in the neutral K245 state. Here, we studied the inhibition of the conductance by NFX in pH
i-insensitive mutants to address the hypothesis that the fenestration can also be found open in the neutral state. Effectively, as shown in
Figure 4E, the mutants TASK2-K245A and TASK2-K245V are inhibited by NFX similarly to the native TASK2, implying that NFX can bind the neutral state. Since NFX binds to the down-state, that is, when the fenestrations are open, the channel can exist with open fenestrations in the neutral K245 state. Summing up, these results indicate that, in the neutral K245 state, the channel can be found in the open and in the closed fenestration states. Our simulations provide an indication that the closed fenestration state might be more stable, as the closure of one of the fenestrations was observed in the T2Tre2OO-K245
0 simulation (
Figure 3B).
Clear evidence exists of the stability of the open fenestration state when K245 is protonated. The simulations of the T2Tre2OO-K245
+ were stable, and the pKa of the K245 residues was the greatest, supporting the stability of the protonated state when the fenestrations are open. The average pKa computed from the two down-state (neutral and protonated) simulations was consistent with the experimental pKa, suggesting that the equilibrium of protonation and deprotonation of the lysine occurs in the open state (
Table 1).
The demonstration of the existence of a significant population, or not, of the closed fenestrations with the protonated K245 is, however, difficult. Our equilibrium simulations indicated that there are significant fluctuations of the structure in these conditions (
Figure 2B), suggesting a possible instability, although the opening of the fenestrations was not observed in the timescale of the simulations (
Figure 3). Experimentally, it is clear that the channel is inhibited at pH
i ≲ 8.0 [
7], when K245 is expected to be protonated. This inhibition can be direct via the introduction of electrostatic repulsion, but also mediated by the shift in equilibrium from the closed to the open state upon protonation.
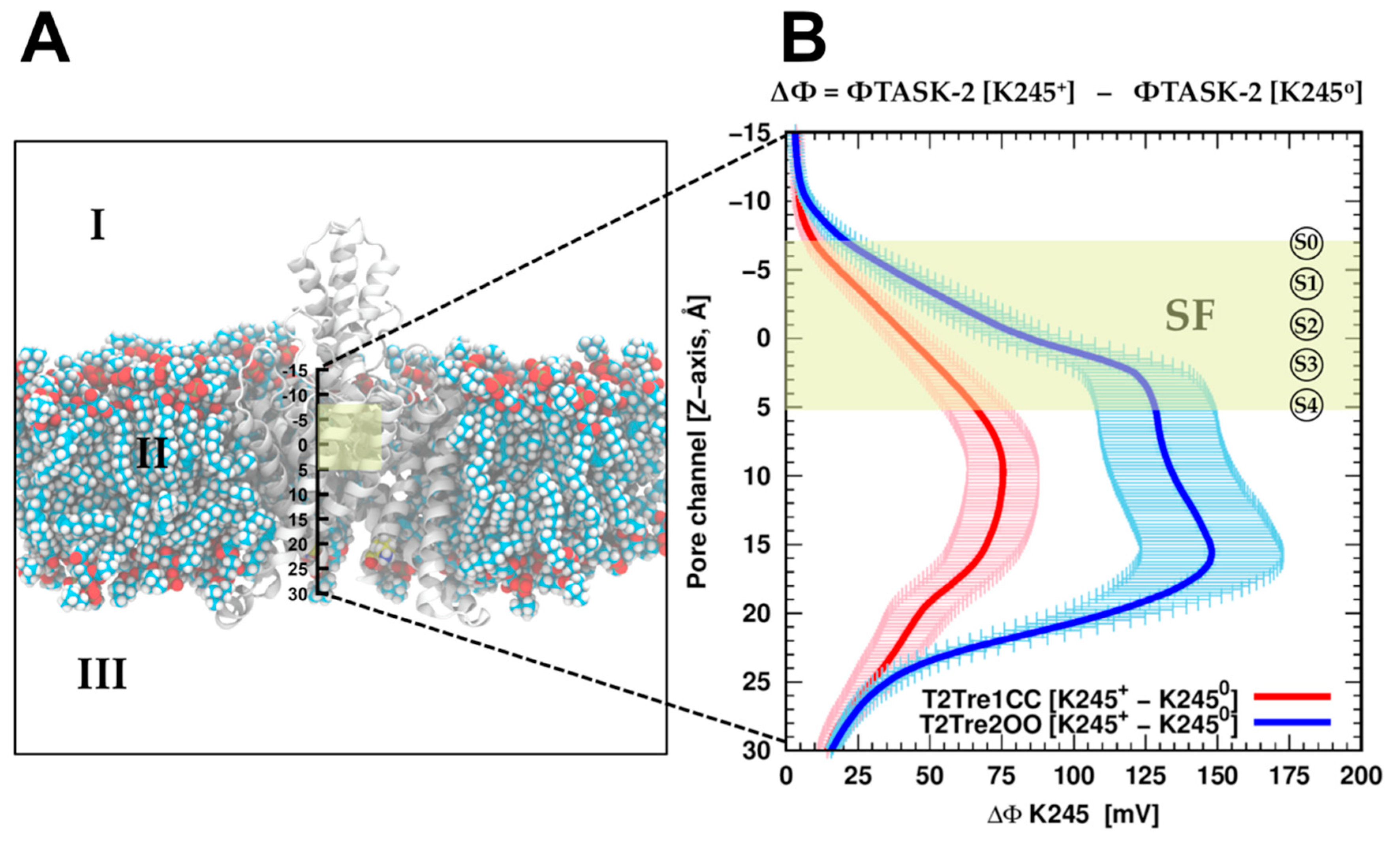
