Curcumin Loaded Nanoliposomes Localization by Nanoscale Characterization
Abstract
1. Introduction
2. Results and Discussion
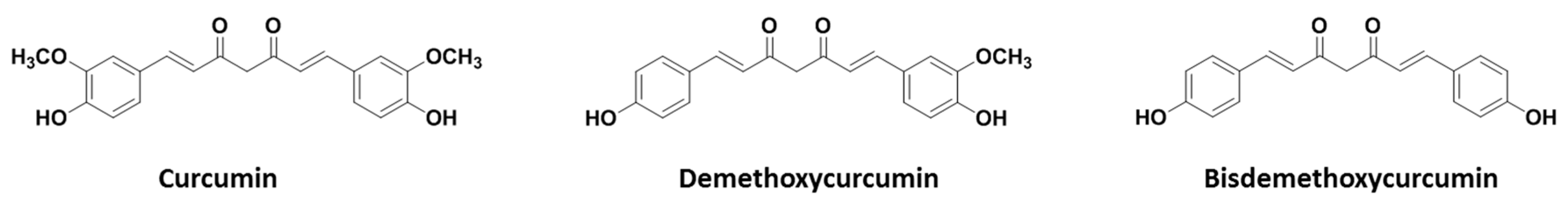
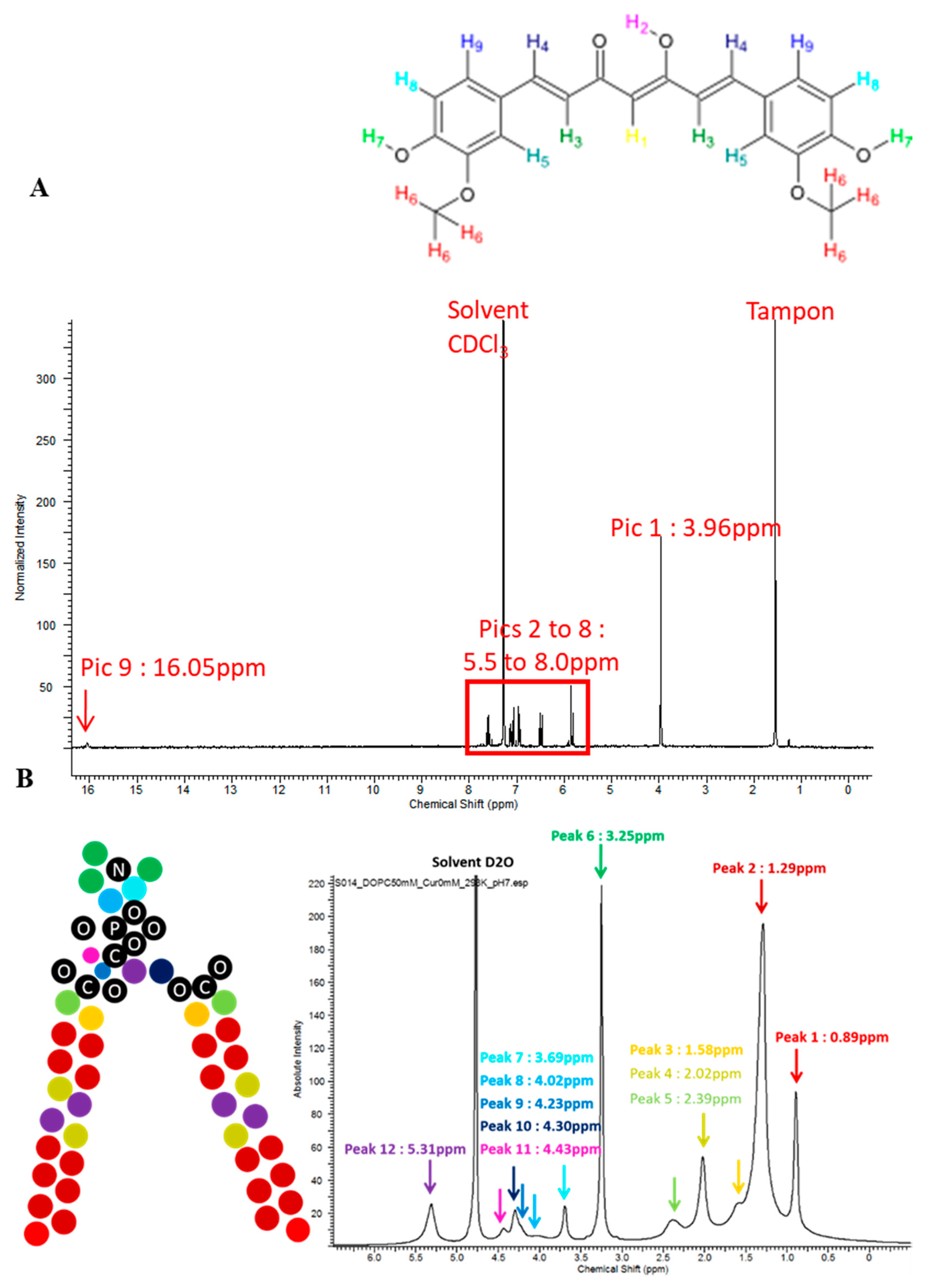
2.1. Curcumin Purity
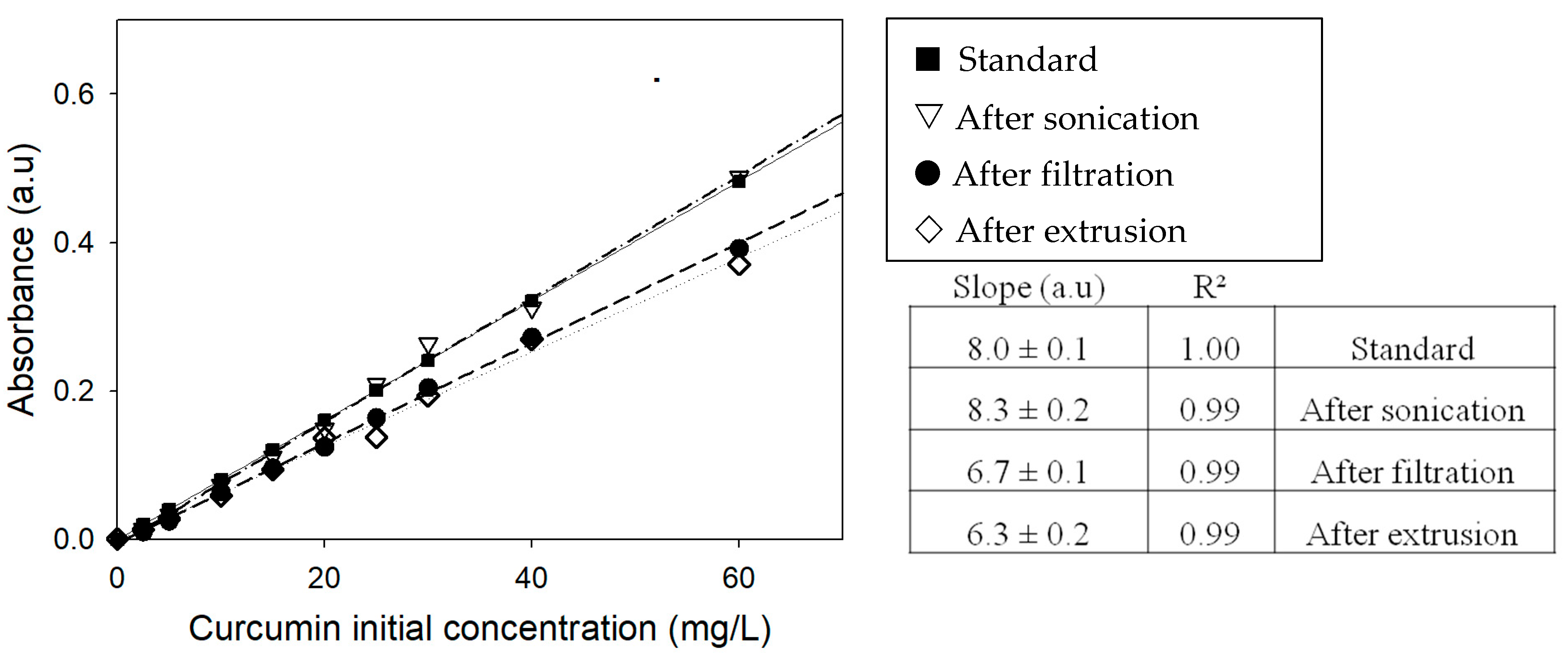
2.2. UV-Visible
2.3. Size and ζ-Potential Measurements
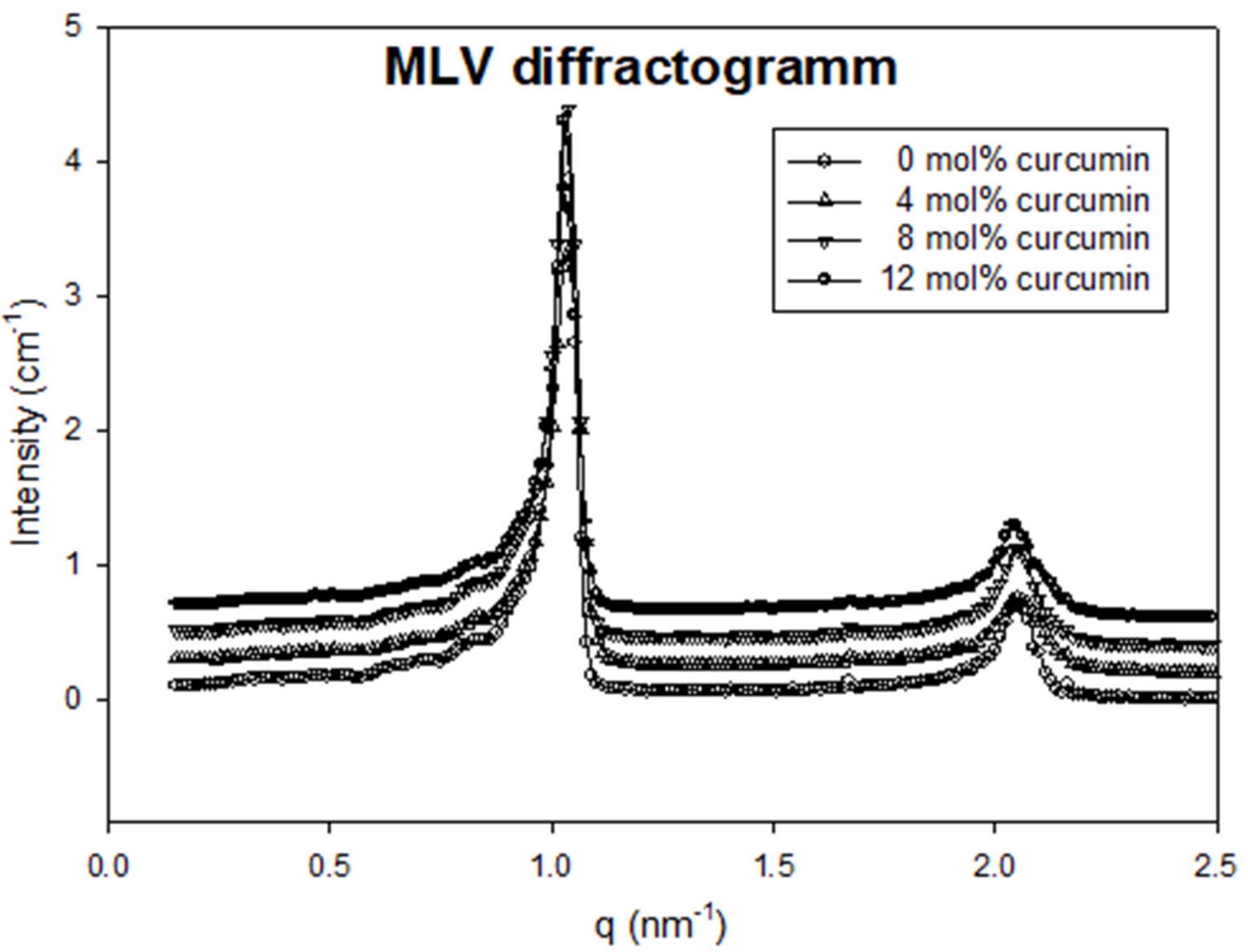
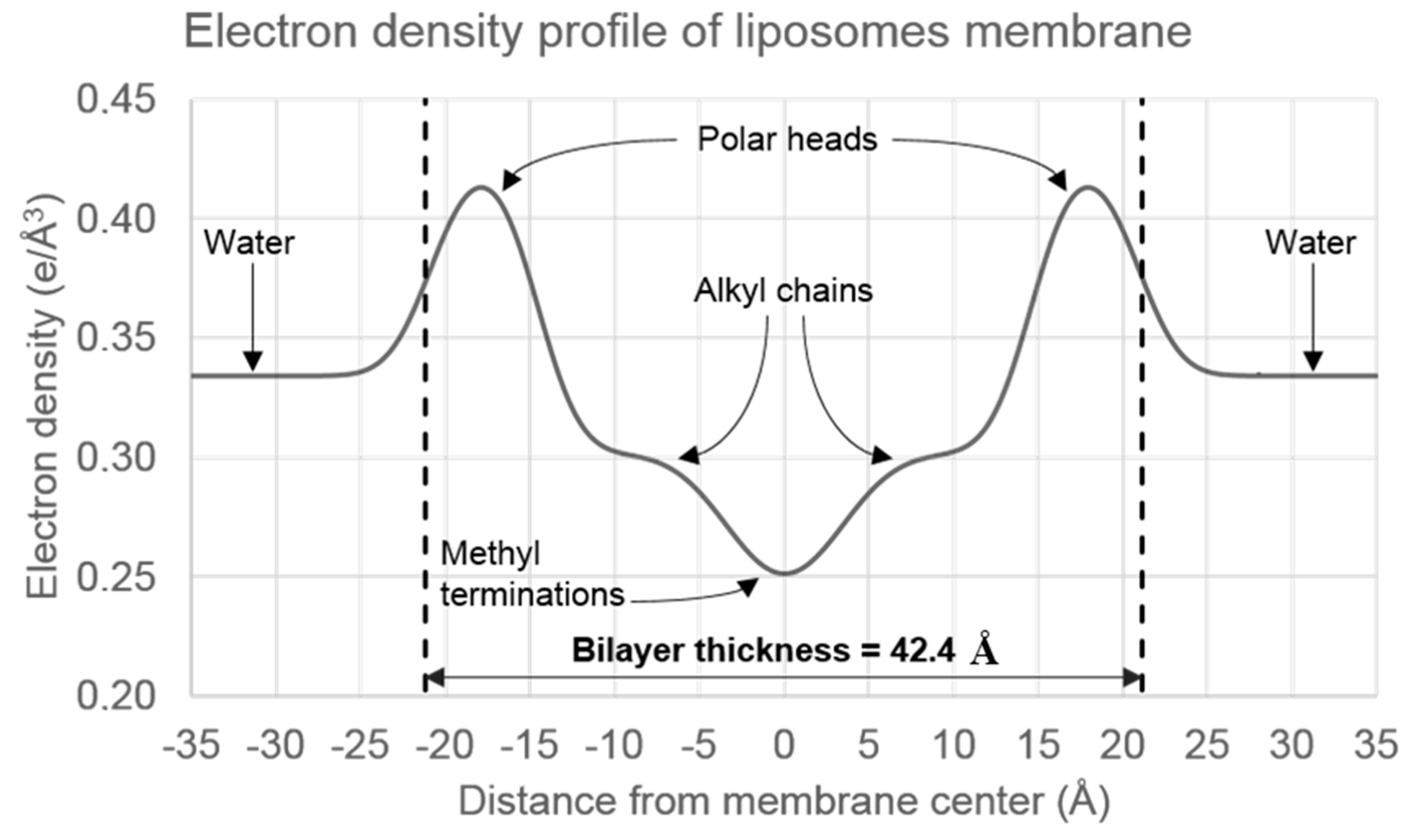
2.4. Nanostructure of Curcumin-Liposome Complexes
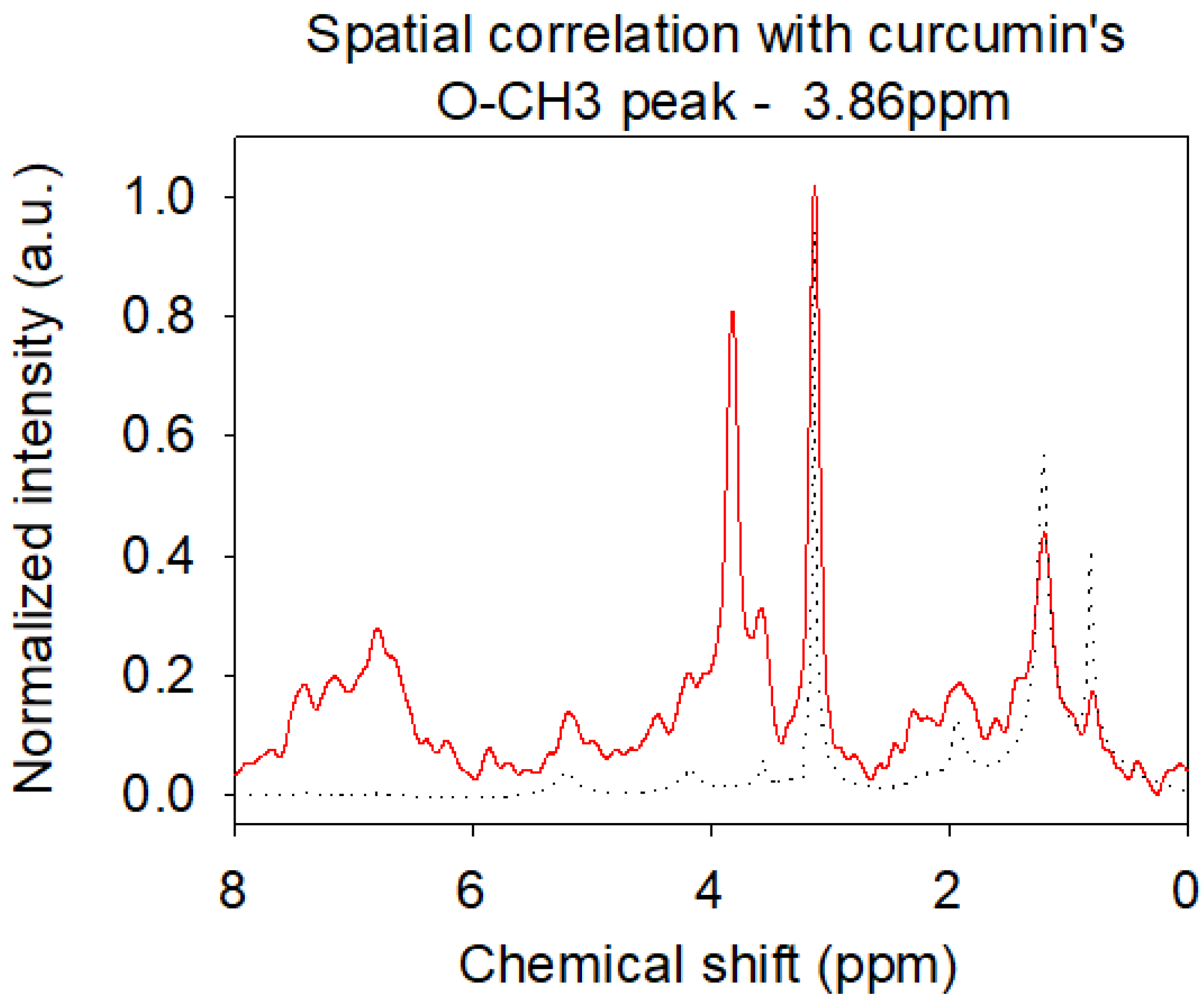
2.5. Localization of Curcumin in the Bilayer by Nuclear Magnetic Resonance (NMR)
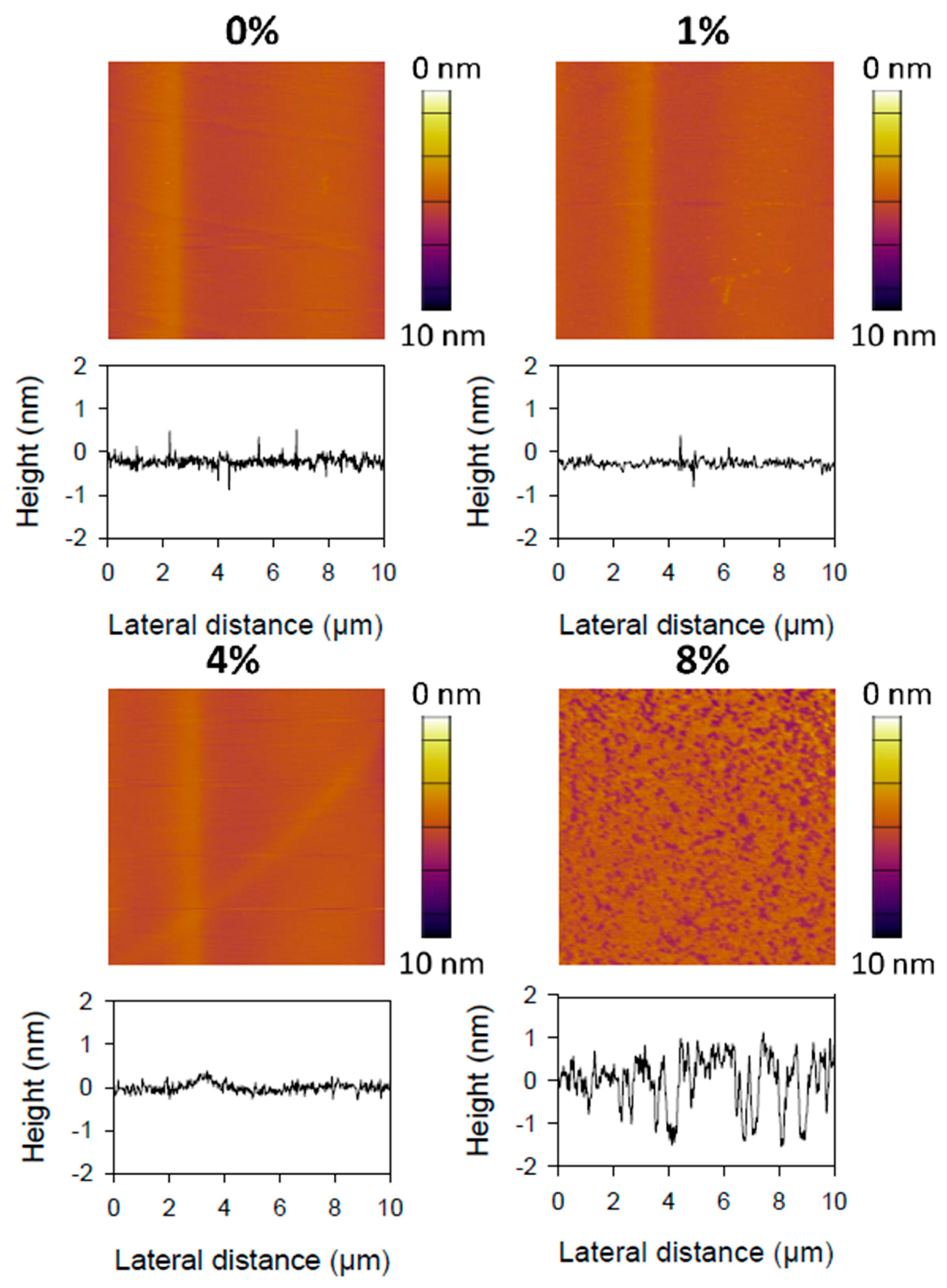
2.6. Topography of Curcumin-Liposome Membranes
2.7. Insertion Mechanism and Structural Changes
3. Materials and Methods
3.1. Reagents
3.2. Liposome Preparation
3.3. Size Measurements
3.4. HPLC Analyses
3.5. UV-Visible Absorbance Measurements
3.6. 1H NMR
3.7. SAXS Experiments
3.8. AFM Imaging and Force Measurements
4. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Soleimani, V.; Sahebkar, A.; Hosseinzadeh, H. Turmeric (Curcuma longa) and its major constituent (curcumin) as nontoxic and safe substances: Review. Phytother. Res. 2018, 32, 985–995. [Google Scholar] [CrossRef] [PubMed]
- Biswas, T.K.; Mukherjee, B. Plant medicines of Indian origin for wound healing activity: A review. Int. J. Low. Extrem. Wounds 2003, 2, 25–39. [Google Scholar] [CrossRef] [PubMed]
- Willenbacher, E.; Khan, S.Z.; Mujica, S.C.A.; Trapani, D.; Hussain, S.; Wolf, D.; Willenbacher, W.; Spizzo, G.; Seeber, A. Curcumin: New Insights into an Ancient Ingredient against Cancer. IJMS 2019, 20, 1808. [Google Scholar] [CrossRef] [PubMed]
- Aggarwal, B.B.; Ichikawa, H.; Garodia, P.; Weerasinghe, P.; Sethi, G.; Bhatt, I.D.; Pandey, M.K.; Shishodia, S.; Nair, M.G. From traditional Ayurvedic medicine to modern medicine: Identification of therapeutic targets for suppression of inflammation and cancer. Expert Opin. Ther. Targets 2006, 10, 87–118. [Google Scholar] [CrossRef]
- Chen, Y.; Lu, Y.; Lee, R.J.; Xiang, G. Nano Encapsulated Curcumin: And Its Potential for Biomedical Applications. IJN 2020, 15, 3099–3120. [Google Scholar] [CrossRef]
- Iurciuc-Tincu, C.-E.; Atanase, L.I.; Ochiuz, L.; Jérôme, C.; Sol, V.; Martin, P.; Popa, M. Curcumin-loaded polysaccharides-based complex particles obtained by polyelectrolyte complexation and ionic gelation. I-Particles obtaining and characterization. Int. J. Biol. Macromol. 2020, 147, 629–642. [Google Scholar] [CrossRef]
- Kuang, G.; Zhang, Q.; He, S.; Liu, Y. Curcumin-loaded PEGylated mesoporous silica nanoparticles for effective photodynamic therapy. RSC Adv. 2020, 10, 24624–24630. [Google Scholar] [CrossRef]
- Rai, M.; Ingle, A.P.; Pandit, R.; Paralikar, P.; Anasane, N.; Santos, C.A.D. Curcumin and curcumin-loaded nanoparticles: Antipathogenic and antiparasitic activities. Expert Rev. Anti. Infect Ther. 2020, 18, 367–379. [Google Scholar] [CrossRef]