To evaluate if the protonation of K245 can introduce a direct inhibition of the conductance by direct electrostatic interactions, the effect of K245 protonation on the electrostatic potential of the channel was calculated. Adding the proton K245 increased the electrostatic potential, as expected and shown in
Figure 6B, for the channel in both the open fenestration state and the closed fenestration state. Therefore, adding the proton works against the presence of potassium ions in the channel in both states from the point of view of direct electrostatic interactions. However, this effect is much greater when the fenestrations are open than when they are closed. This difference stems from the orientation of the K245 side chains in each fenestration state. As shown in
Figure S4A, in the closed state, the side chains of the K245 residues cannot adopt an orientation in which it points inward into the channel. Thus, from the analysis of direct electrostatic interactions, the inhibitory effect of protonating the K245 side chains is expected to be greater with the fenestrations open. From this point of view, the inhibition of the conductance at low pH
i results from the coupled effects of protonation and fenestration opening.
Therefore, assuming that this electrostatic repulsion impairs the entrance of the ions into the channel, the K245 protonation would decrease the conductivity of the channels in the open state of the fenestrations more than in the closed state. In other words, the sensitivity of the channel in its closed fenestration state to the protonation of K245 is relatively small. This must be combined with the fact that the pKa of K245 in the closed fenestration state is lower and, thus, it is less likely that this residue is protonated in this state. Summing up, we expect that protonation is less common for the closed fenestration state and, when it occurs, it affects the electrostatic potential of the channel less than for the open state, favoring ion conduction. By contrast, K245 is found more frequently in the protonated state if the fenestrations are open and, additionally, the effect of protonation is relatively large on the electrostatic potential of the channel, with both effects leading to decreased ion transport.