- Hasan, M.; Latifi, S.; Kahn, C.; Tamayol, A.; Habibey, R.; Passeri, E.; Linder, M.; Arab-Tehrany, E. The Positive Role of Curcumin-Loaded Salmon Nanoliposomes on the Culture of Primary Cortical Neurons. Mar. Drugs 2018, 16, 218. [Google Scholar] [CrossRef]
- Hasan, M.; Belhaj, N.; Benachour, H.; Barberi-Heyob, M.; Kahn, C.J.F.; Jabbari, E.; Linder, M.; Arab-Tehrany, E. Liposome encapsulation of curcumin: Physico-chemical characterizations and effects on MCF7 cancer cell proliferation. Int. J. Pharm. 2014, 461, 519–528. [Google Scholar] [CrossRef]
- Campani, V.; Scotti, L.; Silvestri, T.; Biondi, M.; De Rosa, G. Skin permeation and thermodynamic features of curcumin-loaded liposomes. J. Mater. Sci. Mater. Med. 2020, 31, 18. [Google Scholar] [CrossRef] [PubMed]
- Luisa, D.P.-A.M.; Griselda, R.-M.; Valentín, M.-L.; Carmina, O.-S.; Cristina, V.-M.; Jj, M.; Maykel, G.-T.; David, Q.-G.; Roberto, S.-S.; Gerardo, L.-G. Curcumin-loaded poly-ε-caprolactone nanoparticles show antioxidant and cytoprotective effects in the presence of reactive oxygen species. J. Bioact. Compat. Polym. 2020, 35, 270–285. [Google Scholar] [CrossRef]
- Iurciuc-Tincu, C.-E.; Cretan, M.S.; Purcar, V.; Popa, M.; Daraba, O.M.; Atanase, L.I.; Ochiuz, L. Drug Delivery System Based on pH-Sensitive Biocompatible Poly(2-vinyl pyridine)-b-poly(ethylene oxide) Nanomicelles Loaded with Curcumin and 5-Fluorouracil. Polymers 2020, 12, 1450. [Google Scholar] [CrossRef] [PubMed]