2.4. Reaction Rates from Free Energy Profiles of Ion Transport
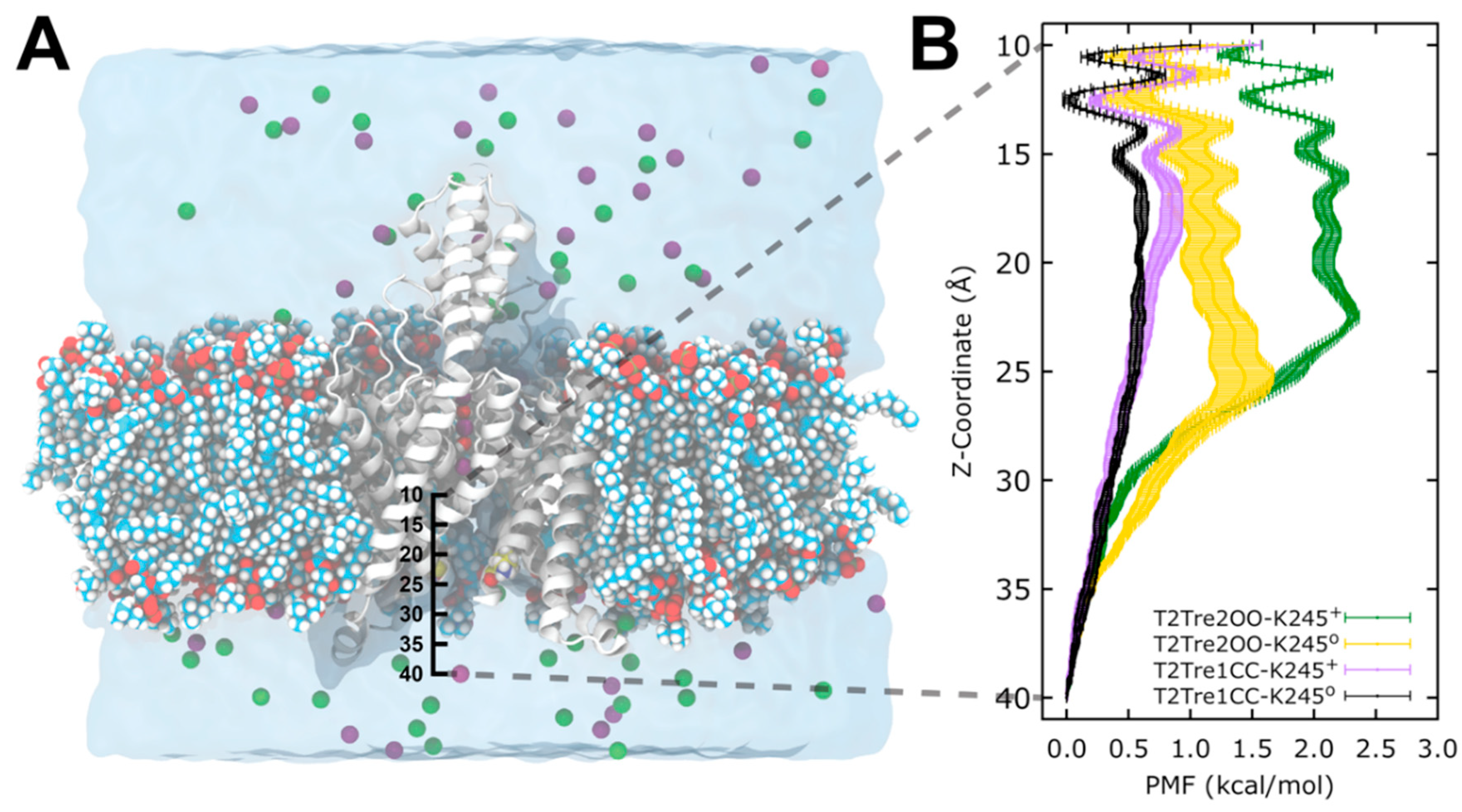
The above analysis assumes that the major barrier for conduction, under K245+ and open fenestration conditions, is the occupancy of the permeation pathway by the ions. This assumption can be supported by molecular dynamics simulations. We performed free energy calculations of K+ entrance into the channels in open and closed fenestrations and in K245 protonated and deprotonated states. Potentials of mean force (PMFs) were obtained for the passage of the ions from the cytosolic solution to the central cavity of the channel, as a function of the fenestration state and K245 charge.
In
Figure 7, we see that, for the channel with closed fenestrations (black and violet curves), the major barriers for ion conductance were within the central cavity of the channel (
Z < 20 Å), while, in the open state, there was an additional and dominant barrier at the channel entrance from the intracellular side (
Z ~ 25 Å). This dominant barrier was further increased with the protonation of the K245 residue (green plot).
From the PMFs of
Figure 7, it is possible to obtain estimates of the relative ion permeation rates, assuming that the maximum barriers sampled are determinant. In a first approximation, it is possible to use transition state theory to compute the rates of ion transport from the highest free energy barrier. The relative rates of reaction of two mechanisms can be computed by
where
k2 and
k1 are the two rate constants, and Δ
G2 and Δ
G1 are the free energy barriers computed relative to the same reference state.