- Sahab-Negah, S.; Ariakia, F.; Jalili-Nik, M.; Afshari, A.R.; Salehi, S.; Samini, F.; Rajabzadeh, G.; Gorji, A. Curcumin Loaded in Niosomal Nanoparticles Improved the Anti-tumor Effects of Free Curcumin on Glioblastoma Stem-like Cells: An In Vitro Study. Mol. Neurobiol. 2020, 57, 3391–3411. [Google Scholar] [CrossRef]
- Dostert, G.; Kahn, C.J.F.; Menu, P.; Mesure, B.; Cleymand, F.; Linder, M.; Velot, é.; Arab-Tehrany, E. Nanoliposomes of Marine Lecithin, a New Way to Deliver TGF- β 1. J. Biomater. Tissue Eng. 2017, 7, 1163–1170. [Google Scholar] [CrossRef]
- Belhaj, N.; Arab-Tehrany, E.; Loing, E.; Bézivin, C. Skin delivery of hydrophilic molecules from liposomes and polysaccharide-coated liposomes. Int. J. Cosmet. Sci. 2017, 39, 435–441. [Google Scholar] [CrossRef]
- Yaroslavov, A.A.; Sybachin, A.V.; Zaborova, O.V.; Migulin, V.A.; Samoshin, V.V.; Ballauff, M.; Kesselman, E.; Schmidt, J.; Talmon, Y.; Menger, F.M. Capacious and programmable multi-liposomal carriers. Nanoscale 2015, 7, 1635–1641. [Google Scholar] [CrossRef]
- Teo, B.M.; Hosta-Rigau, L.; Lynge, M.E.; Städler, B. Liposome-containing polymer films and colloidal assemblies towards biomedical applications. Nanoscale 2014, 6, 6426. [Google Scholar] [CrossRef]
- Elkhoury, K.; Russell, C.S.; Sanchez-Gonzalez, L.; Mostafavi, A.; Williams, T.J.; Kahn, C.; Peppas, N.A.; Arab-Tehrany, E.; Tamayol, A. Soft-Nanoparticle Functionalization of Natural Hydrogels for Tissue Engineering Applications. Adv. Healthc. Mater. 2019, 1900506. [Google Scholar] [CrossRef]