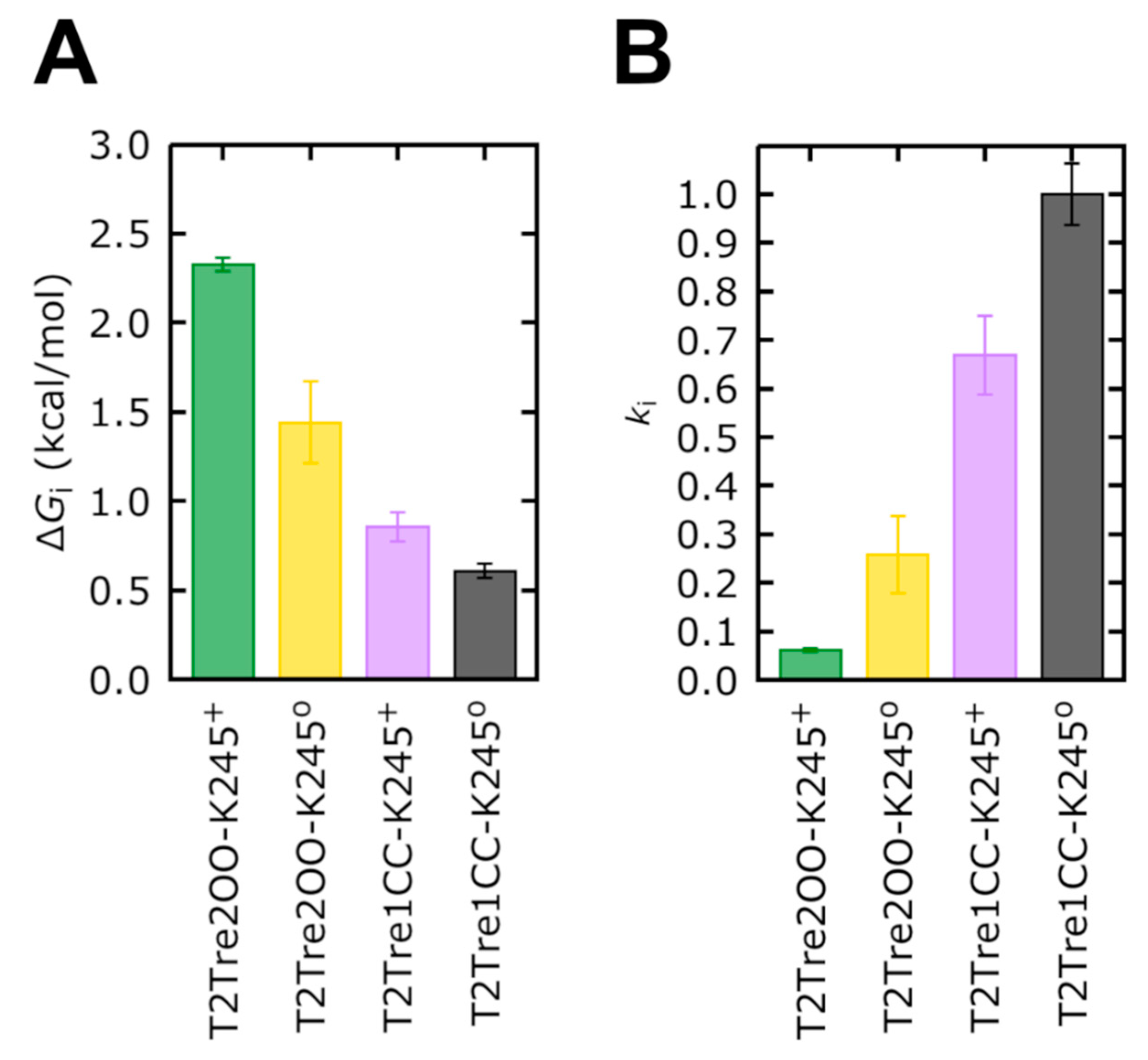
Firstly, the free energy profiles of the closed fenestration states (
Figure 7B) differed by at most 0.25 kcal/mol (
Figure 8A). This free energy barrier difference implies rates of ion transport which differed between the two closed states by at most a factor of ~1.5 (
Figure 8B).
The opening of the fenestrations introduces greater barriers, however. Here, considering as the reference state the ion in the cytosolic solution (reaction coordinate
Z = 40 Å), we obtained that the highest barrier observed for the channel with open fenestrations and a neutral K245 (yellow curve in
Figure 7B) was of 1.44 kcal/mol, which is 0.84 kcal/mol greater than the maximum free energy barrier of the system with the fenestrations in the closed state with neutral K245 (black bar in
Figure 8A). This implies that ion transport through T2Tre2OO-K245
0 is ~4.0 times slower than for T2Tre1CC-K245
0 due to this barrier (
Figure 8B). Thus, the opening of the fenestrations reduces the ion transport rate by about four times, according to the PMFs obtained here. The opening of the fenestrations, when coupled to the protonation of the K245 residue, led to the highest PMF values shown in
Figure 7B. This PMF calculation displays a maximum free energy barrier of 2.32 kcal/mol (
Figure 8A), which is 1.72 kcal/mol greater than the barrier of T2Tre1CC-K245
0. The transport rate of the system with fenestrations closed in the deprotonated state is, thus, ~16 times greater than the system with the open fenestrations in the protonated state (
Figure 8B).
The combined effects of protonation and fenestration opening result, therefore, in a reduction of channel conductance (as measured by the ratio of transport rates) of about 16-fold. The experimental inhibition of the conductance obtained by reducing the pH from −9.5 to −6.5 is of about 10 times for the wild-type TASK2 [
7], which is similar to our result. Therefore, the inhibition observed experimentally by varying the pH appears to be the combined effect of protonation and fenestration opening.
In summary, the free energy profiles of ion permeation from the cytosolic solution to the channel central cavity predict rates of ion transport in qualitative agreement with experimental observations. These results support the hypothesis that the major barrier for conductance is the entrance of ions in the channel, which is modulated by the fenestration aperture state and charge of K245. These effects cannot be easily decoupled in the experimental set-up as the protonation of K245 induces the opening of the fenestrations.
3. Discussion
TASK-2 is a member of the K
2P channel family activated by alkaline intracellular [
7] and extracellular pH [
5,
23]. The pH
o-insensitive TASK-2-R224A mutant is not altered in its pH
i dependence, suggesting that the pH
i dependence of TASK-2 occurs independently of the SF gating responsible for pH
o dependence [
7]. Neutralization of a lysine residue (K245) located at the C-terminal end of TM4 abolishes gating by pH
i [
7]. Previously, our group hypothesized that a neutral K245 favors the hydrophobic interactions that close the fenestrations, but K245
+ would preclude the fenestration closing [
1].
Here, 20 K
2P channels structures were crystallized from the TREK, TWIK, and TASK subfamilies. They served as blueprints in this work to model the TASK-2 channel with closed and open fenestrations (
Figure 1). We propose that K245
+ in a closed state of the fenestrations might disrupt the TASK-2 channel function, altering the SF. A clear insight of stability loss was observed in the RMSD analysis of the selectivity filter of T2Tre1CC-K245
+ molecular dynamics simulation (
Figure 2). Previously, it was observed that activators of K
2P channels stimulate function by reducing the dynamics of the SF [
28]. Conversely, a decrease in pH
i will protonate the K245 residue, promoting the dynamics of the SF—in a closed state of the fenestrations—and moving TASK-2 to a non-conducting channel.
A stability loss was observed also in the RMSD analysis of the whole protein (
Figure S2) in T2Tre1CC-K245
+ MD simulations. Root-mean-square fluctuation (RMSF) analysis (
Figure S3) and this could explain how the protonation of K245 at TM4 disrupts the selectivity filter. A charged K245 alters the apposition of TM4 and TM2, inducing the movement on both segments. These fluctuations were observed in T2Tre1CC-K245
+ MD simulations and could alter the SF which is directly above TM2 and TM4. An RMSF increase can also be observed in the TM4 segment in the MD simulations based on the T2Tre2OO model. However, the T2Tre2OO model does not include the C-terminal region, and the reason for these fluctuations could be the shortened model, ending exactly with TM4.