- Karthikeyan, A.; Senthil, N.; Min, T. Nanocurcumin: A Promising Candidate for Therapeutic Applications. Front. Pharmacol. 2020, 11, 487. [Google Scholar] [CrossRef]
- Jacquot, A.; Francius, G.; Razafitianamaharavo, A.; Dehghani, F.; Tamayol, A.; Linder, M.; Arab-Tehrany, E. Morphological and Physical Analysis of Natural Phospholipids-Based Biomembranes. PLoS ONE 2014, 9, e107435. [Google Scholar] [CrossRef] [PubMed]
- Bianchi, A.; Velot, É.; Kempf, H.; Elkhoury, K.; Sanchez-Gonzalez, L.; Linder, M.; Kahn, C.; Arab-Tehrany, E. Nanoliposomes from Agro-Resources as Promising Delivery Systems for Chondrocytes. IJMS 2020, 21, 3436. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Elkhoury, K.; Barbieux, C.; Linder, M.; Grandemange, S.; Tamayol, A.; Francius, G.; Arab-Tehrany, E. Effects of Bioactive Marine-Derived Liposomes on Two Human Breast Cancer Cell Lines. Mar. Drugs 2020, 18, 211. [Google Scholar] [CrossRef] [PubMed]
- Kadri, R.; Bacharouch, J.; Elkhoury, K.; Ben Messaoud, G.; Kahn, C.; Desobry, S.; Linder, M.; Tamayol, A.; Francius, G.; Mano, J.F.; et al. Role of active nanoliposomes in the surface and bulk mechanical properties of hybrid hydrogels. Mater. Today Bio. 2020, 6, 100046. [Google Scholar] [CrossRef] [PubMed]
- Mu, Q.; Kievit, F.M.; Kant, R.J.; Lin, G.; Jeon, M.; Zhang, M. Anti-HER2/neu peptide-conjugated iron oxide nanoparticles for targeted delivery of paclitaxel to breast cancer cells. Nanoscale 2015, 7, 18010–18014. [Google Scholar] [CrossRef]
- Cadinoiu, A.N.; Rata, D.M.; Atanase, L.I.; Daraba, O.M.; Gherghel, D.; Vochita, G.; Popa, M. Aptamer-Functionalized Liposomes as a Potential Treatment for Basal Cell Carcinoma. Polymers 2019, 11, 1515. [Google Scholar] [CrossRef] [PubMed]
- Elkhoury, K.; Koçak, P.; Kang, A.; Arab-Tehrany, E.; Ellis Ward, J.; Shin, S.R. Engineering Smart Targeting Nanovesicles and Their Combination with Hydrogels for Controlled Drug Delivery. Pharmaceutics 2020, 12, 849. [Google Scholar] [CrossRef]
- Hasan, M.; Elkhoury, K.; Belhaj, N.; Kahn, C.; Tamayol, A.; Barberi-Heyob, M.; Arab-Tehrany, E.; Linder, M. Growth-Inhibitory Effect of Chitosan-Coated Liposomes Encapsulating Curcumin on MCF-7 Breast Cancer Cells. Mar. Drugs 2020, 18, 217. [Google Scholar] [CrossRef]
- Hasan, M.; Elkhoury, K.; Kahn, C.J.F.; Arab-Tehrany, E.; Linder, M. Preparation, Characterization, and Release Kinetics of Chitosan-Coated Nanoliposomes Encapsulating Curcumin in Simulated Environments. Molecules 2019, 24, 2023. [Google Scholar] [CrossRef]
- Sarkar, N.; Bose, S. Liposome-Encapsulated Curcumin-Loaded 3D Printed Scaffold for Bone Tissue Engineering. ACS Appl. Mater. Interfaces 2019, 11, 17184–17192. [Google Scholar] [CrossRef]
- Stevens, M.M.; Honerkamp-Smith, A.R.; Keller, S.L. Solubility Limits of Cholesterol, Lanosterol, Ergosterol, Stigmasterol, and β-Sitosterol in Electroformed Lipid Vesicles. Soft Matter 2010, 6, 5882–5890. [Google Scholar] [CrossRef] [PubMed]