The hypothesis that K245
+ would preclude the fenestration closing [
1] is supported by comparison of the dimensions of the fenestrations during TASK-2 MD simulations (
Figure 3). Similar to the 100-ns MD simulations published earlier for TREK-2 [
14], TASK-1 [
29,
30], or TWIK-1 [
25] channels, the T2Tre2OO model changed into the up-state during MD simulations and the side fenestrations closed but only in the T2Tre2OO-K245
o simulations (
Figure 3B,
Video S1). The fenestration closing was prevented during T2Tre2OO-K245
+ MD simulations. The bottleneck radius of the fenestrations significantly decreased in the T2Tre2OO-K245
0 MD simulations, while K245
+ stabilized the bottleneck radius of the side fenestrations at 2.35 Å over the 100-ns MD simulations (
Figure 3B). Thus, K245
+ precludes closure of the side fenestrations and the transformation of the channel to the up-state. Nevertheless, closure of TASK-2 channels fenestrations was only observed here by MD simulations. Our model must be tested in the future to demonstrate if TASK-2 channels gate in a similar way to other K
2P channel subfamilies and whether they physiologically change into an up-state-like conformation.
To validate the presence of open fenestrations in TASK-2 channels, we used electrophysiological measurements applying norfluoxetine to the channels. NFX is the active metabolite of fluoxetine (Prozac) and binds within intramembrane fenestrations of TREK-2 channels [
14]. McClenaghan et al. [
31] reported that the activation by membrane stretch but not by pH
i produces a loss of NFX sensitivity in the TREK-2 channel. NFX also blocks TASK-2 with an IC
50 = 17 ± 5 µM (
Figure 4B). A docking simulation of NFX in T2Tre2OO model shows that NFX can occupy a similar position in the fenestrations of the TASK-2 channel as it does in the TREK-2 structure (
Figure 4C). However, mutants should be done in the future in the residues highlighted with black boxes in
Figure 4D to determine if NFX binds in TASK-2 in a homologous pocket of the TREK-2 channel. We measured also the inhibition by NFX in two functional mutants of the TASK-2 channel (TASK2-K245A and TASK2-K245V) which are insensitive to pH
i [
7], because the 245th residue is not titratable in the mutants. Both mutants exhibit the same inhibition by NFX as the wild-type TASK-2 channel (
Figure 4E), suggesting that a deprotonated state of the channel could exist also with open fenestrations. In fact, we suggest that the protonation/deprotonation events of K245 take place with open fenestrations. The average of the predicted pK
a values for T2Tre2OO simulations (T2Tre2OO-K245
+ and T2Tre2OO-K245
0) was 7.93 ± 0.98 units which is similar to the reported pK
1/2 of K245 in the TASK-2 channel [
7] (~8 units) (
Table 1).
Once the fenestrations are opened and K245 is protonated, the current block of TASK-2 channels could occur in two different ways which are not mutually exclusive: (1) K245
+ generates a positive electrostatic potential (ΔΦ) distribution within the ion pathway (
Figure 6) preventing the ion flux, or (2) K245
+ induces a higher number of lipid molecules protruding trough the side fenestrations, toward the central cavity of the channel, blocking physically the flux (
Figure 5). Positive potential imparted upon the conducting pore blocking the K
+ ion occupation is a common mechanism of K
2P channel pH sensors [
5,
32] whereas, in the TRAAK channel structure, in the open state of the fenestrations, lipids accessing the internal cavity through these openings prevent ion occupancy [
17].
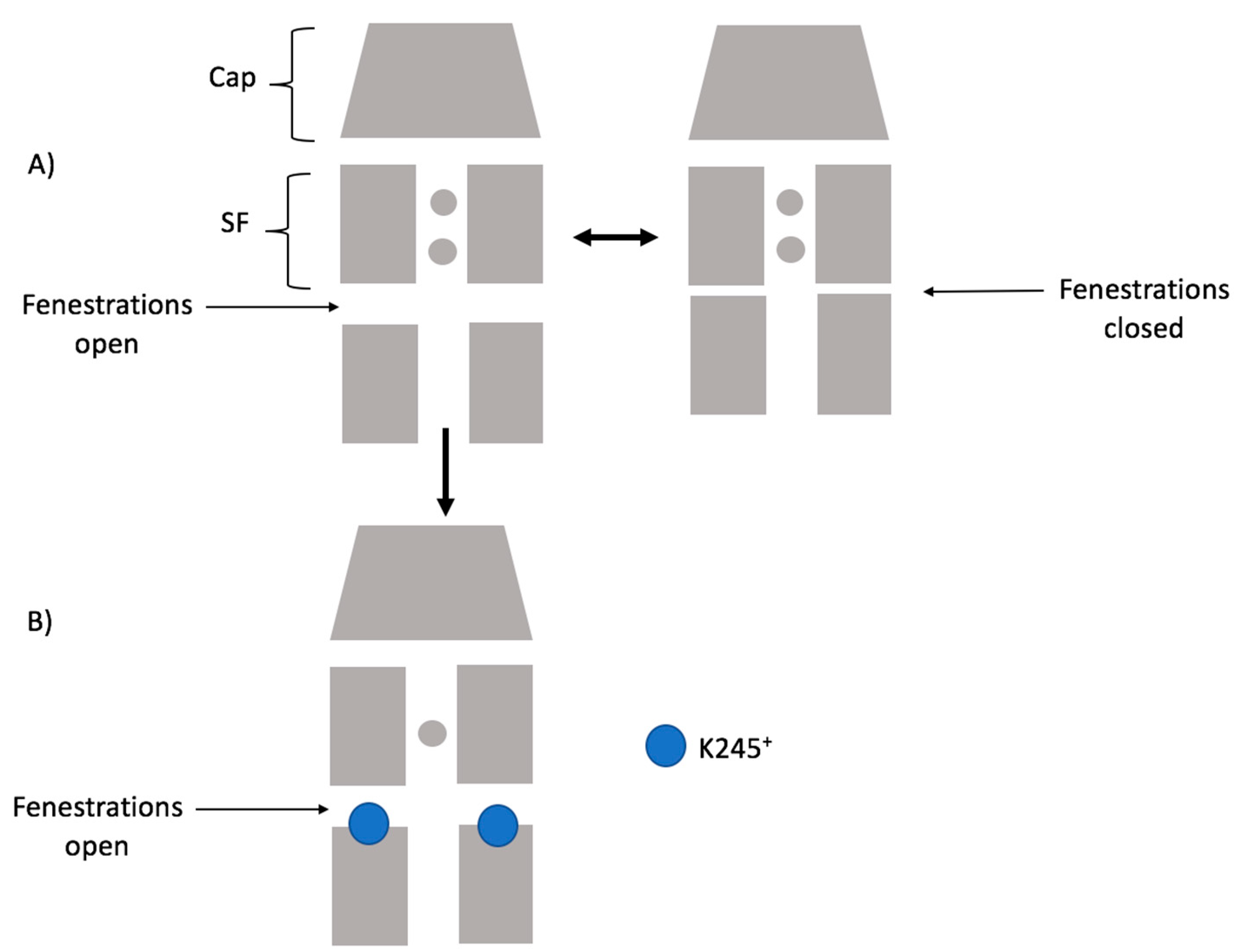
Considering our results, the scheme below (
Figure 9) summarizes how K245
+ would preclude the fenestration closing in TASK-2. Firstly, the channel could switch from a down-state to an up-state. Secondly, in the down-state, K245 protonates and prevents K
+ flux. Under this scheme, the TASK-2 channel could exist in three states in terms of the intracellular pH sensing mechanism: TASK-2 with closed fenestrations and a neutral K245 residue (T2…CC-K245
0), TASK-2 with open fenestrations and a neutral K245 residue (T2…OO-K245
0), and TASK-2 with open fenestrations and a charged K245 residue (T2…OO-K245
+).
Figure 7 and
Figure 8 show that ion flux in the T2…OO-K245
+ system is about 1 kcal/mol less favorable than in T2…OO-K245
0. The most conductive state of TASK-2 is T2…CC-K245
0. This is in agreement with Brohawn et al.’s [
16,
17] hypothesis. They proposed that the closed state of the fenestrations is the conductive state of the K
2P channels. Also, Brennecke and de Groot [
33] studied—using molecular dynamics simulations—the conductance of the K
2P channel TREK-2 and found that the down-state is less conductive than the up-state. We did not include the T2…CC-K245
+ in our scheme because the closed channel is unstable if protonated (
Figure 2A).