- Kopeć, W.; Telenius, J.; Khandelia, H. Molecular dynamics simulations of the interactions of medicinal plant extracts and drugs with lipid bilayer membranes. FEBS J. 2013, 280, 2785–2805. [Google Scholar] [CrossRef] [PubMed]
- Kollmitzer, B.; Heftberger, P.; Rappolt, M.; Pabst, G. Monolayer spontaneous curvature of raft-forming membrane lipids. Soft Matter 2013, 9, 10877. [Google Scholar] [CrossRef] [PubMed]
- Callan-Jones, A.; Bassereau, P. Curvature-driven membrane lipid and protein distribution. Curr. Opin. Solid State Mater. Sci. 2013, 17, 143–150. [Google Scholar] [CrossRef]
- Koller, D.; Lohner, K. The role of spontaneous lipid curvature in the interaction of interfacially active peptides with membranes. Biochim. Biophys. Acta Biomembr. 2014, 1838, 2250–2259. [Google Scholar] [CrossRef]
- Gavara, N.; Chadwick, R.S. Determination of the elastic moduli of thin samples and adherent cells using conical atomic force microscope tips. Nat. Nanotechnol. 2012, 7, 733–736. [Google Scholar] [CrossRef]
- Sneddon, I.N. The relation between load and penetration in the axisymmetric boussinesq problem for a punch of arbitrary profile. Int. J. Eng. Sci. 1965, 3, 47–57. [Google Scholar] [CrossRef]
- Polyakov, P.; Soussen, C.; Duan, J.; Duval, J.F.L.; Brie, D.; Francius, G. Automated Force Volume Image Processing for Biological Samples. PLoS ONE 2011, 6, e18887. [Google Scholar] [CrossRef]

| mol% of Curcumin | Rupture Force (nN) | Elastic Modulus (MPa) |
|---|---|---|
| 0 | 6.3 ± 0.5 | 57.3 ± 28.4 |
| 1 | 6.9 ± 0.7 | 45.3 ± 18.2 |
| 4 | 7.6 ± 0.9 | 48.2 ± 14.6 |
| 8 | 7.2 ± 0.8 | 42.1 ± 14.5 |
| 12 | 2.7 ± 0.3 | 41.1 ± 8.4 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Arab-Tehrany, E.; Elkhoury, K.; Francius, G.; Jierry, L.; Mano, J.F.; Kahn, C.; Linder, M. Curcumin Loaded Nanoliposomes Localization by Nanoscale Characterization. Int. J. Mol. Sci. 2020, 21, 7276. https://doi.org/10.3390/ijms21197276
Arab-Tehrany E, Elkhoury K, Francius G, Jierry L, Mano JF, Kahn C, Linder M. Curcumin Loaded Nanoliposomes Localization by Nanoscale Characterization. International Journal of Molecular Sciences. 2020; 21(19):7276. https://doi.org/10.3390/ijms21197276
Chicago/Turabian StyleArab-Tehrany, Elmira, Kamil Elkhoury, Gregory Francius, Loic Jierry, Joao F. Mano, Cyril Kahn, and Michel Linder. 2020. "Curcumin Loaded Nanoliposomes Localization by Nanoscale Characterization" International Journal of Molecular Sciences 21, no. 19: 7276. https://doi.org/10.3390/ijms21197276
APA StyleArab-Tehrany, E., Elkhoury, K., Francius, G., Jierry, L., Mano, J. F., Kahn, C., & Linder, M. (2020). Curcumin Loaded Nanoliposomes Localization by Nanoscale Characterization. International Journal of Molecular Sciences, 21(19), 7276. https://doi.org/10.3390/ijms21197276








